Submitted:
14 January 2026
Posted:
15 January 2026
You are already at the latest version
Abstract
An archaeological exposure near Hachum, featuring a Ditch profile interpreted as part of a Neolithic earthwork, was characterized using DNA analyses of bacterial 16S rRNA from soil samples. The results showed that the middle and lower parts of the Ditch fill could be clearly distinguished from each other and from the surrounding area based on the composition of soil bacterial DNA. Genera detected predominantly in the lower part of the Ditch suggest that, after the Ditch was completed, organic matter, animal dung, and possibly even human feces were accumulated at the bottom. The investigations demonstrate that analyses of soil bacterial communities can provide valuable insights into the history and function of a Neolithic earthwork and, more generally, represent an important additional source of information for interpreting archaeological contexts that are devoid of or poor in finds.
Keywords:
1. Introduction
2. Experimental
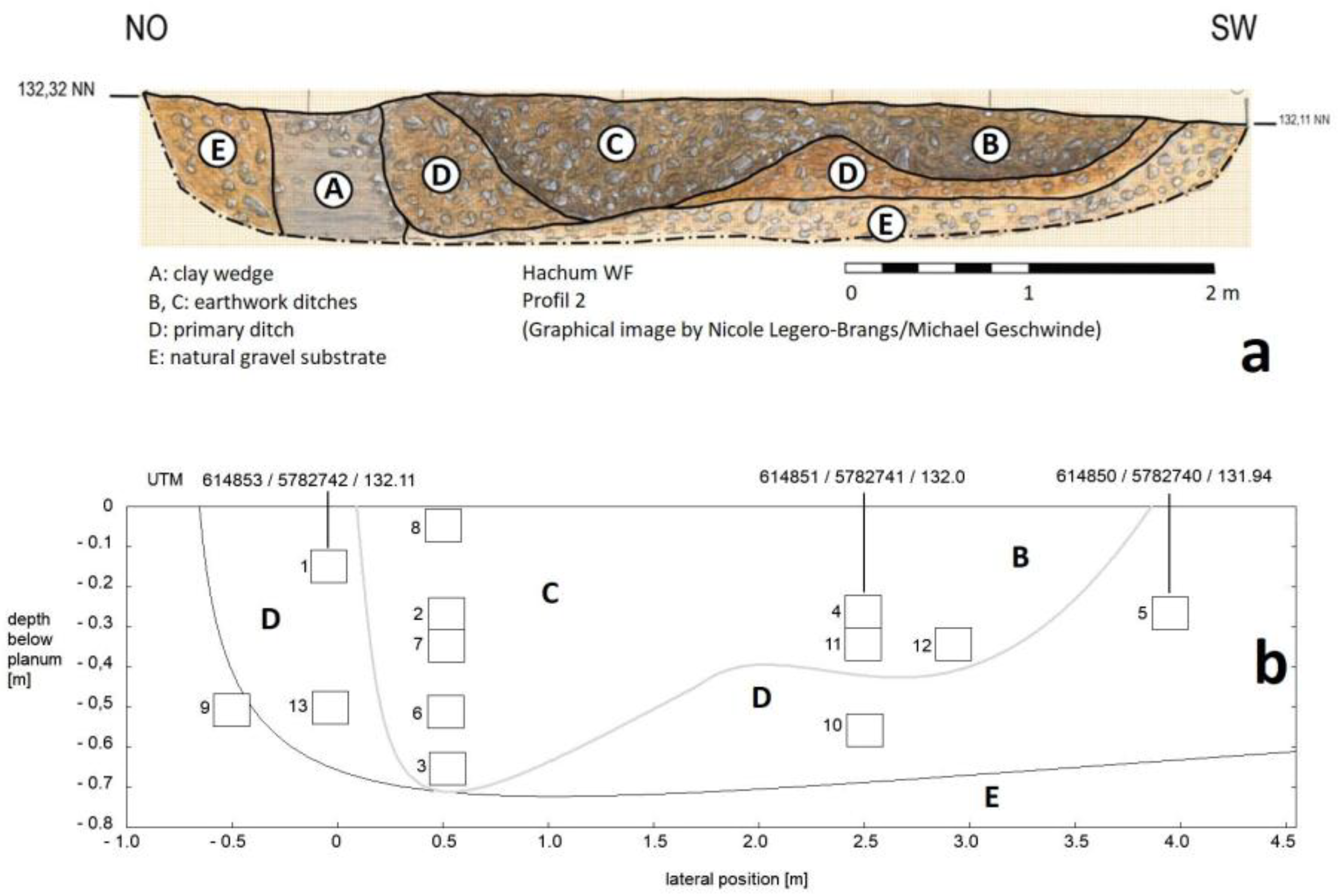
2.1. Archaeological Situation
2.2. Soil Samples and Sequencing
| No. | Lab-intern label | depth (below planum) | Lateral position |
| 1 | HC1 | 15 cm | transition region N |
| 2 | HC2 | 25 cm | central Ditch profile |
| 3 | HC3 | 70 cm | central Ditch profile |
| 4 | HC4 | 30 cm | side Ditch |
| 5 | HC5 | 30 cm | transition region S |
| 6 | HC6 | 50 cm | central Ditch profile |
| 7 | HC7 | 35 cm | central Ditch profile |
| 8 | HC8 | 10 cm | central Ditch profile |
| 9 | HC9 | 55 cm | outside Ditch |
| 10 | HC10 | 60 cm | transition region S |
| 11 | HC11 | 40 cm | side Ditch |
| 12 | HC12 | 40 cm | side Ditch |
| 13 | HC13 | 50 cm | transition region N |
2.3. Data Processing
3. Results and Discussion
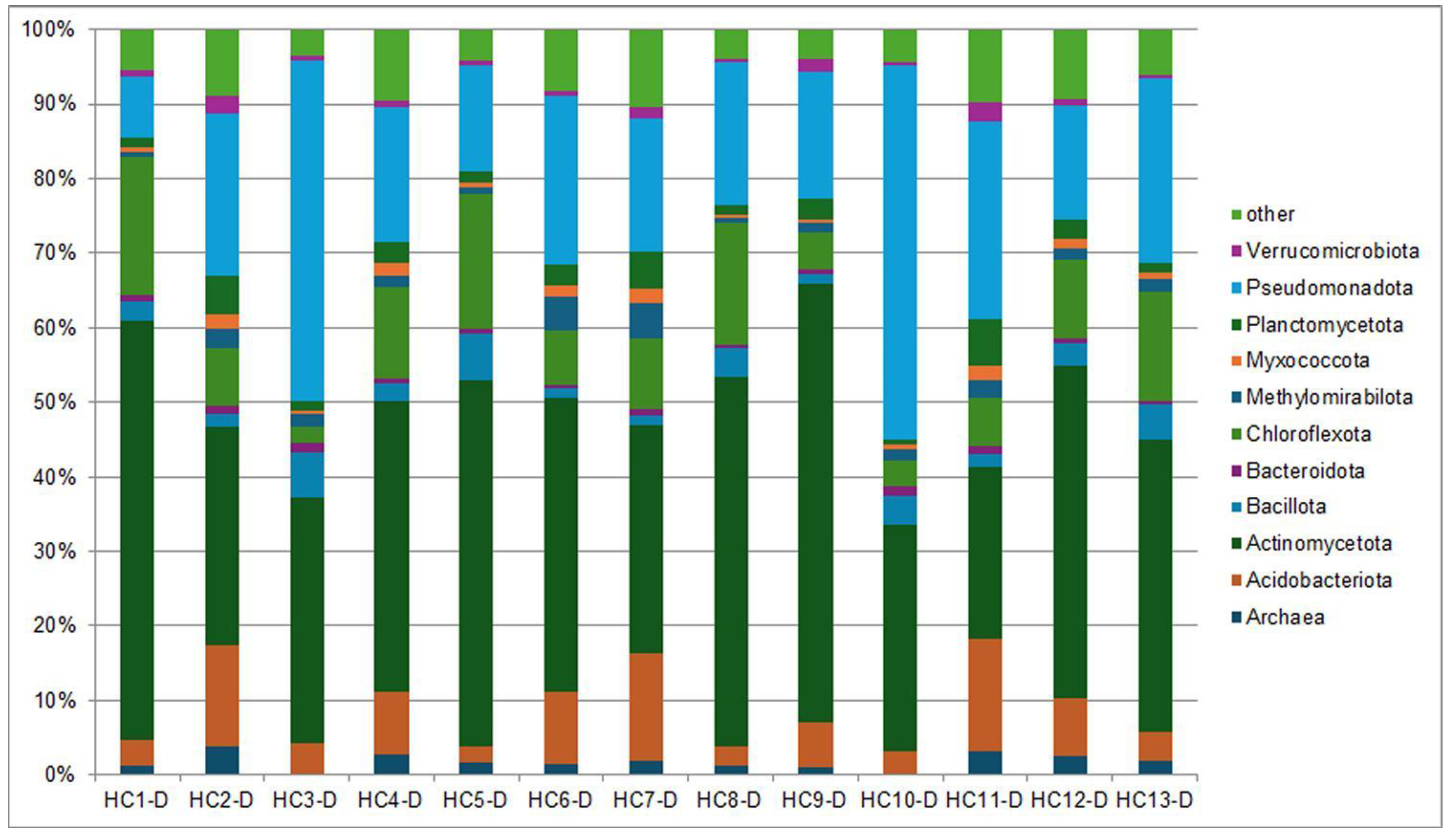
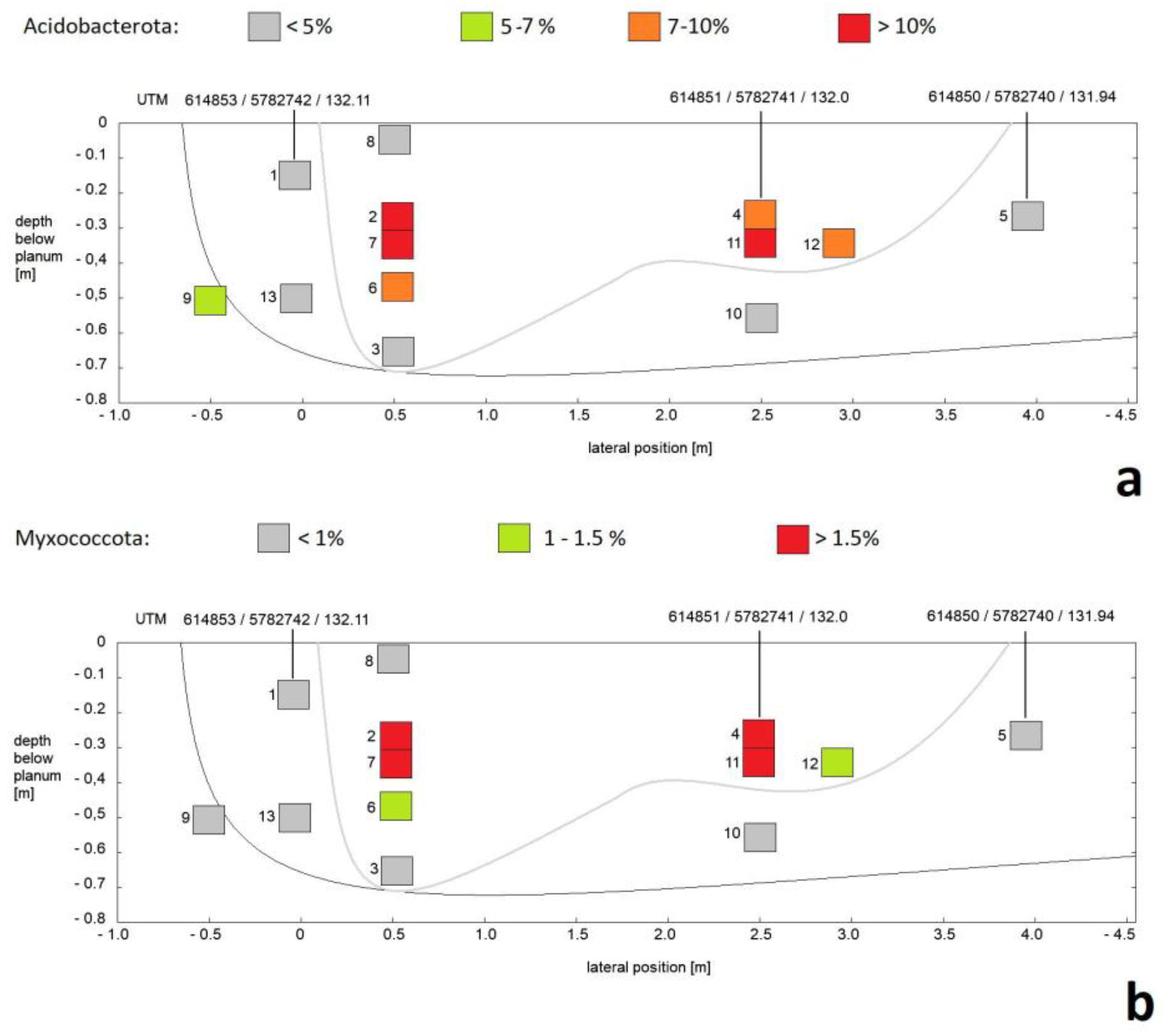
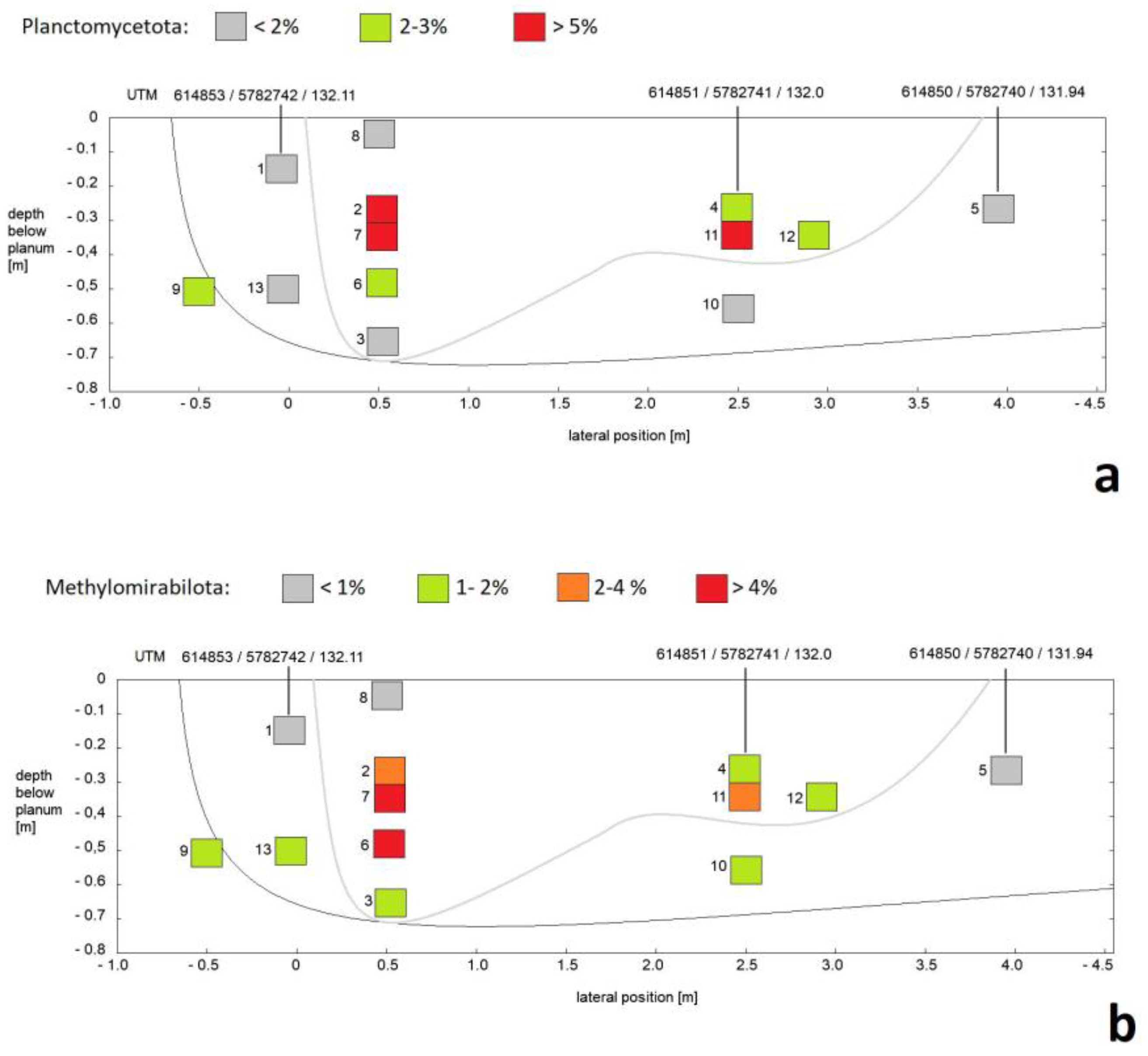
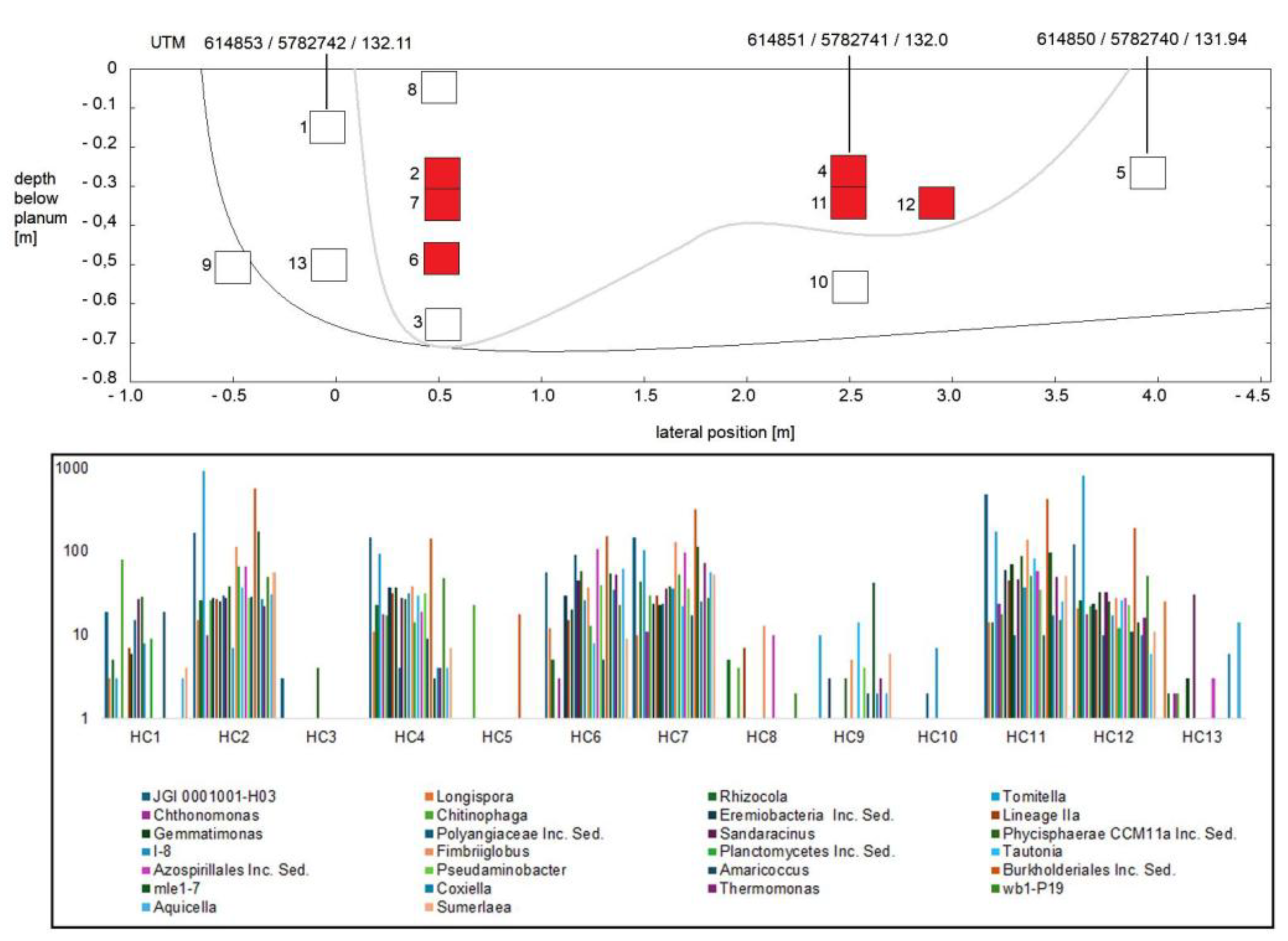
3.1. Composition of Soil Bacterial Communities by Phyla
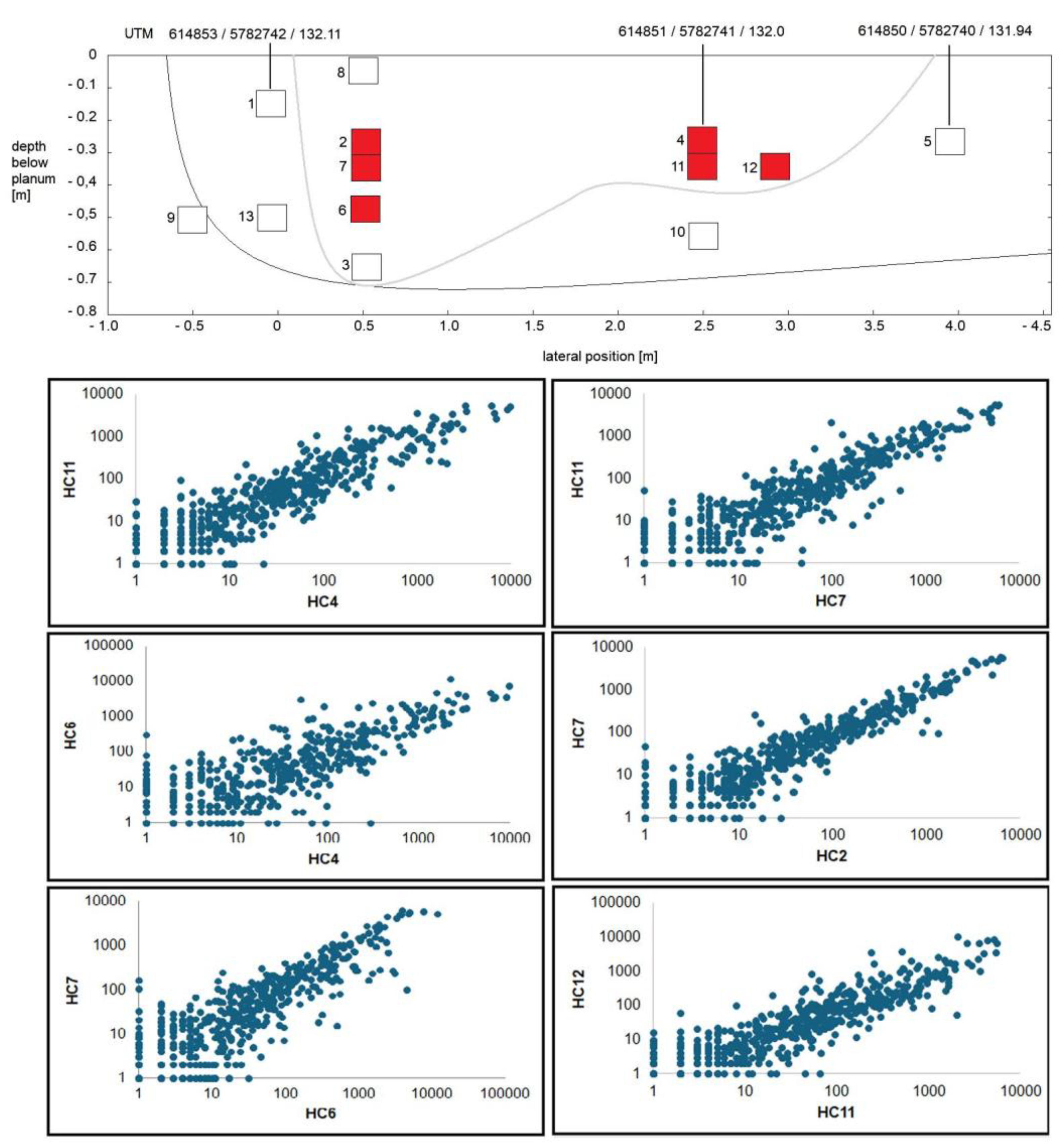
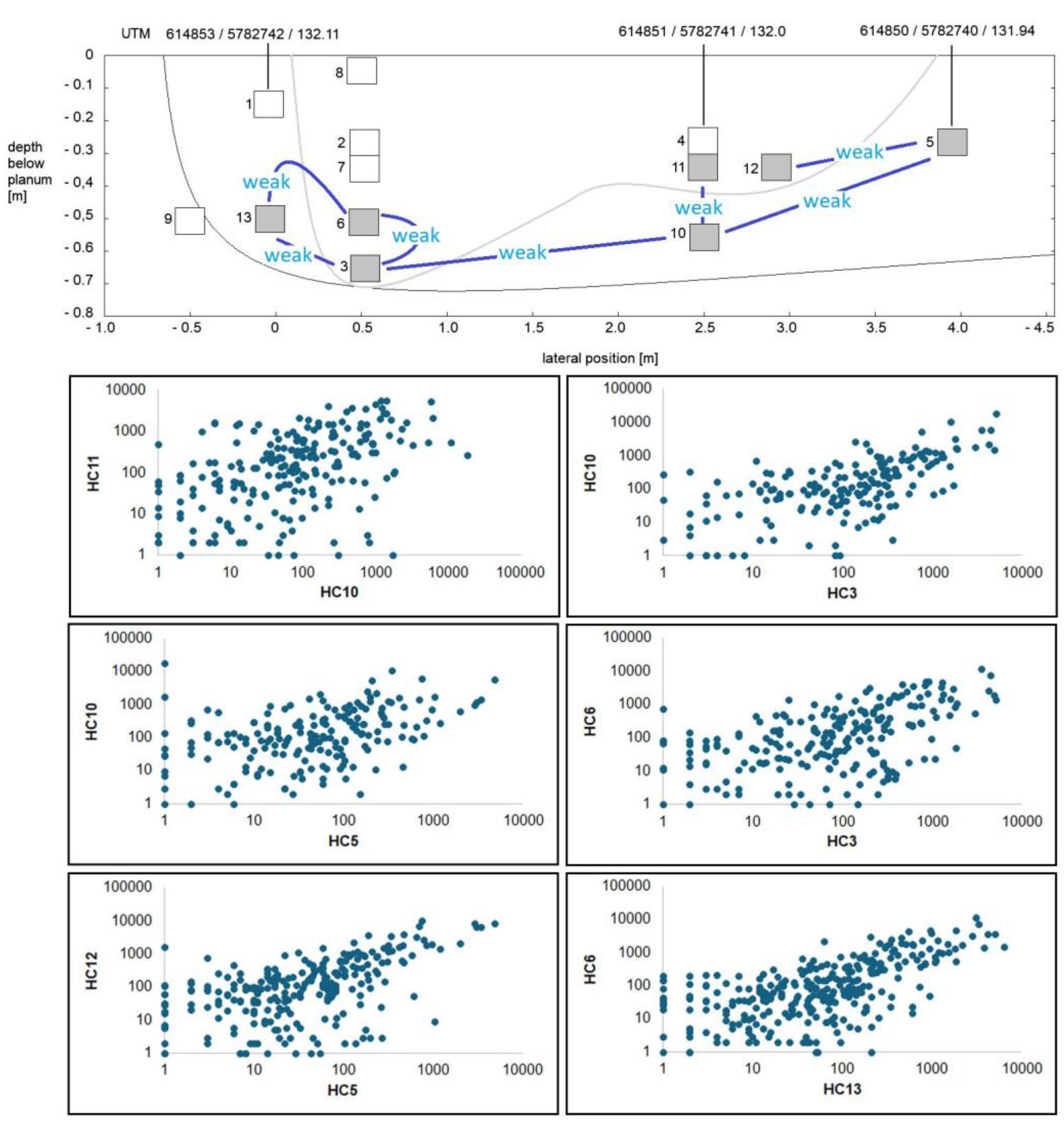
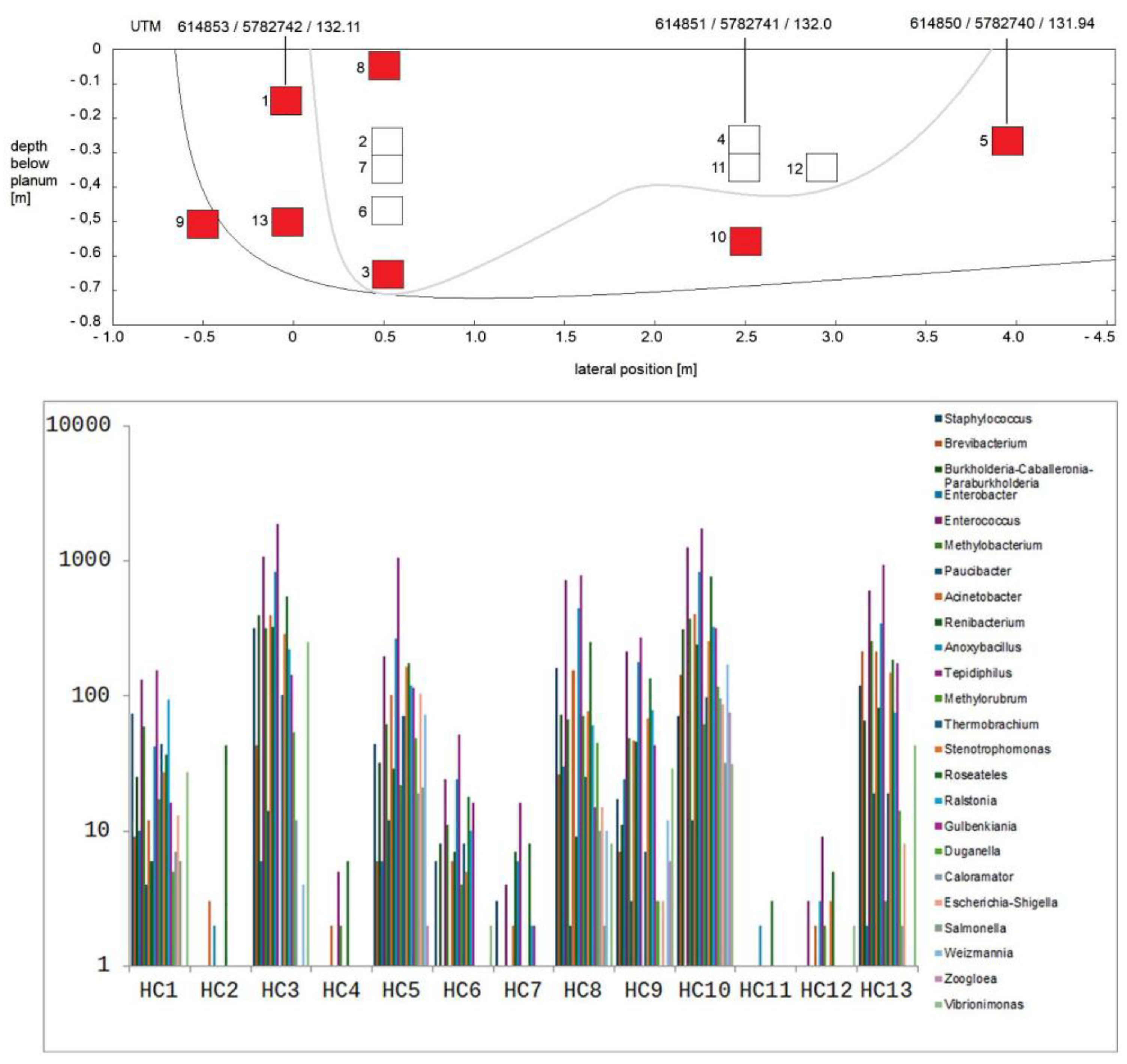
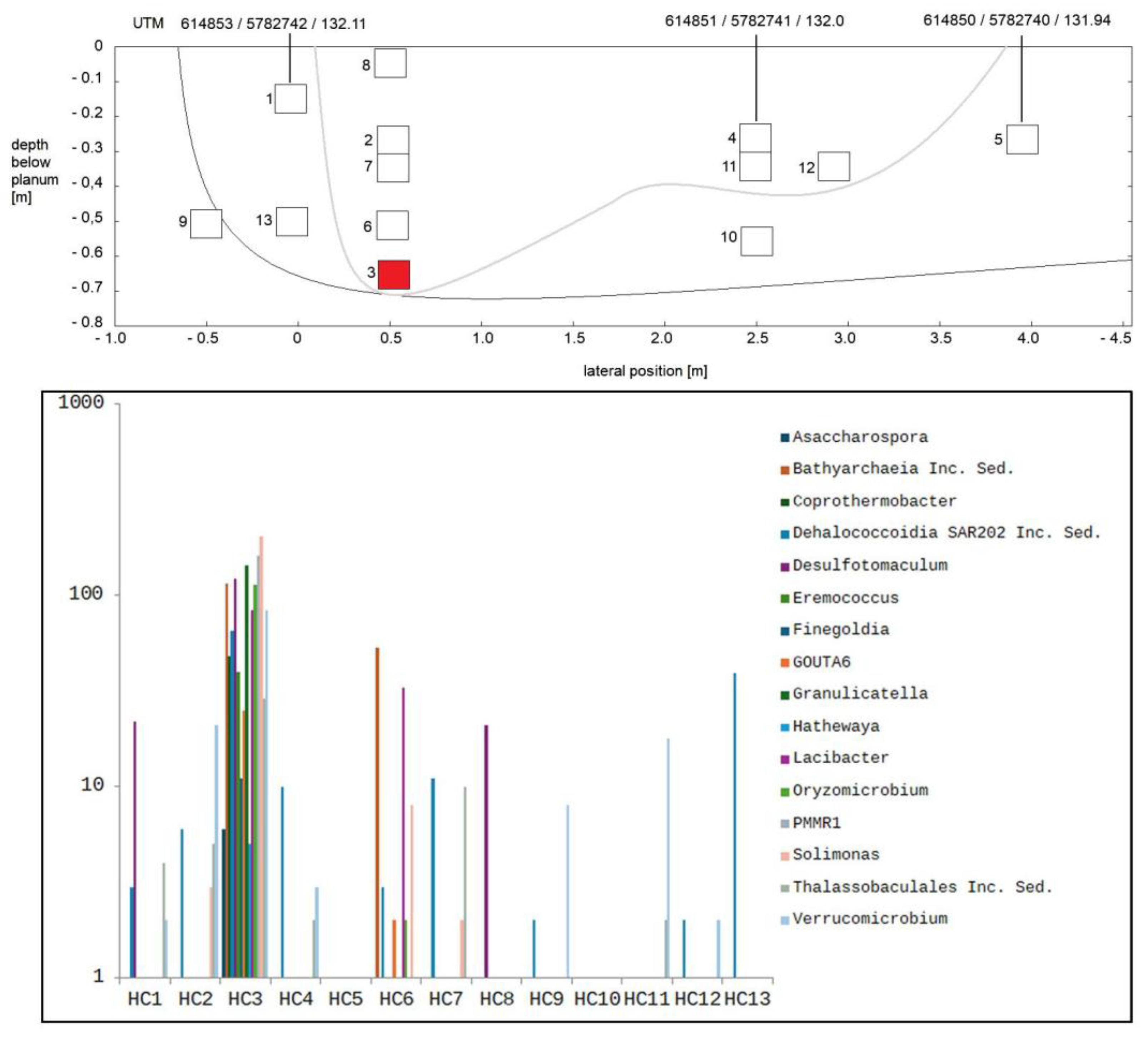
3.2. Composition of Soil Bacterial Communities by Operational Taxonomical Units (OTUs)
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Haak, W.; et al. Ancient DNA from European early Neolithic farmers reveals their Near Eastern affinities. PLoS Biol. 2010, 8, e1000536. [Google Scholar] [CrossRef] [PubMed]
- Lazaridis, I.; et al. Ancient human genomes suggest three ancestral populations for present-day Europeans. Nature 2014, 513, 409–413. [Google Scholar] [CrossRef] [PubMed]
- Haak, W.; et al. Massive migration from the steppe was a source for Indo-European languages in Europe. Nature 2015, 522, 207. [Google Scholar] [CrossRef]
- Warinner, Ch.; Herbig, A.; Mann, A.; Yates, J.A.F.; Weiß, C.L.; Burbano, H.A.; Orlando, L.; Krause, J. A robust framework for microbial archaeology. Ann. Rev. Genom. Hum. Genet. 2017, 18, 321–356. [Google Scholar] [CrossRef]
- Warinner, C. An archaeology of microbes. J. Anthropol. Res. 2022, 10, 721976. [Google Scholar] [CrossRef]
- Oberreich, M.; Steinhoff-Knopp, B.; Burkhard, B.; Kleemann, J. The Research Gap between Soil Biodiversity and Soil-Related Cultural Ecosystem Services. Soil Systems 2024, 8, 97. [Google Scholar] [CrossRef]
- Dergacheva, M. Ecological function of soil humus. Eurasian Soil Sci. 2001, 34, 100–105. [Google Scholar]
- Jones, S.E.; Lennon, J.T. Dormancy contributes to the maintenance of microbial diversity. PNAS 2010, 107, 5881–5886. [Google Scholar] [CrossRef]
- Benito, B.M.; Gil-Romera, G.; Birks, H.J.B. Ecological memory at millennial time-scales: the importance of data constraints, species longevity and niche features. Ecography 2019, 43, 04772. [Google Scholar] [CrossRef]
- Gonze, D.; Coyte, K.; Lahti, L.; Faust, K. Microbial communities as dynamical system. Curr. Opin. Microbiol. 2018, 44, 41–49. [Google Scholar] [CrossRef] [PubMed]
- Khalighi, M.; Gonze, D.; Faust, K.; Sommeria-Klein, G.; Lathi, L. Quantifiying the impact of ecological memory on the dynamics of interacting communities. PLOS Comput. Biol. 2022, 1009396. [Google Scholar] [CrossRef]
- Khalighi, M.; Gonze, D.; Faust, K.; Sommeria-Klein, G.; Lathi, L. Quantifiying the impact of ecological memory on the dynamics of interacting communities. PLOS Comput. Biol. 2022, 1009396. [Google Scholar] [CrossRef]
- Jones, M.; Brown, T. Agricultural origins: the evidence of modern and ancient DNA. Holocene 2000, 10, 769–776. [Google Scholar] [CrossRef]
- Marfennia, O.E.; Ivanova, A.E.; Sacharov, D.S. The mycologicl properties of medieval cultur layers as a form of ‘soil biological memory’ about urbanization. J. Soils Sediments 2008, 8, 340–348. [Google Scholar] [CrossRef]
- Philips, A.; Stolarek, I.; Kuczkowska, B.; et al. Comprehensive analysis of microorganisms accompanying human archaeological remains. GigaScience 2017, 6, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Borisov, A.V.; Demkina, T.S.; Kashirskaya, N.N.; Khomutova, T.E.; Chernysheva, E.V. Changes in the past soil-forming conditions and human activity in soil biological memory: microbial and enzyme components. Eurasian Soil Sci. 2021, 54, 1078–1088. [Google Scholar] [CrossRef]
- Schmid, M.W.; VanMoorsel, S.J.; Hahkl, T.; DeLuca, E.; DeDeyn, G.B.; Wagg, C.; Niklaus, P.A.; Schmid, B. Effects of plant community history, soil legacy and plant diversity on soil microbial communities. J. Ecology 2021, 109, 3007–3023. [Google Scholar] [CrossRef]
- Zhongjun, J. Effects of agricultural land use on the differentiation of nitrifier communities and functional patterns from natural terrestrial ecosystems. Science of The Total Environment 2022, 835, 155568. [Google Scholar] [CrossRef]
- Devendrapandi, G.; Balu, R.; Ayyappan, K.; Ayyamperula, R.; Alhammadi, S.; Lavanya, M.; Senthilkumar, R.; Karthika, P.C. Unearthing Earth’s secrets: Exploring the environmental legacy of contaminants in soil, water, and sediments. Environ. Res. 2024, 249, 118246. [Google Scholar] [CrossRef]
- Köhler, J.M.; Kalensee, F.; Günther, P.M.; Schüler, T.; Cao, J. The local ecological memory of soil: majority and minority components of bacterial communities in prehistoric urns from Schöps (Germany). Int. J. Environ. Res. 2018, 12, 575–684. [Google Scholar] [CrossRef]
- Margesin, R.; Siles, J.A.; Cajthaml, T.; Ohlinger, B.; Kistler, E. Microbiology meets archaeology: soil microbial communities reveal different human activities at archaic Monte Iato (Sixth century BC). Microbial Ecol. 2017, 73, 925–938. [Google Scholar] [CrossRef] [PubMed]
- Köhler, J.M.; Ehrhardt, L.; Günther, P.M.; Cao, J. Soil bacteria in archaeology: What could rank abundance functions tell us about ancient human impact on microbial communities? Microorganisms 2024, 12, 2243. [Google Scholar] [CrossRef]
- Chernysheva, E.; Korobov, D.; Borisov, A. Thermophilic microorganisms in arable land around medieval archaeological sites in Northern Caucasus, Russia: Novel evidence of past manuring practices. Geoarchaeol. Intern. J. 2017, 32, 494–501. [Google Scholar] [CrossRef]
- Wegner, C.E.; Liesack, W. Unexpected dominance of elusive acidobacteria in early industrial soft coal slags. Frontiers Microbiol. 2017, 8, 1023. [Google Scholar] [CrossRef] [PubMed]
- Wolinska, A.; Wlodarczyk, K.; Kuzniar, A.; Marzec-Grzadziel, A.; Grzadziel, J.; Galazka, A.; Uzarowicz, L. Soil microbial community profiling and bacterial metabolic activity of technosols as an effect of soil properties following land reclamation: a case study from abandoned iron sulphide and uranium mine in Rudki (south-central Poland). Agronomy 2020, 10, 1795. [Google Scholar] [CrossRef]
- Sabin, S.; Yeh, H.Y.; Pluskowski, A.; Clamer, C.; Mitchell, P.D.; Bos, K.I. Estimating molecular preservation of the intestinal microbiome via metagenomics analyses of latrine sediments from two medieval cities. Phil. Trans. Royal Soc. B 2020, 375, 20190576. [Google Scholar] [CrossRef]
- Orr, C. Archaeological soil from Roman occupational layers can be differentiated by microbial and chemical signatures. Ecosystems and Biodiversity 2013, 3, 1129040. [Google Scholar]
- Köhler, J.M.; Ehrhardt, L.; Günther, P.M.; Cao, J.; Konecny, A. Bacterial Communities from the Antique Roman City of Carnuntum (Austria):16S r-RNA-Based Comparison of Soil Samples from Different Archaeological Horizons. Preprints 2024. [Google Scholar] [CrossRef]
- Köhler, J.M.; Ehrhardt, L.; Cao, J.; Möller, F.; Schüler, T.; Günther, P.M. Beta-Diversity Enhancement by archaeological structures: bacterial communities of an historical tannery area of the city of Jena (Germany) reflected the ancient human impact. Ecologies 2023, 4, 325–343. [Google Scholar] [CrossRef]
- Cao, J; Chande, C; Kalensee, F; Schüler, T; Köhler, JM. Microfluidically supported characterization of responses of Rhodococcus erythropolis strains isolated from different soils on Cu, Ni-, and Co-stress. Braz. J. Microbiol. 2021, 52, 1405–1415. [Google Scholar] [CrossRef] [PubMed]
- Geschwinde, M.; Raetzel-Fabian, D. EWBSL. Eine Fallstudie zu den jungneolithischen Erdwerken am Nordrand der Mittelgebirge. Beiträge zur Archäologie in Niedersachsen 2009, 14. [Google Scholar]
- Knoche, B. Riten, Routen, Rinder – das jungneolithische Erdwerk von Soest im Wegenetz eines extensiven Viehwirtschaftssystems. In Neue Forschungen zum Neolithikum in Soest und am Hellweg; Soester Beiträge zur Archäologie; Melzer, Walter, Ed.; 2013; Volume 13, pp. 119–274. [Google Scholar]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glockner, F.O. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, D590–D596. [Google Scholar] [CrossRef]
- Klindworth, A.; Pruesse, E.; Schwee, T.; Peplies, J.; Quast, C.; Horn, M.; Glockner, F.O. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013, 41, e1. [Google Scholar] [CrossRef]
- Yilmaz, P.; Parfrey, L.-W.; Yarza, P.; Gerken, J.; Pruesse, E.; Quast, C.; Schweer, T.; Peplies, J.; Ludwig, W.; Glockner, F.O. The SILVA and “All-species Living Tree Project (LTP)” taxonomic frameworks. Nucleic Acid Res. 2014, 42, D643–D648. [Google Scholar] [CrossRef] [PubMed]
- Ettwig, K.; et al. Nitrite-driven anaerobic methane oxdation by oxygenic bacteria. Nature 2010, 464, 543–548. [Google Scholar] [CrossRef]
- Maszenan, A.M.; Seviour, R.J.; Patel, B.K.; Rees, G.N.; McDougall, B.M. Amaricoccus gen. nov., a gram-negative coccus occurring in regular packages or tetrads, isolated from activated sludge biomass, and descriptions of Amaricoccus veronensis sp. nov., Amaricoccus tamworthensis sp. nov., Amaricoccus macauensis sp. nov., and Amaricoccus kaplicensis sp. Nov. Int. J. Syst. Bacteriol. 1997, 47, 727–734. [Google Scholar]
- Mohr, K.I.; Garcia, R.O.; Gerth, K.; Irschi, H.; Müller, R. Sandaracinus amylolyticus gen. nov., sp. nov., a starch-degrading soil myxobacterium, and description of Sandaracinaceae fam. Nov. Int. J. Syst. Bacteriol. 2012, 62, 1192–1198. [Google Scholar] [CrossRef]
- Kadnikov, V.V.; Mardanov, A.V.; Beletsky, A.V.; Rakitin, A.L.; Karnachuk, O.V.; Ravin, N.V. Phylogeny and physiology of candidate phylum BRC1 inferred from the first complete metagenome-assembled genome obtained from deep subsurface aquifer. Syst. Appl. Microbiol. 2019, 42, 67–76. [Google Scholar] [CrossRef] [PubMed]
- Dixon, B. Cheese, toes and mosquitoes. British Med. J. 1996, 312, 72609132. [Google Scholar] [CrossRef]
- Vaz-Moreira, I.; Nobre, M.F.; Nunes, O.C.; Manai, C.M. Gulbenkiania mobilis gen. nov., sp. nov., isolated from treated municipal wastewater. Int. J. Syst. Evol. Microbiol. 2007, 57, 1108–1112. [Google Scholar] [CrossRef]
- Collins, M.D.; Jovita, M.R.; Lawson, P.A.; Foster, G. Characterization of a novel gram-positive, catalase-negative coccus from horses: description of Eremococcus coleocola gen. nov., sp. Nov. Int. J. Syst. Bacteriol. 1999, 49, 1381–1385. [Google Scholar] [CrossRef]
- Etchebehere, C.; Pavan, M.E.; Zorzópulos, J.; Soubes, M.; Muxi, L. Coprothermobacter platensis sp. nov., a new anaerobic proteolytic thermophilic bacterium isolated from an anaerobic mesophilic sludge. Int. J. Syst. Evol. Microbiol. 1998, 49, 1297–1304. [Google Scholar] [CrossRef]
- Liu, C.-T.; et al. Oryzomicrobium terrae gen. nov., sp. nov., of the family Rhodocyclaceae isolated from paddy soil. Int. J. Syst. Evol. Microbiol. 2017, 67, 001565. [Google Scholar] [CrossRef]
- Watanabe, M.; Kojima, H.; Fukui, M. Review of Desulfotomaculum species and proposal of the genera Desulfallas gen. nov., Desulfofundulus gen. nov., Desulfofarcimen gen. nov. and Desulfohalotomaculum gen. nov. Int. J. Syst. Evlo. Microbiol. 2018, 68, 2891–2899. [Google Scholar] [CrossRef] [PubMed]
- Oren, A.; Garrity, G.M. Valid publication of the names of forty-two phyla of prokaryotes. Int. J. Syst. Evol. Microbiol. 2021, 10, 5056. [Google Scholar] [CrossRef] [PubMed]
- Gerritsen, J.; Fuentes, S.; Grievink, W.; VanNiftrik, L.; Tindall, J.B.; Timmermann, H.M.; Rijkers, G.T.; Smidt, H. Characterization of Romboutsia ilealis gen. nov., sp. nov., isolated from the gastro-intestinal tract of a rat, and proposal for the reclassification of five closely related members of the genus Clostridium into the genera Romboutsia gen. nov., Intestinibacter gen. nov., Terrisporobacter gen. nov. and Asaccharospora gen. nov. Int. J. Syst. Evlo. Microbiol. 2014, 64, 1600–1616. [Google Scholar]
- Gupta, S.; Garg, M.; Misra, S.; Singhal, S. Granulicatella adiacens abscess: Two rare cases and review. J. Lab. Physicians 2018, 10, 121–123. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



