Submitted:
14 January 2026
Posted:
15 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Clinical Samples
2.2. Extraction and Amplification
2.3. Obtaining Consensus Sequences of the HIV-1 pol and env genes
2.4. Analysis of Consensus Sequences of the HIV-1 Pol Gene
2.5. Analysis of Consensus Sequences of the HIV-1 Env Gene and CXCR4 Cell Tropism Prediction
2.6. Statistical Analysis
3. Results
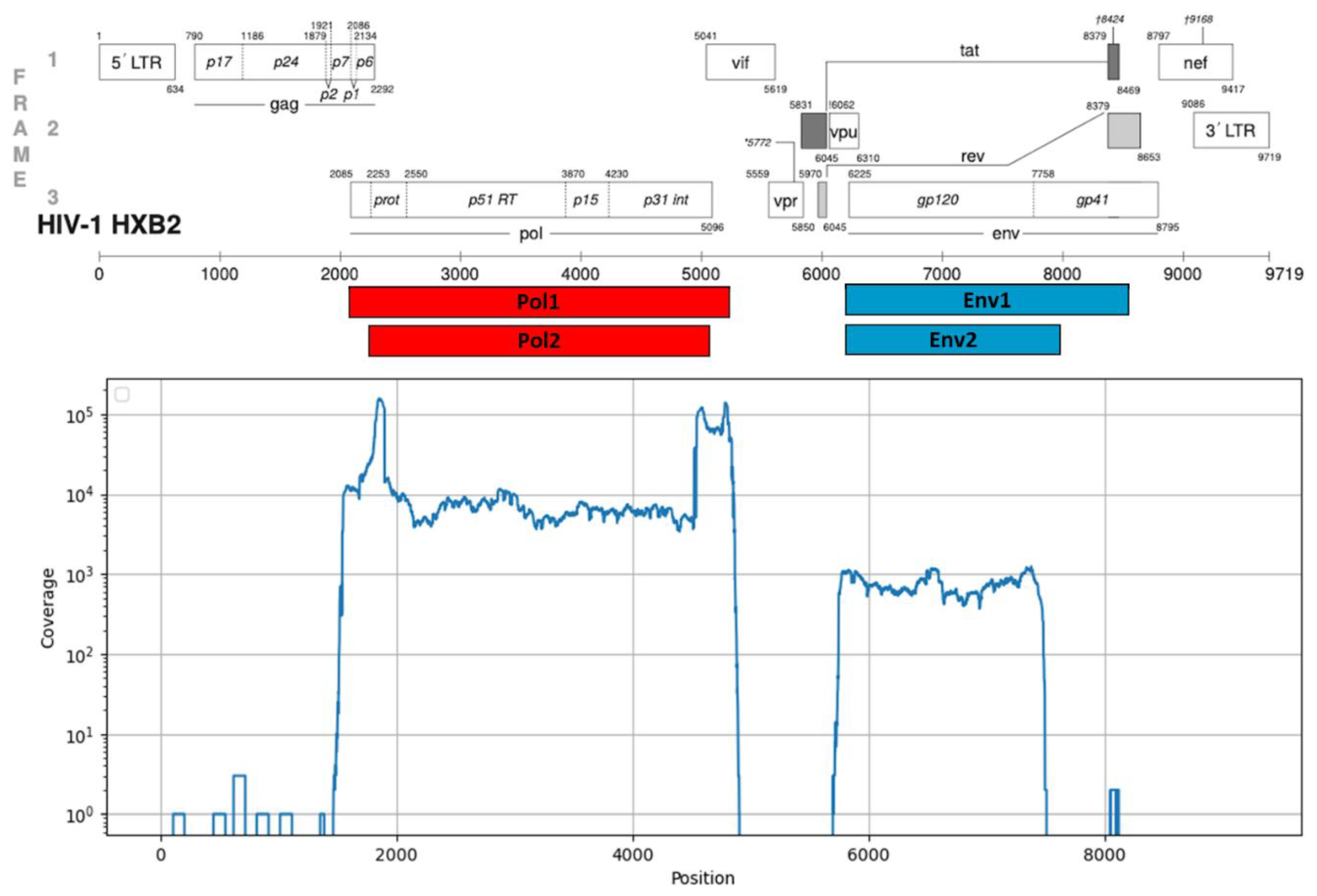
3.1. Testing of the NGS Protocol for HIV-1 Pol and Env Genes
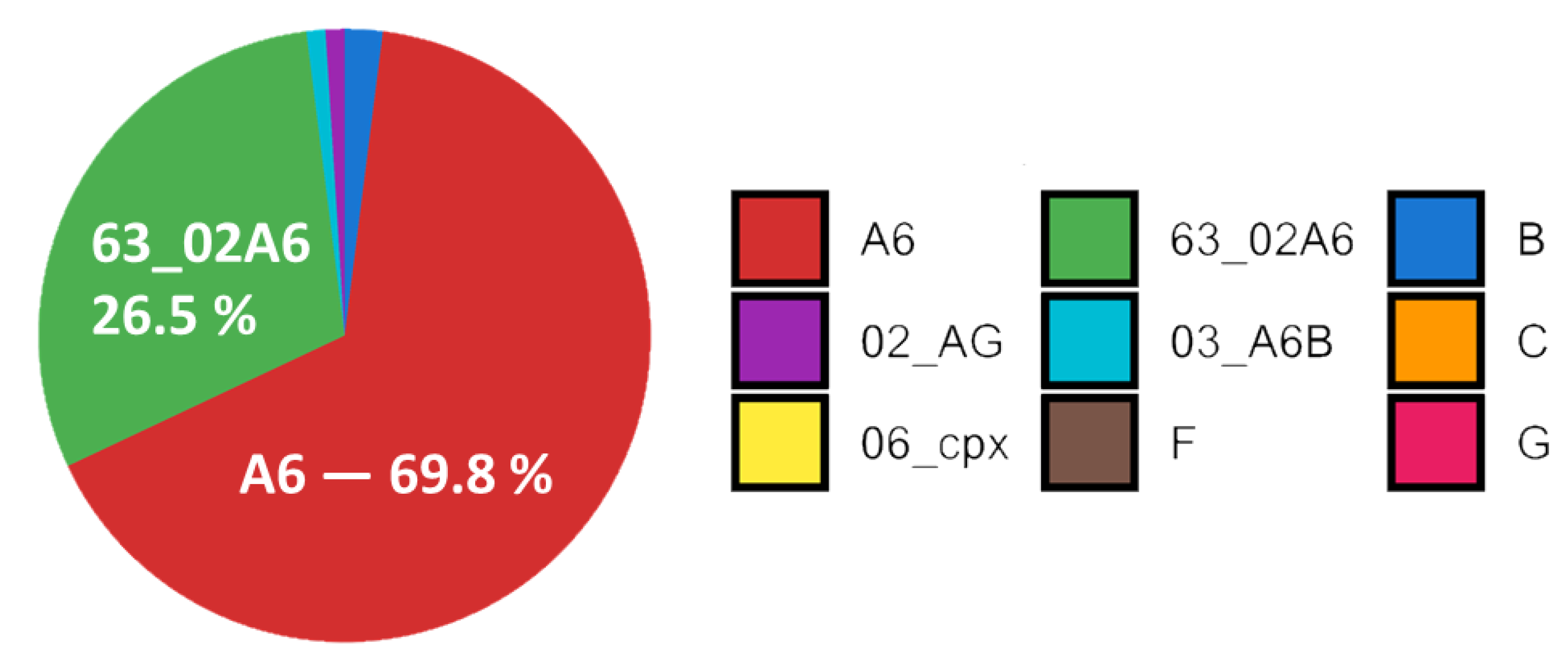
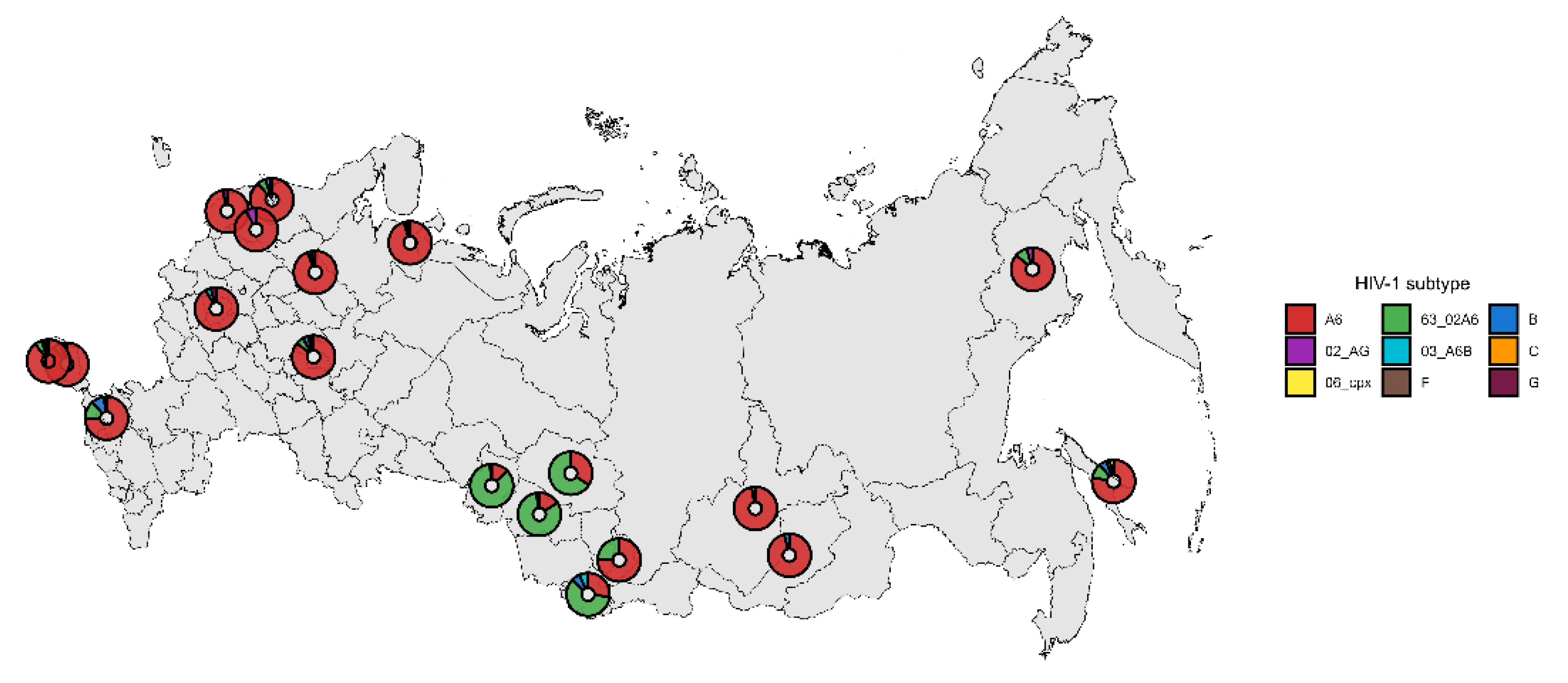
3.2. Subtyping of HIV-1 Viruses
3.3. Analysis of Drug Resistance Mutations
| Mutation | Total (n = 1888) Abs. / % |
ART Experience (n = 1466) | ||
|---|---|---|---|---|
| No (n = 411) Abs. / % |
Yes (n = 1055) Abs. / % |
p | ||
| A62V | 506 / 26,80 | 64 / 15,57 | 334 / 31,66 | < 0,001 |
| M184V1* | 234 / 12,39 | 6 / 1,46 | 210 / 19,91 | < 0,001 |
| K103N1 | 206 / 10,91 | 28 / 6,81 | 149 / 14,12 | 0,013 |
| S68G | 176 / 9,32 | 32 / 7,79 | 114 / 10,81 | 1,000 |
| V90I | 162 / 8,58 | 20 / 4,87 | 115 / 10,90 | 0,037 |
| E138A | 158 / 8,37 | 31 / 7,54 | 91 / 8,63 | 1,000 |
| G190S1 | 135 / 7,15 | 7 / 1,70 | 116 / 11,00 | < 0,001 |
| K65R1 | 114 / 6,04 | 0 / 0,00 | 105 / 9,95 | < 0,001 |
| V106I | 113 / 5,99 | 9 / 2,19 | 93 / 8,82 | 0,001 |
| K101E1 | 108 / 5,72 | 5 / 1,22 | 95 / 9,00 | < 0,001 |
| Y181C1 | 87 / 4,61 | 2 / 0,49 | 76 / 7,20 | < 0,001 |
| M184I1 | 58 / 3,07 | 1 / 0,24 | 54 / 5,12 | < 0,001 |
| H221Y | 46 / 2,44 | 1 / 0,24 | 41 / 3,89 | 0,002 |
| V179E | 42 / 2,22 | 4 / 0,97 | 29 / 2,75 | 1,000 |
| E138K | 36 / 1,91 | 0 / 0,00 | 31 / 2,94 | 0,004 |
| P225H1 | 34 / 1,80 | 1 / 0,24 | 31 / 2,94 | 0,037 |
| Y115F1 | 31 / 1,64 | 0 / 0,00 | 28 / 2,65 | 0,012 |
| D67N1 | 29 / 1,54 | 2 / 0,49 | 26 / 2,46 | 0,675 |
| Y318F | 28 / 1,48 | 0 / 0,00 | 26 / 2,46 | 0,019 |
3.4. Cell Receptor Tropism Prediction
| Rule | Number of positive samples (proportion from the total number of samples) | |
|---|---|---|
| A net charge rule of ≥+5 and total number of charged amino acids in V3-loop of ≥ 8 | 23 (2.5%) | |
| Loss of the N-linked glycosylation site in V3-loop and a net charge of ≥+4 | 11 (1.2%) | |
| R or K at position 11 and/or K at position 25 of V3-loop (11/25 rule) | 10 (1.1%) | |
| R at position 25 of V3-loop and a net charge of ≥+5 | 5 (0.5%) |
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| SDRM | Surveillance Drug Resistance Mutations |
| ART | Anti-Retroviral Therapy |
| PIs | Protease inhibitors |
| NRTIs | Nucleoside reverse transcriptase inhibitors |
| NNRTIs | Non-nucleoside reverse transcriptase inhibitors |
| INSTIs | Integrase strand transfer inhibitors |
References
- Keller SC, Yehia BR, Eberhart MG, Brady KA. Accuracy of definitions for linkage to care in persons living with HIV. J Acquir Immune Defic Syndr. 2013 Aug 15;63(5):622-30. PMID: 23614992; PMCID: PMC3796149. [CrossRef]
- Samji H, Cescon A, Hogg RS, Modur SP, Althoff KN, Buchacz K, Burchell AN, Cohen M, Gebo KA, Gill MJ, Justice A, Kirk G, Klein MB, Korthuis PT, Martin J, Napravnik S, Rourke SB, Sterling TR, Silverberg MJ, Deeks S, Jacobson LP, Bosch RJ, Kitahata MM, Goedert JJ, Moore R, Gange SJ; North American AIDS Cohort Collaboration on Research and Design (NA-ACCORD) of IeDEA. Closing the gap: increases in life expectancy among treated HIV-positive individuals in the United States and Canada. PLoS One. 2013 Dec 18;8(12):e81355. PMID: 24367482; PMCID: PMC 3867319. [CrossRef]
- Wensing AM, Calvez V, Ceccherini-Silberstein F, Charpentier C, Günthard HF, Paredes R, Shafer RW, Richman DD. 2022 update of the drug resistance mutations in HIV-1. Top Antivir Med. 2022 Oct;30(4):559-574. PMID: 36375130; PMCID: PMC9681141.
- Puertas MC, Ploumidis G, Ploumidis M, Fumero E, Clotet B, Walworth CM, Petropoulos CJ, Martinez-Picado J. Pan-resistant HIV-1 emergence in the era of integrase strand-transfer inhibitors: a case report. Lancet Microbe. 2020 Jul;1(3):e130-e135. Epub 2020 Jun 11. PMID: 35544263. [CrossRef]
- King JR, Wynn H, Brundage R, Acosta EP. Pharmacokinetic enhancement of protease inhibitor therapy. Clin Pharmacokinet. 2004;43(5):291-310. PMID: 15080763. [CrossRef]
- Larder B. Mechanisms of HIV-1 drug resistance. AIDS. 2001;15 Suppl 5:S27-34. PMID: 11816171. [CrossRef]
- Finzi D, Hermankova M, Pierson T, Carruth LM, Buck C, Chaisson RE, Quinn TC, Chadwick K, Margolick J, Brookmeyer R, Gallant J, Markowitz M, Ho DD, Richman DD, Siliciano RF. Identification of a reservoir for HIV-1 in patients on highly active antiretroviral therapy. Science. 1997 Nov 14;278(5341):1295-300. PMID: 9360927. [CrossRef]
- Bruner KM, Wang Z, Simonetti FR, Bender AM, Kwon KJ, Sengupta S, Fray EJ, Beg SA, Antar AAR, Jenike KM, Bertagnolli LN, Capoferri AA, Kufera JT, Timmons A, Nobles C, Gregg J, Wada N, Ho YC, Zhang H, Margolick JB, Blankson JN, Deeks SG, Bushman FD, Siliciano JD, Laird GM, Siliciano RF. A quantitative approach for measuring the reservoir of latent HIV-1 proviruses. Nature. 2019 Feb;566(7742):120-125. Epub 2019 Jan 30. PMID: 30700913; PMCID: PMC6447073. [CrossRef]
- Manyana S, Gounder L, Pillay M, Manasa J, Naidoo K, Chimukangara B. HIV-1 Drug Resistance Genotyping in Resource Limited Settings: Current and Future Perspectives in Sequencing Technologies. Viruses. 2021 Jun 11;13(6):1125. PMID: 34208165; PMCID: PMC8230827. [CrossRef]
- Ávila-Ríos S, Parkin N, Swanstrom R, Paredes R, Shafer R, Ji H, Kantor R. Next-Generation Sequencing for HIV Drug Resistance Testing: Laboratory, Clinical, and Implementation Considerations. Viruses. 2020 Jun 5;12(6):617. PMID: 32516949; PMCID: PMC7354449. [CrossRef]
- Weber J, Volkova I, Sahoo MK, Tzou PL, Shafer RW, Pinsky BA. Prospective Evaluation of the Vela Diagnostics Next-Generation Sequencing Platform for HIV-1 Genotypic Resistance Testing. J Mol Diagn. 2019 Nov;21(6):961-970. Epub 2019 Aug 2. PMID: 31382033; PMCID: PMC7152740. [CrossRef]
- Raymond S, Nicot F, Abravanel F, Minier L, Carcenac R, Lefebvre C, Harter A, Martin-Blondel G, Delobel P, Izopet J. Performance evaluation of the Vela Dx Sentosa next-generation sequencing system for HIV-1 DNA genotypic resistance. J Clin Virol. 2020 Jan; 122:104229. Epub 2019 Nov 26. PMID: 31809945. [CrossRef]
- Wymant C. et al. Easy and accurate reconstruction of whole HIV genomes from short-read sequence data with shiver. Virus evolution. 2018 May; 4 (1). pp.vey007.. [CrossRef]
- Available online: https://github.com/hivdb/sierra.
- Bennett DE, Camacho RJ, Otelea D, Kuritzkes DR, Fleury H, Kiuchi M, et al. (2009) Drug Resistance Mutations for Surveillance of Transmitted HIV-1 Drug-Resistance: 2009 Update. PLoS ONE 4(3): e4724. [CrossRef]
- Bailey AJ, Rhee SY, Shafer RW. Integrase Strand Transfer Inhibitor Resistance in Integrase Strand Transfer Inhibitor-Naive Persons. AIDS Res Hum Retroviruses. 2021 Oct;37(10):736-743. Epub 2021 Apr 15. PMID: 33683148; PMCID: PMC8665799. [CrossRef]
- Riemenschneider, M., Cashin, K., Budeus, B. et al. Genotypic Prediction of Co-receptor Tropism of HIV-1 Subtypes A and C. Sci Rep 6, 24883 (2016). [CrossRef]
- Esbjörnsson, J., Månsson, F., Martínez-Arias, W. et al. Frequent CXCR4 tropism of HIV-1 subtype A and CRF02_AG during late-stage disease - indication of an evolving epidemic in West Africa. Retrovirology 7, 23 (2010). [CrossRef]
- Raymond, Stéphaniea; Delobel, Pierrea; Mavigner, Mauda; Cazabat, Michellea; Souyris, Corinnea; Sandres-Sauné, Karinea; Cuzin, Lised; Marchou, Brunob; Massip, Patriceb; Izopet, Jacquesa. Correlation between genotypic predictions based on V3 sequences and phenotypic determination of HIV-1 tropism. AIDS 22(14):p F11-F16, September 12, 2008. |. [CrossRef]
- Available online: https://hivfrenchresistance.org/hiv-french-resistance-hiv-tropism/.
- Sede MM, Moretti FA, Laufer NL, Jones LR, Quarleri JF (2014) HIV-1 Tropism Dynamics and Phylogenetic Analysis from Longitudinal Ultra-Deep Sequencing Data of CCR5- and CXCR4-Using Variants. PLoS ONE 9(7): e102857. [CrossRef]
- Kirichenko, A.; Kireev, D.; Lapovok, I.; Shlykova, A.; Lopatukhin, A.; Pokrovskaya, A.; Bobkova, M.; Antonova, A.; Kuznetsova, A.; Ozhmegova, E.; et al. HIV-1 Drug Resistance among Treatment-Naïve Patients in Russia: Analysis of the National Database, 2006–2022. Viruses 2023, 15, 991. [CrossRef]
- Kirichenko A.A., Kireev D.E., Lopatukhin A.E., Murzakova A.V., Lapovok I.A., Ladnaya N.N., Pokrovsky V.V. Prevalence And Structure Of Hiv-1 Drug Resistance Among Treatment Naïve Patients Since The Introduction Of Antiretroviral Therapy In The Russian Federation. HIV Infection and Immunosuppressive Disorders. 2019;11(2):75-83. (In Russ.) . [CrossRef]
- Lapovok I. et al. Prevalence of HIV-1 drug resistance mutations among ART-naïve patients in Russia from 2005 to 2015. 14th European Meeting on HIV & Hepatitis, Rome, Italy. 2016. pp. 25-27.
- Singh A, Sunpath H, Green TN, Padayachi N, Hiramen K, Lie Y, Anton ED, Murphy R, Reeves JD, Kuritzkes DR, Ndung’u T. Drug resistance and viral tropism in HIV-1 subtype C-infected patients in KwaZulu-Natal, South Africa: implications for future treatment options. J Acquir Immune Defic Syndr. 2011 Nov 1;58(3):233-40. PMID: 21709569; PMCID: PMC3196677. [CrossRef]
- Williams A, Menon S, Crowe M, Agarwal N, Biccler J, Bbosa N, Ssemwanga D, Adungo F, Moecklinghoff C, Macartney M, Oriol-Mathieu V. Geographic and Population Distributions of Human Immunodeficiency Virus (HIV)-1 and HIV-2 Circulating Subtypes: A Systematic Literature Review and Meta-analysis (2010-2021). J Infect Dis. 2023 Nov 28;228(11):1583-1591. PMID: 37592824; PMCID: PMC10681860. [CrossRef]
| Mutation | Total (n = 1888) Abs. / % |
ART Experience (n = 1466) | ||
|---|---|---|---|---|
| No (n = 411) Abs. / % |
Yes (n = 1055) Abs. / % |
p | ||
| M46I1 | 15 / 0,79 | 4 / 0,97 | 10 / 0,95 | 1,000 |
| K43T | 10 / 0,53 | 2 / 0,49 | 5 / 0,47 | 1,000 |
| L33F | 10 / 0,53 | 1 / 0,24 | 4 / 0,38 | 1,000 |
| Q58E | 6 / 0,32 | 0 / 0,00 | 5 / 0,47 | 1,000 |
| V11I | 4 / 0,21 | 0 / 0,00 | 2 / 0,19 | 1,000 |
| I54S1 | 4 / 0,21 | 1 / 0,24 | 2 / 0,19 | 1,000 |
| F53L1 | 3 / 0,16 | 0 / 0,00 | 2 / 0,19 | 1,000 |
| I47V1 | 2 / 0,11 | 0 / 0,00 | 2 / 0,19 | 1,000 |
| I54V1 | 2 / 0,11 | 0 / 0,00 | 1 / 0,09 | 1,000 |
| M46L1 | 2 / 0,11 | 1 / 0,24 | 1 / 0,09 | 1,000 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




