Submitted:
13 January 2026
Posted:
14 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Vaccines Design
2.2. 3D Structure Prediction
2.3. Plasmid Construction
2.4. Bacterial Strain and Transformation
2.5. Colony PCR Verification
2.6. Vaccine Production
2.7. Post-Inactivation Testing
2.8. Flow Cytometry
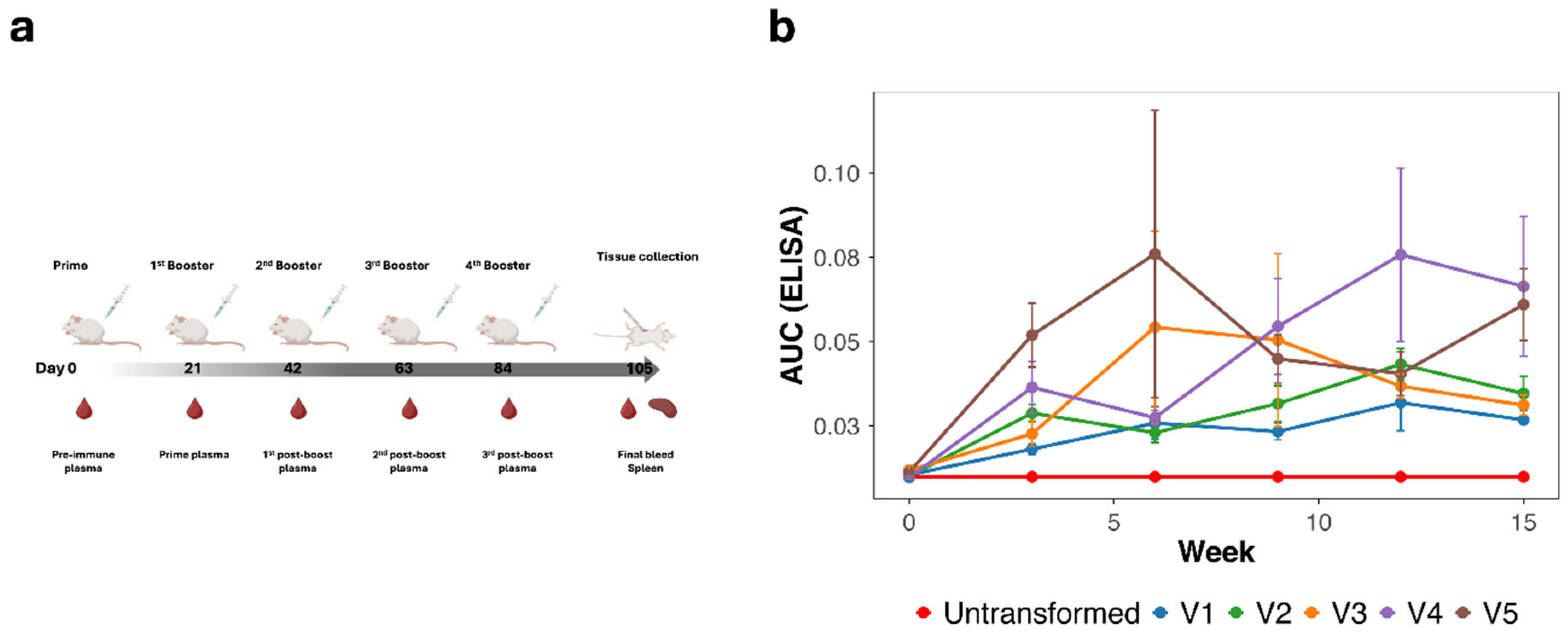
2.9. Mouse Immunization
2.10. Enzyme-Linked Immunosorbent Assay (ELISA)
2.11. Neutralization Assays
2.12. Statistical Analysis
2.13. Ethics, Animal Welfare, and Biosafety
3. Results
3.1. Scaffold-MPER KWC/GRB Vaccine Design
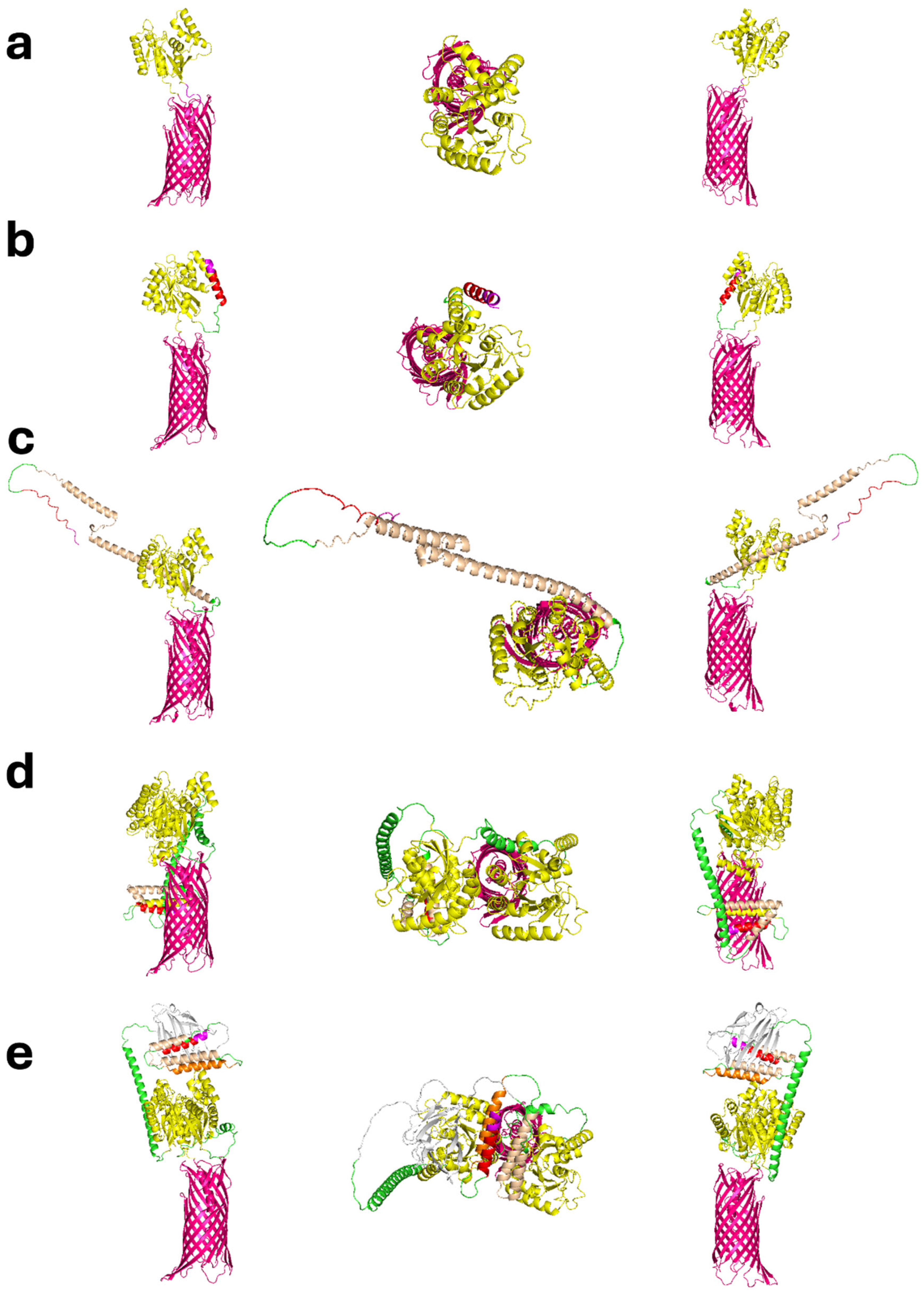
3.2. Structural Modeling and Predicted Architecture
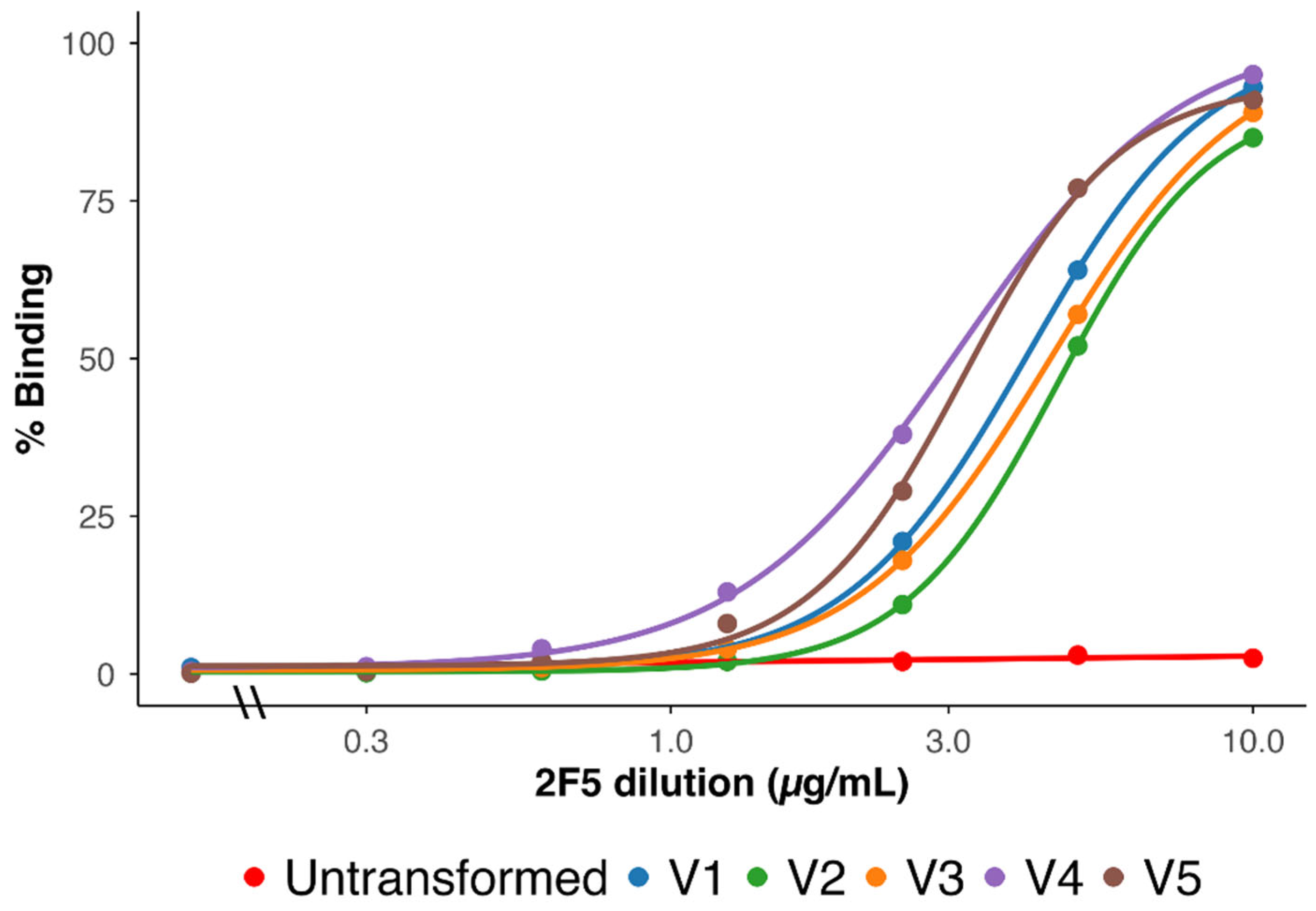
3.3. Surface Expression and bNAb 2F5 Recognition
3.4. Induction of MPER-Specific Antibodies (ELISA)
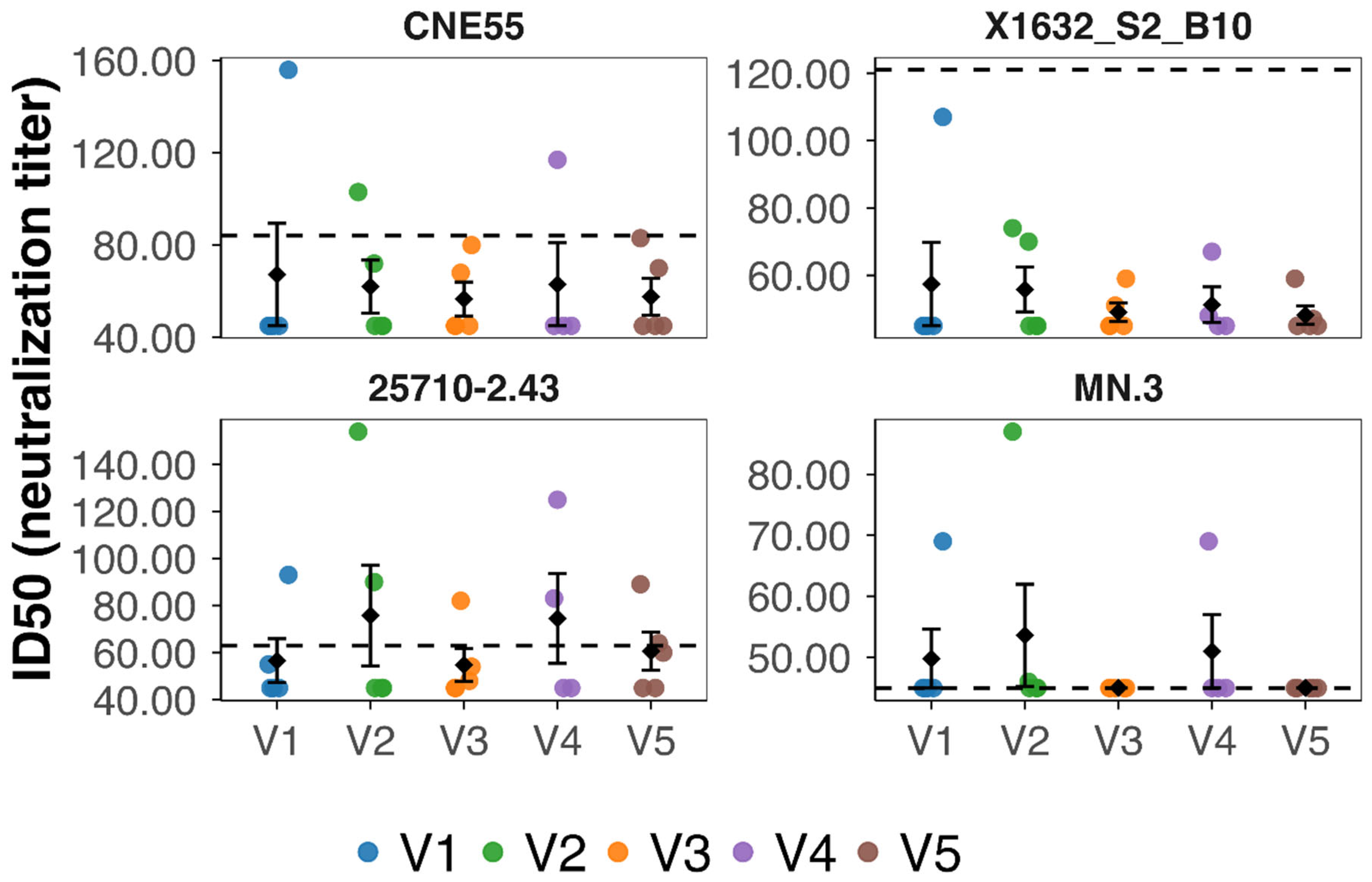
3.5. Neutralizing Activity Against HIV-1 Pseudoviruses
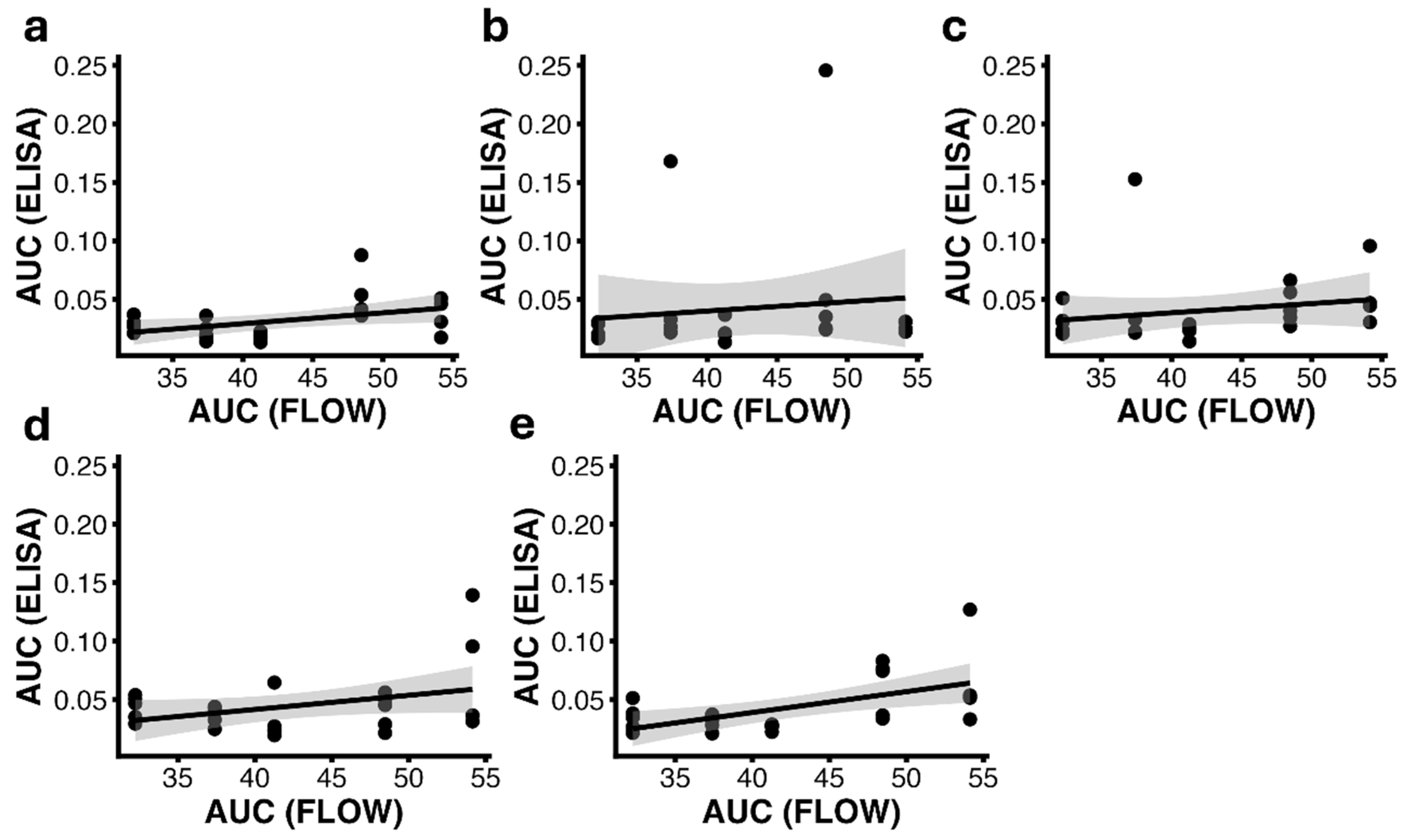
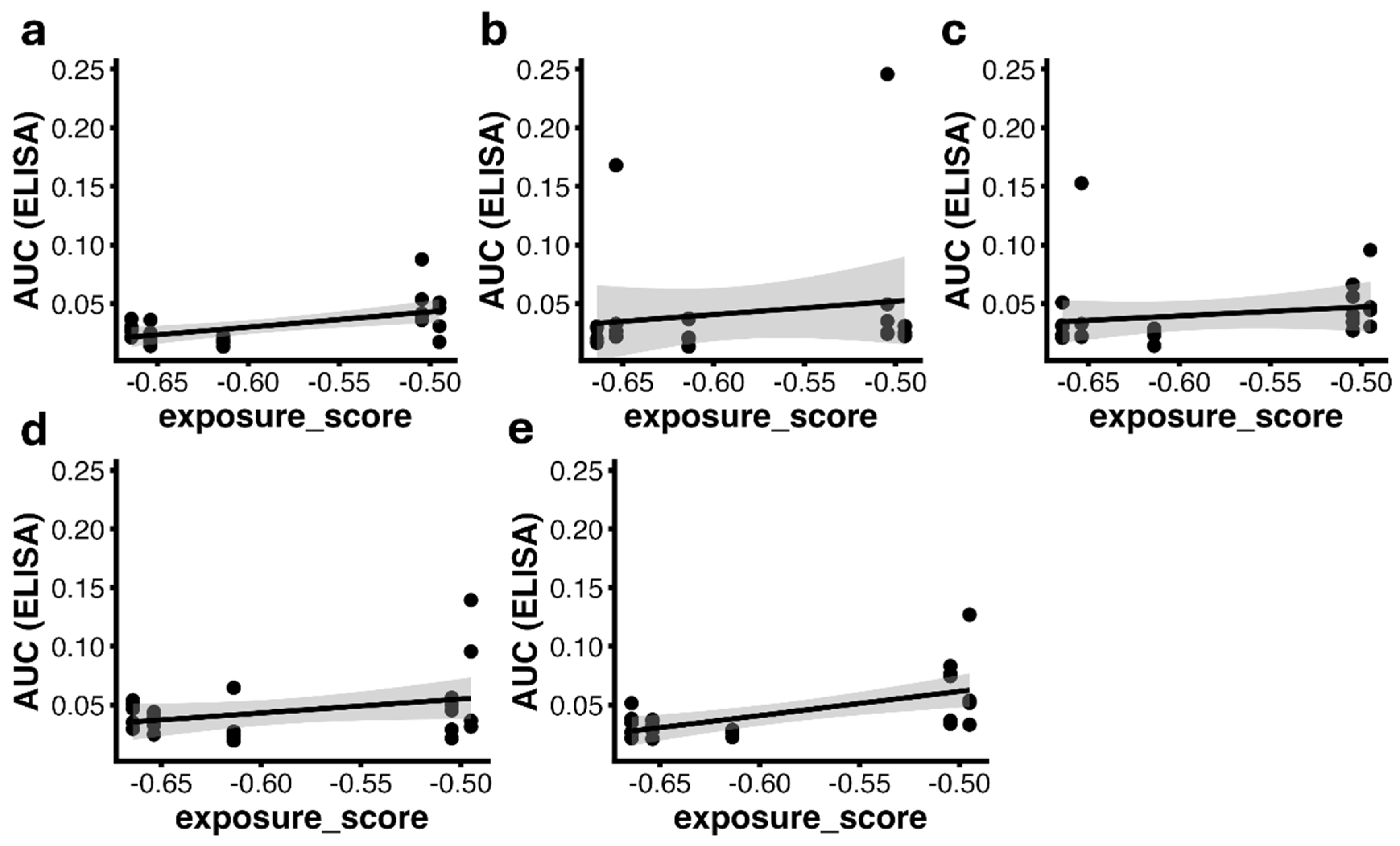
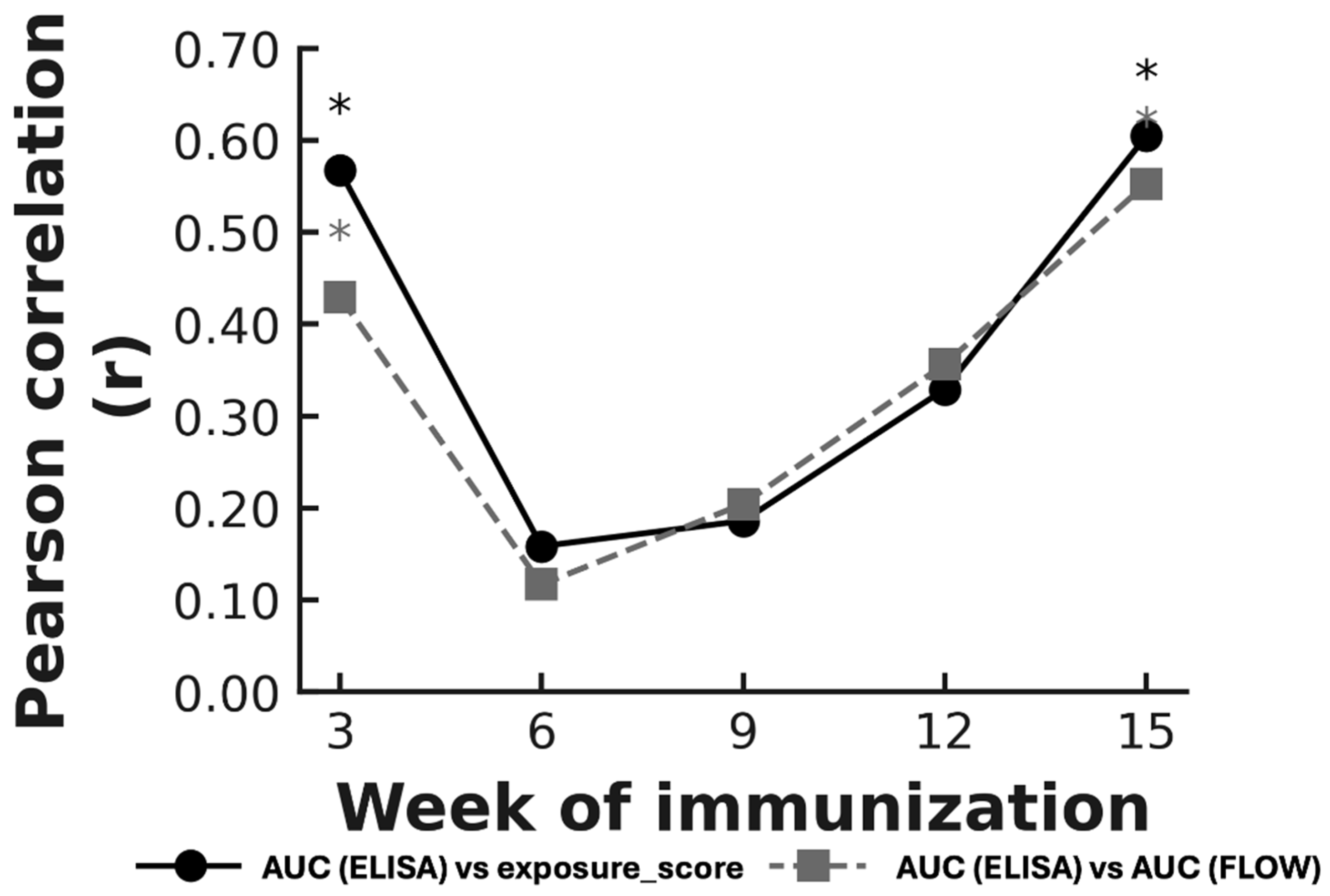
3.6. Cross-Assay Correlations Between Antigen Exposure and Humoral Response
3.7. Relationship Between Surface Antigen Display and Neutralizing Activity Induction
3.8. Relationship Between MPER-Specific Antibody Binding and Neutralization
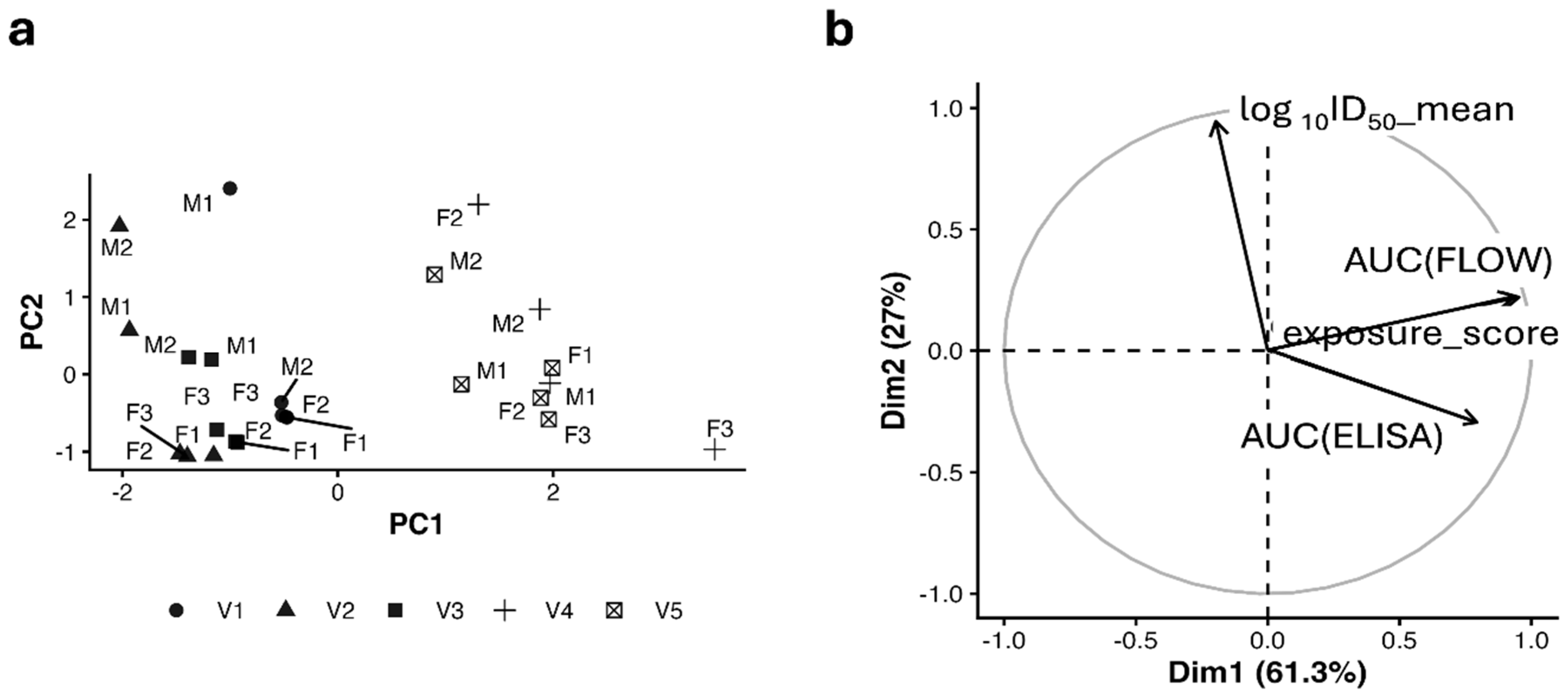
3.9. Integrated Multivariate Modeling of Display, Binding, and Neutralization
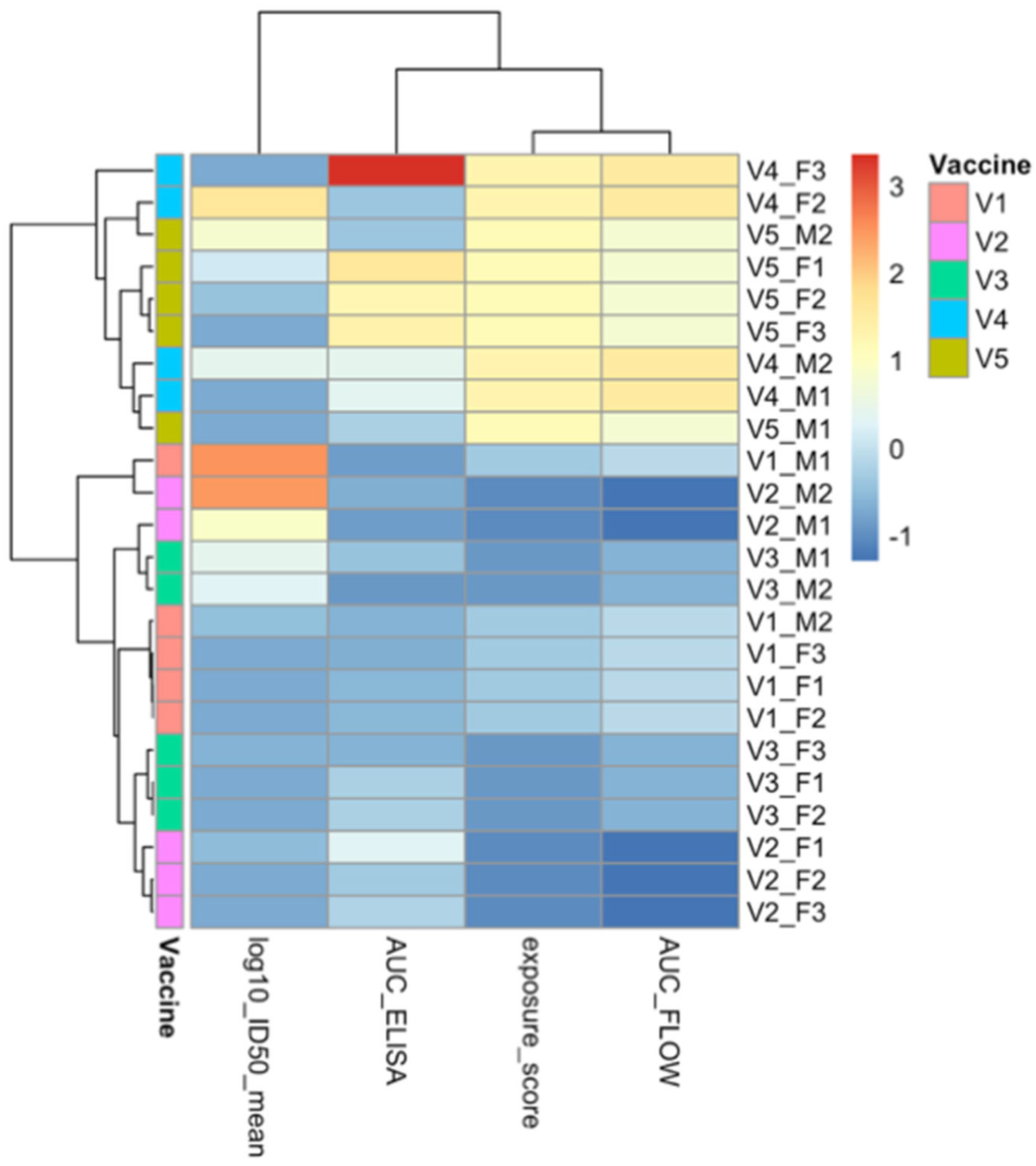
3.10. Integrated Correlation Network and Data-Driven Clustering
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| 3AGJ | Scaffolded MPER antigen based on PDB 3AGJ |
| 3DA | DADADA N-terminal motif |
| AEAAAK | Rigid α-helical linker motif AEAAAK (EAAAK repeats) |
| AIDA | Adhesin Involved in Diffuse Adherence |
| AIDA-I | Adhesin Involved in Diffuse Adherence I autotransporter |
| AQQASSS | Semi-flexible α-helical linker motif AQQASSS |
| AT | Autotransporter |
| ATs | Autotransporters |
| AUC | Area Under the Curve |
| AUC(ELISA) | ELISA area under the curve |
| AUC(FLOW) | Flow cytometry area under the curve |
| BCR | B-cell receptor |
| BSA | Bovine serum albumin |
| BSL-2 | Biosafety level 2 |
| CFU | Colony-forming unit |
| DBTL | Design–Build–Test–Learn |
| EC50 | Half-maximal effective concentration |
| ELISA | Enzyme-Linked Immunosorbent Assay |
| Env | Envelope glycoprotein |
| FBS | Fetal bovine serum |
| FDR | False discovery rate |
| FLIC | Flagellin fragment (TLR5 agonist) |
| FP | Fusion Peptide |
| FP-like | Fusion peptide-like |
| GC | Germinal center |
| HBSS | Hank’s Balanced Salt Solution |
| HET3 | Four-way crossbred HET3 mouse stock |
| HIV | Human Immunodeficiency Virus |
| HIV-1 | Human Immunodeficiency Virus type 1 |
| HRP | Horseradish peroxidase |
| ID50 | 50% inhibitory dilution titer |
| ID80 | 80% inhibitory dilution titer |
| IgG | Immunoglobulin G |
| KWC/GRB | Killed Whole-Cell Genome-Reduced Bacteria |
| LB | Luria–Bertani broth/agar |
| ME5125 | Genome-reduced Escherichia coli strain ME5125 |
| MG1655 | Escherichia coli strain MG1655 |
| MPER | Membrane-Proximal External Region |
| NCI | National Cancer Institute |
| NIAID | National Institute of Allergy and Infectious Diseases |
| NIFA | National Institute of Food and Agriculture |
| NIH | National Institutes of Health |
| OD600 | Optical density at 600 nm |
| OX40L | OX40 ligand |
| PADRE | Pan DR Epitope (universal CD4+ T-helper epitope) |
| PBS | Phosphate-buffered saline |
| PC1 | Principal component 1 |
| PC2 | Principal component 2 |
| PCA | Principal component analysis |
| PEDV | Porcine epidemic diarrhea virus |
| RAU | Relative antibody units |
| RLU | Relative luminescence units |
| ROC | Receiver operating characteristic |
| SEM | Standard error of the mean |
| SOC | Super Optimal broth with Catabolite repression |
| SPF | Specific-pathogen-free |
| TBS | Tris-buffered saline |
| TBST | Tris-buffered saline with Tween-20 |
| TLR2 | Toll-like receptor 2 |
| TLR2/4 | Toll-like receptors 2 and 4 |
| TLR4 | Toll-like receptor 4 |
| TLR5 | Toll-like receptor 5 |
| TMB | Tetramethylbenzidine |
| TZM-bl | HIV-1 reporter cell line TZM-bl |
| USA | United States of America |
| USDA | United States Department of Agriculture |
| UVA | University of Virginia |
| V1 | Scaffold-MPER vaccine 1 |
| V2 | Scaffold-MPER vaccine 2 |
| V3 | Scaffold-MPER vaccine 3 |
| V4 | Scaffold-MPER vaccine 4 |
| V5 | Scaffold-MPER vaccine 5 |
| bnAb | Broadly neutralizing antibody |
| bnAbs | Broadly neutralizing antibodies |
| gp41 | Glycoprotein 41 |
| rSIP | Recombinant surface immunogenic protein |
References
- Plotkin, S. Vaccines: Past, Present and Future. Nat Med 2005, 11, S5–S11. [Google Scholar]
- Plotkin, S.A.; Gilbert, P.B. Nomenclature for Immune Correlates of Protection After Vaccination. Clinical Infectious Diseases 2012, 54, 1615–1617. [Google Scholar] [CrossRef] [PubMed]
- Greenwood, B. The Contribution of Vaccination to Global Health: Past, Present and Future. Phil. Trans. R. Soc. B 2014, 369, 20130433. [Google Scholar] [CrossRef]
- Pardi, N.; Hogan, M.J.; Porter, F.W.; Weissman, D. mRNA Vaccines — a New Era in Vaccinology. Nat Rev Drug Discov 2018, 17, 261–279. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Sreenivasan, C.; Sheng, Z.; Zhai, S.-L.; Wollman, J.W.; Luo, S.; Huang, C.; Gao, R.; Wang, Z.; Kaushik, R.S.; et al. A Recombinant Chimeric Influenza Virus Vaccine Expressing the Consensus H3 Hemagglutinin Elicits Broad Hemagglutination Inhibition Antibodies against Divergent Swine H3N2 Influenza Viruses. Vaccine 2023, 41, 6318–6326. [Google Scholar] [CrossRef] [PubMed]
- Hotez, P.J.; Bottazzi, M.E. Whole Inactivated Virus and Protein-Based COVID-19 Vaccines. Annu. Rev. Med. 2022, 73, 55–64. [Google Scholar] [CrossRef]
- Broadbent, A.J.; Subbarao, K. Influenza Virus Vaccines: Lessons from the 2009 H1N1 Pandemic. Curr Opin Virol 2011, 1, 254–262. [Google Scholar] [CrossRef]
- Corey, L.; Mascola, J.R.; Fauci, A.S.; Collins, F.S. A Strategic Approach to COVID-19 Vaccine R&D. Science 2020, 368, 948–950. [Google Scholar] [CrossRef]
- Ofek, G.; Guenaga, F.J.; Schief, W.R.; Skinner, J.; Baker, D.; Wyatt, R.; Kwong, P.D. Elicitation of Structure-Specific Antibodies by Epitope Scaffolds. Proc. Natl. Acad. Sci. U.S.A. 2010, 107, 17880–17887. [Google Scholar] [CrossRef]
- Correia, B.E.; Ban, Y.-E.A.; Holmes, M.A.; Xu, H.; Ellingson, K.; Kraft, Z.; Carrico, C.; Boni, E.; Sather, D.N.; Zenobia, C.; et al. Computational Design of Epitope-Scaffolds Allows Induction of Antibodies Specific for a Poorly Immunogenic HIV Vaccine Epitope. Structure 2010, 18, 1116–1126. [Google Scholar] [CrossRef]
- Zhou, T.; Zhu, J.; Yang, Y.; Gorman, J.; Ofek, G.; Srivatsan, S.; Druz, A.; Lees, C.R.; Lu, G.; Soto, C.; et al. Transplanting Supersites of HIV-1 Vulnerability. PLoS ONE 2014, 9, e99881. [Google Scholar] [CrossRef]
- Rujas, E.; Leaman, D.P.; Insausti, S.; Ortigosa-Pascual, L.; Zhang, L.; Zwick, M.B.; Nieva, J.L. Functional Optimization of Broadly Neutralizing HIV-1 Antibody 10E8 by Promotion of Membrane Interactions. J Virol 2018, 92, e02249-17. [Google Scholar] [CrossRef]
- Eslami, M.; Fadaee Dowlat, B.; Yaghmayee, S.; Habibian, A.; Keshavarzi, S.; Oksenych, V.; Naderian, R. Next-Generation Vaccine Platforms: Integrating Synthetic Biology, Nanotechnology, and Systems Immunology for Improved Immunogenicity. Vaccines 2025, 13, 588. [Google Scholar] [CrossRef]
- Barrett, A.; Stanberry, L. Vaccines for Biodefense and Emerging and Neglected Diseases; Academic Press: London, 2009; ISBN 978-0-12-369408-9. [Google Scholar]
- Buck, P.O.; Gomes, D.A.; Beck, E.; Kirson, N.; Mattera, M.; Carroll, S.; Ultsch, B.; Jayasundara, K.; Uhart, M.; Garrison, L.P., Jr. New Vaccine Platforms—Novel Dimensions of Economic and Societal Value and Their Measurement. Vaccines 2024, 12, 234. [Google Scholar] [CrossRef]
- Park, H.; Otte, A.; Park, K. Evolution of Drug Delivery Systems: From 1950 to 2020 and Beyond. Journal of Controlled Release 2022, 342, 53–65. [Google Scholar] [CrossRef]
- Graham, B.S.; Sullivan, N.J. Emerging Viral Diseases from a Vaccinology Perspective: Preparing for the next Pandemic. Nat Immunol 2018, 19, 20–28. [Google Scholar] [CrossRef] [PubMed]
- Burton, D.R.; Hangartner, L. Broadly Neutralizing Antibodies to HIV and Their Role in Vaccine Design. Annual Review of Immunology 2016, 34, 635–659. [Google Scholar] [CrossRef]
- McCoy, L.E.; Burton, D.R. Identification and Specificity of Broadly Neutralizing Antibodies against HIV. Immunological Reviews 2017, 275, 11–20. [Google Scholar] [CrossRef] [PubMed]
- Holmgren, J.; Czerkinsky, C. Mucosal Immunity and Vaccines. Nat Med 2005, 11, S45–S53. [Google Scholar] [CrossRef]
- Lycke, N. Recent Progress in Mucosal Vaccine Development: Potential and Limitations. Nat Rev Immunol 2012, 12, 592–605. [Google Scholar] [CrossRef] [PubMed]
- Franquelim, H.G.; Chiantia, S.; Veiga, A.S.; Santos, N.C.; Schwille, P.; Castanho, M.A. Anti-HIV-1 Antibodies 2F5 and 4E10 Interact Differently with Lipids to Bind Their Epitopes. Aids 2011, 25, 419–428. [Google Scholar] [CrossRef]
- Kong, R.; Xu, K.; Zhou, T.; Acharya, P.; Lemmin, T.; Liu, K.; Ozorowski, G.; Soto, C.; Taft, J.D.; Bailer, R.T.; et al. Fusion Peptide of HIV-1 as a Site of Vulnerability to Neutralizing Antibody. Science 2016, 352, 828–833. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Yang, Z.-Y.; Li, Y.; Hogerkorp, C.-M.; Schief, W.R.; Seaman, M.S.; Zhou, T.; Schmidt, S.D.; Wu, L.; Xu, L.; et al. Rational Design of Envelope Identifies Broadly Neutralizing Human Monoclonal Antibodies to HIV-1. Science 2010, 329, 856–861. [Google Scholar] [CrossRef] [PubMed]
- Irimia, A.; Serra, A.M.; Sarkar, A.; Jacak, R.; Kalyuzhniy, O.; Sok, D.; Saye-Francisco, K.L.; Schiffner, T.; Tingle, R.; Kubitz, M.; et al. Lipid Interactions and Angle of Approach to the HIV-1 Viral Membrane of Broadly Neutralizing Antibody 10E8: Insights for Vaccine and Therapeutic Design. PLoS Pathog 2017, 13, e1006212. [Google Scholar] [CrossRef]
- Burton, D.R.; Poignard, P.; Stanfield, R.L.; Wilson, I.A. Broadly Neutralizing Antibodies Present New Prospects to Counter Highly Antigenically Diverse Viruses. Science 2012, 337, 183–186. [Google Scholar] [CrossRef]
- Wang, Q.; Zhang, L. Broadly Neutralizing Antibodies and Vaccine Design against HIV-1 Infection. Front. Med. 2020, 14, 30–42. [Google Scholar] [CrossRef]
- Williams, W.B.; Wiehe, K.; Saunders, K.O.; Haynes, B.F. Strategies for Induction of HIV-1 Envelope-Reactive Broadly Neutralizing Antibodies. Journal of the International AIDS Society 2021, 24, e25831. [Google Scholar] [CrossRef]
- Brouwer, P.J.M.; Antanasijevic, A.; Berndsen, Z.; Yasmeen, A.; Fiala, B.; Bijl, T.P.L.; Bontjer, I.; Bale, J.B.; Sheffler, W.; Allen, J.D.; et al. Enhancing and Shaping the Immunogenicity of Native-like HIV-1 Envelope Trimers with a Two-Component Protein Nanoparticle. Nat Commun 2019, 10. [Google Scholar] [CrossRef]
- Ou, L.; Kong, W.-P.; Chuang, G.-Y.; Ghosh, M.; Gulla, K.; O’Dell, S.; Varriale, J.; Barefoot, N.; Changela, A.; Chao, C.W.; et al. Preclinical Development of a Fusion Peptide Conjugate as an HIV Vaccine Immunogen. Sci Rep 2020, 10, 3032. [Google Scholar] [CrossRef]
- Huang, J.; Ofek, G.; Laub, L.; Louder, M.K.; Doria-Rose, N.A.; Longo, N.S.; Imamichi, H.; Bailer, R.T.; Chakrabarti, B.; Sharma, S.K.; et al. Broad and Potent Neutralization of HIV-1 by a Gp41-Specific Human Antibody. Nature 2012, 491, 406–412. [Google Scholar] [CrossRef]
- van Gils, M.J.; van den Kerkhof, T.L.G.M.; Ozorowski, G.; Cottrell, C.A.; Sok, D.; Pauthner, M.; Pallesen, J.; de Val, N.; Yasmeen, A.; de Taeye, S.W.; et al. An HIV-1 Antibody from an Elite Neutralizer Implicates the Fusion Peptide as a Site of Vulnerability. Nat Microbiol 2016, 2, 16199. [Google Scholar] [CrossRef]
- Domene, C.; Wiley, B.; Insausti, S.; Rujas, E.; Nieva, J.L. Distinctive Membrane Accommodation Traits Underpinning the Neutralization Activity of HIV-1 Antibody against MPER. Mol. Pharmaceutics 2025, 22, 2494–2508. [Google Scholar] [CrossRef]
- Pinto, D.; Fenwick, C.; Caillat, C.; Silacci, C.; Guseva, S.; Dehez, F.; Chipot, C.; Barbieri, S.; Minola, A.; Jarrossay, D.; et al. Structural Basis for Broad HIV-1 Neutralization by the MPER-Specific Human Broadly Neutralizing Antibody LN01. Cell Host & Microbe 2019, 26, 623–637.e8. [Google Scholar] [CrossRef]
- Barouch, D.H. Challenges in the Development of an HIV-1 Vaccine. Nature 2008, 455, 613–619. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.H.; Rerks-Ngarm, S.; Excler, J.-L.; Michael, N.L. HIV Vaccines—Lessons Learned and the Way Forward. Curr Opin HIV AIDS 2010, 5, 428–434. [Google Scholar] [CrossRef] [PubMed]
- Kong, R.; Duan, H.; Sheng, Z.; Xu, K.; Acharya, P.; Chen, X.; Cheng, C.; Dingens, A.S.; Gorman, J.; Sastry, M.; et al. Antibody Lineages with Vaccine-Induced Antigen-Binding Hotspots Develop Broad HIV Neutralization. Cell 2019, 178, 567–584.e19. [Google Scholar] [CrossRef]
- Liu, H.; Su, X.; Si, L.; Lu, L.; Jiang, S. The Development of HIV Vaccines Targeting Gp41 Membrane-Proximal External Region (MPER): Challenges and Prospects. Protein & Cell 2018, 9, 596–615. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Cheng, C.; Santo, J.L.D.; Shen, C.-H.; Bylund, T.; Henry, A.R.; Howe, C.A.; Hwang, J.; Morano, N.C.; Morris, D.J.; et al. Potent and Broad HIV-1 Neutralization in Fusion Peptide-Primed SHIV-Infected Macaques. Cell 2024, 187, 7214–7231.e23. [Google Scholar] [CrossRef]
- Cheng, C.; Xu, K.; Kong, R.; Chuang, G.-Y.; Corrigan, A.R.; Geng, H.; Hill, K.R.; Jafari, A.J.; O’Dell, S.; Ou, L.; et al. Consistent Elicitation of Cross-Clade HIV-Neutralizing Responses Achieved in Guinea Pigs after Fusion Peptide Priming by Repetitive Envelope Trimer Boosting. PLoS ONE 2019, 14, e0215163. [Google Scholar] [CrossRef]
- Guenaga, J.; Dosenovic, P.; Ofek, G.; Baker, D.; Schief, W.R.; Kwong, P.D.; Karlsson Hedestam, G.B.; Wyatt, R.T. Heterologous Epitope-Scaffold Prime∶Boosting Immuno-Focuses B Cell Responses to the HIV-1 Gp41 2F5 Neutralization Determinant. PLoS ONE 2011, 6, e16074. [Google Scholar] [CrossRef]
- Ward, A.B.; Wilson, I.A. The HIV -1 Envelope Glycoprotein Structure: Nailing down a Moving Target. Immunological Reviews 2017, 275, 21–32. [Google Scholar] [CrossRef] [PubMed]
- Azoitei, M.L.; Ban, Y.-E.A.; Julien, J.-P.; Bryson, S.; Schroeter, A.; Kalyuzhniy, O.; Porter, J.R.; Adachi, Y.; Baker, D.; Pai, E.F.; et al. Computational Design of High-Affinity Epitope Scaffolds by Backbone Grafting of a Linear Epitope. Journal of Molecular Biology 2012, 415, 175–192. [Google Scholar] [CrossRef]
- Rantalainen, K.; Berndsen, Z.T.; Antanasijevic, A.; Schiffner, T.; Zhang, X.; Lee, W.-H.; Torres, J.L.; Zhang, L.; Irimia, A.; Copps, J.; et al. HIV-1 Envelope and MPER Antibody Structures in Lipid Assemblies. Cell Reports 2020, 31, 107583. [Google Scholar] [CrossRef] [PubMed]
- Zeng, X.; Vidlund, J.; Gillespie, B.; Cao, L.; Agga, G.E.; Lin, J.; Dego, O.K. Evaluation of Immunogenicity of Enterobactin Conjugate Vaccine for the Control of Escherichia Coli Mastitis in Dairy Cows. Journal of Dairy Science 2023, 106, 7147–7163. [Google Scholar] [CrossRef]
- Bhattacharya, S.K.; Sur, D.; Ali, M.; Kanungo, S.; You, Y.A.; Manna, B.; Sah, B.; Niyogi, S.K.; Park, J.K.; Sarkar, B.; et al. 5 Year Efficacy of a Bivalent Killed Whole-Cell Oral Cholera Vaccine in Kolkata, India: A Cluster-Randomised, Double-Blind, Placebo-Controlled Trial. The Lancet Infectious Diseases 2013, 13, 1050–1056. [Google Scholar] [CrossRef] [PubMed]
- Ferreras, E.; Chizema-Kawesha, E.; Blake, A.; Chewe, O.; Mwaba, J.; Zulu, G.; Poncin, M.; Rakesh, A.; Page, A.-L.; Stoitsova, S.; et al. Single-Dose Cholera Vaccine in Response to an Outbreak in Zambia. N Engl J Med 2018, 378, 577–579. [Google Scholar] [CrossRef]
- Jose, J. Autodisplay: Efficient Bacterial Surface Display of Recombinant Proteins. Appl Microbiol Biotechnol 2006, 69, 607–614. [Google Scholar] [CrossRef]
- Henderson, I.R.; Navarro-Garcia, F.; Nataro, J.P. The Great Escape: Structure and Function of the Autotransporter Proteins. Trends in Microbiology 1998, 6, 370–378. [Google Scholar] [CrossRef]
- Henderson, I.R.; Navarro-Garcia, F.; Desvaux, M.; Fernandez, R.C.; Ala’Aldeen, D. Type V Protein Secretion Pathway: The Autotransporter Story. Microbiol Mol Biol Rev 2004, 68, 692–744. [Google Scholar] [CrossRef]
- Leyton, D.L.; Rossiter, A.E.; Henderson, I.R. From Self Sufficiency to Dependence: Mechanisms and Factors Important for Autotransporter Biogenesis. Nat Rev Microbiol 2012, 10, 213–225. [Google Scholar] [CrossRef] [PubMed]
- Benz, I.; Schmidt, M.A. AIDA-I, the Adhesin Involved in Diffuse Adherence of the Diarrhoeagenic Escherichia Coli Strain 2787 (O126:H27), Is Synthesized via a Precursor Molecule. Molecular Microbiology 1992, 6, 1539–1546. [Google Scholar] [CrossRef]
- Cotter, S.E.; Surana, N.K.; Grass, S.; St Geme, J.W., 3rd. Trimeric Autotransporters Require Trimerization of the Passenger Domain for Stability and Adhesive Activity. J Bacteriol 2006, 188, 5400–5407. [Google Scholar] [CrossRef]
- Meng, G.; Surana, N.K.; St Geme, J.W., 3rd; Waksman, G. Structure of the Outer Membrane Translocator Domain of the Haemophilus Influenzae Hia Trimeric Autotransporter. EMBO J 2006, 25, 2297–2304. [Google Scholar] [CrossRef]
- Surana, N.K.; Cutter, D.; Barenkamp, S.J.; St Geme, J.W., 3rd. The Haemophilus Influenzae Hia Autotransporter Contains an Unusually Short Trimeric Translocator Domain. J Biol Chem 2004, 279, 14679–14685. [Google Scholar] [CrossRef] [PubMed]
- Maeda, D.L.N.F.; Tian, D.; Yu, H.; Dar, N.; Rajasekaran, V.; Meng, S.; Mahsoub, H.M.; Sooryanarain, H.; Wang, B.; Heffron, C.L.; et al. Killed Whole-Genome Reduced-Bacteria Surface-Expressed Coronavirus Fusion Peptide Vaccines Protect against Disease in a Porcine Model. Proceedings of the National Academy of Sciences 2021, 118, e2025622118. [Google Scholar] [CrossRef]
- Quintero-Barbosa, J.S.; Song, Y.; Mehl, F.; Mathur, S.; Livingston, L.; Shen, X.; Montefiori, D.C.; Tan, J.; Zeichner, S.L. Engineering Enhanced Immunogenicity of Surface-Displayed Immunogens in a Killed Whole-Cell Genome-Reduced Bacterial Vaccine Platform Using Class I Viral Fusion Peptides. Vaccines 2025, 14, 14. [Google Scholar] [CrossRef]
- Alexander, J.; del Guercio, M.-F.; Maewal, A.; Qiao, L.; Fikes, J.; Chesnut, R.W.; Paulson, J.; Bundle, D.R.; DeFrees, S.; Sette, A. Linear PADRE T Helper Epitope and Carbohydrate B Cell Epitope Conjugates Induce Specific High Titer IgG Antibody Responses1. The Journal of Immunology 2000, 164, 1625–1633. [Google Scholar] [CrossRef] [PubMed]
- Del Guercio, M.-F.; Alexander, J.; Kubo, R.T.; Arrhenius, T.; Maewal, A.; Appella, E.; Hoffman, S.L.; Jones, T.; Valmori, D.; Sakaguchi, K. Potent Immunogenic Short Linear Peptide Constructs Composed of B Cell Epitopes and Pan DR T Helper Epitopes (PADRE) for Antibody Responses in Vivo. Vaccine 1997, 15, 441–448. [Google Scholar] [CrossRef]
- Lu, Y.; Swartz, J.R. Functional Properties of Flagellin as a Stimulator of Innate Immunity. Sci Rep 2016, 6, 18379. [Google Scholar] [CrossRef]
- Sharma, P.; Levy, O.; Dowling, D.J. The TLR5 Agonist Flagellin Shapes Phenotypical and Functional Activation of Lung Mucosal Antigen Presenting Cells in Neonatal Mice. Front. Immunol. 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Rady, H.F.; Dai, G.; Huang, W.; Shellito, J.E.; Ramsay, A.J. Flagellin Encoded in Gene-Based Vector Vaccines Is a Route-Dependent Immune Adjuvant. PLOS ONE 2016, 11, e0148701. [Google Scholar] [CrossRef]
- Murthy, K.G.; Deb, A.; Goonesekera, S.; Szabó, C.; Salzman, A.L. Identification of Conserved Domains in Salmonella Muenchen Flagellin That Are Essential for Its Ability to Activate TLR5 and to Induce an Inflammatory Response in Vitro. Journal of Biological Chemistry 2004, 279, 5667–5675. [Google Scholar] [CrossRef]
- Diaz-Dinamarca, D.A.; Manzo, R.A.; Soto, D.A.; Avendaño-Valenzuela, M.J.; Bastias, D.N.; Soto, P.I.; Escobar, D.F.; Vasquez-Saez, V.; Carrión, F.; Pizarro-Ortega, M.S.; et al. Surface Immunogenic Protein of Streptococcus Group B Is an Agonist of Toll-Like Receptors 2 and 4 and a Potential Immune Adjuvant. Vaccines 2020, 8, 29. [Google Scholar] [CrossRef] [PubMed]
- Asokan, M.; Rudicell, R.S.; Louder, M.; McKee, K.; O’Dell, S.; Stewart-Jones, G.; Wang, K.; Xu, L.; Chen, X.; Choe, M.; et al. Bispecific Antibodies Targeting Different Epitopes on the HIV-1 Envelope Exhibit Broad and Potent Neutralization. J Virol 2015, 89, 12501–12512. [Google Scholar] [CrossRef]
- Taylor, D.N.; Treanor, J.J.; Sheldon, E.A.; Johnson, C.; Umlauf, S.; Song, L.; Kavita, U.; Liu, G.; Tussey, L.; Ozer, K.; et al. Development of VAX128, a Recombinant Hemagglutinin (HA) Influenza-Flagellin Fusion Vaccine with Improved Safety and Immune Response. Vaccine 2012, 30, 5761–5769. [Google Scholar] [CrossRef]
- Croft, M. Control of Immunity by the TNFR-Related Molecule OX40 (CD134). Annu. Rev. Immunol. 2010, 28, 57–78. [Google Scholar] [CrossRef]
- Amet, N.; Lee, H.-F.; Shen, W.-C. Insertion of the Designed Helical Linker Led to Increased Expression of Tf-Based Fusion Proteins. Pharm Res 2009, 26, 523–528. [Google Scholar] [CrossRef]
- Jeong, W.H.; Lee, H.; Song, D.H.; Eom, J.-H.; Kim, S.C.; Lee, H.-S.; Lee, H.; Lee, J.-O. Connecting Two Proteins Using a Fusion Alpha Helix Stabilized by a Chemical Cross Linker. Nat Commun 2016, 7, 11031. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Zaro, J.; Shen, W.-C. Fusion Protein Linkers: Property, Design and Functionality. Adv Drug Deliv Rev 2013, 65, 1357–1369. [Google Scholar] [CrossRef] [PubMed]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly Accurate Protein Structure Prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- Goddard, T.D.; Huang, C.C.; Meng, E.C.; Pettersen, E.F.; Couch, G.S.; Morris, J.H.; Ferrin, T.E. UCSF ChimeraX: Meeting Modern Challenges in Visualization and Analysis. Protein Science 2018, 27, 14–25. [Google Scholar] [CrossRef]
- Meng, E.C.; Goddard, T.D.; Pettersen, E.F.; Couch, G.S.; Pearson, Z.J.; Morris, J.H.; Ferrin, T.E. UCSF ChimeraX: Tools for Structure Building and Analysis. Protein Science 2023, 32, e4792. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Meng, E.C.; Couch, G.S.; Croll, T.I.; Morris, J.H.; Ferrin, T.E. UCSF ChimeraX: Structure Visualization for Researchers, Educators, and Developers. Protein Science 2021, 30, 70–82. [Google Scholar] [CrossRef]
- Hashimoto, M.; Ichimura, T.; Mizoguchi, H.; Tanaka, K.; Fujimitsu, K.; Keyamura, K.; Ote, T.; Yamakawa, T.; Yamazaki, Y.; Mori, H.; et al. Cell Size and Nucleoid Organization of Engineered Escherichia Coli Cells with a Reduced Genome. Molecular Microbiology 2005, 55, 137–149. [Google Scholar] [CrossRef]
- Kato, J.; Hashimoto, M. Construction of Consecutive Deletions of the Escherichia Coli Chromosome. Molecular Systems Biology 2007, 3, 132. [Google Scholar] [CrossRef] [PubMed]
- Seaman, M.S.; Janes, H.; Hawkins, N.; Grandpre, L.E.; Devoy, C.; Giri, A.; Coffey, R.T.; Harris, L.; Wood, B.; Daniels, M.G.; et al. Tiered Categorization of a Diverse Panel of HIV-1 Env Pseudoviruses for Assessment of Neutralizing Antibodies. J Virol 2010, 84, 1439–1452. [Google Scholar] [CrossRef] [PubMed]
- Montefiori, D.C. Measuring HIV Neutralization in a Luciferase Reporter Gene Assay. In HIV Protocols; Prasad, V.R., Kalpana, G.V., Eds.; Humana Press: Totowa, NJ, 2009; pp. 395–405. ISBN 978-1-59745-170-3. [Google Scholar]
- Li, M.; Gao, F.; Mascola, J.R.; Stamatatos, L.; Polonis, V.R.; Koutsoukos, M.; Voss, G.; Goepfert, P.; Gilbert, P.; Greene, K.M.; et al. Human Immunodeficiency Virus Type 1 Env Clones from Acute and Early Subtype B Infections for Standardized Assessments of Vaccine-Elicited Neutralizing Antibodies. Journal of Virology 2005, 79, 10108–10125. [Google Scholar] [CrossRef]
- Buzon, V.; Natrajan, G.; Schibli, D.; Campelo, F.; Kozlov, M.M.; Weissenhorn, W. Crystal Structure of HIV-1 Gp41 Including Both Fusion Peptide and Membrane Proximal External Regions. PLoS Pathog 2010, 6, e1000880. [Google Scholar] [CrossRef]
- Kamdem Toukam, D.; Tenbusch, M.; Stang, A.; Temchura, V.; Storcksdieck Genannt Bonsmann, M.; Grewe, B.; Koch, S.; Meyerhans, A.; Nchinda, G.; Kaptue, L.; et al. Targeting Antibody Responses to the Membrane Proximal External Region of the Envelope Glycoprotein of Human Immunodeficiency Virus. PLoS ONE 2012, 7, e38068. [Google Scholar] [CrossRef]
- Shao, S.; Huang, W.-C.; Lin, C.; Hicar, M.D.; LaBranche, C.C.; Montefiori, D.C.; Lovell, J.F. An Engineered Biomimetic MPER Peptide Vaccine Induces Weakly HIV Neutralizing Antibodies in Mice. Ann Biomed Eng 2020, 48, 1991–2001. [Google Scholar] [CrossRef]
- Pancera, M.; Zhou, T.; Druz, A.; Georgiev, I.S.; Soto, C.; Gorman, J.; Huang, J.; Acharya, P.; Chuang, G.-Y.; Ofek, G.; et al. Structure and Immune Recognition of Trimeric Pre-Fusion HIV-1 Env. Nature 2014, 514, 455–461. [Google Scholar] [CrossRef]
- Reardon, P.N.; Sage, H.; Dennison, S.M.; Martin, J.W.; Donald, B.R.; Alam, S.M.; Haynes, B.F.; Spicer, L.D. Structure of an HIV-1–Neutralizing Antibody Target, the Lipid-Bound Gp41 Envelope Membrane Proximal Region Trimer. Proc. Natl. Acad. Sci. U.S.A. 2014, 111, 1391–1396. [Google Scholar] [CrossRef]
- Krebs, S.J.; Kwon, Y.D.; Schramm, C.A.; Law, W.H.; Donofrio, G.; Zhou, K.H.; Gift, S.; Dussupt, V.; Georgiev, I.S.; Schätzle, S.; et al. Longitudinal Analysis Reveals Early Development of Three MPER-Directed Neutralizing Antibody Lineages from an HIV-1-Infected Individual. Immunity 2019, 50, 677–691.e13. [Google Scholar] [CrossRef] [PubMed]
- Bachmann, M.F.; Jennings, G.T. Vaccine Delivery: A Matter of Size, Geometry, Kinetics and Molecular Patterns. Nat Rev Immunol 2010, 10, 787–796. [Google Scholar] [CrossRef]
- Ols, S.; Lenart, K.; Arcoverde Cerveira, R.; Miranda, M.C.; Brunette, N.; Kochmann, J.; Corcoran, M.; Skotheim, R.; Philomin, A.; Cagigi, A.; et al. Multivalent Antigen Display on Nanoparticle Immunogens Increases B Cell Clonotype Diversity and Neutralization Breadth to Pneumoviruses. Immunity 2023, 56, 2425–2441.e14. [Google Scholar] [CrossRef]
- Irvine, D.J.; Read, B.J. Shaping Humoral Immunity to Vaccines through Antigen-Displaying Nanoparticles. Current Opinion in Immunology 2020, 65, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Victora, G.D.; Nussenzweig, M.C. Germinal Centers. Annu. Rev. Immunol. 2012, 30, 429–457. [Google Scholar] [CrossRef] [PubMed]
- Mesin, L.; Ersching, J.; Victora, G.D. Germinal Center B Cell Dynamics. Immunity 2016, 45, 471–482. [Google Scholar] [CrossRef]
- Kelly, H.G.; Tan, H.-X.; Juno, J.A.; Esterbauer, R.; Ju, Y.; Jiang, W.; Wimmer, V.C.; Duckworth, B.C.; Groom, J.R.; Caruso, F.; et al. Self-Assembling Influenza Nanoparticle Vaccines Drive Extended Germinal Center Activity and Memory B Cell Maturation. JCI Insight 2020, 5, e136653. [Google Scholar] [CrossRef] [PubMed]
- Ursin, R.L.; Dhakal, S.; Liu, H.; Jayaraman, S.; Park, H.-S.; Powell, H.R.; Sherer, M.L.; Littlefield, K.E.; Fink, A.L.; Ma, Z.; et al. Greater Breadth of Vaccine-Induced Immunity in Females than Males Is Mediated by Increased Antibody Diversity in Germinal Center B Cells. mBio 2022, 13, e01839-22. [Google Scholar] [CrossRef]
- de Taeye, S.W.; Ozorowski, G.; Torrents de la Peña, A.; Guttman, M.; Julien, J.-P.; van den Kerkhof, T.L.G.M.; Burger, J.A.; Pritchard, L.K.; Pugach, P.; Yasmeen, A.; et al. Immunogenicity of Stabilized HIV-1 Envelope Trimers with Reduced Exposure of Non-Neutralizing Epitopes. Cell 2015, 163, 1702–1715. [Google Scholar] [CrossRef]
- Sok, D.; Burton, D.R. Recent Progress in Broadly Neutralizing Antibodies to HIV. Nat Immunol 2018, 19, 1179–1188. [Google Scholar] [CrossRef]
- Eisen, H.N.; Siskind, G.W. Variations in Affinities of Antibodies during the Immune Response*. Biochemistry 1964, 3, 996–1008. [Google Scholar] [CrossRef] [PubMed]
- Krammer, F.; Palese, P. Advances in the Development of Influenza Virus Vaccines. Nat Rev Drug Discov 2015, 14, 167–182. [Google Scholar] [CrossRef]
- Setliff, I.; Shiakolas, A.R.; Pilewski, K.A.; Murji, A.A.; Mapengo, R.E.; Janowska, K.; Richardson, S.; Oosthuysen, C.; Raju, N.; Ronsard, L.; et al. High-Throughput Mapping of B Cell Receptor Sequences to Antigen Specificity. Cell 2019, 179, 1636–1646.e15. [Google Scholar] [CrossRef]
- Kong, R.; Duan, H.; Sheng, Z.; Xu, K.; Acharya, P.; Chen, X.; Cheng, C.; Dingens, A.S.; Gorman, J.; Sastry, M.; et al. Antibody Lineages with Vaccine-Induced Antigen-Binding Hotspots Develop Broad HIV Neutralization. Cell 2019, 178, 567–584.e19. [Google Scholar] [CrossRef] [PubMed]
- Conti, S.; Kaczorowski, K.J.; Song, G.; Porter, K.; Andrabi, R.; Burton, D.R.; Chakraborty, A.K.; Karplus, M. Design of Immunogens to Elicit Broadly Neutralizing Antibodies against HIV Targeting the CD4 Binding Site. Proc Natl Acad Sci U S A 2021, 118, e2018338118. [Google Scholar] [CrossRef] [PubMed]
- Rojas, E.R.; Billings, G.; Odermatt, P.D.; Auer, G.K.; Zhu, L.; Miguel, A.; Chang, F.; Weibel, D.B.; Theriot, J.A.; Huang, K.C. The Outer Membrane Is an Essential Load-Bearing Element in Gram-Negative Bacteria. Nature 2018, 559, 617–621. [Google Scholar] [CrossRef]
- Nicolay, T.; Vanderleyden, J.; Spaepen, S. Autotransporter-Based Cell Surface Display in Gram-Negative Bacteria. Crit Rev Microbiol 2015, 41, 109–123. [Google Scholar] [CrossRef]
- Pugach, P.; Ozorowski, G.; Cupo, A.; Ringe, R.; Yasmeen, A.; De Val, N.; Derking, R.; Kim, H.J.; Korzun, J.; Golabek, M.; et al. A Native-Like SOSIP.664 Trimer Based on an HIV-1 Subtype B Env Gene. J Virol 2015, 89, 3380–3395. [Google Scholar] [CrossRef]
- Gray, E.S.; Madiga, M.C.; Moore, P.L.; Mlisana, K.; Karim, S.S.; Binley, J.M.; Shaw, G.M.; Mascola, J.R.; Morris, L. Broad HIV-1 Neutralization Mediated by Plasma Antibodies against the Gp41 Membrane Proximal External Region. J Virol 2009. [Google Scholar] [CrossRef]
- Schiffner, T.; Phung, I.; Ray, R.; Irimia, A.; Tian, M.; Swanson, O.; Lee, J.H.; Lee, C.-C.D.; Marina-Zárate, E.; Cho, S.Y.; et al. Vaccination Induces Broadly Neutralizing Antibody Precursors to HIV Gp41. Nat Immunol 2024, 25, 1073–1082. [Google Scholar] [CrossRef] [PubMed]
- Xu, K.; Acharya, P.; Kong, R.; Cheng, C.; Chuang, G.-Y.; Liu, K.; Louder, M.K.; O’Dell, S.; Rawi, R.; Sastry, M.; et al. Epitope-Based Vaccine Design Yields Fusion Peptide-Directed Antibodies That Neutralize Diverse Strains of HIV-1. Nat Med 2018, 24, 857–867. [Google Scholar] [CrossRef] [PubMed]
- Croft, M. The Role of TNF Superfamily Members in T-Cell Function and Diseases. Nat Rev Immunol 2009, 9, 271–285. [Google Scholar] [CrossRef]
- Watts, T.H. TNF/TNFR FAMILY MEMBERS IN COSTIMULATION OF T CELL RESPONSES. Annu. Rev. Immunol. 2005, 23, 23–68. [Google Scholar] [CrossRef]
- Shen, C.H.; DeKosky, B.J.; Guo, Y.; Xu, K.; Gu, Y.; Kilam, D.; Ko, S.H.; Kong, R.; Liu, K.; Louder, M.K.; et al. VRC34-Antibody Lineage Development Reveals How a Required Rare Mutation Shapes the Maturation of a Broad HIV-Neutralizing Lineage. Cell Host Microbe 2020, 27, 531–543 e6. [Google Scholar] [CrossRef]
- Gao, G.; Wieczorek, L.; Peachman, K.K.; Polonis, V.R.; Alving, C.R.; Rao, M.; Rao, V.B. Designing a Soluble near Full-Length HIV-1 Gp41 Trimer. J Biol Chem 2013, 288, 234–246. [Google Scholar] [CrossRef]
- Krebs, S.J.; McBurney, S.P.; Kovarik, D.N.; Waddell, C.D.; Jaworski, J.P.; Sutton, W.F.; Gomes, M.M.; Trovato, M.; Waagmeester, G.; Barnett, S.J.; et al. Multimeric Scaffolds Displaying the HIV-1 Envelope MPER Induce MPER-Specific Antibodies and Cross-Neutralizing Antibodies When Co-Immunized with Gp160 DNA. PLoS ONE 2014, 9, e113463. [Google Scholar] [CrossRef]
- Meuskens, I.; Saragliadis, A.; Leo, J.C.; Linke, D. Type V Secretion Systems: An Overview of Passenger Domain Functions. Front. Microbiol. 2019, 10, 1163. [Google Scholar] [CrossRef]
- van Ulsen, P.; Zinner, K.M.; Jong, W.S.P.; Luirink, J. On Display: Autotransporter Secretion and Application. FEMS Microbiol Letters 2018, 365. [Google Scholar] [CrossRef] [PubMed]
| Component | Name | Sequence | Abbreviation | Function | Ref |
| Motif | DADADA Box | DADADA | 3DA | Increases expression of autotransporter-expressed recombinant passenger proteins | [57] |
| Immunomodulators | Pan DR Epitope non-cognate T-cell antigen | AKFVAAWTLKAAA | PADRE | Enhances the adaptive immune response by providing broad T-helper cell epitope coverage, thereby improving T-cell activation and memory | [57,58] |
| Flagellin from Salmonella enterica (short fragment) | GALNEINNNLQRVRELAVQSANSTNSQSDLDSIQAEITQ | FLIC | TLR5 agonist | [59,60,61,62,65] | |
| Grp A Strep Recombinant Surface Immunogenic Protein | CDTTWTARTVSEVKADLVKQDNK | rSIP | TLR2 and 4 agonists | [63] | |
| OX40L peptide | SSSPAKDPPIQRLRGAVTRCEDGQLFISSYKNEYQTMEVQNNSVVIKCDGLYIIYLKGSFFQEVKIDLHFREDHNPISIPMLNDGRRIVFTVVASLAFKDKVYLTVNAPDTLCEHLQINDGELIVVQLTPGYCAPEGSYHSTVNQVPL | OX40L | T-cell co-stimulation | [66] | |
| Antigen | 3AGJ | KPHMNLVVIGHVDHGKSTLVGHLLARLGYIEWFKLTNLLWQARARGKGSFGFAWILDKMKEERERGITIDLTFMKFETKKYVFTIIDAPGHRDFVKNMITGASQADAAILVVSARKGEFEAGMSTEGQTREHLLLARTMGIEQIIVAVNKMDAPDVNYDQKRYEFVVSVLKKFMKGLGYQVDKIPFIPVSAWKGDNLIERSPNMPWYNGPTLVEALDQLQPPAK | 3AGJ | HIV MPER target | [11] |
| Linkers | GSx4 | GSGSGSGS | L1 | Shorter linker with non-charged, hydrophilic amino acids | [69] |
| AQQASSS×3 | AQQASSSAQQASSSAQQASSS | L2 | Semi-flexible alpha helical linker between scaffold repeats | [68] | |
| (EAAAK)4–L–(EAAAK)4 | AEAAAKEAAAKEAAAKEAAAKALEAEAAAKEAAAKEAAAKEAAAKA | L3 | Rigid linker separating immunomodulator from antigen | [67] |
| Vaccine | Composition (N→C) |
| V1 | 3DA–3AGJ |
| V2 | 3DA–PADRE–GS×4–3AGJ |
| V3 | 3DA–PADRE–GS×4–FLIC–GS×4–3AGJ |
| V4 | 3DA–PADRE–GS×4–rSIP–GS×4–short FLIC–(EAAAK)4–L–(EAAAK)4–3AGJ–AQQASSS×3–3AGJ |
| V5 | 3DA–PADRE–GS×4–rSIP–GS×4–short FLIC–GS×4–OX40L–(EAAAK)4–L–(EAAAK)4–3AGJ–AQQASSS×3–3AGJ |
| Vaccine | Animal | AUC(ELISA) Week 0 | AUC(ELISA) Week 3 | AUC(ELISA) Week 6 | AUC(ELISA) Week 9 | AUC(ELISA) Week 12 | AUC(ELISA) Week 15 | exposure_score | AUC(FLOW) |
| V1 | F1 | 0.01 | 0.01 | 0.01 | 0.01 | 0.02 | 0.03 | −0.62 | 41.27 |
| F2 | 0.01 | 0.02 | 0.04 | 0.02 | 0.03 | 0.03 | |||
| F3 | 0.02 | 0.02 | 0.02 | 0.02 | 0.03 | 0.03 | |||
| M1 | 0.01 | 0.02 | 0.04 | 0.03 | 0.02 | 0.02 | |||
| M2 | 0.01 | 0.02 | 0.02 | 0.03 | 0.06 | 0.03 | |||
| V2 | F1 | 0.01 | 0.02 | 0.02 | 0.03 | 0.05 | 0.05 | −0.67 | 32.23 |
| F2 | 0.01 | 0.04 | 0.03 | 0.02 | 0.05 | 0.03 | |||
| F3 | 0.01 | 0.03 | 0.03 | 0.05 | 0.05 | 0.04 | |||
| M1 | 0.01 | 0.03 | 0.02 | 0.02 | 0.03 | 0.02 | |||
| M2 | 0.01 | 0.03 | 0.02 | 0.03 | 0.04 | 0.03 | |||
| V3 | F1 | 0.02 | 0.04 | 0.21 | 0.15 | 0.04 | 0.04 | −0.65 | 37.39 |
| F2 | 0.01 | 0.02 | 0.03 | 0.02 | 0.03 | 0.04 | |||
| F3 | 0.01 | 0.02 | 0.02 | 0.02 | 0.04 | 0.03 | |||
| M1 | 0.01 | 0.03 | 0.03 | 0.02 | 0.04 | 0.03 | |||
| M2 | 0.01 | 0.01 | 0.02 | 0.03 | 0.02 | 0.02 | |||
| V4 | F2 | 0.01 | 0.02 | 0.02 | 0.04 | 0.04 | 0.03 | −0.49 | 54.14 |
| F3 | 0.01 | 0.03 | 0.03 | 0.04 | 0.14 | 0.13 | |||
| M1 | 0.01 | 0.05 | 0.03 | 0.09 | 0.03 | 0.05 | |||
| M2 | 0.01 | 0.05 | 0.03 | 0.03 | 0.10 | 0.05 | |||
| V5 | F1 | 0.01 | 0.05 | 0.04 | 0.06 | 0.06 | 0.08 | −0.50 | 48.46 |
| F2 | 0.01 | 0.09 | 0.05 | 0.04 | 0.05 | 0.07 | |||
| F3 | 0.01 | 0.04 | 0.25 | 0.07 | 0.05 | 0.08 | |||
| M1 | 0.01 | 0.04 | 0.02 | 0.03 | 0.03 | 0.04 | |||
| M2 | 0.01 | 0.04 | 0.03 | 0.03 | 0.02 | 0.04 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




