Submitted:
13 January 2026
Posted:
14 January 2026
You are already at the latest version
Abstract
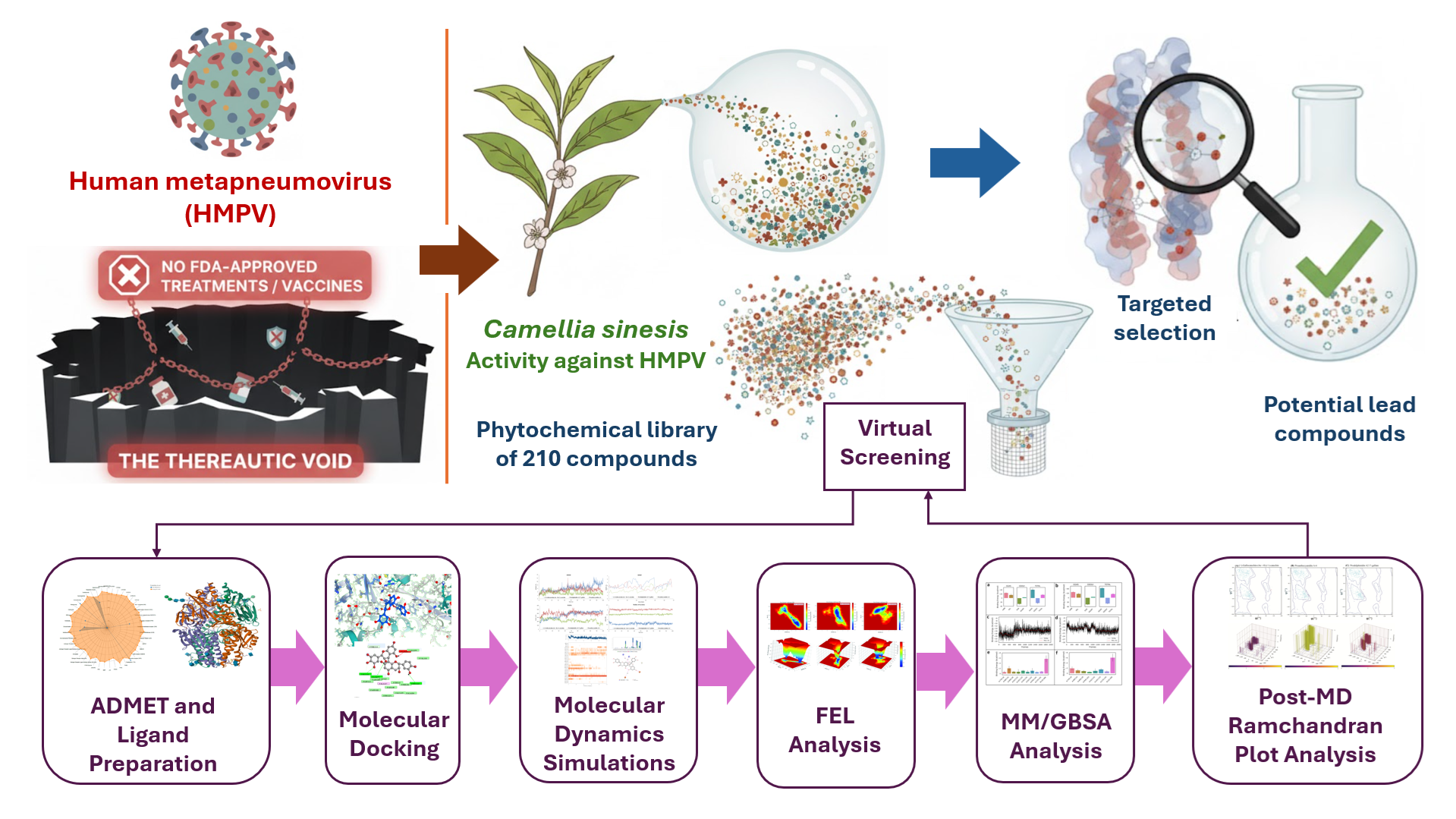
Keywords:
1. Introduction
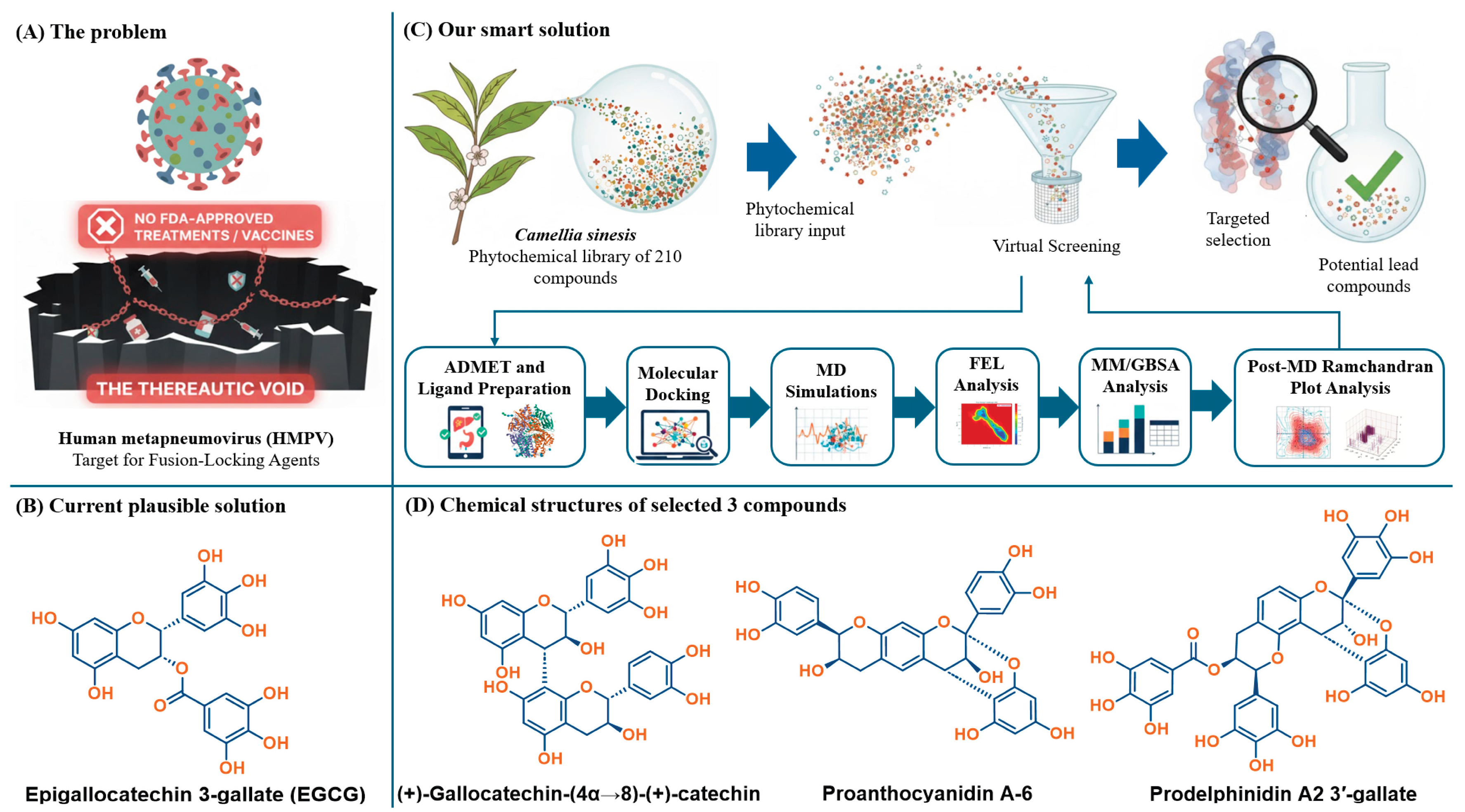
2. Results and Discussion
2.1. Theoretical Binding Stabilization and Multi-Criteria ADMET Filtration of Phytochemicals
| MW | Vol | TPSA | logS | logD | logP | MDCK | PAMPA | hia | hERG | Ames | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| (+)-gallocatechin-(4α→8)-(+)-catechin | 594.140 | 558.733 | 240.990 | -2.898 | 1.465 | 1.010 | -4.883 | 0.670 | 0.009 | 0.158 | 0.259 |
| Prodelphinidin A2 3-gallate | 760.130 | 696.097 | 316.980 | -4.153 | 0.421 | 0.608 | -4.910 | 0.941 | 0.000 | 0.069 | 0.740 |
| Proanthocyanidin A-6 | 576.130 | 541.386 | 209.760 | -3.238 | 1.650 | 1.468 | -4.875 | 0.546 | 0.000 | 0.110 | 0.712 |
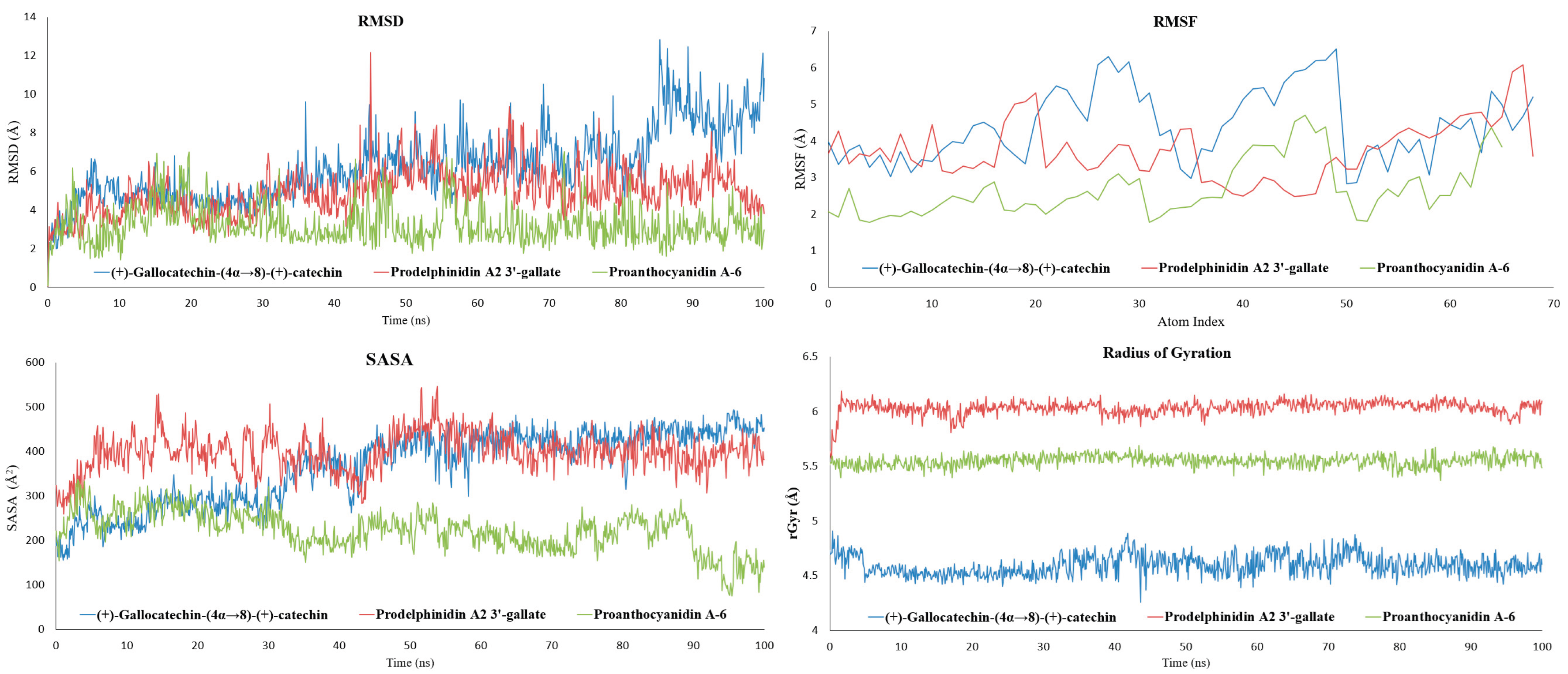
2.2. Molecular Dynamics Simulation and Trajectory Analysis
2.2.1. Proanthocyanidin A-6 as the Most Stabilizing Complex
2.2.2. Stable Anchoring with Flexible Sampling of Prodelphinidin A2 3′-Gallate
2.2.3. Adaptability to Loop Motion of (+)-Gallocatechin-(4α→8)-(+)-Catechin
2.2.4. Ligand Atomic Fluctuations (L-RMSF)
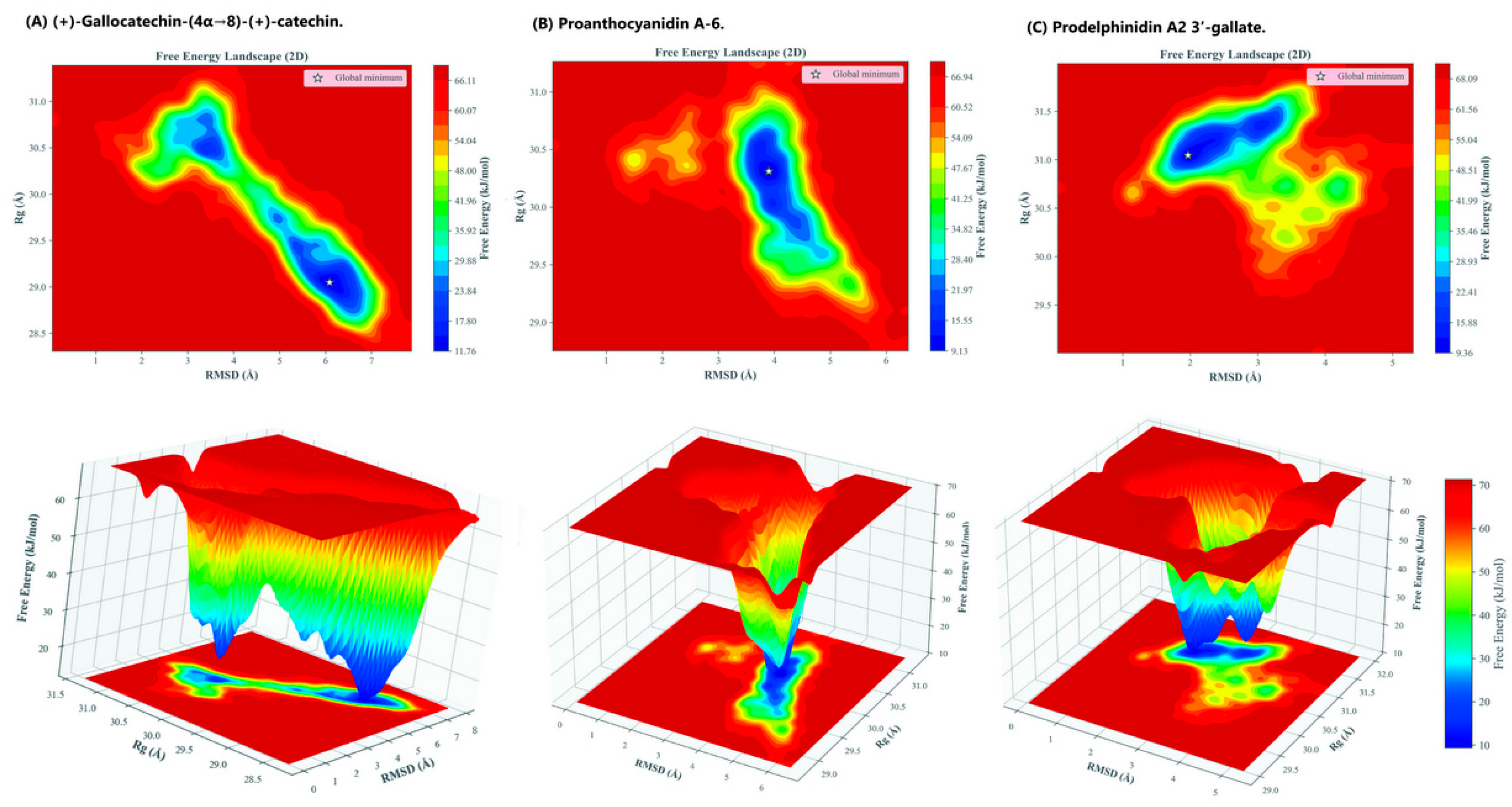
2.3. Free Energy Landscape (FEL) Analysis
2.4. Thermodynamic Stability Analysis (MM/GBSA and MM/PBSA)
2.4.1. Electrostatic and van der Waals Contributions:
2.4.2. Solvation Penalty Management:
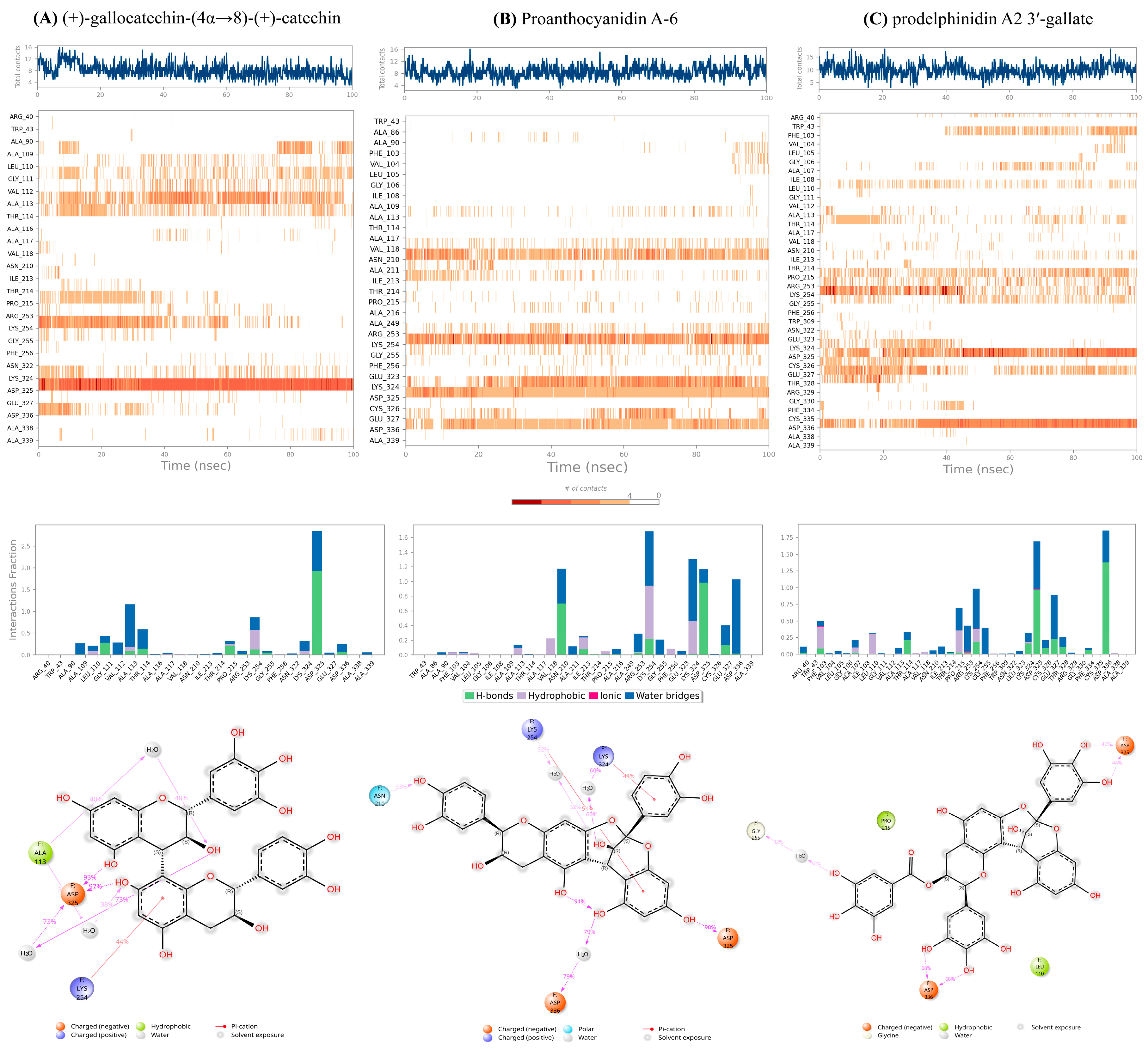
2.5. Mechanistic and Structure-Dynamics Implications (Fusion-Locking)
2.5.1. Targeted Anchoring:
2.5.2. Restricted Flexibility:
2.5.3. Structural Chemotype Influence:
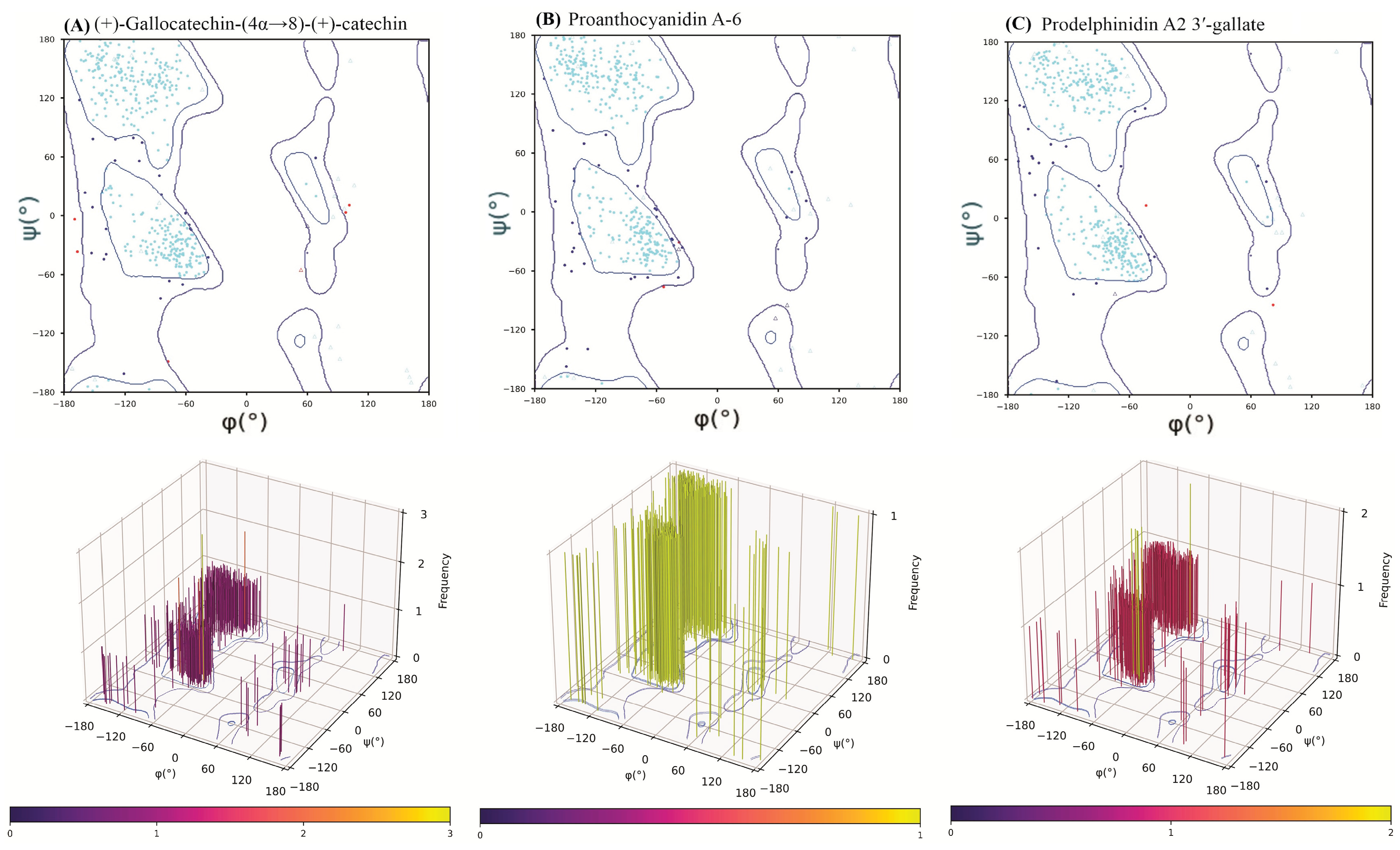
2.6. Post-MD Ramachandran Plot Analysis
3. Materials and Methods
3.1. Target and Ligand Preparation
3.2. Docking-Based Virtual Screening and ADMET Filtration
3.3. Molecular Dynamics (MD) Simulation Setup
3.4. Free Energy Landscape (FEL) Analysis
3.5. Binding Free Energy Calculations (MM/GBSA and MM/PBSA)
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mohammadi, K.; Faramarzi, S.; Yaribash, S.; Valizadeh, Z.; Rajabi, E.; Ghavam, M.; Samiee, R.; Karim, B.; Salehi, M.; Seifi, A.; Shafaati, M. Human metapneumovirus (hMPV) in 2025: emerging trends and insights from community and hospital-based respiratory panel analyses—a comprehensive review. Virol. J. 2025, 22, 150. [Google Scholar] [CrossRef]
- van den Hoogen, B. G.; de Jong, J. C.; Groen, J.; Kuiken, T.; de Groot, R.; Fouchier, R. A.; Osterhaus, A. D. A. A newly discovered human pneumovirus isolated from young children with respiratory tract disease. Nat. Med. 2001, 7, 719–724. [Google Scholar] [CrossRef]
- Principi, N.; Fainardi, V.; Esposito, S. Human Metapneumovirus: A Narrative Review on Emerging Strategies for Prevention and Treatment. Viruses 2025, 17, 1140. [Google Scholar] [CrossRef]
- Twumasi, S.; Ansah, R. O.; Ayirebi, A. A.; Antonio, D. N. M.; Asafoakaa, Y. A.; Tawiah, E.; Opoku, A. Human Metapneumovirus: A Review of Its Epidemiology, Clinical Features, Public Health Implications and Treatment Options. Rev. Med. Virol. 2025, 35, e70043. [Google Scholar] [CrossRef]
- Branche, A. R.; Edwards, K. M. Review: Knowledge Gained and Gaps in Understanding in the 25 Years Since Human Metapneumovirus Was First Identified as a Cause of Human Disease. J. Infect. Dis. 2025, 232, S1–S9. [Google Scholar] [CrossRef]
- Krüger, N.; Laufer, S. A.; Pillaiyar, T. An overview of progress in human metapneumovirus (hMPV) research: Structure, function, and therapeutic opportunities. Drug Discovery Today 2025, 30, 104364. [Google Scholar] [CrossRef]
- Battles, M. B.; Más, V.; Olmedillas, E.; Cano, O.; Vázquez, M.; Rodríguez, L.; Melero, J. A.; McLellan, J. S. Structure and immunogenicity of pre-fusion-stabilized human metapneumovirus F glycoprotein. Nat. Commun. 2017, 8, 1528. [Google Scholar] [CrossRef]
- Chang, A.; Masante, C.; Buchholz, U. J.; Dutch, R. E. Human metapneumovirus (HMPV) binding and infection are mediated by interactions between the HMPV fusion protein and heparan sulfate. J. Virol. 2012, 86, 3230–3243. [Google Scholar] [CrossRef] [PubMed]
- Atanasov, A. G.; Zotchev, S. B.; Dirsch, V. M.; Supuran, C. T.; the International Natural Product Sciences Taskforce. Natural products in drug discovery: advances and opportunities. Nat. Rev. Drug Discov. 2021, 20, 200–216. [Google Scholar] [CrossRef]
- Butler, M. S. The Role of Natural Product Chemistry in Drug Discovery. J. Nat. Prod. 2004, 67, 2141–2153. [Google Scholar] [CrossRef] [PubMed]
- Dzobo, K. The Role of Natural Products as Sources of Therapeutic Agents for Innovative Drug Discovery. Compr. Pharmacol. 2022, 408–422. [Google Scholar] [CrossRef]
- Li, Z.; Chen, K.; Rose, P.; Zhu, Y. Z. Natural products in drug discovery and development: Synthesis and medicinal perspective of leonurine. Front. Chem. 2022, 10, 1036329. [Google Scholar] [CrossRef] [PubMed]
- Zhao, T.; Li, C.; Wang, S.; Song, X. Green Tea (Camellia sinensis): A Review of Its Phytochemistry, Pharmacology, and Toxicology. Molecules 2022, 27, 3909. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y. Q.; Li, Q. S.; Zheng, X. Q.; Lu, J. L.; Liang, Y. R. Antiviral Effects of Green Tea EGCG and Its Potential Application against COVID-19. Molecules 2021, 26, 3962. [Google Scholar] [CrossRef]
- Zhang, Z.; Hao, M.; Zhang, X.; He, Y.; Chen, X.; Taylor, E. W.; Zhang, J. Potential of green tea EGCG in neutralizing SARS-CoV-2 Omicron variant with greater tropism toward the upper respiratory tract. Trends Food Sci. Technol. 2023, 132, 40–53. [Google Scholar] [CrossRef]
- Mirmojarabian, S.; Karimi, A.; Lorigooini, Z.; Javadi-Farsani, F.; Soltani, A.; Moradi, M.-T. Phytochemical properties and antiviral effect of green tea (Camellia sinensis) extract on adenovirus in vitro. J. Shahrekord Univ. Med. Sci. 2022, 24, 104–110. [Google Scholar] [CrossRef]
- Karimi, A.; Asadi-Samani, M.; Altememy, D.; Moradi, M.-T. Anti-influenza and anti-inflammatory effects of green tea (Camellia sinensis L.) extract. Future Nat. Prod. 2022, 8, 59–64. [Google Scholar] [CrossRef]
- Rowe, C. A.; Nantz, M. P.; Bukowski, J. F.; Percival, S. S. Specific formulation of Camellia sinensis prevents cold and flu symptoms and enhances gamma,delta T cell function: a randomized, double-blind, placebo-controlled study. J. Am. Coll. Nutr. 2007, 26, 445–452. [Google Scholar] [CrossRef]
- Kharisma, V. D.; Kharisma, S. D.; Ansori, A. N. M.; Kurniawan, H. P.; Witaningrum, A. M.; Fadholly, A.; Tacharina, M. R. Antiretroviral effect simulation from black tea (Camellia sinensis) via dual inhibitors mechanism in HIV-1 and its social perspective in Indonesia. Res. J. Pharm. Technol. 2021, 14, 455–460. [Google Scholar] [CrossRef]
- Chaudhary, P.; Mitra, D.; Das Mohapatra, P. K.; Docea, A. O.; Mon Myo, E.; Janmeda, P.; Martorell, M.; Iriti, M.; Ibrayeva, M.; Sharifi-Rad, J.; Santini, A.; Romano, R.; Calina, D.; Cho, W. C. Camellia sinensis: Insights on its molecular mechanisms of action towards nutraceutical, anticancer potential and other therapeutic applications. Arabian J. Chem. 2023, 16, 104680. [Google Scholar] [CrossRef]
- Dubey, A.; Kumar, M.; Tufail, A.; Dwivedi, V. D.; Ragusa, A. Unlocking antiviral potentials of traditional plants: A multi-method computational study against human metapneumovirus (HMPV). J. Infect. Public Health 2025, 18, 102885. [Google Scholar] [CrossRef]
- Conceição, P. J. P.; Ayusso, G. M.; Carvalho, T.; Duarte Lima, M. L.; Marinho, M. d. S.; Moraes, F. R.; Galán-Jurado, P. E.; González-Santamaría, J.; Bittar, C.; Zhang, B.; Jardim, A. C. G.; Rahal, P.; Calmon, M. F. In Vitro Evaluation of the Antiviral Activity of Polyphenol (−)-Epigallocatechin-3-Gallate (EGCG) Against Mayaro Virus. Viruses 2025, 17, 258. [Google Scholar] [CrossRef]
- Meshram, V. D.; Balaji, R.; Saravanan, P.; Subbamanda, Y.; Deeksha, W.; Bajpai, A.; Joshi, H.; Bhargava, A.; Patel, B. K. Computational Insights Into the Mechanism of EGCG's Binding and Inhibition of the TDP-43 Aggregation. Chem. Biol. Drug Des. 2024, 104, e14640. [Google Scholar] [CrossRef] [PubMed]
- Vivek-Ananth, R. P.; Mohanraj, K.; Sahoo, A. K.; Samal, A. IMPPAT 2.0: An Enhanced and Expanded Phytochemical Atlas of Indian Medicinal Plants. ACS Omega 2023, 8, 8827–8845. [Google Scholar] [CrossRef] [PubMed]
- Mohanraj, K.; Karthikeyan, B. S.; Vivek-Ananth, R. P.; Bharath Chand, R. P.; Aparna, S. R.; Mangalapandi, P.; Samal, A. IMPPAT: A curated database of Indian Medicinal Plants, Phytochemistry And Therapeutics. Sci. Rep. 2018, 8, 4329. [Google Scholar] [CrossRef]
- Liu, Y.; Yang, X.; Gan, J.; Chen, S.; Xiao, Z. X.; Cao, Y. CB-Dock2: improved protein–ligand blind docking by integrating cavity detection, docking and homologous template fitting. Nucleic Acids Res. 2022, 50, W159–W164. [Google Scholar] [CrossRef]
- Banerjee, P.; Kemmler, E.; Dunkel, M.; Preissner, R. ProTox 3.0: a webserver for the prediction of toxicity of chemicals. Nucleic Acids Res. 2024, 52, W513–W520. [Google Scholar] [CrossRef]
- Bowers, K. J.; Chow, D. E.; Xu, H.; Dror, R. O.; Eastwood, M. P.; Gregersen, B. A.; Klepeis, J. L.; Kolossvary, I.; Moraes, M. A.; Sacerdoti, F. D.; et al. Scalable Algorithms for Molecular Dynamics Simulations on Commodity Clusters. In SC '06: Proceedings of the 2006 ACM/IEEE Conference on Supercomputing; IEEE, 2006; pp. 43–43. [Google Scholar] [CrossRef]
- Roos, K.; Wu, C.; Damm, W.; Reboul, M.; Stevenson, J. M.; Lu, C.; Dahlgren, M. K.; Mondal, S.; Chen, W.; Wang, L.; Abel, R.; Friesner, R. A.; Harder, E. D. OPLS3e: Extending Force Field Coverage for Drug-Like Small Molecules. J. Chem. Theory Comput. 2019, 15, 1863–1874. [Google Scholar] [CrossRef]
- Farago, O. Langevin thermostat for robust configurational and kinetic sampling. Physica A: Stat. Mech. Its Appl. 2019, 534, 122210. [Google Scholar] [CrossRef]
- Ke, Q.; Gong, X.; Liao, S.; Duan, C.; Li, L. Effects of thermostats/barostats on physical properties of liquids by molecular dynamics simulations. J. Mol. Liq. 2022, 365, 120116. [Google Scholar] [CrossRef]
- Kräutler, V.; van Gunsteren, W. F.; Hünenberger, P. H. A fast SHAKE algorithm to solve distance constraint equations for small molecules in molecular dynamics simulations. Comput. Chem. 2001, 22, 501–508. [Google Scholar] [CrossRef]
- Darden, T.; Perera, L.; Li, L.; Pedersen, L. New tricks for modelers from the crystallography toolkit: the particle mesh Ewald algorithm and its use in nucleic acid simulations. Structure 1999, 7, R55–R60. [Google Scholar] [CrossRef] [PubMed]
- Prada-Gracia, D.; Gómez-Gardeñes, J.; Echenique, P.; Falo, F. Exploring the Free Energy Landscape: From Dynamics to Networks and Back. PLoS Comput. Biol. 2009, 5, e1000415. [Google Scholar] [CrossRef] [PubMed]
- Al-Khafaji, Khattab; Taskin Tok, T. Molecular dynamics simulation, free energy landscape and binding free energy computations in exploration the anti-invasive activity of amygdalin against metastasis. Comput. Methods Programs Biomed. 2020, 195, 105660. [Google Scholar] [CrossRef] [PubMed]
- Roux, B.; Chipot, C. Editorial Guidelines for Computational Studies of Ligand Binding Using MM/PBSA and MM/GBSA Approximations Wisely. J. Phys. Chem. B 2024, 128, 12027–12029. [Google Scholar] [CrossRef] [PubMed]
- Genheden, S.; Ryde, U. The MM/PBSA and MM/GBSA methods to estimate ligand-binding affinities. Expert Opin. Drug Discov. 2015, 10, 449–461. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Bo, Z.; Xu, T.; Xu, B.; Wang, D.; Zheng, H. Uni-GBSA: an open-source and web-based automatic workflow to perform MM/GB(PB)SA calculations for virtual screening. Briefings Bioinform. 2023, 24, bbad218. [Google Scholar] [CrossRef] [PubMed]
- Kumar, M.; Rathore, R. S. RamPlot: a webserver to draw 2D, 3D and assorted Ramachandran (ϕ, ψ) maps. J. Appl. Crystallogr. 2025, 58, 630–636. [Google Scholar] [CrossRef]
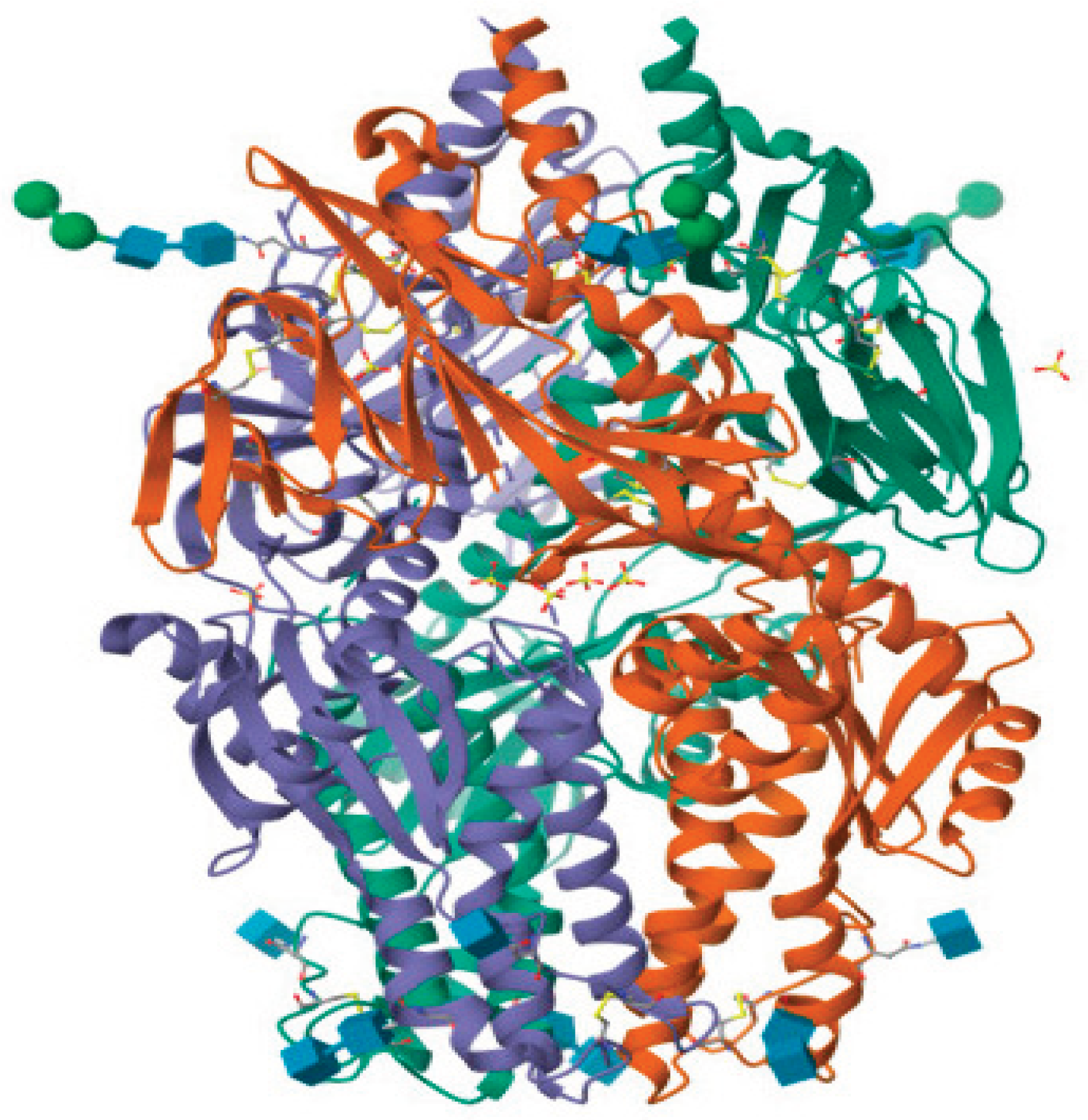
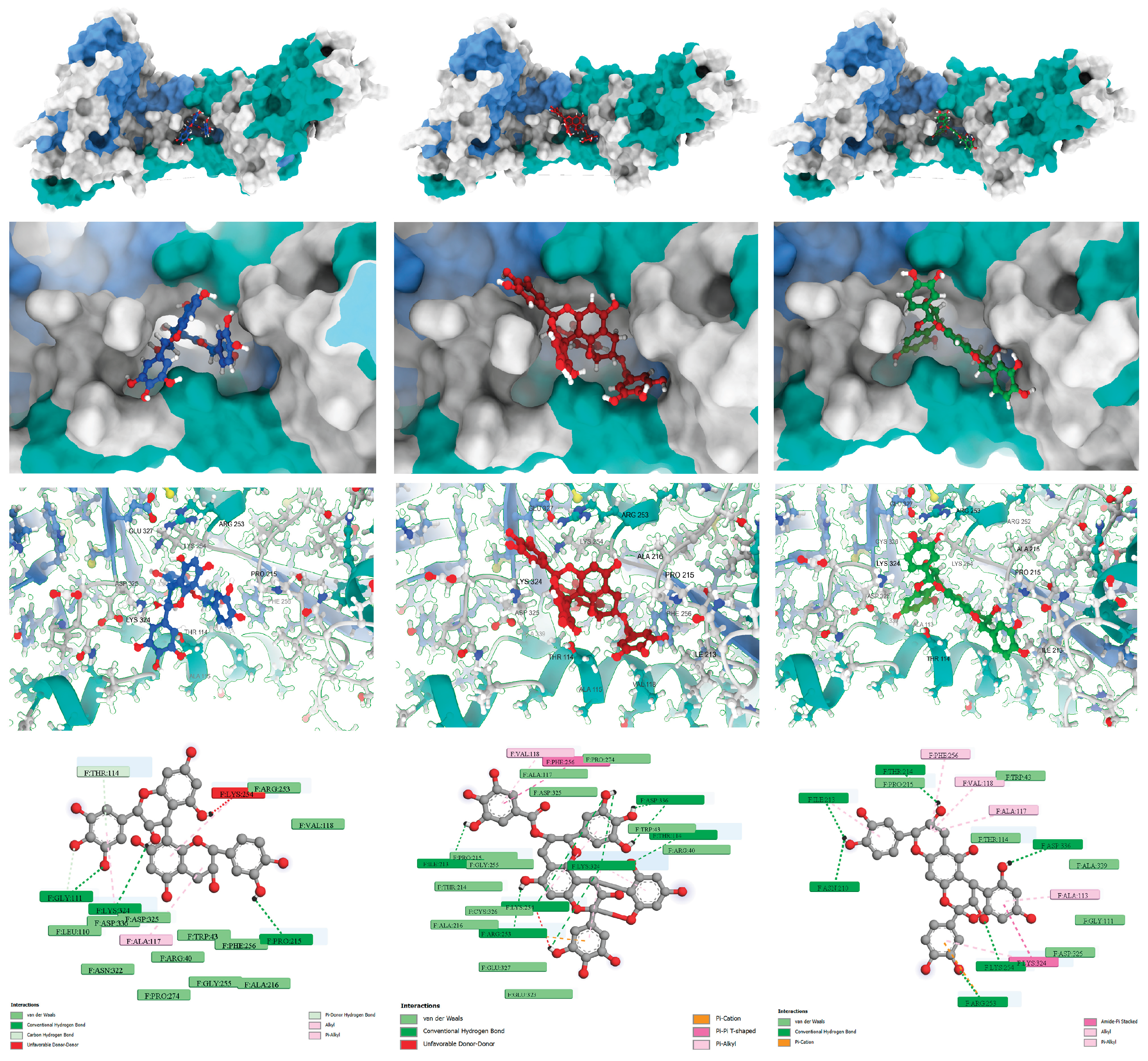
| Compound | MM/GBSA (kcal/mol) | MM/PBSA (kcal/mol) |
| (+)-Gallocatechin-(4α→8)-(+)-catechin | −33.17 | −10.92 |
| Proanthocyanidin A-6 | −25.94 | +18.01 |
| Prodelphinidin A2 3′-gallate | −42.43 | −6.24 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




