Submitted:
09 January 2026
Posted:
12 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Dataset
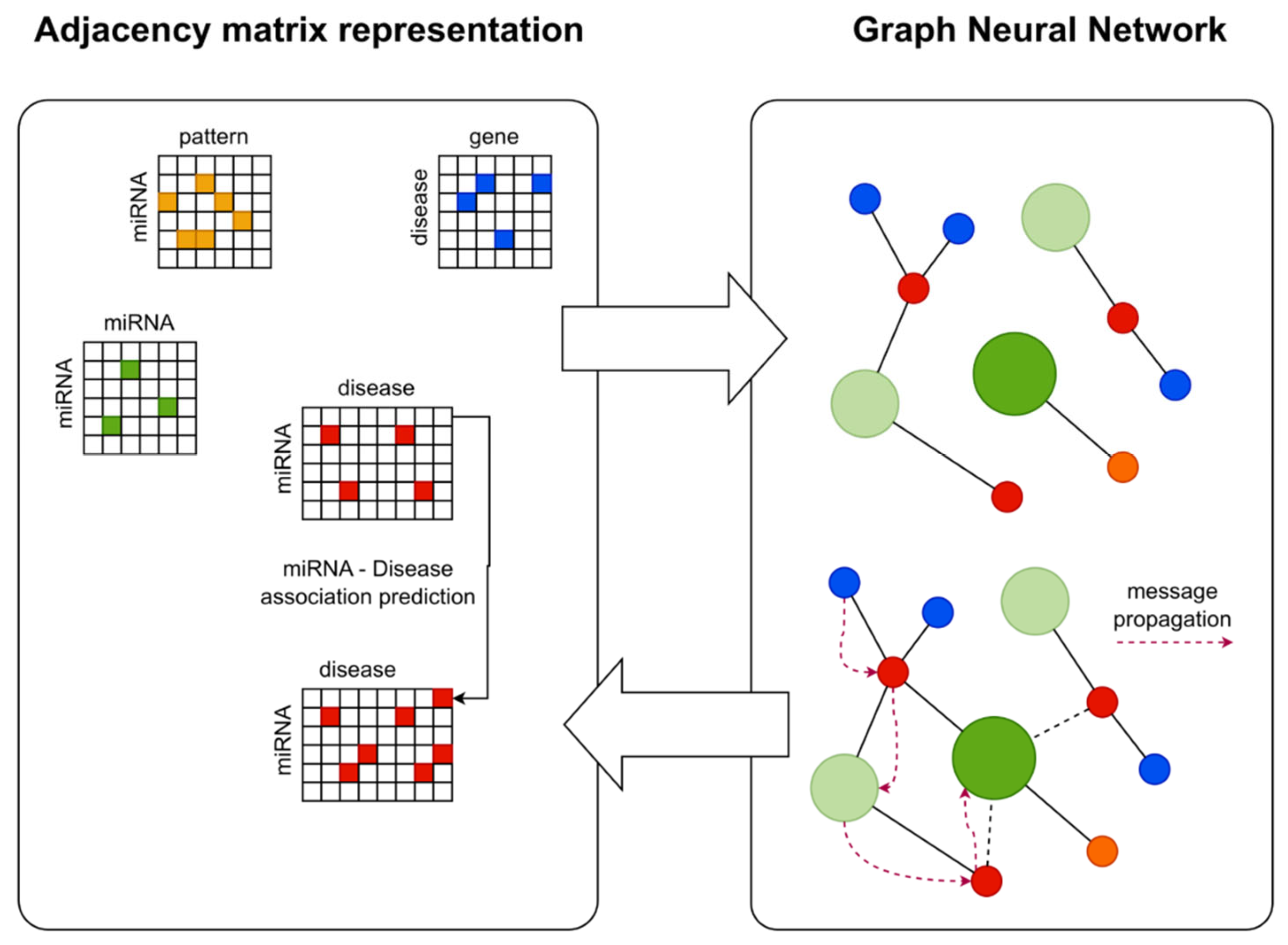
2.2. Graph Neural Network Architecture
2.3. Training and Validation
3. Results
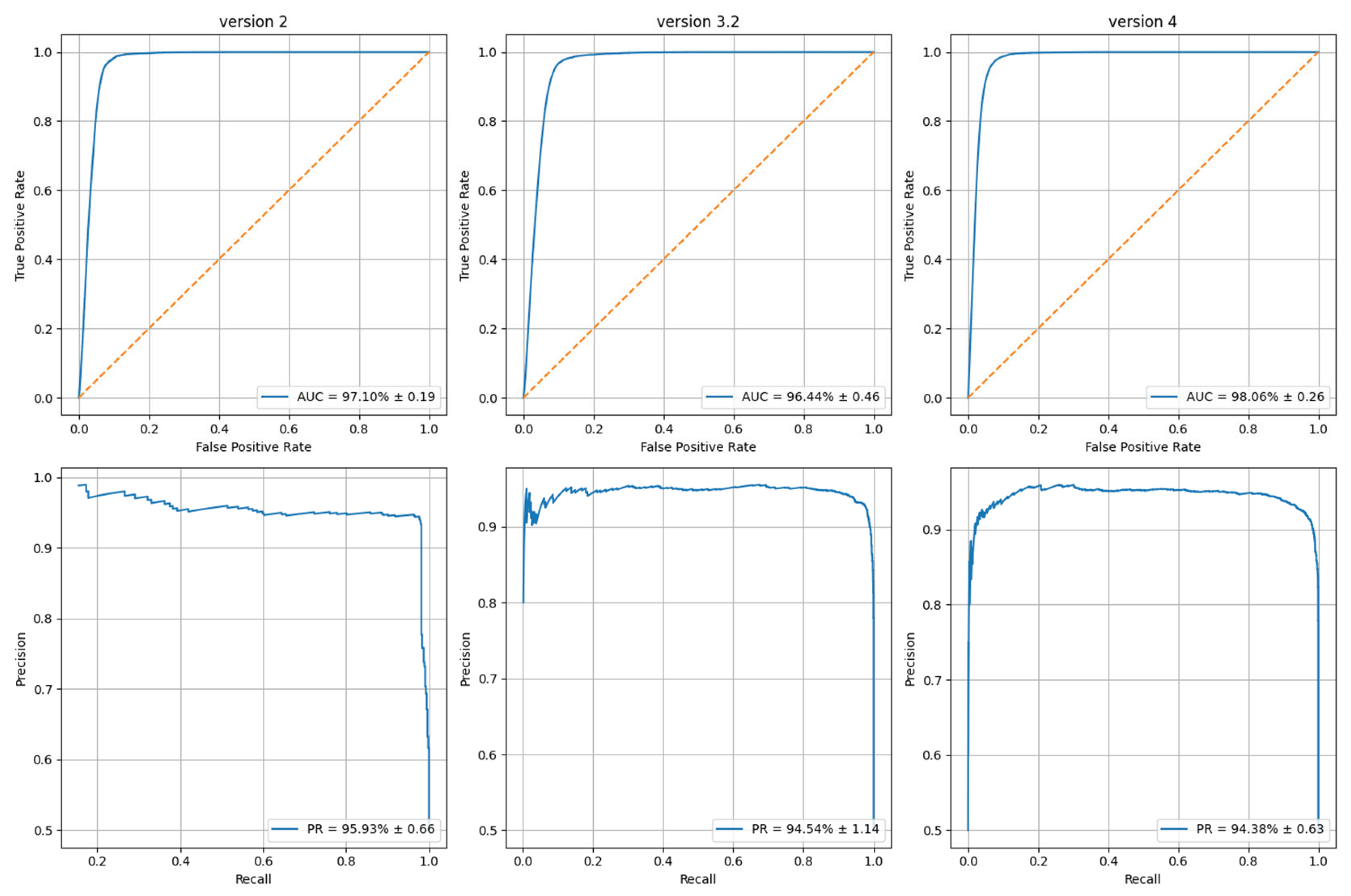
3.1. Evaluation Metrics
3.2. Comparison with Existing Methods
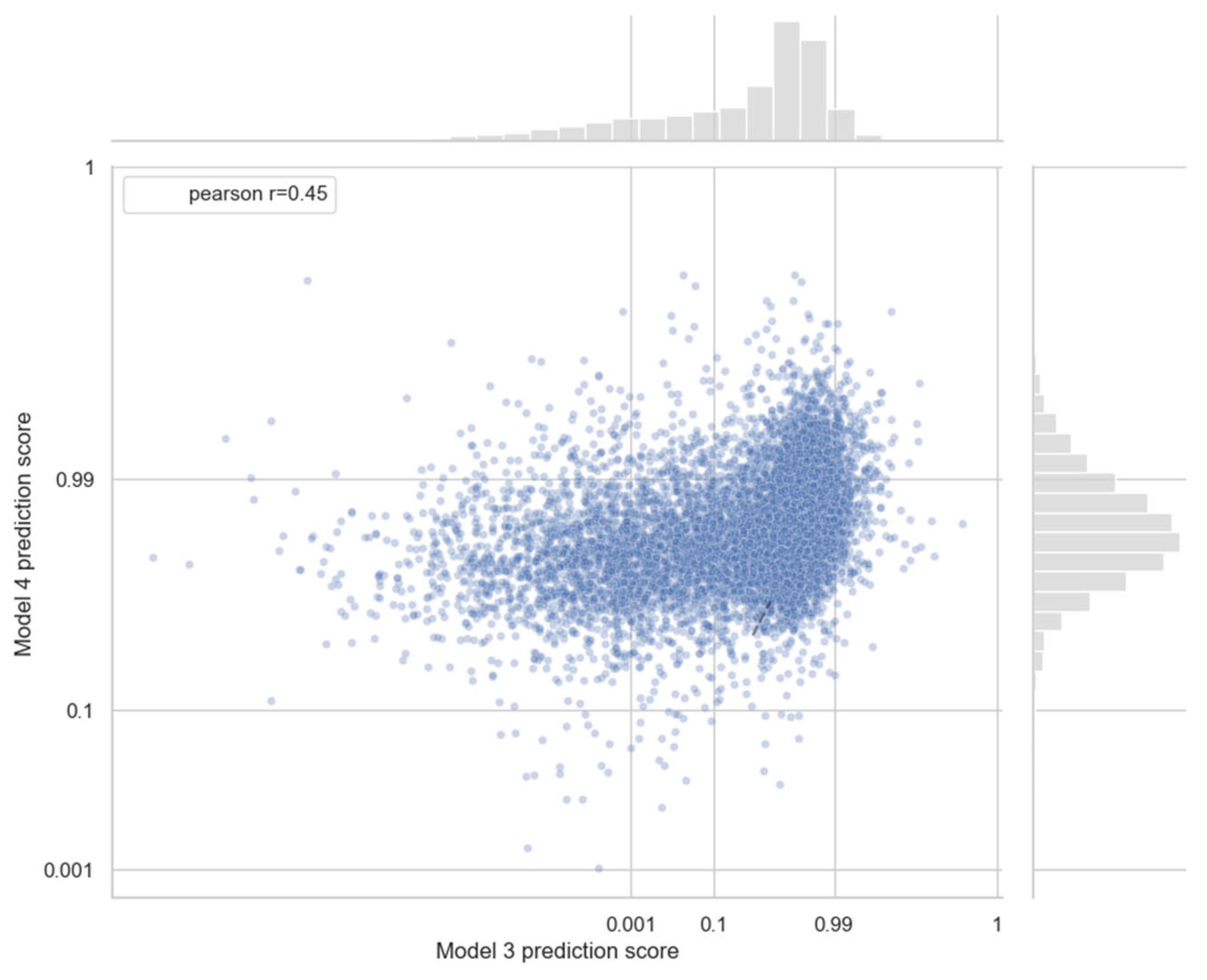
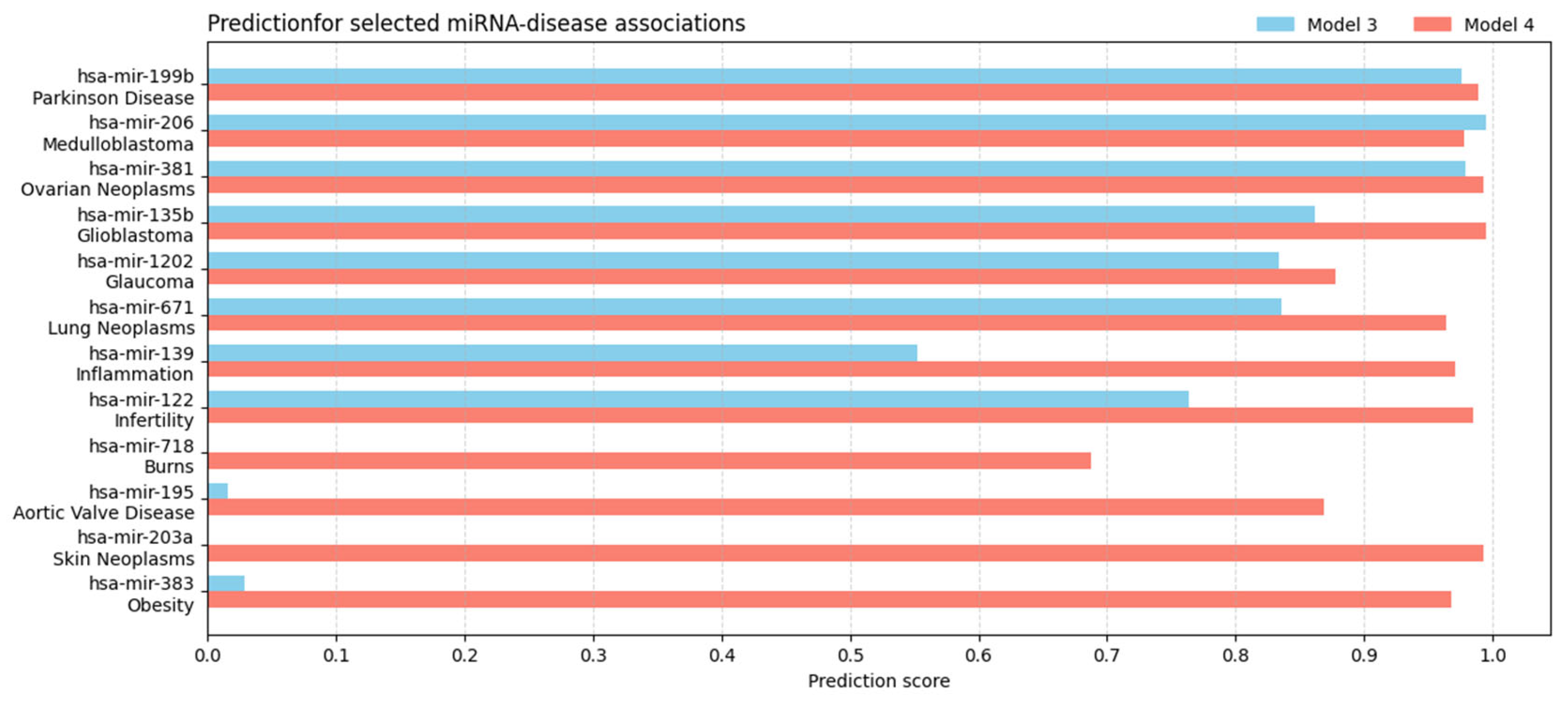
3.3. Analysis of Newly Predicted Associations
3.4. Ablation Analysis
4. Discussion and Conclusions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ambros, V. The functions of animal microRNAs. Nature 2004, 431, 350–355. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Bartel, DP. Metazoan MicroRNAs. Cell. 2018, 173(1), 20–51. [Google Scholar] [CrossRef]
- Eulalio, A; Huntzinger, E; Izaurralde, E. Getting to the Root of miRNA-Mediated Gene Silencing. Cell. 2008, 132, 9–14. [Google Scholar] [CrossRef] [PubMed]
- Meister, G.; Tuschl, T. Mechanisms of gene silencing by double-stranded RNA. Nature 2004, 431, 343–349. [Google Scholar] [CrossRef]
- Vasudevan, S.; Tong, Y.; Steitz, J.A. Switching from repression to activation: microRNAs can up-regulate translation. Science 2007, 318, 1931–1934. [Google Scholar] [CrossRef]
- de Rooij, LA; Mastebroek, DJ; Ten Voorde, N; van der Wall, E; van Diest, PJ; Moelans, CB. The microRNA lifecycle in health and cancer. Cancers 2022, 14(23), 5748. [Google Scholar] [CrossRef]
- Wightman, B.; Ha, I.; Ruvkun, G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 1993, 75, 855–862. [Google Scholar] [CrossRef] [PubMed]
- Griffiths-Jones, S; Saini, HK; van Dongen, S; Enright, AJ. miRBase: tools for microRNA genomics. Nucleic acids research 2008, 36, D154–158. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Alaimo, S; Giugno, R; Pulvirenti, A. ncPred: ncRNA-Disease Association Prediction through Tripartite Network-Based Inference. Front Bioeng Biotechnol. 2014, 2, 71. [Google Scholar] [CrossRef]
- Calin, GA; Croce, CM. MicroRNA signatures in human cancers. Nature reviews Cancer 2006, 6, 857–866. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Zhou, SS; Jin, JP; Wang, JQ; Zhang, ZG; Freedman, JH; Zheng, Y; Cai, L. miRNAS in cardiovascular diseases: potential biomarkers, therapeutic targets and challenges. Acta Pharmacologica Sinica 2018, 39(7), 1073–84. [Google Scholar] [CrossRef] [PubMed]
- Li, S; Lei, Z; Sun, T. The role of microRNAs in neurodegenerative diseases: a review. Cell biology and toxicology 2023, 39(1), 53–83. [Google Scholar] [CrossRef] [PubMed]
- Rottiers, V; Näär, AM. MicroRNAs in metabolism and metabolic disorders. Nature reviews Molecular cell biology 2012, 13(4), 239–50. [Google Scholar] [CrossRef]
- Liu, Z; Sall, A; Yang, D. MicroRNA: An emerging therapeutic target and intervention tool. International journal of molecular sciences 2008, 9, 978–999. [Google Scholar] [CrossRef] [PubMed Central]
- Ye, JW; Xu, MC; Tian, XK; Cai, S; Zeng, S. Research advances in the detection of miRNA. J Pharm Anal. 2019, 9(4), 217–26. [Google Scholar] [CrossRef]
- Chen, X; Xie, D; Zhao, Q; You, ZH. MicroRNAs and complex diseases: from experimental results to computational models. Briefings in bioinformatics 2019, 20, 515–539. [Google Scholar] [CrossRef] [PubMed]
- Zeng, X; Ding, N; Rodríguezpatón, A; Lin, Z; Ju, Y. Prediction of MicroRNA–disease Associations by Matrix Completion. Current Proteomics 2016, 13, 151–157. [Google Scholar] [CrossRef]
- Li, Y; Qiu, C; Tu, J; Geng, B; Yang, J; et al. HMDD v2.0: a database for experimentally supported human microRNA and disease associations. Nucleic acids Res. 2014, 42, D1070–1074. [Google Scholar] [CrossRef]
- Huang, Z; Shi, JC; Gao, YX; Cui, CM; Zhang, S; Li, JW; Zhou, Y; Cui, QH. HMDD v3.0: a database for experimentally supported human microRNA-disease associations. Nucleic Acids Res. 2019, 47(D1), D1013–7. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z; Ren, F; Liu, CN; He, SM; Sun, G; Gao, QA; Yao, L; Zhang, YD; Miao, RY; Cao, Y; et al. dbDEMC: a database of differentially expressed miRNAs in human cancers. Bmc Genom 2010, 11. [Google Scholar] [CrossRef]
- Jiang, Q; Wang, Y; Hao, Y; Juan, L; Teng, M; et al. miR2Disease: a manually curated database for microRNA deregulation in human disease. Nucleic acids research 2009, 37, D98–104. [Google Scholar] [CrossRef]
- Pasquier, C; Gardes, J. Prediction of miRNA-disease associations with a vector space model. Scientific reports 2016, 6, 27036. [Google Scholar] [CrossRef]
- Bandyopadhyay, S; Mitra, R; Maulik, U; Zhang, MQ. Development of the human cancer microRNA network. Silence 2010, 1, 6. [Google Scholar] [CrossRef]
- Shi, H; Xu, J; Zhang, G; Xu, L; Li, C; et al. Walking the interactome to identify human miRNA-disease associations through the functional link between miRNA targets and disease genes. Bmc Systems Biology 2013, 7, 101. [Google Scholar] [CrossRef] [PubMed Central]
- Mørk, S; Pletscherfrankild, S; Palleja, CA; Gorodkin, J; Jensen, LJ. Protein-driven inference of miRNA-disease associations; Bioinformatics: Oxford, England, 2014; Volume 30, pp. 392–397. [Google Scholar]
- Chen, X; Jiang, ZC; Xie, D; Huang, DS; Zhao, Q; et al. A novel computational model based on super-disease and miRNA for potential miRNA-disease association prediction. Molecular bioSystems 2017, 13, 1202–1212. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Chen, X; Wu, QF; Yan, GY. RKNNMDA: ranking-based KNN for MiRNA-disease association prediction. RNA Biol. 2017, 14(7), 952–62. [Google Scholar] [CrossRef] [PubMed]
- Ha, J. SMAP: Similarity-based matrix factorization framework for inferring miRNA-disease association. Knowl. Based Syst. 2023, 263, 110295. [Google Scholar] [CrossRef]
- Jiang, Q; Hao, Y; Wang, G; Juan, L; Zhang, T; et al. Prioritization of disease microRNAs through a human phenome-microRNAome network. BMC systems biology 2010, 4 Suppl 1, pmid:20522252. [Google Scholar] [CrossRef]
- You, ZH; Huang, ZA; Zhu, Z; Yan, GY; Li, ZW; et al. PBMDA: A novel and effective path-based computational model for miRNA-disease association prediction. PLoS computational biology 2017, 13, pmid:28339468. [Google Scholar] [CrossRef]
- Yu, H; Chen, X; Lu, L. Large-scale prediction of microRNA-disease associations by combinatorial prioritization algorithm. Scientific reports 2017, 7, 43792. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Liu, Q. Generalized matrix factorization based on weighted hypergraph learning for microbe-drug association prediction. Computers in Biology and Medicine 2022, 145, 105503. [Google Scholar] [CrossRef] [PubMed]
- Xuan, P; Han, K; Guo, MZ; Guo, YH; Li, JB; Ding, J; Liu, Y; Dai, QG; Li, J; Teng, ZX; et al. Prediction of microRNAs Associated with Human Diseases Based on Weighted k Most Similar Neighbors. PLoS ONE 2013, 8(8), pmid:23950912. [Google Scholar] [CrossRef]
- Chen, X; Liu, MX; Yan, GY. RWRMDA: predicting novel human microRNA-disease associations. Molecular Biosystems 2012, 8, 2792–2798. [Google Scholar] [CrossRef] [PubMed Central]
- Xuan, P; Han, K; Guo, Y; Li, J; Li, X; et al. Prediction of potential disease-associated microRNAs based on random walk. Bioinformatics (Oxford, England) 2015, 31, 1805–1815. [Google Scholar] [CrossRef] [PubMed Central]
- Chen, X; Yan, CC; Zhang, X; You, ZH; Deng, LX; Liu, Y; Zhang, YD; Dai, QH. WBSMDA: within and between score for MiRNA-disease association prediction. Sci Rep. 2016, 6, 1–9. [Google Scholar] [CrossRef]
- Chen, X; Clarence, YC; Zhang, X; You, ZH; Huang, YA; et al. HGIMDA: Heterogeneous graph inference for miRNA-disease association prediction. Oncotarget 2016, 7, 65257–65269. [Google Scholar] [CrossRef]
- Chen, X; Zhou, Z; Zhao, Y. ELLPMDA: Ensemble learning and link prediction for miRNA-disease association prediction. RNA biology 2018, 15, 807–818. [Google Scholar] [CrossRef] [PubMed]
- Ha, J. Graph Convolutional Network with Neural Collaborative Filtering for Predicting miRNA-Disease Association. Biomedicines 2025, 9, 1152. [Google Scholar] [CrossRef]
- Jin, Z.; Wang, M.; Tang, C.; Zheng, X.; Zhang, W.; Sha, X.; An, S. Predicting miRNA-disease association via graph attention learning and multiplex adaptive modality fusion. Comput. Biol. Med. 2024, 169, 107904. [Google Scholar] [CrossRef]
- Xu, J; Li, CX; Lv, JY; Li, YS; Xiao, Y; et al. Prioritizing Candidate Disease miRNAs by Topological Features in the miRNA Target-Dysregulated Network: Case Study of Prostate Cancer. Molecular Cancer Therapeutics 2011, 10, 1857–1866. [Google Scholar] [PubMed Central]
- Chen, X; Yan, CC; Zhang, X; Li, Z; Deng, L; et al. RBMMMDA: predicting multiple types of disease-microRNA associations. Scientific reports 2015, 5, 13877. [Google Scholar]
- Chen, X; Yan, GY. Semi-supervised learning for potential human microRNA-disease associations inference. Sci Rep. 2014, 4, pmid:24975600. [Google Scholar] [CrossRef]
- Li, JQ; Rong, ZH; Chen, X; Yan, GY; You, ZH. MCMDA: Matrix completion for MiRNA-disease association prediction. Oncotarget 2017, 8, 21187–21199. [Google Scholar] [CrossRef] [PubMed Central]
- Gu, C; Li, X. Prediction of disease-related miRNAs by voting with multiple classifiers. BMC bioinformatics 2023, 24(1), 177. [Google Scholar] [CrossRef]
- Ning, Q; Zhao, YM; Gao, J; Chen, C; Li, X; Li, TT; Yin, MH. AMHMDA: attention aware multi-view similarity networks and hypergraph learning for miRNA-disease associations identification. Brief Bioinform. 2023, 24(2). [Google Scholar]
- Peng, JJ; Hui, WW; Li, QQ; Chen, BL; Hao, JY; Jiang, QH; Shang, XQ; Wei, ZY. A learning-based framework for miRNA-disease association identification using neural networks. Bioinformatics 2019, 35(21), 4364–71. [Google Scholar] [PubMed]
- Yu, L.; Yu, Z.G.; Han, G.S.; Li, J.; Anh, V. Heterogeneous types of miRNA-disease associations stratified by multi-layer network embedding and prediction. Biomedicines 2021, 9, 1152. [Google Scholar] [CrossRef]
- Available online: https://www.disgenet.org/.
- Bodenreider, O. The unified medical language system (UMLS): integrating biomedical terminology. Nucleic acids research 2004, 32 suppl_1, D267–70. [Google Scholar] [CrossRef]
- Needleman, SB; Wunsch, CD. A general method applicable to the search for similarities in the amino acid sequence of two proteins. Journal of molecular biology 1970, 48(3), 443–53. [Google Scholar] [CrossRef] [PubMed]
- Gilmer, J; Schoenholz, SS; Riley, PF; Vinyals, O; Dahl, GE. Neural message passing for quantum chemistry. InInternational conference on machine learning 2017 Jul 17 (pp. 1263-1272). Pmlr.
- Ma, YJ. DeepMNE: deep multi-network embedding for lncRNA-disease association prediction. IEEE J Biomed Health 2022, 26(7), 3539–49. [Google Scholar] [CrossRef]
- Ma, Y; Ma, Y. Hypergraph-based logistic matrix factorization for metabolite–disease interaction prediction. Bioinformatics 2022, 38(2), 435–43. [Google Scholar] [CrossRef] [PubMed]
- Barbato, A; Iuliano, A; Volpe, M; D’Alterio, R; Brillante, S; Massa, F; De Cegli, R; Carrella, S; Salati, M; Russo, A; Russo, G. Integrated genomics identifies miR-181/TFAM pathway as a critical driver of drug resistance in melanoma. International Journal of Molecular Sciences 2021, 22(4), 1801. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y; Xu, W; Yang, Y; Zhang, Z. miRNA-93-5p promotes gemcitabine resistance in pancreatic cancer cells by targeting the PTEN-mediated PI3K/Akt signaling pathway. Annals of Clinical & Laboratory Science 2021, 51(3), 310–20. [Google Scholar]
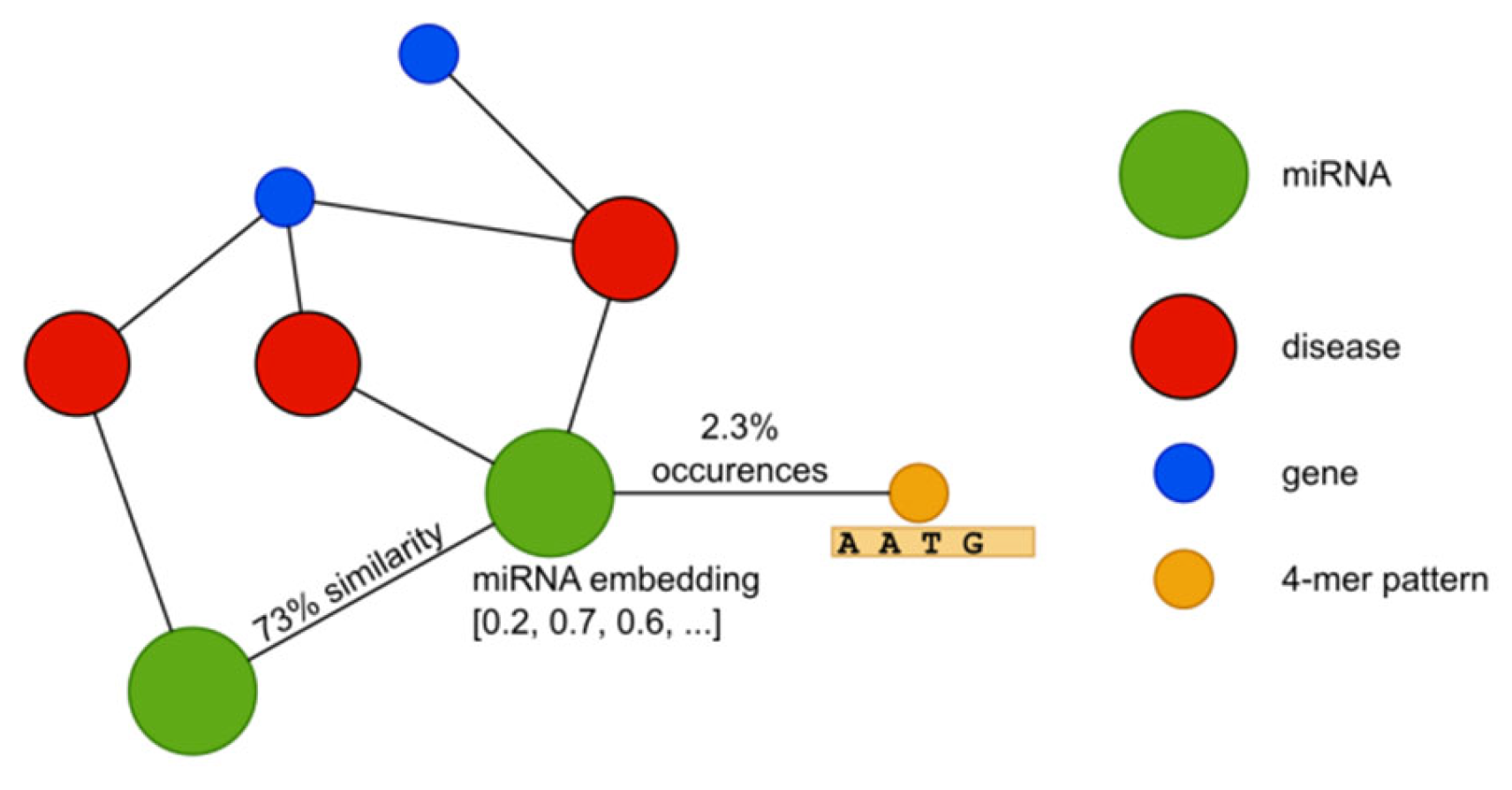
| version 2 | version 3.2 | version 4 | |
| nodes | |||
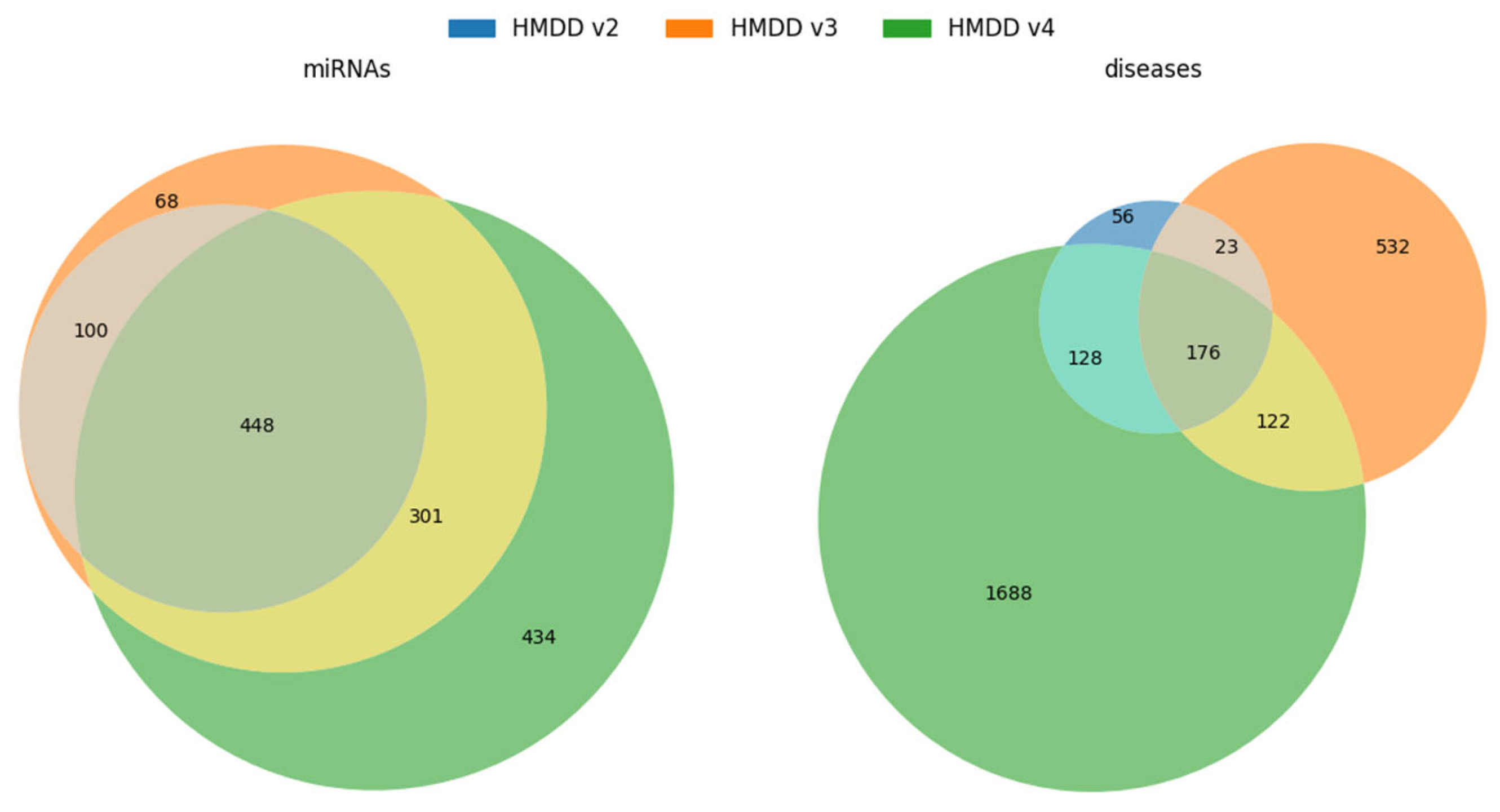
| miRNAs | 548 | 917 | 1183 |
| diseases | 383 | 853 | 2114 |
| genes | 6356 | 6356 | 6356 |
| patterns (4-mers) | 256 | 256 | 256 |
| edges | |||
| miRNA–disease | 6331 (3.02%) |
15161 (1.94%) |
24074 (0.96%) |
| miRNA–miRNA similarity |
58814 (19.58%) |
133958 (15.93%) |
209186 (14.95%) |
| disease–gene |
11977 0.49%) |
13683 (0.25%) |
18617 (0.14%) |
| miRNA–pattern |
36602 (24.27%) |
58695 (25.0%) |
73515 (26.09%) |
| Method | Precision | Recall | F1-Score | AUCROC | AUPR |
| SVM | 83.69 ± 0.85 | 83.71 ± 1.43 | 83.70 ± 0.75 | 90.91 ± 0.31 | 90.57 ± 0.36 |
| GBDT | 83.69 ± 1.07 | 84.90 ± 0.57 | 84.29 ± 0.54 | 91.72 ± 0.34 | 91.38 ± 0.39 |
| RF | 84.24 ± 1.08 | 83.54 ± 1.31 | 83.88 ± 0.91 | 91.41 ± 0.49 | 91.23 ± 0.47 |
| XGBoost | 84.71 ± 0.90 | 84.86 ± 0.99 | 84.78 ± 0.76 | 91.91 ± 0.39 | 91.65 ± 0.45 |
| ELMDA | 84.85 ± 1.39 | 85.36 ± 1.01 | 85.10 ± 0.94 | 92.29 ± 0.35 | 92.17 ± 0.31 |
| MDA-CF | - | - | - | 92.58 | - |
| TCRWMDA | - | - | - | 92.09 | - |
| WBSMDA | - | - | - | 81.85 | - |
| ABMDA | - | - | - | 90.45 | - |
| ICFMDA | N.A. | N.A. | N.A. | 90.23 | N.A. |
| P.A. v2 | 92.02 ± 0.90 | 96.19 ± 0.90 | 94.06 ± 0.63 | 97.10 ± 0.19 | 95.93 ± 0.66 |
| P.A. v3 | 91.49 ± 1.09 | 94.79 ± 1.02 | 93.11 ± 0.92 | 96.44 ± 0.46 | 94.54 ± 1.14 |
| P.A. v4 | 94.94 ± 0.57 | 90.56 ± 1.79 | 92.70 ± 1.01 | 98.06 ± 0.26 | 94.38 ± 0.63 |
| Dropped edge | Decrease AUC |
| miRNA–miRNA similarity | 5.4 % |
| disease–gene | 11.2 % |
| miRNA–pattern | 3.4 % |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




