Submitted:
08 January 2026
Posted:
09 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Complete Methodological Description of the Proposed Pipeline
2.1. Computational Formulation of the Problem
2.2. Overview of the Methodological Pipeline
2.3. Intrinsic Challenges of Morphological Classification
2.3.1. Imbalance between Morphological Classes
2.3.2. Impact of Imbalance on Training and Decision-Making
2.3.3. Structural Variability across Images
2.4. Methodological Implications
3. Dataset Construction and Labeling Strategy
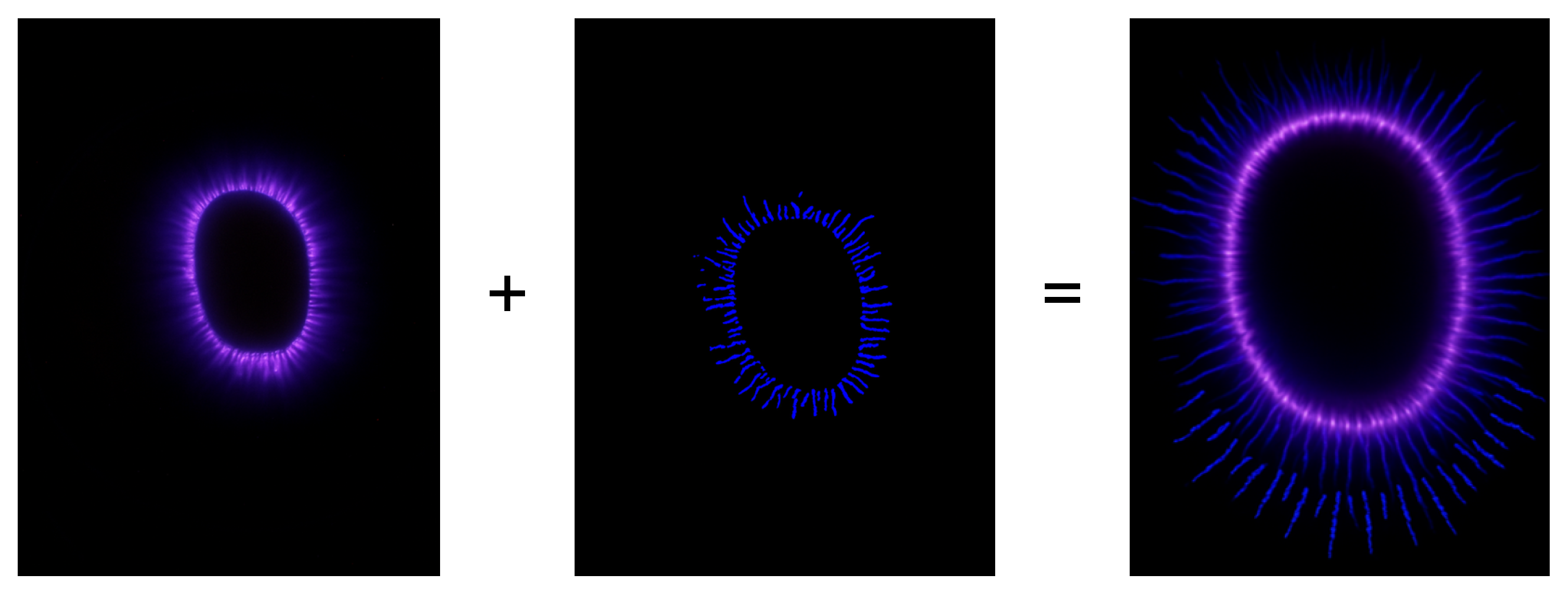
3.1. Operational Definitions of Morphological Patterns
3.2. Data Origin and Dataset Volumetry
3.3. Labeling Strategy and XOR Consistency Rule
3.4. Initial Analysis of Class Imbalance
3.5. Reproducibility and Dataset Construction Protocol
4. Materials and Methods
4.1. Image Preprocessing and Patch Extraction
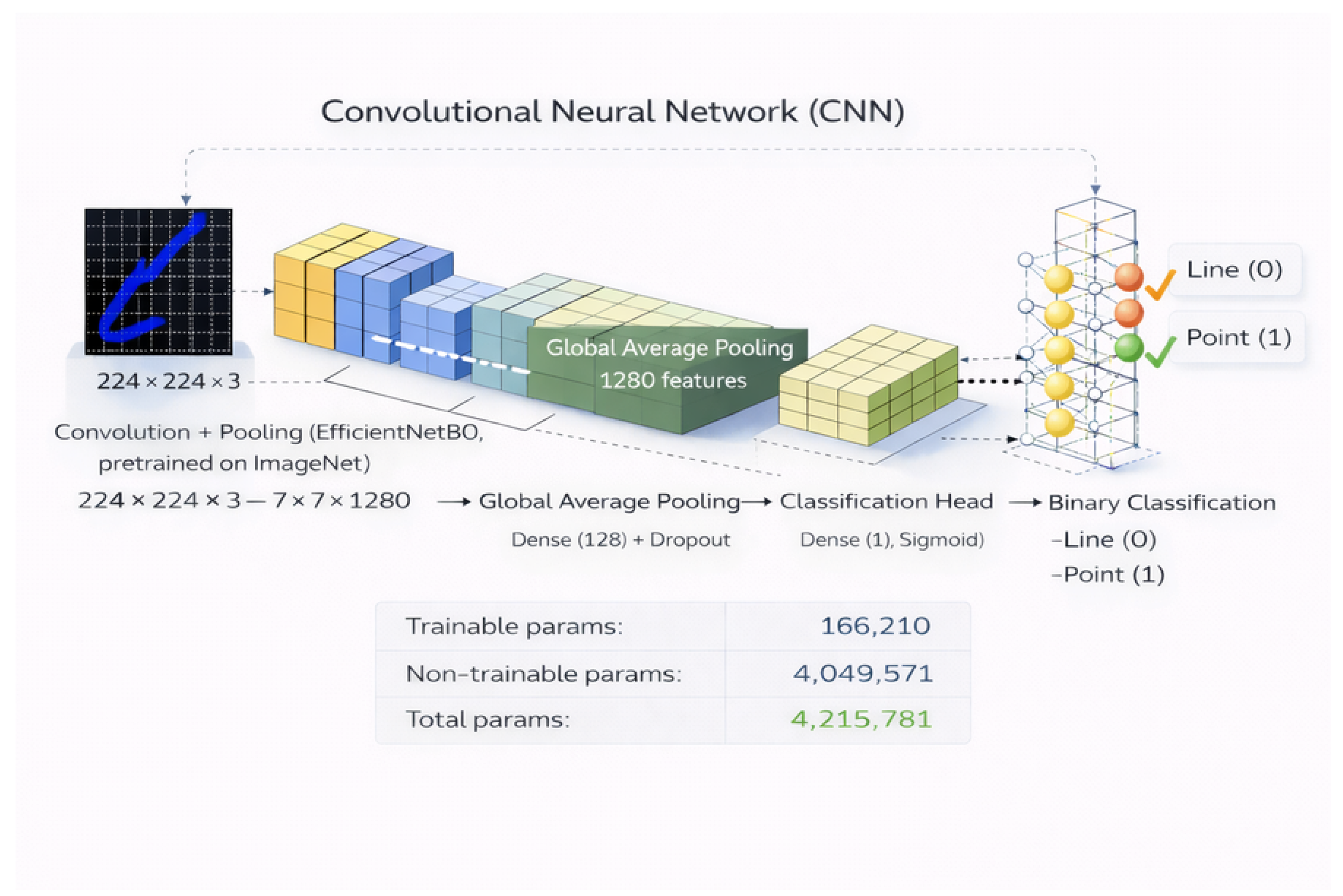
4.2. CNN Architecture and Training Protocol
4.3. Data Splitting Strategy and Leakage Prevention
4.4. Class Imbalance Handling
4.5. Decision Threshold Calibration Protocol
5. Results
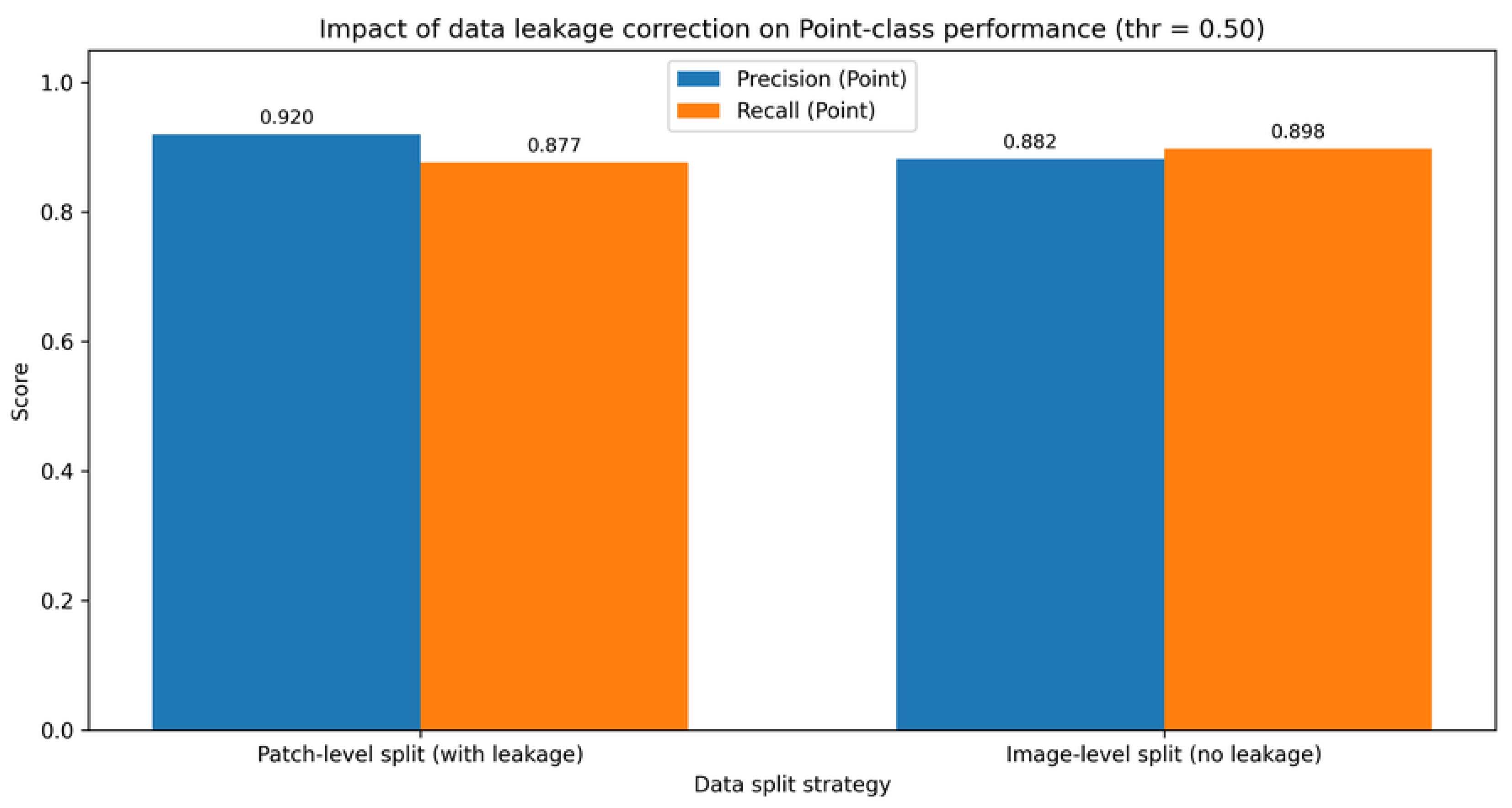
5.1. Impact of Image-Level Data Splitting
5.2. Decision Threshold Calibration Results
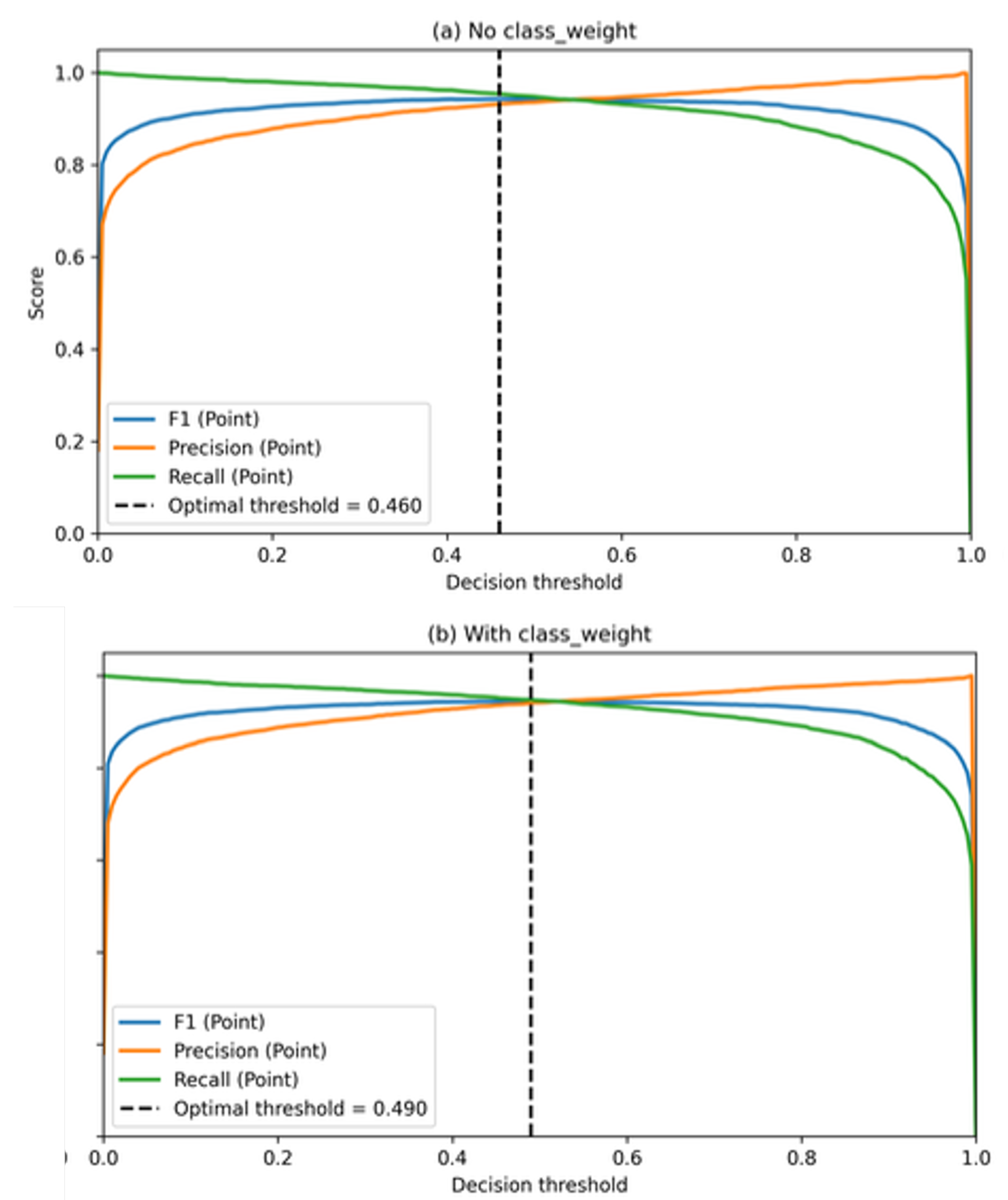
| Configuration | Optimal threshold | Precision (P) | Recall (P) | F1-score (P) |
|---|---|---|---|---|
| No class_weight | 0.460 | 0.932 | 0.954 | 0.943 |
| With class_weight | 0.490 | 0.941 | 0.948 | 0.944 |
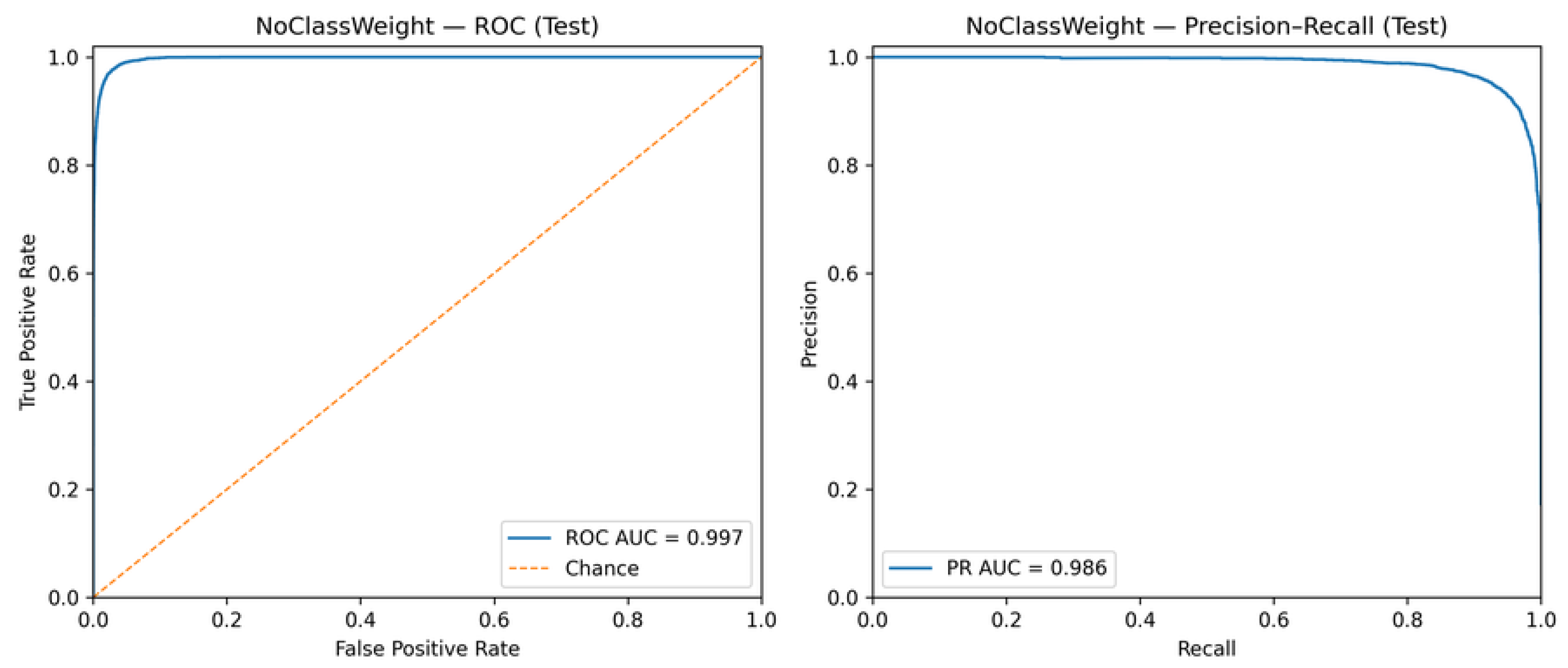
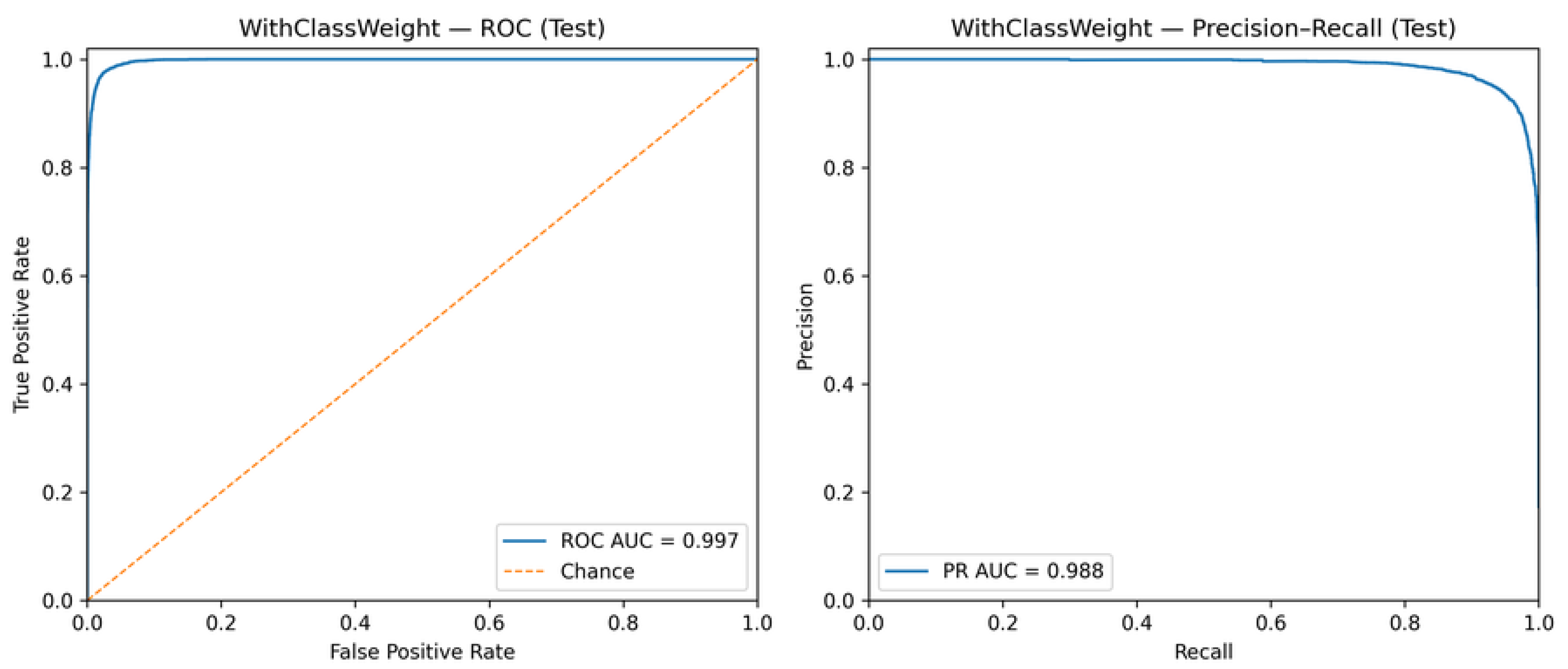
5.3. ROC and Precision–Recall Analysis
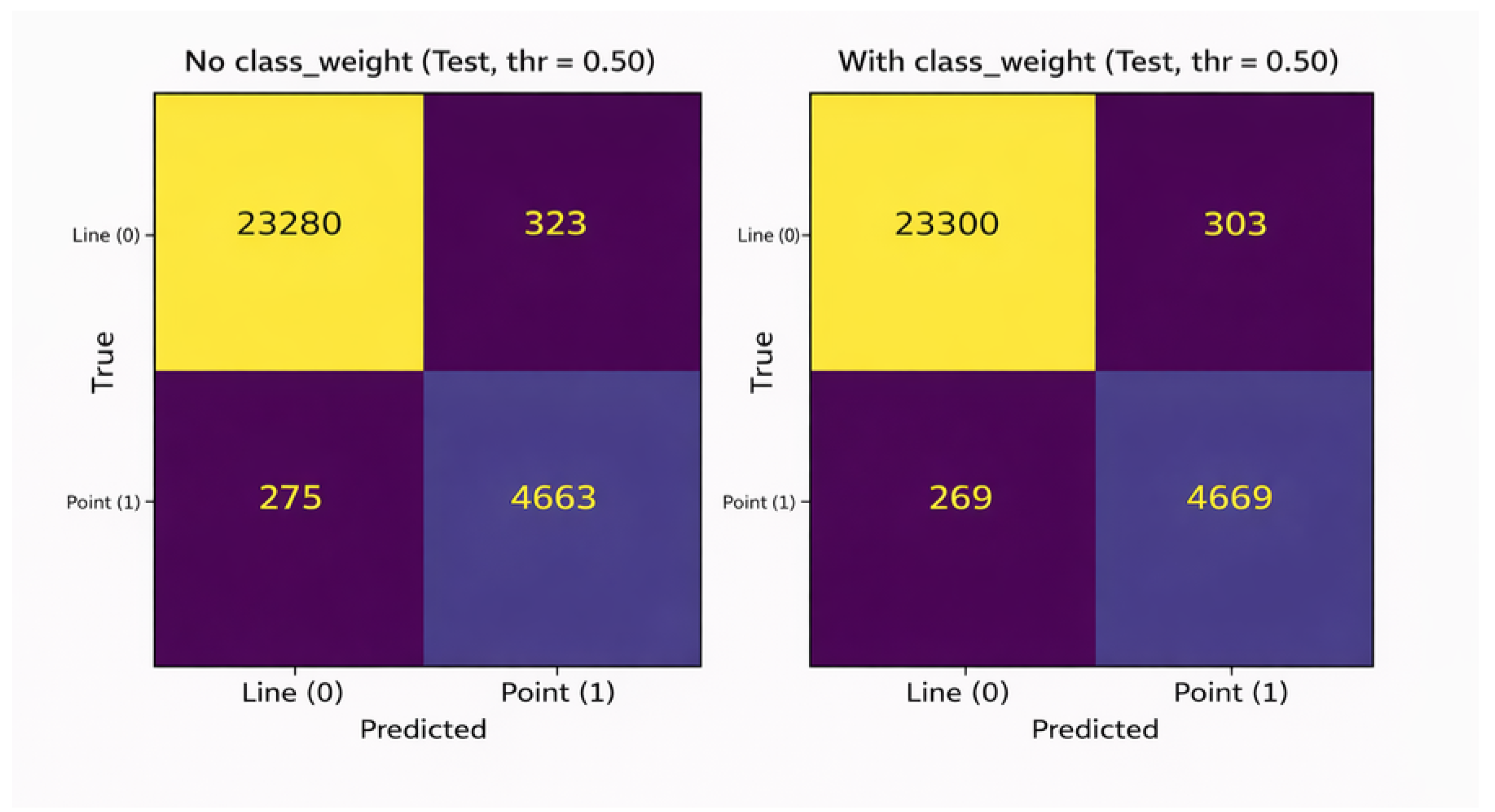
5.4. Confusion Matrix Analysis
5.5. Image-Level Aggregation Results
6. Structural Variability Across Images
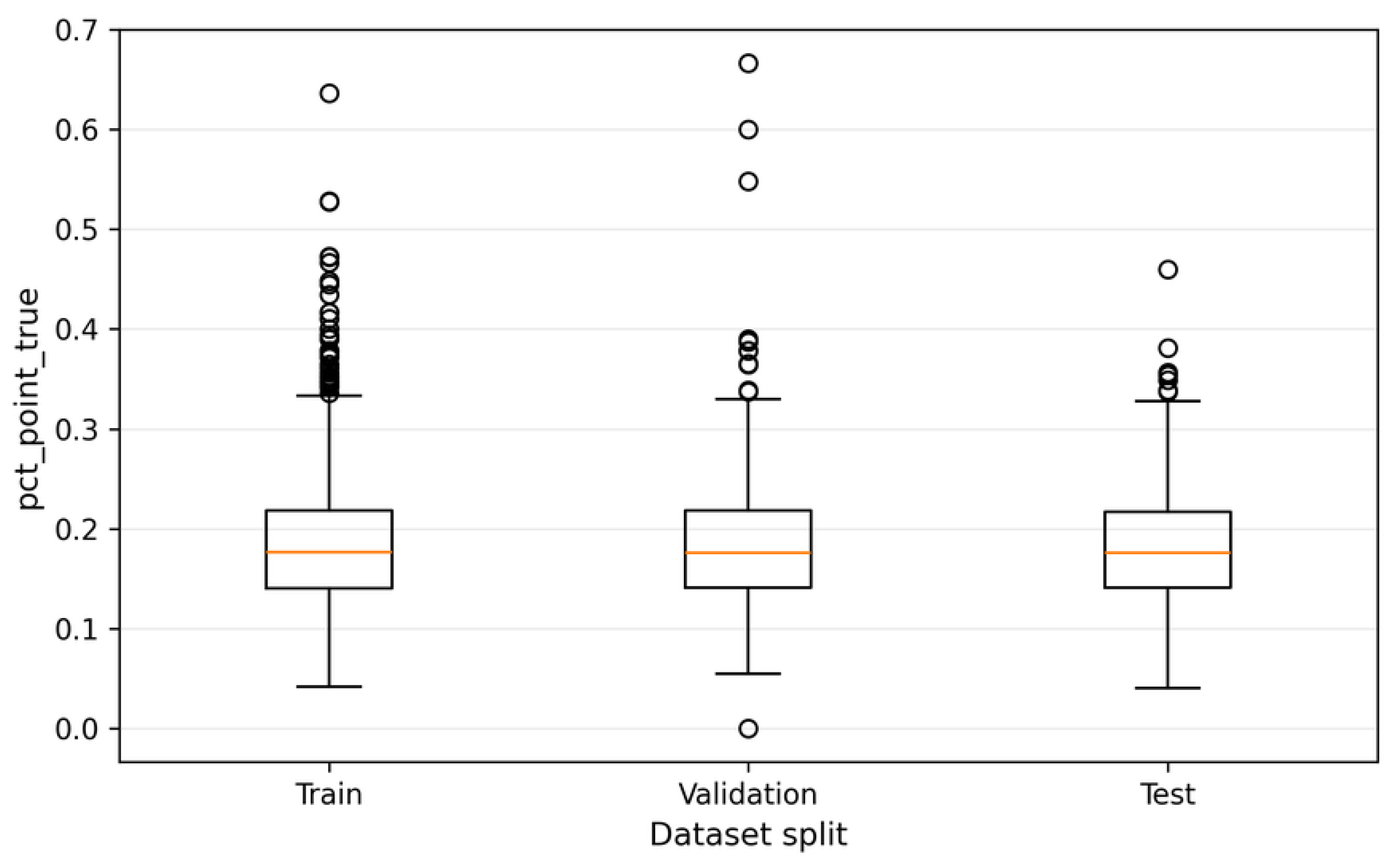
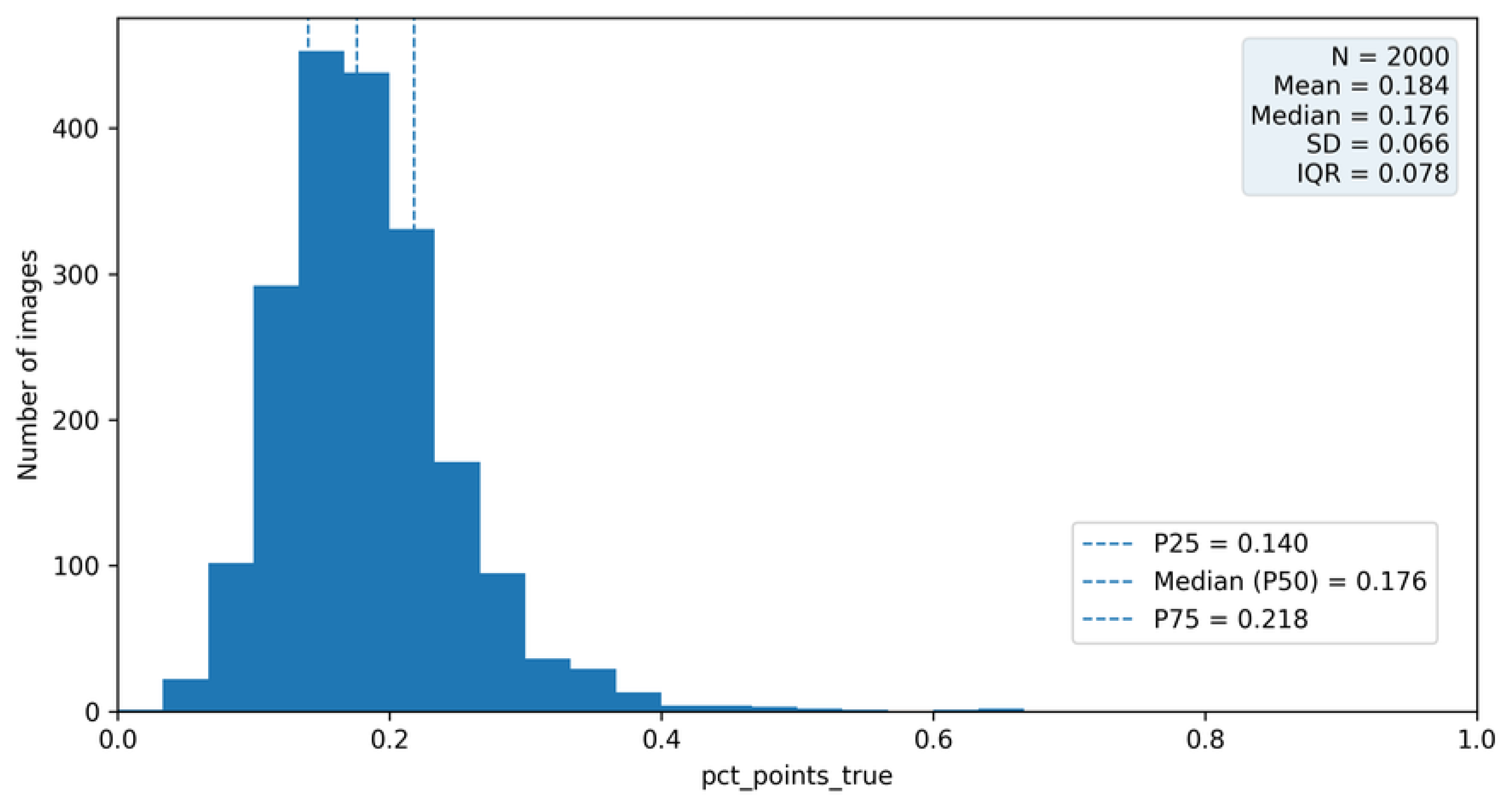
6.1. Definition of the pct_point_true Metric
6.2. Distribution and Interpretation of Structural Variability
7. Integrated Results Analysis
8. Discussion
9. Limitations and Implications
10. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CNN | Convolutional Neural Network |
| ROC | Receiver Operating Characteristic |
| PR | Precision–Recall |
| DoG | Difference of Gaussians |
References
- Pehek, J.O.; Kyler, H.J.; Faust, D.L. Image Modulation in Corona Discharge Photography. Science 1976, 194, 263–270. [Google Scholar] [CrossRef]
- Korotkov, K.G.; Matravers, P.; Orlov, D.V.; Williams, B.O. Application of Electrophoton Capture (EPC) Analysis Based on Gas Discharge Visualization (GDV) Technique in Medicine: A Systematic Review. The Journal of Alternative and Complementary Medicine 2010, 16, 13–25. [Google Scholar] [CrossRef]
- Shichkina, Y.; Fatkieva, R.; Sychev, A.; Kazak, A. Method for Detecting Pathology of Internal Organs Using Bioelectrography. Diagnostics 2024, 14, 991. [Google Scholar] [CrossRef]
- Komura, D.; Ishikawa, S. Machine Learning Methods for Histopathological Image Analysis. Computational and Structural Biotechnology Journal 2018, 16, 34–42. [Google Scholar] [CrossRef]
- Zhou, S.; Greenspan, H.; Shen, D. Deep Learning for Medical Image Analysis: A Survey. Neurocomputing 2021, 452, 631–647. [Google Scholar]
- Gonzalez, R.C.; Woods, R.E. Digital Image Processing, 4 ed.; Pearson, 2018. [Google Scholar]
- Wang, J.; Yang, Y.; Chen, X. Patch-based image classification: A review. Pattern Recognition 2019, 88, 32–48. [Google Scholar]
- Litjens, G.; Kooi, T.; Bejnordi, B.E.; et al. A Survey on Deep Learning in Medical Image Analysis. Medical Image Analysis 2017, 42, 60–88. [Google Scholar] [CrossRef] [PubMed]
- Roberts, M.; Driggs, D.; Thorpe, M.; et al. Common Pitfalls and Recommendations for Using Machine Learning to Detect and Prognosticate for COVID-19 Using Chest Radiographs and CT Scans. Nature Machine Intelligence 2021, 3, 199–217. [Google Scholar] [CrossRef]
- Haralick, R.M.; Shanmugam, K.; Dinstein, I. Textural features for image classification. IEEE Transactions on Systems, Man, and Cybernetics 1973, SMC-3, 610–621. [Google Scholar] [CrossRef]
- Carbonneau, M.A.; Cheplygina, V.; Granger, E.; Gagnon, G. Multiple Instance Learning: A Survey of Problem Characteristics and Applications. Pattern Recognition 2018, 77, 329–353. [Google Scholar] [CrossRef]
- Ojala, T.; Pietikäinen, M.; Mäenpää, T. Multiresolution gray-scale and rotation invariant texture classification with local binary patterns. IEEE Transactions on Pattern Analysis and Machine Intelligence 2002, 24, 971–987. [Google Scholar] [CrossRef]
- Campanella, G.; Hanna, M.G.; Geneslaw, L.; et al. Clinical-Grade Computational Pathology Using Weakly Supervised Deep Learning on Whole Slide Images. Nature Medicine 2019, 25, 1301–1309. [Google Scholar] [CrossRef]
- Kather, J.N.; Krisam, J.; Charoentong, P.; et al. Predicting Survival from Colorectal Cancer Histology Slides Using Deep Learning. PLOS Medicine 2019, 16, e1002730. [Google Scholar] [CrossRef] [PubMed]
- Bulten, W.; Pinckaers, H.; van Boven, H.; et al. Automated Deep-Learning System for Gleason Grading of Prostate Cancer Using Biopsies. The Lancet Oncology 2020, 21, 233–241. [Google Scholar] [CrossRef] [PubMed]
- He, H.; Garcia, E.A. Learning from imbalanced data. IEEE Transactions on Knowledge and Data Engineering 2009, 21, 1263–1284. [Google Scholar] [CrossRef]
- Saito, T.; Rehmsmeier, M. The precision–recall plot is more informative than the ROC plot when evaluating binary classifiers on imbalanced datasets. PLOS ONE 2015, 10, e0118432. [Google Scholar] [CrossRef]
- Guo, C.; Pleiss, G.; Sun, Y.; Weinberger, K.Q. On Calibration of Modern Neural Networks. In Proceedings of the 34th International Conference on Machine Learning (ICML); 2017; pp. 1321–1330. [Google Scholar]
- Korotkov, K. Human Energy Field: Study with GDV Bioelectrography; CRC Press, 2012. [Google Scholar]
- LeCun, Y.; Bengio, Y.; Hinton, G. Deep learning. Nature 2015, 521, 436–444. [Google Scholar] [CrossRef] [PubMed]
- Japkowicz, N.; Stephen, S. The Class Imbalance Problem: A Systematic Study. Intelligent Data Analysis 2002, 6, 429–449. [Google Scholar] [CrossRef]
- Davis, J.; Goadrich, M. The Relationship between Precision–Recall and ROC Curves. In Proceedings of the Proceedings of the 23rd International Conference on Machine Learning, 2006; pp. 233–240. [Google Scholar]
- Hand, D.J. Measuring Classifier Performance: A Coherent Alternative to the Area Under the ROC Curve. Machine Learning 2009, 77, 103–123. [Google Scholar] [CrossRef]
- Frénay, B.; Verleysen, M. Classification in the Presence of Label Noise: A Survey. IEEE Transactions on Neural Networks and Learning Systems 2014, 25, 845–869. [Google Scholar] [CrossRef] [PubMed]
- Mongan, J.; Moy, L.; Kahn, C.E.J. Checklist for Artificial Intelligence in Medical Imaging (CLAIM): A Guide for Authors and Reviewers. Radiology: Artificial Intelligence 2020, 2, e200029. [Google Scholar] [CrossRef] [PubMed]
- Szeliski, R. Computer Vision: Algorithms and Applications; Springer, 2010. [Google Scholar]
- Soille, P. Morphological Image Analysis: Principles and Applications; Springer, 2003. [Google Scholar]
- Serra, J. Image Analysis and Mathematical Morphology; Academic Press, 1982. [Google Scholar]
- Marr, D.; Hildreth, E. Theory of Edge Detection. Proceedings of the Royal Society of London B 1980, 207, 187–217. [Google Scholar] [CrossRef]
- Lindeberg, T. Scale-Space Theory: A Basic Tool for Analyzing Structures at Different Scales. Journal of Applied Statistics 1994, 21, 225–270. [Google Scholar] [CrossRef]
- Lowe, D.G. Distinctive Image Features from Scale-Invariant Keypoints. International Journal of Computer Vision 2004, 60, 91–110. [Google Scholar] [CrossRef]
- Yosinski, J.; Clune, J.; Bengio, Y.; Lipson, H. How Transferable Are Features in Deep Neural Networks? Advances in Neural Information Processing Systems 2014, 27. [Google Scholar]
- Deng, J.; Dong, W.; Socher, R.; Li, L.J.; Li, K.; Fei-Fei, L. ImageNet: A Large-Scale Hierarchical Image Database. In Proceedings of the Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2009; pp. 248–255. [Google Scholar]
- Russakovsky, O.; Deng, J.; Su, H.; et al. ImageNet Large Scale Visual Recognition Challenge. International Journal of Computer Vision 2015, 115, 211–252. [Google Scholar] [CrossRef]
- Lin, M.; Chen, Q.; Yan, S. Network In Network. arXiv 2013, arXiv:1312.4400. [Google Scholar]
- Goodfellow, I.; Bengio, Y.; Courville, A. Deep Learning; MIT Press, 2016. [Google Scholar]
- Wolff, J.; Büchel, C.; Kather, J.N. Leakage in Biomedical Image Classification: Sources, Consequences and Remedies. Nature Machine Intelligence 2022, 4, 371–380. [Google Scholar]
- King, G.; Zeng, L. Logistic Regression in Rare Events Data. Political Analysis 2001, 9, 137–163. [Google Scholar] [CrossRef]
- Cui, Y.; Jia, M.; Lin, T.; Song, Y.; Belongie, S. Class-Balanced Loss Based on Effective Number of Samples. In Proceedings of the Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2019; pp. 9268–9277. [Google Scholar] [CrossRef]
- Lin, T.; Goyal, P.; Girshick, R.; He, K.; Dollár, P. Focal Loss for Dense Object Detection. In Proceedings of the Proceedings of the IEEE International Conference on Computer Vision (ICCV), 2017; pp. 2980–2988. [Google Scholar] [CrossRef]
- Niculescu-Mizil, A.; Caruana, R. Predicting Good Probabilities with Supervised Learning. In Proceedings of the Proceedings of the 22nd International Conference on Machine Learning (ICML), 2005; pp. 625–632. [Google Scholar]
- Fawcett, T. An Introduction to ROC Analysis. Pattern Recognition Letters 2006, 27, 861–874. [Google Scholar] [CrossRef]
- Forman, G.; Scholz, M. Apples-to-Apples in Cross-Validation Studies: Pitfalls in Classifier Performance Measurement. SIGKDD Explorations Newsletter 2010, 12, 49–57. [Google Scholar] [CrossRef]
- Kohavi, R. A Study of Cross-Validation and Bootstrap for Accuracy Estimation and Model Selection. In Proceedings of the Proceedings of the 14th International Joint Conference on Artificial Intelligence (IJCAI), 1995; pp. 1137–1143. [Google Scholar]


| Statistic | Value |
|---|---|
| Number of images | 2,000 |
| Number of patches | 191,964 |
| Mean patches per image | 95.98 |
| Standard deviation | 25.13 |
| Minimum | 3 |
| 1st Quartile (Q1) | 81 |
| Median (Q2) | 97 |
| 3rd Quartile (Q3) | 112 |
| Maximum | 200 |
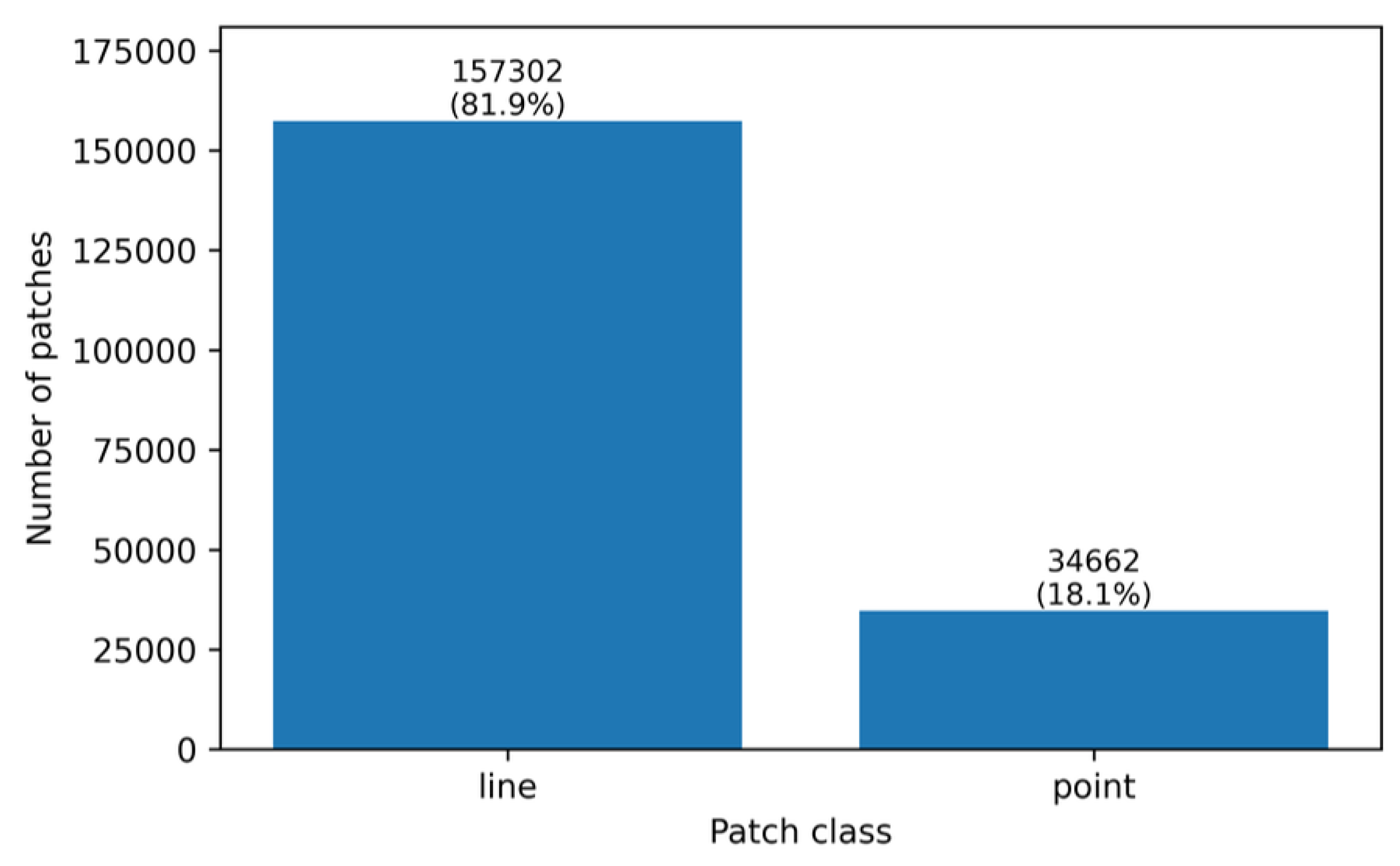
| Class | Number of patches | Percentage |
|---|---|---|
| Line | 157,302 | 81.94% |
| Point | 34,662 | 18.06% |
| Total | 191,964 | 100.00% |
| Stage | Parameter | Value |
|---|---|---|
| Preprocessing (edges / mask) | Ring threshold percentile () | 85 |
| Ring band width () | 18 | |
| Inner margin () | 9 | |
| Inner edge band () | 12 | |
| Gaussian blur kernel | ||
| Gaussian blur sigma | 1.3 | |
| DoG sigma1 | 0.40 | |
| DoG sigma2 | 1.90 | |
| DoG threshold percentile | 65 | |
| Morphological opening kernel | 2 | |
| Minimum blob area | 25 | |
| Resize max side (pixels) | 900 | |
| IoU acceptance threshold | 0.40 | |
| Adaptive thresholding | Target P95 | 210.0 |
| Scale factor (K) | 2.5 | |
| Threshold min / max | 65 / 72 | |
| Density factor | 1.75 | |
| Components factor | 2.0 | |
| Connectedness criterion | IoU-based filtering | |
| Binary mask generation | Enabled | |
| Patch extraction | Minimum component area | 30 |
| Padding around component | 5 | |
| CNN input and split | Input size | |
| Batch size | 32 | |
| Random seed | 42 | |
| Split strategy | Stratified by image_id | |
| Validation fraction | 0.15 | |
| Test fraction | 0.15 | |
| CNN training | Loss function | Binary cross-entropy |
| Optimizer (initial) | Adam () | |
| Optimizer (fine-tuning) | Adam () | |
| Epochs (initial) | 15 | |
| Epochs (fine-tuning) | 10 | |
| Early stopping | Enabled (patience = 5) | |
| Dropout rate | 0.2 | |
| Weight decay | Not applied | |
| Decision calibration | Output activation | Sigmoid |
| Threshold sweep | 201 values in | |
| Sweep step | 0.005 | |
| Selection criterion | Max F1 on validation set |
| Split strategy | Acc. | Prec. (P) | Rec. (P) | F1 (P) | Supp. (P) | FP (L→P) | FN (P→L) |
|---|---|---|---|---|---|---|---|
| Patch-level split (with leakage) | 0.9644 | 0.9202 | 0.8769 | 0.8981 | 5144 | 391 | 633 |
| Image-level split (no leakage) | 0.9600 | 0.8819 | 0.8979 | 0.8898 | 5240 | 630 | 535 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




