Submitted:
05 January 2026
Posted:
07 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
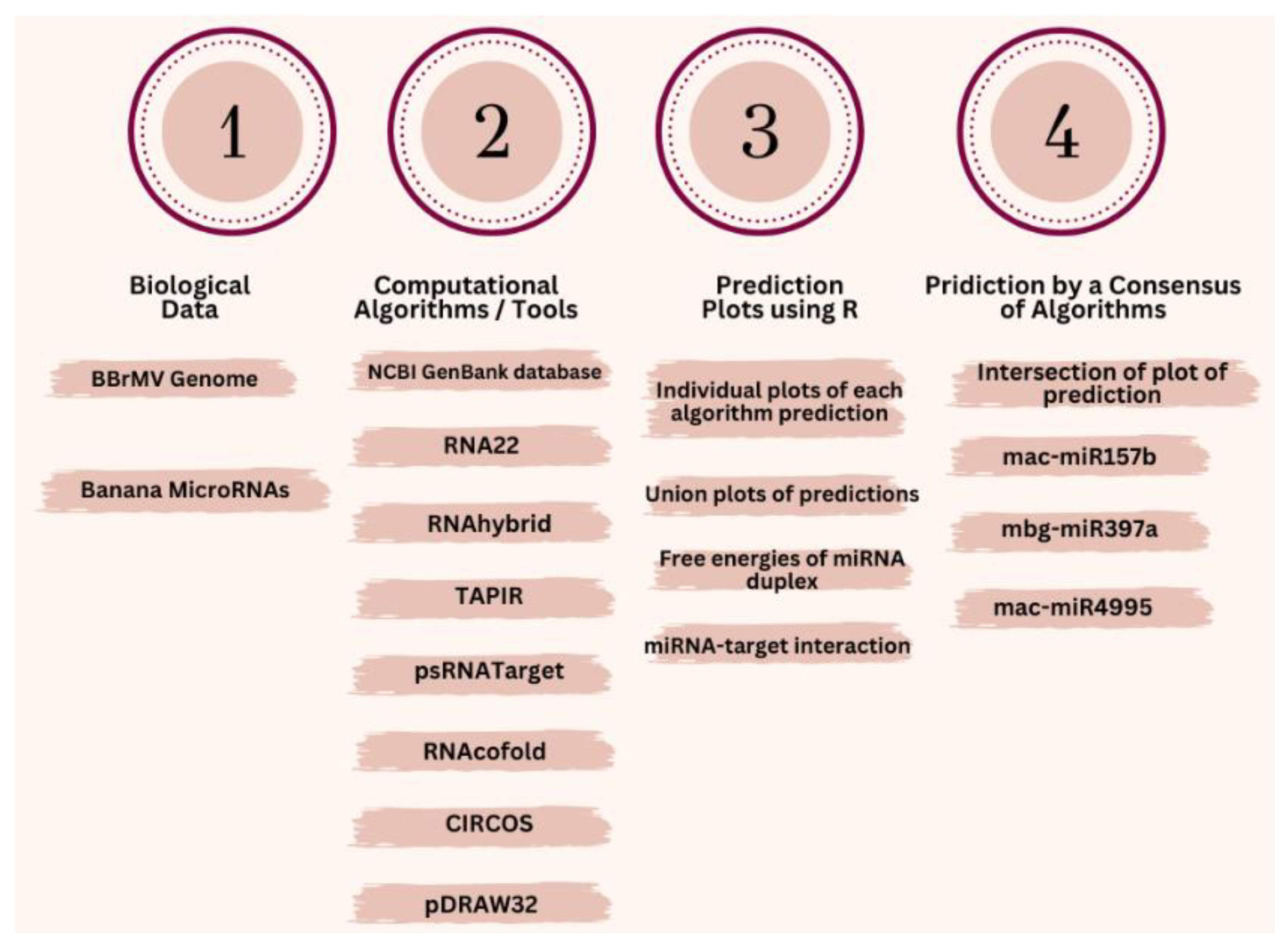
2.1. Banana MicroRNAs and BBrMV Genome Sequence
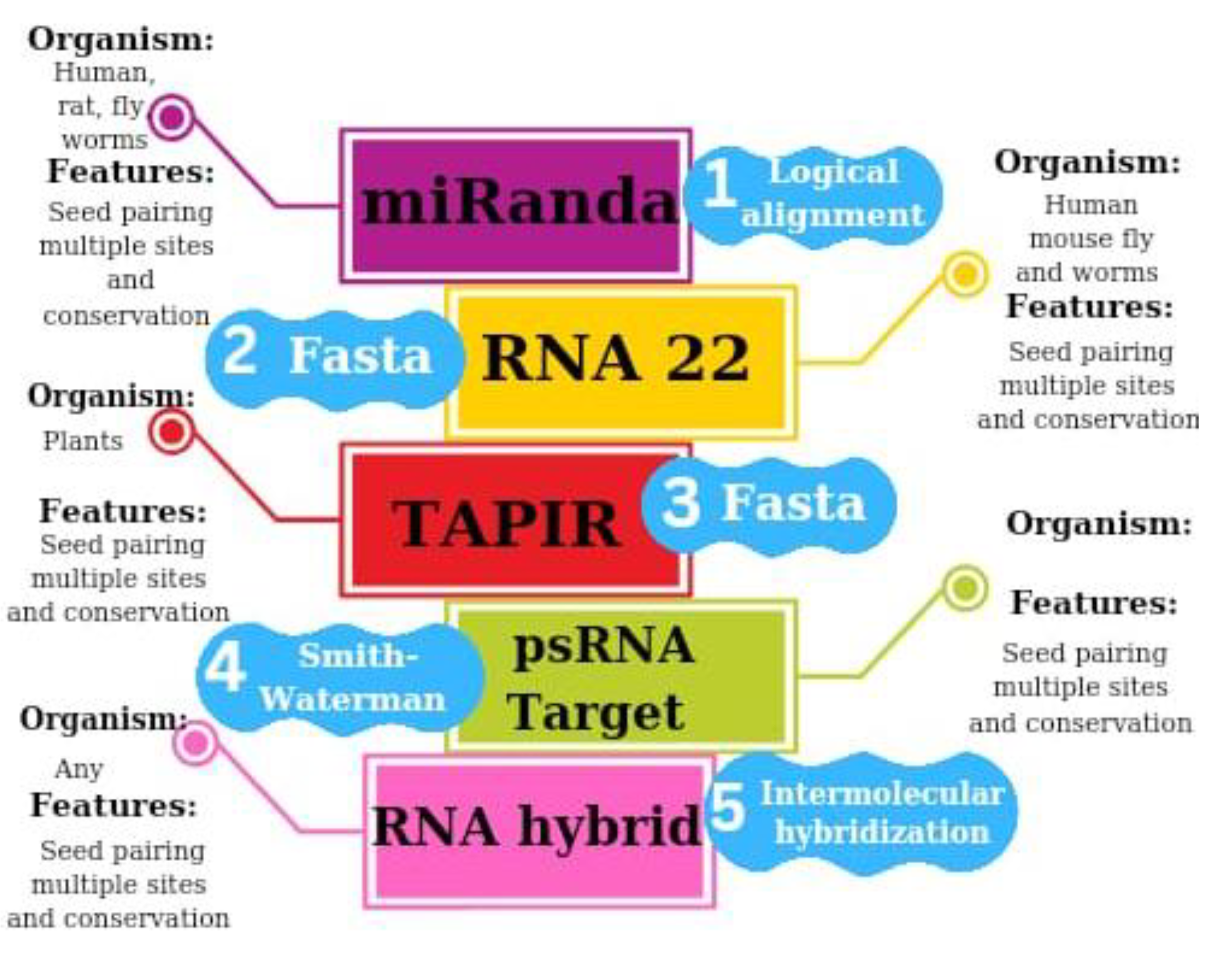
2.2. RNA22 Algorithm
2.3. RNAhybrd Algorithm
2.4. TAPIR Algorithm
2.5. psRNATarget Algorithm
2.6. Mapping of miRNAs-Target Interaction
2.7. RNAcofold Algorithm
2.8. Statistical Analysis
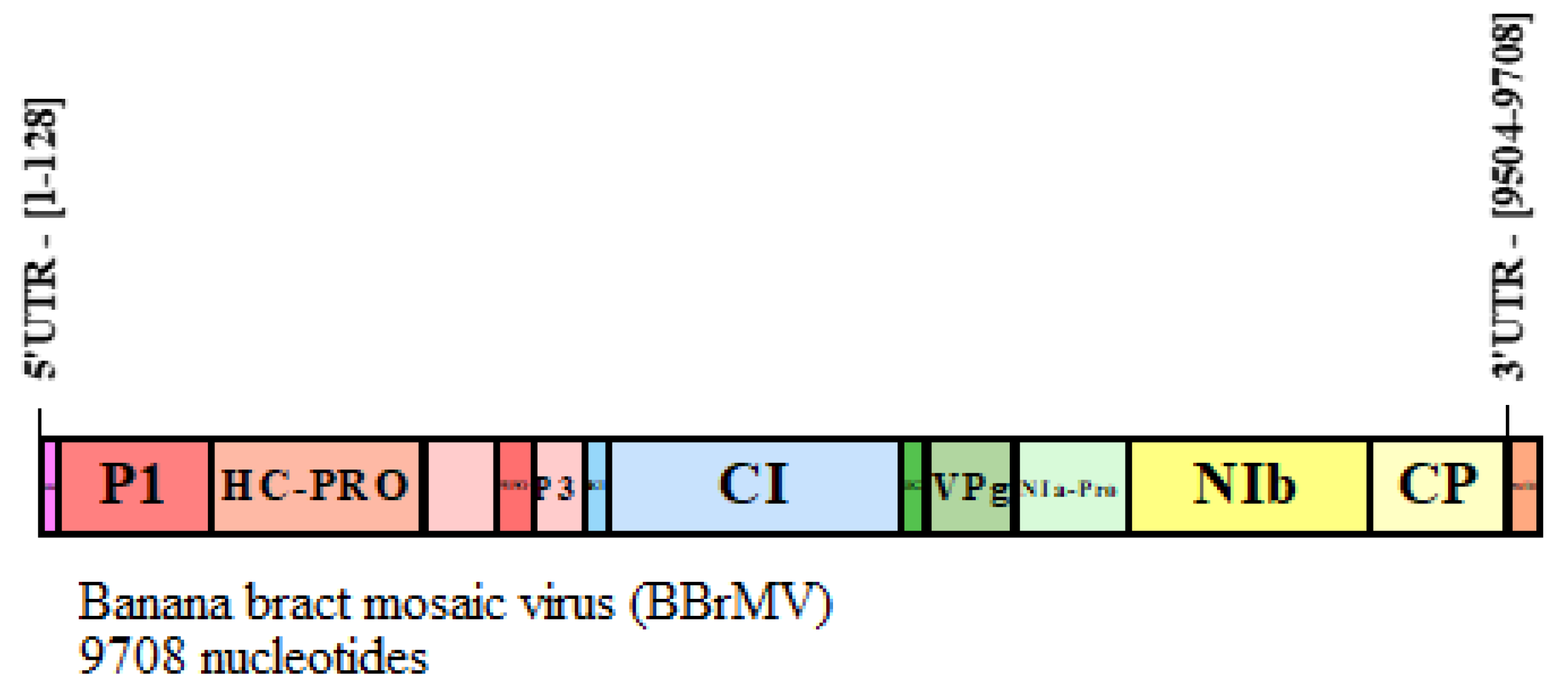
2.9. BBrMV Genome Annotation
3. Results
3.1. Banana Locus-Derived miRNA-mRNA Interactive Pairs for the BBrMV Genome
3.2. Predicted Targets for P1 ORF of BBrMV
3.3. Helper-Component Proteinase (HC-Pro) of BBrMV Genome
3.4. Membrane Associated Protein (P3) of BBrMV
3.5. Banana miRNAs Targeting 6K1
3.6. Cylindrical Inclusion Protein (CI) of BBrMV Genome
3.7. Banana miRNAs Targeting 6K2
3.8. Viral Protein Genome-Linked (VPg) of the BBrMV
3.9. Nuclear Inclusion-a Protease (NIa-Pro) of BBrMV Genome
3.10. Nuclear Inclusion-b Protease (NIb) of BBrMV Genome
3.10.1. Coat Protein (CP) of BBrMV Genome
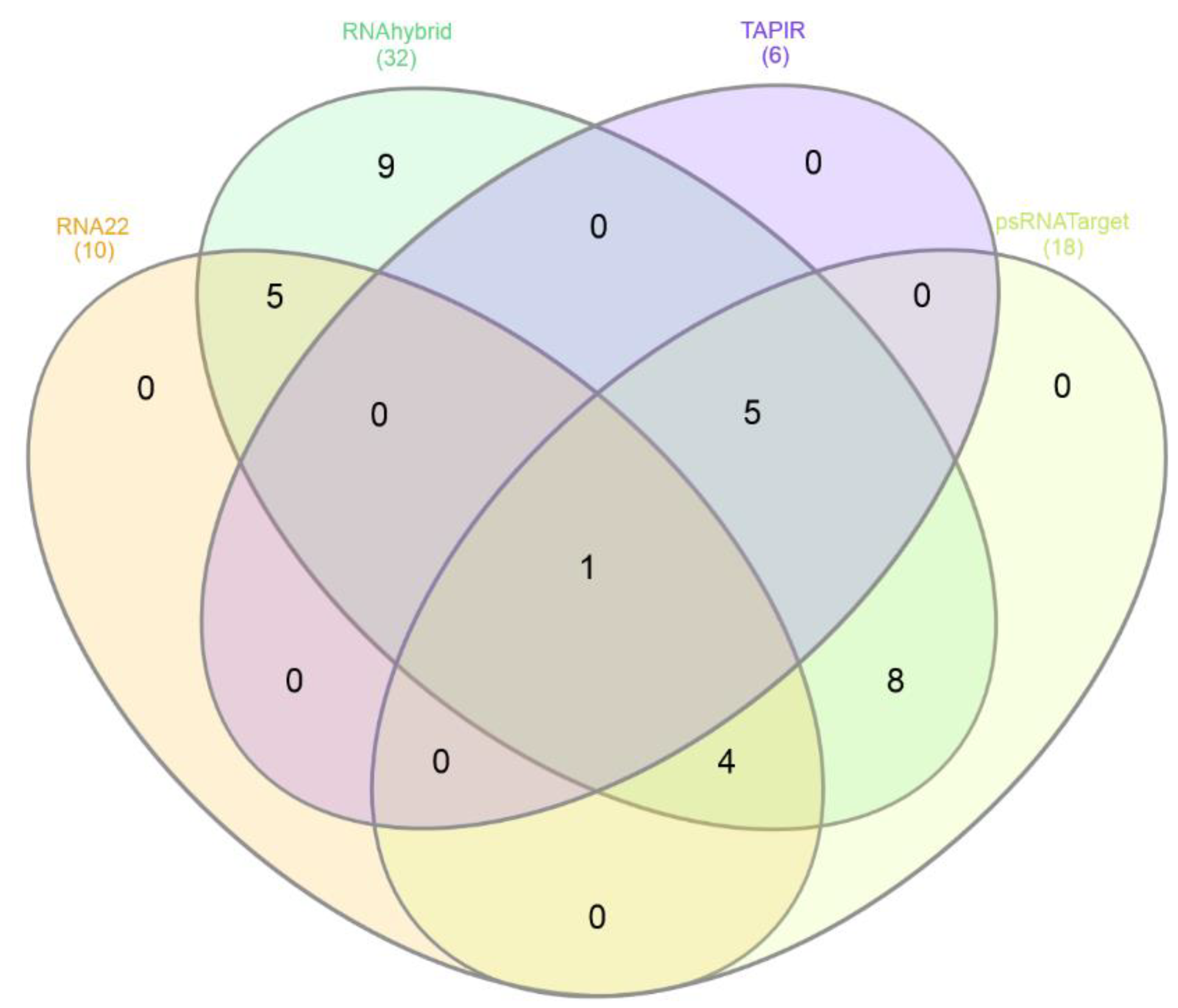
3.10.2. Consensus and Unique miRNAs Prediction
3.10.3. Integrated Analysis of miRNA-Target Network
3.11. Assessment of Free Energy of Interaction (ΔG)
4. Discussion
5. Challenges and Limitations
6. Conclusion and Future Directions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Maseko, K.H.; Regnier, T.; Meiring, B.; Wokadala, O.C.; Anyasi, T.A. Musa species variation, production, and the application of its processed flour: A review. Scientia Horticulturae 2024, 325, 112688. [Google Scholar] [CrossRef]
- Pereira, A.; Maraschin, M. Banana (Musa spp) from peel to pulp: ethnopharmacology, source of bioactive compounds and its relevance for human health. Journal of ethnopharmacology 2015, 160, 149–163. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Yu, S.; Cheng, Z.; Chang, X.; Yun, Y.; Jiang, M.; Chen, X.; Wen, X.; Li, H.; Zhu, W. Origin and evolution of the triploid cultivated banana genome. Nature genetics 2024, 56, 136–142. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.-R.; Liu, X.; Arshad, R.; Wang, X.; Li, W.-M.; Zhou, Y.; Ge, X.-J. Telomere-to-telomere haplotype-resolved reference genome reveals subgenome divergence and disease resistance in triploid Cavendish banana. Horticulture research 2023, 10, uhad153. [Google Scholar] [CrossRef]
- Selvarajan, R.; Balasubramanian, V.; Priyanka, P.; Jebakumar, R.M.; Selvam, K.P.; Uma, S. Evidence of seed transmission of Banana bract mosaic virus in Musa synthetic diploid H-201, a possible threat to banana breeding. European Journal of Plant Pathology 2020, 156, 1181–1187. [Google Scholar] [CrossRef]
- Bhat, A.; Pamitha, N.; Gopika, A.; Biju, C. Complete genome sequencing of banana bract mosaic virus isolate infecting cardamom revealed its closeness to banana infecting isolate from India. VirusDisease 2018, 29, 212–215. [Google Scholar] [CrossRef]
- Bajczyk, M.; Jarmolowski, A.; Jozwiak, M.; Pacak, A.; Pietrykowska, H.; Sierocka, I.; Swida-Barteczka, A.; Szewc, L.; Szweykowska-Kulinska, Z. Recent insights into plant miRNA biogenesis: multiple layers of miRNA level regulation. Plants 2023, 12, 342. [Google Scholar] [CrossRef]
- Bravo-Vázquez, L.A.; Castro-Pacheco, A.M.; Pérez-Vargas, R.; Velázquez-Jiménez, J.F.; Paul, S. The Emerging Applications of Artificial MicroRNA-Mediated Gene Silencing in Plant Biotechnology. Non-coding RNA 2025, 11, 19. [Google Scholar] [CrossRef]
- Khalid, A.; Zhang, X.; Ji, H.; Yasir, M.; Farooq, T.; Dai, X.; Li, F. Large artificial microRNA cluster genes confer effective resistance against multiple tomato yellow leaf curl viruses in transgenic tomato. Plants 2023, 12, 2179. [Google Scholar] [CrossRef]
- Zhou, L.; Yuan, Q.; Ai, X.; Chen, J.; Lu, Y.; Yan, F. Transgenic rice plants expressing artificial miRNA targeting the rice stripe virus MP gene are highly resistant to the virus. Biology 2022, 11, 332. [Google Scholar] [CrossRef]
- Bhakta, S.; Tak, H.; Ganapathi, T.R. Exploring diverse roles of micro RNAs in banana: Current status and future prospective. Physiologia Plantarum 2021, 173, 1323–1334. [Google Scholar] [CrossRef]
- Chai, J.; Feng, R.; Shi, H.; Ren, M.; Zhang, Y.; Wang, J. Bioinformatic identification and expression analysis of banana microRNAs and their targets. Plos one 2015, 10, e0123083. [Google Scholar] [CrossRef]
- Peng, K.; Kong, X.; Li, Y.; Dalmay, T.; Han, Y.; Jiang, Y.; Qu, H.; Zhu, H. Cu-miRNA-mediated redox homeostasis explains the differences in cold tolerance between AAA and ABB banana varieties. Journal of Advanced Research 2025. [Google Scholar] [CrossRef]
- Ashraf, M.A.; Ali, B.; Fareed, M.; Sardar, A.; Saeed, E.; Islam, S.; Bano, S.; Yu, N. In Silico Identification of Banana High-Confidence MicroRNA Binding Sites Targeting Banana Streak GF Virus. 2024. [Google Scholar] [CrossRef]
- Kong, X.; Peng, K.; Shan, Y.; Yun, Z.; Dalmay, T.; Duan, X.; Jiang, Y.; Qu, H.; Zhu, H. Transcriptional regulation of miR528-PPO module by miR156 targeted SPLs orchestrates chilling response in banana. Molecular Horticulture 2025, 5, 2. [Google Scholar] [CrossRef] [PubMed]
- Sayers, E.W.; Beck, J.; Bolton, E.E.; Brister, J.R.; Chan, J.; Connor, R.; Feldgarden, M.; Fine, A.M.; Funk, K.; Hoffman, J. Database resources of the National Center for Biotechnology Information in 2025. Nucleic acids research 2025, 53, D20–D29. [Google Scholar] [CrossRef]
- Miranda, K.C.; Huynh, T.; Tay, Y.; Ang, Y.-S.; Tam, W.-L.; Thomson, A.M.; Lim, B.; Rigoutsos, I. A pattern-based method for the identification of MicroRNA binding sites and their corresponding heteroduplexes. Cell 2006, 126, 1203–1217. [Google Scholar] [CrossRef]
- Loher, P.; Rigoutsos, I. Interactive exploration of RNA22 microRNA target predictions. Bioinformatics 2012, 28, 3322–3323. [Google Scholar] [CrossRef] [PubMed]
- Krüger, J.; Rehmsmeier, M. RNAhybrid: microRNA target prediction easy, fast and flexible. Nucleic acids research 2006, 34, W451–W454. [Google Scholar] [CrossRef]
- Bonnet, E.; He, Y.; Billiau, K.; Van de Peer, Y. TAPIR, a web server for the prediction of plant microRNA targets, including target mimics. Bioinformatics 2010, 26, 1566–1568. [Google Scholar] [CrossRef]
- Dai, X.; Zhuang, Z.; Zhao, P.X. psRNATarget: a plant small RNA target analysis server (2017 release). Nucleic acids research 2018, 46, W49–W54. [Google Scholar] [CrossRef]
- Dai, X.; Zhao, P.X. psRNATarget: a plant small RNA target analysis server. Nucleic acids research 2011, 39, W155–W159. [Google Scholar] [CrossRef]
- Krzywinski, M.; Schein, J.; Birol, I.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: an information aesthetic for comparative genomics. Genome research 2009, 19, 1639–1645. [Google Scholar] [CrossRef]
- Bernhart, S.H.; Tafer, H.; Mückstein, U.; Flamm, C.; Stadler, P.F.; Hofacker, I.L. Partition function and base pairing probabilities of RNA heterodimers. Algorithms for Molecular Biology 2006, 1, 3. [Google Scholar] [CrossRef]
- Gandrud, C. Reproducible research with R and R studio; Chapman and Hall/CRC, 2018. [Google Scholar]
- Pasin, F.; Simon-Mateo, C.; García, J.A. The hypervariable amino-terminus of P1 protease modulates potyviral replication and host defense responses. PLoS Pathogens 2014, 10, e1003985. [Google Scholar] [CrossRef] [PubMed]
- Saha, S.; Lõhmus, A.; Dutta, P.; Pollari, M.; Mäkinen, K. Interplay of HCPro and CP in the regulation of potato virus A RNA expression and encapsidation. Viruses 2022, 14, 1233. [Google Scholar] [CrossRef]
- Nunna, H.; Qu, F.; Tatineni, S. P3 and NIa-Pro of turnip mosaic virus are independent elicitors of superinfection exclusion. Viruses 2023, 15, 1459. [Google Scholar] [CrossRef] [PubMed]
- Bera, S.; Arena, G.D.; Ray, S.; Flannigan, S.; Casteel, C.L. The potyviral protein 6K1 reduces plant proteases activity during turnip mosaic virus infection. Viruses 2022, 14, 1341. [Google Scholar] [CrossRef] [PubMed]
- Sorel, M.; García, J.A.; German-Retana, S. The Potyviridae cylindrical inclusion helicase: a key multipartner and multifunctional protein. Molecular plant-microbe interactions 2014, 27, 215–226. [Google Scholar] [CrossRef]
- Zhang, H.; Cheng, G.; Yang, Z.; Wang, T.; Xu, J. Identification of sugarcane host factors interacting with the 6K2 protein of the sugarcane mosaic virus. International Journal of Molecular Sciences 2019, 20, 3867. [Google Scholar] [CrossRef]
- Andrade, F.A.d.; Luna-Aragão, M.A.d.; Ferreira, J.D.C.; Souza, F.F.; Rocha Oliveira, A.C.d.; Costa, A.F.d.; Aragão, F.J.L.; Santos-Silva, C.A.d.; Benko-Iseppon, A.M.; Pandolfi, V. Deciphering Cowpea Resistance to Potyvirus: Assessment of eIF4E Gene Mutations and Their Impact on the eIF4E-VPg Protein Interaction. Viruses 2025, 17, 1050. [Google Scholar] [CrossRef]
- Zhuang, X.; Zhang, W.; Xu, S.; Yang, W.; Yin, J.; Zhou, T.; Kundu, J.K.; Xu, K. Protease activity of NIa-Pro determines systemic pathogenicity of clover yellow vein virus. Virology 2025, 604, 110417. [Google Scholar] [CrossRef]
- Rajamaki, M.-L.; Valkonen, J.P. Control of nuclear and nucleolar localization of nuclear inclusion protein a of picorna-like Potato virus A in Nicotiana species. The Plant Cell 2009, 21, 2485–2502. [Google Scholar] [CrossRef]
- Ge, L.; Jia, M.; Shan, H.; Gao, W.; Jiang, L.; Cui, H.; Cheng, X.; Uzest, M.; Zhou, X.; Wang, A. Viral RNA polymerase as a SUMOylation decoy inhibits RNA quality control to promote potyvirus infection. Nature Communications 2025, 16, 157. [Google Scholar] [CrossRef] [PubMed]
- Shen, W.; Shi, Y.; Dai, Z.; Wang, A. The RNA-dependent RNA polymerase NIb of potyviruses plays multifunctional, contrasting roles during viral infection. Viruses 2020, 12, 77. [Google Scholar] [CrossRef] [PubMed]
- Hervás, M.; Ciordia, S.; Navajas, R.; García, J.A.; Martínez-Turiño, S. Common and strain-specific post-translational modifications of the potyvirus Plum pox virus coat protein in different hosts. Viruses 2020, 12, 308. [Google Scholar] [CrossRef]
- Brodersen, P.; Sakvarelidze-Achard, L.; Bruun-Rasmussen, M.; Dunoyer, P.; Yamamoto, Y.Y.; Sieburth, L.; Voinnet, O. Widespread translational inhibition by plant miRNAs and siRNAs. Science 2008, 320, 1185–1190. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Zhang, L.; Yang, Y.; Schmid, M.; Wang, Y. miRNA mediated regulation and interaction between plants and pathogens. International Journal of Molecular Sciences 2021, 22, 2913. [Google Scholar] [CrossRef]
- Riffo-Campos, Á.L.; Riquelme, I.; Brebi-Mieville, P. Tools for sequence-based miRNA target prediction: what to choose? International journal of molecular sciences 2016, 17. [Google Scholar] [CrossRef]
- Ashraf, M.A.; Ali, B.; Brown, J.K.; Shahid, I.; Yu, N. In silico identification of cassava genome-encoded MicroRNAs with predicted potential for targeting the ICMV-Kerala begomoviral pathogen of cassava. Viruses 2023, 15, 486. [Google Scholar] [CrossRef]
- Ashraf, M.A.; Shahid, I.; Brown, J.K.; Yu, N. An Integrative Computational Approach for Identifying Cotton Host Plant MicroRNAs with Potential to Abate CLCuKoV-Bur Infection. Viruses 2025, 17, 399. [Google Scholar] [CrossRef]
- Latif, M.F.; Tan, J.; Zhang, W.; Yang, W.; Zhuang, T.; Lu, W.; Qiu, Y.; Du, X.; Zhuang, X.; Zhou, T. Transgenic expression of artificial microRNA targeting soybean mosaic virus P1 gene confers virus resistance in plant. Transgenic Research 2024, 33, 149–157. [Google Scholar] [CrossRef]
- Al-Roshdi, M.R.; Ammara, U.; Khan, J.; Al-Sadi, A.M.; Shahid, M.S. Artificial microRNA-mediated resistance against Oman strain of tomato yellow leaf curl virus. Frontiers in Plant Science 2023, 14, 1164921. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, A.C.; Bovolenta, L.A.; Nachtigall, P.G.; Herkenhoff, M.E.; Lemke, N.; Pinhal, D. Combining results from distinct microRNA target prediction tools enhances the performance of analyses. Frontiers in genetics 2017, 8, 59. [Google Scholar] [CrossRef]
- Ghoshal, A.; Shankar, R.; Bagchi, S.; Grama, A.; Chaterji, S. MicroRNA target prediction using thermodynamic and sequence curves. BMC genomics 2015, 16, 999. [Google Scholar] [CrossRef]
- Thody, J.; Moulton, V.; Mohorianu, I. PAREameters: a tool for computational inference of plant miRNA–mRNA targeting rules using small RNA and degradome sequencing data. Nucleic Acids Research 2020, 48, 2258–2270. [Google Scholar] [CrossRef] [PubMed]
- Pinzón, N.; Li, B.; Martinez, L.; Sergeeva, A.; Presumey, J.; Apparailly, F.; Seitz, H. microRNA target prediction programs predict many false positives. Genome research 2017, 27, 234–245. [Google Scholar] [CrossRef]
- Kertesz, M.; Iovino, N.; Unnerstall, U.; Gaul, U.; Segal, E. The role of site accessibility in microRNA target recognition. Nature genetics 2007, 39, 1278–1284. [Google Scholar] [CrossRef]
- Golyshev, V.; Pyshnyi, D.; Lomzov, A. Calculation of energy for RNA/RNA and DNA/RNA duplex formation by molecular dynamics simulation. Molecular Biology 2021, 55, 927–940. [Google Scholar] [CrossRef]
- Chai, M.; Wu, X.; Liu, J.; Fang, Y.; Luan, Y.; Cui, X.; Zhou, X.; Wang, A.; Cheng, X. P3N-PIPO interacts with P3 via the shared N-terminal domain to recruit viral replication vesicles for cell-to-cell movement. Journal of Virology 2020, 94. [Google Scholar] [CrossRef]
- Luan, H.; Shine, M.; Cui, X.; Chen, X.; Ma, N.; Kachroo, P.; Zhi, H.; Kachroo, A. The potyviral P3 protein targets eukaryotic elongation factor 1A to promote the unfolded protein response and viral pathogenesis. Plant Physiology 2016, 172, 221–234. [Google Scholar] [CrossRef] [PubMed]
- Saha, K.; Ihearahu, O.C.; Agbor, V.E.; Evans, T.; Naitchede, L.H.S.; Ray, S.; Ude, G. In Silico Genome-Wide Profiling of Conserved miRNAs in AAA, AAB, and ABB Groups of Musa spp.: Unveiling MicroRNA-Mediated Drought Response. International Journal of Molecular Sciences 2025. [Google Scholar] [CrossRef] [PubMed]
- Patel, P.; Yadav, K.; Srivastava, A.K.; Suprasanna, P.; Ganapathi, T.R. Overexpression of native Musa-miR397 enhances plant biomass without compromising abiotic stress tolerance in banana. Scientific Reports 2019, 9, 16434. [Google Scholar] [CrossRef] [PubMed]
- Dash, P.K.; Gupta, P.; Pradhan, S.K.; Shasany, A.K.; Rai, R. Analysis of homologous regions of small RNAs MIR397 and MIR408 reveals the conservation of microsynteny among rice crop-wild relatives. Cells 2022, 11, 3461. [Google Scholar] [CrossRef]
- Huang, S.; Zhou, J.; Gao, L.; Tang, Y. Plant miR397 and its functions. Functional Plant Biology 2020, 48, 361–370. [Google Scholar] [CrossRef]
- Xue, M.; Arvy, N.; German-Retana, S. The mystery remains: How do potyviruses move within and between cells? Molecular Plant Pathology 2023, 24, 1560–1574. [Google Scholar] [CrossRef]
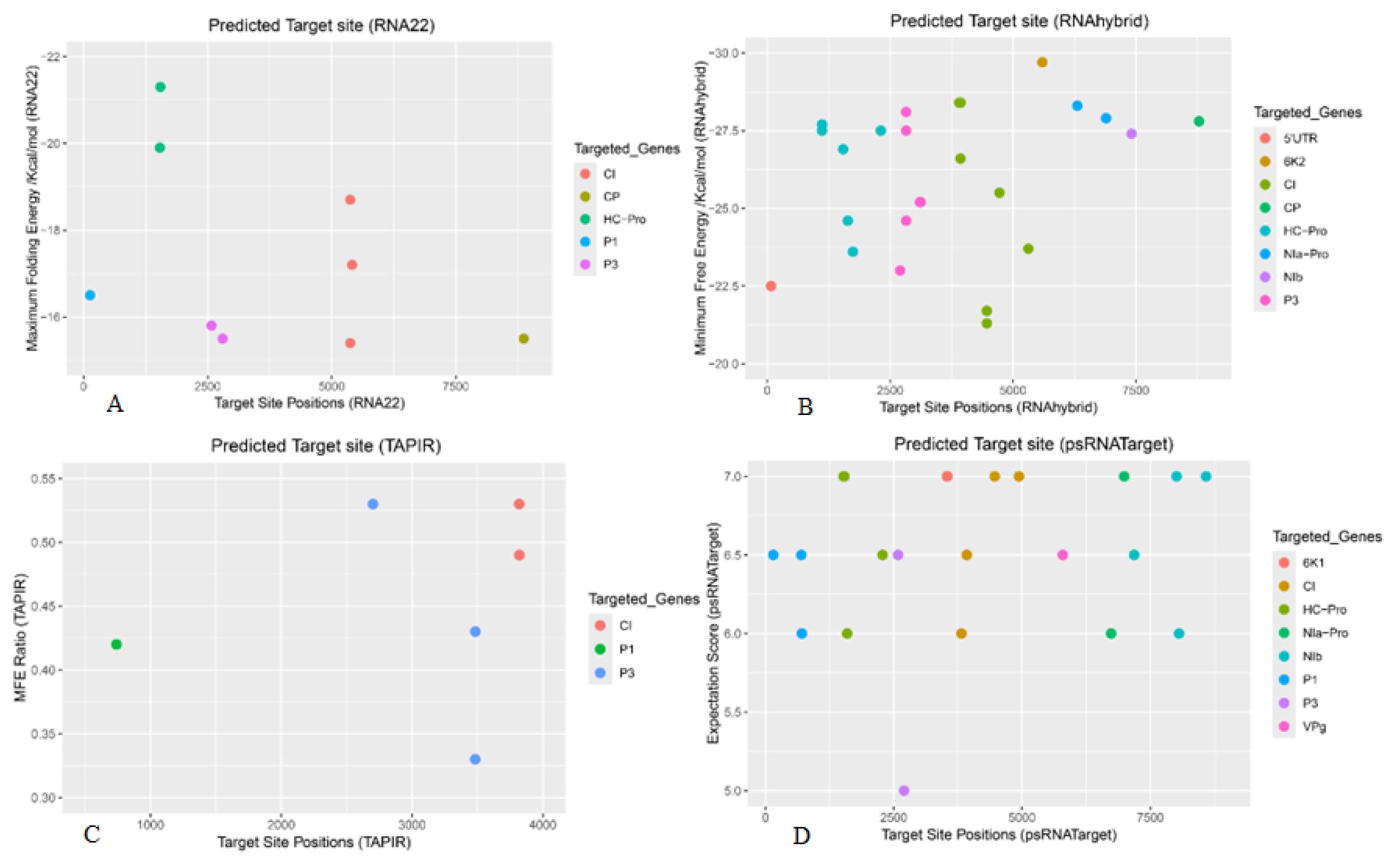
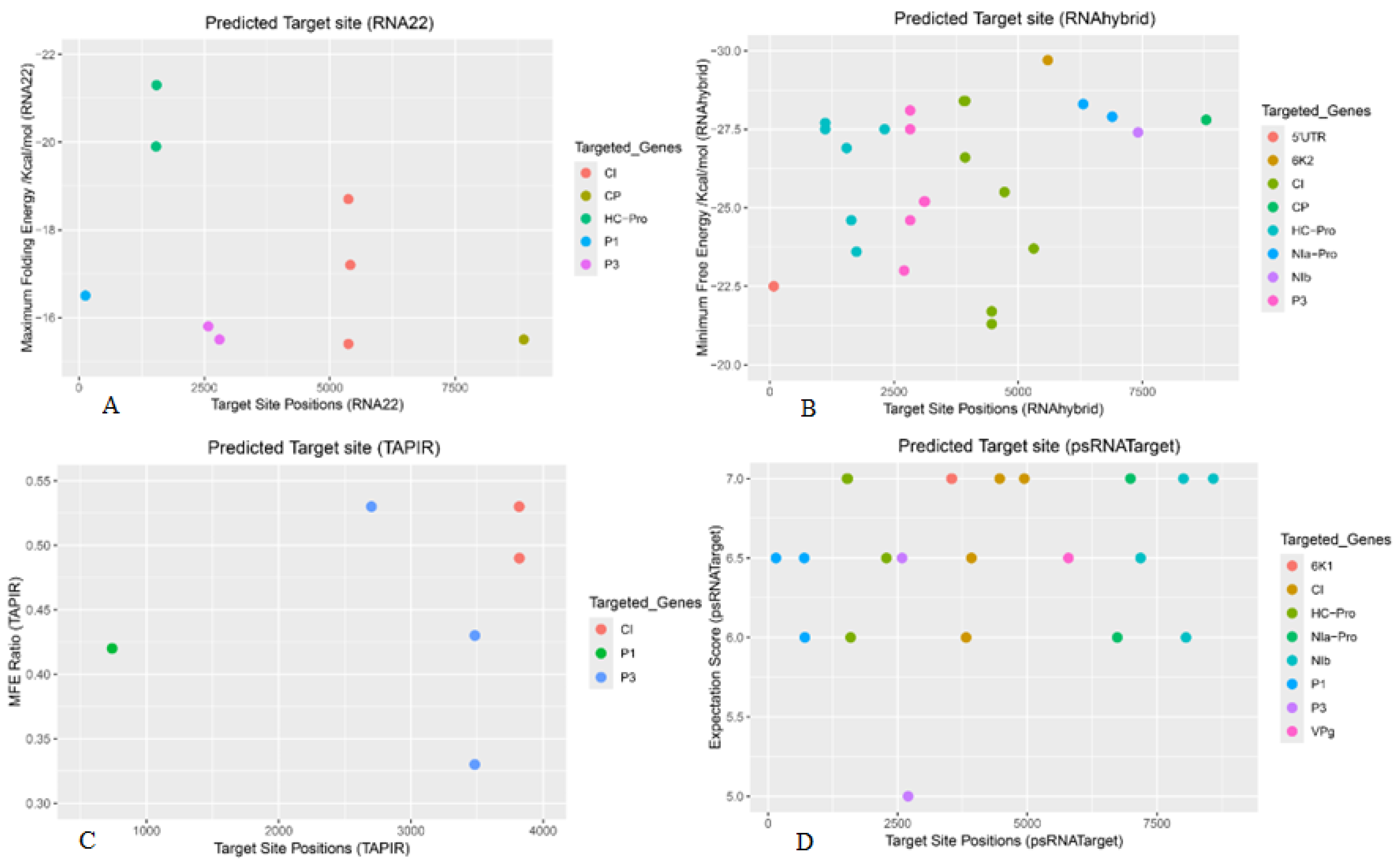
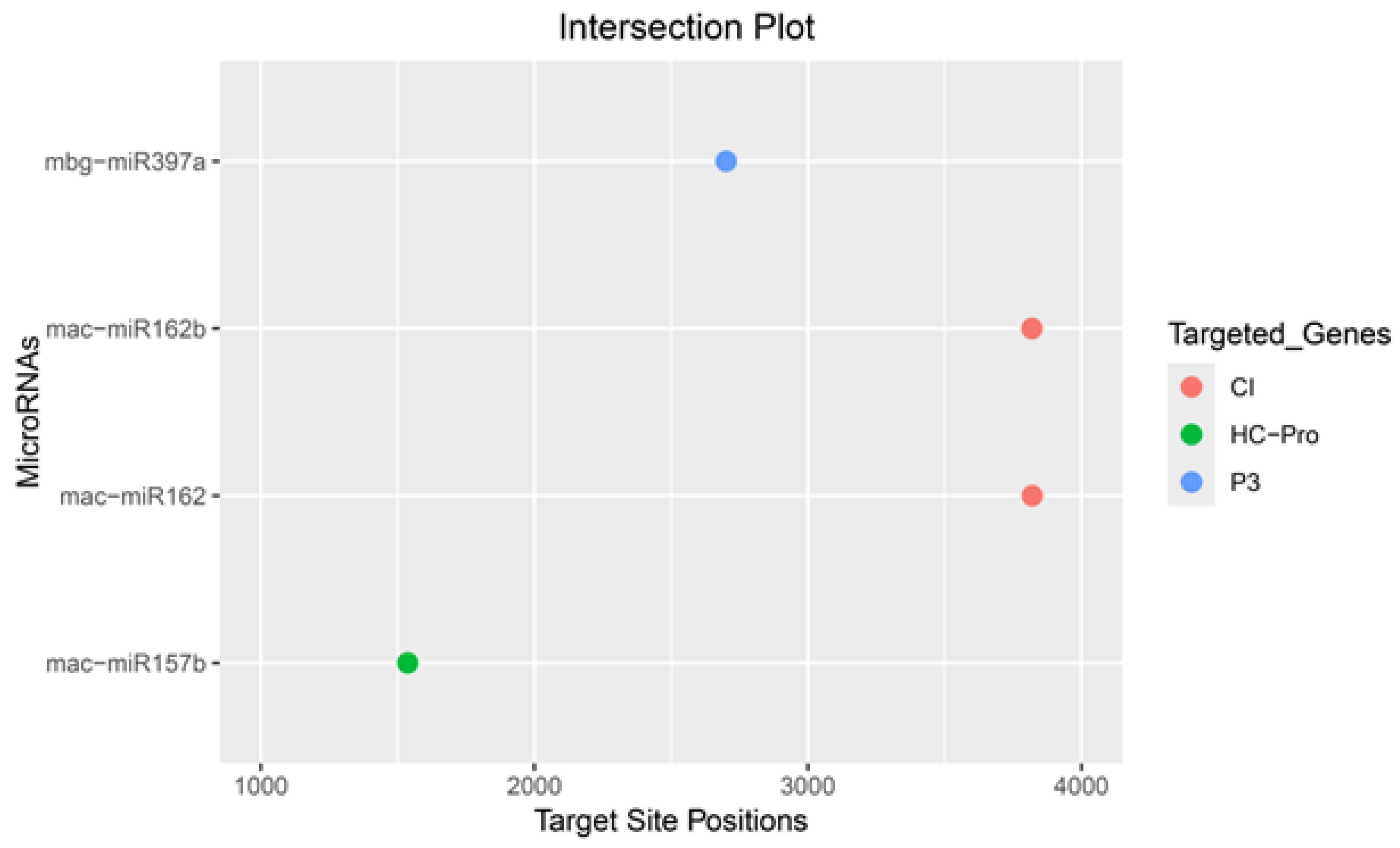
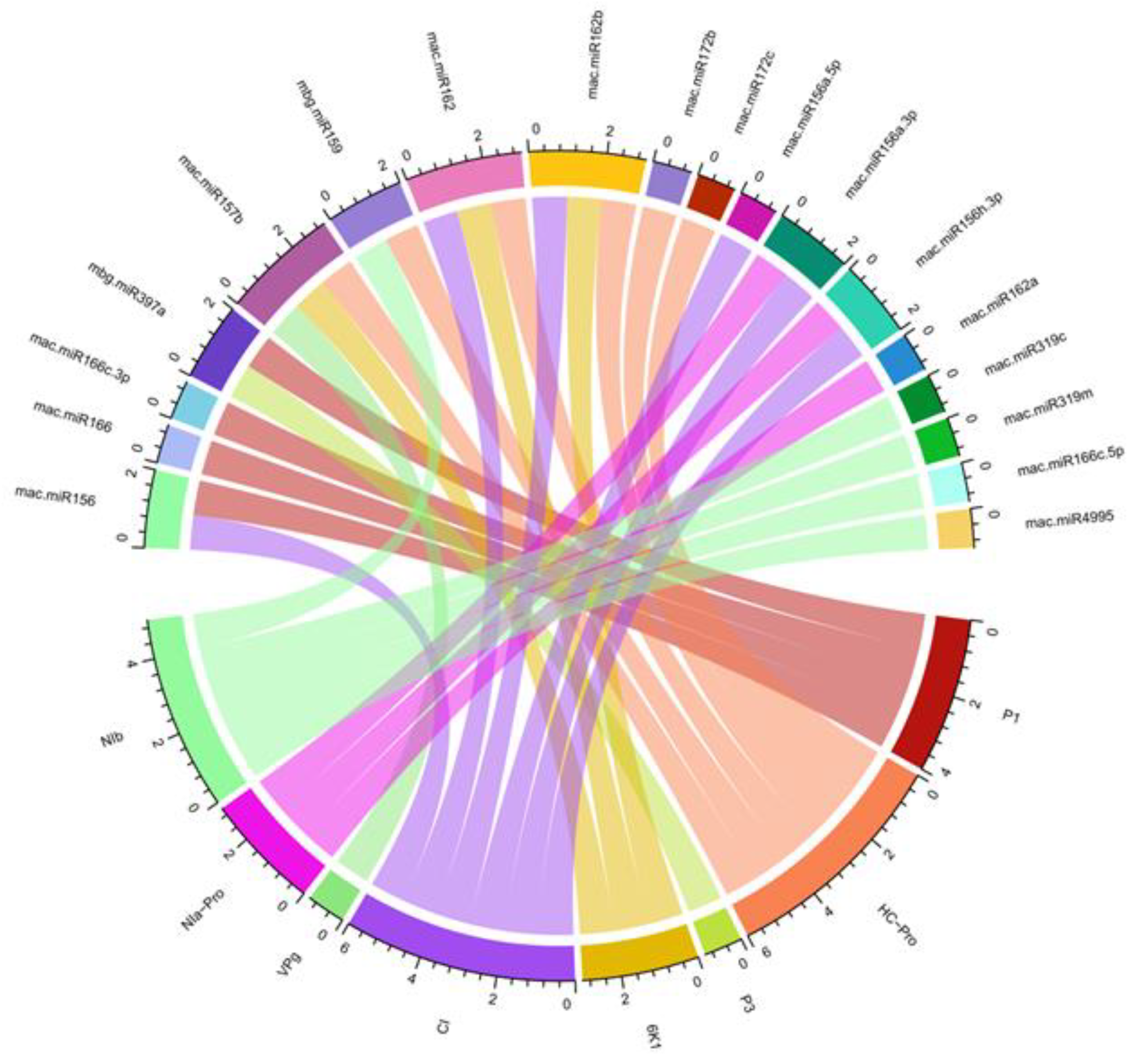
| BBrMV Gene | RNA22 | RNAhybrid | TAPIR | psRNATarget |
|---|---|---|---|---|
| P1 | mbg-miR399a1 | mac-miR4995 | mac-miR156, mac-miR166, mac-miR166c-3p and mbg-miR397a | |
| HC-Pro | mac-miR157b and mac-miR160 (a, g-5p) | mac-miR157b, mac-miR160g, mac-miR166 (b, c-5p), mac-miR172 (b, c) and mac-miR4995 | mbg-miR157b, mbg-miR159, mac-miR162, mac-miR162b and mac-miR172 (b, c) | |
| P3 | mac-miR166 (b, c-5p) and mac-miR4995 | mac-miR156a-5p, mac-miR159, mac-miR319 (c, m) and mac-miR397a | mac-miR172 (b, c) and mbg-miR397a | mbg-miR397a |
| 6K1 | mac-miR157b, mac-miR162 mac-miR162b | |||
| CI | mac-miR166, mac-miR166 (b, c-5p) and mac-miR166c-3p | mac-miR156 (a-3p, h-3p), mac-miR156 (d, g), mac-miR157 (b-5p), mac-miR160 (a, g-5p), mac-miR162, mac-miR162a, mac-miR162b and mbg-miR399a1 | mac-miR162 and mac-miR162b | mac-miR156, mac-miR156a-5p, mac-miR156 (a-3p, h-3p), mac-miR162 and mac-miR162b |
| 6K2 | mac-miR164e | |||
| VPg | mac-miR157b | |||
| NIa-Pro | mac-miR169h, mac-miR166 and mac-miR166c-3p | mac-miR156 (a-3p, h-3p) and mac-miR162a | ||
| NIb-Pro | mac-miR169h, mac-miR166 and mac-miR166c-3p | mbg-miR159, mac-miR319c, mac-miR319m, mac-miR166c-5p and mac-miR4995 | ||
| CP | mbg-miR399a | mac-miR167 (c, d) | ||
| 5’UTR | mac-miR156 |
| Banana miRNAs ID |
Site/Gene RNA22 |
Site/Gene RNAhybrid |
Site/Gene TAPIR |
Site/Gene psRNATarget |
MFE * RNA22 |
MFE** RNAhybrid |
MFE** Ratio TAPIR |
Expectation psRNATarget |
|---|---|---|---|---|---|---|---|---|
| mac-miR157b | 1537 (HC-Pro) | 1544(HC-Pro) | 1537(HC-Pro) | −19.90 | −26.90 | 7.00 | ||
| mac-miR162 | 4725(CI) | 3820(CI) | 3820(CI) | −15.30 | −25.50 | 0.49 | 6.00 | |
| mac-miR162b | 4725(CI) | 3820(CI) | 3820(CI) | −15.30 | −25.50 | 0.49 | 6.00 | |
| mbg-miR397a | 2703(P3) | 2703(P3) | 2701(P3) | −23.00 | 0.50 | 5.00 |
| Banana miRNAs ID |
Mature Sequence (5′–3′) |
Predicted Targets ORF(s) |
Binding Sites (nt) |
Mode of Inhibition |
|---|---|---|---|---|
| mac-miR157b | GCUCUCUAUGCUUCUGUCAUCA | HC-Pro | 1537-1559 | Cleavage |
| mac-miR162 | UCGAUAAACCGCUGCGUCCA | CI | 3820-3839 | Cleavage |
| mac-miR162b | UCGAUAAACCGCUGCGUCCAG | CI | 3820-3839 | Cleavage |
| mbg-miR397a | UCAUUGAGUGCAGCGUUGAUG | P3 | 2701-2721 | Cleavage |
| miRNA ID | miRNA–mRNA Sequence (5′–3′) |
ΔG Duplex (Kcal/mol) |
ΔG Binding (Kcal/mol) |
Genomic Coordinates |
|---|---|---|---|---|
| mac-miR157b | 5′ GCUCUCUAUGCUUCUGUCAUCA 3′ 5′ AATTTTCTGAAGCATCAGAGAGC 3′ |
−21.18 | −13.39 | 1537-1559 |
| mac-miR162 | 5′ UCGAUAAACCGCUGCGUCCA 3′ 5′ CGTGTTGTAGCGCATTATCGA 3′ |
−17.84 | −14.86 | 3820-3839 |
| mac-miR162b | 5′ UCGAUAAACCGCUGCGUCCAG 3′ 5′ CGTGTTGTAGCGCATTATCGA3′ |
−17.99 | −14.82 | 3820-3839 |
| mbg-miR397a | 5′ UCAUUGAGUGCAGCGUUGAUG 3′ 5′ TCACAACATTGCATTCATTGG 3′ |
−19.35 | −17.25 | 2701-2721 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




