Submitted:
06 January 2026
Posted:
07 January 2026
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
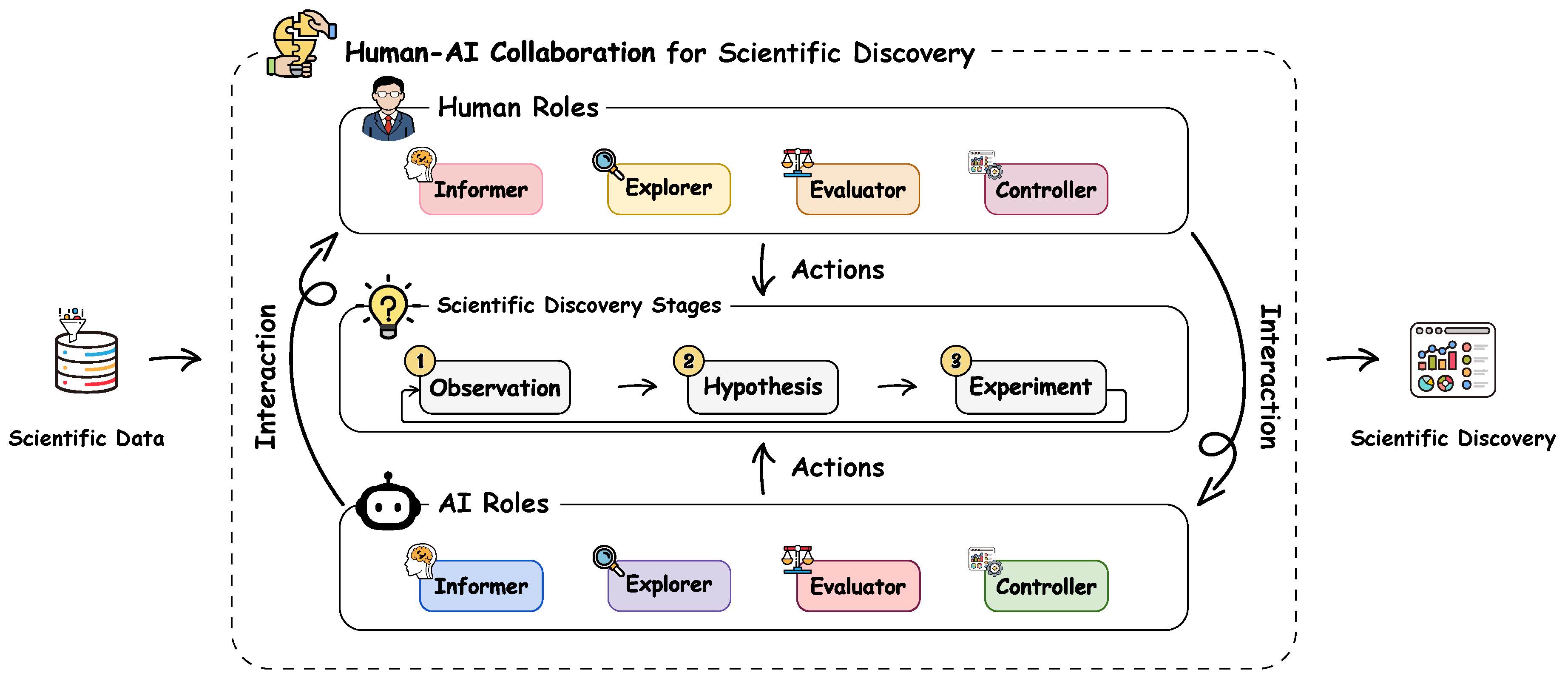
- We presented a comprehensive survey that systematically reviews human-AI collaboration specifically in scientific discovery.
- We introduced a novel taxonomy that defines the four roles of human and AI and characterized their distinct collaboration patterns across the three stages of scientific discovery.
- We identified critical challenges and future pathways for building human-AI partnerships in the scientific discovery process.
2. Methodology
2.1. Paper Collection
2.2. Paper Coding
3. Taxonomy
3.1. Roles of Human and AI
- ⋄Informer. The Informer synthesizes, distills, or articulates key information, insights, or constraints from raw data or intermediate analyses to guide the actions of other roles. For instance, the AI-Informer in THALIS extracts temporal patterns from longitudinal symptom records in cancer therapy (Floricel et al. 2022). This provides a summarized trajectory view for experts to analyze patient responses to treatment. Similarly, in ISHMAP for Mars rover operations (Wright et al. 2023), the human-Informer marks instrument states and operational events on the telemetry timeline. The AI uses these annotations to reduce false alarms during state changes and to highlight unexpected signals.
- ⋄Explorer. The Explorer operates within the space of data patterns, hypotheses, or experimental designs to explore promising candidates or directions. Compared with Informer, whose output provides low-level data insights, the Explorer directly generates candidates tailored to the specific needs of each stage in the scientific discovery process. For instance, in the hAE interface (Pratiush et al. 2024), the AI-Explorer searches the parameter space to select the next experimental conditions for electron microscopy. ChemVA (Sabando et al. 2021) enables the human-Explorer to interactively navigate a projected chemical space to identify molecular targets.
- ⋄Evaluator. Once artifacts are proposed, their scientific merit must be rigorously evaluated and even refined. The Evaluator assesses observed patterns, hypotheses, or experimental designs based on predefined criteria, evidence, and domain constraints, revising them as necessary to meet quality standards. For instance, in RetroLens (Shi et al. 2023a), chemists serve as the human-Evaluator. Specifically, they can assess AI-predicted synthetic routes for chemical feasibility or refine the synthetic steps by themselves.
- ⋄Controller. The Controller oversees the scientific discovery workflow to ensure correct and constraint-compliant execution, intervening when necessary to adjust procedures and handle runtime exceptions. This role is central to BIA (Xin et al. 2024), where the AI-Controller orchestrates the execution of complex bioinformatics toolchains, dynamically handling errors and modifying the workflow logic to ensure successful completion.
3.2. Common Collaboration Patterns Within Each Stage of Scientific Discovery
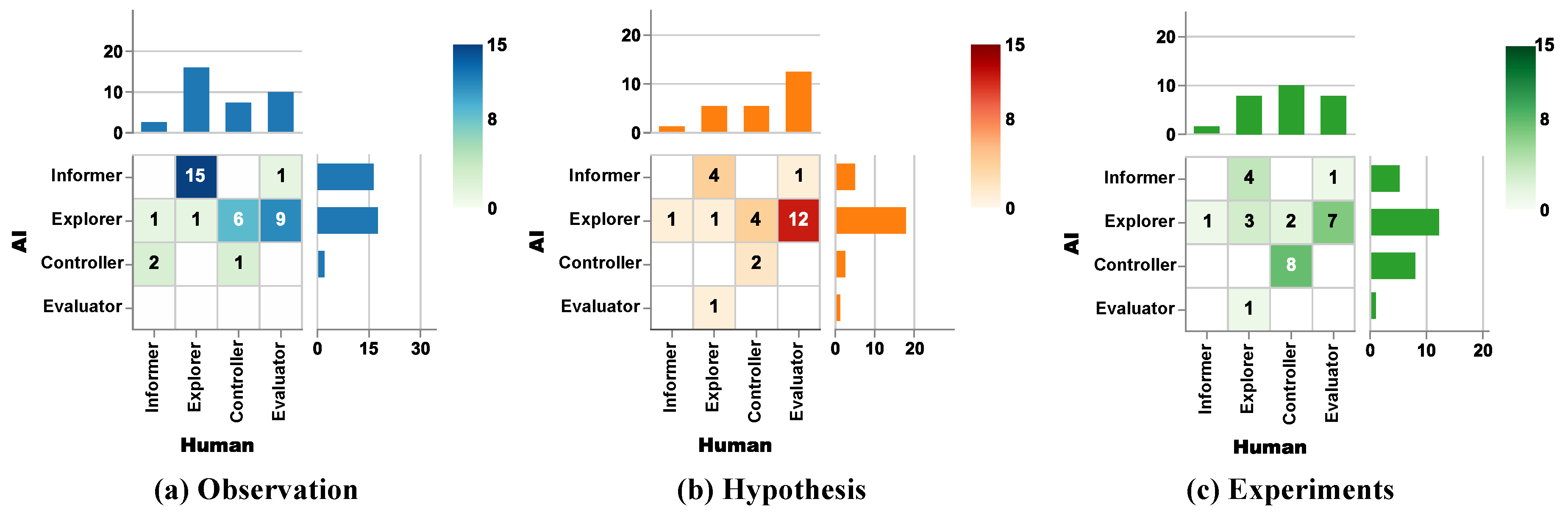
3.2.1. Observation Stage
3.2.2. Hypothesis Stage
3.2.3. Experiment Stage
3.2.4. Role Differences Across Three Stages
4. Discussion
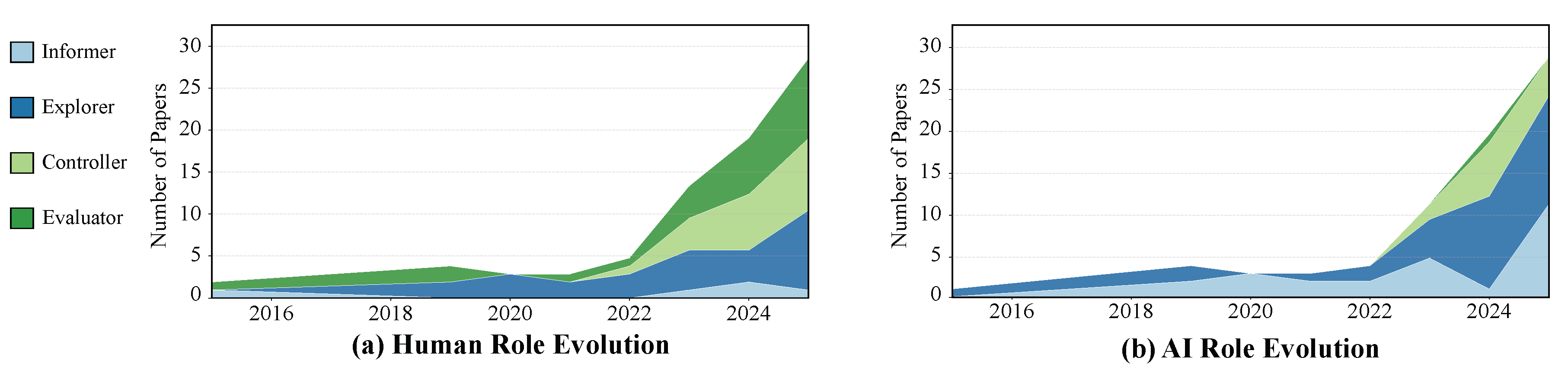
4.1. From Asymmetric Growth to Symbiotic Evolution
4.2. Generative Interfaces for Supporting Human Involvement
4.3. Adaptive Role Assignments Between Human and AI
4.4. Empowering Embodied AI in Scientific Experiments
4.5. Long-Term Implications for Leveraging AI in Scientific Discovery
5. Conclusion
Limitations
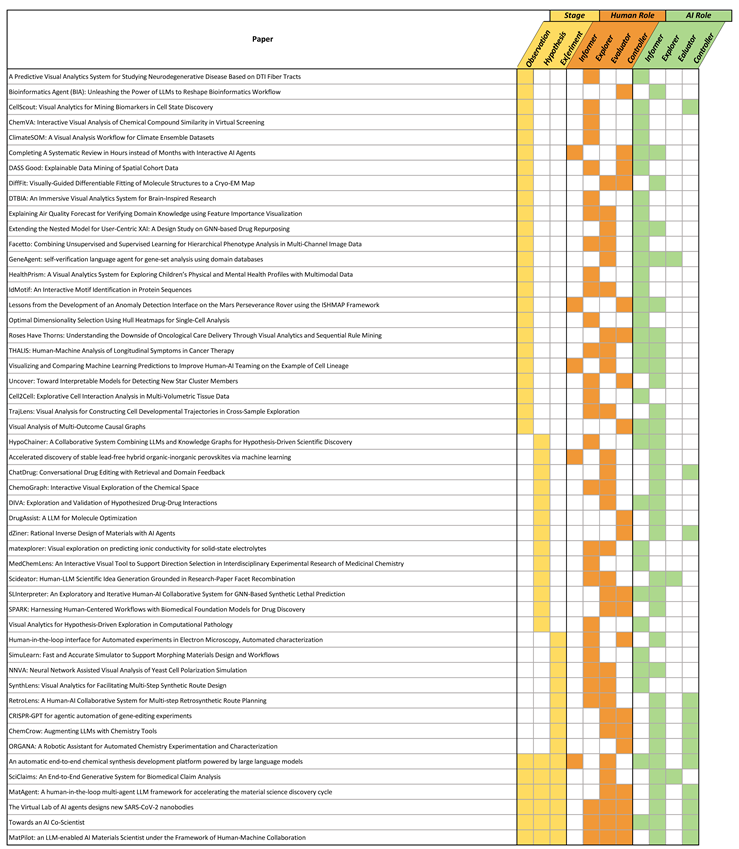
Appendix A. Coding Results
References
- Mehrad Ansari, Jeffrey Watchorn, Carla E. Brown, and Joseph S Brown. 2024. dZiner: Rational Inverse Design of Materials with AI Agents. In AI for Accelerated Materials Design - NeurIPS 2024.
- Adib Bazgir, Rama chandra Praneeth Madugula, and Yuwen Zhang. 2025. MatAgent: A human-in-the-loop multi-agent LLM framework for accelerating the material science discovery cycle. In AI for Accelerated Materials Design - ICLR 2025.
- Daniil A. Boiko, Robert MacKnight, Ben Kline, and Gabe Gomes. 2023. Autonomous chemical research with large language models. Nature, 624(7992):570–578. [CrossRef]
- Andres M. Bran, Sam Cox, Oliver Schilter, Carlo Baldassari, Andrew D. White, and Philippe Schwaller. 2024. Augmenting large language models with chemistry tools. Nature Machine Intelligence, 6(5):525–535. [CrossRef]
- Jiaqi Chen, Yanzhe Zhang, Yutong Zhang, Yijia Shao, and Diyi Yang. 2025. Generative interfaces for language models. arXiv preprint arXiv:2508.19227.
- Yiming Chen, Chen Zhang, Danqing Luo, Luis Fernando D’Haro, Robby Tan, and Haizhou Li. 2024. Unveiling the Achilles’ Heel of NLG Evaluators: A Unified Adversarial Framework Driven by Large Language Models. In Findings of the Association for Computational Linguistics: ACL 2024, pages 1359–1375.
- A. Corvò, H. S. Garcia Caballero, M. A. Westenberg, M. A. van Driel, and J. J. van Wijk. 2021. Visual Analytics for Hypothesis-Driven Exploration in Computational Pathology. IEEE Transactions on Visualization and Computer Graphics, 27(10):3851–3866. [CrossRef]
- Kourosh Darvish, Marta Skreta, Yuchi Zhao, Naruki Yoshikawa, Sagnik Som, Miroslav Bogdanovic, Yang Cao, Han Hao, Haoping Xu, Alán Aspuru-Guzik, Animesh Garg, and Florian Shkurti. 2025. ORGANA: A robotic assistant for automated chemistry experimentation and characterization. Matter, 8(2):101897. [CrossRef]
- Mengjie Fan, Jinlu Yu, Daniel Weiskopf, Nan Cao, Huai-Yu Wang, and Liang Zhou. 2025. Visual Analysis of Multi-Outcome Causal Graphs. IEEE Transactions on Visualization and Computer Graphics, 31(1):656–666. [CrossRef]
- Carla Floricel, Nafiul Nipu, Mikayla Biggs, Andrew Wentzel, Guadalupe Canahuate, Lisanne Van Dijk, Abdallah Mohamed, C.David Fuller, and G.Elisabeta Marai. 2022. THALIS: Human-Machine Analysis of Longitudinal Symptoms in Cancer Therapy. IEEE Transactions on Visualization and Computer Graphics, 28(1):151–161.
- Carla Floricel, Andrew Wentzel, Abdallah Mohamed, C.David Fuller, Guadalupe Canahuate, and G.Elisabeta Marai. 2024. Roses Have Thorns: Understanding the Downside of Oncological Care Delivery Through Visual Analytics and Sequential Rule Mining. IEEE Transactions on Visualization and Computer Graphics, 30(1):1227–1237. [CrossRef]
- Juraj Gottweis, Wei-Hung Weng, Alexander Daryin, Tao Tu, Anil Palepu, Petar Sirkovic, Artiom Myaskovsky, Felix Weissenberger, Keran Rong, Ryutaro Tanno, and 1 others. 2025. Towards an AI Co-Scientist. arXiv preprint arXiv:2502.18864.
- Subhashis Hazarika, Haoyu Li, Ko-Chih Wang, Han-Wei Shen, and Ching-Shan Chou. 2020. NNVA: Neural Network Assisted Visual Analysis of Yeast Cell Polarization Simulation. IEEE Transactions on Visualization and Computer Graphics, 26(1):34–44.
- Juyeon Heo, Miao Xiong, Christina Heinze-Deml, and Jaya Narain. 2025. Do LLMs estimate uncertainty well in instruction-following? In The Thirteenth International Conference on Learning Representations.
- Jiayi Hong, Ross Maciejewski, Alain Trubuil, and Tobias Isenberg. 2024. Visualizing and Comparing Machine Learning Predictions to Improve Human-AI Teaming on the Example of Cell Lineage. IEEE Transactions on Visualization and Computer Graphics, 30(4):1956–1969. [CrossRef]
- Chen Huang, Yang Deng, Wenqiang Lei, Jiancheng Lv, Tat-Seng Chua, and Jimmy Huang. 2025. How to Enable Effective Cooperation Between Humans and NLP Models: A Survey of Principles, Formalizations, and Beyond. In Proceedings of the 63rd Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers), pages 466–488.
- Peter Jansen, Oyvind Tafjord, Marissa Radensky, Pao Siangliulue, Tom Hope, Bhavana Dalvi Mishra, Bodhisattwa Prasad Majumder, Daniel S Weld, and Peter Clark. 2025. CodeScientist: End-to-end semi-automated scientific discovery with code-based experimentation. pages 13370–13467.
- Haejin Jeong, Hyoung-oh Jeong, Semin Lee, and Won-Ki Jeong. 2025. Optimal Dimensionality Selection Using Hull Heatmaps for Single-Cell Analysis. Computer Graphics Forum, 44(6):e70151.
- Haoran Jiang, Shaohan Shi, Yunjie Yao, Chang Jiang, and Quan Li. 2025a. HypoChainer: a Collaborative System Combining LLMs and Knowledge Graphs for Hypothesis-Driven Scientific Discovery. IEEE Transactions on Visualization and Computer Graphics, pages 1–11.
- Haoran Jiang, Shaohan Shi, Shuhao Zhang, Jie Zheng, and Quan Li. 2025b. SLInterpreter: An Exploratory and Iterative Human-AI Collaborative System for GNN-Based Synthetic Lethal Prediction. IEEE Transactions on Visualization and Computer Graphics, 31(1):919–929. [CrossRef]
- Zhihan Jiang, Handi Chen, Rui Zhou, Jing Deng, Xinchen Zhang, Running Zhao, Cong Xie, Yifang Wang, and Edith C.H. Ngai. 2024. HealthPrism: A Visual Analytics System for Exploring Children’s Physical and Mental Health Profiles with Multimodal Data . IEEE Transactions on Visualization & Computer Graphics, 30(01):1205–1215. [CrossRef]
- Tabassum Kakar, Xiao Qin, Elke A. Rundensteiner, Lane Harrison, Sanjay K. Sahoo, and Suranjan De. 2019. DIVA: Exploration and Validation of Hypothesized Drug-Drug Interactions. In Computer Graphics Forum, volume 38, pages 95–106. [CrossRef]
- Bharat Kale, Austin Clyde, Maoyuan Sun, Arvind Ramanathan, Rick Stevens, and Michael E. Papka. 2023. ChemoGraph: Interactive Visual Exploration of the Chemical Space. volume 42, pages 13–24. [CrossRef]
- Yuya Kawakami, Daniel Cayan, Dongyu Liu, and Kwan-Liu Ma. 2025. ClimateSOM: a Visual Analysis Workflow for Climate Ensemble Datasets. IEEE Transactions on Visualization and Computer Graphics, pages 1–11.
- Robert Krueger, Johanna Beyer, Won-Dong Jang, Nam Wook Kim, Artem Sokolov, Peter K. Sorger, and Hanspeter Pfister. 2020. Facetto: Combining Unsupervised and Supervised Learning for Hierarchical Phenotype Analysis in Multi-Channel Image Data. IEEE Transactions on Visualization and Computer Graphics, 26(1):227–237. [CrossRef]
- Bum Chul Kwon, Simona Rabinovici-Cohen, Beldine Moturi, Ruth Mwaura, Kezia Wahome, Oliver Njeru, Miguel Shinyenyi, Catherine Wanjiru, Sekou Remy, William Ogallo, Itai Guez, Partha Suryanarayanan, Joseph Morrone, Shreyans Sethi, Seung-Gu Kang, Tien Huynh, Kenney Ng, Diwakar Mahajan, Hongyang Li, and 4 others. 2024. SPARK: harnessing human-centered workflows with biomedical foundation models for drug discovery. IJCAI ’24.
- Shengchao Liu, Jiongxiao Wang, Yijin Yang, Chengpeng Wang, Ling Liu, Hongyu Guo, and Chaowei Xiao. 2024. Conversational Drug Editing Using Retrieval and Domain Feedback. In The Twelfth International Conference on Learning Representations (ICLR).
- Shuaihua Lu, Qionghua Zhou, Yixin Ouyang, Yilv Guo, Qiang Li, and Jinlan Wang. 2018. Accelerated Discovery of Stable Lead-Free Hybrid Organic-Inorganic Perovskites via Machine Learning. Nature Communications, 9(1):3405.
- Deng Luo, Zainab Alsuwaykit, Dawar Khan, Ondřej Strnad, Tobias Isenberg, and Ivan Viola. 2025. DiffFit: Visually-Guided Differentiable Fitting of Molecule Structures to a Cryo-EM Map. IEEE Transactions on Visualization and Computer Graphics, 31(1):558–568. [CrossRef]
- Stéphane Luro, Laurie Potvin-Trottier, Burak Okumus, and Johan Paulsson. 2020. Isolating live cells after high-throughput, long-term time-lapse microscopy. Nature Methods, 17:93–100. [CrossRef]
- Yubo Ma, Zhibin Gou, Junheng Hao, Ruochen Xu, Shuohang Wang, Liangming Pan, Yujiu Yang, Yixin Cao, and Aixin Sun. 2024. SciAgent: Tool-augmented language models for scientific reasoning. In Proceedings of the 2024 Conference on Empirical Methods in Natural Language Processing, pages 15701–15736.
- Vikram Mohanty, Jude Lim, and Kurt Luther. 2025. What Lies Beneath? Exploring the Impact of Underlying AI Model Updates in AI-Infused Systems. In Proceedings of the 2025 CHI Conference on Human Factors in Computing Systems, CHI ’25.
- Eric Mörth, Kevin Sidak, Zoltan Maliga, Torsten Möller, Nils Gehlenborg, Peter Sorger, Hanspeter Pfister, Johanna Beyer, and Robert Krüger. 2025. Cell2Cell: Explorative Cell Interaction Analysis in Multi-Volumetric Tissue Data. IEEE Transactions on Visualization and Computer Graphics, 31(1):569–579. [CrossRef]
- Ziqi Ni, Yahao Li, Kaijia Hu, Kunyuan Han, Ming Xu, Xingyu Chen, Fengqi Liu, Yicong Ye, and Shuxin Bai. 2024. MatPilot: An LLM-Enabled AI Materials Scientist Under the Framework of Human-Machine Collaboration. arXiv preprint arXiv:2411.08063.
- Raúl Ortega and Jose Manuel Gomez-Perez. 2025. SciClaims: An end-to-end generative system for biomedical claim analysis. In Proceedings of the 2025 Conference on Empirical Methods in Natural Language Processing: System Demonstrations, pages 141–154.
- Reshika Palaniyappan Velumani, Meng Xia, Jun Han, Chaoli Wang, ALEXIS K LAU, and Huamin Qu. 2022. AQX: Explaining Air Quality Forecast for Verifying Domain Knowledge using Feature Importance Visualization. In Proceedings of the 27th International Conference on Intelligent User Interfaces, page 720–733.
- Ji Hwan Park, Vikash Prasad, Sydney Newsom, Fares Najar, and Rakhi Rajan. 2024. IdMotif: An Interactive Motif Identification in Protein Sequences. IEEE Computer Graphics and Applications, 44(3):114–125. [CrossRef]
- Utkarsh Pratiush, Gerd Duscher, and Sergei Kalinin. 2024. Human-in-the-loop interface for Automated experiments in Electron Microscopy, Automated characterization. In AI for Accelerated Materials Design - NeurIPS 2024.
- Jiansu Pu, Hui Shao, Boyang Gao, Zhengguo Zhu, Yanlin Zhu, Yunbo Rao, and Yong Xiang. 2022. matExplorer: Visual Exploration on Predicting Ionic Conductivity for Solid-state Electrolytes. IEEE Transactions on Visualization and Computer Graphics, 28(1):65–75. [CrossRef]
- Rui Qiu, Shijie Chen, Yu Su, Po-Yin Yen, and Han Wei Shen. 2025. Completing a systematic review in hours instead of months with interactive AI agents. In Proceedings of the 63rd Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers), pages 31559–31593.
- Yuanhao Qu, Kaixuan Huang, Ming Yin, Kanghong Zhan, Dyllan Liu, Di Yin, Henry C. Cousins, William A. Johnson, Xiaotong Wang, Mihir Shah, and 1 others. 2025. CRISPR-GPT for agentic automation of gene-editing experiments. Nature Biomedical Engineering.
- Niroop Channa Rajashekar, Yeo Eun Shin, Yuan Pu, Sunny Chung, Kisung You, Mauro Giuffre, Colleen E Chan, Theo Saarinen, Allen Hsiao, Jasjeet Sekhon, Ambrose H Wong, Leigh V Evans, Rene F. Kizilcec, Loren Laine, Terika Mccall, and Dennis Shung. 2024. Human-Algorithmic Interaction Using a Large Language Model-Augmented Artificial Intelligence Clinical Decision Support System. In Proceedings of the 2024 CHI Conference on Human Factors in Computing Systems, CHI ’24.
- Sebastian Ratzenböck, Verena Obermüller, Torsten Möller, João Alves, and Immanuel M. Bomze. 2023. Uncover: Toward Interpretable Models for Detecting New Star Cluster Members. IEEE Transactions on Visualization and Computer Graphics, 29(9):3855–3872.
- Chandan K. Reddy and Parshin Shojaee. 2025. Towards scientific discovery with generative AI: progress, opportunities, and challenges. In Proceedings of the Thirty-Ninth AAAI Conference on Artificial Intelligence. AAAI Press. [CrossRef]
- MohammadHossein Rezaei, Yicheng Fu, Phil Cuvin, Caleb Ziems, Yanzhe Zhang, Hao Zhu, and Diyi Yang. 2025. EgoNormia: Benchmarking Physical-Social Norm Understanding. In Findings of the Association for Computational Linguistics: ACL 2025, pages 19256–19283.
- María Virginia Sabando, Pavol Ulbrich, Matías Selzer, Jan Byška, Jan Mičan, Ignacio Ponzoni, Axel J. Soto, María Luján Ganuza, and Barbora Kozlíková. 2021. ChemVA: Interactive Visual Analysis of Chemical Compound Similarity in Virtual Screening. IEEE Transactions on Visualization and Computer Graphics, 27(2):891–901. [CrossRef]
- Samuel Schmidgall, Yusheng Su, Ze Wang, Ximeng Sun, Jialian Wu, Xiaodong Yu, Jiang Liu, Michael Moor, Zicheng Liu, and Emad Barsoum. 2025. Agent laboratory: Using LLM agents as research assistants. In Findings of the Association for Computational Linguistics: EMNLP 2025, pages 5977–6043.
- Rui Sheng, Xingbo Wang, Jiachen Wang, Xiaofu Jin, Zhonghua Sheng, Zhenxing Xu, Suraj Rajendran, Huamin Qu, and Fei Wang. 2025a. Trialcompass: Visual analytics for enhancing the eligibility criteria design of clinical trials. IEEE Transactions on Visualization and Computer Graphics, pages 1–11.
- Rui Sheng, Zelin Zang, Jiachen Wang, Yan Luo, Zixin Chen, Yan Zhou, Shaolun Ruan, and Huamin Qu. 2025b. CellScout: Visual Analytics for Mining Biomarkers in Cell State Discovery. IEEE Transactions on Visualization and Computer Graphics, pages 1–16.
- Chuhan Shi, Yicheng Hu, Shenan Wang, Shuai Ma, Chengbo Zheng, Xiaojuan Ma, and Qiong Luo. 2023a. RetroLens: A Human-AI Collaborative System for Multi-step Retrosynthetic Route Planning.
- Chuhan Shi, Fei Nie, Yicheng Hu, Yige Xu, Lei Chen, Xiaojuan Ma, and Qiong Luo. 2023b. MedChemLens: An Interactive Visual Tool to Support Direction Selection in Interdisciplinary Experimental Research of Medicinal Chemistry. IEEE Transactions on Visualization and Computer Graphics, 29(1):63–73.
- Kyle Swanson, Wesley Wu, Nash L Bulaong, John E Pak, and James Zou. 2025. The Virtual Lab of AI agents designs new SARS-CoV-2 nanobodies. Nature, 646(8085):716–723.
- Jiabin Tang, Lianghao Xia, Zhonghang Li, and Chao Huang. 2025. AI-researcher: Autonomous scientific innovation. In The Thirty-ninth Annual Conference on Neural Information Processing Systems.
- Hanchen Wang, Tianfan Fu, Yuanqi Du, Wenhao Gao, Kexin Huang, Ziming Liu, Payal Chandak, Shengchao Liu, Peter Van Katwyk, Andreea Deac, and 1 others. 2023a. Scientific discovery in the age of artificial intelligence. Nature, 620(7972):47–60. [CrossRef]
- Qianwen Wang, Kexin Huang, Payal Chandak, Marinka Zitnik, and Nils Gehlenborg. 2023b. Extending the Nested Model for User-Centric XAI: A Design Study on GNN-based Drug Repurposing. IEEE Transactions on Visualization and Computer Graphics, 29(1):1266–1276. [CrossRef]
- Qingyun Wang, Rylan Schaeffer, Fereshte Khani, Filip Ilievski, and 1 others. 2025a. Scideator: Human-LLM Scientific Idea Generation Grounded in Research-Paper Facet Recombination. Transactions on Machine Learning Research.
- Qipeng Wang, Shaolun Ruan, Rui Sheng, Yong Wang, Min Zhu, and Huamin Qu. 2025b. Trajlens: Visual analysis for constructing cell developmental trajectories in cross-sample exploration. IEEE Transactions on Visualization and Computer Graphics, pages 1–11.
- Qipeng Wang, Rui Sheng, Shaolun Ruan, Xiaofu Jin, Chuhan Shi, and Min Zhu. 2025c. SynthLens: Visual Analytics for Facilitating Multi-Step Synthetic Route Design. IEEE Transactions on Visualization and Computer Graphics, 31(10):7647–7660.
- Xingbo Wang, Samantha L. Huey, Rui Sheng, Saurabh Mehta, and Fei Wang. 2025d. SciDaSynth: Interactive Structured Data Extraction From Scientific Literature With Large Language Model. Campbell Systematic Reviews, 21(1):e70073.
- Zhizheng Wang, Qiao Jin, Chih-Hsuan Wei, Shubo Tian, Po-Ting Lai, Qingqing Zhu, Chi-Ping Day, Christina Ross, Robert Leaman, and Zhiyong Lu. 2025e. GeneAgent: Self-verification language agent for gene-set analysis using domain databases. Nature Methods, 22(8):1677–1685. [CrossRef]
- Andrew Wentzel, Carla Floricel, Guadalupe Canahuate, Mohamed A. Naser, Abdallah S. Mohamed, Clifton D. Fuller, Lisanne van Dijk, and G. Elisabeta Marai. 2023. DASS Good: Explainable Data Mining of Spatial Cohort Data. Computer Graphics Forum, 42(3):283–295. [CrossRef]
- Austin P Wright, Peter Nemere, Adrian Galvin, Duen Horng Chau, and Scott Davidoff. 2023. Lessons from the development of an anomaly detection interface on the mars perseverance rover using the ishmap framework. In Proceedings of the 28th International Conference on Intelligent User Interfaces, page 91–105.
- Peter M. Wright, Ian B. Seiple, and Andrew G. Myers. 2014. The evolving role of chemical synthesis in antibacterial drug discovery. Angewandte Chemie International Edition, 53(34):8840–8869. [CrossRef]
- Qiujie Xie, Qingqiu Li, Zhuohao Yu, Yuejie Zhang, Yue Zhang, and Linyi Yang. 2025. An Empirical Analysis of Uncertainty in Large Language Model Evaluations. In The Thirteenth International Conference on Learning Representations.
- Qi Xin, Quyu Kong, Hongyi Ji, Yue Shen, Yuqi Liu, Yan Sun, Zhilin Zhang, Zhaorong Li, Xunlong Xia, Bing Deng, and Yinqi Bai. 2024. BioInformatics Agent (BIA): Unleashing the Power of Large Language Models to Reshape Bioinformatics Workflow. bioRxiv.
- Chaoqing Xu, Tyson Neuroth, Takanori Fujiwara, Ronghua Liang, and Kwan-Liu Ma. 2023. A Predictive Visual Analytics System for Studying Neurodegenerative Disease Based on DTI Fiber Tracts. IEEE Transactions on Visualization and Computer Graphics, 29(4):2020–2035. [CrossRef]
- Humphrey Yang, Kuanren Qian, Haolin Liu, Yuxuan Yu, Jianzhe Gu, Matthew McGehee, Yongjie Jessica Zhang, and Lining Yao. 2020. SimuLearn: Fast and Accurate Simulator to Support Morphing Materials Design and Workflows. In Proceedings of the 33rd Annual ACM Symposium on User Interface Software and Technology, UIST ’20, page 71–84.
- Jun-Hsiang Yao, Mingzheng Li, Jiayi Liu, Yuxiao Li, Jielin Feng, Jun Han, Qibao Zheng, Jianfeng Feng, and Siming Chen. 2025. DTBIA: An Immersive Visual Analytics System for Brain-Inspired Research. IEEE Transactions on Visualization and Computer Graphics, 31(6):3796–3808.
- Shunyu Yao, Jeffrey Zhao, Dian Yu, Nan Du, Izhak Shafran, Karthik Narasimhan, and Yuan Cao. 2023. ReAct: Synergizing Reasoning and Acting in Language Models. In International Conference on Learning Representations. [CrossRef]
- Fei Ye, Zaixiang Zheng, Dongyu Xue, Yuning Shen, Lihao Wang, Yiming Ma, Yan Wang, Xinyou Wang, Xiangxin Zhou, and Quanquan Gu. 2025a. Proteinbench: A holistic evaluation of protein foundation models. In International Conference on Representation Learning, volume 2025, pages 29857–29891.
- Geyan Ye, Xibao Cai, Houtim Lai, Xing Wang, Junhong Huang, Longyue Wang, Wei Liu, and Xiangxiang Zeng. 2025b. DrugAssist: a large language model for molecule optimization. Briefings in Bioinformatics, 26(1):bbae693.
- Yang Zhang, Zhewei Wei, Ye Yuan, Chongxuan Li, and Wenbing Huang. 2024a. EquiPocket: an e(3)-equivariant geometric graph neural network for ligand binding site prediction. In Proceedings of the 41st International Conference on Machine Learning, volume 235, pages 60021–60039.
- Yu Zhang, Xiusi Chen, Bowen Jin, Sheng Wang, Shuiwang Ji, Wei Wang, and Jiawei Han. 2024b. A Comprehensive Survey of Scientific Large Language Models and Their Applications in Scientific Discovery. In Proceedings of the 2024 Conference on Empirical Methods in Natural Language Processing, pages 8783–8817.
- Tianshi Zheng, Zheye Deng, Hong Ting Tsang, Weiqi Wang, Jiaxin Bai, Zihao Wang, and Yangqiu Song. 2025. From Automation to Autonomy: A Survey on Large Language Models in Scientific Discovery. pages 17744–17761.
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



