Submitted:
05 January 2026
Posted:
07 January 2026
You are already at the latest version
Abstract
Germline pathogenic variants influence breast cancer risk and clinical behavior, yet data from Indonesian populations remain scarce. In this pilot cross-sectional study, 31 newly diagnosed Indonesian women with luminal A or luminal B breast cancer underwent germline testing using a 113-gene hereditary cancer panel, with variants classified according to ACMG criteria and correlated with clinicopathological features. Pathogenic or likely pathogenic variants were identified in 9 of 31 patients (29.0%), most frequently involving BRCA2, PALB2, and RECQL4. A significantly higher frequency of pathogenic/likely pathogenic variants was observed among patients with mixed invasive histology (3/3) compared with those with invasive carcinoma of no special type (6/27; p = 0.019), and a positive first-degree family history of breast cancer was also associated with pathogenic/likely pathogenic variant status (p = 0.022). This study provides the first description of germline pathogenic variants in Indonesian luminal breast cancer and suggests that mixed invasive histology and first-degree family cancer history may represent enrichment signals for hereditary predisposition, warranting validation in larger cohorts.
Keywords:
1. Introduction
2. Materials and Methods
2.1. Statistical Analysis
3. Results
3.1. Patient Characteristics
3.2. Germline Variant Profiles
3.3. Association Between Germline Variants and Clinical Variables
4. Discussion
4.1. Pathogenic and Likely-Pathogenic Variants and Their Clinical Correlations
4.2. Family History and Variant Status
4.3. Implications for Histopathological Interpretation
4.4. Study Strengths and Limitations
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ferlay J, Ervik M, Lam F, Laversanne M, Colombet M, Mery L, Piñeros M, Znaor A, Soerjomataram I, Bray F (2024). Global Cancer Observatory: Cancer Today. Lyon, France: International Agency for Research on Cancer. Available from: https://gco.iarc.who.int/today, accessed [15 June 2025].
- Sibghatullah H, Verma H, Sangi S, Memon F, Soomro S. Retrospective Five-Year Survival Evaluation: Analyzing Breast Cancer Molecular Subtypes through Comprehensive Clinical Profiles and Prognostic Survival Trends. Transylvanian Review. 2024 Nov 8;32(3).
- Thi-Qar, T.-Q.D.O.E.I.; Al-Hilali, K.A.; Aljawher, R.Q.; Faris, S.A. Protein expression of estrogen, progesterone, and human epidermal growth factor receptors in young Iraqi women with breast cancer. Ukr. Biochem. J. 2024, 96, 38–43, . [CrossRef]
- Testa, U.; Castelli, G.; Pelosi, E. Breast Cancer: A Molecularly Heterogenous Disease Needing Subtype-Specific Treatments. Med Sci. 2020, 8, 18, . [CrossRef]
- Nik-Zainal, S.; Davies, H.; Staaf, J.; Ramakrishna, M.; Glodzik, D.; Zou, X.; Martincorena, I.; Alexandrov, L.B.; Martin, S.; Wedge, D.C.; et al. Landscape of somatic mutations in 560 breast cancer whole-genome sequences. Nature 2016, 534, 47–54, . [CrossRef]
- Ajaz, S.; Zaidi, S.-E.; Ali, S.; Siddiqa, A.; Memon, M.A. Germline Mutation Analysis in Sporadic Breast Cancer Cases With Clinical Correlations. Front. Genet. 2022, 13, 820610, . [CrossRef]
- Waks, A.G.; Kim, D.; Jain, E.; Snow, C.; Kirkner, G.J.; Rosenberg, S.M.; Oh, C.; Poorvu, P.D.; Ruddy, K.J.; Tamimi, R.M.; et al. Somatic and Germline Genomic Alterations in Very Young Women with Breast Cancer. Clin. Cancer Res. 2022, 28, 2339–2348, . [CrossRef]
- Momozawa, Y.; Iwasaki, Y.; Parsons, M.T.; Kamatani, Y.; Takahashi, A.; Tamura, C.; Katagiri, T.; Yoshida, T.; Nakamura, S.; Sugano, K.; et al. Germline pathogenic variants of 11 breast cancer genes in 7,051 Japanese patients and 11,241 controls. Nat. Commun. 2018, 9, 1–7, . [CrossRef]
- Sukpan, P.; Kanokwiroon, K.; Sriplung, H.; Laochareonsuk, W.; Choochuen, P.; Auseng, N.; Wanawanakorn, K.; Sangkhathat, S. Prevalence of Pathogenic Germline Mutations in 13 Hereditary Cancer-Related Genes in Breast Cancer Patients in Narathiwat Province, Thailand. Asian Pac. J. Cancer Prev. 2023, 24, 525–530, . [CrossRef]
- Zhang, J.; Wang, N.; Zheng, T.; Lu, T.; Zhang, R.; Ran, R.; Li, K.; Huang, Y.; Xie, F.; Zhang, Y.; et al. Germline Mutational Landscape in Chinese Patients With Advanced Breast Cancer. Front. Oncol. 2022, 12, 745796, . [CrossRef]
- Biancolella, M.; Ouédraogo, N.L.M.; Zongo, N.; Zohoncon, T.M.; Testa, B.; Rizzacasa, B.; Latini, A.; Conte, C.; Compaore, T.R.; Ouedraogo, C.M.R.-N.; et al. Breast cancer in West Africa: molecular analysis of BRCA genes in early-onset breast cancer patients in Burkina Faso. Hum. Genom. 2021, 15, 1–11, . [CrossRef]
- Ardern-Jones, A.; Kenen, R.; Eeles, R. Too much, too soon? Patients and health professionals' views concerning the impact of genetic testing at the time of breast cancer diagnosis in women under the age of 40. Eur. J. Cancer Care 2005, 14, 272–281, . [CrossRef]
- Budzik, M.P.; Fudalej, M.M.; Badowska-Kozakiewicz, A.M. Histopathological analysis of mucinous breast cancer subtypes and comparison with invasive carcinoma of no special type. Sci. Rep. 2021, 11, 1–9, . [CrossRef]
- Rechsteiner, A.; Dietrich, D.; Varga, Z. Prognostic relevance of mixed histological subtypes in invasive breast carcinoma: a retrospective analysis. J. Cancer Res. Clin. Oncol. 2022, 149, 4967–4978, . [CrossRef]
- Dayan, D.; Lukac, S.; Rack, B.; Ebner, F.; Fink, V.; Leinert, E.; Veselinovic, K.; Schütze, S.; El Taie, Z.; Janni, W.; et al. Effect of histological breast cancer subtypes invasive lobular versus non-special type on survival in early intermediate-to-high-risk breast carcinoma: results from the SUCCESS trials. Breast Cancer Res. 2023, 25, 1–15, . [CrossRef]
- Arora, A.; Agarwal, D.; Abdel-Fatah, T.M.; Lu, H.; Croteau, D.L.; Moseley, P.; A Aleskandarany, M.; Green, A.R.; Ball, G.; A Rakha, E.; et al. RECQL4 helicase has oncogenic potential in sporadic breast cancers. J. Pathol. 2015, 238, 495–501, . [CrossRef]
- Yan, K.; Gao, Y.; Heller, S.L. Breast Cancer Screening Utilization and Outcomes in Women With Neurofibromatosis Type 1. Clin. Breast Cancer 2023, 23, e200–e205, . [CrossRef]
- Keske, A.; Weisman, P.; Ospina-Romero, M.; Raut, P.; Smith-Simmer, K.; Zakas, A.L.; Flynn, C.; Xu, J. Breast cancers in monoallelic MUTYH germline mutation carriers have clinicopathological features overlapping with those in BRCA1 germline mutation carriers. Breast Cancer Res. Treat. 2023, 204, 151–158, . [CrossRef]
- Fang, H.; Nie, L.; Chi, Z.; Liu, J.; Guo, D.; Lu, X.; Hei, T.K.; Balajee, A.S.; Zhao, Y. RecQL4 Helicase Amplification Is Involved in Human Breast Tumorigenesis. PLOS ONE 2013, 8, e69600, . [CrossRef]
- Balajee, A.S. Human RecQL4 as a Novel Molecular Target for Cancer Therapy. Cytogenet. Genome Res. 2021, 161, 305–327, . [CrossRef]
- Pylkäs, K.; Vuorela, M.; Otsukka, M.; Kallioniemi, A.; Jukkola-Vuorinen, A.; Winqvist, R. Rare Copy Number Variants Observed in Hereditary Breast Cancer Cases Disrupt Genes in Estrogen Signaling and TP53 Tumor Suppression Network. PLOS Genet. 2012, 8, e1002734, . [CrossRef]
- Liu, Y.; Yang, H.; Fu, X.; Zhong, L.; Xu, P.; Fang, F.; Liu, Y.; Li, Q.; Yan, Y.; Wei, S.; et al. BRCA2, PALB2, RECQL4 Germline Pathogenic Variants, and Somatic TP53 Mutation in Triple Metachronous Malignancies: A Case Report and Literature Review. Int. Med Case Rep. J. 2024, ume 17, 23–29, . [CrossRef]
- O Seminog, O.; Goldacre, M.J. Risk of benign tumours of nervous system, and of malignant neoplasms, in people with neurofibromatosis: population-based record-linkage study. Br. J. Cancer 2012, 108, 193–198, . [CrossRef]
- Dischinger, P.S.; Tovar, E.A.; Essenburg, C.J.; Madaj, Z.B.; Gardner, E.E.; Callaghan, M.E.; Turner, A.N.; Challa, A.K.; Kempston, T.; Eagleson, B.; et al. NF1 deficiency correlates with estrogen receptor signaling and diminished survival in breast cancer. npj Breast Cancer 2018, 4, 1–14, . [CrossRef]
- Ntowe, K.W.; Thomas, S.M.; Dalton, J.C.; Olunuga, E.; Wang, T.; Chiba, A.; Plichta, J.K. Demographics and Clinical Decision Making in Patients with Germline Moderate Penetrance Non-BRCA Mutations in Breast Cancer Related Genes. Ann. Surg. Oncol. 2024, 31, 7290–7300, . [CrossRef]
- Stewart, D.R.; Korf, B.R.; Nathanson, K.L.; Stevenson, D.A.; Yohay, K. Care of adults with neurofibromatosis type 1: a clinical practice resource of the American College of Medical Genetics and Genomics (ACMG). Anesthesia Analg. 2018, 20, 671–682, . [CrossRef]
- Tung, N.; Domchek, S.M.; Stadler, Z.; Nathanson, K.L.; Couch, F.; Garber, J.E.; Offit, K.; Robson, M.E. Counselling framework for moderate-penetrance cancer-susceptibility mutations. Nat. Rev. Clin. Oncol. 2016, 13, 581–588, . [CrossRef]
- Kairupan, C.; Scott, R.J. Base excision repair and the role of MUTYH. Hered. Cancer Clin. Pr. 2007, 5, 199–209, . [CrossRef]
- Boesaard, E.P.; Vogelaar, I.P.; Bult, P.; AP Wauters, C.; van Krieken, J.H.J.; Ligtenberg, M.J.; van der Post, R.S.; Hoogerbrugge, N. Germline MUTYH gene mutations are not frequently found in unselected patients with papillary breast carcinoma. Hered. Cancer Clin. Pr. 2014, 12, 1–4, . [CrossRef]
- Cheadle, J.P.; Sampson, J.R. MUTYH-associated polyposis—From defect in base excision repair to clinical genetic testing. DNA Repair 2007, 6, 274–279, . [CrossRef]
- Nassar, A.; Zekri, A.-R.N.; Kamel, M.M.; Elberry, M.H.; Lotfy, M.M.; Seadawy, M.G.; Hassan, Z.K.; Soliman, H.K.; Lymona, A.M.; Youssef, A.S.E.-D. Frequency of Pathogenic Germline Mutations in Early and Late Onset Familial Breast Cancer Patients Using Multi-Gene Panel Sequencing: An Egyptian Study. Genes 2022, 14, 106, . [CrossRef]
- Wasielewski, M.; Out, A.A.; Vermeulen, J.; Nielsen, M.; Ouweland, A.v.D.; Tops, C.M.J.; Wijnen, J.T.; Vasen, H.F.A.; Weiss, M.M.; Klijn, J.G.M.; et al. Increased MUTYH mutation frequency among Dutch families with breast cancer and colorectal cancer. Breast Cancer Res. Treat. 2010, 124, 635–641, . [CrossRef]
- Easton, D.F.; Pharoah, P.D.; Antoniou, A.C.; Tischkowitz, M.; Tavtigian, S.V.; Nathanson, K.L.; Devilee, P.; Meindl, A.; Couch, F.J.; Southey, M.; et al. Gene-Panel Sequencing and the Prediction of Breast-Cancer Risk. New Engl. J. Med. 2015, 372, 2243–2257, . [CrossRef]
| Variable | Mean (SD) | n (%) |
| Age (year) | 47.87 (9.52) | |
| ≤ 40 | 7 (22.6) | |
| >40 | 24 (77.4) | |
| Education level | ||
| No education | 2 (6.5) | |
| Primary school | 3 (9.7) | |
| Middle school | 1 (3.2) | |
| Senior high school | 16 (51.6) | |
| Diploma and above | 9 (29) | |
| History of Cigarette Smoking | ||
| No history | 24 (77.4) | |
| Passive smoker | 7 (22.6) | |
| Age of Menarche | 13.23 (1.89) | |
| Menopausal Status when the symptoms emerged | ||
| Pre-menopause | 20 (64.5) | |
| Menopause | 11 (35.5) | |
| Parity | ||
| Nulliparity | 1 (3.2) | |
| Primiparity | 4 (12.9) | |
| Multiparity | 26 (83.9) | |
| History of hormonal contraception usage | ||
| No | 14 (45.2) | |
| Yes | 17 (54.8) | |
| History of first-degree family member(s) with breast cancer | ||
| No | 29 (93.5) | |
| Yes | 2 (6.5) | |
| History of first-degree family member(s) with ovarian cancer | ||
| No | 31 (100) | |
| Yes | 0 (0) | |
| BMI (Asia-Pacific population) | 25.18 (4.99) | |
| Underweight | 2 (6.5) | |
| Normoweight | 10 (32.3) | |
| Overweight | 3 (9.7) | |
| Obese | 16 (51.6) | |
| Stage at diagnosis | ||
| I | 1 (3.2) | |
| II | 5 (16.1) | |
| IIIA | 2 (6.5) | |
| IIIB | 8 (25.8) | |
| IV | 15 (48.4) | |
| Subtype | ||
| Luminal A | 5 (16.1) | |
| Luminal B, HER2-negative | 11 (35.5) | |
| Luminal B, HER2-positive | 15 (48.4) | |
| Ki67 index | 36.87 (23.16) | |
| Low (<20) | 6 (19.4) | |
| High (≥20) | 25 (80.6) | |
| Histological type | ||
| Invasive Breast Carcinoma, NST | 27 (87.1) | |
| Lobular Invasive Carcinoma | 1 (3.2) | |
| Mixed Invasive Carcinoma | 3 (9.7) | |
| Histological grade | ||
| Well differentiated (grade I) | 1 (3.2) | |
| Moderately differentiated (grade II) | 12 (38.7) | |
| Poorly differentiated (grade III) | 18 (58.1) | |
| Hemoglobin level at diagnosis (g/dL) | 11.87 (1.39) | |
| Leukocyte count (x 10³ cells/µL) | 8.31 (2.92) | |
| Platelet count (x 10³ cells/µL) | 314.32 (88.93) | |
| Germline-variant finding | ||
| No finding | 4 (12.9) | |
| Variant of Uncertain Significance (VUS) | 18 (58.1) | |
| Likely-pathogenic | 3 (9.7) | |
| Pathogenic | 6 (19.4) |
| Patient’s Code | Main Finding | Gene(s) | Nucleotide Change | Amino Acid Change | Other findings | |
| Gene | Classification | |||||
| BR01 | VUS | MEN1 | ||||
| BR02 | VUS | DICER1, NF2 | ||||
| BR03 | Pathogenic | CDKN2A | c.159G>C | p.Met53Ile | BLM, MUTYH | VUS |
| BR04 | VUS | BARD1, PDGFRA, TERT, XRCC2 | ||||
| BR05 | Likely-Pathogenic | ERCC3 | c.1730del | p.Lys577AsnfsTer34 | XPC | VUS |
| BR06 | Pathogenic | BRCA2 | c.262_263del | p.Leu88AlafsTer12 | EPCAM | VUS |
| BR07 | VUS | ATM, TERT | ||||
| BR08 | VUS | PTCH1 | ||||
| BR09 | Likely-Pathogenic | RECQL4 | c.2881del | p.His961Thrfs*83 | FANCE | VUS |
| Pathogenic | SPINK1 | c.101A>G | p.Asn34Ser | |||
| BR10 | VUS | PALB2, SMARCA4 | ||||
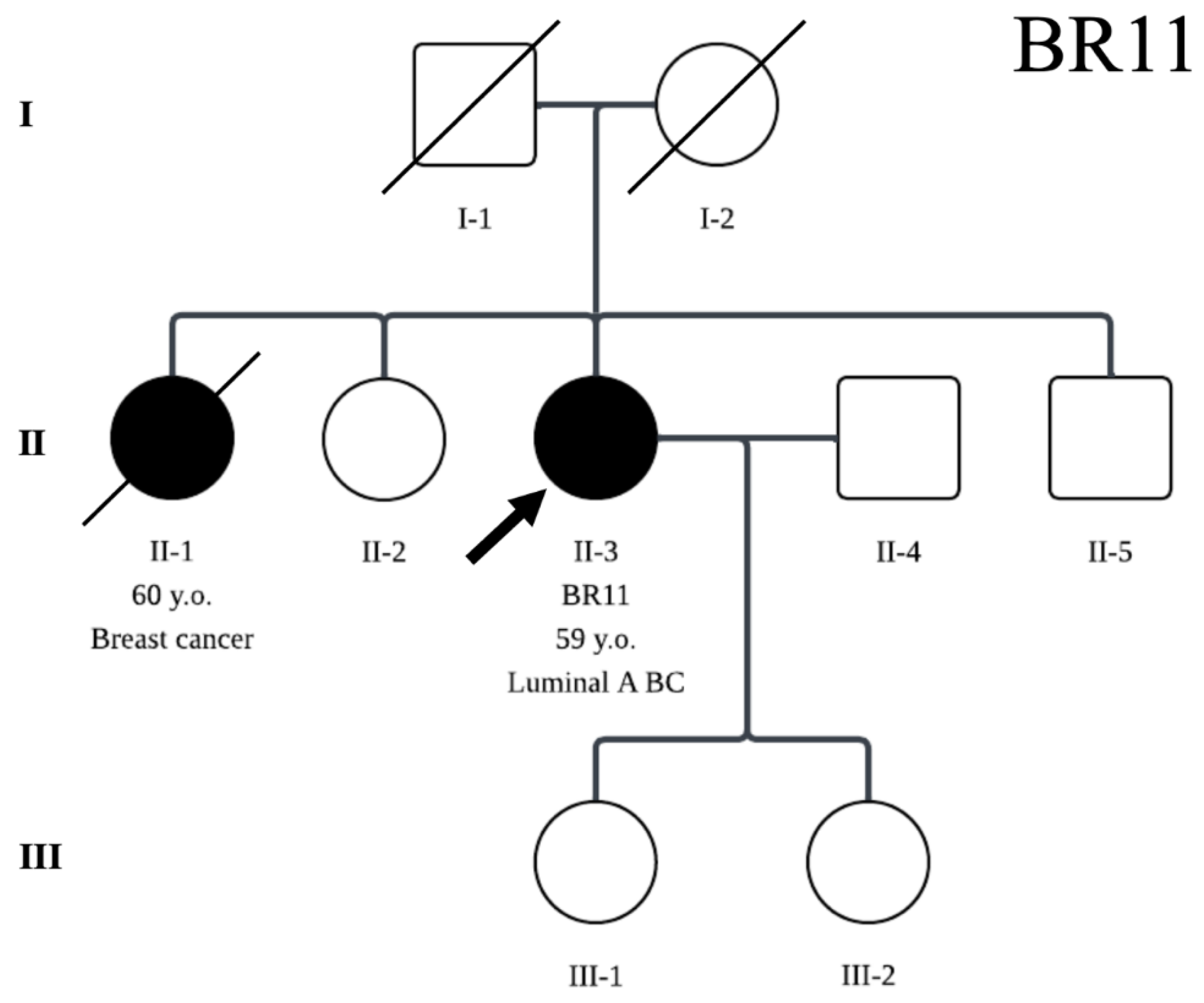
| BR11 | Likely-Pathogenic | RECQL4 | c.1342C>T | p.Pro448Ser | BRCA2, RAD51 | |
| BR12 | No Finding | |||||
| BR13 | VUS | KIF1B | ||||
| BR14 | No Finding | |||||
| BR15 | VUS | BRD10, POLE, TCHHL1 | ||||
| BR16 | VUS | KIF1B, POLE, RAD50 | ||||
| BR17 | No Finding | |||||
| BR18 | VUS | BARD1, BRIP1, FANCI, HOXB13, POLE | ||||
| BR19 | VUS | BARD1 | ||||
| BR20 | No Finding | |||||
| BR21 | VUS | NTHL1 | ||||
| BR22 | VUS | FANCE, FANCI | ||||
| BR23 | VUS | RECQL4 | ||||
| BR24 | Likely-Pathogenic | NF1 | c.3817A>G | p.Thr1273Ala | CTAGE1, RB1 | VUS |
| BR25 | VUS | BARD1, MSH3 | ||||
| BR26 | Pathogenic | PALB2 | c.1168dup | p.Ser390PhefsTer11 | CTRC | VUS |
| BR27 | Pathogenic | BRCA2 | c.8639_8640del | p.Thr2880AsnfsTer26 | BARD1, MSH3, SMARCA4 | |
| BR28 | VUS | SLX4 | ||||
| BR29 | VUS | TSC2, ATM, CHEK2, SDHA | ||||
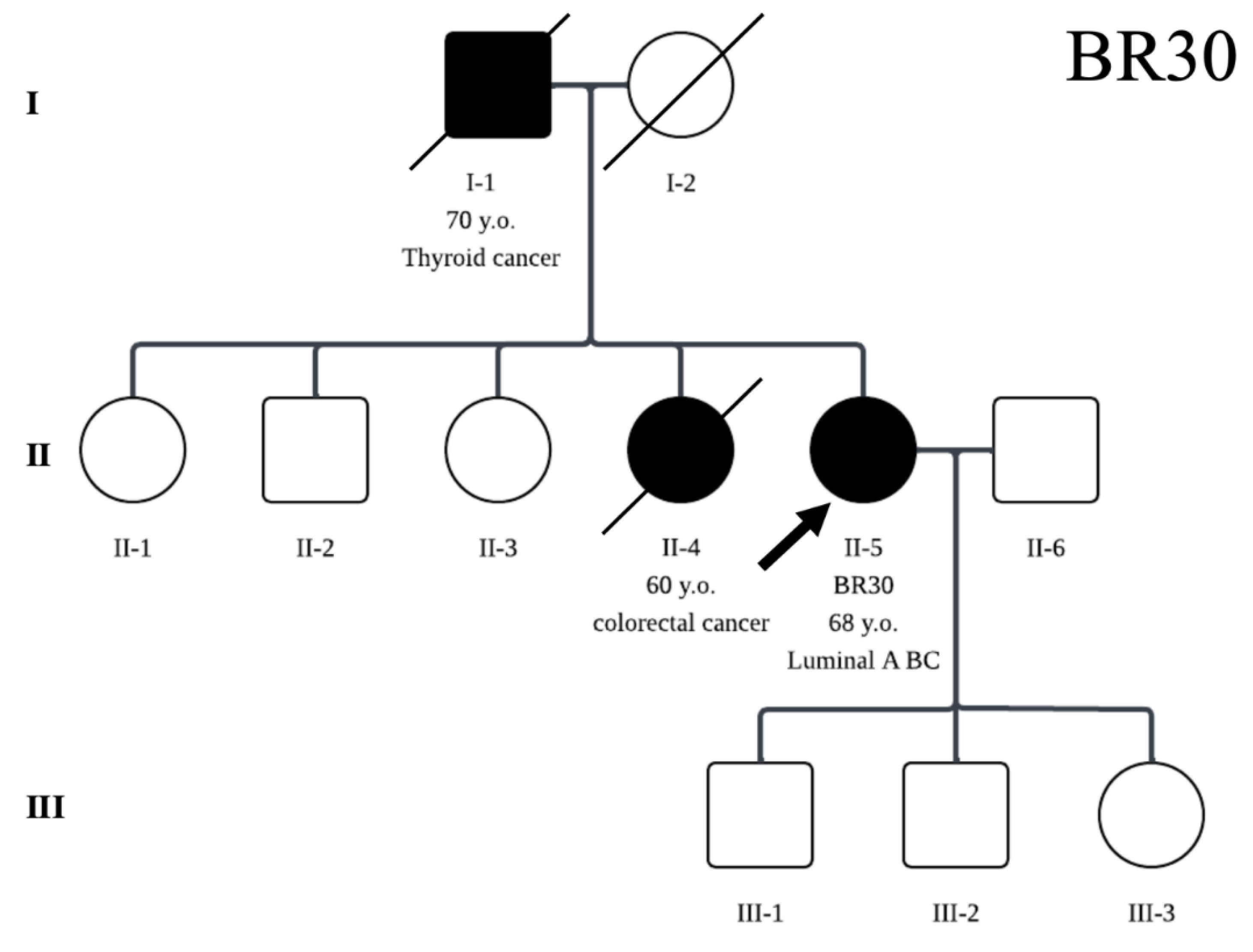
| BR30 | Pathogenic | MUTYH | c.383G>A | p.Trp128Ter | ||
| BR31 | VUS | RAD51D | ||||
| Variable | Germline-Variant Finding | OR (95% CI) | p* | p** | |
| No Finding and VUS (%) | Likely- and Pathogenic (%) | ||||
| Age | |||||
| Young age (≤ 40) | 5 (71.4) | 2 (28.6) | 1.03 (0.16–6.62) | 0.976 | 1.000 |
| >40 | 17 (70.8) | 7 (29.2) | ref | ||
| History of first-degree family member(s) with breast cancer | |||||
| No | 22 (75.8) | 7 (24.2) | ref | 0.022 | 0.077 |
| Yes | 0 | 2 (100) | 15.0 (1.1–204.5)¹ | ||
| Stage at diagnosis | |||||
| Early stage (I-IIIA) | 6 (75) | 2 (25) | 1.31 (0.21–8.18) | 0.771 | 1.000 |
| Advanced stage (IIIB-IV) | 16 (69.5) | 7 (30.5) | ref | ||
| Subtype | |||||
| Luminal A | 2 (40) | 3 (60) | 0.20 (0.03–1.49) | 0.096 | 0.131 |
| Luminal B | 20 (76.9) | 6 (23.1) | ref | ||
| Ki67 index | |||||
| Low | 3 (50) | 3 (50) | 0.32 (0.05–1.99) | 0.208 | 0.320 |
| High | 19 (76) | 6 (24) | ref | ||
| Histological type | |||||
| Invasive Breast Carcinoma, NST | 21 (77.7) | 6 (22.3) | ref | 0.019 | 0.019 |
| Lobular Invasive Carcinoma | 1 (100) | 0 | 1.10 (0.04–30.4) | ||
| Mixed Invasive Carcinoma | 0 | 3 (100) | 23.15 (1.05–506.3) | ||
| Histopathological grade | |||||
| Well differentiated (grade I) | 1 (100) | 0 | ref | 0.133 | 0.133 |
| Moderately differentiated (grade II) | 6 (50) | 6 (50) | 3.0 (0.10–89.4) | ||
| Poorly differentiated (grade III) | 15 (83.3) | 3 (16.7) | 0.68 (0.02–20.5) | ||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




