Submitted:
25 January 2026
Posted:
26 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
2.1. Consensus Meta-QTL Hotspots
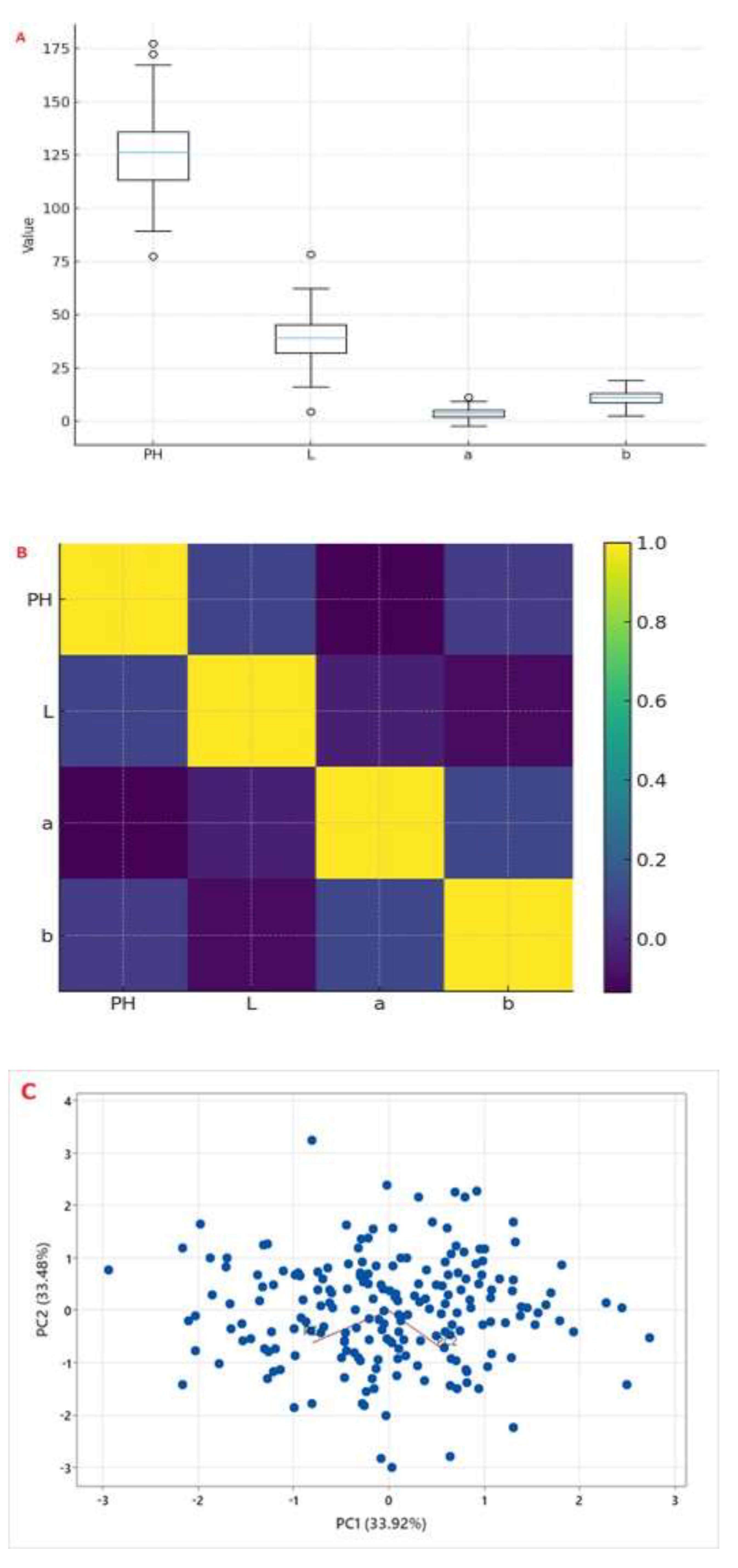
2.2. Phenotypic Variation and Heritability
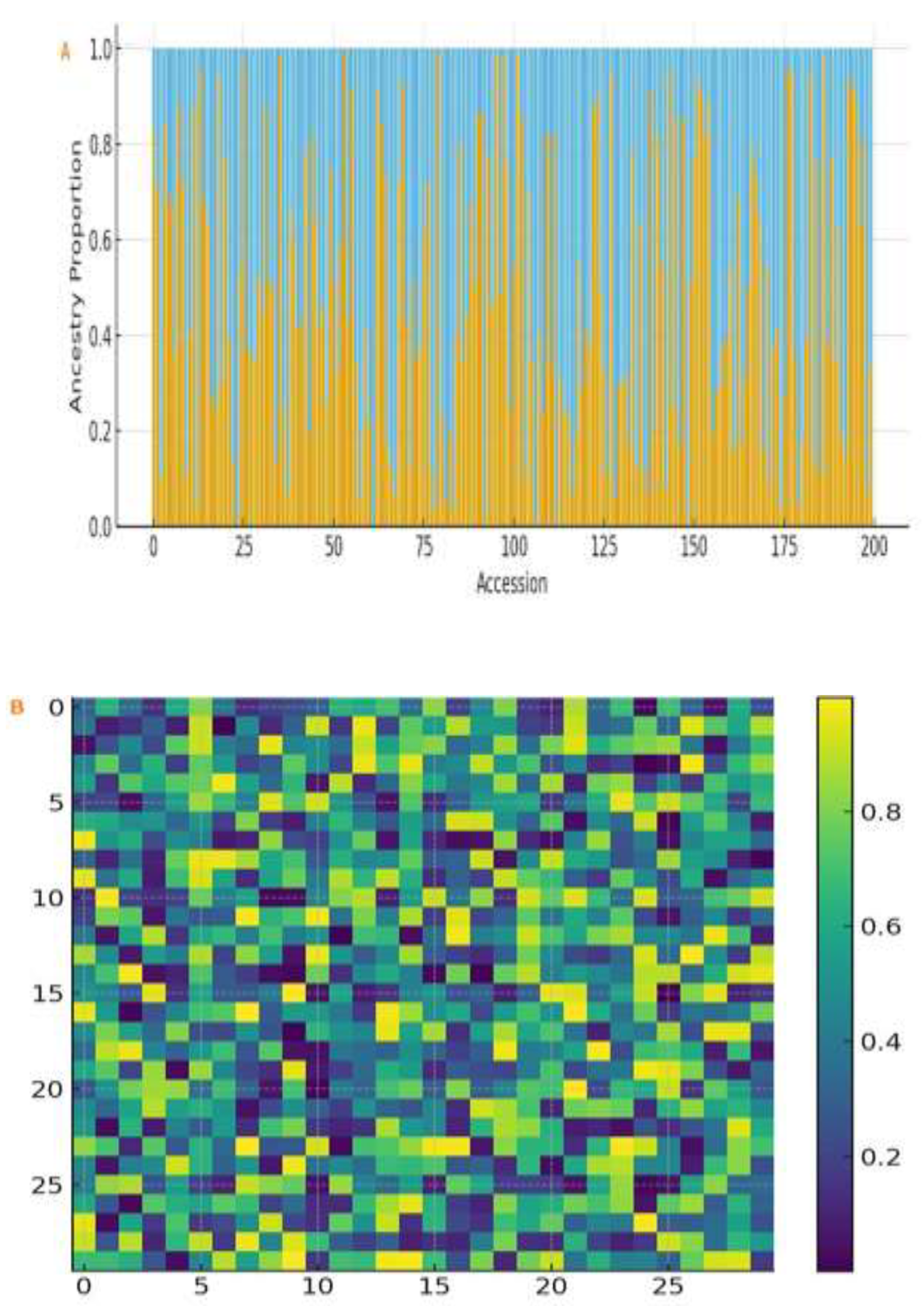
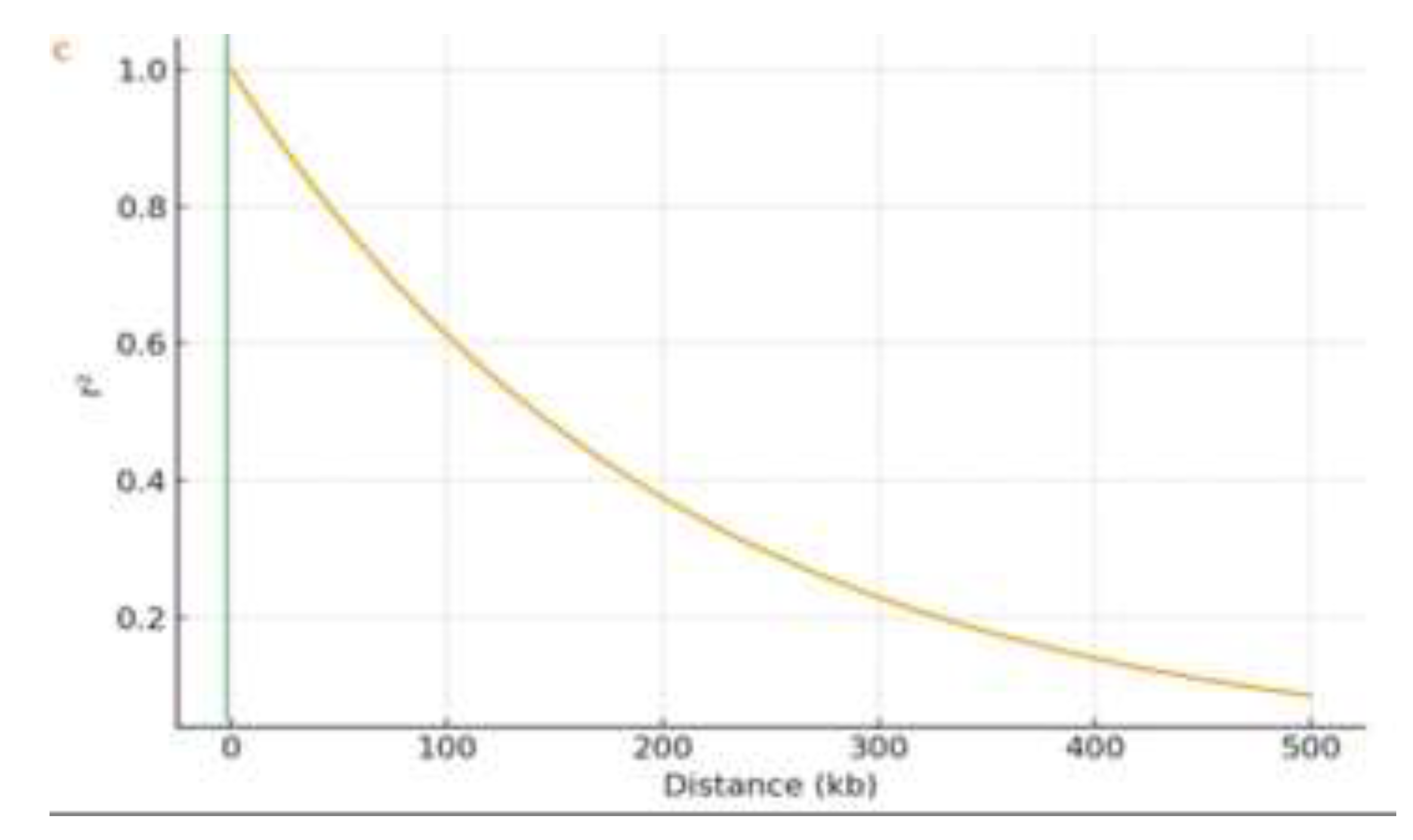
2.3. Population Structure, Kinship and Linkage Disequilibrium
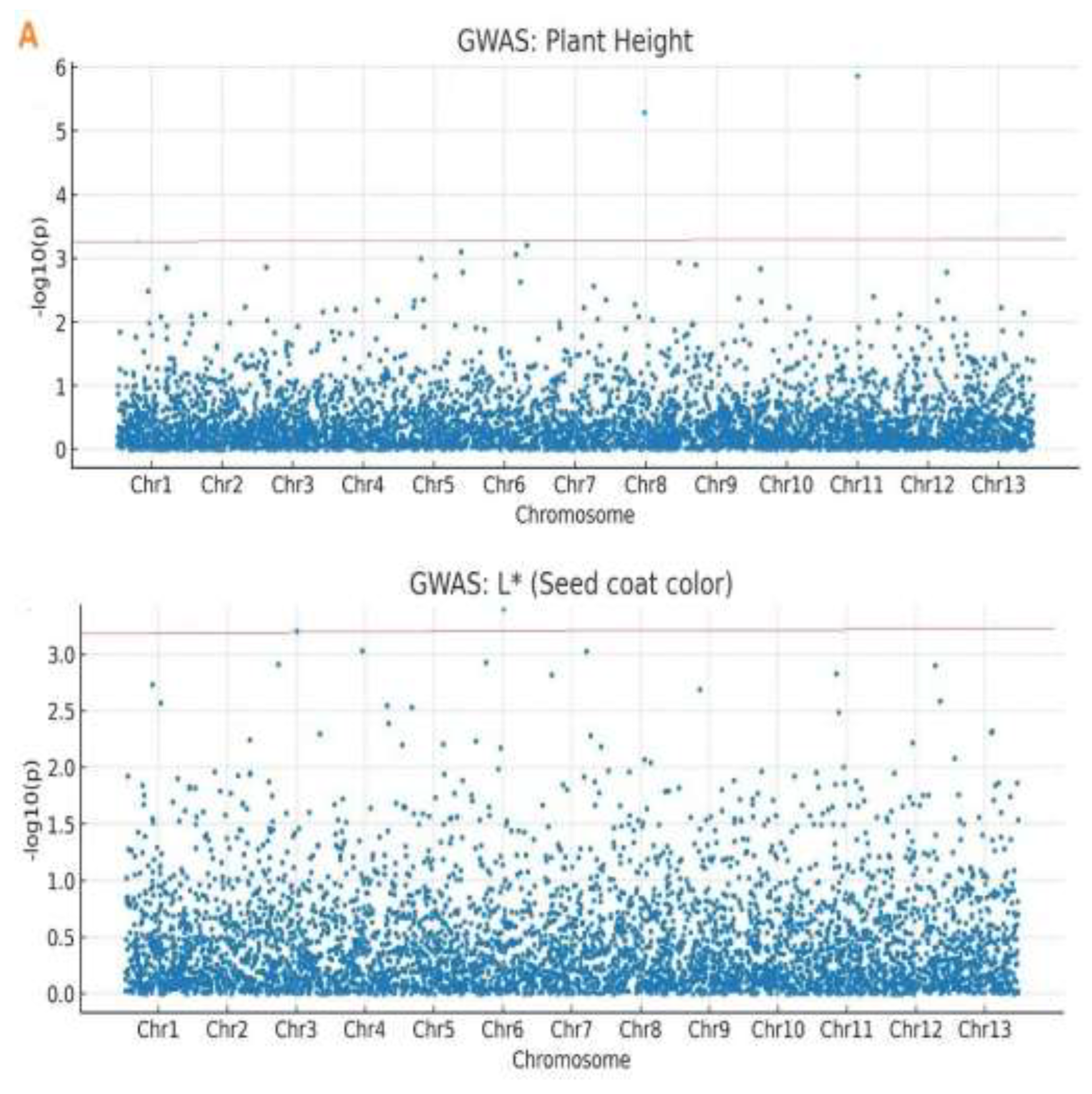
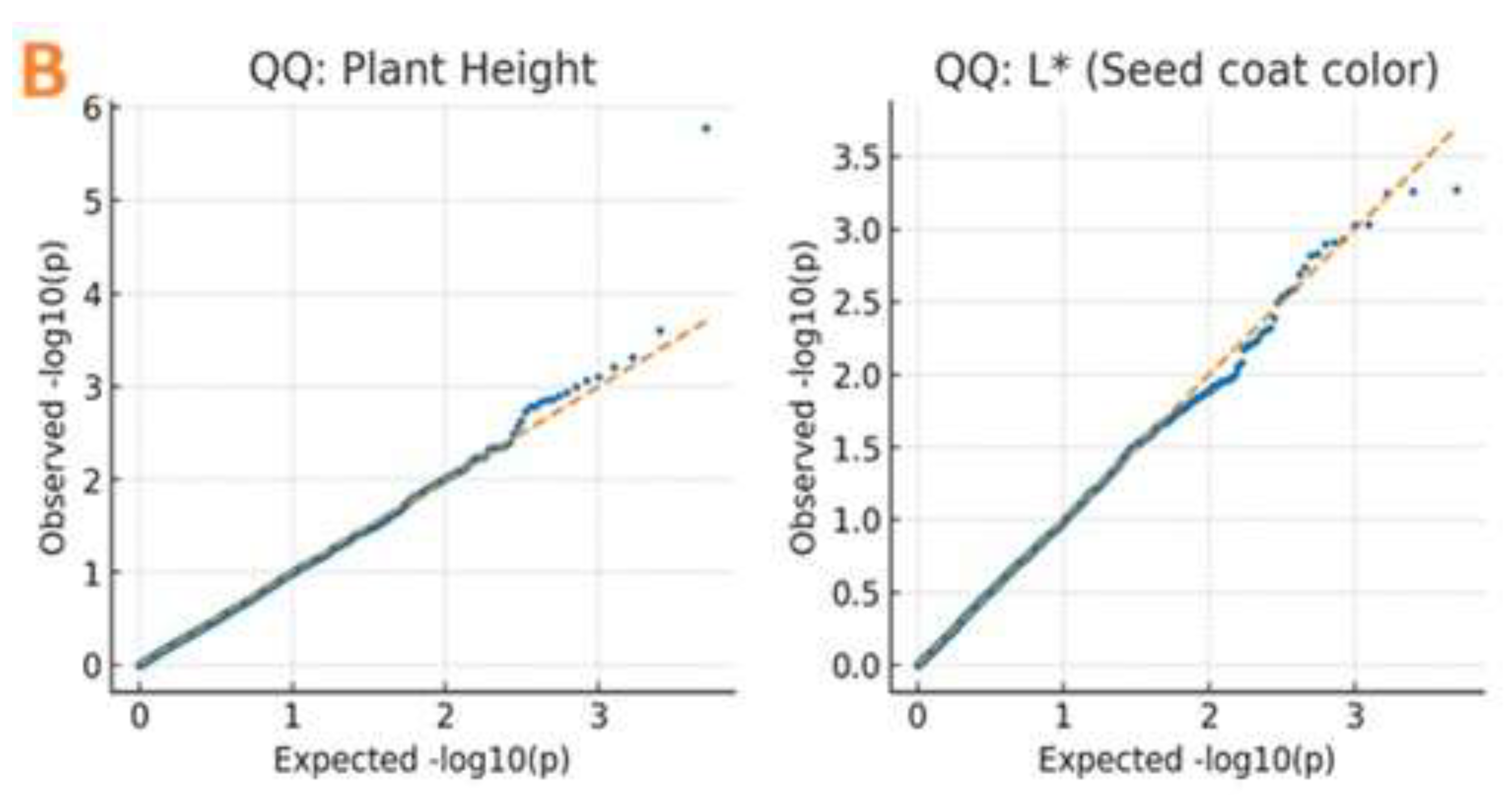
2.4. Genome-Wide Association Study
2.5. Comparative Genomic Analysis
2.6. Prioritization of High-Priority Candidate Genes
3. Discussion
3.1. Validation of Genomic Regions and Discovery of New Alleles
3.2. Hormonal Regulation of Plant Architecture
3.3. Transcriptional Networks Behind Seed Coat Color
3.4. Population Structure and LD Decay
3.5. From Discovery to Application: A Molecular Toolkit for Sesame Breeding
3.6. Limitations and Future Directions
4. Materials and Methods
4.1. Global Meta-QTL Analysis
4.2. Plant Materials and Field Experimental Design
4.3. Phenotyping
4.4. SNP Data Processing
4.5. Comparative Genomic Analysis
4.6. Population Structure, Kinship and Linkage Disequilibrium Analysis
4.7. Genome-Wide Association Analysis
4.8. Candidate Gene Identification and In Silico Functional Annotation
4.9. Phenotypic Data Analysis
5. Conclusion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AIC | Akaike Information Criterion |
| AICc | Corrected Akaike Information Criterion |
| AP2/ERF | APETALA2/ETHYLENE RESPONSE FACTOR |
| BIC | Bayesian Information Criterion |
| BSA | Bulked Segregant Analysis |
| BWA-MEM | Burrows-Wheeler Aligner – Maximal Exact Matches (bioinformatics tool) |
| Chr | Chromosome |
| cM | Centimorgan |
| CR-400 | Model of the Konica Minolta Chroma Meter |
| CV | Coefficient of Variation |
| CYP90B1 | Cytochrome P450 90B1 (DWF4 gene) |
| DIR | DIRigent (gene family) |
| DOF | DNA-binding One Zinc Finger |
| EBI | Ethiopian Biodiversity Institute |
| F₂ | Second Filial Generation |
| F₃ | Third Filial Generation |
| F₇ | Seventh Filial Generation |
| F₈ | Eighth Filial Generation |
| FAO | Food and Agriculture Organization |
| FarmCPU | Fixed and Random Model Circulating Probability Unification |
| FEM | Fixed-Effect Model |
| FST | Fixation Index (population genetic statistic) |
| GAPIT | Genome Association and Prediction Integrated Tool |
| GATK | Genome Analysis Toolkit |
| GBS | Genotyping-by-Sequencing |
| GFF3 | General Feature Format version 3 |
| GWAS | Genome-Wide Association Study |
| H² | Broad-sense Heritability |
| HTRX | Haplotype Trend Regression with eXclusion |
| IBS | Identity-by-State |
| K | Number of genetic clusters (in population structure) |
| KASP | Kompetitive Allele-Specific PCR |
| kb | Kilobase |
| LD | Linkage Disequilibrium |
| LOD | Logarithm of Odds |
| LOESS | Locally Estimated Scatterplot Smoothing |
| L* | Lightness (CIELAB color space parameter) |
| a* | Green–Red component (CIELAB color space parameter) |
| b* | Blue–Yellow component (CIELAB color space parameter) |
| MAF | Minor Allele Frequency |
| MAS | Marker-Assisted Selection |
| Mb | Megabase |
| MYB | v-MYB avian myeloblastosis viral oncogene homolog (transcription factor family) |
| bHLH | Basic Helix-Loop-Helix (transcription factor family) |
| NCBI nr | National Center for Biotechnology Information non-redundant (database) |
| PCA | Principal Component Analysis |
| PEG | Polyethylene Glycol |
| PH | Plant Height |
| PLINK | Whole genome association analysis toolset |
| PPO | Polyphenol Oxidase |
| PVE | Phenotypic Variance Explained |
| Q-matrix | Ancestry proportion matrix (from population structure) |
| Q-Q plot | Quantile-Quantile plot |
| QTL | Quantitative Trait Locus/Loci |
| r | Number of replicates |
| r² | Squared correlation coefficient (measure of LD) |
| RAD-seq | Restriction-site Associated DNA Sequencing |
| REM | Random-Effect Model (mentioned as background model in FarmCPU) |
| RIL | Recombinant Inbred Line |
| SBP | SQUAMOSA Promoter-Binding Protein |
| SCC | Seed Coat Color |
| SIACS9 | Sesamum indicum 1-aminocyclopropane-1-carboxylic acid synthase 9 |
| SICEN2 | Sesamum indicum Centroradialis 2 |
| Sindi | Sesamum indicum (gene prefix in genome annotation) |
| SLAF | Specific-Length Amplified Fragment |
| SLUBI | Swedish University of Agricultural Sciences Bioinformatics Infrastructure |
| SLU | Swedish University of Agricultural Sciences |
| SNP | Single Nucleotide Polymorphism |
| SSD | Single-Seed Descent |
| SSR | Simple Sequence Repeat |
| STY8 | Serine/Threonine-protein kinase STY8 |
| TASSEL | Trait Analysis by aSSociation, Evolution and Linkage |
| TF | Transcription Factor |
| VCF | Variant Call Format |
| VCFtools | Variant Call Format tools |
| WARC | Werer Agricultural Research Center |
| WRKY | Transcription factor family named after the conserved WRKY domain |
References
- Wei, W.; Zhang, Y.; Lü, H.; Li, D.; Wang, L.; Zhang, X. Association Analysis for Quality Traits in a Diverse Panel of Chinese Sesame (Sesamum indicum L.) Germplasm. J. Integr. Plant Biol. 2013, 55, 745–758. [Google Scholar] [CrossRef]
- Andargie, M.; Vinas, M.; Rathgeb, A.; Möller, E.; Karlovsky, P. Lignans of Sesame (Sesamum indicum L.): A Comprehensive Review. Molecules 2021, 26, 883. [Google Scholar] [CrossRef] [PubMed]
- Dossa, K.; Diouf, D.; Wang, L.; Wei, X.; Zhang, Y.; Niang, M.; Fonceka, D.; Yu, J.; Mmadi, M.A.; Yehouessi, L.W.; et al. The Emerging Oilseed Crop Sesamum indicum Enters the “Omics” Era. Front. Plant Sci. 2017, 8, 1154. [Google Scholar] [CrossRef]
- Rauf, S.; Basharat, T.; Gebeyehu, A.; Elsafy, M.; Rahmatov, M.; Ortiz, R.; Kaya, Y. Sesame, an Underutilized Oilseed Crop: Breeding Achievements and Future Challenges. Plants 2024, 13, 2662. [Google Scholar] [CrossRef]
- FAOSTAT. World Food and Agriculture - Statistical Yearbook 2024. 2024. Available online: https://www.fao.org/faostat/ (accessed on 21 September 2025).
- Tesfaye, T.; Tesfaye, K.; Keneni, G.; Alemu, T.; Alemu, A. Genome-Wide Association Study for Yield-Related Traits in Sesame (Sesamum Indicum). Plant Breed. 2022, 141, 246–256. [Google Scholar] [CrossRef]
- Varshney, R.K.; Bohra, A.; Yu, J.; Graner, A.; Zhang, Q.; Sorrells, M.E. Designing Future Crops: Genomics-Assisted Breeding Comes of Age. Trends Plant Sci. 2021, 26, 631–649. [Google Scholar] [CrossRef] [PubMed]
- Varshney, R.K.; Graner, A.; Sorrells, M.E. Genomics-Assisted Breeding for Crop Improvement. Trends Plant Sci. 2005, 10, 621–630. [Google Scholar] [CrossRef]
- Weldemichael, M.Y.; Gebremedhn, H.M.; Runo, S.; Gandhi, H. Molecular Breeding for Stress Tolerance in Sesame. Mol. Genet. Genom. 2025, 300, 69. [Google Scholar] [CrossRef]
- Berhe, M.; Dossa, K.; You, J.; Mboup, P.A.; Diallo, I.N.; Diouf, D.; Zhang, X.; Wang, L. Genome-Wide Association Study and Its Applications in the Non-Model Crop Sesamum indicum. BMC Plant Biol. 2021, 21, 283. [Google Scholar] [CrossRef]
- Debnath, A.J.; Ernst, D.; Harenčár, Ľ.; Kučka, M.; Basu, D.; Sikdar, S.R. A Review on Yield Improvement of the Important Oilseed Sesame (Sesamum indicum L.) using Biotechnology. Sesamum indicum 2024. [Google Scholar] [CrossRef]
- Teklu, D.H.; Shimelis, H.; Abady, S. Genetic Improvement in Sesame (Sesamum indicum L.): Progress and Outlook: A Review. Agronomy 2022, 12, 2144. [Google Scholar] [CrossRef]
- Wang, L.; Yu, S.; Tong, C.; Zhao, Y.; Liu, Y.; Song, C.; Zhang, Y.; Zhang, X.; Wang, Y.; Hua, W.; et al. Genome Sequencing of the High Oil Crop Sesame Provides Insight into Oil Biosynthesis. Genome Biol. 2014, 15, R39. [Google Scholar] [CrossRef]
- Mondal, R.; Kumar, A.; Gnanesh, B.N. Crop Germplasm: Current Challenges, Physiological-Molecular Perspective, and Advanced Strategies Towards Development of Climate-Resilient Crops. Heliyon 2023, 9, e12879. [Google Scholar] [CrossRef]
- Teklu, D.H.; Shimelis, H.; Tesfaye, A.; Mashilo, J.; Zhang, X.; Zhang, Y.; Dossa, K.; Shayanowako, A.I.T. Genetic Variability and Population Structure of Ethiopian Sesame (Sesamum indicum L.) Germplasm Assessed through Phenotypic Traits and Simple Sequence Repeats Markers. Plants 2021, 10, 1129. [Google Scholar] [CrossRef]
- Harlan, J.R. Crops and Man; American Society of Agronomy and Crop Science Society of America: Madison, WI, USA, 1992. [Google Scholar]
- Nayar, N.M.; Mehra, K. Sesame: Its Uses, Botany, Cytogenetics, and Origin. Econ. Bot. 1970, 24, 20–31. [Google Scholar] [CrossRef]
- Gedifew, S.; Demelash, H.; Abate, A.; Abebe, T.D. Association of Quantitative Traits and Genetic Diversity in Ethiopian Sesame (Sesamum indicum L.) Genotypes. Heliyon 2024, 10, e26623. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Dossa, K.; Zhang, Y.; Wei, X.; Wang, L.; Zhang, Y.; Liu, A.; Zhou, R.; Zhang, X. GWAS Uncovers Differential Genetic Bases for Drought and Salt Tolerances in Sesame at the Germination Stage. Genes 2018, 9, 87. [Google Scholar] [CrossRef]
- Wu, J.; Cheng, J.-H.; Yang, F.-C. Transcriptional Regulation of Anthocyanin Biosynthesis in Plants. Chin. J. Cell Biol. 2006, 28, 453–456. [Google Scholar]
- Cui, C.; Liu, Y.; Liu, Y.; Cui, X.; Sun, Z.; Du, Z.; Wu, K.; Jiang, X.; Mei, H.; Zheng, Y. Genome-Wide Association Study of Seed Coat Color in Sesame (Sesamum indicum L.). PLoS ONE 2021, 16, e0251526. [Google Scholar] [CrossRef]
- Wang, L.; Dossou, S.S.K.; Wei, X.; Zhang, Y.; Li, D.; Yu, J.; Zhang, X. Transcriptome Dynamics during Black and White Sesame (Sesamum indicum L.) Seed Development and Identification of Candidate Genes Associated with Black Pigmentation. Genes 2020, 11, 1399. [Google Scholar] [CrossRef]
- Zhang, H.; Miao, H.; Wei, L.; Li, C.; Zhao, R.; Wang, C. Genetic Analysis and QTL Mapping of Seed Coat Color in Sesame (Sesamum indicum L.). PLoS ONE 2013, 8, e63898. [Google Scholar] [CrossRef] [PubMed]
- Moazzami, A.A.; Kamal-Eldin, A. Sesame Seed Is a Rich Source of Dietary Lignans. J. Am. Oil Chem. Soc. 2006, 83, 719–723. [Google Scholar] [CrossRef]
- Teklu, D.H.; Shimelis, H.; Tesfaye, A.; Abady, S. Appraisal of the Sesame Production Opportunities and Constraints, and Farmer-Preferred Varieties and Traits, in Eastern and Southwestern Ethiopia. Sustainability 2021, 13, 11202. [Google Scholar] [CrossRef]
- Du, H.; Zhang, H.; Wei, L.; Li, C.; Duan, Y.; Wang, H. A High-Density Genetic Map Constructed Using Specific Length Amplified Fragment (SLAF) Sequencing and QTL Mapping of Seed-Related Traits in Sesame (Sesamum indicum L.). BMC Plant Biol. 2019, 19, 588. [Google Scholar] [CrossRef]
- Elsafy, M.; Badawi, W.; Ibrahim, A.; Hafiz Baillo, E.; Bajgain, P.; Abdelhalim, T.S.; Rahmatov, M. Genome-Wide Association Scan and Candidate Gene Analysis for Seed Coat Color in Sesame (Sesamum indicum L.). Front. Plant Sci. 2025, 16, 1541656. [Google Scholar] [CrossRef] [PubMed]
- Guden, B.; Qureshi, M.; Wang, L.; Li, H.; Sinare, B.; Uzun, B.; Yol, E. Molecular Breeding to Design Ideal Sesame Crop. In Allele Mining for Genomic Designing of Oilseed Crops; Kole, C., Ed.; CRC Press: Boca Raton, FL, USA, 2024; pp. 217–238. [Google Scholar]
- Wang, L.; Xia, Q.; Zhang, Y.; Zhu, X.; Zhu, X.; Li, D.; Ni, X.; Gao, Y.; Xiang, H.; Wei, X.; et al. Updated Sesame Genome Assembly and Fine Mapping of Plant Height and Seed Coat Color QTLs Using a New High-Density Genetic Map. BMC Genom. 2016, 17, 31. [Google Scholar] [CrossRef]
- Myles, S.; Peiffer, J.; Brown, P.J.; Ersoz, E.S.; Zhang, Z.; Costich, D.E.; Buckler, E.S. Association Mapping: Critical Considerations Shift from Genotyping to Experimental Design. Plant Cell 2009, 21, 2194–2202. [Google Scholar] [CrossRef]
- Bhat, J.A.; Ali, S.; Salgotra, R.K.; Mir, Z.A.; Dutta, S.; Jadon, V.; Tyagi, A.; Mushtaq, M.; Jain, N.; Singh, P.K.; et al. Genomic Selection in the Era of Next-Generation Sequencing for Complex Traits in Plant Breeding. Front. Genet. 2016, 7, 221. [Google Scholar] [CrossRef]
- Liang, J.; Sun, J.; Ye, Y.; Yan, X.; Yan, T.; Rao, Y.; Zhou, H.; Le, M. QTL Mapping of PEG-Induced Drought Tolerance at the Early Seedling Stage in Sesame Using Whole Genome Re-Sequencing. PLoS ONE 2021, 16, e0247681. [Google Scholar] [CrossRef] [PubMed]
- Sheng, C.; Song, S.; Zhou, R.; Li, D.; Gao, Y.; Cui, X.; Tang, X.; Zhang, Y.; Tu, J.; Zhang, X. QTL-seq and Transcriptome Analysis Disclose Major QTL and Candidate Genes Controlling Leaf Size in Sesame (Sesamum indicum L.). Front. Plant Sci. 2021, 12, 580846. [Google Scholar] [CrossRef]
- Wu, K.; Liu, H.; Yang, M.; Tao, Y.; Ma, H.; Wu, W.; Zuo, Y.; Zhao, Y. High-Density Genetic Map Construction and QTLs Analysis of Grain Yield-Related Traits in Sesame (Sesamum indicum L.) Based on RAD-Seq Technology. BMC Plant Biol. 2014, 14, 274. [Google Scholar] [CrossRef]
- Wang, L.; Yu, J.; Zhang, Y.; You, J.; Zhang, X.; Wang, L. Sinbase 2.0: An Updated Database to Study Multi-Omics in Sesamum indicum. Plants 2021, 10, 272. [Google Scholar] [CrossRef]
- Sosnowski, O.; Charcosset, A.; Joets, J. BioMercator V3: An Upgrade of Genetic Map Compilation and Quantitative Trait Loci Meta-Analysis Algorithms. Bioinformatics 2012, 28, 2082–2083. [Google Scholar] [CrossRef]
- Wang, M; Huang, J; Liu, S; Liu, X; Li, R; Luo, J; Fu, Z. Improved assembly and annotation of the sesame genome. DNA Res. 2022, 29(6), dsac041. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Alexander, D.H.; Novembre, J.; Lange, K. Fast Model-Based Estimation of Ancestry in Unrelated Individuals. Genome Res. 2009, 19, 1655–1664. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Lawson, D.J. HTRX: An R Package for Learning Non-Contiguous Haplotypes Associated with a Phenotype. Bioinform. Adv. 2023, 3, vbad038. [Google Scholar] [CrossRef]
- Bradbury, P.J.; Zhang, Z.; Kroon, D.E.; Casstevens, T.M.; Ramdoss, Y.; Buckler, E.S. TASSEL: Software for Association Mapping of Complex Traits in Diverse Samples. Bioinformatics 2007, 23, 2633–2635. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Huang, M.; Fan, B.; Buckler, E.S.; Zhang, Z. Iterative Usage of Fixed and Random Effect Models for Powerful and Efficient Genome-Wide Association Studies. PLoS Genet. 2016, 12, e1005767. [Google Scholar] [CrossRef]
- Jones, P.; Binns, D.; Chang, H.-Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. InterProScan 5: Genome-Scale Protein Function Classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef] [PubMed]
- Kaur, S.; Das, A.; Sheoran, S.; Rakshit, S. QTL Meta-Analysis: An Approach to Detect Robust and Precise QTL. Trop. Plant Biol. 2023, 16, 225–243. [Google Scholar] [CrossRef]
- Wei, X; Gong, H; Yu, J; Liu, P; Wang, L; Zhang, Y; Zhang, X. SesameFG: an integrated database for the functional genomics of sesame. Sci Rep. 2017, 7(1), 2342. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Dubois, M.; Van den Broeck, L.; Inzé, D. The Pivotal Role of Ethylene in Plant Growth. Trends Plant Sci. 2018, 23, 311–323. [Google Scholar] [CrossRef]
- Mei, H.; Liu, Y.; Cui, C.; Hu, C.; Xie, F.; Zheng, L.; Du, Z.; Wu, K.; Jiang, X.; Zheng, Y. QTL Mapping of Yield-Related Traits in Sesame. Mol. Breed. 2021, 41, 43. [Google Scholar] [CrossRef]
- Müller, M.; Munné-Bosch, S. Ethylene Response Factors: A Key Regulatory Hub in Hormone and Stress Signaling. Plant Physiol. 2015, 169, 32–41. [Google Scholar] [CrossRef]
- Choe, S.; Dilkes, B.P.; Fujioka, S.; Takatsuto, S.; Sakurai, A.; Feldmann, K.A. The DWF4 Gene of Arabidopsis Encodes a Cytochrome P450 That Mediates Multiple 22α-Hydroxylation Steps in Brassinosteroid Biosynthesis. Plant Cell 1998, 10, 231–243. [Google Scholar] [CrossRef] [PubMed]
- Yanagisawa, S. Dof Domain Proteins: Plant-Specific Transcription Factors Associated with Diverse Phenomena Unique to Plants. Plant Cell Physiol. 2004, 45, 386–391. [Google Scholar] [CrossRef] [PubMed]
- Cortés, A.J.; López-Hernández, F. Harnessing Crop Wild Diversity for Climate Change Adaptation. Genes 2021, 12, 783. [Google Scholar] [CrossRef]
- Nolan, T.M.; Vukašinović, N.; Liu, D.; Russinova, E.; Yin, Y. Brassinosteroids: Multidimensional Regulators of Plant Growth, Development, and Stress Responses. Plant Cell 2020, 32, 295–318. [Google Scholar] [CrossRef]
- Achard, P.; Vriezen, W.H.; Van Der Straeten, D.; Harberd, N.P. Ethylene Regulates Arabidopsis Development via the Modulation of DELLA Protein Growth Repressor Function. Plant Cell 2003, 15, 2816–2825. [Google Scholar] [CrossRef] [PubMed]
- Shahzadi, I.; Ahmad, A.; Noreen, Z.; Akram, W.; Yasin, N.A.; Khan, W.U. Brassinosteroid and Ethylene-Mediated Cross Talk in Plant Growth and Development. In Brassinosteroids Signalling: Intervention with Phytohormones and Their Relationship in Plant Adaptation to Abiotic Stresses; Aftab, T., Ed.; Springer: Singapore, 2022; pp. 117–136. [Google Scholar]
- Langham, D. Shatter Resistance in Sesame. In Sesame Improvement by Induced Mutations; Final reports of an FAO/IAEA coordinated research project. 1993-1998; International Atomic Energy Agency: Vienna, Austria, 2001; pp. 51–61. [Google Scholar]
- Dossou, S.S.K.; Luo, Z.; Deng, Q.; Zhou, R.; Zhang, Y.; Li, D.; Li, H.; Tozo, K.; You, J.; Wang, L. Biochemical and Molecular Insights into Variation in Sesame Seed Antioxidant Capability as Revealed by Metabolomics and Transcriptomics Analysis. Antioxidants 2024, 13, 514. [Google Scholar] [CrossRef]
- Wang, J.; Zhang, Z. GAPIT Version 3: Boosting Power and Accuracy for Genomic Association and Prediction. Genom. Proteom. Bioinform. 2021, 19, 629–640. [Google Scholar] [CrossRef] [PubMed]
- Heslot, N.; Jannink, J.L.; Sorrells, M.E. Perspectives for Genomic Selection Applications and Research in Plants. Crop Sci. 2015, 55, 1–12. [Google Scholar] [CrossRef]
- Sharma, R.; Yang, C.J.; Rossi, N.; Irving, E.; Tuffin, A.; Aliki, H.; Powell, W.; Dawson, I.K. Integrating Molecular Genetics with Plant Breeding to Deliver Impact. Plant Physiol. 2025, 198, kiaf087. [Google Scholar] [CrossRef] [PubMed]
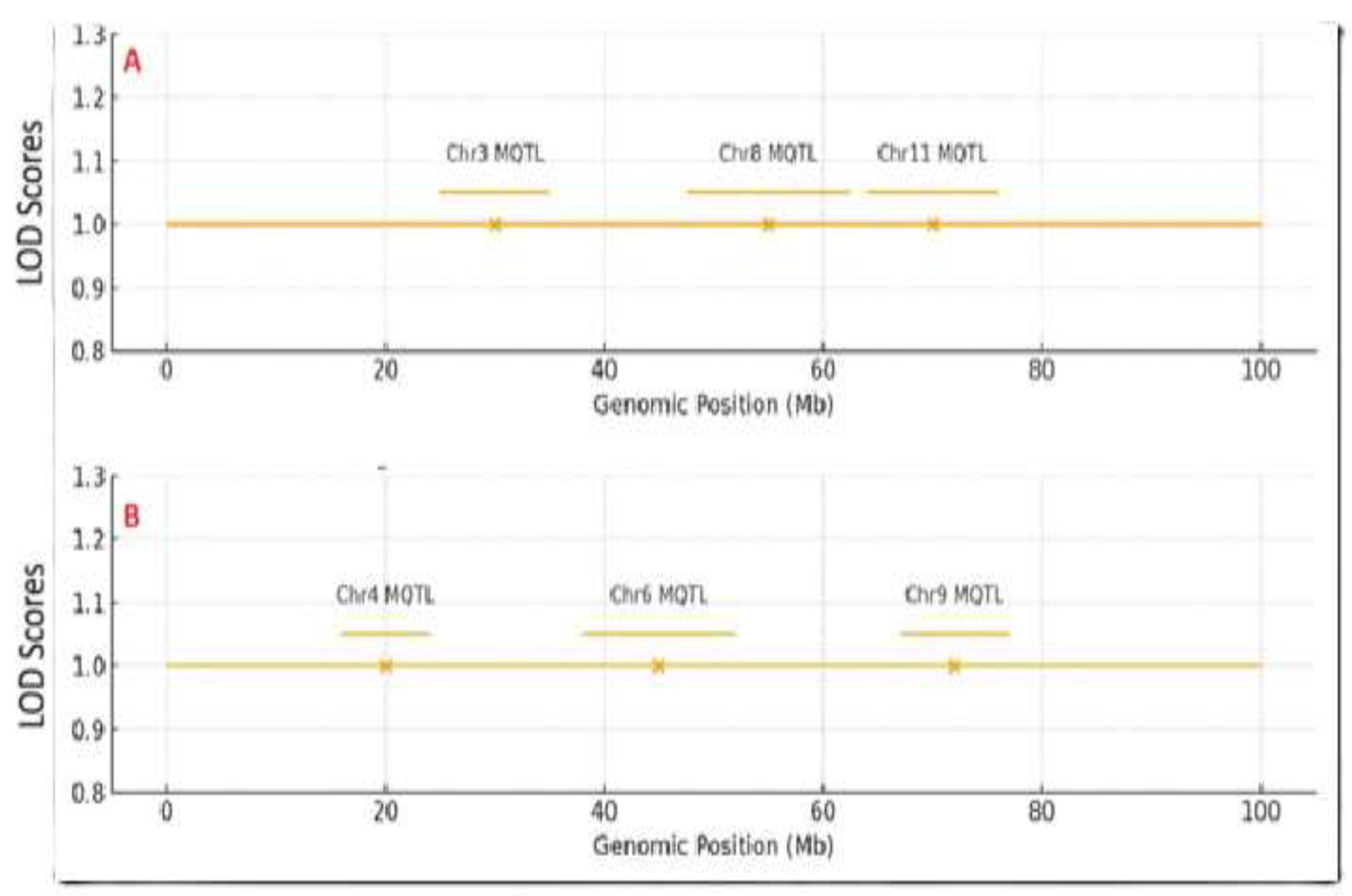
| Trait | Meta-QTL hotspot region | Peak position (Mb) | Closely linked markers | Number of QTL | PVE range (%) | Key candidate genes/ References |
|---|---|---|---|---|---|---|
| Plant Height | Chr03: ~25-35 cM | 28.4 | Sindi.03G185, Sindi.03G192 | 5 | 9.44 – 15.10 |
SICEN2 [46], SIACS9 [32] |
| Plant Height | Chr08: ~175-180 cM | 177.8 | Sindi.08G774, Sindi.I08G781 | 4 | 12.80 – 71.41 | qFCHLG08-2 [46], CYP90B1 (this study) |
| Plant Height | Chr11: ~185-190 cM | 187.2 | Sindi11G945, Sindi11G952 | 4 | 11.23 – 18.50 | qPLLG11-1 [46], AP2/ERF (this study) |
| Seed Coat Color | Chr04: ~45-55 cM | 48.6 | Sindi04G332, Sindi04G339 | 6 | 5.62 – 23.10 | qSC6-4-1 [29], DIR gene family [21] |
| Seed Coat Color | Chr06: ~0.7-0.85 cM | 1.21 | Sindi06G058, Sindi06G065 | 5 | 8.50 – 25.50 | qBSCchr6 [43], PPO [44], WRKY (this study) |
| Seed Coat Color | Chr09: ~88-92 cM | 90.1 | Sindi09G441, Sindi09G448 | 4 | 10.15 – 32.88 | qSC6-9 [26], MYB/bHLH [21,22] |
| Trait | SNP marker | Chr. | Position (bp) | p-value | -log₁₀(p) | PVE (%) | Allelic effect |
|---|---|---|---|---|---|---|---|
| Plant Height | Chr11_1877114 | 11 | 1,877,114 | 1.24 × 10⁻⁶ | 5.91 | 14.20 | -8.45 |
| Plant Height | Chr08_1771424 | 8 | 1,771,424 | 3.89 × 10⁻⁶ | 5.41 | 12.80 | 7.21 |
| L* | Chr12_16523829 | 12 | 16,523,829 | 2.17 × 10⁻³ | 2.66 | 6.32 | -3.95 |
| a* | Chr06_27694080 | 6 | 27,694,080 | 7.84 × 10⁻⁷ | 6.11 | 8.95 | -1.72 |
| a* | Chr03_15960455 | 3 | 15,960,455 | 4.25 × 10⁻⁴ | 3.37 | 7.05 | 1.42 |
| b* | Chr13_345249 | 13 | 345,249 | 1.48 × 10⁻³ | 2.83 | 6.08 | -4.71 |
| Trait | SNP marker | Candidate gene |
Distance to SNP (kb) | Putative function | Sequence identity (%) |
|---|---|---|---|---|---|
| Plant Height | Chr11_1877114 | Sindi.11G025000 | 12.4 | AP2/ERF domain-containing protein | 95.2 |
| Plant Height | Chr08_1771424 | Sindi.08G015600 | 8.7 | Cytochrome P450 CYP90B1 (Brassinosteroid biosynthesis) | 88.7 |
| a* | Chr06_27694080 | Sindi.06G123400 | 15.2 | WRKY transcription factor 23 | 94.3 |
| L* | Chr12_16523829 | Sindi.12G045200 | 22.8 | Squamosa promoter-binding protein 1 | 97.8 |
| a* | Chr03_15984975 | Sindi.03G078100 | 18.5 | DOF zinc finger protein DOF3.1 | 82.1 |
| a* | Chr03_26242291 | Sindi.03G090200 | 31.7 | Serine/threonine-protein kinase STY8 | 98.5 |
| b* | Chr09_22387055 | Sindi.09G078500 | 26.3 | Salicylic acid-binding protein 2 | 96.7 |
| Chr. | Key QTL region | PVE range (%) | Population | Key candidates | Reference |
|---|---|---|---|---|---|
| 4 | qSCa-4.1, qscca*4 (∼78-81 cM) | 8.56–23.10 | RIL, F₃ | DIR gene family | [26,29] |
| 6 | qBSCchr6 (∼2.1 cM) | Major QTL | RIL (BSA) | 13 candidate brown seed locus | [43] |
| 6 | Meta-QTL hotspot | 8.50–25.50 | Meta-analysis | PPO, WRKY TFs | This study (Table 1) |
| 9 | qsccY9, qsccZ9 (∼90-104 cM) | 32.88–33.25 | F₃ | MYB, bHLH TFs | [26] |
| 9 | Meta-QTL hotspot | 10.15–32.88 | Meta-analysis | MYB/bHLH complex | Table 1 |
| 12 | qsccZ12 | 5.58 | F₃ | – | [26] |
| 12, 13 | Novel GWAS associations | 6.22–6.51 | Ethiopian panel | SBP-like, Kinase STY8 | Table 1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




