Submitted:
03 January 2026
Posted:
05 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Theoretical Developments
2.1. Bayesian Inference and Cell Decision-Making
| Symbol | Description |
|---|---|
| X | Internal/phenotypic random variable |
| Y | External/environmental random variable |
| x | Internal/phenotypic states realization |
| y | External/environmental states realization |
| Posterior probability distribution of X given Y | |
| Likelihood of perceived environmental state given internal state | |
| Percieved probability distribution of X at time t | |
| Marginal percieved probability distribution of Y | |
| Real distribution of microenvironment Y | |
| True joint distribution between X and Y | |
| True conditional distribution of Y given X | |
| Relaxation time for adaptation | |
| Phenotypic displacement due to adaptation | |
| Drift velocity of the phenotypes during adaptation | |
| D | Diffusion constant (; from physical diffusion noise) |
| Standard deviation of noise from physical diffusion | |
| Standard deviation of noise from imperfect decision-making | |
| Proliferation (or growth) rate | |
| Average proliferation rate | |
| Deterministic term of environmental relation | |
| Gaussian distributed microenvironmental noise | |
| Intrinsic-extrinsic correlation coefficient | |
| Expected Y over true distribution of Y | |
| Expected Y over percieved distribution of Y | |
| Expected X over the distribution of X | |
| Standardized internal variable | |
| Standardized external variable | |
| Expected under the true conditional | |
| Signal-to-noise ratio (SNR) for internal variable | |
| Steady-state probability distribution of phenotypes | |
| Z | Normalization constant (partition function) |
2.1.1. Case-I: Cell State Adaptation Limit ( )
2.1.2. Case-II: Operating Around the Expected Phenotypic State Limit ()
2.2. Effective Fokker-Planck Equation for Phenotypic Dynamics
3. Results
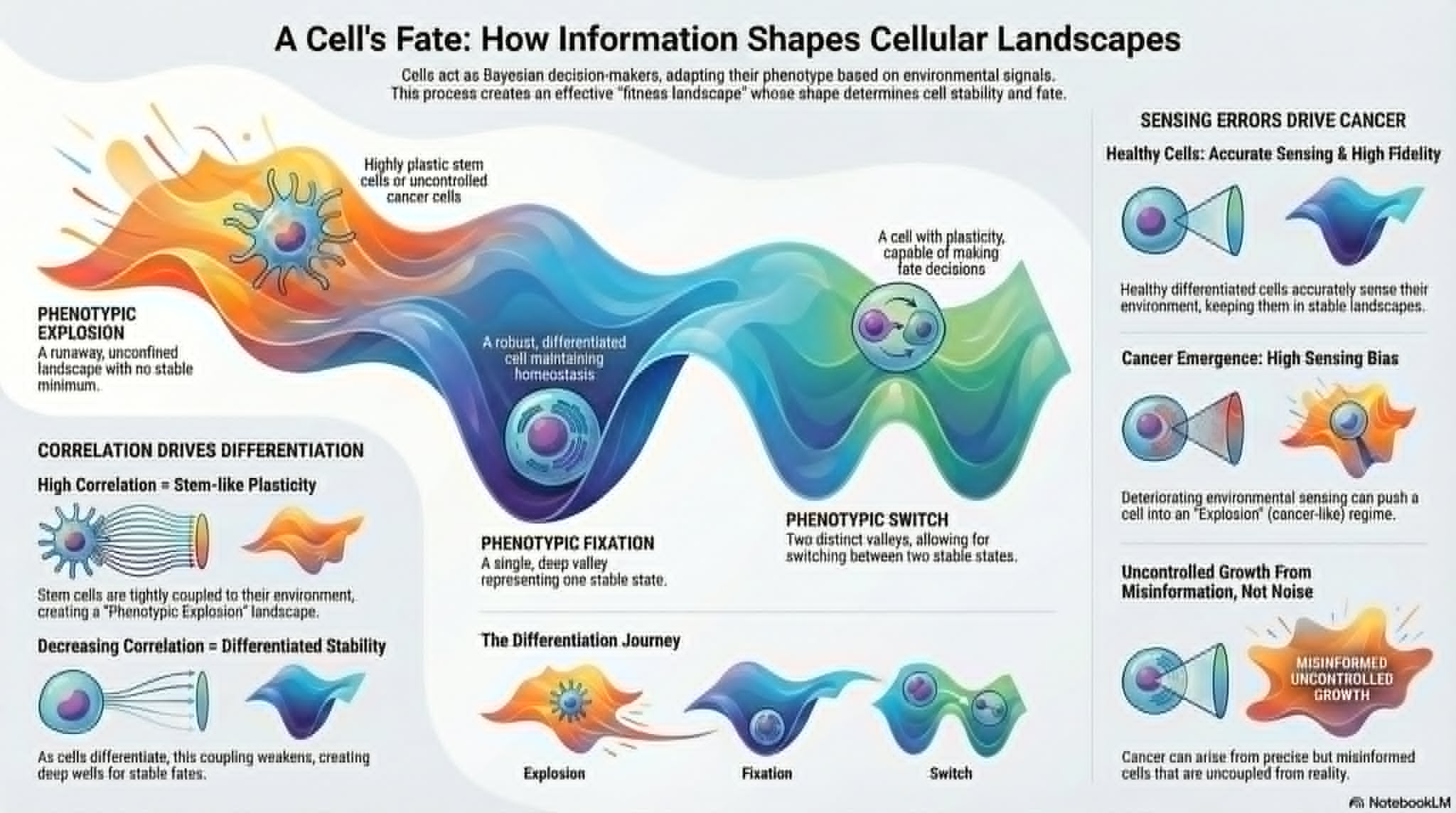
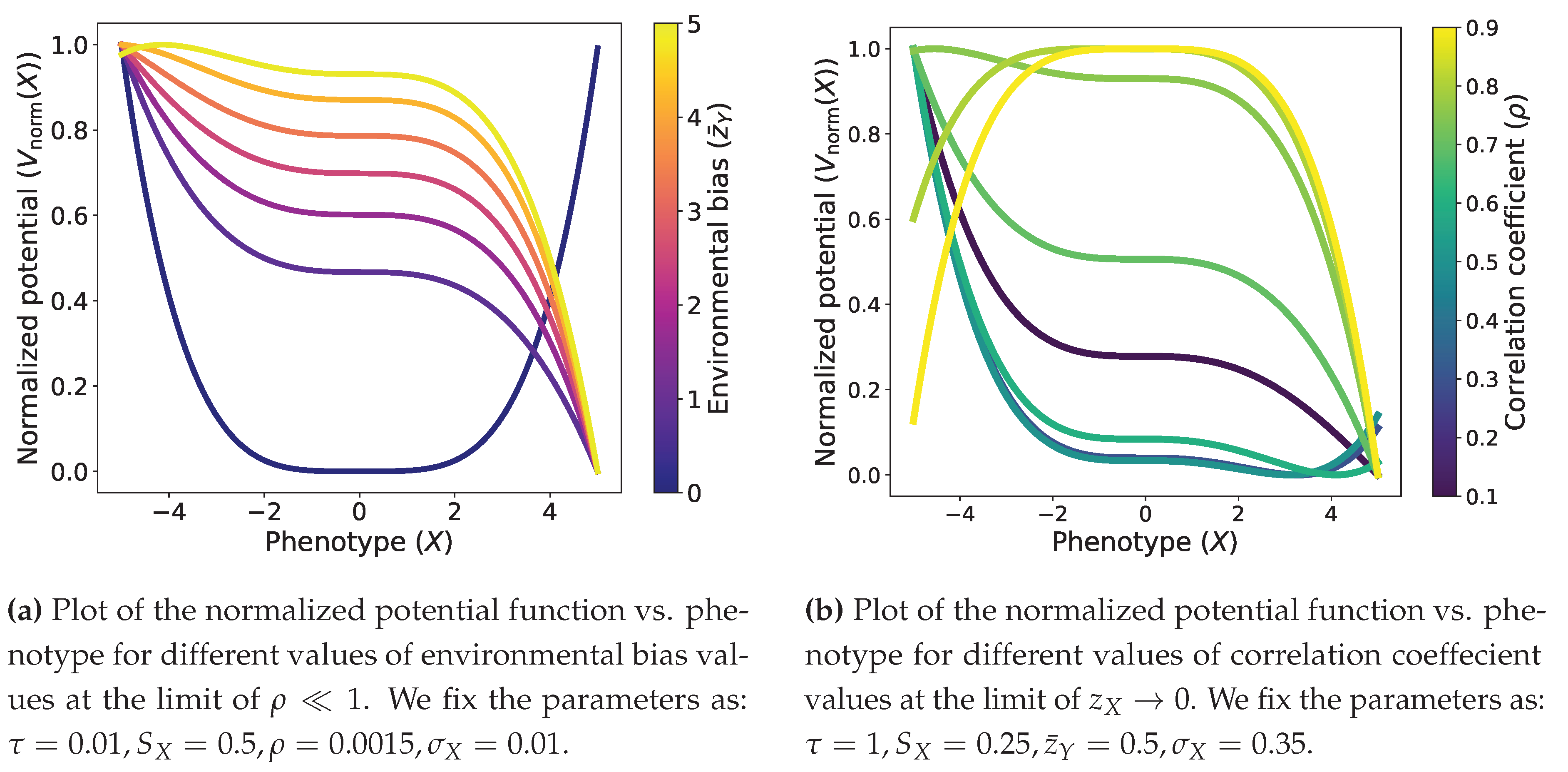
3.1. Bayesian Adaptation Shapes Phenotypic Fitness Landscapes
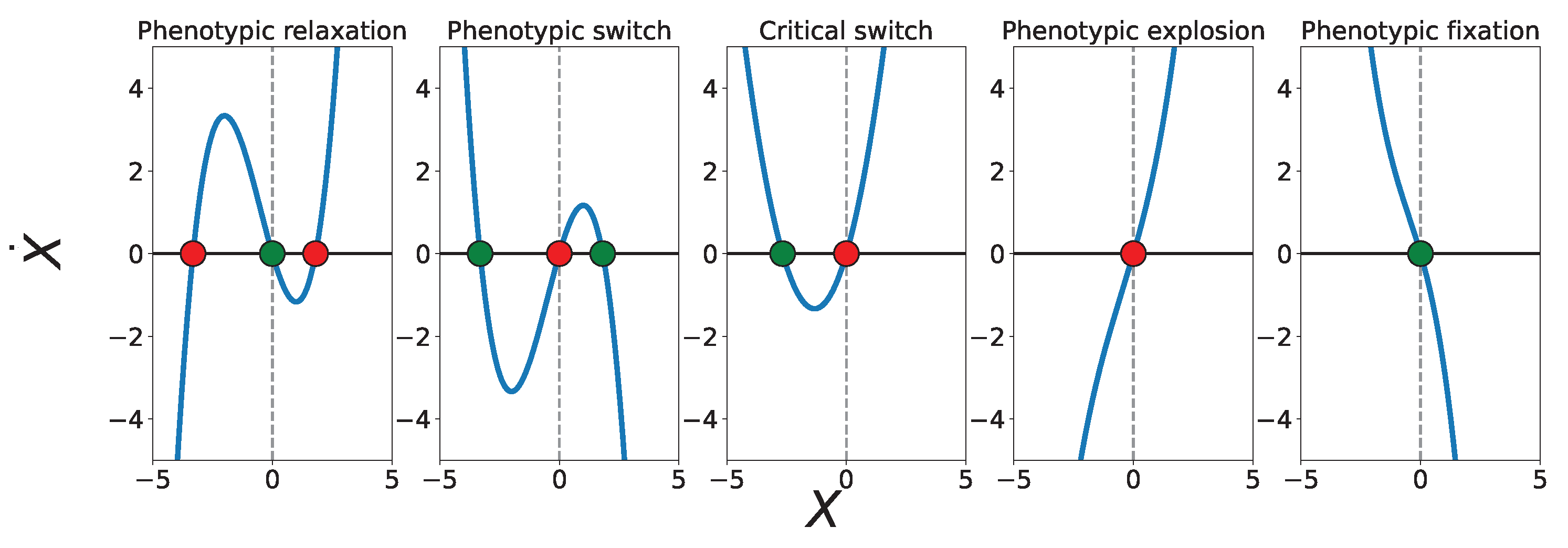
Fixed points and stability.
Classification of phenotypic landscapes.
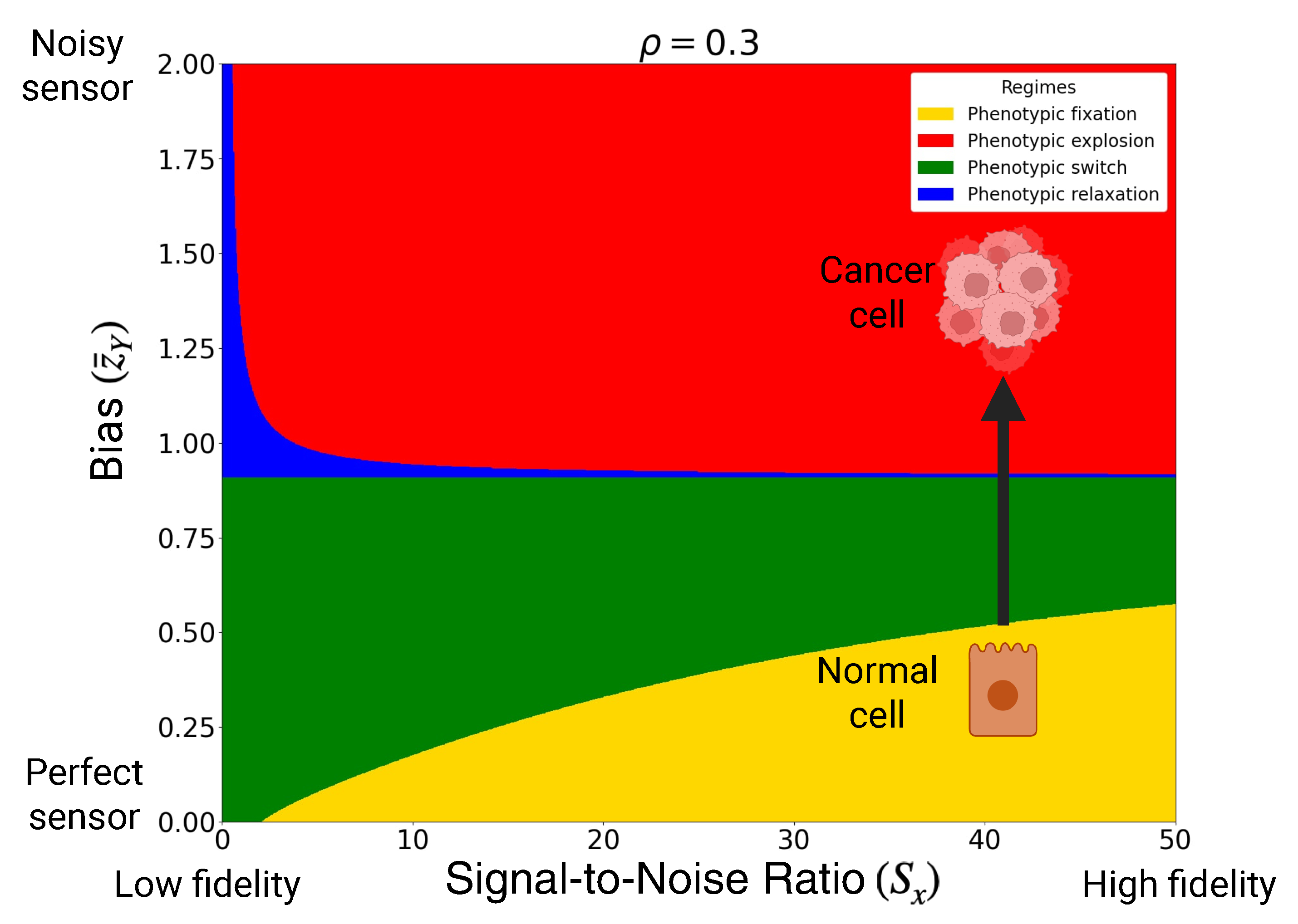
- Phenotypic fixation (homeostatic monostability): and . The potential has a single deep minimum at and all trajectories relax toward a stable homeostatic phenotype.
- Phenotypic switch (bistable landscape): and with . Two stable minima coexist and are separated by an unstable fixed point, enabling decision-like switching between discrete phenotypic states.
- Critical switch (marginal stability): or . One stable and one unstable fixed point coexist, corresponding to a near-critical landscape at the boundary between bistability and monostability.
- Phenotypic relaxation (canalizing monostability):, , and . The landscape possesses a single stable minimum, but nearby unstable fixed points generate slow relaxation and enhanced sensitivity to noise.
- Phenotypic explosion (runaway landscape): and with . The origin is unstable and the potential fails to confine the dynamics at large amplitudes, leading to runaway phenotypic amplification.
Role of the coefficients.
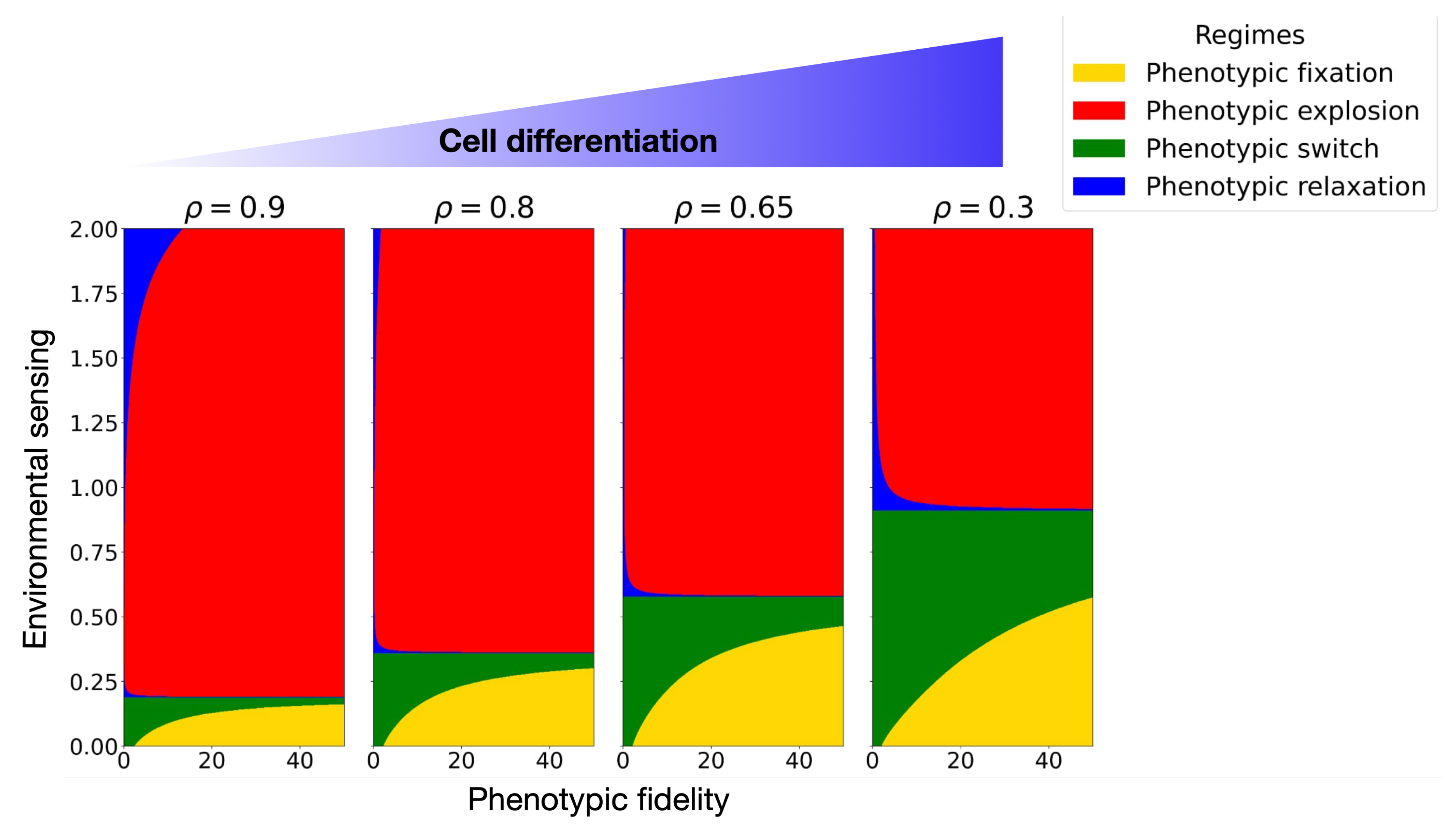
3.2. Correlation-Driven Emergence of Cell Fate Decisions: From Stem to Differentiated Cells
3.3. Microenvironmental Sensing Deterioration as Pathway to Cancer
3.4. Proliferation, Tissue Homeostasis and Carcinogenesis
Linear proliferation: the case .
- If , adaptation alone already stabilises . A range of values of preserves , corresponding to robust homeostasis.
- If , adaptation alone would make unstable. A suitable choice of can further enhance instability (make more positive).
Beyond linear proliferation: curvature and growth control.
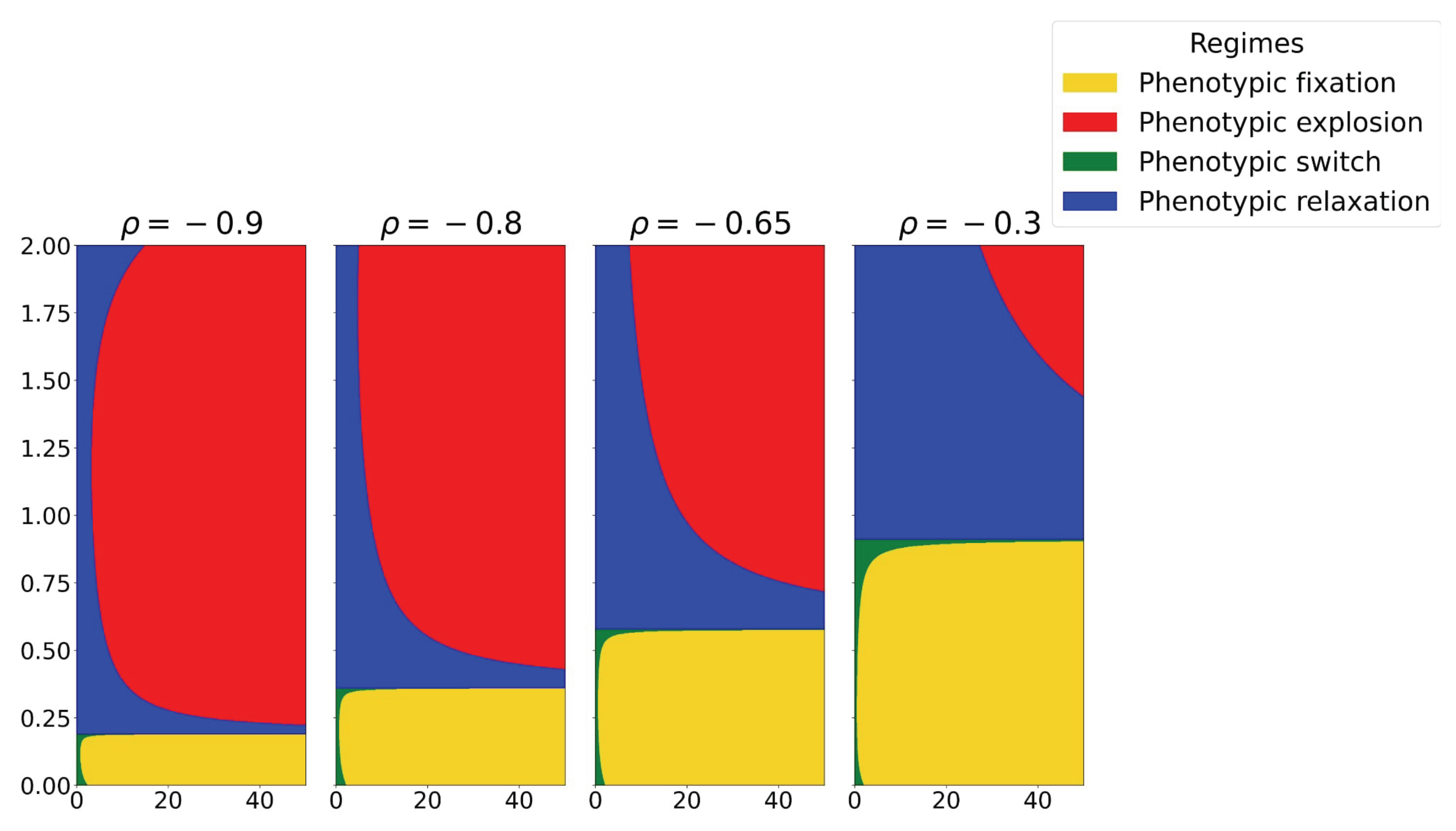
3.5. Negative Correlation-Driven Bifurcation: The Robustness-Plasticity Trade-Off
4. Discussion
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Simon, H.A. The new science of management decision.; Harper & Brothers: New York, 1960.
- Hatzikirou, H. Statistical mechanics of cell decision-making: the cell migration force distribution. J. Mech. Behav. Mater. 2018, 27, 1–7. [CrossRef]
- Pujar, A.A.; Barua, A.; Singh, D.; Roy, U.; Jolly, M.K.; Hatzikirou, H. Lattice-based microenvironmental uncertainty driven phenotypic decision-making: a comparison with Notch-Delta-Jagged signaling. bioRxiv 2021, p. 2021.11.16.468748. [CrossRef]
- Barua, A.; Nava-Sedeño, J.M.; Meyer-Hermann, M.; Hatzikirou, H. A least microenvironmental uncertainty principle (LEUP) as a generative model of collective cell migration mechanisms. Sci. Rep. 2020, 10, 22371. [CrossRef]
- Barua, A.; Beygi, A.; Hatzikirou, H. Close to Optimal Cell Sensing Ensures the Robustness of Tissue Differentiation Process: The Avian Photoreceptor Mosaic Case, 2021. [CrossRef]
- Barua, A.; Syga, S.; Mascheroni, P.; Kavallaris, N.; Meyer-Hermann, M.; Deutsch, A.; Hatzikirou, H. Entropy-driven cell decision-making predicts `fluid-to-solid’ transition in multicellular systems. New J. Phys. 2020, 22, 123034. [CrossRef]
- Perkins, T.J.; Swain, P.S. Strategies for cellular decision-making. Molecular Systems Biology 2009, 5, 326. [CrossRef]
- Balázsi, G.; van Oudenaarden, A.; Collins, J. Cellular Decision Making and Biological Noise: From Microbes to Mammals. Cell 2011, 144, 910–925. [CrossRef]
- Elowitz, M.B.; Levine, A.J.; Siggia, E.D.; Swain, P.S. Stochastic Gene Expression in a Single Cell. Science 2002, 297, 1183–1186. [CrossRef]
- Raj, A.; van Oudenaarden, A. Nature, Nurture, or Chance: Stochastic Gene Expression and Its Consequences. Cell 2008, 135, 216–226. [CrossRef]
- Alon, U. Network motifs: theory and experimental approaches. Nature Reviews Genetics 2007, 8, 450–461. [CrossRef]
- Bialek, W. Biophysics: Searching for Principles; Princeton University Press: Princeton, NJ, 2012.
- Shyam, S.; S, N.N.; Anand, V.; Jolly, M.K.; Hari, K. Mutually inhibiting teams of nodes: A predictive framework for structure–dynamics relationships in gene regulatory networks. Physical Biology 2025, 22, 066008. [CrossRef]
- BV, H.; Billakurthi, H.S.; Adigwe, S.; Hari, K.; Levine, H.; Gedeon, T.; Jolly, M.K. Emergent dynamics of cellular decision making in multi-node mutually repressive regulatory networks. Journal of The Royal Society Interface 2025, 22, 20250190, [https://royalsocietypublishing.org/rsif/article-pdf/doi/10.1098/rsif.2025.0190/2826204/rsif.2025.0190.pdf]. [CrossRef]
- Jolly, M.K.; Boareto, M.; Lu, M.; Onuchic, J.N.; Clementi, C.; Ben-Jacob, E. Operating principles of Notch–Delta–Jagged module of cell–cell communication. New Journal of Physics 2015, 17, 055021. [CrossRef]
- Ito, K.; Ito, K. Metabolism and the Control of Cell Fate Decisions and Stem Cell Renewal. Annual Review of Cell and Developmental Biology 2016, 32, 399–409. [CrossRef]
- Chu, X.; Wang, J. Insights into the cell fate decision-making processes from chromosome structural reorganizations. Biophysics Reviews 2022, 3, 041402. [CrossRef]
- Zorzan, I.; López, A.R.; Malyshava, A.; Ellis, T.; Barberis, M. Synthetic designs regulating cellular transitions: Fine-tuning of switches and oscillators. Current Opinion in Systems Biology 2021, 25, 11–26. [CrossRef]
- Li, S.; Liu, Q.; Wang, E.; Wang, J. Global quantitative understanding of non-equilibrium cell fate decision-making in response to pheromone. iScience 2023, 26, 107885. [CrossRef]
- Cheong, R.; Rhee, A.; Wang, C.J.; Nemenman, I.; Levchenko, A. Information Transduction Capacity of Noisy Biochemical Signaling Networks. Science 2011, 334, 354–358. [CrossRef]
- Huang, S.; Eichler, G.; Bar-Yam, Y.; Ingber, D.E. Cell Fates as High-Dimensional Attractor States of a Complex Gene Regulatory Network. Phys. Rev. Lett. 2005, 94, 128701. [CrossRef]
- Yao, Y.; Wang, C. Dedifferentiation: inspiration for devising engineering strategies for regenerative medicine. npj Regenerative Medicine 2020, 5. [CrossRef]
- Takahashi, K.; Yamanaka, S. Induction of Pluripotent Stem Cells from Mouse Embryonic and Adult Fibroblast Cultures by Defined Factors. Cell 2006, 126, 663–676. [CrossRef]
- Barua, A.; Hatzikirou, H. Cell Decision Making through the Lens of Bayesian Learning. Entropy 2023, 25, 609, [2301.06941]. [CrossRef]
- Sourjik, V.; Berg, H.C. Receptor sensitivity in bacterial chemotaxis. Proceedings of the National Academy of Sciences 2002, 99, 123–127, [https://www.pnas.org/content/99/1/123.full.pdf]. [CrossRef]
- Ueda, M.; Shibata, T. Stochastic Signal Processing and Transduction in Chemotactic Response of Eukaryotic Cells. Biophys, J. 2007, 93, 11–20. [CrossRef]
- Lancaster, H.O. The Structure of Bivariate Distributions. Annals of Mathematical Statistics 1958, 29, 719–736. [CrossRef]
- Hoeffding, W. A Class of Statistics with Asymptotically Normal Distribution. Annals of Mathematical Statistics 1948, 19, 293–325. [CrossRef]
- NIST Digital Library of Mathematical Functions. Hermite Polynomials. https://dlmf.nist.gov/18, 2024. Accessed: 2024-05-22.
- Risken, H. The Fokker-Planck equation: methods of solution and applications, 2. ed., 3rd printing ed.; Springer series in synergetics, Springer: Berlin, 1996; p. 472. 18.
- Hanahan, D. Hallmarks of Cancer: New Dimensions. Cancer Discovery 2022, 12, 31–46. [CrossRef]
- Bowsher, C.G.; Swain, P.S. Environmental sensing, information transfer, and cellular decision-making. Current Opinion in Biotechnology 2014, 28, 149–155. [CrossRef]
- McClatchey, A.I.; Yap, A.S. Contact inhibition (of proliferation) redux. Current Opinion in Cell Biology 2012, 24, 685–694. [CrossRef]
- Cognet, G.; Muir, A. Identifying metabolic limitations in the tumor microenvironment. Science Advances 2024, 10, 37–39. [CrossRef]
- Boareto, M.; Jolly, M.K.; Lu, M.; Onuchic, J.N.; Clementi, C.; Ben-Jacob, E. Jagged–Delta asymmetry in Notch signaling can give rise to a Sender/Receiver hybrid phenotype. Proceedings of the National Academy of Sciences 2015, 112, E402–E409. [CrossRef]
- Antebi, Y.E.; Reich-Zeliger, S.; Hart, Y.; Mayo, A.; Eizenberg, I.; Rimer, J.; Putheti, P.; Pe’er, D.; Friedman, N. Mapping differentiation under mixed culture conditions reveals a tunable continuum of T cell fates. PLoS biology 2013, 11. [CrossRef]
- Tomasetti, C.; Vogelstein, B. Variation in cancer risk among tissues can be explained by the number of stem cell divisions. Science 2015, 347, 78–81. [CrossRef]
- Nowak, D.; Stewart, D.; Koeffler, H.P. Differentiation therapy of leukemia: 3 Decades of development. Blood 2009, 113, 3655–3665. [CrossRef]
- Cao, M.; Nawalaniec, K.; Ajay, A.K.; Luo, Y.; Moench, R.; Jin, Y.; Xiao, S.; Hsiao, L.L.; Waaga-Gasser, A.M. PDE4D targeting enhances anti-tumor effects of sorafenib in clear cell renal cell carcinoma and attenuates MAPK/ERK signaling in a CRAF-dependent manner. Translational Oncology 2022, 19, 101377. [CrossRef]
- Vinyard, M.E.; Rasmussen, A.W.; Li, R.; Klein, A.M.; Getz, G.; Pinello, L. Learning cell dynamics with neural differential equations. Nature Machine Intelligence 2025, 7, 1969–1984. [CrossRef]
- Barua, A.; Hatzikirou, H.; Abe, S. Geodesic learning. Physica A: Statistical Mechanics and its Applications 2025, 669, 130539. [CrossRef]
| Phenotypic regime | Potential landscape | ||||
| Phenotypic relaxation | any | Two barriers, finite well | |||
| Phenotypic switch | any | Two wells, finite barrier | |||
| Critical switch | any | any | Vanishing barrier | ||
| Phenotypic explosion | Runaway, no minimum | ||||
| Phenotypic fixation | Single deep global well |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




