Submitted:
02 January 2026
Posted:
04 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
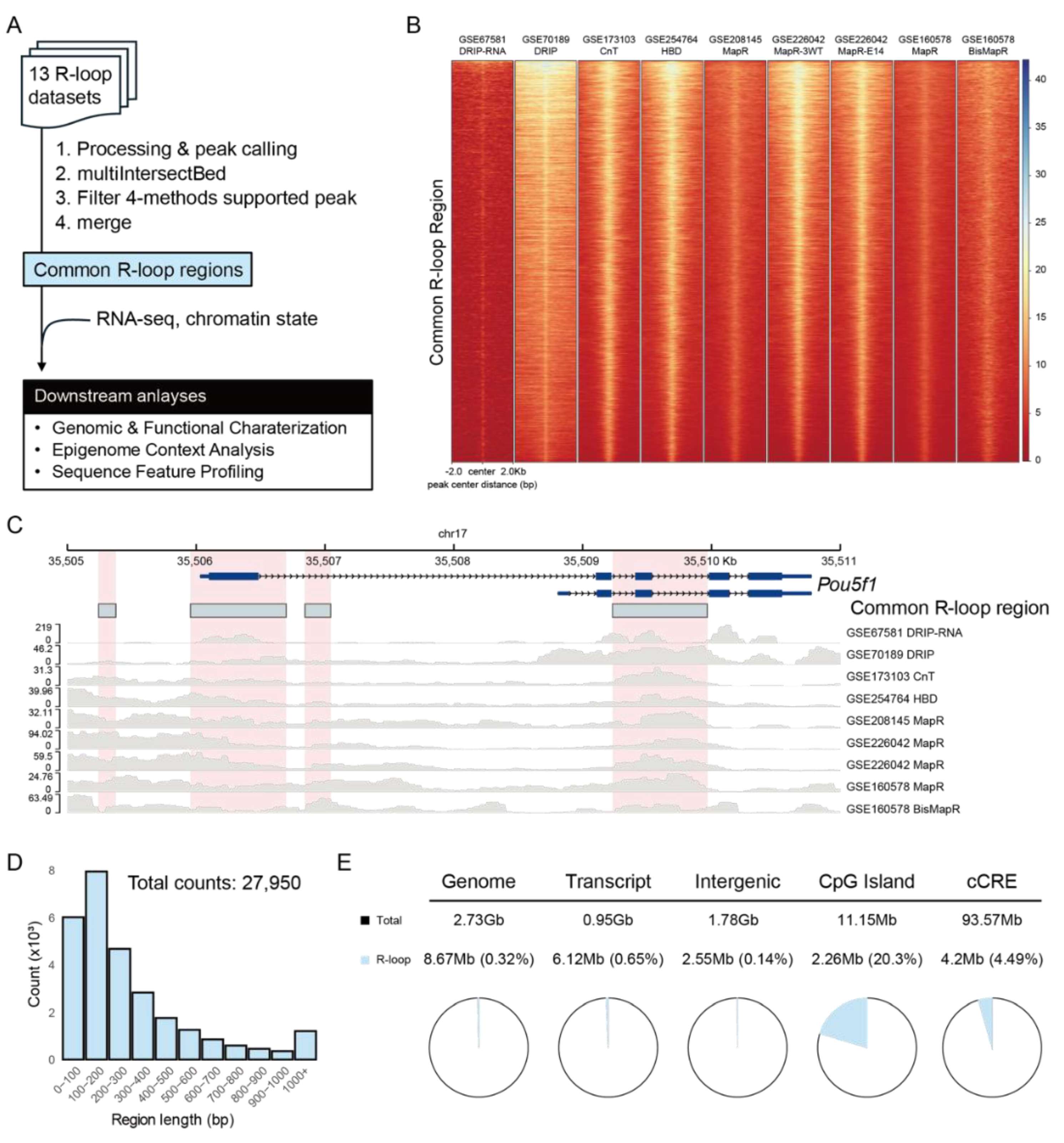
2.1. Integration of R-Loop Datasets Defines Common R-Loop Regions
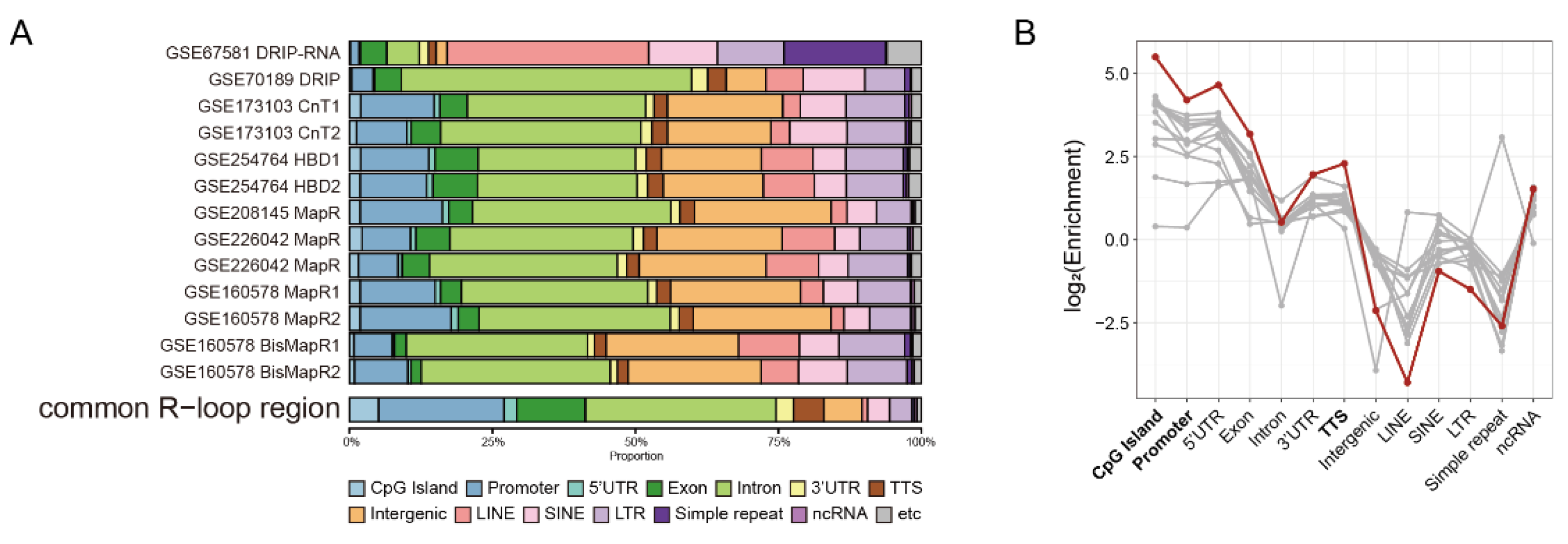
2.2. Common R-Loops Are Preferentially Enriched at Genic Regions
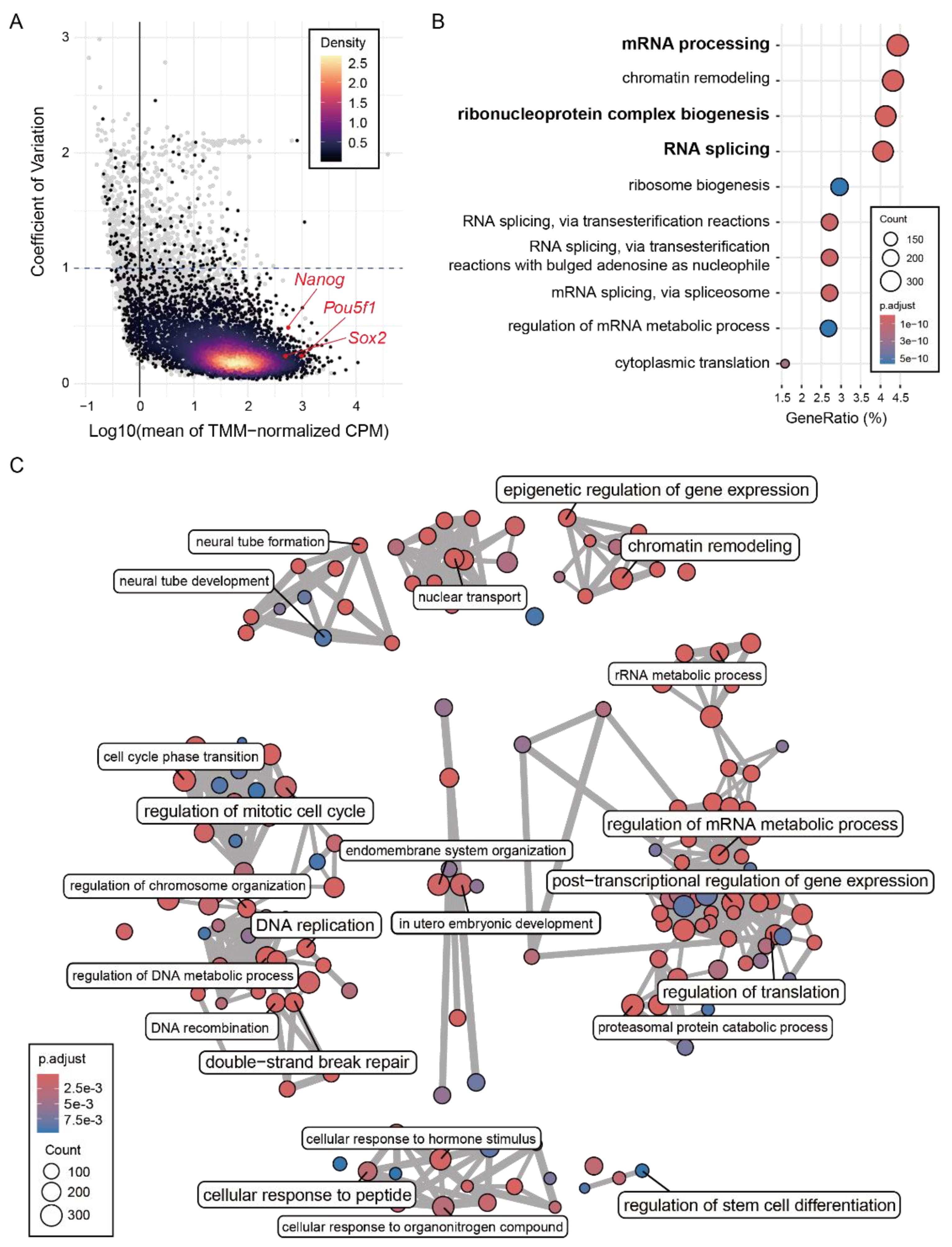
2.3. Common R-Loops Are Associated with Highly Expressed RNA-Regulatory Genes
2.4. Common R-Loops Exhibit Uniform Positional Patterns Independent of Gene Function
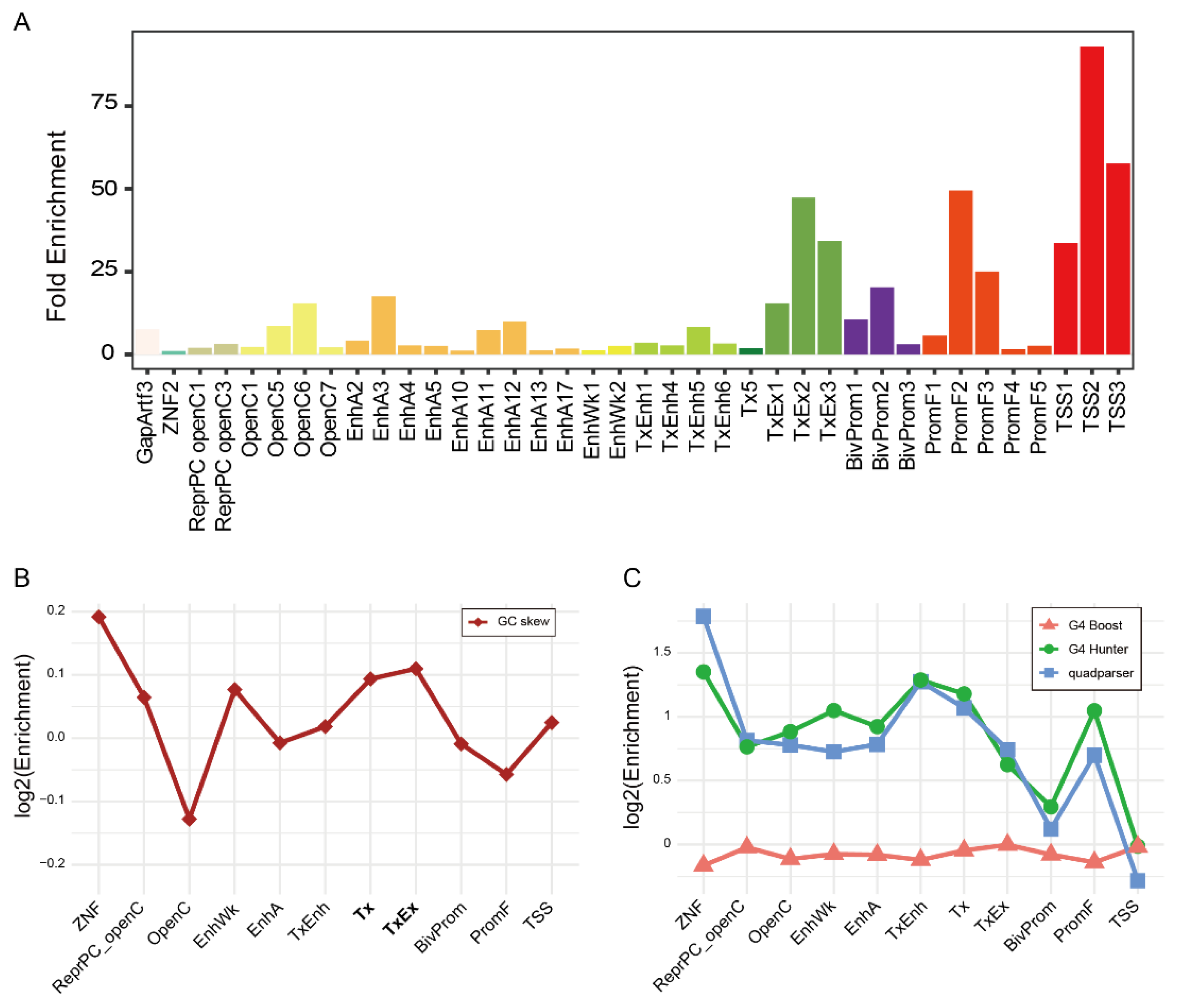
2.5. Common R-Loops Are Enriched in Transcriptionally Active and Regulatory Chromatin States
2.6. GC Skew as a Sequence Signature of R-Loops in Transcribed Chromatin States
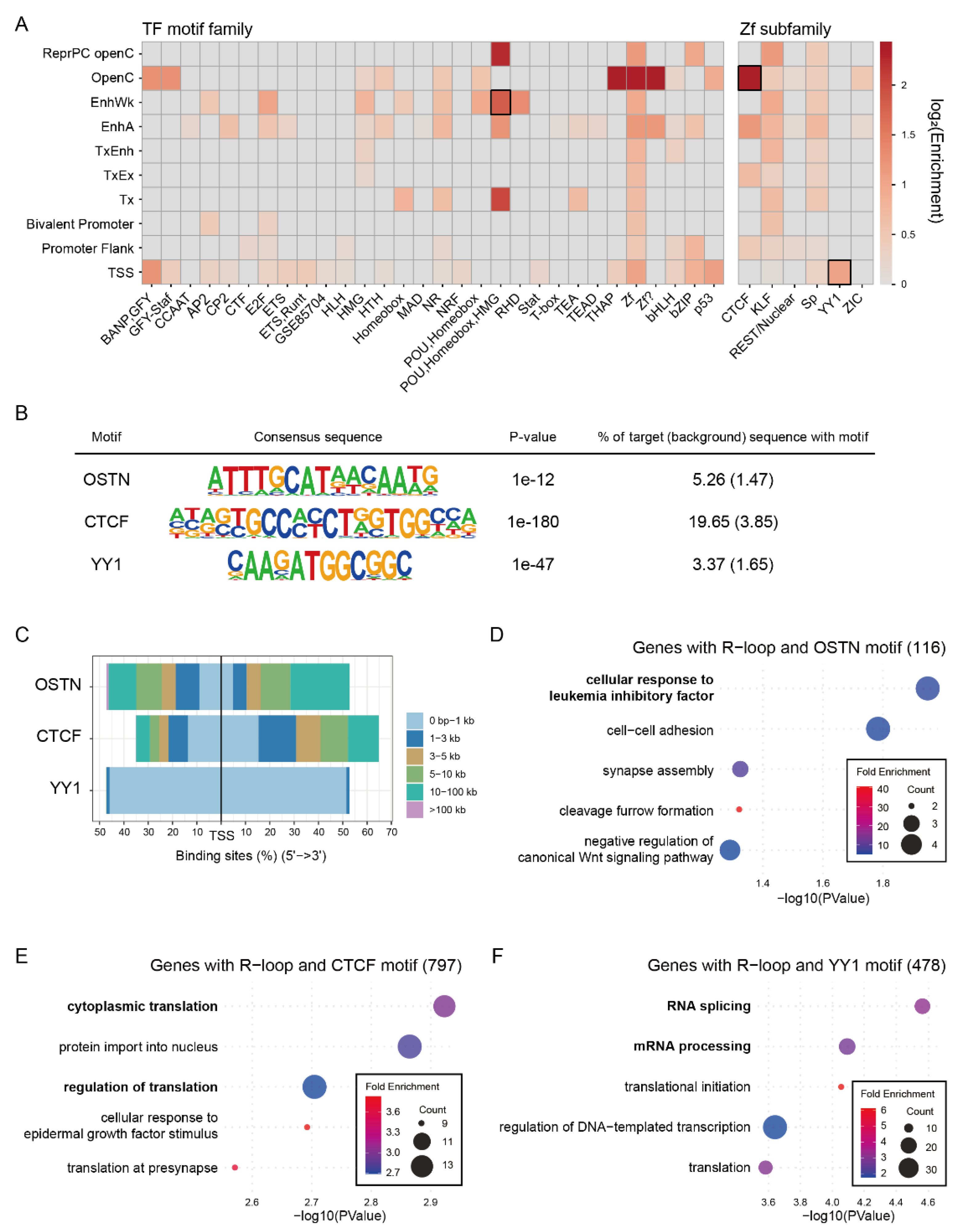
2.7. Chromatin State-Dependent Motif Enrichment Delineates the Regulatory Context of Common R-Loop Regions
3. Discussion
4. Materials and Methods
Curation of R-Loop Datasets
Public Data Processing
Identification of Common R-Loop Regions
Gene Annotation and Functional Enrichment Analysis
Chromatin State Annotation
Construction of Chromatin State-Specific R-Loop Rich Regions
Nucleotide Skewness Analysis
G-Quadruplex Motif Analysis
Transcription Factor Motif Enrichment Analysis
Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| mESC | Mouse embryonic stem cell |
| DRIP | DNA-RNA immunoprecipitation |
| HBD | Hybrid binding domain |
| GO | Gene ontology |
| GOBP | Gene ontology biological process |
| GOCC | Gene ontology cellular component |
| GOMF | Gene ontology molecular function |
| cCRE | Candidate cis regulatory element |
| TTS | Transcription termination site |
| LINE | Long interspersed nuclear element |
| SINE | Short interspersed nuclear element |
| LTR | Long terminal repeat |
| ncRNA | Non-coding RNA |
| TSS | Transcription start site |
| PromF | Promoter flank |
| BivProm | Bivalent promoter |
| TxEx | Transcription and exon |
| Tx | Transcription |
| TxEnh | Transcribed enhancer |
| EnhA | Active enhancer |
| OpenC | Open chromatin |
| ReprPC openC | Polycomb repressed and open chromatin |
| ZNF | Zinc finger genes |
| GapArtf | Assembly gaps and artifacts |
| OSTN | OCT4-SOX2-TCF-NANOG |
| NR | Nuclear receptor |
| Zf | Zinc finger |
References
- Thomas, M.; White, R.L.; Davis, R.W. Hybridization of RNA to double-stranded DNA: formation of R-loops. Proc Natl Acad Sci U S A 1976, 73, 2294–2298. [Google Scholar] [CrossRef]
- White, R.L.; Hogness, D.S. R loop mapping of the 18S and 28S sequences in the long and short repeating units of Drosophila melanogaster rDNA. Cell 1977, 10, 177–192. [Google Scholar] [CrossRef] [PubMed]
- Aguilera, A.; Garcia-Muse, T. R loops: from transcription byproducts to threats to genome stability. Mol Cell 2012, 46, 115–124. [Google Scholar] [CrossRef]
- Santos-Pereira, J.M.; Aguilera, A. R loops: new modulators of genome dynamics and function. Nat Rev Genet 2015, 16, 583–597. [Google Scholar] [CrossRef]
- Petermann, E.; Lan, L.; Zou, L. Sources, resolution and physiological relevance of R-loops and RNA-DNA hybrids. Nat Rev Mol Cell Biol 2022, 23, 521–540. [Google Scholar] [CrossRef]
- Lee, S.Y.; Kwak, M.J.; Kim, J.J. R-loops: a key driver of inflammatory responses in cancer. Exp Mol Med 2025, 57, 1455–1466. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; et al. R-loop and diseases: the cell cycle matters. Mol Cancer 2024, 23, 84. [Google Scholar] [CrossRef]
- Lee, H.G.; et al. Site-specific R-loops induce CGG repeat contraction and fragile X gene reactivation. Cell 2023, 186, 2593–2609 e18. [Google Scholar] [CrossRef]
- Zhang, S.; et al. Aberrant R-loop-mediated immune evasion, cellular communication, and metabolic reprogramming affect cancer progression: a single-cell analysis. Mol Cancer 2024, 23, 11. [Google Scholar] [CrossRef]
- Ouyang, Y.; et al. Dysregulation of R-loop homeostasis shapes the immunosuppressive microenvironment and induces malignant progression in melanoma. Apoptosis 2025, 30, 131–148. [Google Scholar] [CrossRef]
- Xu, C.; et al. R-loop-dependent promoter-proximal termination ensures genome stability. Nature 2023, 621, 610–619. [Google Scholar]
- Castillo-Guzman, D.; Chedin, F. Defining R-loop classes and their contributions to genome instability. DNA Repair (Amst) 2021, 106, 103182. [Google Scholar] [CrossRef] [PubMed]
- Moore, M.J.; Proudfoot, N.J. Pre-mRNA processing reaches back to transcription and ahead to translation. Cell 2009, 136, 688–700. [Google Scholar] [CrossRef] [PubMed]
- Allison, D.F.; Wang, G.G. R-loops: formation, function, and relevance to cell stress. Cell Stress 2019, 3, 38–46. [Google Scholar] [CrossRef]
- Luna, R.; Gomez-Gonzalez, B.; Aguilera, A. RNA biogenesis and RNA metabolism factors as R-loop suppressors: a hidden role in genome integrity. Genes Dev 2024, 38, 504–527. [Google Scholar] [CrossRef]
- Xu, C.; et al. R-loop resolution promotes co-transcriptional chromatin silencing. Nat Commun 2021, 12, 1790. [Google Scholar] [CrossRef]
- Qiu, Y.; et al. R-loops’ m6A modification and its roles in cancers. Mol Cancer 2024, 23, 232. [Google Scholar] [CrossRef] [PubMed]
- Chedin, F. Nascent Connections: R-Loops and Chromatin Patterning. Trends Genet 2016, 32, 828–838. [Google Scholar] [CrossRef]
- Kim, A.; Wang, G.G. R-loop and its functions at the regulatory interfaces between transcription and (epi)genome. Biochim Biophys Acta Gene Regul Mech 2021, 1864, 194750. [Google Scholar] [CrossRef]
- Goulielmaki, E.; et al. The splicing factor XAB2 interacts with ERCC1-XPF and XPG for R-loop processing. Nat Commun 2021, 12, 3153. [Google Scholar] [CrossRef]
- Jin, B.; et al. MEN1 is a regulator of alternative splicing and prevents R-loop-induced genome instability through suppression of RNA polymerase II elongation. Nucleic Acids Res 2023, 51, 7951–7971. [Google Scholar] [CrossRef]
- Stratigi, K.; Siametis, A.; Garinis, G.A. Looping forward: exploring R-loop processing and therapeutic potential. FEBS Lett 2025, 599, 244–266. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; et al. The Augmented R-Loop Is a Unifying Mechanism for Myelodysplastic Syndromes Induced by High-Risk Splicing Factor Mutations. Mol Cell 2018, 69, 412–425 e6. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.S.; et al. R-Loop Accumulation in Spliceosome Mutant Leukemias Confers Sensitivity to PARP1 Inhibition by Triggering Transcription-Replication Conflicts. Cancer Res 2024, 84, 577–597. [Google Scholar] [CrossRef]
- Chakraborty, P.; Huang, J.T.J.; Hiom, K. DHX9 helicase promotes R-loop formation in cells with impaired RNA splicing. Nat Commun 2018, 9, 4346. [Google Scholar] [CrossRef]
- Bhandari, J.; et al. The molecular chaperone ALYREF promotes R-loop resolution and maintains genome stability. J Biol Chem 2024, 300, 107996. [Google Scholar] [CrossRef]
- Merida-Cerro, J.A.; et al. Cellular imbalance of specific RNA-binding proteins associates with harmful R-loops. PLoS Genet 2025, 21, e1011491. [Google Scholar] [CrossRef]
- Jayakumar, S.; et al. PSIP1/LEDGF reduces R-loops at transcription sites to maintain genome integrity. Nat Commun 2024, 15, 361. [Google Scholar] [CrossRef]
- Belotserkovskii, B.P.; et al. R-loop generation during transcription: Formation, processing and cellular outcomes. DNA Repair (Amst) 2018, 71, 69–81. [Google Scholar] [CrossRef]
- Ginno, P.A.; et al. R-loop formation is a distinctive characteristic of unmethylated human CpG island promoters. Mol Cell 2012, 45, 814–825. [Google Scholar] [CrossRef]
- Sanz, L.A.; Chedin, F. High-resolution, strand-specific R-loop mapping via S9.6-based DNA-RNA immunoprecipitation and high-throughput sequencing. Nat Protoc 2019, 14, 1734–1755. [Google Scholar] [CrossRef]
- Yan, Q.; et al. Mapping Native R-Loops Genome-wide Using a Targeted Nuclease Approach. Cell Rep 2019, 29, 1369–1380 e5. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; et al. Genomic profiling of native R loops with a DNA-RNA hybrid recognition sensor. Sci Adv 2021, 7. [Google Scholar] [CrossRef] [PubMed]
- Wulfridge, P.; Sarma, K. A nuclease- and bisulfite-based strategy captures strand-specific R-loops genome-wide. Elife 2021, 10. [Google Scholar] [CrossRef] [PubMed]
- Chen, P.B.; et al. R loops regulate promoter-proximal chromatin architecture and cellular differentiation. Nat Struct Mol Biol 2015, 22, 999–1007. [Google Scholar] [CrossRef]
- Miller, H.E.; et al. Quality-controlled R-loop meta-analysis reveals the characteristics of R-loop consensus regions. Nucleic Acids Res 2022, 50, 7260–7286. [Google Scholar] [CrossRef]
- Yu, K.; et al. R-loops at immunoglobulin class switch regions in the chromosomes of stimulated B cells. Nat Immunol 2003, 4, 442–451. [Google Scholar] [CrossRef]
- Duquette, M.L.; et al. Intracellular transcription of G-rich DNAs induces formation of G-loops, novel structures containing G4 DNA. Genes Dev 2004, 18, 1618–1629. [Google Scholar] [CrossRef]
- Chen, L.; et al. R-ChIP Using Inactive RNase H Reveals Dynamic Coupling of R-loops with Transcriptional Pausing at Gene Promoters. Mol Cell 2017, 68, 745–757 e5. [Google Scholar] [CrossRef]
- McLean, C.Y.; et al. GREAT improves functional interpretation of cis-regulatory regions. Nat Biotechnol 2010, 28, 495–501. [Google Scholar] [CrossRef]
- Garcia-Muse, T.; Aguilera, A. R Loops: From Physiological to Pathological Roles. Cell 2019, 179, 604–618. [Google Scholar] [CrossRef]
- Ginno, P.A.; et al. GC skew at the 5’ and 3’ ends of human genes links R-loop formation to epigenetic regulation and transcription termination. Genome Res 2013, 23, 1590–1600. [Google Scholar] [CrossRef] [PubMed]
- Vu, H.; Ernst, J. Universal chromatin state annotation of the mouse genome. Genome Biol 2023, 24, 153. [Google Scholar] [CrossRef] [PubMed]
- Wulfridge, P.; et al. G-quadruplexes associated with R-loops promote CTCF binding. Mol Cell 2023, 83, 3064–3079 e5. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; et al. YY1 interacts with guanine quadruplexes to regulate DNA looping and gene expression. Nat Chem Biol 2021, 17, 161–168. [Google Scholar] [CrossRef]
- Wulfridge, P.; Sarma, K. Intertwining roles of R-loops and G-quadruplexes in DNA repair, transcription and genome organization. Nat Cell Biol 2024, 26, 1025–1036. [Google Scholar] [CrossRef]
- Chambers, I.; Tomlinson, S.R. The transcriptional foundation of pluripotency. Development 2009, 136, 2311–2322. [Google Scholar] [CrossRef]
- Niwa, H.; et al. A parallel circuit of LIF signalling pathways maintains pluripotency of mouse ES cells. Nature 2009, 460, 118–122. [Google Scholar] [CrossRef]
- Mosler, T.; et al. R-loop proximity proteomics identifies a role of DDX41 in transcription-associated genomic instability. Nat Commun 2021, 12, 7314. [Google Scholar] [CrossRef]
- Wu, T.; et al. Characterization of R-Loop-Interacting Proteins in Embryonic Stem Cells Reveals Roles in rRNA Processing and Gene Expression. Mol Cell Proteomics 2021, 20, 100142. [Google Scholar] [CrossRef]
- Yan, Q.; et al. Proximity labeling identifies a repertoire of site-specific R-loop modulators. Nat Commun 2022, 13, 53. [Google Scholar] [CrossRef] [PubMed]
- Beagan, J.A.; et al. YY1 and CTCF orchestrate a 3D chromatin looping switch during early neural lineage commitment. Genome Res 2017, 27, 1139–1152. [Google Scholar] [CrossRef]
- Weintraub, A.S.; et al. YY1 Is a Structural Regulator of Enhancer-Promoter Loops. Cell 2017, 171, 1573–1588 e28. [Google Scholar] [CrossRef] [PubMed]
- Khan, E.S.; Danckwardt, S. Pathophysiological Role and Diagnostic Potential of R-Loops in Cancer and Beyond. Genes (Basel) 2022, 13. [Google Scholar] [CrossRef]
- Gatti, V.; et al. Senataxin and R-loops homeostasis: multifaced implications in carcinogenesis. Cell Death Discov 2023, 9, 145. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; et al. Unraveling R-loops: The hidden drivers of inflammation and immune dysregulation. Medicine (Baltimore) 2025, 104, e42833. [Google Scholar] [CrossRef]
- Sanz, L.A.; et al. Prevalent, Dynamic, and Conserved R-Loop Structures Associate with Specific Epigenomic Signatures in Mammals. Mol Cell 2016, 63, 167–178. [Google Scholar] [CrossRef]
- Lyu, J.; et al. Genome-wide mapping of G-quadruplex structures with CUT&Tag. Nucleic Acids Res 2022, 50, e13. [Google Scholar]
- Huang, X.; et al. ZFP281 controls transcriptional and epigenetic changes promoting mouse pluripotent state transitions via DNMT3 and TET1. Dev Cell 2024, 59, 465–481 e6. [Google Scholar] [CrossRef]
- Liu, T.; et al. Genome-wide mapping of native co-localized G4s and R-loops in living cells. Elife 2024, 13. [Google Scholar] [CrossRef]
- Chen, S.; et al. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef]
- Chen, S. Ultrafast one-pass FASTQ data preprocessing, quality control, and deduplication using fastp. Imeta 2023, 2, e107. [Google Scholar] [CrossRef]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat Methods 2012, 9, 357–359. [Google Scholar] [CrossRef]
- Danecek, P.; et al. Twelve years of SAMtools and BCFtools. Gigascience 2021, 10. [Google Scholar] [CrossRef]
- Quinlan, A.R.; Hall, I.M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 2010, 26, 841–842. [Google Scholar] [CrossRef] [PubMed]
- Dobin, A.; et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Ramirez, F.; et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res 2016, 44, W160–5. [Google Scholar] [CrossRef]
- Zhang, Y.; et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol 2008, 9, R137. [Google Scholar] [CrossRef]
- Liao, Y.; Smyth, G.K.; Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef] [PubMed]
- Yu, G.; Wang, L.G.; He, Q.Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 2015, 31, 2382–2383. [Google Scholar] [CrossRef]
- Yu, G.; et al. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 2012, 16, 284–287. [Google Scholar] [CrossRef]
- Huang da, W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 2009, 4, 44–57. [Google Scholar] [CrossRef]
- Sherman, B.T.; et al. DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res 2022, 50, W216–W221. [Google Scholar] [CrossRef]
- Ernst, J.; Kellis, M. Chromatin-state discovery and genome annotation with ChromHMM. Nat Protoc 2017, 12, 2478–2492. [Google Scholar] [CrossRef] [PubMed]
- Huppert, J.L.; Balasubramanian, S. Prevalence of quadruplexes in the human genome. Nucleic Acids Res 2005, 33, 2908–2916. [Google Scholar] [CrossRef]
- Bedrat, A.; Lacroix, L.; Mergny, J.L. Re-evaluation of G-quadruplex propensity with G4Hunter. Nucleic Acids Res 2016, 44, 1746–1759. [Google Scholar] [CrossRef] [PubMed]
- Cagirici, H.B.; Budak, H.; Sen, T.Z. G4Boost: a machine learning-based tool for quadruplex identification and stability prediction. BMC Bioinformatics 2022, 23, 240. [Google Scholar] [CrossRef]
- Heinz, S.; et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol Cell 2010, 38, 576–589. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



