Submitted:
31 December 2025
Posted:
01 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Conjugation of NH₂-Functionalized Quantum Dots with Thiolated DNA Probes (DPP/DP-SH or AP-SH)
2.2. Immobilization of Thiolated Capture DNA on NH₂-Functionalized Glass Slides (CPP/CP-SH)
2.3. Dengue DNA or RNA Detection
3. Results
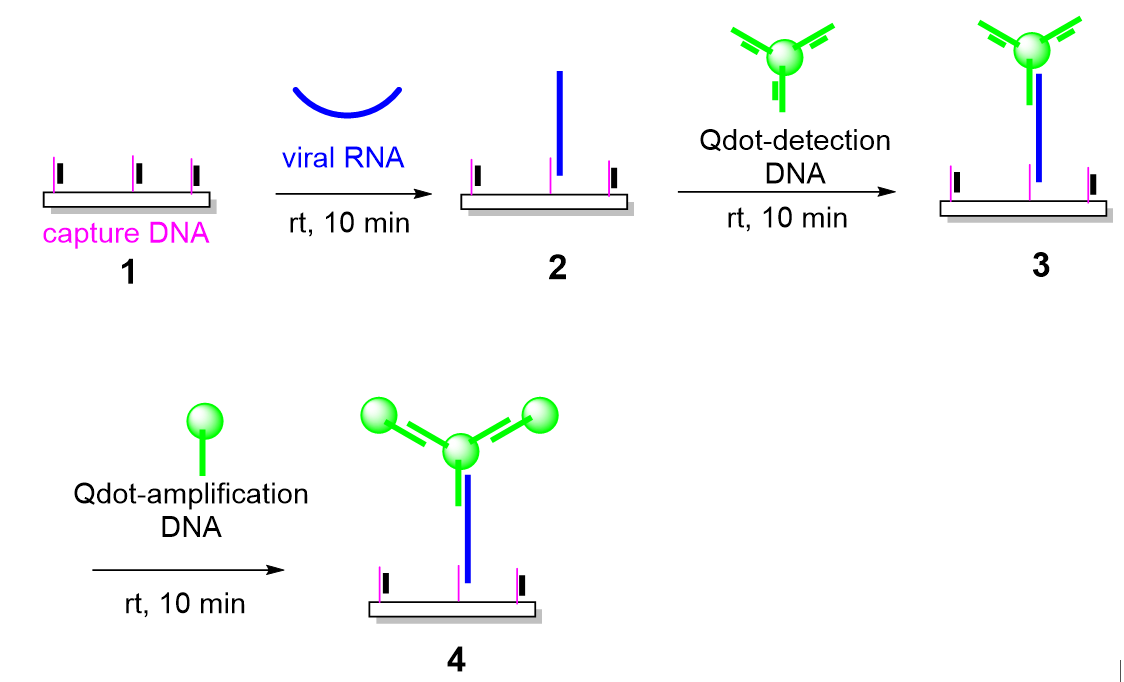
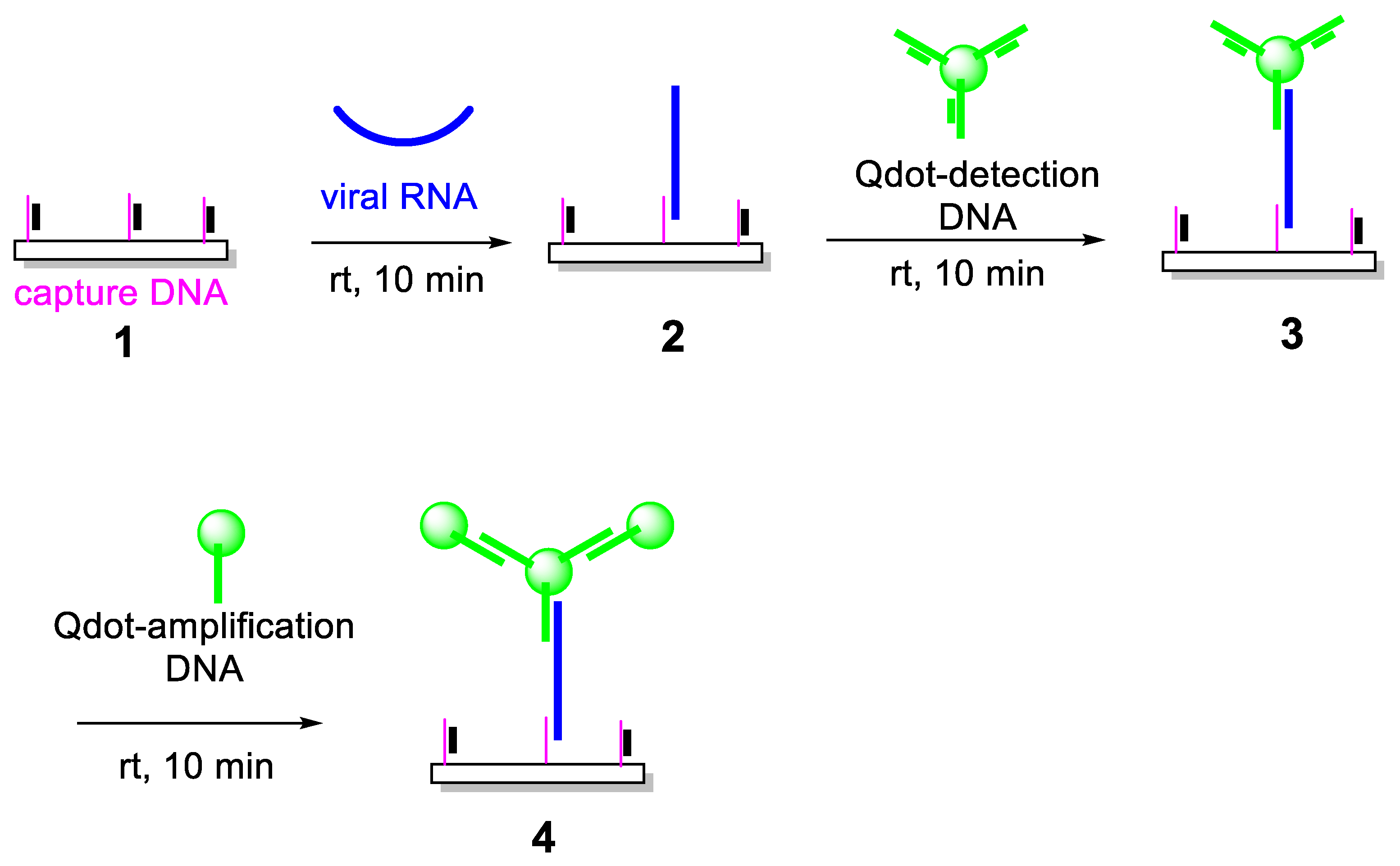
3.1. Design and Principle of an Enzyme-Free, Quantum Dot–Based RNA Detection Platform
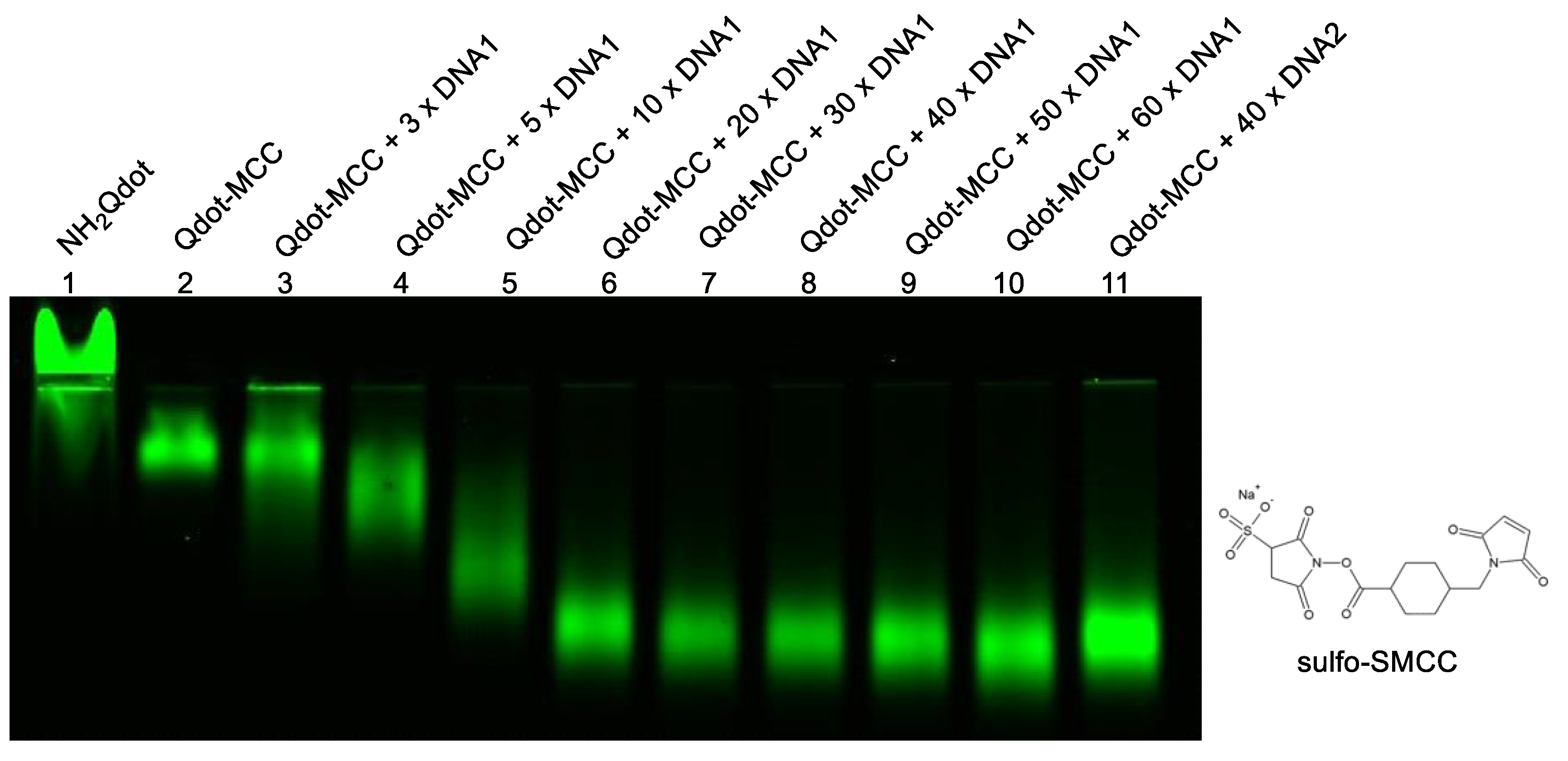
3.2. Preparation and Characterization of DNA–Quantum Dot Conjugates
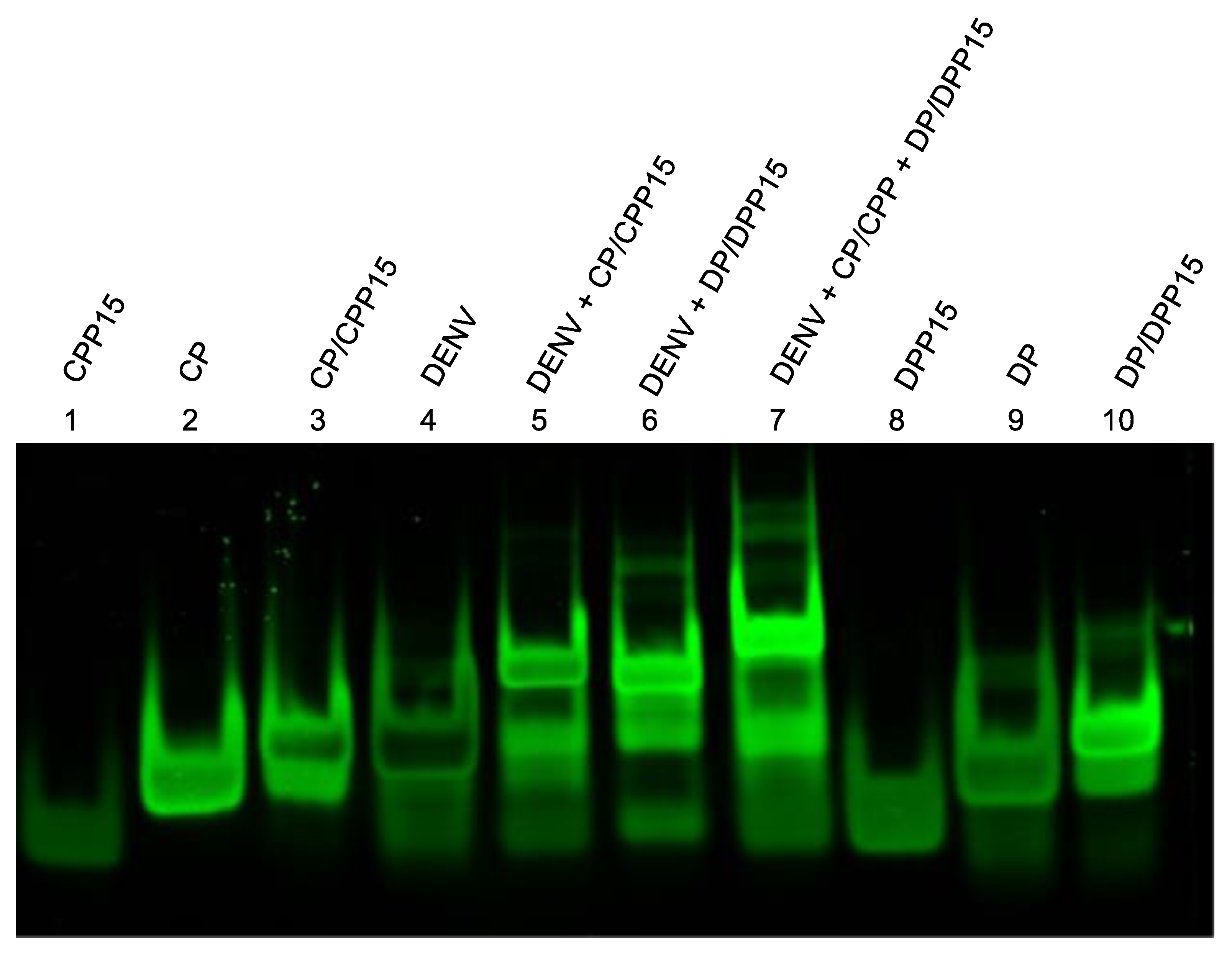
3.3. Validation of Toehold-Mediated Strand Displacement Reactions
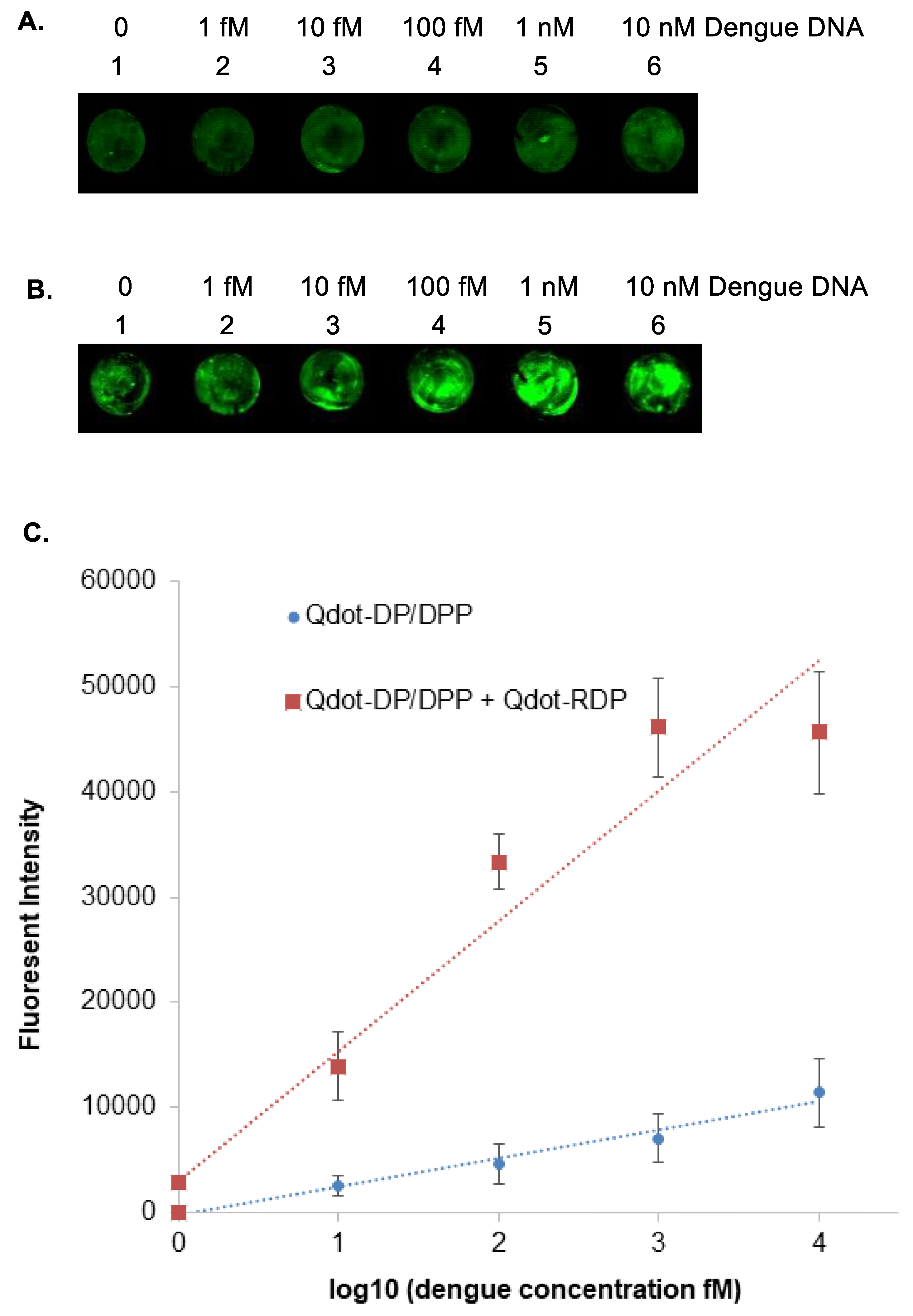
3.4. Sensitive Detection of DNA Targets
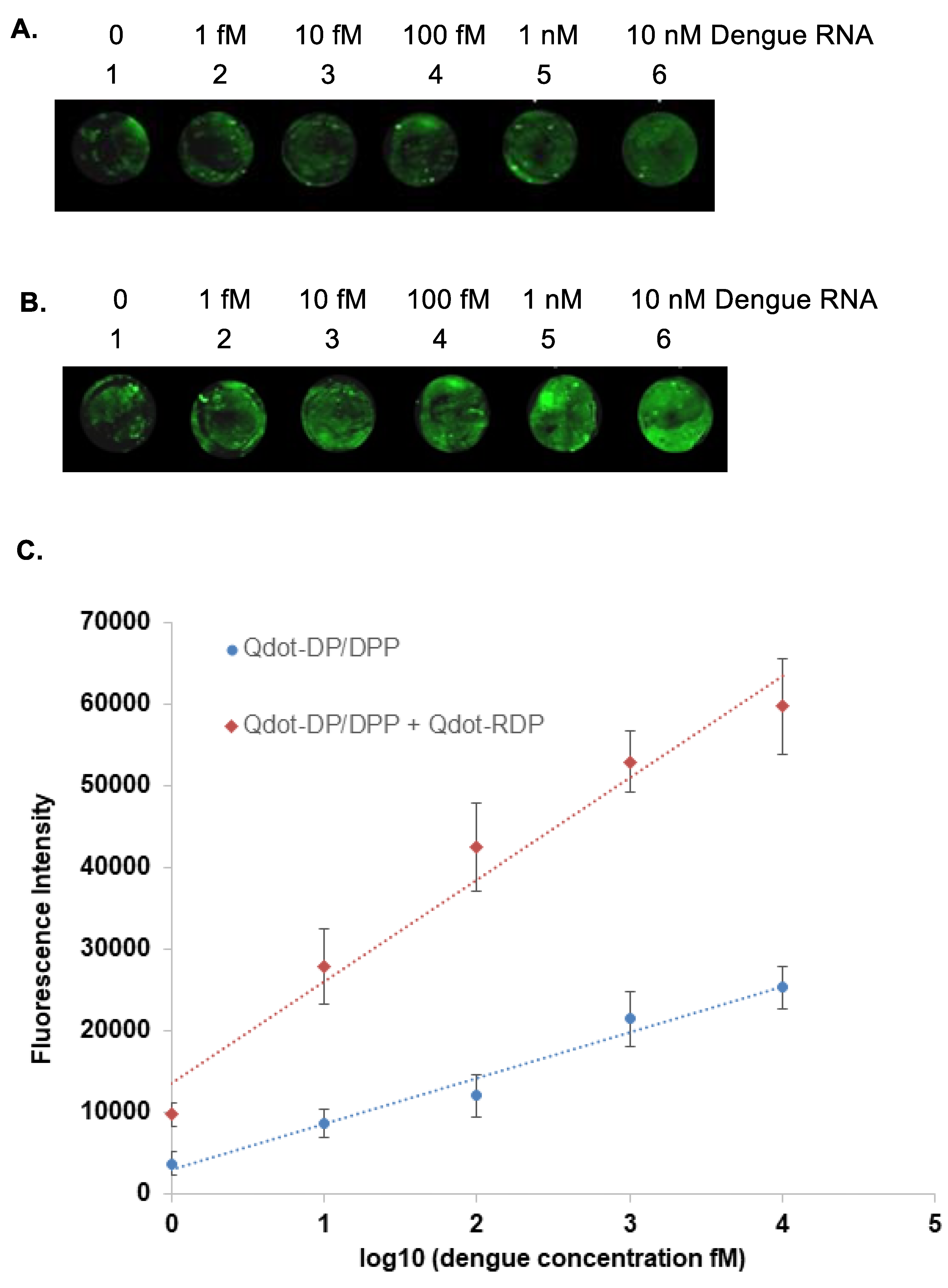
3.5. Preparation and Detection of Dengue RNA Targets
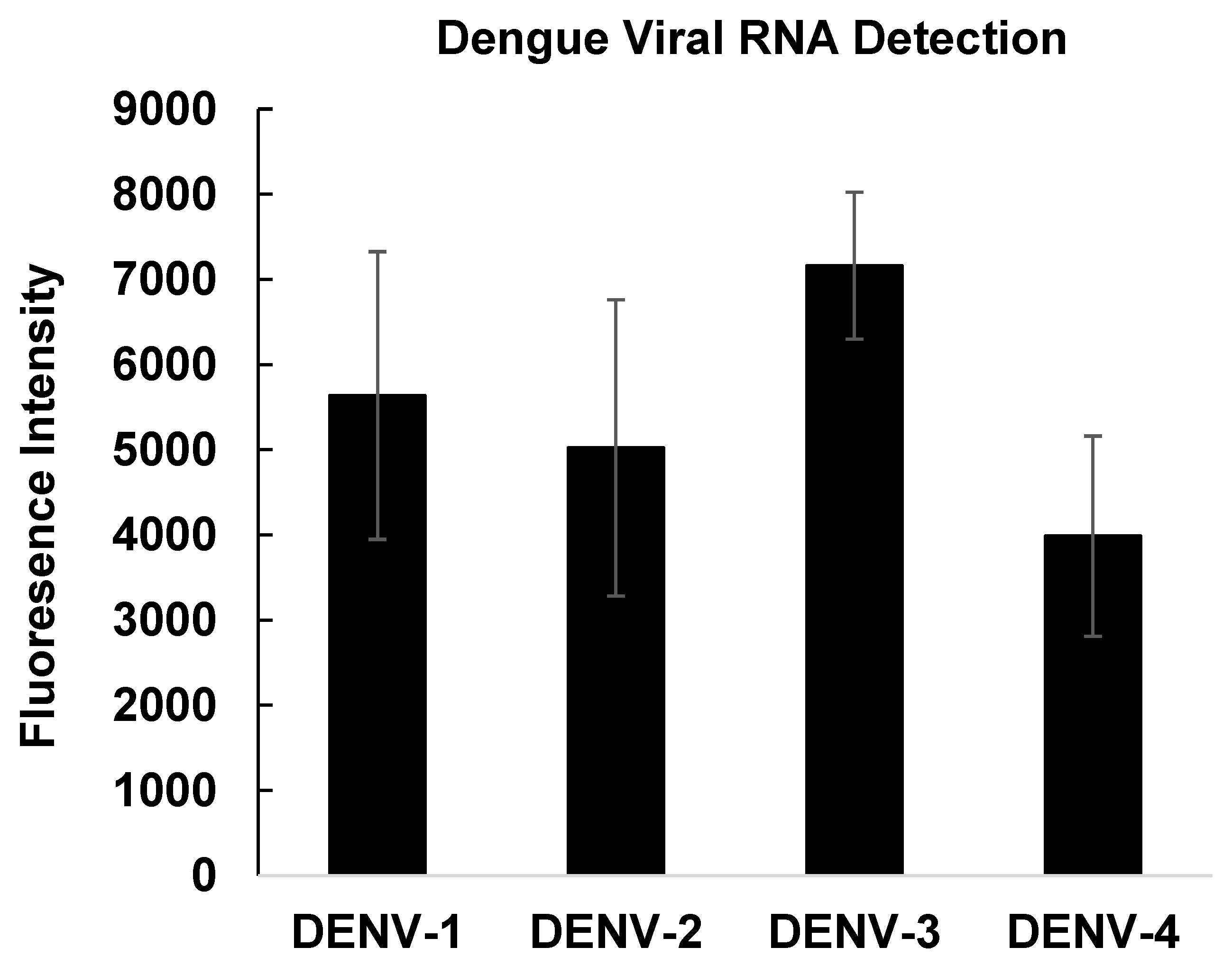
3.6. Detection of Dengue RNA in Contrived Clinical Samples
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data availability Statement
Conflicts of Interest
Abbreviations
| DENV | Dengue virus |
| EMCS | Maleimidocaproyloxysuccinimide ester |
| Qdot | Quantum dot |
| RT-PCR | Reverse transcription–polymerase chain reaction |
| TMDR | Toehold-mediated strand displacement reactions |
References
- World Health Organization. Dengue: Guidelines for Diagnosis, Treatment, Prevention and Control; World Health Organization: Geneva, Switzerland, 2009. [Google Scholar]
- Halstead, S.B. Pathogenesis of dengue: Challenges to molecular biology. Science 1988, 239, 476–481. [Google Scholar] [CrossRef]
- Shu, P.Y.; Huang, J.H. Current advances in dengue diagnosis. Clin. Diagn. Lab. Immunol. 2004, 11, 642–650. [Google Scholar] [CrossRef]
- Lanciotti, R.S.; Calisher, C.H.; Gubler, D.J.; Chang, G.J.; Vorndam, A.V. Rapid detection and typing of dengue viruses from clinical samples using reverse transcriptase–polymerase chain reaction. J. Clin. Microbiol. 1992, 30, 545–551. [Google Scholar] [CrossRef]
- Callahan, J.D.; Wu, S.J.; Dion-Schultz, A.; et al. Development and evaluation of serotype- and group-specific fluorogenic reverse transcriptase PCR (TaqMan) assays for dengue virus. J. Clin. Microbiol. 2001, 39, 4119–4124. [Google Scholar] [CrossRef]
- Wu, S.J.; Lee, E.M.; Putvatana, R.; et al. Detection of dengue viral RNA using a nucleic acid sequence-based amplification assay. J. Clin. Microbiol. 2001, 39, 2794–2798. [Google Scholar] [CrossRef]
- Carter, M.J.; Emary, K.R.; Moore, C.E.; et al. Rapid diagnostic tests for dengue virus infection in febrile Cambodian children: Diagnostic accuracy and incorporation into diagnostic algorithms. PLoS Negl. Trop. Dis. 2015, 9, e0003424. [Google Scholar] [CrossRef] [PubMed]
- Blacksell, S.D. Commercial dengue rapid diagnostic tests for point-of-care application: Recent evaluations and future needs. J. Biomed. Biotechnol. 2012, 2012, 151967. [Google Scholar] [CrossRef] [PubMed]
- Yow, K.S.; Aik, J.; Tan, E.Y.; Ng, L.C.; Lai, Y.L. Rapid diagnostic tests for the detection of recent dengue infections: An evaluation of six kits on clinical specimens. PLoS ONE 2021, 16, e0249602. [Google Scholar] [CrossRef] [PubMed]
- Sufi Aiman Sabrina, R.; Muhammad Azami, N.A.; Yap, W.B. Dengue and flavivirus co-infections: Challenges in diagnosis, treatment, and disease management. Int. J. Mol. Sci. 2025, 26, 6609. [Google Scholar] [CrossRef]
- Maeki, T.; Tajima, S.; Ando, N.; et al. Analysis of cross-reactivity among flaviviruses using sera of patients with dengue showed the importance of neutralization tests with paired serum samples for correct interpretation of serological test results for dengue. J. Infect. Chemother. 2023, 29, 469–474. [Google Scholar] [CrossRef]
- Gao, M.; Waggoner, J.J.; Hecht, S.M.; Chen, S. Selective detection of dengue virus serotypes using tandem toehold-mediated displacement reactions. ACS Infect. Dis. 2019, 5, 1907–1914. [Google Scholar] [CrossRef] [PubMed]
- Gao, M.; Daniel, D.; Zou, H.; Chen, S. Rapid detection of a dengue virus RNA sequence with single-molecule sensitivity using tandem toehold-mediated displacement reactions. Chem. Commun. 2018, 54, 968–971. [Google Scholar] [CrossRef]
- Simmel, F.C. Nucleic acid strand displacement—From DNA nanotechnology to translational regulation. RNA Biol. 2023, 20, 154–163. [Google Scholar] [CrossRef]
- Bai, S.; Xu, B.; Wu, J.; Xie, G. Series or parallel toehold-mediated strand displacement and its application in circular RNA detection and logic gates. Biosens. Bioelectron. 2023, 241, 115677. [Google Scholar] [CrossRef] [PubMed]
- Walbrun, A.; Wang, T.; Matthies, M.; Šulc, P.; Simmel, F.C.; Rief, M. Single-molecule force spectroscopy of toehold-mediated strand displacement. Nat. Commun. 2024, 15, 7564. [Google Scholar] [CrossRef] [PubMed]
- Kapadia, J.B.; Kharma, N.; Davis, A.N.; Kamel, N.; Perreault, J. Toehold-mediated strand displacement to measure released product from self-cleaving ribozymes. RNA 2022, 28, 263–273. [Google Scholar] [CrossRef]
- Hwang, M.T.; Landon, P.B.; Lee, J.; et al. Highly specific SNP detection using 2D graphene electronics and DNA strand displacement. Proc. Natl. Acad. Sci. USA 2016, 113, 7088–7093. [Google Scholar] [CrossRef]
- Han, Z.; Guo, A.X.; Luo, T.; Cai, T.; Mirkin, C.A. Biomineralization of semiconductor quantum dots using DNA-functionalized protein nanoreactors. Sci. Adv. 2025, 11, eadv6906. [Google Scholar] [CrossRef]
- Banerjee, A.; Pons, T.; Lequeux, N.; Dubertret, B. Quantum dots–DNA bioconjugates: Synthesis to applications. Interface Focus 2016, 6, 20160064. [Google Scholar] [CrossRef]
- Boeneman, K.; Deschamps, J.R.; Buckhout-White, S.; et al. Quantum dot–DNA bioconjugates: Attachment chemistry strongly influences the resulting composite architecture. ACS Nano 2010, 4, 7253–7266. [Google Scholar] [CrossRef]
- Hernandez, S.; Cardozo, F.; Myers, D.R.; Rojas, A.; Waggoner, J.J. Simple and economical extraction of viral RNA and storage at ambient temperature. Microbiol. Spectr. 2022, 10, e0085922. [Google Scholar] [CrossRef] [PubMed]
- Waggoner, J.J.; Abeynayake, J.; Sahoo, M.K.; et al. Single-reaction, multiplex, real-time RT-PCR for the detection, quantitation, and serotyping of dengue viruses. PLoS Negl. Trop. Dis. 2013, 7, e2116. [Google Scholar] [CrossRef] [PubMed]
- Sikes, H.D.; Hansen, R.R.; Johnson, L.M.; Jenison, R.; Birks, J.W.; Rowlen, K.L.; Bowman, C.N. Using polymeric materials to generate an amplified response to molecular recognition events. Nat. Mater. 2008, 7, 52–56. [Google Scholar] [CrossRef] [PubMed]
- Qian, H.; He, L. Polymeric macroinitiators for signal amplification in AGET ATRP-based DNA detection. Sens. Actuators B Chem. 2010, 150, 594–600. [Google Scholar] [CrossRef]
- Tricou, V.; Minh, N.N.; Farrar, J.; Tran, H.T.; Simmons, C.P. Kinetics of viremia and NS1 antigenemia are shaped by immune status and virus serotype in adults with dengue. PLoS Negl. Trop. Dis. 2011, 5, e1309. [Google Scholar] [CrossRef]
| DNA names | DNA sequences |
|---|---|
| Capture probe (CP-SH): | 5’- T CCC AGC GTC AAT ATG CTG TTT TTT TTT-SH-3’ |
| Protector for Capture probe (CPP): | 5’-AAA CAG CAT ATT GAC-3’ |
| Detection probe (DP-SH): | 5’-SH-TTT TTT GAG ACA GCA GGA TCT CTG GTC T-3’ |
| Protector for Detection probe (DPP): | 5’-AGA TCC TGC TGT CTC-3’ |
| Amplification probe (AP-SH): | 5’-SH-TTT TTT GAC CAG AGA TCC TGC TGT CTC-3’ |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




