Submitted:
28 December 2025
Posted:
29 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
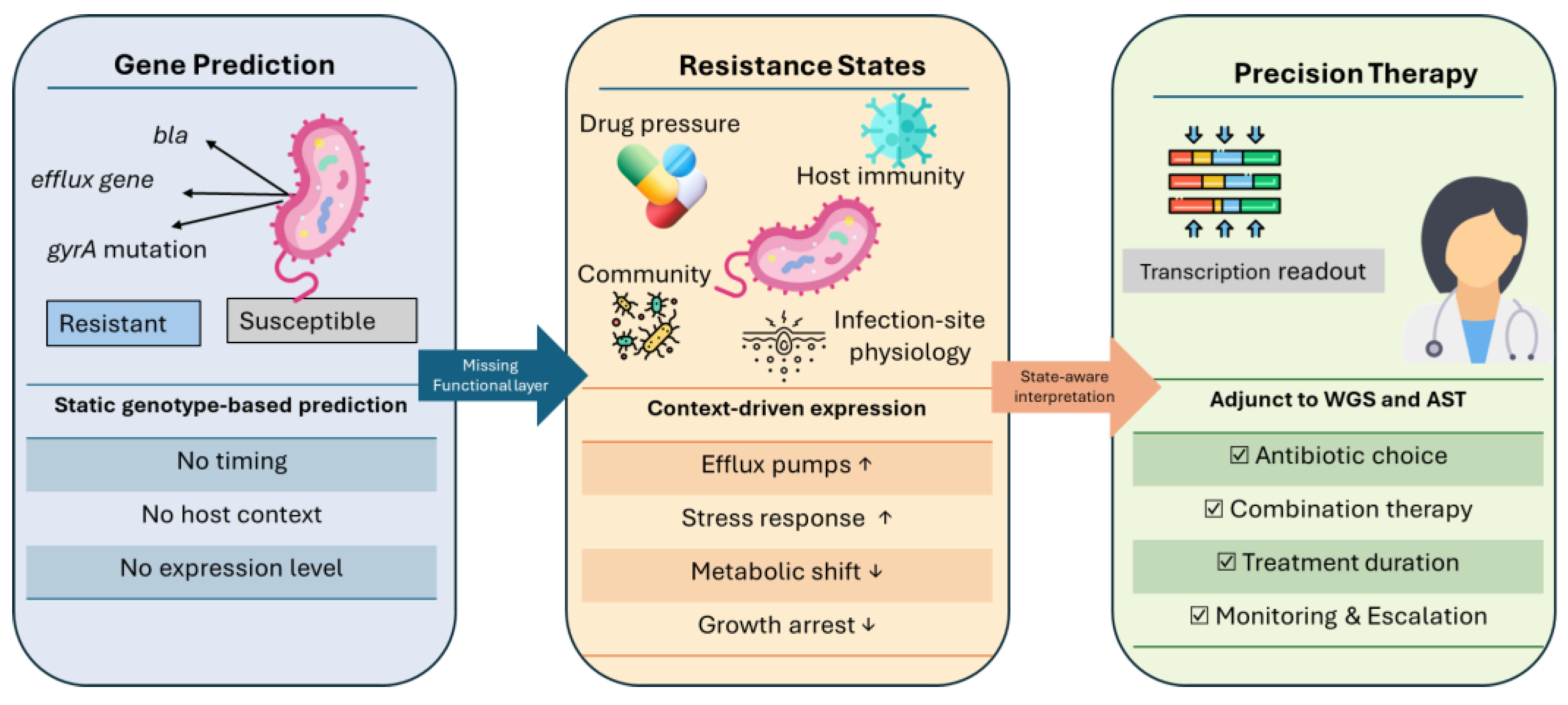
2. Resistance Genes vs Resistance States: A Conceptual Shift in Precision AMR
2.1. Defining Resistance States
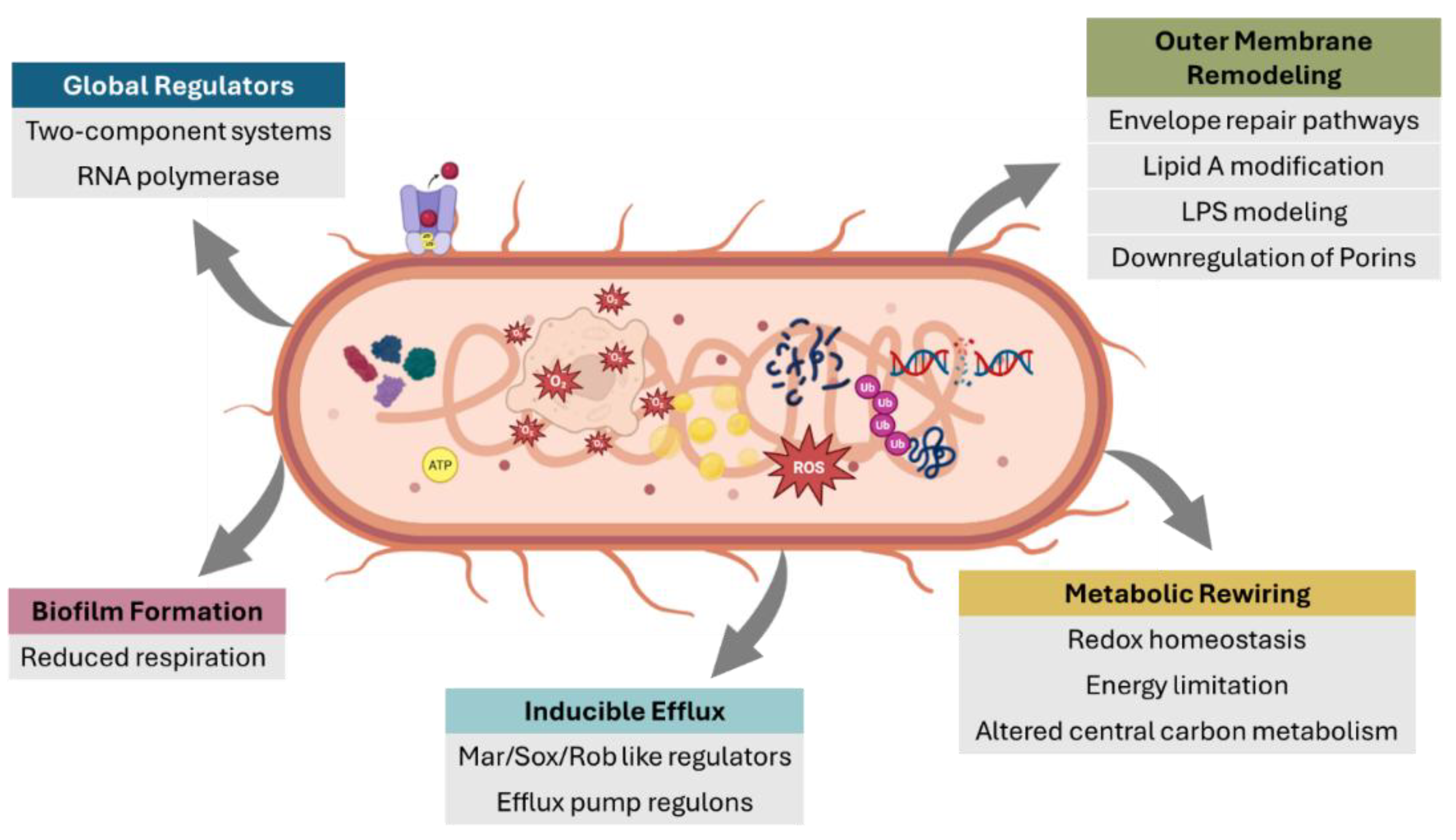
2.2. Determinants of Resistance States
2.3. Implications for Precision Medicine
3. What Pathogen Transcriptomics Adds Beyond Genomics
3.1. Mechanistic Insights Uniquely Revealed by Transcriptomics
3.2. Transcriptomic Signatures Under Antibiotic Stress: Empirical Evidence
3.3. Distinguishing In Vitro and Host-Relevant Transcriptional States
3.4. Classifying Resistance States: Toward Minimal Predictive Signatures
3.5. Limitations and Technical Considerations
4. Host and Environmental Modulation of Resistance States
4.1. Host Immune Pressures and Transcriptional Adaptation
4.2. Infection Site Physiology and Spatial Heterogeneity
4.3. Microbiome-Mediated Modulation of Resistance States
4.4. Evidence from Host-Relevant and Dual RNA-Seq Studies
4.5. Implications for Precision Antimicrobial Therapy
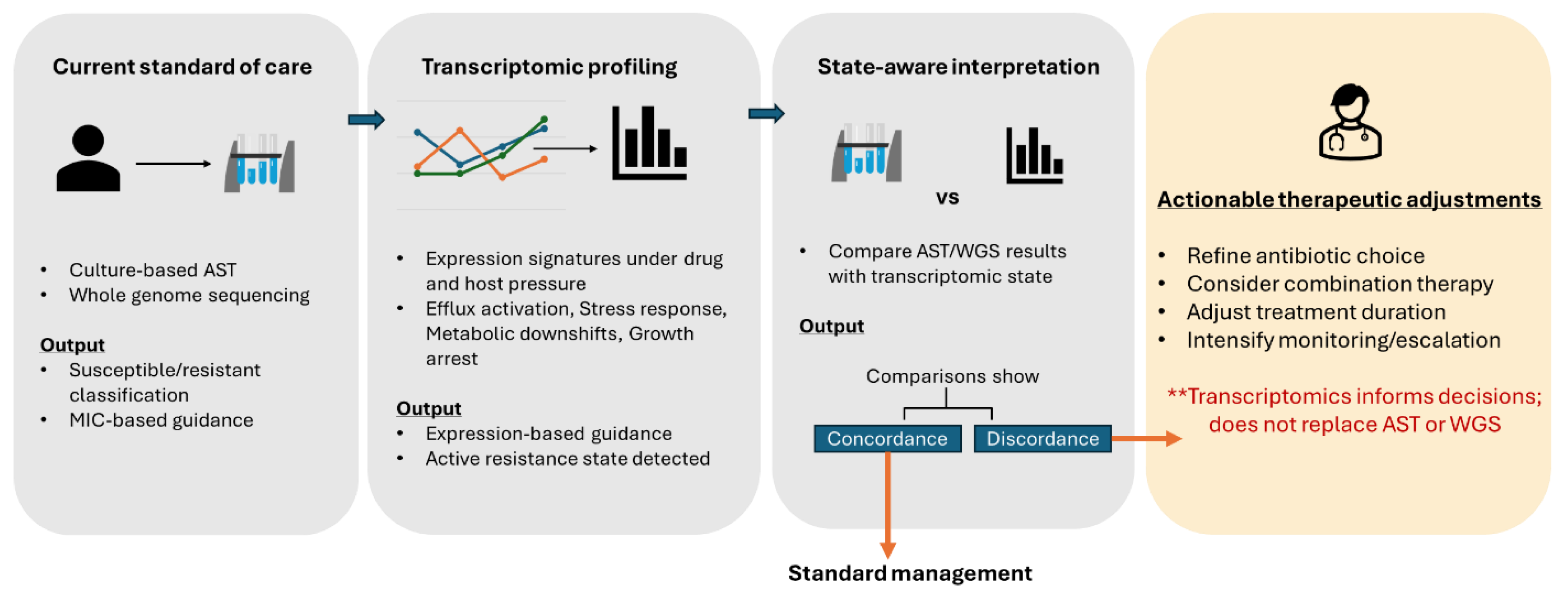
5. Transcriptomics-Informed Precision AMR Diagnostics
5.1. Moving Beyond Binary Resistance Classification
5.2. Expression-Based Classifiers and Resistance State Signatures
5.3. Validation Requirements for Clinical Translation
5.4. Practical Considerations for Implementation
5.5. Ethical and Regulatory Considerations
6. Implications for Personalized Antimicrobial Therapy
6.1. Antibiotic Selection and Therapeutic Stratification
6.2. Optimizing Treatment Duration and Monitoring Response
6.3. Preventing Resistance Emergence Through State-Aware Therapy
6.4. Clinical Feasibility and Translational Boundaries
7. Knowledge Gaps and Future Directions
7.1. Defining and Standardizing Resistance States
7.2. Improving Physiological Relevance of Transcriptomic Models
7.3. Validation Across Clinical Diversity
7.4. Balancing Model Complexity and Interpretability
7.5. Ethical, Logistical, and Implementation Challenges
8. Conclusions
Conflicts of Interest
References
- Andersson, D.I.; Hughes, D. Microbiological effects of sublethal levels of antibiotics. Nat. Rev. Microbiol. 2014, 12, 465–478. [Google Scholar] [CrossRef]
- Avican, K.; Aldahdooh, J.; Togninalli, M.; Mahmud, A.K.M.F.; Tang, J.; Borgwardt, K.M.; Rhen, M.; Fällman, M. RNA atlas of human bacterial pathogens uncovers stress dynamics linked to infection. Nat. Commun. 2021, 12, 3282. [Google Scholar] [CrossRef] [PubMed]
- Balaban, N.Q.; Helaine, S.; Lewis, K.; Ackermann, M.; Aldridge, B.; Andersson, D.I.; Brynildsen, M.P.; Bumann, D.; Camilli, A.; Collins, J.J.; et al. Definitions and guidelines for research on antibiotic persistence. Nat. Rev. Microbiol. 2019, 17, 441–448. [Google Scholar] [CrossRef] [PubMed]
- Band, V.I.; Weiss, D.S. Heteroresistance: A cause of unexplained antibiotic treatment failure? PLOS Pathog. 2019, 15, e1007726. [Google Scholar] [CrossRef]
- Van Belkum, A.; Burnham, C.-A.D.; Rossen, J.W.A.; Mallard, F.; Rochas, O.; Dunne, W.M., Jr. Innovative and rapid antimicrobial susceptibility testing systems. Nat. Rev. Microbiol. 2020, 18, 299–311. [Google Scholar] [CrossRef]
- Bie, L.; Zhang, M.; Wang, J.; Fang, M.; Li, L.; Xu, H.; Wang, M. Comparative Analysis of Transcriptomic Response of Escherichia coli K-12 MG1655 to Nine Representative Classes of Antibiotics. Microbiol. Spectr. 2023, 11, e0031723. [Google Scholar] [CrossRef]
- Brauner, A.; Fridman, O.; Gefen, O.; Balaban, N.Q. Distinguishing between resistance, tolerance and persistence to antibiotic treatment. Nat. Rev. Microbiol. 2016, 14, 320–330. [Google Scholar] [CrossRef]
- Buffie, C.G.; Pamer, E.G. Microbiota-mediated colonization resistance against intestinal pathogens. Nat. Rev. Immunol. 2013, 13, 790–801. [Google Scholar] [CrossRef]
- Byron, S.A.; Van Keuren-Jensen, K.R.; Engelthaler, D.M.; Carpten, J.D.; Craig, D.W. Translating RNA sequencing into clinical diagnostics: opportunities and challenges. Nat. Rev. Genet. 2016, 17, 257–271. [Google Scholar] [CrossRef]
- Chappell, J.; Takahashi, M.K.; Lucks, J.B. Creating small transcription activating RNAs. Nat. Chem. Biol. 2015, 11, 214–220. [Google Scholar] [CrossRef] [PubMed]
- Ching, T.; Himmelstein, D.S.; Beaulieu-Jones, B.K.; Kalinin, A.A.; Do, B.T.; Way, G.P.; Ferrero, E.; Agapow, P.-M.; Zietz, M.; Hoffman, M.M.; et al. Opportunities and obstacles for deep learning in biology and medicine. J. R. Soc. Interface 2018, 15. [Google Scholar] [CrossRef]
- Collins, F.S.; Varmus, H. A New Initiative on Precision Medicine. New Engl. J. Med. 2015, 372, 793–795. [Google Scholar] [CrossRef]
- Croucher, N.J.; Thomson, N.R. Studying bacterial transcriptomes using RNA-seq. Curr. Opin. Microbiol. 2010, 13, 619–624. [Google Scholar] [CrossRef]
- Dawan, J.; Ahn, J. Bacterial Stress Responses as Potential Targets in Overcoming Antibiotic Resistance. Microorganisms 2022, 10, 1385. [Google Scholar] [CrossRef] [PubMed]
- Didelot, X.; Maiden, M.C. Impact of recombination on bacterial evolution. Trends Microbiol. 2010, 18, 315–322. [Google Scholar] [CrossRef]
- El Meouche, I.; Jain, P.; Jolly, M.K.; Capp, J.-P. Drug tolerance and persistence in bacteria, fungi and cancer cells: Role of non-genetic heterogeneity. Transl. Oncol. 2024, 49, 102069. [Google Scholar] [CrossRef]
- Ellington, M.; Ekelund, O.; Aarestrup, F.; Canton, R.; Doumith, M.; Giske, C.; Grundman, H.; Hasman, H.; Holden, M.; Hopkins, K.; et al. The role of whole genome sequencing in antimicrobial susceptibility testing of bacteria: report from the EUCAST Subcommittee. Clin. Microbiol. Infect. 2017, 23, 2–22. [Google Scholar] [CrossRef]
- Fang, F.C.; Frawley, E.R.; Tapscott, T.; Vázquez-Torres, A. Bacterial Stress Responses during Host Infection. Cell Host Microbe 2016, 20, 133–143. [Google Scholar] [CrossRef] [PubMed]
- Hassall, J.; Coxon, C.; Patel, V.C.; Goldenberg, S.D.; Sergaki, C. Limitations of current techniques in clinical antimicrobial resistance diagnosis: examples and future prospects. npj Antimicrob. Resist. 2024, 2, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Hu, W.; Zhai, Z.Y.; Huang, Z.Y.; Chen, Z.M.; Zhou, P.; Li, X.X.; Yang, G.H.; Bao, C.J.; You, L.J.; Cui, X.B.; et al. Dual RNA sequencing of Helicobacter pylori and host cell transcriptomes reveals ontologically distinct host-pathogen interaction. mSystems 2024, 9, e0020624. [Google Scholar] [CrossRef]
- Joubert, I.A.; Mullally, C.; Litton, E.; Raby, E.; Chopra, A.; Strunk, T.; Wang, P.; Currie, A. Dual RNA isolation from blood: an optimized protocol for host and bacterial RNA purification for dual RNA-sequencing analysis in whole blood sepsis samples. Microb. Genom. 2025, 11, 001501. [Google Scholar] [CrossRef]
- Kirchner, M.; Nunez-Garcia, J.; Duggett, N.; Gosling, R.J.; Anjum, M.F. Use of transcriptomics and genomics to assess the effect of disinfectant exposure on the survival and resistance of Escherichia coli O157:H7, a human pathogen. Front. Microbiol. 2024, 15, 1477683. [Google Scholar] [CrossRef] [PubMed]
- Levin-Reisman, I.; Brauner, A.; Ronin, I.; Balaban, N.Q. Epistasis between antibiotic tolerance, persistence, and resistance mutations. Proc. Natl. Acad. Sci. 2019, 116, 14734–14739. [Google Scholar] [CrossRef]
- Lewis, K. Persister Cells. Annu. Rev. Microbiol. 2010, 64, 357–372. [Google Scholar] [CrossRef] [PubMed]
- Liao, J.; Tan, J.; Li, X.; Huang, D.; Wang, L.; Zhu, L.; Chen, Z.; Gan, L.; Liu, W.; Jin, X. Combined transcriptome and metabolome analysis revealed the antimicrobial mechanism of Griseorhodin C against Methicillin-resistant Staphylococcus aureus. Sci. Rep. 2024, 14, 30242. [Google Scholar] [CrossRef]
- Lobritz, M.A.; Belenky, P.; Porter, C.B.M.; Gutierrez, A.; Yang, J.H.; Schwarz, E.G.; Dwyer, D.J.; Khalil, A.S.; Collins, J.J. Antibiotic efficacy is linked to bacterial cellular respiration. Proc. Natl. Acad. Sci. 2015, 112, 8173–8180. [Google Scholar] [CrossRef] [PubMed]
- Rendón, J.M.; Lang, B.; Llorens, M.R.; Tartaglia, G.G.; Burgas, M.T. DualSeqDB: the host–pathogen dual RNA sequencing database for infection processes. Nucleic Acids Res. 49, D687–D693. [CrossRef]
- Martínez, J.L.; Rojo, F. Metabolic regulation of antibiotic resistance. FEMS Microbiol. Rev. 2011, 35, 768–789. [Google Scholar] [CrossRef]
- A McDevitt, C.; Ogunniyi, A.D.; Valkov, E.; Lawrence, M.C.; Kobe, B.; McEwan, A.G.; Paton, J.C. A Molecular Mechanism for Bacterial Susceptibility to Zinc. PLOS Pathog. 2011, 7, e1002357. [Google Scholar] [CrossRef]
- Ransom, E.M.; Potter, R.F.; Dantas, G.; Burnham, C.-A.D. Genomic Prediction of Antimicrobial Resistance: Ready or Not, Here It Comes! Clin. Chem. 2020, 66, 1278–1289. [Google Scholar] [CrossRef]
- Shahid, U. Microbiome-guided precision medicine: Mechanistic insights, multi-omics integration, and translational horizons. J. Precis. Med. Heal. Dis. 2025, 3. [Google Scholar] [CrossRef]
- Shahid, U. Advances in RNA therapeutics: Classes, innovations and clinical applications. J. Precis. Med. Heal. Dis. 2025, 3, 100016. [Google Scholar] [CrossRef]
- Shahid, U.; Hooi, S.L.; Lim, S.Y.; Aris, A.M.; Khor, B.C.; Ayub, Q.; Tan, H.S. Metagenomic surveillance of microbial community and antibiotic resistant genes associated with Malaysian wastewater during the COVID-19 pandemic. Curr. Genet. 2024, 71, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Shahreen, N.; Shahid, S.A.; Subhani, M.; Al-Siyabi, A.; Saha, R. Minimal gene signatures enable high-accuracy prediction of antibiotic resistance in Pseudomonas aeruginosa. npj Syst. Biol. Appl. 2025, 11, 108. [Google Scholar] [CrossRef] [PubMed]
- Stewart, P.S.; Franklin, M.J. Physiological heterogeneity in biofilms. Nat. Rev. Microbiol. 2008, 6, 199–210. [Google Scholar] [CrossRef] [PubMed]
- Su, M.; Satola, S.W.; Read, T.D. Genome-Based Prediction of Bacterial Antibiotic Resistance. J. Clin. Microbiol. 2019, 57, 1–15. [Google Scholar] [CrossRef]
- Toprak, E.; Veres, A.; Michel, J.-B.; Chait, R.; Hartl, D.L.; Kishony, R. Evolutionary paths to antibiotic resistance under dynamically sustained drug selection. Nat. Genet. 2011, 44, 101–105. [Google Scholar] [CrossRef]
- Westermann, A.J.; Gorski, S.A.; Vogel, J. Dual RNA-seq of pathogen and host. Nat. Rev. Microbiol. 2012, 10, 618–630. [Google Scholar] [CrossRef]
- Windels, E.M.; Michiels, J.E.; Fauvart, M.; Wenseleers, T.; Van den Bergh, B.; Michiels, J. Bacterial persistence promotes the evolution of antibiotic resistance by increasing survival and mutation rates. ISME J. 2019, 13, 1239–1251. [Google Scholar] [CrossRef]
| Dimension | Gene-Centric AMR Framework | Resistance-State Framework |
|---|---|---|
| Primary unit of analysis | Presence/absence of resistance genes or mutations | Transcriptionally regulated functional states |
| Temporal resolution | Static | Dynamic and time-dependent |
| Sensitivity to environment | Limited | High (host, antibiotic exposure, microbiome, site physiology) |
| Captures inducible resistance | Poorly | Explicitly |
| Explains tolerance and persistence | Largely no | Yes |
| Predictive power for treatment outcome | Moderate and population-averaged | Potentially patient-specific |
| Diagnostic modality | WGS, PCR-based assays | Transcriptomics (whole or targeted panels) |
| Clinical interpretation | Binary (resistant/susceptible) | State-aware and probabilistic |
| Precision medicine compatibility | Limited | High |
| Resistance state | Dominant transcriptomic features | Biological interpretation | Potential clinical relevance |
|---|---|---|---|
| Antibiotic tolerance | Upregulation of global stress-response regulons; reduced expression of growth-associated genes | Transient survival strategy without genetic resistance | Explains treatment failure despite in vitro susceptibility |
| Persistence | Suppression of replication, transcription, and translation programs | Entry into dormant or low-activity subpopulation | Predicts relapse and prolonged infection |
| Inducible resistance | Conditional expression of resistance determinants (e.g., efflux systems, modifying enzymes) | Context-dependent activation of resistance pathways | Missed by static genotypic diagnostics |
| Metabolic adaptation | Reprogramming of central carbon metabolism and redox balance | Reduced antibiotic lethality through altered physiology | Identifies metabolic vulnerabilities for adjuvant therapy |
| Biofilm-associated resistance | Expression of matrix synthesis genes; altered metabolic and stress-response profiles | Spatial and physiological protection from antibiotics | Relevant to chronic and device-associated infections |
| Host-induced resistance states | Activation of oxidative, nitrosative, or nutrient-limitation stress responses | Resistance behavior shaped by immune and tissue context | Explains discordance between in vitro and in vivo response |
| Diagnostic dimension | What transcriptomics can reveal | Added value beyond genomics | Key limitations / considerations |
|---|---|---|---|
| Resistance pathway activation | Active expression of efflux systems, stress-response regulons, metabolic adaptations | Distinguishes expressed vs silent resistance potential | Expression is context- and time-dependent |
| Early adaptive responses | Transcriptional changes preceding phenotypic resistance | Enables detection of pre-resistance or tolerance states | Requires precise timing of sampling |
| Host-influenced resistance states | Immune- and environment-induced expression programs | Captures in vivo-relevant resistance behavior | Host RNA dominance in clinical samples |
| Treatment monitoring | Dynamic shifts in resistance-associated expression during therapy | Potential marker of treatment response or persistence | Longitudinal sampling often impractical |
| Diagnostic stratification | Compact expression signatures associated with survival outcomes | Supports state-aware risk stratification | Limited cross-cohort validation |
| Clinical feasibility | Targeted expression panels (vs whole transcriptome) | Reduced cost and turnaround time | Requires standardization and regulatory approval |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




