Submitted:
27 December 2025
Posted:
29 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Biological Roles of Base Pair Formation
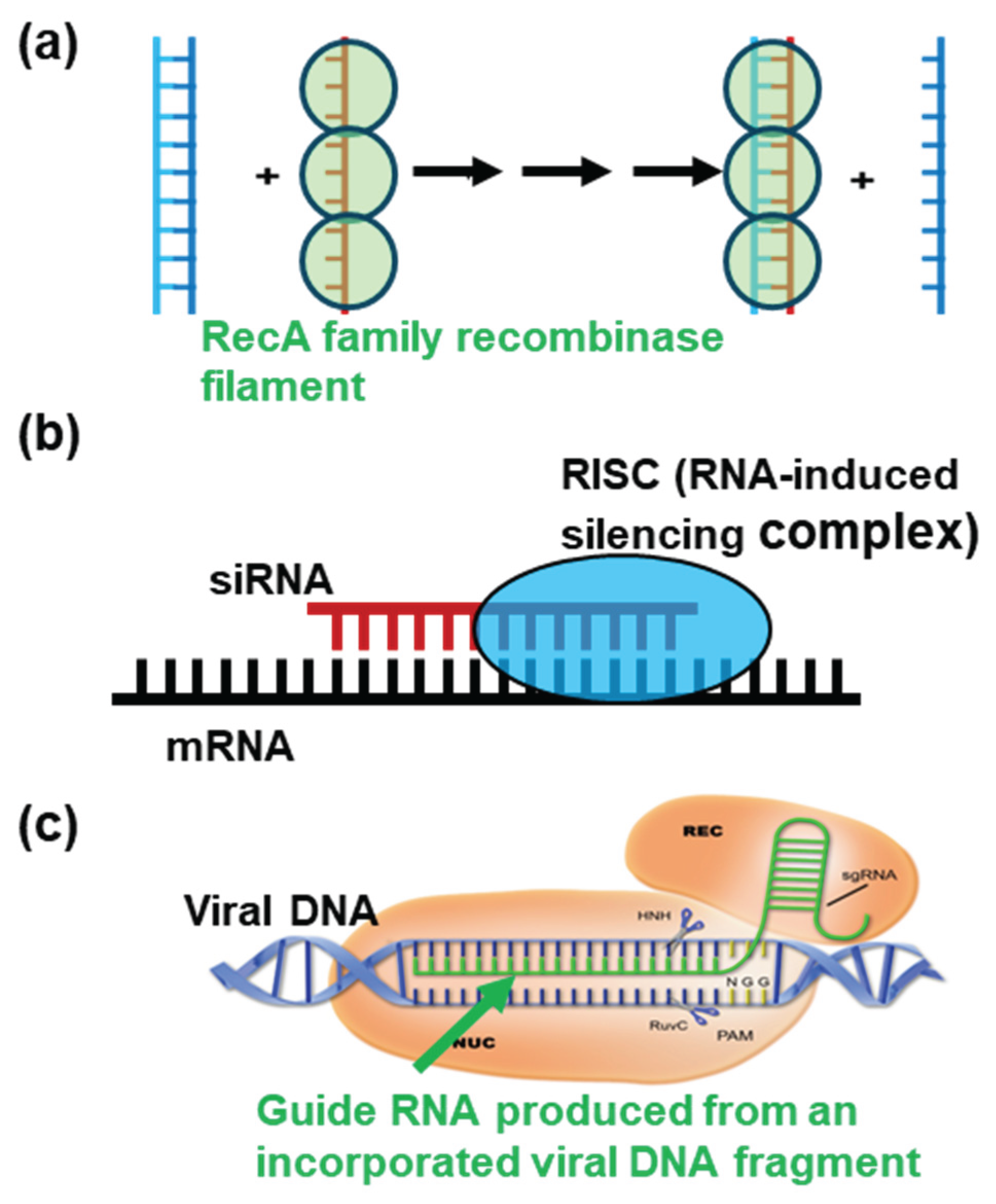
2.1. Homologous Recombination for DNA Repair and Meiosis
2.2. Regulation of Translation by siRNA and microRNA
2.3. Disruption of Virus DNA by the CRISPR–Cas9 System
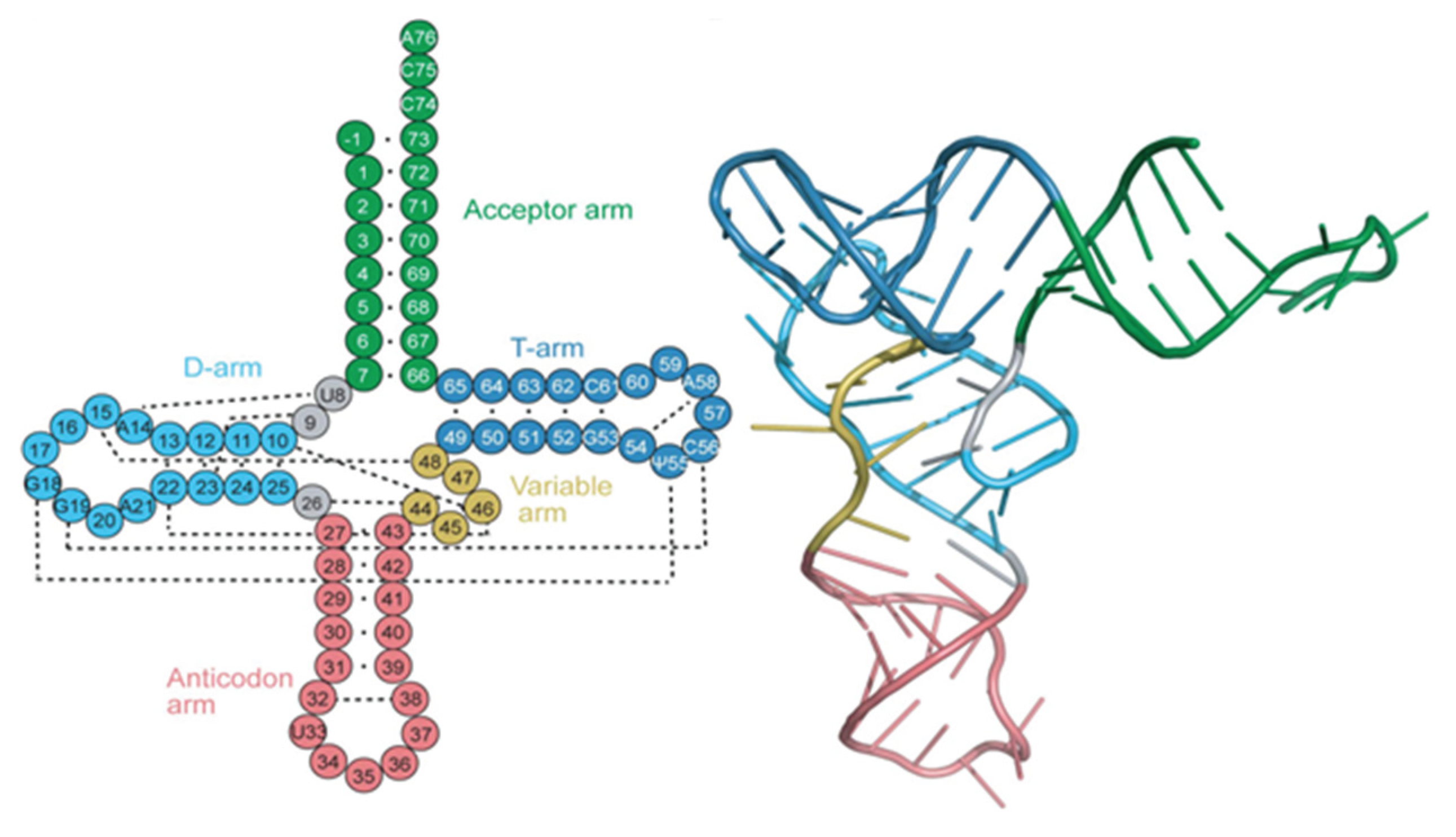
2.4. Structure of Non-Coding RNA
2.5. Size and Recognition Accuracy of Sequence Complementarity in the Reactions
3. Biotechnological Applications Using Base Pair Formation
3.1. Antisense- and siRNA-Based Gene Regulation and Drugs
3.2. Homologous Recombination- and CRISPR–Cas9-Assisted Gene Editing
3.3. DNA Hybridization
3.4. PCR for Amplification of a Specific DNA Segment (or Fragment)
3.5. DNA Nanodevices
3.6. Aptamers (Chemical Antibodies)
3.7. Problems with Applications
4. Molecular Mechanism of Complementary Base Recognition
4.1. Existence of Factors Other than the Hydrogen Bond
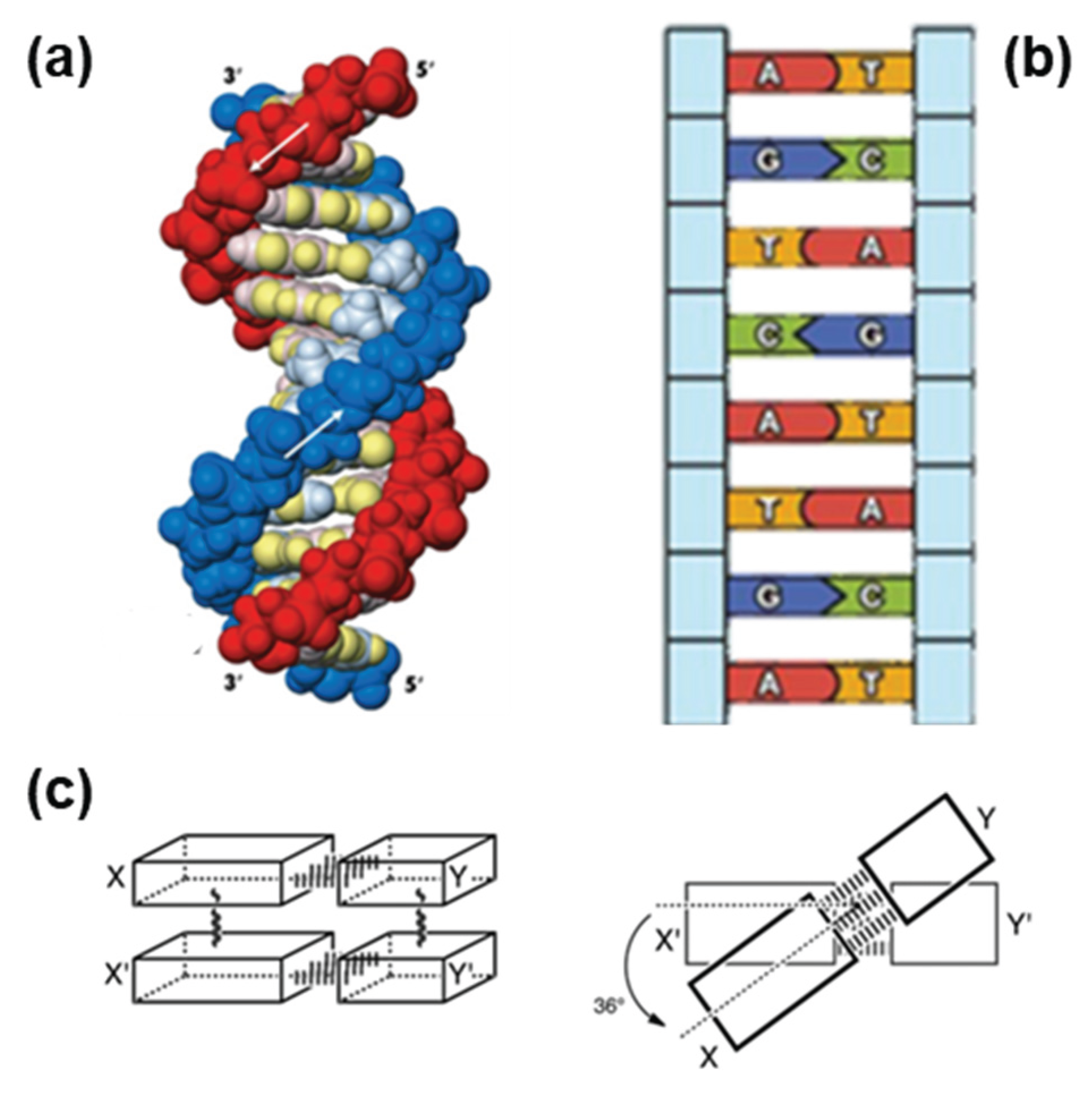
4.2. Geometry of Base Pairs
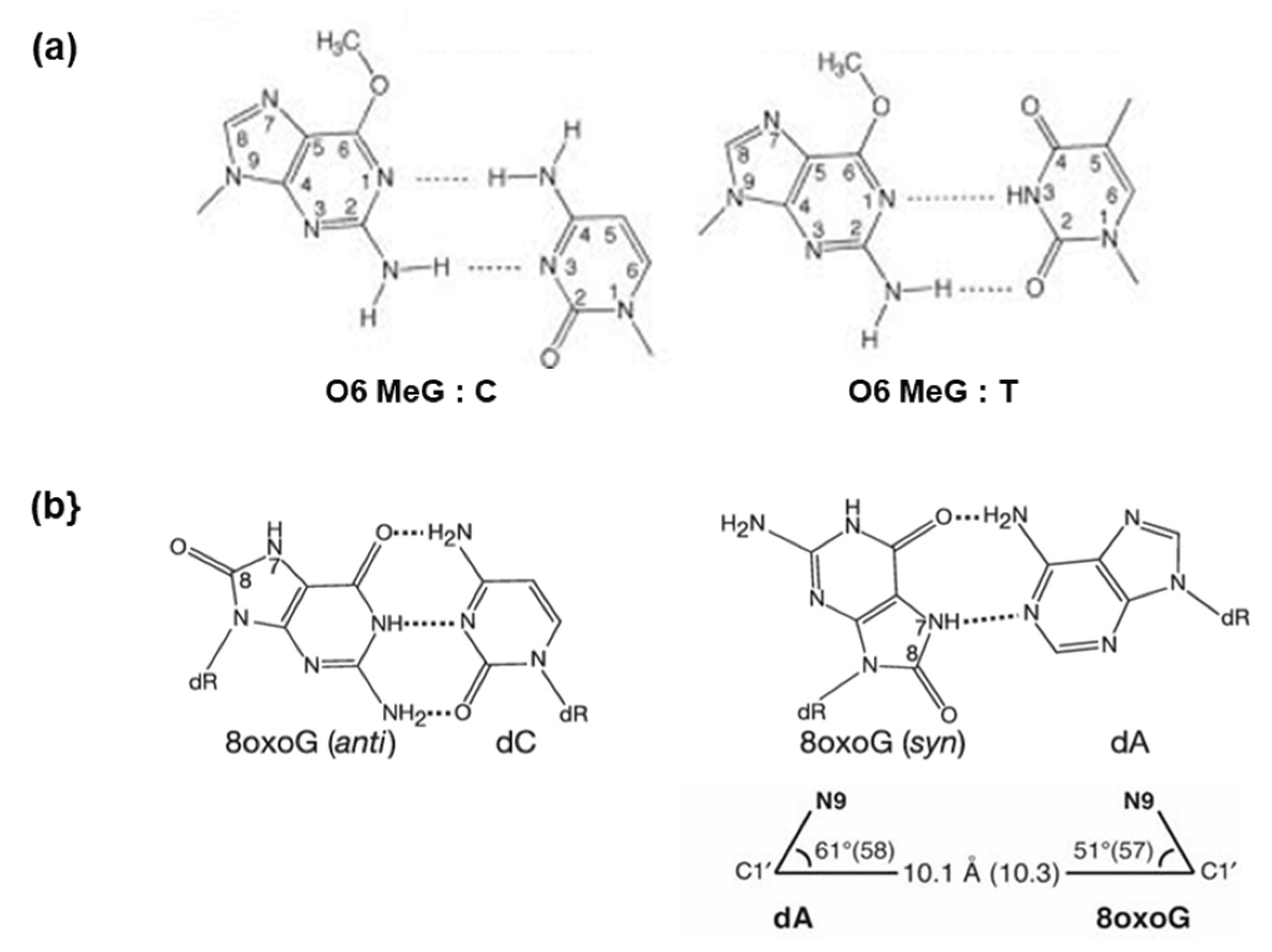
4.3. The Geometry of Base Pairs Explains the Mutagenic Effects of O6-methyl G and 8-oxo G
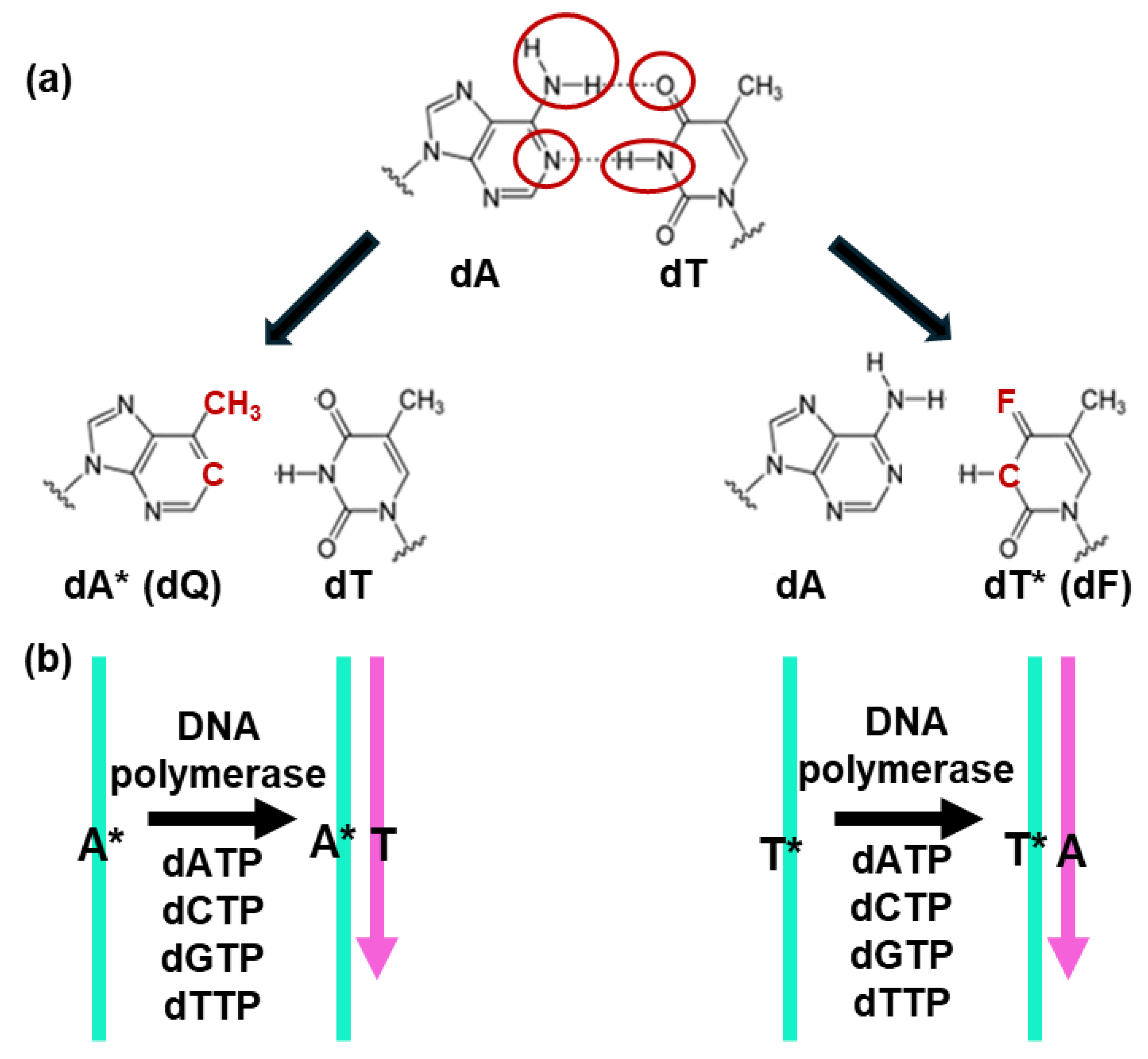
4.4. Artificial Base Pairs Without Hydrogen Bond Formation
4.5. Role of Base Stacking
5. Discussion
5.1. Kinetic Analyses
5.2. Existence of Multiple Mechanisms
5.3. Limitations of Perturbing Approaches
5.4. Examining the Strand Separation Step
6. Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- Watson, J.; Crick, F. Molecular Structure of Nucleic Acids: A Structure for Deoxyribose Nucleic Acid. Nature 1953, 171, 737–738. [CrossRef]
- Crick, F. On protein synthesis. Symposia of the Society for Experimental Biology 1958, 12, 138–163.
- Meselson, M.; Stahl, F. The replication of DNA in Escherichia coli. Proc Natl Acad Sci USA 1958, 44, 671–682.
- Cairns, J. The bacterial chromosome and its manner of replication as seen by autoradiography. J Mol Biol 1961, 6, 208–213.
- Lehman, I.R.; Bessman, M.J; Simms, E.S.; Kornberg, A. Enzymatic synthesis of deoxyribonucleic acid. I. Preparation of substrates and partial purification of an enzyme from Escherichia coli. J Biol Chem 1958, 233, 163–170.
- Weiss, S.B.; Gladstone, L.A. A mammalian system for the incorporation of cytidine triphosphate into ribonucleic acid. J. Am. Chem. Soc. 1958, 81, 4118-4119. [CrossRef]
- Chapeville, F.; Lipmann, F.; Ehrenstein, G.V.; Weisblum, B.; Ray Jr, W.J.; Benzer, S. On the role of soluble ribonucleic acid in coding for amino acids. Proc Natl Acad Sci USA 1962, 48, 1086–1092.
- Matthaei, H.J.; Jones, O.W.; Martin, R.G.; Nirenberg, M.W. Characteristics and Composition of RNA Coding Units. Proc Natl Acad Sci USA 1962, 48, 666–677. [CrossRef]
- Fire, A,; Xu, S. Montgomery, M.K.; Kostas, S.A.; Driver, S.E.; Mello, C.C. Potent and specific genetic interference by double-stranded RNA in caenorhabditis elegans. Nature 1998, 391, 806-811. [CrossRef]
- Dana, H.; Chalbatani, G.M,; Mahmoodzadeh, H.; Karimloo, R.; Rezaiean, O.; Moradzadeh, A.; Mehmandoost, N.; Moazzen, F.; Mazraeh, A.; Marmari, V.; Ebrahimi, M.; Rashno, M.M.; Abadi, S.J.; Gharagouzlo, E. Molecular Mechanisms and Biological Functions of siRNA. Int J Biomed Sci. 2017, 13, 48-57. [CrossRef]
- Wright, W.D.; Shah, S.S.; Heyer, W.D. Homologous recombination and the repair of DNA double-strand breaks. J Biol Chem. 2018, 6, 10524-10535. [CrossRef]
- Chakraborty, S.; Schirmeisen, K.; Lambert, S.A. The multifaceted functions of homologous recombination in dealing with replication-associated DNA damages. DNA Repair, 2023, 129, 103548. [CrossRef]
- Székvölgyi, L.; Nicolas, A. From meiosis to postmeiotic events: Homologous recombination is obligatory but flexible. FEBS J. 2010, 277, 571-589. [CrossRef]
- Mojica, F.J.; Díez-Villaseñor C, García-Martínez J, Soria E. Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J Mol Evol. 2005, 60, 174-82. [CrossRef]
- Kleppe, K.; Ohtsuka, E.; Kleppe, R.; Molineux, I.; Khorana, H.G. Studies on polynucleotides. XCVI. Repair replications of short synthetic DNA's as catalyzed by DNA polymerases. J Mol Biol. 1971, 56 (2): 341–61. [CrossRef]
- Kang, H.; Ga, Y.J.; Kim, S.H. et al. Small interfering RNA (siRNA)-based therapeutic applications against viruses: principles, potential, and challenges. J Biomed Sci 2023, 30, 88. [CrossRef]
- Chi, X.; Gatti, P.; Papoian, T. Safety of antisense oligonucleotide and siRNA-based therapeutics, Drug Discovery Today, 2017, 22, 823-833. [CrossRef]
- Kang, H., Ga, Y.J., Kim, S.H., et al. Small interfering RNA (siRNA)-based therapeutic applications against viruses: principles, potential, and challenges. J Biomed Sci 2023, 30, 88. [CrossRef]
- Cox, D.B.; Platt, R.J.; Zhang, F. Therapeutic genome editing: prospects and challenges. Nature Medicine. 2015, 21, 121–131. [CrossRef]
- Fan, Q., Yang, L., & Chao, J. Recent Advances in Dynamic DNA Nanodevice. Chemistry, 2023, 5, 1781-1803. [CrossRef]
- Moran, S.; Ren, R.X.; Kool, E.T. A thymidine triphosphate shape analog lacking Watson–Crick pairing ability is replicated with high sequence selectivity Proc. Natl. Acad. Sci. USA. 1997, 94, 10506-10511. [CrossRef]
- Morales, J.C.; Kool, E. T. Efficient replication between non-hydrogen-bonded nucleoside shape analogs. Nature Structural Biology, 1998, 5, 950-954. [CrossRef]
- Kool, E.T.; Sintim, H.O. The Difluorotoluene Debate — A Decade Later. Chemical Communications 2006, 38, 3665-75.
- Tikhomirova, A.; Beletskaya, I.V.; Chalikian, T.V. Stability of DNA Duplexes Containing GG, CC, AA, and TT Mismatches Biochemistry 2006, 45, 10563-10571. [CrossRef]
- Bartel, D.P. Metazoan MicroRNAs. Cell. 2018, 173, 20–51. [CrossRef]
- O'Brien, J.; Hayder, H.; Zayed, Y.; Peng, C. Overview of MicroRNA Biogenesis, Mechanisms of Actions, and Circulation. Front Endocrinol (Lausanne). 2018, 9, 402. [CrossRef]
- Komor, A.C.; Kim, Y.B.; Packer, M.S. Zuris, J.A.; Liu, D.R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature, 2016, 533, 420-424. [CrossRef]
- Li, T. ; Yang, Y. ; Qi, H. et al. CRISPR/Cas9 therapeutics: progress and prospects. Sig Transduct Target Ther 2023, 8, 36. [CrossRef]
- Biela, A.; Hammermeister, A.; Kaczmarczyk, I.; Walczak, M.; Koziej, L.; Lin, T.; Glatt, S. The diverse structural modes of tRNA binding and recognition. J Biol Chem, 2023, 299, 104966. [CrossRef]
- Leitão, A. L.; Enguita, F.J.; Leitão, A.L.; Enguita, F.J. (2025). The Unpaved Road of Non-Coding RNA Structure–Function Relationships: Current Knowledge, Available Methodologies, and Future Trends. Non-Coding RNA 2025, 11, 20. [CrossRef]
- Sloane, D.L.; Goodman, M.F.; Echols, H. The fidelity of base selection by the polymerase subunit of DNA polymerase III holoenzyme. Nucleic Acids Res. 1988, 16, 6465-75. [CrossRef]
- Anand, R.; Beach, A.; Li, K.; Haber, J. Rad51-mediated double-strand break repair and mismatch correction of divergent substrates. Nature 2017, 453, 489–494.
- Takahashi, M. RecA kinetically selects homologous DNA by testing a five- or six-nucleotide matching sequence and deforming the second DNA. Quarterly Rev. Biophys. 2018, 51, e11. [CrossRef]
- Kunkel, T.A.; Bebenek, K. DNA replication fidelity. Annu Rev Biochem. 2000; 69,497-529.
- Imashimizu, M.; Oshima, T.; Lubkowska, L.; Kashlev, M. Direct assessment of transcription fidelity by high-resolution RNA sequencing. Nucleic Acids Res. 2013; 41, 9090–9104. [CrossRef]
- Fijalkowska, I.J.; Schaaper, R.M.; Jonczyk, P. DNA replication fidelity in Escherichia coli: a multi-DNA polymerase affair. FEMS Microbiol Rev. 2012, 36,1105-21. [CrossRef]
- Han, H. RNA Interference to Knock Down Gene Expression. Methods Mol Biol. 2018; 1706:293-302. [CrossRef]
- Lauffer, M.C. Possibilities and limitations of antisense oligonucleotide therapies for the treatment of monogenic disorders. Communications Medicine, 2024, 4, 6. [CrossRef]
- Ratilainen, T.; Lincoln, P.; Nordén, B. A simple model for gene targeting. Biophys J. 2001, 81, 2876-85. [CrossRef]
- Neumeier, J.; Meister, G. SiRNA Specificity: RNAi Mechanisms and Strategies to Reduce Off-Target Effects. Frontiers in Plant Science, 2021, 11, 526455. [CrossRef]
- Hoshijima, K.; Jurynec, M.J.; Grunwald, D.J. Precise genome editing by homologous recombination. Methods Cell Biol. 2016, 135:121-47.
- Hoheisel, J.D. Application of hybridization techniques to genome mapping and sequencing. Trends in Genetics, 1994, 10, 79-83. [CrossRef]
- Lao, A.I.K.; Su, X.; Aung, K.M.M. SPR study of DNA hybridization with DNA and PNA probes under stringent conditions. Biosensors and Bioelectronics, 2009, 24,1717-1722. [CrossRef]
- Furst, A.L.; Klass, S.H.; Francis, M.B. DNA Hybridization to Control Cellular Interactions. Trends in Biochemical Sciences, 2019, 44, 342-350. [CrossRef]
- Bej, A.K.; Mahbubani, M.H.; Atlas, R.M. Amplification of Nucleic Acids by Polymerase Chain Reaction (PCR) and Other Methods and their Applications. Critl Rev Biochem Mol Biol, 1991, 26, 301–334. [CrossRef]
- Fan, Q.; Yang, L.; Chao, J. Recent Advances in Dynamic DNA Nanodevice. Chemistry, 2023, 5, 1781-1803. [CrossRef]
- Guan, C.; Zhu, X.; Feng, C. DNA Nanodevice-Based Drug Delivery Systems. Biomolecules. 2021, 11, 1855. [CrossRef]
- Kong, L.; Kim, S.; Wang, C.; Lee, S.Y.; Oh, S.; Lee, S.; Jo, S.; Kim, T. Understanding nucleic acid sensing and its therapeutic applications. Experimental & Molecular Medicine, 2023, 55, 2320-2331. [CrossRef]
- Sherman, W.B.; Seeman, N.C. A precisely controlled DNA biped walking device Nano Letters 2004, 4, 1203–1207.
- Parker, J. Computing with DNA. EMBO Rep. 2003, 4, 7-10.
- Shraim, A.S.; Abdel Majeed, B.A.; Al-Binni, M.A.; Hunaiti, A. Therapeutic Potential of Aptamer-Protein Interactions. ACS Pharmacol Transl Sci. 2022, 5, 1211-1227. [CrossRef]
- Nimjee, S.M.; White, R.R.; Becker, R.C.; Sullenger, B.A. Aptamers as Therapeutics. Annu Rev Pharmacol Toxicol. 2017, 57:61-79.
- Yang, W., Ran, C., Lian, X., Wang, Z., Du, Z., Bing, T., Zhang, Y., & Tan, W. Aptamer-based targeted drug delivery and disease therapy in preclinical and clinical applications. Advanced Drug Delivery Reviews, 2025, 226, 115680. [CrossRef]
- Ali Zaidi, S.S.; Fatima, F., Ali Zaidi, S.A.; Zhou, D.; Deng, W.; Liu, S. Engineering siRNA therapeutics: challenges and strategies. J Nanobiotechnol 2023, 21, 381. [CrossRef]
- Guo, C.; Ma, X.; Gao, F; Guo, Y. Off-target effects in CRISPR/Cas9 gene editing. Front Bioeng Biotechnol. 2023, 11, 1143157. [CrossRef]
- Cai, Y.; Yu, X.; Hu, S.; Yu, J. A Brief Review on the Mechanisms of miRNA Regulation. Genomics, Proteomics & Bioinformatics, 2009, 7, 147-154. [CrossRef]
- Moran, S.; Ren, R.X.; Kool, E.T. A thymidine triphosphate shape analog lacking Watson–Crick pairing ability is replicated with high sequence selectivity. Proc. Natl. Acad. Sci. USA. 1997, 94,10506-10511. [CrossRef]
- Delaney, J.C.; Henderson, P.T.; Helquist, S.A.; Morales, J.C.; Essigmann, J.M.; Kool, E.T. High-fidelity in vivo replication of DNA base shape mimics without Watson–Crick hydrogen bonds. Proc. Natl. Acad. Sci. USA 2003, 100, 4469-4473. [CrossRef]
- Chavarria, D.; Ramos-Serrano, A.; Hirao, I.; Berdis, A.J. Exploring the roles of nucleobase desolvation and shape complementarity during the misreplication of O6-methylguanine. J. Mol. Biol. 2011, 412, 325-339. [CrossRef]
- Hsu. G.W.; Ober, M.; Carell, T.; Beese, L.S. Error-prone replication of oxidatively damaged DNA by a high-fidelity DNA polymerase. Nature 2004, 431, 217-221. [CrossRef]
- Yamashige, R.; Kimoto, M.; Takezawa, Y.; Sato, A.; Mitsui, T.; Yokoyama, S.; Hirao, I. Highly specific unnatural base pair systems as a third base pair for PCR amplification. Nucleic Acids Res. 2012, 40, 2793–2806. [CrossRef]
- Guckian, K.M.; Krugh, T.R.; Kool, E.T., Solution structure of a nonpolar, non-hydrogen-bonded base pair surrogate in DNA. J. Am. Chem. Soc. 2000, 122, 6841–6847. [CrossRef]
- Feng, B.; Sosa, R.P.; Mårtensson, A.K.F.; Jiang, K.; Tong, A.; Dorfman, K.D.; Takahashi, M.; Lincoln. P.; Bustamante, C.J.; Westerlund, F.; Nordén, B. Hydrophobic catalysis and a potential biological role of DNA unstacking induced by environment effects. Proc. Natl. Acad. Sci. USA 2019, 116, 17169-17174. [CrossRef]
- Nordén, B.; Brown, T.; Feng, B. Mismatch detection in homologous strand exchange amplified by hydrophobic effects. Biopolymers, 2021, 112, e23426. [CrossRef]
- Takahashi, M.; Ito, K.; Iwasaki, H.; Nordén, B. Linear dichroism reveals the perpendicular orientation of DNA bases in the RecA and Rad51 recombinase filaments: a possible mechanism for the strand exchange reaction. Chirality 2024, 36, e23664. [CrossRef]
- Freudenthal, B.D.; Beard, W.A.; Wilson, S.H. Structures of dNTP intermediate states during DNA polymerase active site assembly. Structure, 2012, 20, 1829-1837. [CrossRef]
- Ito, K.; Murayama, Y.; Takahashi, M.; Iwasaki, H. Two three-strand intermediates are processed during Rad51-driven DNA strand exchange. Nature Struct. & Mol. Biol. 2018, 25, 29-36. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



