Submitted:
23 December 2025
Posted:
24 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
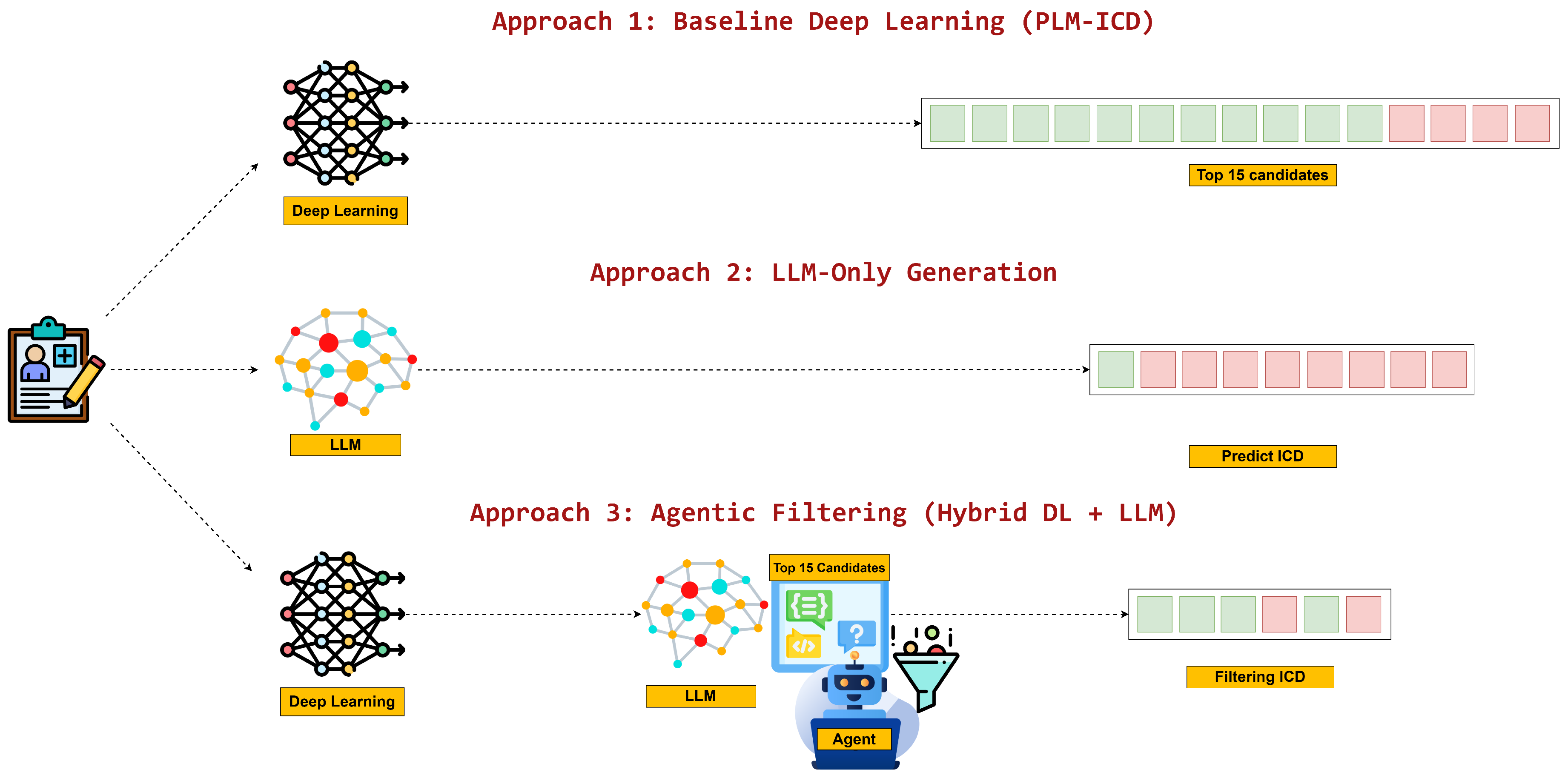
- Deep Learning Automation: Specialized PLM-ICD models generate 15-code lists directly from discharge summaries, mirroring conventional fixed-output automation.
- Direct LLM Automation: Standalone LLMs attempt to write complete ICD code sets from raw text, selecting both the codes and the number of predictions without outside assistance.
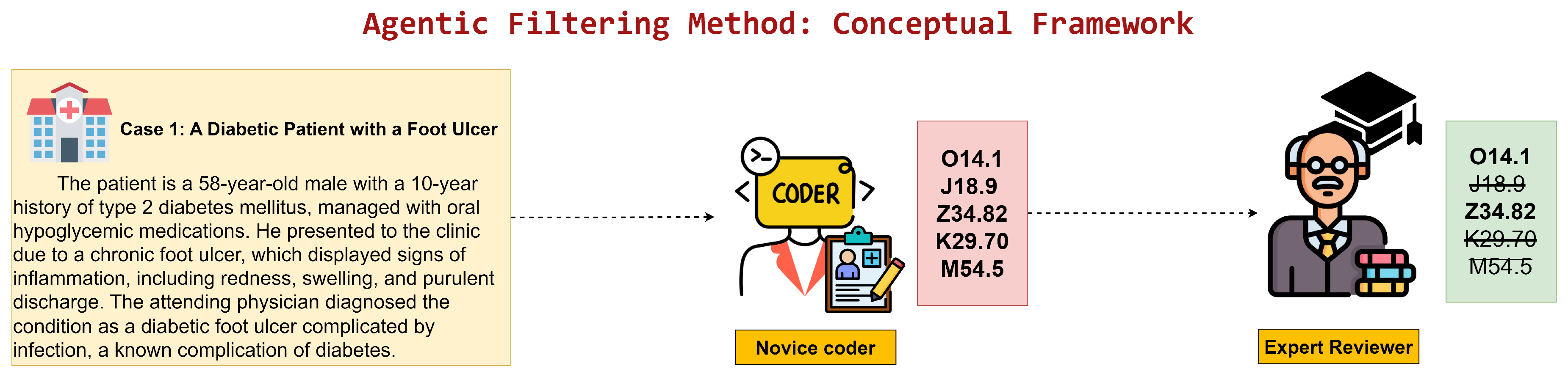
- Agentic Filtering Automation: A filtering-centric workflow in which candidate codes are proposed by a deep learning model and then validated or discarded by an LLM agent, shifting LLM effort toward quality assurance rather than generation.
- Head-to-head evaluation of three automation paradigms: The comparative behavior of conventional deep learning, direct LLM generation, and agentic filtering is quantified on a shared dataset with consistent metrics.
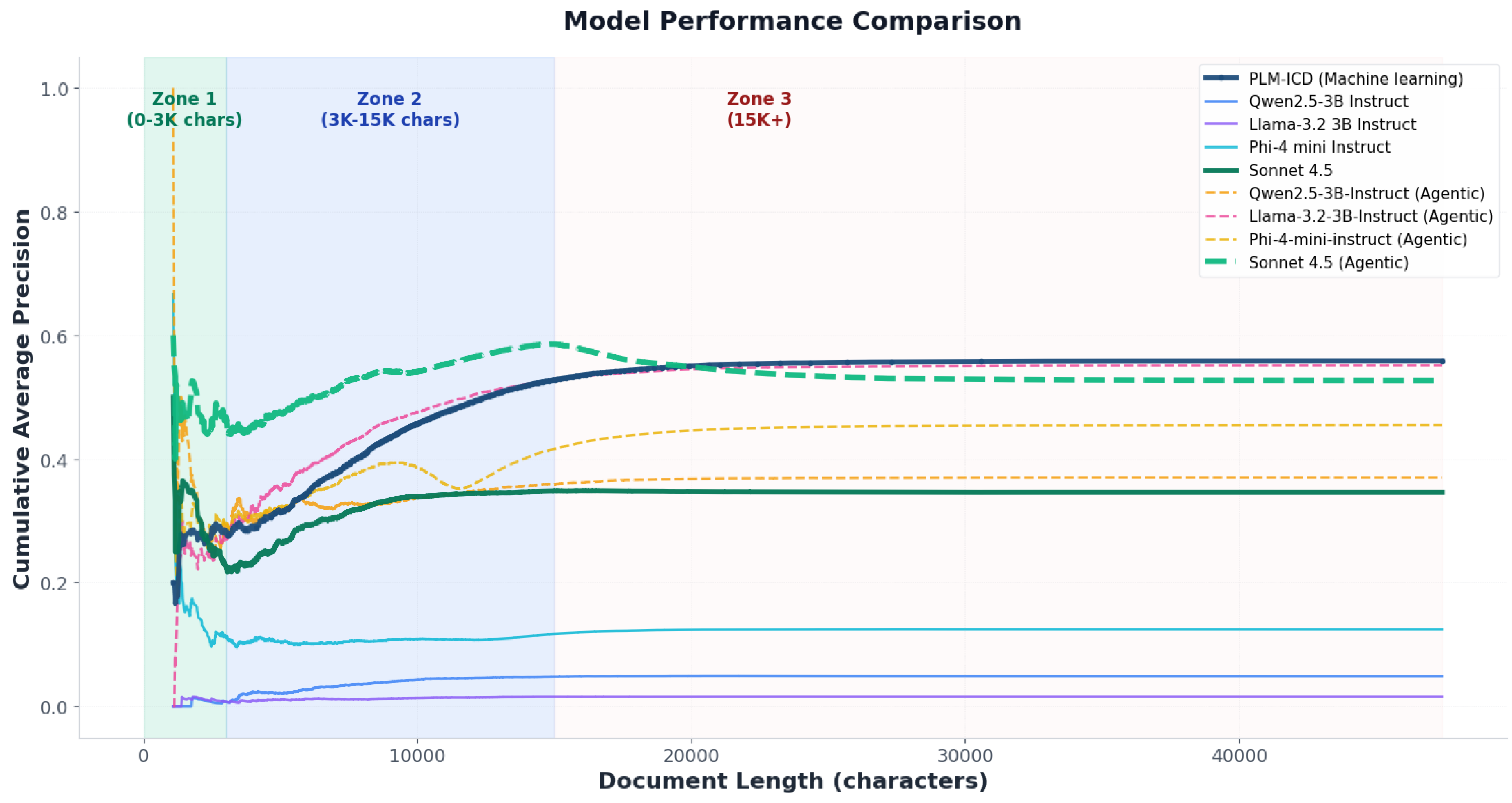
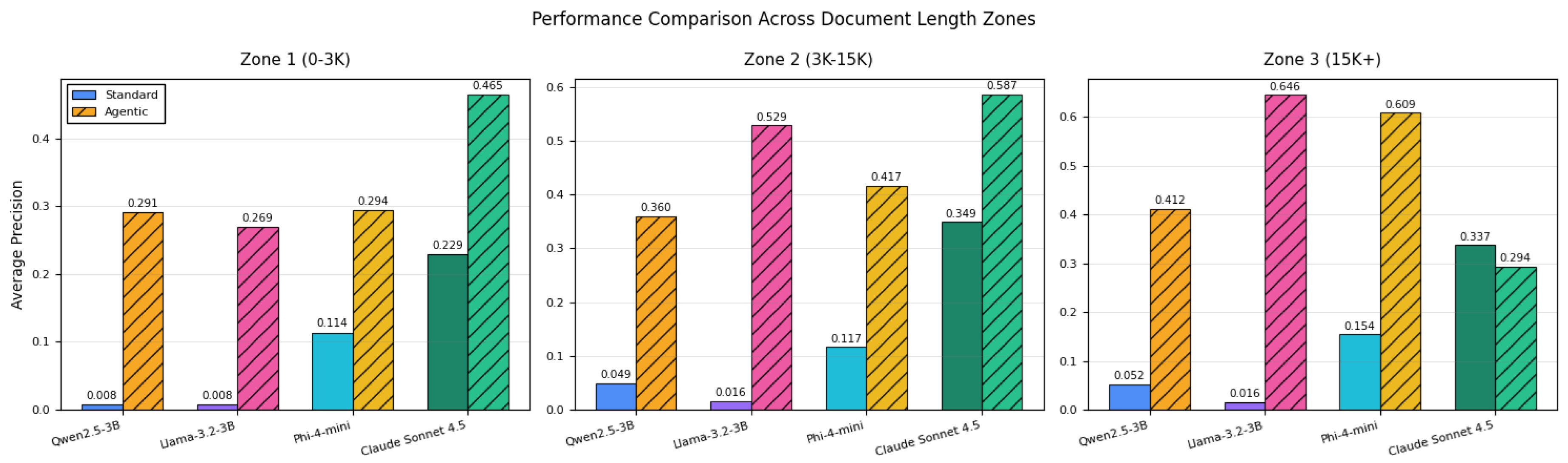
- Guidelines for document-length-aware implementation: Performance is mapped across short, medium, and long discharge summaries, providing operational guidance on when each approach is preferable.
- Evidence for filtering-centric agentic design: The study demonstrates that decomposing tasks and assigning LLMs to verification delivers the largest quality gains while preserving interpretability.
2. Related Work
2.1. Traditional Machine Learning Approaches
2.2. Deep Learning for ICD Coding
- Limited interpretability: Neural architectures work as black boxes, providing minimal insight into coding reasoning.
- Fixed prediction number: A predetermined list length (e.g., the top 15 codes) is still emitted by most models, even when the case clearly calls for more or fewer predictions.
- Difficulty with rare codes: Models struggle with long-tail distribution of infrequent diagnostic codes [29].
- Lack of uncertainty measurement: Predictions lack calibrated confidence estimates for clinical decision support.
2.3. Large Language Models in Healthcare
- Precision and reliability: Do LLMs achieve precision comparable to specialized models trained on large-scale clinical coding datasets?
- Consistency and determinism: Do LLM outputs remain stable across identical inputs—critical for clinical implementation?
- Optimal integration strategies: Should LLMs replace or complement existing specialized models?
2.4. Prompt Engineering for Medical Validation
- Clearly define the validation task and decision rules.
- Supply the necessary clinical snippets and ICD-10 descriptions so evidence remains localized.
- Request structured outputs (for example, Accept/Reject plus justification) so downstream parsers can automate follow-up actions.
- Provide few-shot exemplars that demonstrate correct validation logic.
- Constrain formatting for consistent, machine-checkable responses.
2.5. Research Gap and Contribution
3. Materials & Methods
3.1. Deep Learning Automation
3.2. Direct LLM Automation
3.3. Agentic Filtering Automation
3.4. Hardware Configuration
3.5. Language Models Evaluated
3.6. Evaluation Metrics and Research Scope
- Precision@15 (P@15): Applied to the baseline approach which produces exactly 15 predictions. It measures the proportion of correct codes among the 15 outputs:
- Precision (P): Applied to approaches with variable output sizes (LLM-only and agentic). It measures the proportion of correct predictions:
- Baseline: Fixed output of 15 codes (evaluated with P@15)
- LLM-only: Variable output size decided autonomously by the model (evaluated with P)
- Agentic: Selects variable number of codes from baseline’s 15 candidates through LLM-based filtering (evaluated with P)
4. Results
4.1. Dataset and Experimental Setup
4.2. Evaluation Dataset
4.3. Overall Performance Comparison
4.3.1. Key Performance Observations
4.3.2. Document-Length Sensitivity
5. Discussions
5.1. Interpretation of Findings
5.2. Implications for Automating Clinical Workflows Using Agentic AI
5.3. Comparison with Related Work
5.4. Precision-Oriented Design Rationale
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ICD | International Classification of Diseases |
| ICD-10 | International Classification of Diseases, 10th Revision |
| LLM | Large Language Model |
| PLM | Pretrained Language Model |
| PLM-ICD | Pretrained Language Model for ICD |
| AI | Artificial Intelligence |
| NLP | Natural Language Processing |
| MIMIC-IV | Medical Information Mart for Intensive Care IV |
| ML | Machine Learning |
| DUA | Data Use Agreement |
| CITI | Collaborative Institutional Training Initiative |
| IRB | Institutional Review Board |
| TF-IDF | Term Frequency-Inverse Document Frequency |
| BPE | Byte Pair Encoding |
| CNN | Convolutional Neural Network |
| RNN | Recurrent Neural Network |
| GPU | Graphics Processing Unit |
| mAP | Mean Average Precision |
| ROC | Receiver Operating Characteristic |
References
- World Health Organization. International Statistical Classification of Diseases and Related Health Problems 10th Revision (ICD-10), 5 ed.; World Health Organization: Geneva, Switzerland, 2016. [Google Scholar]
- Centers for Medicare; Medicaid Services; National Center for Health Statistics. ICD-10-CM Official Guidelines for Coding and Reporting FY 2024 . In Available from the Centers for Medicare and Medicaid Services and the National Center for Health Statistics; 2023. [Google Scholar]
- Henderson, T.; Shepheard, J.; Sundararajan, V. Quality of Diagnosis and Procedure Coding in ICD-10 Administrative Data. Medical Care 2006, 44, 1011–1019. [Google Scholar] [CrossRef]
- Edin, J.; Junge, A.; Havtorn, J.D.; Borgholt, L.; Maistro, M.; Ruotsalo, T.; Maaløe, L. Automated Medical Coding on MIMIC-III and MIMIC-IV: A Critical Review and Replicability Study. In Proceedings of the Proceedings of the 46th International ACM SIGIR Conference on Research and Development in Information Retrieval, ACM, Taipei, Taiwan, 2023; pp. 2572–2582. [Google Scholar] [CrossRef]
- Devlin, J.; Chang, M.W.; Lee, K.; Toutanova, K. BERT: Pre-training of Deep Bidirectional Transformers for Language Understanding. In Proceedings of the Proceedings of the 2019 Conference of the North American Chapter of the Association for Computational Linguistics, 2019; Association for Computational Linguistics; pp. 4171–4186. [Google Scholar] [CrossRef]
- Liu, Y.; Ott, M.; Goyal, N.; Du, J.; Joshi, M.; Chen, D.; Levy, O.; Lewis, M.; Zettlemoyer, L.; Stoyanov, V. RoBERTa: A Robustly Optimized BERT Pretraining Approach. arXiv 2019, arXiv:1907.11692. [Google Scholar] [CrossRef]
- Lee, J.; Yoon, W.; Kim, S.; Kim, D.; Kim, S.; So, C.H.; Kang, J. BioBERT: a pre-trained biomedical language representation model for biomedical text mining. Bioinformatics 2019, 36, 1234–1240. [Google Scholar] [CrossRef]
- Alsentzer, E.; Murphy, J.; Boag, W.; Weng, W.H.; Jindi, D.; Naumann, T.; McDermott, M. Publicly Available Clinical BERT Embeddings. In Proceedings of the Proceedings of the 2nd Clinical Natural Language Processing Workshop, 2019; Association for Computational Linguistics. [Google Scholar] [CrossRef]
- Huang, C.W.; Tsai, S.C.; Chen, Y.N. PLM-ICD: Automatic ICD Coding with Pretrained Language Models. In Proceedings of the Proceedings of the 4th Clinical Natural Language Processing Workshop, Association for Computational Linguistics, Seattle, WA, 2022; pp. 10–20. [Google Scholar] [CrossRef]
- Mosqueira-Rey, E.; Hernandez-Pereira, E.; Alonso-Rios, D.; Bobes-Bascaran, J.; Fernandez-Leal, A. Human-in-the-loop machine learning: a state of the art. Artificial Intelligence Review 2022, 56, 3005–3054. [Google Scholar] [CrossRef]
- Lu, Z.; Peng, Y.; Cohen, T.; Ghassemi, M.; Weng, C.; Tian, S. Large language models in biomedicine and health: current research landscape and future directions. Journal of the American Medical Informatics Association 2024, 31, 1801–1811. [Google Scholar] [CrossRef]
- Nori, H.; King, N.; McKinney, S.M.; Carignan, D.; Horvitz, E. Capabilities of GPT-4 on Medical Challenge Problems. arXiv 2023, arXiv:2303.13375. [Google Scholar] [CrossRef]
- Singhal, K.; Azizi, S.; Tu, T.; Mahdavi, S.S.; Wei, J.; Chung, H.W.; Scales, N.; Tanwani, A.; Cole-Lewis, H.; Pfohl, S.; et al. Large language models encode clinical knowledge. Nature 2023, 620, 172–180. [Google Scholar] [CrossRef]
- Perotte, A.; Pivovarov, R.; Natarajan, K.; Weiskopf, N.; Wood, F.; Elhadad, N. Diagnosis code assignment: models and evaluation metrics. Journal of the American Medical Informatics Association 2014, 21, 231–237. [Google Scholar] [CrossRef] [PubMed]
- Masud, J.H.B.; Kuo, C.C.; Yeh, C.Y.; Yang, H.C.; Lin, M.C. Applying Deep Learning Model to Predict Diagnosis Code of Medical Records. Diagnostics 2023, 13, 2297. [Google Scholar] [CrossRef]
- Li, H.; Mourad, A.; Zhuang, S.; Koopman, B.; Zuccon, G. Pseudo Relevance Feedback with Deep Language Models and Dense Retrievers: Successes and Pitfalls. arXiv 2022, arXiv:cs. [Google Scholar] [CrossRef]
- Manning, C.D.; Raghavan, P.; Schütze, H. Introduction to Information Retrieval; Cambridge University Press: Cambridge, UK, 2008. [Google Scholar] [CrossRef]
- Mikolov, T.; Chen, K.; Corrado, G.; Dean, J. Efficient Estimation of Word Representations in Vector Space. arXiv 2013, arXiv:1301.3781. [Google Scholar] [CrossRef]
- Otter, D.W.; Medina, J.R.; Kalita, J.K. A Survey of the Usages of Deep Learning for Natural Language Processing. IEEE Transactions on Neural Networks and Learning Systems 2021, 32, 604–624. [Google Scholar] [CrossRef]
- Sammani, A.; Bagheri, A.; van der Heijden, P.G.M.; te Riele, A.S.J.M.; Baas, A.F.; Oosters, C.A.J.; Oberski, D.; Asselbergs, F.W. Automatic Multilabel Detection of ICD-10 Codes in Dutch Cardiology Discharge Letters Using Neural Networks. npj Digital Medicine 2021, 4. [Google Scholar] [CrossRef]
- Xie, P.; Xing, E. A Neural Architecture for Automated ICD Coding. Proceedings of the Proceedings of the 56th Annual Meeting of the Association for Computational Linguistics 2018, Volume 1, 1066–1076. [Google Scholar] [CrossRef]
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, L.; Polosukhin, I. Attention is All you Need. Advances in Neural Information Processing Systems 2017, 30, 5998–6008. [Google Scholar] [CrossRef]
- Mullenbach, J.; Wiegreffe, S.; Duke, J.; Sun, J.; Eisenstein, J. Explainable Prediction of Medical Codes from Clinical Text. Proceedings of the Proceedings of the 2018 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies. Association for Computational Linguistics 2018, Vol. 1, 1101–1111. [Google Scholar] [CrossRef]
- Vu, T.; Nguyen, D.Q.; Nguyen, A. A Label Attention Model for ICD Coding from Clinical Text. In Proceedings of the Proceedings of the 29th International Joint Conference on Artificial Intelligence, 2020; pp. 3335–3341. [Google Scholar] [CrossRef]
- Liu, L.; Perez-Concha, O.; Nguyen, A.; Bennett, V.; Jorm, L. Hierarchical label-wise attention transformer model for explainable ICD coding. Journal of Biomedical Informatics 2022, 133, 104161. [Google Scholar] [CrossRef]
- Li, F.; Yu, H. ICD Coding from Clinical Text Using Multi-Filter Residual Convolutional Neural Network. Proceedings of the AAAI Conference on Artificial Intelligence 2020, 34, 8180–8187. [Google Scholar] [CrossRef]
- Qiu, W.; Wu, Y.; Li, Y.; Niu, K.; Zeng, M.; Li, M. DILM-ICD: A Deep Iterative Learning Model for Automatic ICD Coding. In Proceedings of the 2023 IEEE International Conference on Bioinformatics and Biomedicine (BIBM). IEEE, 2023; pp. 1394–1399. [Google Scholar] [CrossRef]
- Zhang, W.; Yan, J.; Wang, X.; Zha, H. Deep Extreme Multi-label Learning. In Proceedings of the Proceedings of the 2018 ACM on International Conference on Multimedia Retrieval. ACM, 2018; pp. 100–107. [Google Scholar] [CrossRef]
- Rios, A.; Kavuluru, R. Few-Shot and Zero-Shot Multi-Label Learning for Structured Label Spaces. In Proceedings of the Proceedings of the 2018 Conference on Empirical Methods in Natural Language Processing, 2018; Association for Computational Linguistics; pp. 3132–3142. [Google Scholar] [CrossRef]
- Touvron, H.; Martin, L.; Stone, K.; Albert, P.; Almahairi, A.; Babaei, Y.; Bashlykov, N.; Batra, S.; Bhargava, P.; Bhosale, S.; et al. Llama 2: Open Foundation and Fine-Tuned Chat Models. arXiv 2023, arXiv:2307.09288. [Google Scholar] [CrossRef]
- Abdin, M.; Aneja, J.; Behl, H.; Bubeck, S.; Eldan, R.; Gunasekar, S.; Harrison, M.; Hewett, R.J.; Javaheripi, M.; Kauffmann, P.; et al. Phi-4 Technical Report. arXiv 2024, arXiv:2412.08905. [Google Scholar] [CrossRef]
- Bai, Y.; Kadavath, S.; Kundu, S.; Askell, A.; Kernion, J.; Jones, A.; Chen, A.; Goldie, A.; Mirhoseini, A.; McKinnon, C.; et al. Constitutional AI: Harmlessness from AI Feedback. arXiv 2022, arXiv:2212.08073. [Google Scholar] [CrossRef]
- Jiang, F.; Jiang, Y.; Zhi, H.; Dong, Y.; Li, H.; Ma, S.; Wang, Y.; Dong, Q.; Shen, H.; Wang, Y. Artificial intelligence in healthcare: past, present and future. Stroke and Vascular Neurology 2017, 2, 230–243. [Google Scholar] [CrossRef]
- Kim, Y.; Jeong, H.; Park, C.; Park, E.; Zhang, H.; Liu, X.; Lee, H.; McDuff, D.; Ghassemi, M.; Breazeal, C.; et al. Tiered Agentic Oversight: A Hierarchical Multi-Agent System for Healthcare Safety. arXiv 2025, arXiv:2506.12482. [Google Scholar] [CrossRef]
- Liu, J.; Liu, F.; Wang, C.; Liu, S. Prompt Engineering in Clinical Practice: Tutorial for Clinicians. Journal of Medical Internet Research 2025, 27, e72644–e72644. [Google Scholar] [CrossRef]
- Sivarajkumar, S.; Kelley, M.; Samolyk-Mazzanti, A.; Visweswaran, S.; Wang, Y. An Empirical Evaluation of Prompting Strategies for Large Language Models in Zero-Shot Clinical Natural Language Processing: Algorithm Development and Validation Study. JMIR Medical Informatics 2024, 12, e55318. [Google Scholar] [CrossRef]
- Wang, L.; Chen, X.; Deng, X.; Wen, H.; You, M.; Liu, W.; Li, Q.; Li, J. Prompt engineering in consistency and reliability with the evidence-based guideline for LLMs. npj Digital Medicine 2024, 7, 41. [Google Scholar] [CrossRef] [PubMed]
- Zaghir, J.; Naguib, M.; Bjelogrlic, M.; Névéol, A.; Tannier, X.; Lovis, C. Prompt Engineering Paradigms for Medical Applications: Scoping Review. Journal of Medical Internet Research 2024, 26, e60501. [Google Scholar] [CrossRef] [PubMed]
- Zaghir, J.; Naguib, M.; Bjelogrlic, M.; Névéol, A.; Tannier, X.; Lovis, C. Prompt engineering paradigms for medical applications: scoping review and recommendations for better practices. arXiv;arXiv version of the paper published in 2024. 2024, arXiv:2405.01249. [Google Scholar] [CrossRef]
- Zouhar, V.; Meister, C.; Gastaldi, J.L.; Du, L.; Vieira, T.; Sachan, M.; Cotterell, R. A Formal Perspective on Byte-Pair Encoding. arXiv 2023, arXiv:2306.16837. [Google Scholar] [CrossRef]
- Bai, J.; Bai, S.; Chu, Y.; Cui, Z.; Dang, K.; Deng, X.; Fan, Y.; Ge, W.; Han, Y.; Huang, F.; et al. Qwen Technical Report. arXiv 2023, arXiv:2309.16609. [Google Scholar] [CrossRef]
- Davis, J.; Goadrich, M. The Relationship Between Precision-Recall and ROC Curves. In Proceedings of the Proceedings of the 23rd International Conference on Machine Learning (ICML ’06), 2006; ACM; pp. 233–240. [Google Scholar] [CrossRef]
- Collins, G.S.; Moons, K.G.; Dhiman, P.; Riley, R.D.; Beam, A.L.; Van Calster, B.; Ghassemi, M.; Liu, X.; Reitsma, J.B.; van Smeden, M.; et al. TRIPOD+AI statement: updated guidance for reporting clinical prediction models that use regression or machine learning methods. BMJ 2024, 385, e078378. [Google Scholar] [CrossRef]
- Johnson, A.; Pollard, T.; Horng, S.; Celi, L.A.; Mark, R. MIMIC-IV-Note: Deidentified free-text clinical notes; PhysioNet, 2023. [Google Scholar] [CrossRef]
- Johnson, A.E.W.; Bulgarelli, L.; Shen, L.; Gayles, A.; Shammout, A.; Horng, S.; Pollard, T.J.; Hao, S.; Moody, B.; Gow, B.; et al. MIMIC-IV, a freely accessible electronic health record dataset. Scientific Data 2023, 10, 1. [Google Scholar] [CrossRef] [PubMed]
- Pham, H.; Wang, G.; Lu, Y.; Florencio, D.; Zhang, C. Understanding Long Documents with Different Position-Aware Attentions. arXiv 2022, arXiv:2208.08201. [Google Scholar] [CrossRef]
- Sim, I.; Gorman, P.; Greenes, R.A.; Haynes, R.B.; Kaplan, B.; Lehmann, H.; Tang, P.C. Clinical Decision Support Systems for the Practice of Evidence-based Medicine. Journal of the American Medical Informatics Association 2001, 8, 527–534. [Google Scholar] [CrossRef]
- Dahmke, H.; Fiumefreddo, R.; Schuetz, P.; De Iaco, R.; Zaugg, C. Tackling alert fatigue with a semi-automated clinical decision support system: quantitative evaluation and end-user survey. Swiss Medical Weekly 2023, 153, 40082. [Google Scholar] [CrossRef] [PubMed]
- Soroush, A.; Glicksberg, B.S.; Zimlichman, E.; Barash, Y.; Freeman, R.; Charney, A.W.; Nadkarni, G.N.; Klang, E. Large Language Models Are Poor Medical Coders – Benchmarking of Medical Code Querying. NEJM AI 2024, 1, AIdbp2300040. [Google Scholar] [CrossRef]

| Approach | Model | Metric | Precision (%) | Avg Out |
|---|---|---|---|---|
| Baseline | PLM-ICD | P@15 | 55.8 | 15.0 |
|
LLM-Only Generation |
Qwen2.5-3B-Instruct | P | 4.9 | 4.9 |
| Llama-3.2-3B-Instruct | P | 1.5 | 5.9 | |
| Phi-4-mini-instruct | P | 12.4 | 7.0 | |
| Sonnet-4.5 | P | 34.6 | 9.8 | |
|
Agentic Filtering |
PLM-ICD + Qwen2.5-3B | P | 37.0 | 2.0 |
| PLM-ICD + Llama-3.2-3B | P | 55.1 | 4.0 | |
| PLM-ICD + Phi-4-mini-instruct | P | 45.5 | 7.0 | |
| PLM-ICD + Sonnet-4.5 | P | 52.6 | 7.9 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




