Submitted:
18 December 2025
Posted:
19 December 2025
You are already at the latest version
Abstract
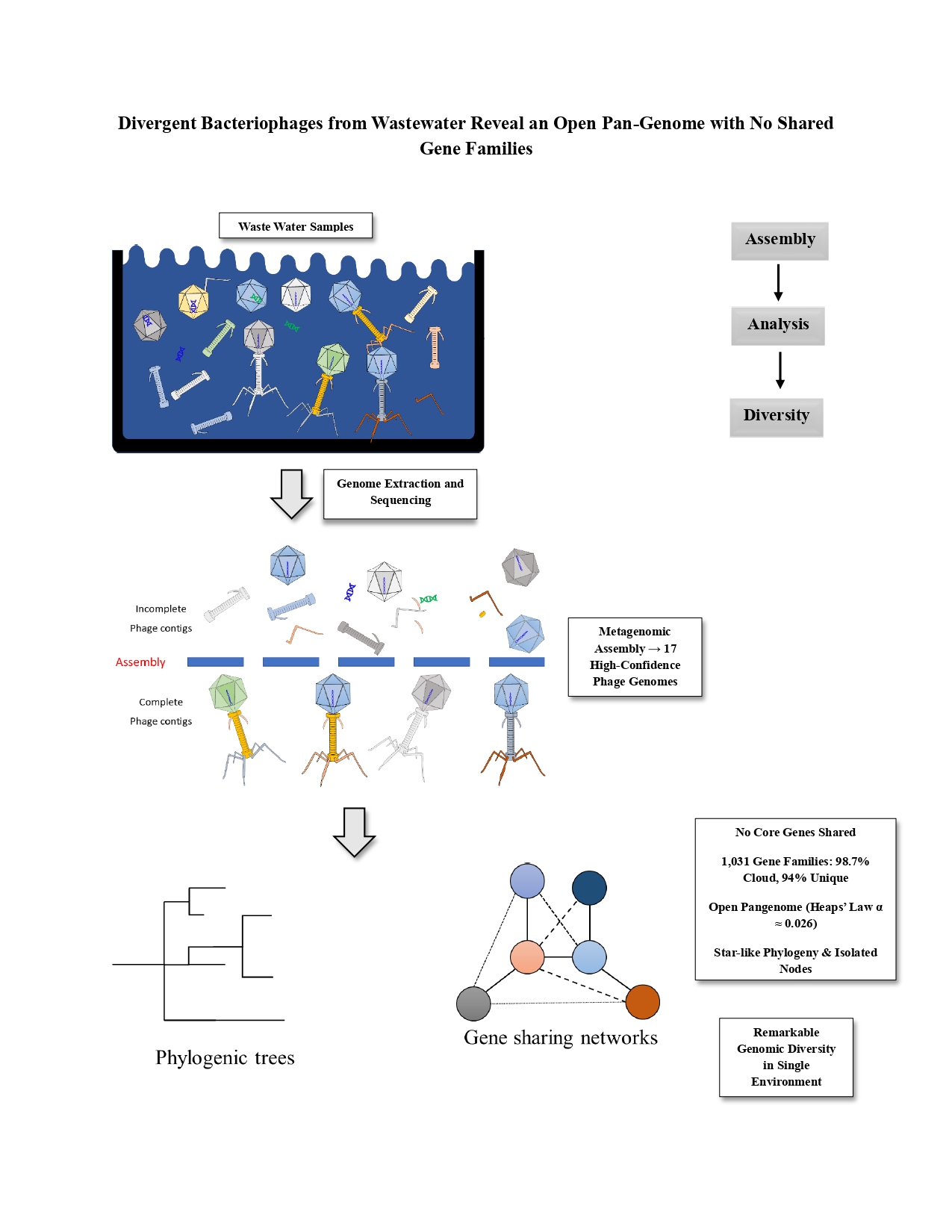
Keywords:
1. Introduction
2. Materials and Methods
2.1. Data Collection
2.2. Data Manipulation
2.3. Phage Prediction and Annotation
2.4. Computing Environment
2.5. Genome Parsing and Protein Extraction
2.6. Clustering Proteins into Gene Families
2.7. Presence–Absence Matrix and Gene Partitions
2.8. Definition of Pangenome Metrics
2.9. Pangenome Accumulation and Openness
2.10. Gene-Content Distances, Clustering, and Heatmaps
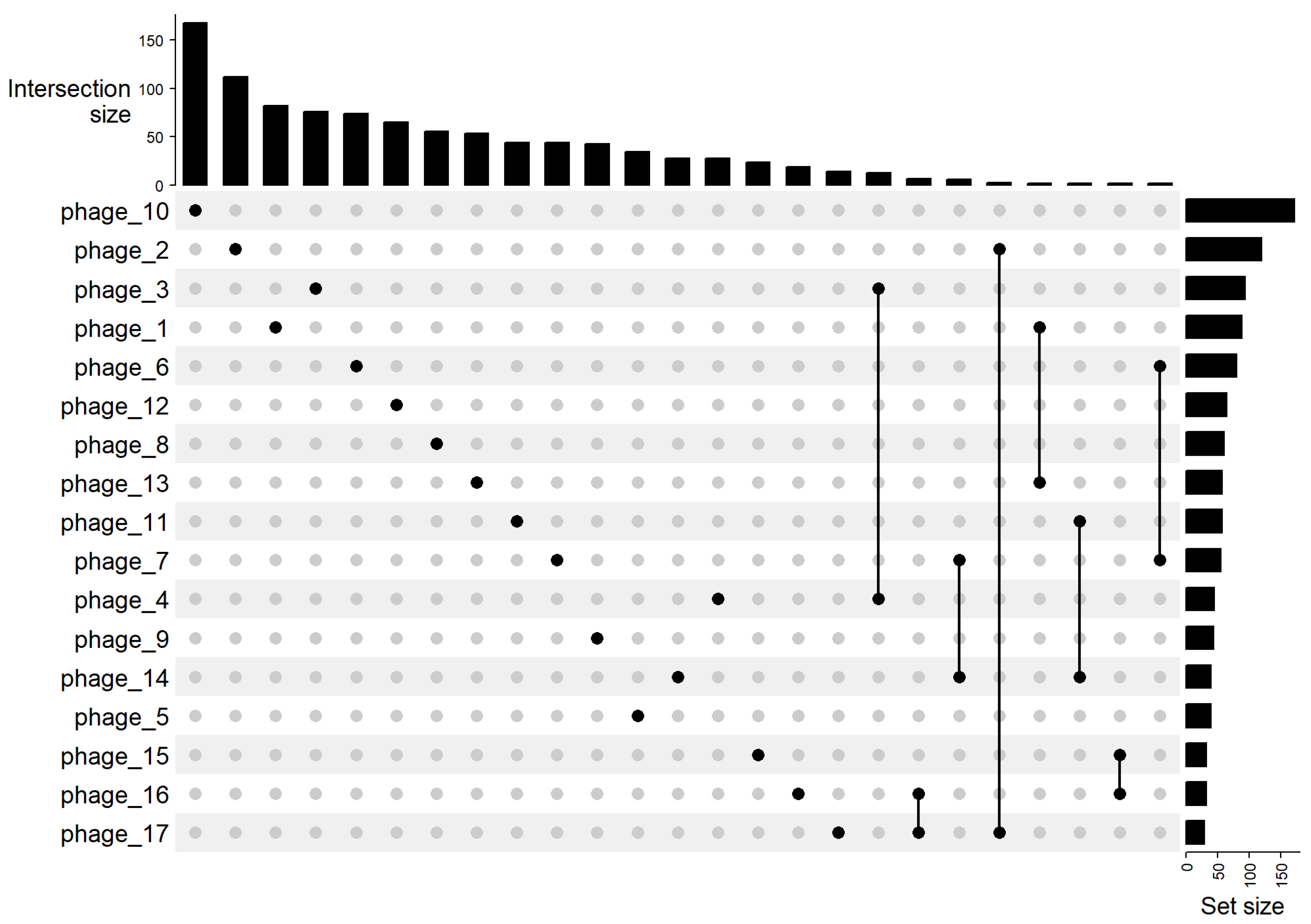
2.11. Intersections of Shared Gene Families
2.12. Genome–Genome Gene-Sharing Network
2.13. Ancestral Reconstruction of Presence/Absence and Branch Dynamics
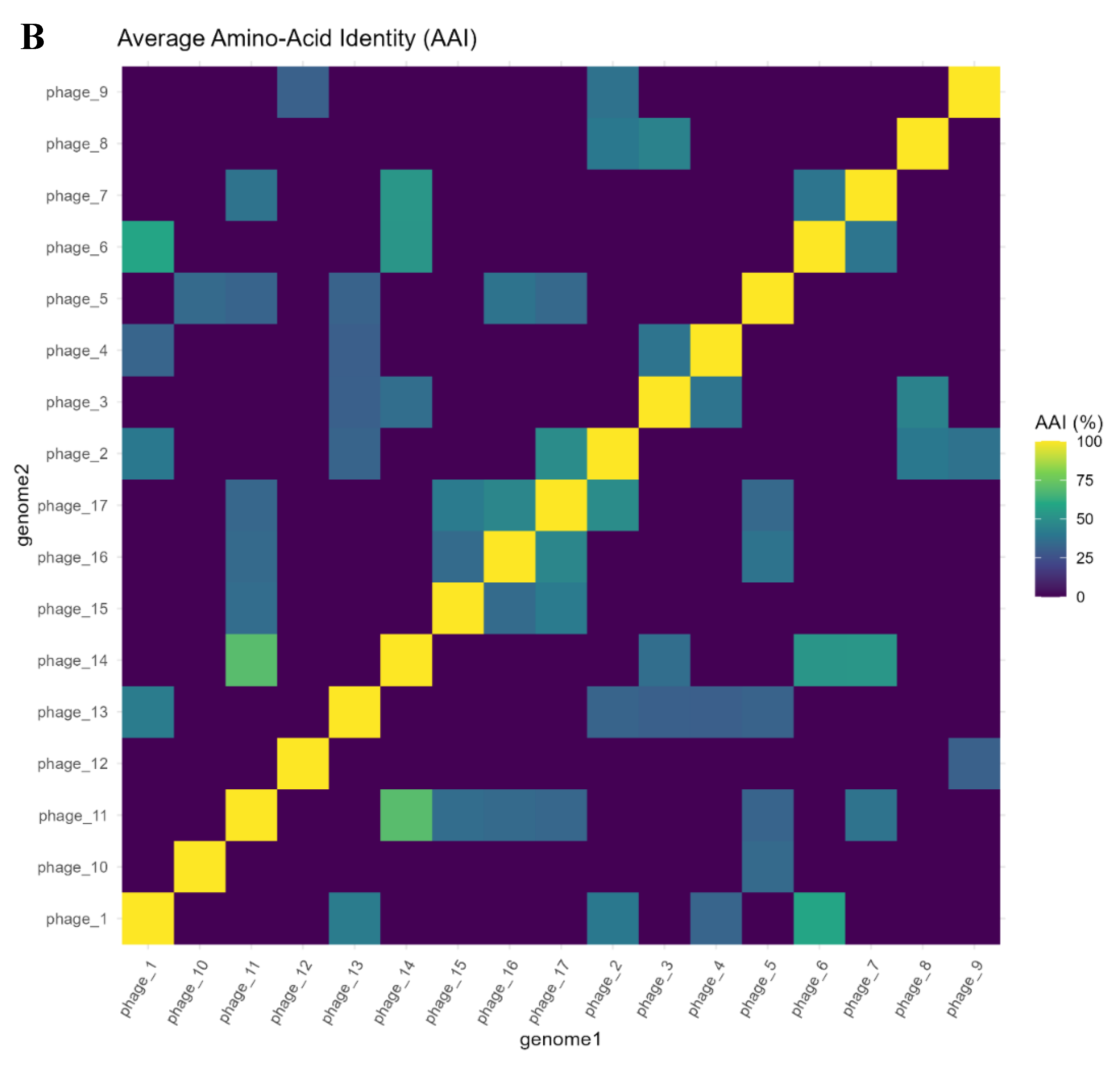
2.14. Proteome-Level Relatedness by AAI
2.15. Visualization and Reporting
3. Results
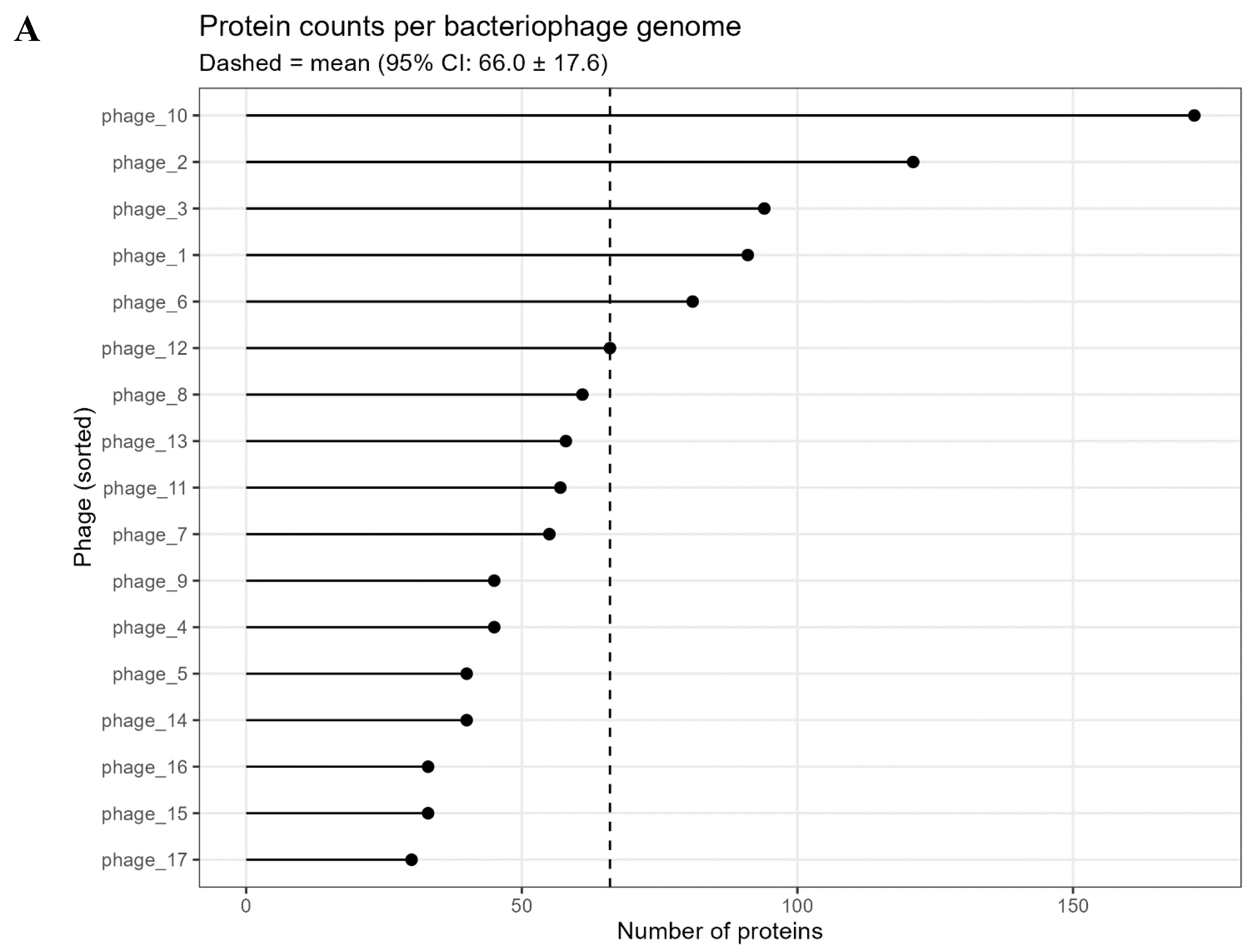
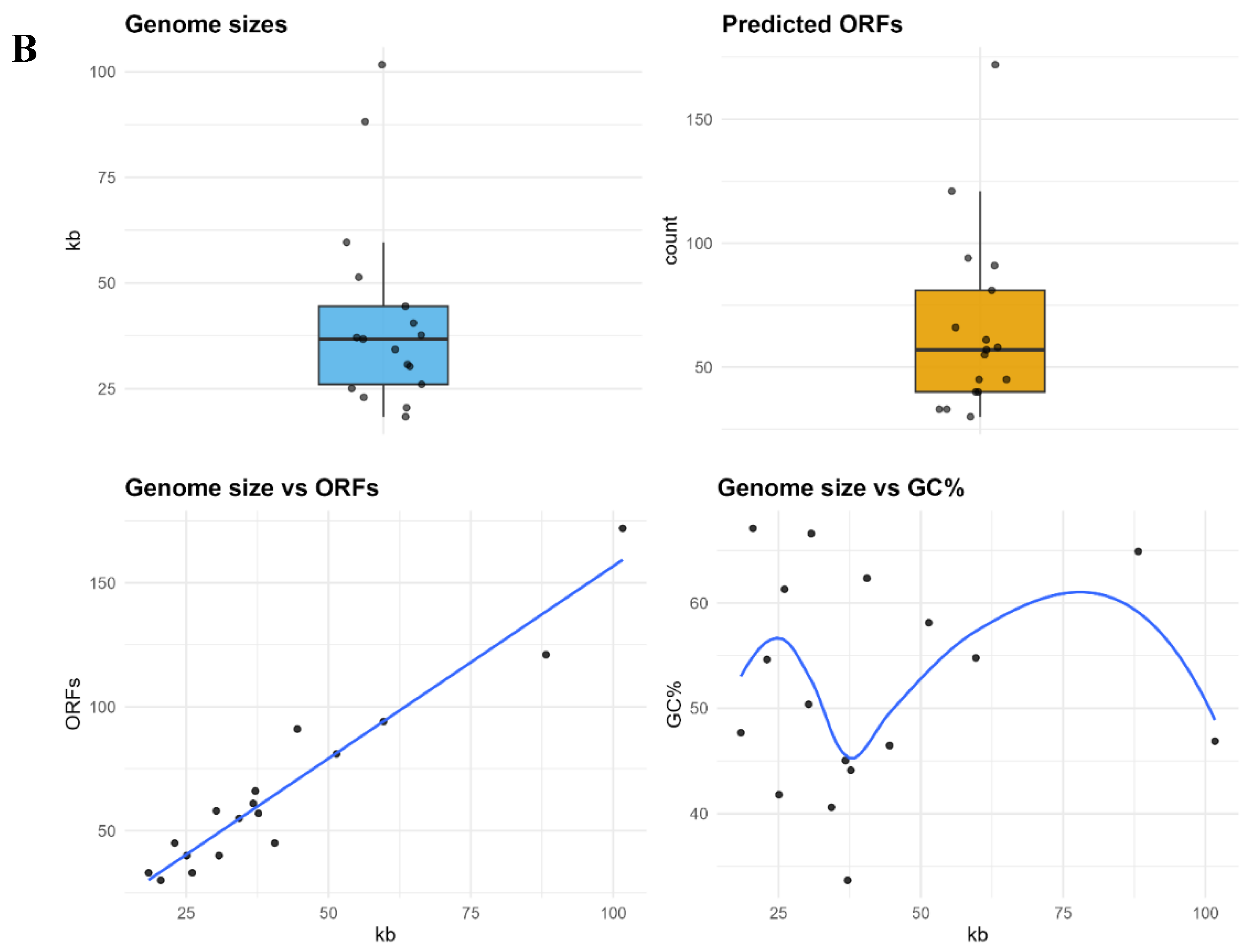
3.1. Phage Identification and Annotation
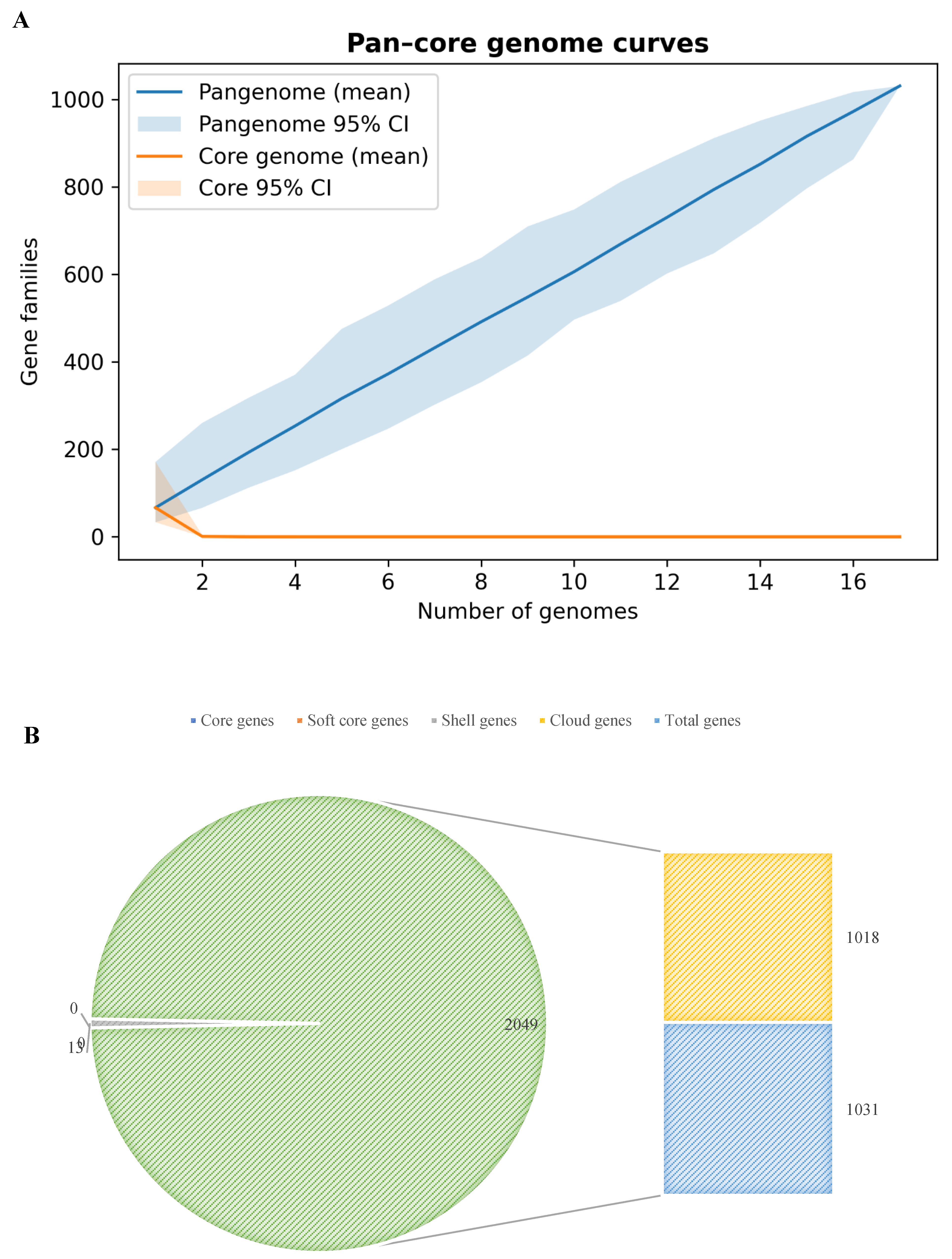
3.2. Pangenome Composition and Statistics
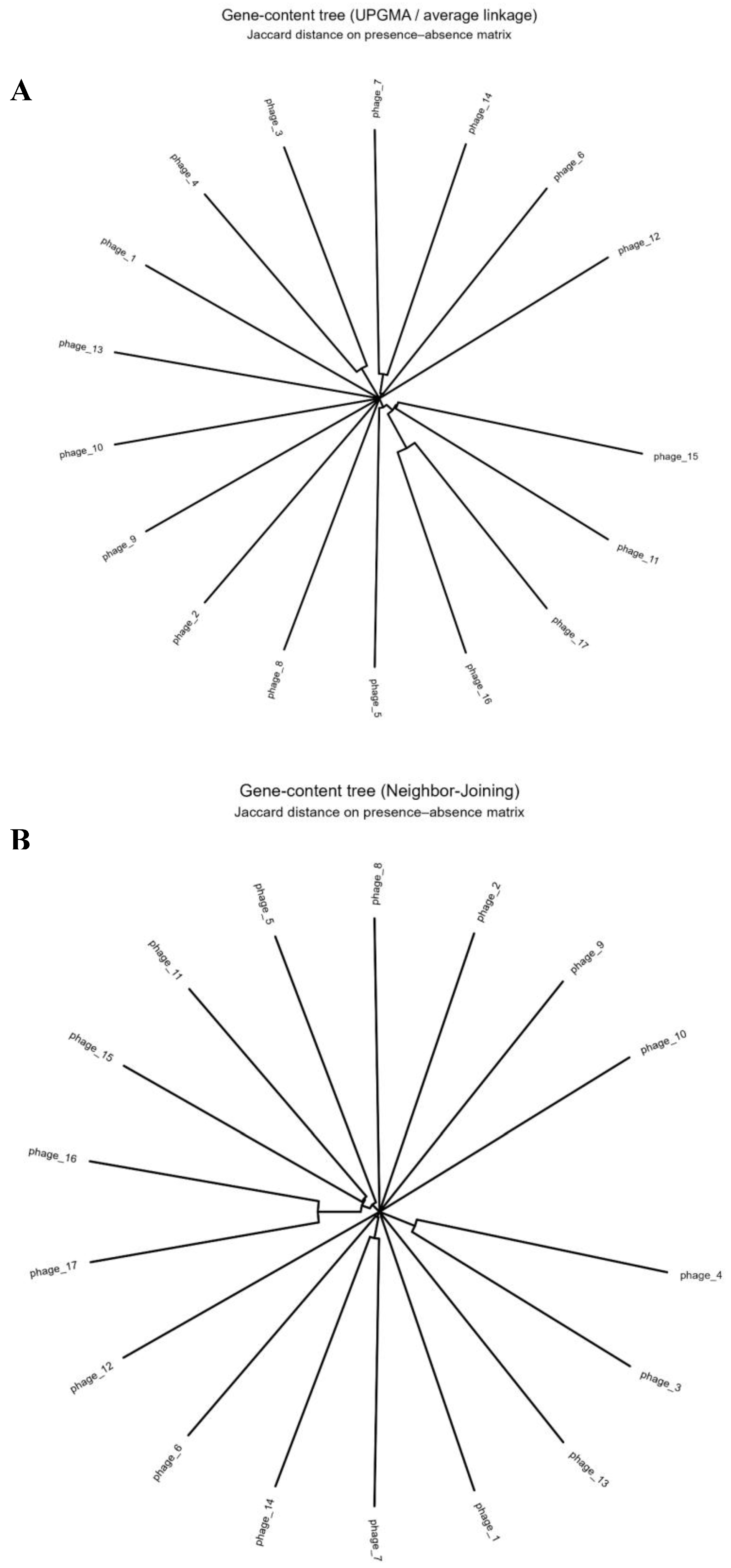
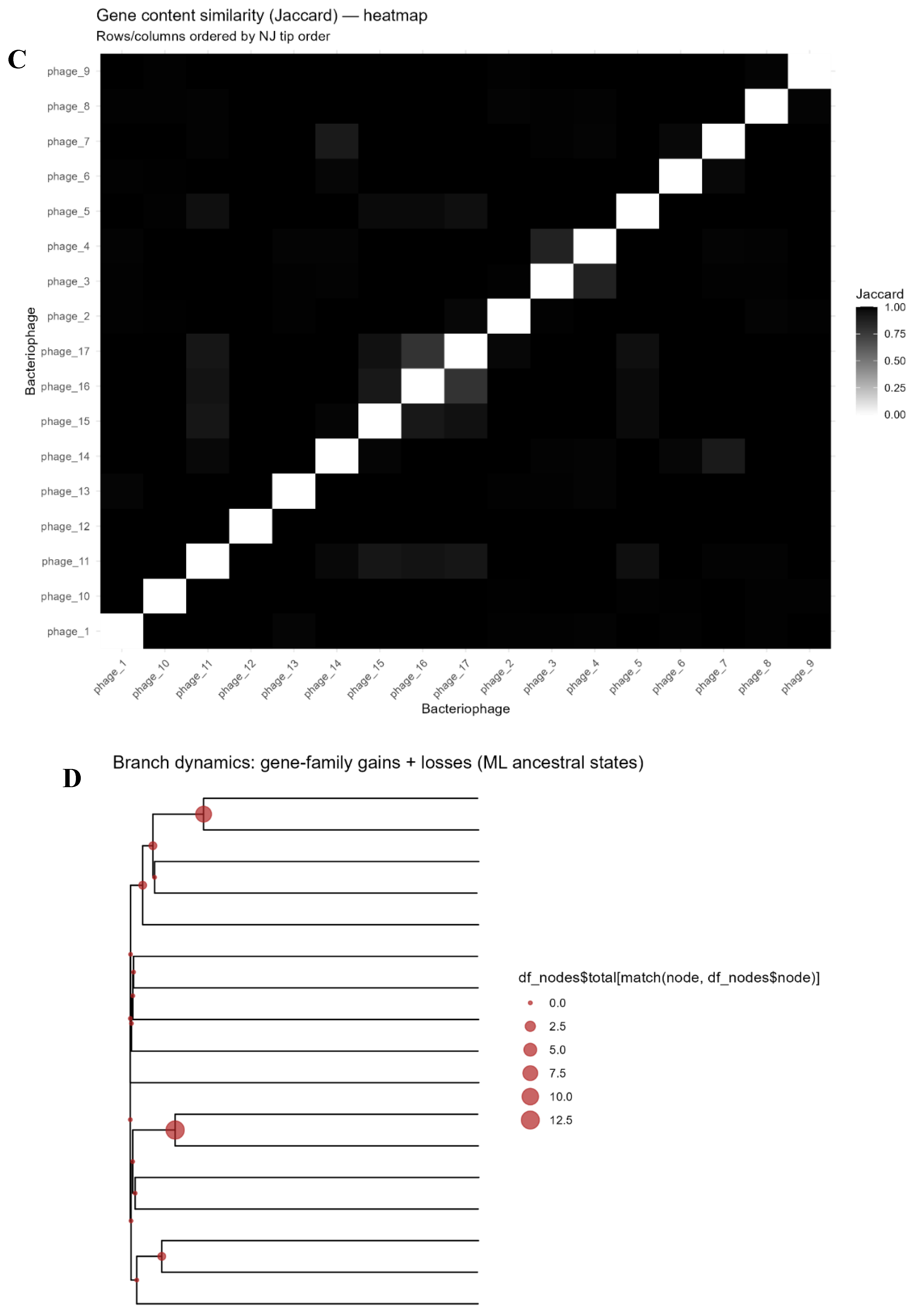
3.3. Gene Content Similarity and Phylogeny
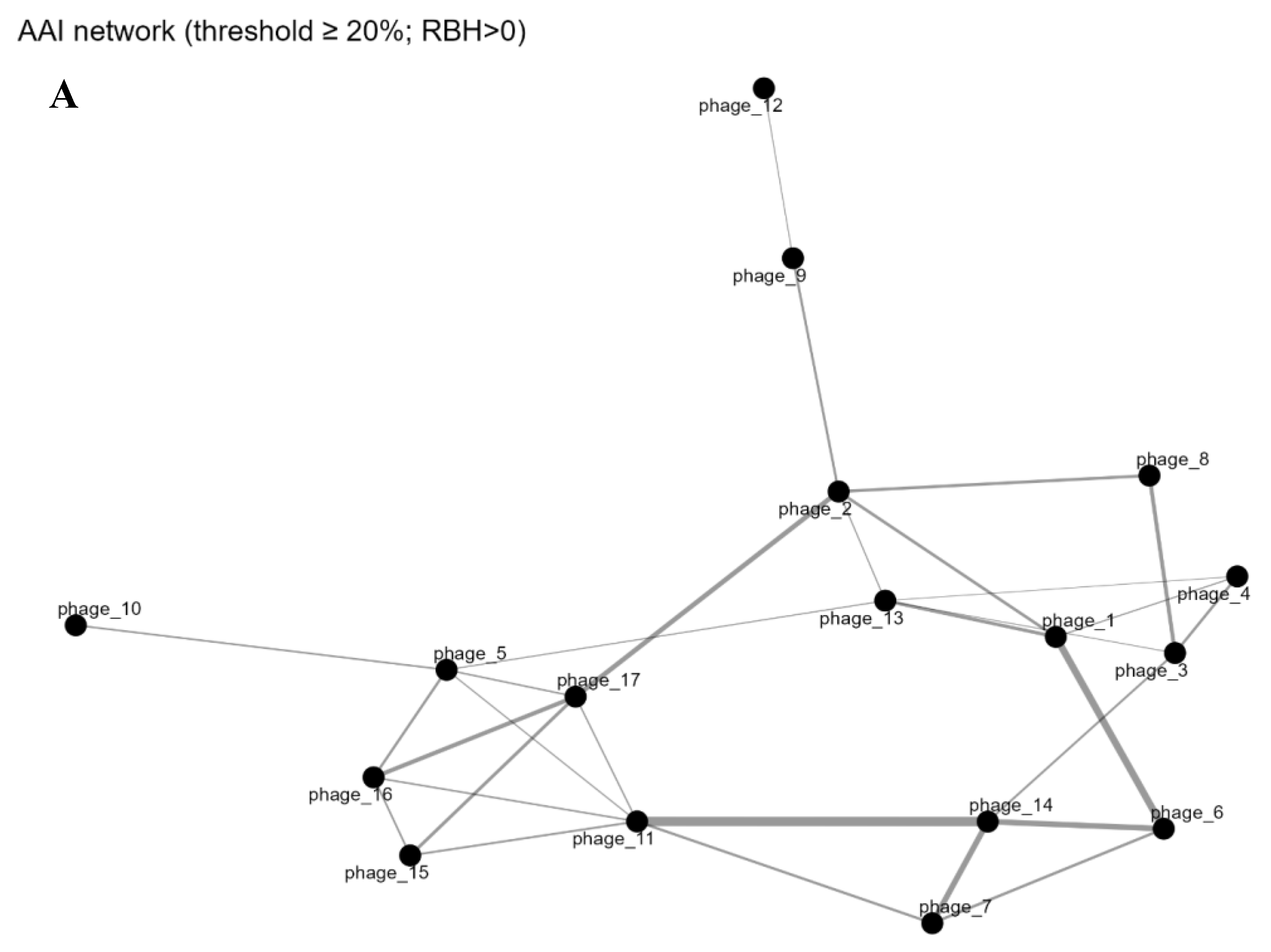
3.4. Gene-Sharing Network and Intersections
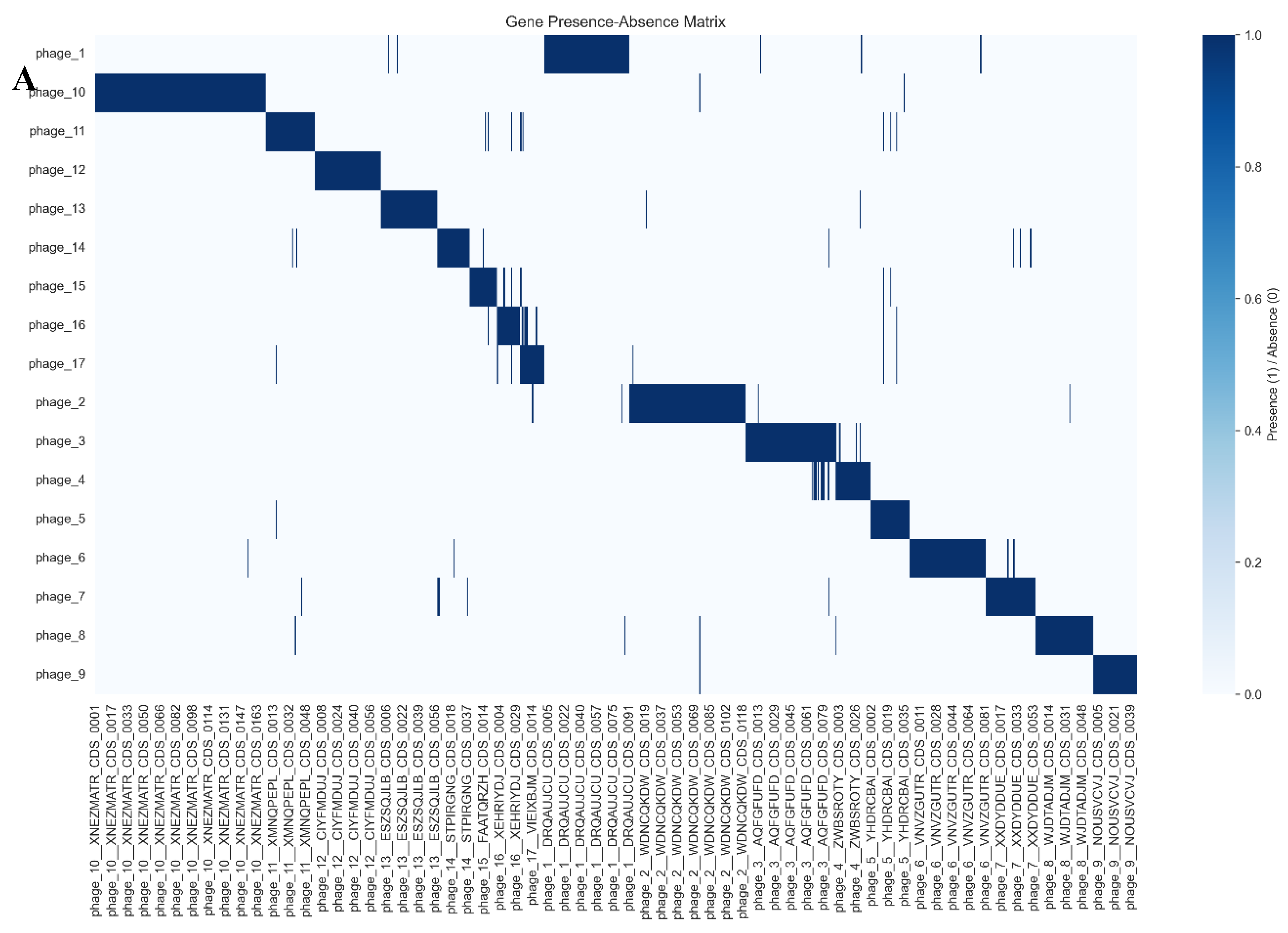

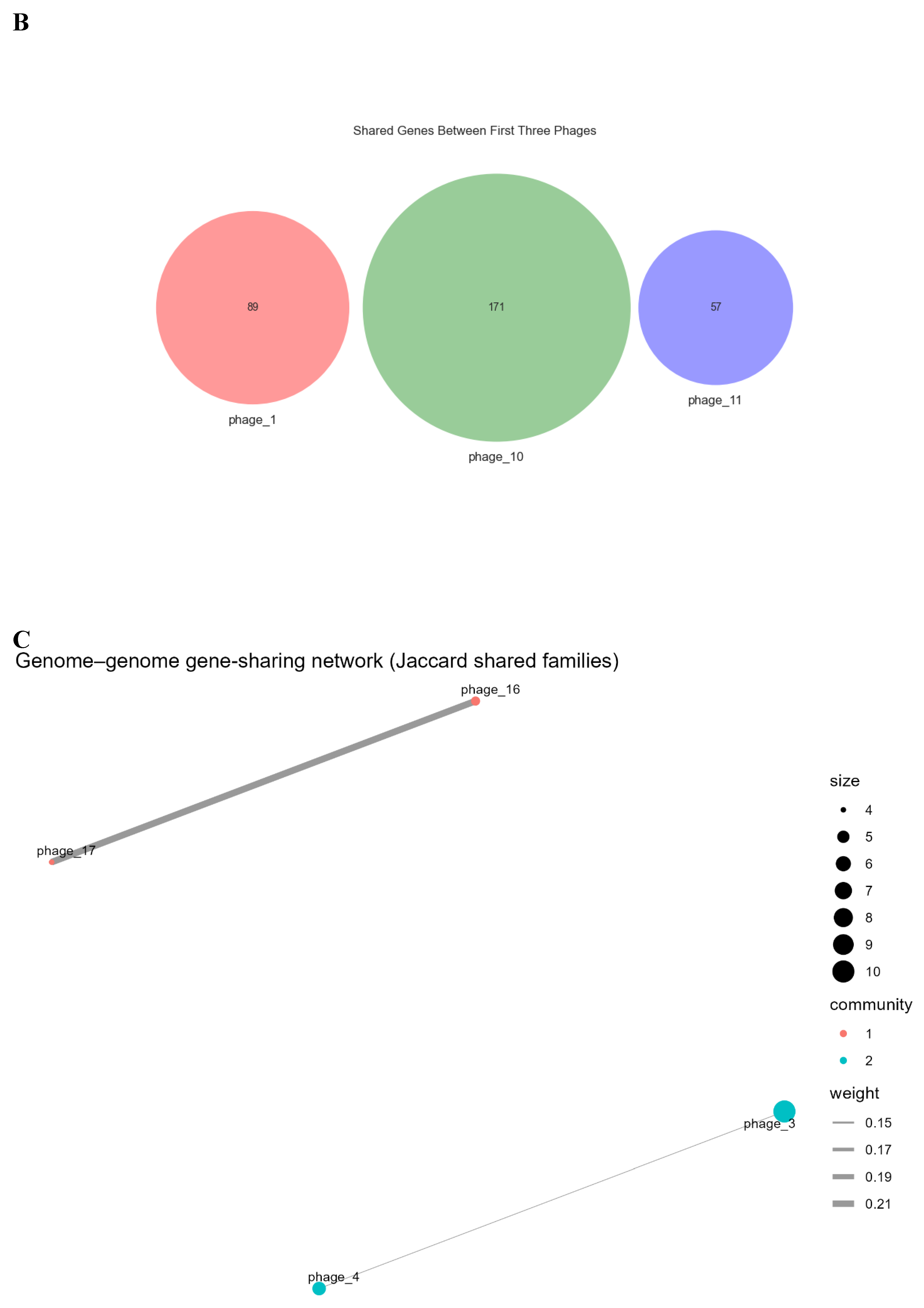
3.5. Proteome-Level Relatedness and Evolutionary Dynamics
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Ethics approval and consent to participate
Availability of data and materials
Consent for publication
Clinical trial
Acknowledgments
Conflicts of Interest Statement
References
- Jurczak-Kurek, A; Gasior, T; Nejman-Faleńczyk, B; Bloch, S; Dydecka, A; Topka, G; et al. Biodiversity of bacteriophages: morphological and biological properties of a large group of phages isolated from urban sewage. Scientific Reports 2016 6:1; Nature Publishing Group, 4 Oct 2016; Volume 6, 1, pp. 1–17. Available online: https://www.nature.com/articles/srep34338.
- Zrelovs, N; Dislers, A; Kazaks, A. Motley Crew: Overview of the Currently Available Phage Diversity. Front Microbiol. Frontiers Media S.A 2020, 11, 579452. [Google Scholar] [CrossRef]
- Fremin, BJ; Bhatt, AS; Kyrpides, NC; Sengupta, A; Sczyrba, A; Maria da Silva, A; et al. Thousands of small, novel genes predicted in global phage genomes. Cell Rep [En ligne]; Elsevier B.V., 21 Jun 2022; Volume 39, 12, p. 110984. Available online: https://pmc.ncbi.nlm.nih.gov/articles/PMC9254267/.
- Benler, S; Yutin, N; Antipov, D; Rayko, M; Shmakov, S; Gussow, AB; et al. Thousands of previously unknown phages discovered in whole-community human gut metagenomes. Microbiome 2021 9:1 [En ligne]. BioMed Central 29 Mar 2021 [cité le 8 Nov 2025, 9(1), 1–17. Available online: https://microbiomejournal.biomedcentral.com/articles/10.1186/s40168-021-01017-w.
- Hatfull, GF. Dark Matter of the Biosphere: the Amazing World of Bacteriophage Diversity. In J Virol [En ligne]; cité le; American Society for Microbiology, 15 Aug 2015; Volume 89, 16, pp. 8107–10, https://journals.asm.org/doi/10.1128/jvi.01340-15. [Google Scholar]
- Pope, WH; Bowman, CA; Russell, DA; Jacobs-Sera, D; Asai, DJ; Cresawn, SG; et al. Whole genome comparison of a large collection of mycobacteriophages reveals a continuum of phage genetic diversity. Elife [En ligne]; eLife Sciences Publications Ltd, 2015; Volume 4, p. e06416. Available online: https://pmc.ncbi.nlm.nih.gov/articles/PMC4408529/.
- Hayes, S; Mahony, J; Nauta, A; Van Sinderen, D. Metagenomic Approaches to Assess Bacteriophages in Various Environmental Niches. Viruses [En ligne]. Viruses. 1 Jun 2017 [cité le 8 Nov 2025, 9. Available online: https://pubmed.ncbi.nlm.nih.gov/28538703/.
- Bellas, CM; Schroeder, DC; Edwards, A; Barker, G; Anesio, AM. Flexible genes establish widespread bacteriophage pan-genomes in cryoconite hole ecosystems. Nature Communications 2020 11:1 [En ligne]; Nature Publishing Group, 2 Sep 2020; Volume 11, 1, pp. 1–10. Available online: https://www.nature.com/articles/s41467-020-18236-8.
- Shirzad-Aski, H; Yazdi, M; Mohebbi, A; Rafiee, M; Soleimani-Delfan, A; Tabarraei, A; et al. Isolation, characterization, and genomic analysis of three novel Herelleviridae family lytic bacteriophages against uropathogenic isolates of Staphylococcus saprophyticus. Virology Journal 2025 22:1 [En ligne]. BioMed Central 2025, 22(1), 1–15. Available online: https://virologyj.biomedcentral.com/articles/10.1186/s12985-025-02710-0.
- Borodovich, T; Shkoporov, AN; Ross, RP; Hill, C. Phage-mediated horizontal gene transfer and its implications for the human gut microbiome. Gastroenterol Rep (Oxf) [En ligne]; Oxford University Press, 2022; Volume 10, p. goac012. Available online: https://pmc.ncbi.nlm.nih.gov/articles/PMC9006064/.
- Sher, S; Khan, HA; Khan, Z; Siddique, MS; Bukhari, DA; Rehman, A. Bacteriophages: Potential Candidates for the Dissemination of Antibiotic Resistance Genes in the Environment. Targets 2025, Vol 3, Page 25 [En ligne]; Multidisciplinary Digital Publishing Institute, 22 Jul 2025; Volume 3, 3, p. 25. Available online: https://www.mdpi.com/2813-3137/3/3/25/htm.
- Moura de Sousa, JA; Pfeifer, E; Touchon, M; Rocha, EPC. Causes and Consequences of Bacteriophage Diversification via Genetic Exchanges across Lifestyles and Bacterial Taxa. In Mol Biol Evol [En ligne]; cité le; Oxford Academic, 19 May 2021; Volume 38, 6, pp. 2497–512. [Google Scholar] [CrossRef]
- Dkhili, S; Ribeiro, M; Slama, K; Ben. A Century of Bacteriophages: Insights, Applications, and Current Utilization. Antibiotics 2025, Vol 14, Page 1080 [En ligne; Multidisciplinary Digital Publishing Institute, 27 Oct 2025; Volume 14, 11, p. 1080. Available online: https://www.mdpi.com/2079-6382/14/11/1080/htm.
- Sahoo, K; Meshram, S. The Evolution of Phage Therapy: A Comprehensive Review of Current Applications and Future Innovations. In Cureus [En ligne; Springer Science and Business Media LLC, 29 Sep 2024; Volume 16, 9, p. e70414. Available online: https://pmc.ncbi.nlm.nih.gov/articles/PMC11519598/.
- Hatfull, GF. Dark Matter of the Biosphere: the Amazing World of Bacteriophage Diversity. J Virol. American Society for Microbiology 2015, 89(16), 8107–10. [Google Scholar] [CrossRef] [PubMed]
- Weitz, JS; Poisot, T; Meyer, JR; Flores, CO; Valverde, S; Sullivan, MB; et al. Phage-bacteria infection networks. Trends Microbiol [En ligne]. Trends Microbiol Feb 2013 [cité le 8 Nov 2025, 21(2), 82–91. Available online: https://pubmed.ncbi.nlm.nih.gov/23245704/. [CrossRef] [PubMed]
- Naureen, Z; Dautaj, A; Anpilogov, K; Camilleri, G; Dhuli, K; Tanzi, B; et al. Bacteriophages presence in nature and their role in the natural selection of bacterial populations. Acta Bio Medica : Atenei Parmensis [En ligne]. Mattioli 1885 2020, 91 Suppl 13, e2020024. Available online: https://pmc.ncbi.nlm.nih.gov/articles/PMC8023132/.
- Ranveer, SA; Dasriya, V; Ahmad, MF; Dhillon, HS; Samtiya, M; Shama, E; et al. Positive and negative aspects of bacteriophages and their immense role in the food chain. npj Science of Food 2024 8:1 [En ligne]; Nature Publishing Group, 3 Jan 2024; Volume 8, 1, pp. 1–13. Available online: https://www.nature.com/articles/s41538-023-00245-8.
- Nurk, S; Meleshko, D; Korobeynikov, A; Pevzner, PA. metaSPAdes: a new versatile metagenomic assembler. In Genome Res [En ligne]; Cold Spring Harbor Laboratory Press, 1 May 2017; Volume 27, 5, pp. 824–34. Available online: http://genome.cshlp.org/content/27/5/824.full.
- Wood, DE; Salzberg, SL. Kraken: Ultrafast metagenomic sequence classification using exact alignments. In Genome Biol [En ligne]; BioMed Central Ltd., 2014; Volume 15, 3, pp. 1–12. Available online: https://genomebiology.biomedcentral.com/articles/10.1186/gb-2014-15-3-r46.
- Lu, J; Rincon, N; Wood, DE; Breitwieser, FP; Pockrandt, C; Langmead, B; et al. Metagenome analysis using the Kraken software suite. Nature Protocols 2022 17:12 [En ligne]; Nature Publishing Group, 28 Sep 2022; Volume 17, 12, pp. 2815–39. Available online: https://www.nature.com/articles/s41596-022-00738-y.
- Wood, DE; Lu, J; Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biol [En ligne]. BioMed Central Ltd. 2019, 20(1), 1–13. Available online: https://genomebiology.biomedcentral.com/articles/10.1186/s13059-019-1891-0.
- Wishart, DS; Han, S; Saha, S; Oler, E; Peters, H; Grant, JR; et al. PHASTEST: faster than PHASTER, better than PHAST. In Nucleic Acids Res [En ligne]; cité le; Oxford Academic, 5 Jul 2023; Volume 51, W1, pp. W443–50. [Google Scholar] [CrossRef]
- Bouras, G; Nepal, R; Houtak, G; Psaltis, AJ; Wormald, PJ; Vreugde, S. Pharokka: a fast scalable bacteriophage annotation tool. In Bioinformatics [En ligne]; cité le; Oxford Academic, 1 Jan 2023; Volume 39, 1. [Google Scholar] [CrossRef]
- Steinegger, M; Söding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. In Nature Biotechnology 2017; Nature Publishing Group, 16 Oct 2017; Volume 35, 11, pp. 1026–8. Available online: https://www.nature.com/articles/nbt.3988.
- Jordan, TC; Burnett, SH; Carson, S; Caruso, SM; Clase, K; DeJong, RJ; et al. A Broadly Implementable Research Course in Phage Discovery and Genomics for First-Year Undergraduate Students. mBio [En ligne] 2014, 5(1), e01051-13. Available online: https://pmc.ncbi.nlm.nih.gov/articles/PMC3950523/. [CrossRef] [PubMed]
- Fu, L; Niu, B; Zhu, Z; Wu, S; Fu; Niu, B; Zhu, Z; Wu, S. Bioinformatics WL, 2012 undefined. CD-HIT: accelerated for clustering the next-generation sequencing data. LiBioinformatics: 2012, 8 Nov 2025. Available online: https://academic.oup.com/bioinformatics/article-abstract/28/23/3150/192160.
- Li, W; Godzik, A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics [En ligne]. Bioinformatics 1 Jul 2006 [cité le 8 Nov 2025], 22(13), 1658–9. Available online: https://pubmed.ncbi.nlm.nih.gov/16731699/. [CrossRef] [PubMed]
- Wang, D; Liu, L; Xu, X; Wang, C; Wang, Y; Deng, Y; et al. Distributions, interactions, and dynamics of prokaryotes and phages in a hybrid biological wastewater treatment system. In Microbiome [En ligne; BioMed Central Ltd, 1 Dec 2024; Volume 12, 1, pp. 1–17. Available online: https://microbiomejournal.biomedcentral.com/articles/10.1186/s40168-024-01853-6.
- Flores, VS; Amgarten, DE; Iha, BKV; Ryon, KA; Danko, D; Tierney, BT; et al. Discovery and description of novel phage genomes from urban microbiomes sampled by the MetaSUB consortium. Scientific Reports 2024 14:1 [En ligne]. Nature Publishing Group 2024, 14(1), 1–14. Available online: https://www.nature.com/articles/s41598-024-58226-0.
| Metric | Value |
|---|---|
| Number of phage genomes | 17 |
| Total predicted proteins (CDS) | 1122 |
| Total gene families (pangenome size) | 1031 |
| Core gene families (present in 100% genomes) | 0 (0%) |
| Soft-core gene families (present in ≥95% genomes) | 0 (0%) |
| Shell gene families (present in 15–<95% genomes) | 13 (1.3%) |
| Cloud gene families (present in <15% genomes) | 1018 (98.7%) |
| Singleton gene families (unique to 1 genome) | 966 (93.7%) |
| Heaps’ law α (pangenome openness) | 0.026 (Open) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




