1. Introduction
TNs are commonly encountered, and their evaluation is predominantly performed using ultrasound (US), which provides a fast, non-invasive, and radiation-free imaging method. Due to its ability to generate real-time, high-resolution images of the thyroid gland, US has become the primary diagnostic tool for assessing thyroid abnormalities [
1,
2]. However, traditional CAD systems used by radiologists for diagnosing TN relied on manually extracted features, such as shape, texture, and margin characteristics. The process of manual feature extraction is labor-intensive and inefficient, and operator-dependent, often introducing variability and limiting large-scale applicability in clinical settings [
3]. Ongoing innovations in DL techniques have greatly enhanced CAD systems. However, when employing DL for TN diagnosis, several challenges arise. First, DL models typically require large labelled datasets for effective training, which are often scarce in the medical imaging domain due to privacy concerns, annotation costs, and limited expert availability. Additionally, training DL models on small datasets can lead to significant overfitting issues, potentially leading to poor generalization on unseen data. Their black-box nature also presents a significant interpretability challenge, making clinical adoption difficult, as clinicians often require transparent and explainable decision-making processes. Lastly, variability in US images, originating from differing hardware settings and user-specific techniques, operator techniques, and patient-specific factors, further complicates model robustness and generalization.
TL has emerged as a promising solution to address these limitations by leveraging pre-trained models, initially trained on large-scale, general-purpose datasets, and fine-tuning them on smaller, domain-specific datasets such as thyroid US images. This approach reduces the dependency on extensive labelled data, mitigates overfitting, and enhances model generalization across diverse imaging conditions. TL also offers the advantage of significantly reducing computational costs and training time, making it more practical for clinical deployment [
4].
This study involves training a DL models based on TL techniques from using a publicly available thyroid US image dataset. TL has gained significant traction in medical imaging thanks to its effectiveness in employing pre-trained features, such as ResNet, DenseNet, and EfficientNet, and adapt them for specific tasks with limited data availability [
5]. These pre-trained models, often trained on large-scale datasets like ImageNet, bring generalized knowledge that can be adapted to address particular medical imaging applications, such as TN classification [
6]. In this study, TL-based models are fine-tuned to classify TN as benign or malignant, based on their severity. TL models tend to converge faster during training and generalize better, owing to the prior knowledge encoded in their layers from large-scale pre-training. This comparison highlights the advantages of using pre-trained networks for medical image analysis, particularly in TN diagnosis, where accurate classification is crucial for proper patient management [
7]. Additionally, in the part of pre-processing, the authors focus on the use of second-generation wavelets, specifically BT (bandlet transform) , which are designed to generate decorrelated coefficients, eliminate redundancy, and preserve essential information. BT are based on geometric principles and are adept at capturing complex structures that are often not apparent when used WT. A key benefit of this geometric approach is its ability to model real-world data more effectively, particularly when the data is non-uniformly sampled, lies on curves, or exists in higher-dimensional spaces like surfaces and manifolds. This novel approach to geometric wavelets includes techniques such as edgelets, ridgelets, curvelets, bandelets, grouplets, shearlets, and wedgelets [
8,
9]. This work makes a unique contribution by integrating with TL. As far as we know, no prior research has explored this particular combination in the context of TC classification.
This work is characterized by the following novel contributions:
The study investigates the integration of TN with TL for enhancing medical image classification. In this framework, is employed to capture complex geometric patterns and spatial dependencies from US images, enabling the extraction of more robust and discriminative feature representations, which in turn improve the accuracy and reliability of TC classification.
TL leverages pre-trained models to improve generalization and reduce the dependency on large labeled datasets. Among the evaluated pre-trained models, the VGG19 architecture achieves the best performance across various assessment criteria, demonstrating its superior capability in extracting discriminative features for the target classification task.
To the best of our knowledge, no prior research has investigated the integration of with TL for TC classification or medical image classification, in contrast to existing studies that have primarily focused on WT and/or TL. This novel integration offers a distinctive and effective solution to address the challenges associated with TC detection in US images.
An overview of the paper’s organization is provided below: Section 2 examines prior research in the field that have utilized WT, , and TL, particularly in the context of TC diagnosis and classification.
Section 3 presents the background and preliminaries necessary to understand the proposed approach, including the theoretical foundations of WT, , and TL.
Section 4 describes the proposed methodology in detail, outlining the key steps of the algorithm.
Section 5 discusses the experimental results, performance evaluation, and comparative analysis. The paper concludes in Section 6, which reflects on the main contributions and explores prospective research paths.
2. Related Work
In recent years, significant efforts have been directed toward diagnosing benign and malignant TC. While many studies initially relied on the FNAB method, the increasing use of US imaging is driven by its accessibility and cost-effectiveness. A range of AI-based approaches have emerged to enhance diagnostic accuracy. The related literature is organized below according to the AI methodology employed.
Several studies have employed DL techniques for TC detection using imaging and clinical data. Vahdati et al. [
10] propose a DL-based method combining YOLOv5 for detection and XGBoost for classification using transverse and longitudinal US views. Similarly, Wang et al. [
11] introduce a model that integrates multimodal MRI data with clinical features to predict lymph node metastasis in papillary thyroid cancer. Gummalla et al. [
12] develop a hybrid framework using a sequential CNN and K-means clustering for classifying thyroid images. Chandana et al. [
13] present a deep CNN model for classifying adenoma, thyroiditis, and cancer based on CT and US scans. Wang et al. [
14] utilize a combination of BLR and CNN for metastasis prediction, integrating genetic mutations and clinical data. Qi et al. [
15] apply Mask R-CNN with ResNet-50 and FPN for detecting gross extrathyroidal extension in TC, outperforming radiologists in accuracy. Finally, Zhang et al. [
16] propose an automated DL-based system (AWT-AA) to differentiate benign and malignant nodules in US imaging, supported by logistic regression analysis.
Other studies utilize ensemble learning to improve model robustness and generalizability. Shah et al. [
17] design a deep ensemble model incorporating LSTM, GRU, and Bi-LSTM for mutation detection in thyroid adenocarcinoma, achieving high diagnostic accuracy from genomic data. Zhang et al. [
18] introduce MC-CNNs along with a weighted averaging ensemble and Faster Apriori for multi-view medical image classification and association rule mining. Zhang et al. [
19] develop a dynamic ensemble TL-based system that integrates multi-view ultrasonography data, featuring a novel weighting mechanism for optimal decision-making.
TL has been applied in several works to enhance generalization on medical datasets. Chen et al. [
20] utilize an improved GoogLeNet model with secondary transfer learning, total variation-based image restoration, and joint training on hospital and public datasets to classify TN. Ma et al. [
21] introduce Mul-DenseNet, a multi-channel DenseNet model pre-trained on ImageNet, to simultaneously segment thyroid and breast lesions in US images. Bakht et al. [
22] apply fine-tuned VGG-19 and AlexNet models with a weighted classification layer to cytology slides, enhancing performance despite class imbalance.
Hybrid and domain-specific architectures have also been explored. Lu et al. [
23] propose a dual-tree complex WT-based CNN for segmenting human thyroid OCT images. The model uses wavelet pooling to preserve texture details and resist noise, improving segmentation robustness. Wang et al. [
24] present SL-FCN with soft labeling, enabling more accurate boundary delineation in TC segmentation tasks compared to hard-label models.
Table 1 provides a comparative summary of the reviewed studies, detailing the techniques used, datasets, performance metrics, and limitations. It also highlights existing research gaps, contextualizing the need for the proposed approach.
3. Preliminaries
3.1. Wavelet Transform
The WT is a powerful mathematical tool used in image processing to analyze images at multiple resolutions. Unlike the Fourier transform, which provides only frequency information, the WT captures both spatial and frequency characteristics, making it highly effective for feature extraction. The WT is based on the concept of analyzing a signal or an image using scaled and shifted versions of a finite-duration function called the mother wavelet. This function is designed to be localized in both time and frequency domains, making wavelets suitable for capturing transient, localized, and multi-scale features within signals and images. The fundamental principle of WT relies on the decomposition of a given function into a set of basis functions derived from a mother wavelet through scaling and translation operations. Mathematically, a wavelet function
satisfies the admissibility condition, which ensures that it has zero mean and is well-localized:
The wavelet basis functions are constructed by modifying the scale and position of the mother wavelet as follows:
where
a is the scaling parameter that controls the frequency resolution and
b is the translation parameter that shifts the wavelet function in time. The continuous WT is given by:
where
represents the wavelet coefficients at different scales and positions. The DWT, which is a computationally efficient variant of continuous WT, employs dyadic scales (
) and integer translations (
) to construct the wavelet basis. This results in an orthogonal or biorthogonal representation that allows hierarchical decomposition of images into different frequency subbands. Using the DWT, an image is decomposed into four distinct frequency subbands: LL, LH, HL, and HH. The LL subband retains the approximation information, primarily reflecting the image’s coarse details, while LH, HL, and HH contain detailed coefficients corresponding to horizontal, vertical, and diagonal details, respectively [
26,
27].
This process is mathematically represented by the system of equations in (
4), where
denotes the original image. The two-dimensional DWT is carried out by first applying the one-dimensional DWT along the rows, followed by its application along the columns.
Where is the scaling function, which generates the LL subband (approximation coefficients), capturing the coarse image details, is the wavelet function, which generates the LH, HL, and HH subbands (detailed coefficients). This decomposition provides a multi-resolution representation of the image, allowing for efficient image processing techniques such as feature extraction.
3.2. Bandlet Transform
The , introduced by Le Pennec and Mallat [
28], is designed to construct a basis that aligns with the geometric structure of an image by locally deforming the spatial domain. This deformation simplifies the structure into a separable basis along a fixed direction—either horizontal or vertical. A key aspect of this transform is the
flow-curve relationship, where the flow in the vertical direction corresponds to curves with non-vertical tangents. This allows for the construction of test bandelets that respect the geometric regularity of each sub-block. Mathematically, the basis function is defined as:
where
defines the curve, and
denotes its slope, interpreted as the optical flow.
The is applied depending on the presence of geometric flow: if a sub-block exhibits no significant geometric variation, it is treated as uniformly regular and processed using a classical separable wavelet basis. Conversely, if geometric variations are detected, bandelet processing is employed. In cases involving singularities, additional Lagrangian-based computations are required.
The cost function governing this transformation is given by:
where f is the original image, is its reconstructed approximation, denotes the number of bits used to encode the geometric flow (optical flow) in sub-block j, and corresponds to the bits used to encode the quantized bandelet coefficients. The parameter is a Lagrangian multiplier that balances rate and distortion, and T is the quantization step.
To efficiently represent the image, a quadtree decomposition is employed. This technique recursively divides the image domain into four quadrants (sub-blocks), denoted as
and
, yielding a hierarchical representation. For each block
S, the goal is to choose the best representation strategy—either by encoding
S as a whole or by subdividing it further. This decision is governed by minimizing the Lagrangian cost:
Here,
represents the cost of encoding the block
S directly without further subdivision, while
is the cumulative cost of encoding its four children:
The term
accounts for the overhead cost of subdivision. This recursive strategy ensures that each sub-block is processed in the most efficient manner, adapting the representation complexity to local image content. To address curved singularities in image structures, a deformation operator is applied to locally realign blocks so that anisotropic features are better captured along either horizontal or vertical directions. This leads to a new orthonormal basis in
, replacing the standard horizontal wavelets
with geometry-adapted functions defined as:
Here,
and
represent the horizontal and vertical spatial coordinates of the image domain, respectively. The function
defines a local geometric flow or deformation that aligns the vertical coordinate
with directional image features such as edges. By warping the coordinate system through
, the basis functions adapt to the underlying structure of the image, improving alignment with anisotropic features. This transformation process, known as
bandeletization, builds an orthonormal basis that is aligned with the image’s geometric structure. By doing so, it replaces traditional wavelet bases with more expressive functions, allowing for enhanced compression and analysis by better respecting geometric regularities in both horizontal and vertical orientations.
The theoretical foundation of relies on their ability to optimize representation in anisotropic geometric image structures, unlike traditional WT that only capture local oscillations. This geometric adaptability enables improved feature extraction, making bandelets particularly effective for DL applications where efficient hierarchical feature representation is crucial [
13]. The final output of the consists of bandelet coefficients that capture the image’s directional and structural information. These coefficients, derived from optimal sub-block partitioning and adaptive transformations, form the bandelet feature vector, which is fed into a DL model. The bandelet feature vector includes multi-scale directional energy distributions, geometric flow descriptors, and localized structural patterns, providing a compact yet expressive representation of the image. These bandelet features are then input into CNNs, or transformer-based models, depending on the application. The hierarchical nature of bandelet-based feature extraction ensures that DL models receive a structurally enriched representation of the data, significantly improving their performance in image classification, segmentation, super-resolution, and medical image analysis. By leveraging the ’s ability to adapt to image geometry, DL architectures can process images more efficiently, achieving higher accuracy while reducing computational overhead [
9,
29].
3.3. Inductive TL
Inductive TL is a machine learning paradigm where a model trained on a source task is adapted to a target task, assuming that both tasks share some structural similarities while differing in their label spaces or distributions. Mathematically, given a labeled source dataset
associated with a task
and a labeled target dataset
associated with a task
, the goal is to learn a target function
by leveraging knowledge from a source function
, where
but
, meaning that while both tasks share a common feature space, they exhibit differences in their distributions or label mappings. The learning process consists of two key steps: (i) pre-training and (ii) fine-tuning. During pre-training, a model
is trained on the source dataset to minimize the loss function:
Where,
refers to the loss function appropriate to the task at hand, typically cross-entropy for classification or mean squared error for regression. The learned parameters
serve as the initialization for the target model
, which is then fine-tuned on the target dataset using
Typically employing strategies such as feature extraction, where early layers of the pre-trained model are frozen while only task-specific layers are updated, or full fine-tuning, where all model parameters are updated but with a lower learning rate to preserve generalizable knowledge. Optimization is commonly performed using gradient descent with an update rule
, where
is the learning rate, ensuring stable adaptation to the new task. To balance knowledge retention and task-specific adaptation, a weighted loss function
can be employed, where
controls the contribution of the source knowledge during training. Inductive TL is widely applied in DL, particularly in computer vision, where CNNs such as ResNet, VGG, or EfficientNet pre-trained on ImageNet are fine-tuned for specialized applications like medical image analysis, object detection, or satellite imagery classification, and in natural language processing [
30].
4. Methodology
This section evaluates the performance of both WT and approaches in enhancing image classification and feature extraction accuracy within the proposed framework. The algorithm is specifically designed to differentiate between malignant and benign TN, aiming to improve diagnostic precision by leveraging geometry-adaptive representations.
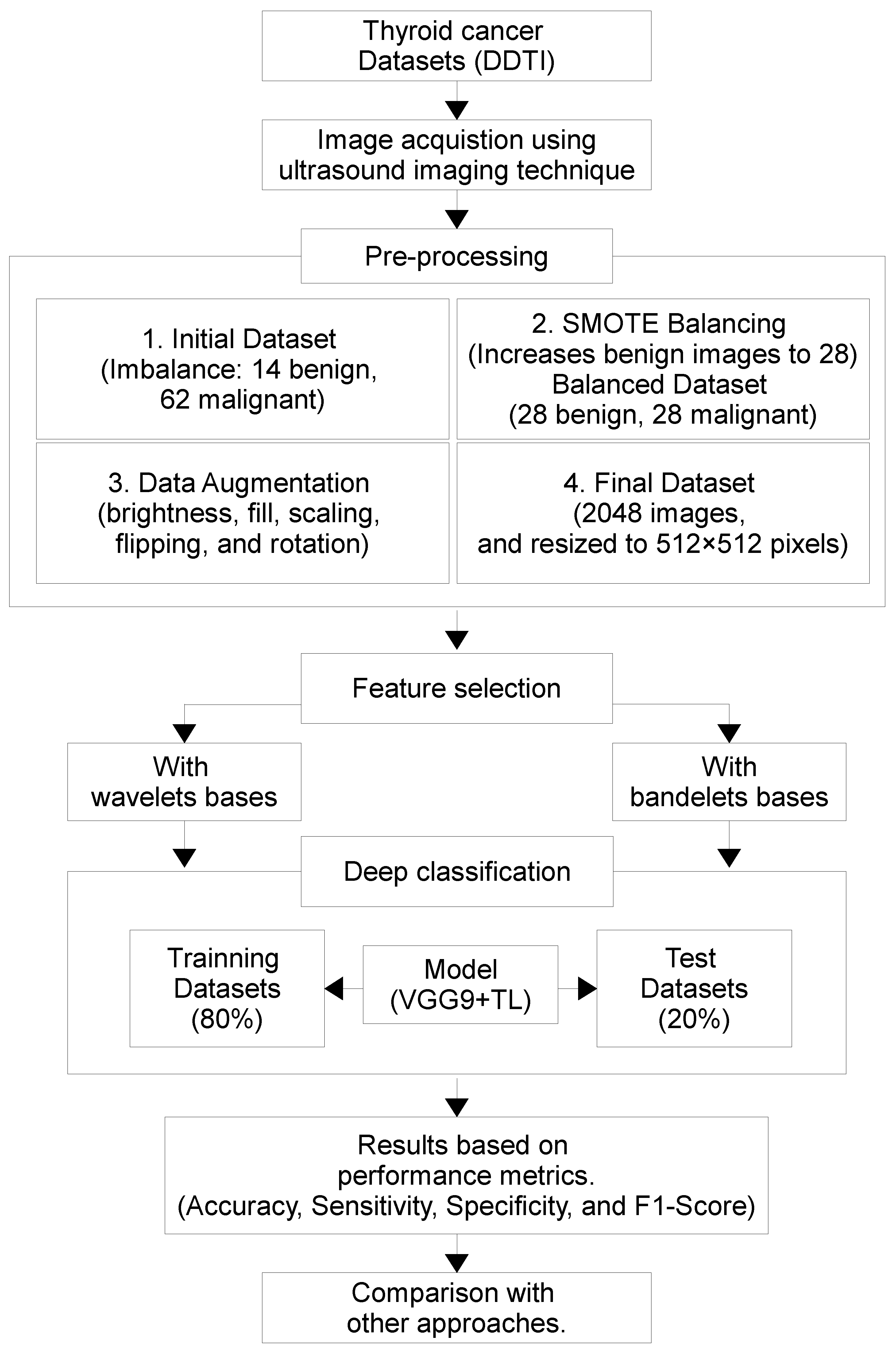
The overall procedural workflow is illustrated in
Figure 1. The process begins with the DDTI dataset, where US images are acquired and pre-processed. Due to class imbalance, SMOTE is applied to augment benign cases and create a balanced dataset, followed by additional DA techniques to further enhance data diversity. Feature selection is then performed using both WT and bases, allowing extraction of rich structural and texture descriptors. These features are subsequently fed into a deep learning classifier using TL from the pre-trained VGG19 model. The dataset is split into training and testing subsets, and the classification model is trained accordingly. Performance is evaluated using key metrics and compared against alternative methods to assess the effectiveness and efficiency of the proposed approach. Algorithm 1 outlines the suggested TN classification algorithm based on DL techniques.
|
Algorithm 1 TN Classification Algorithm |
- 1:
function
LoadDatasets
- 2:
Dataset ← DDTI ▹ 134 images: 14 benign, 62 malignant - 3:
return Dataset - 4:
end function - 5:
function Preprocess(Dataset) - 6:
Benign ← GetBenignImages(Dataset) ▹ 14 images - 7:
Malignant ← GetMalignantImages(Dataset) ▹ 62 images - 8:
New_Benign ← SMOTE(Benign, target = 28) ▹ Oversample benign - 9:
Balanced_Dataset ← Combine(New_Benign, Malignant[0:28]) - 10:
Augmented_Dataset ← Augment(Balanced_Dataset, Techniques = {Brightness, Flip, Rotate, Resize_to_512x512}) - 11:
return Augmented_Dataset ▹ Target: 2048 images - 12:
end function - 13:
function ExtractFeatures(Augmented_Dataset) - 14:
Features ← [] - 15:
for each image in Augmented_Dataset do
- 16:
Bandelet_Features ← ApplyBandeletTransform(image) - 17:
Features ← Add(Bandelet_Features) - 18:
end for
- 19:
return Features - 20:
end function - 21:
function Classify(Augmented_Dataset, Features) - 22:
Train_Data ← Take80Percent(Augmented_Dataset) ▹ 1638 images - 23:
Val_Data ← Take20Percent(Augmented_Dataset) ▹ 410 images - 24:
Models ← {VGG16} ▹ Simplified list - 25:
for each model in Models do
- 26:
LoadPretrained(model) - 27:
FineTune(model, Train_Data, Features) - 28:
Predictions ← Test(model, Val_Data) - 29:
Save(Predictions) - 30:
end for
- 31:
return Predictions - 32:
end function - 33:
function Evaluate(Predictions, Val_Data) - 34:
for each model in Predictions do
- 35:
Accuracy ← CalculateAccuracy(Predictions, Val_Data) - 36:
Sensitivity ← CalculateSensitivity(Predictions, Val_Data) - 37:
Display(model, Accuracy, Sensitivity) - 38:
end for
- 39:
end function - 40:
function
Main
- 41:
Dataset ← LoadDatasets() - 42:
Augmented_Dataset ← Preprocess(Dataset) - 43:
Features ← ExtractFeatures(Augmented_Dataset) - 44:
Predictions ← Classify(Augmented_Dataset, Features) - 45:
Evaluate(Predictions, Augmented_Dataset) - 46:
end function |
4.1. Input TC Datasets
The DDTI dataset [
31], provided by the Universidad Nacional de Colombia and the Instituto de Diagnóstico Médico (IDIME), is an open-access collection of thyroid US images.
Table 2 summarizes its key characteristics. This dataset was selected as the primary source for this study due to its credibility, public availability, and relevance to TN classification tasks. Previously used in related research, such as in [
32,
33], it serves as a reliable benchmark for performance comparison. Its open-access nature addresses the common challenge of restricted medical datasets, which often require complex ethical approvals. Additionally, the dataset is well-annotated, with clear labels distinguishing between benign and malignant cases, ensuring suitability for supervised learning. Although limited in size, the DDTI dataset remains valuable given the scarcity of publicly available, high-quality thyroid US datasets.
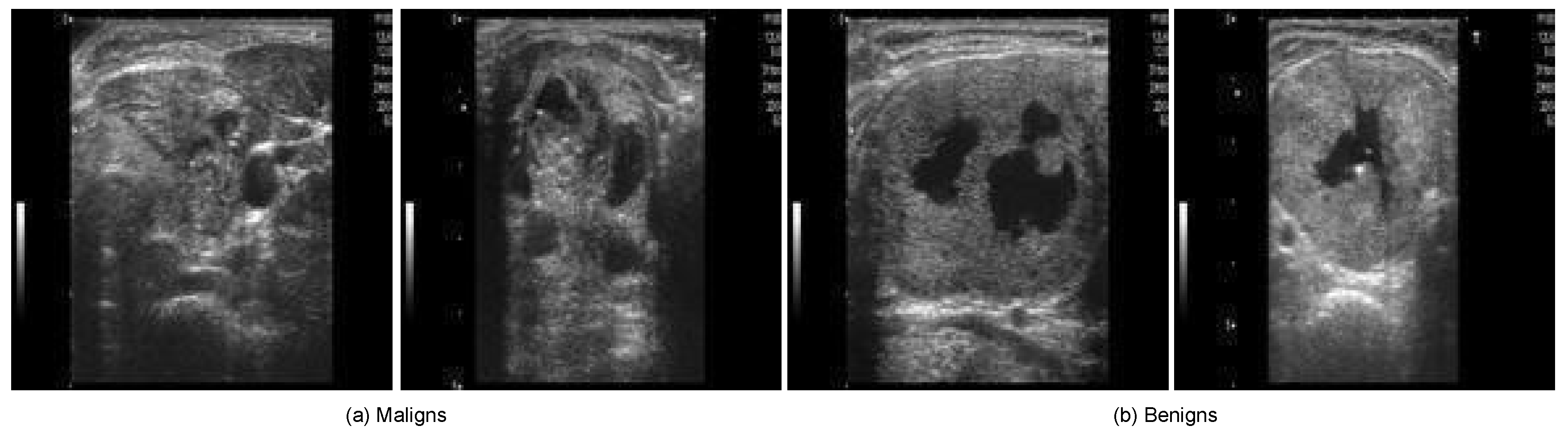
Figure 2 illustrates sample thyroid US images from the DDTI dataset, showcasing both malignant and benign cases.
4.2. Preprocessing
To tackle dataset imbalance in TC classification, a two-step approach was applied: synthetic oversampling followed by DA [
34,
35]. Initially, the dataset comprised 14 benign and 62 malignant images, resulting in a significant class imbalance that could bias model predictions. To mitigate this, the SMOTE was employed to generate additional synthetic benign images by interpolating between existing samples, increasing their count from 14 to 28. This adjustment ensured parity with the 28 malignant images, creating a more balanced dataset for training. Following dataset balancing, various DA techniques were implemented to further expand the dataset to 2048 images, introducing variability to enhance model generalization and robustness. The augmentation process included brightness adjustments to simulate diverse lighting conditions, ensuring the model’s adaptability to varying illumination levels. Nearest-neighbor fill was applied to preserve pixel integrity during transformations, preventing unwanted artifacts. Height scaling was introduced to modify the vertical proportions of images, while horizontal flipping effectively doubled spatial variations. Rotation was employed to alter image orientation, preventing model bias toward specific angles. Finally, a fixed input size of 512×512 pixels was used by resizing all images accordingly, improving consistency and resolution for feature extraction. This preprocessing not only mitigated class imbalance but also enriched dataset diversity, reducing overfitting risks and enhancing the model’s ability to accurately classify benign and malignant cases. By integrating SMOTE with targeted DA, the dataset became more representative and robust, ultimately leading to improved generalization and performance in TC classification.
4.3. Bandelet Feature Selection
adapts to the image’s intrinsic geometry, allowing for a more efficient encoding of directional features and leading to enhanced feature selection. Empirical studies demonstrate that the classification performance of TC images improves significantly when using TL-extracted features compared to WT [
8,
36,
37].
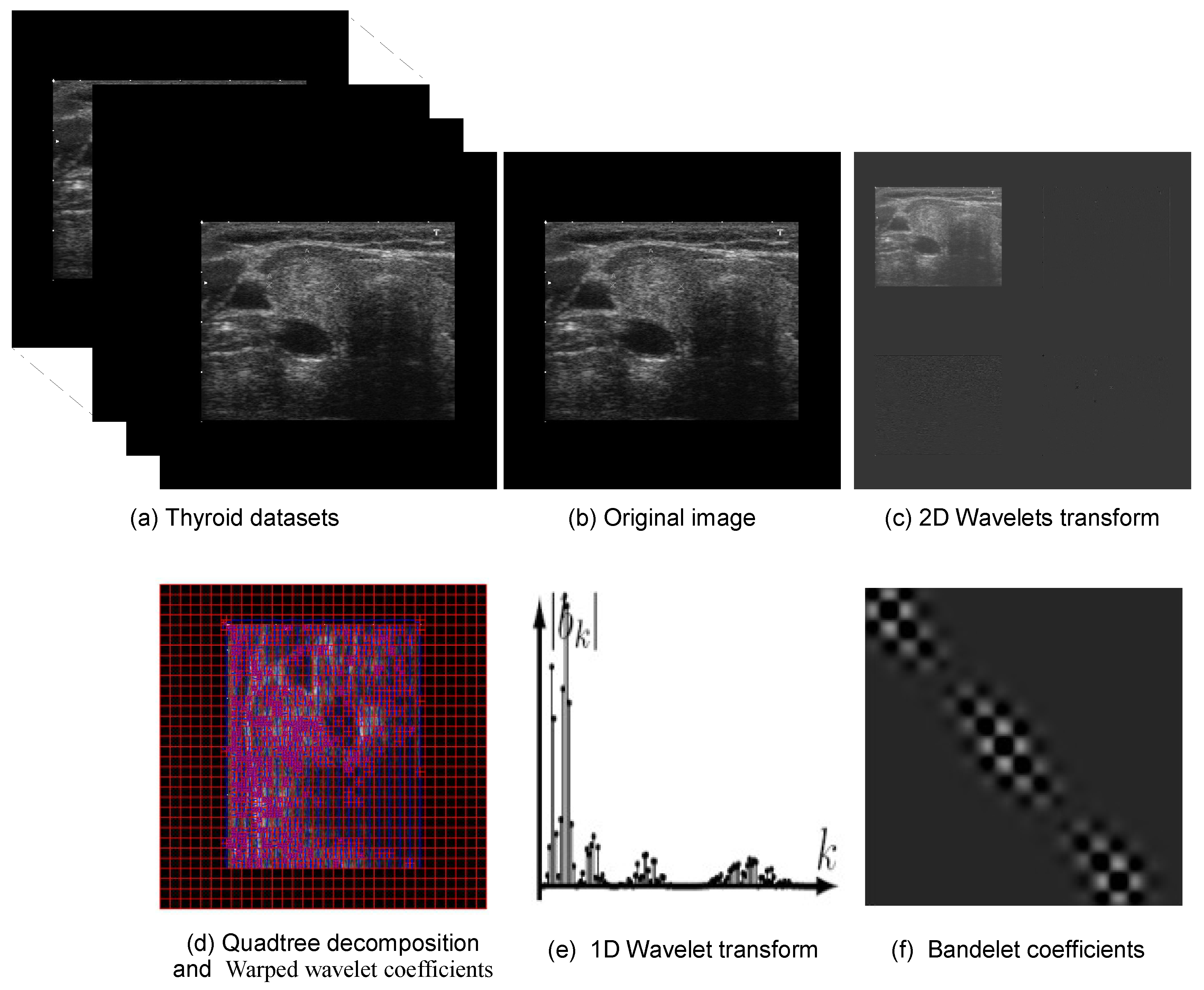
Figure 3 illustrates the sequential steps of the applied to thyroid US images for enhanced feature extraction. The process begins with thyroid nodule images sourced from the DDTI dataset, comprising both benign and malignant cases. Initially, each input image undergoes a classical 2D wavelet transform to decompose it into frequency subbands; however, this transform alone does not account for the geometric orientation of structures within the image. To address this, the image is then recursively partitioned through a quadtree decomposition, segmenting it into smaller blocks based on local homogeneity. For each block, a deformation field—representing the geometric flow—is estimated to align features such as edges or curves. The wavelet coefficients are subsequently warped along this estimated flow, allowing the basis functions to conform to the image’s directional patterns. A 1D wavelet transform is then applied along the deformation direction to further enhance the structural encoding. The final output is a set of bandelet coefficients that capture directional, geometric, and textural information in a sparse and compact representation. These coefficients serve as discriminative features for downstream classification tasks, offering superior structural fidelity compared to traditional wavelet features, particularly in the context of complex and anisotropic patterns found in thyroid nodules.
4.4. Deep Classification
Medical image classification benefits considerably from the adoption of the proposed strategy, where acquiring large, well-annotated datasets is time-consuming and resource-intensive [
30,
38]. Typically, a network trained on a large dataset like ImageNet is either fine-tuned by adjusting its layers or used as a fixed feature extractor, transferring learned representations to the new task. To optimize training and evaluation, the dataset was carefully partitioned following DL best practices, with 80% (1638 samples) allocated for training and 20% (410 samples) designated for validation. The training subset was essential for refining model parameters, improving the ability to extract meaningful feature representations, while the validation subset played a crucial role in ensuring generalization and detecting overfitting by evaluating the model’s performance on unseen data.
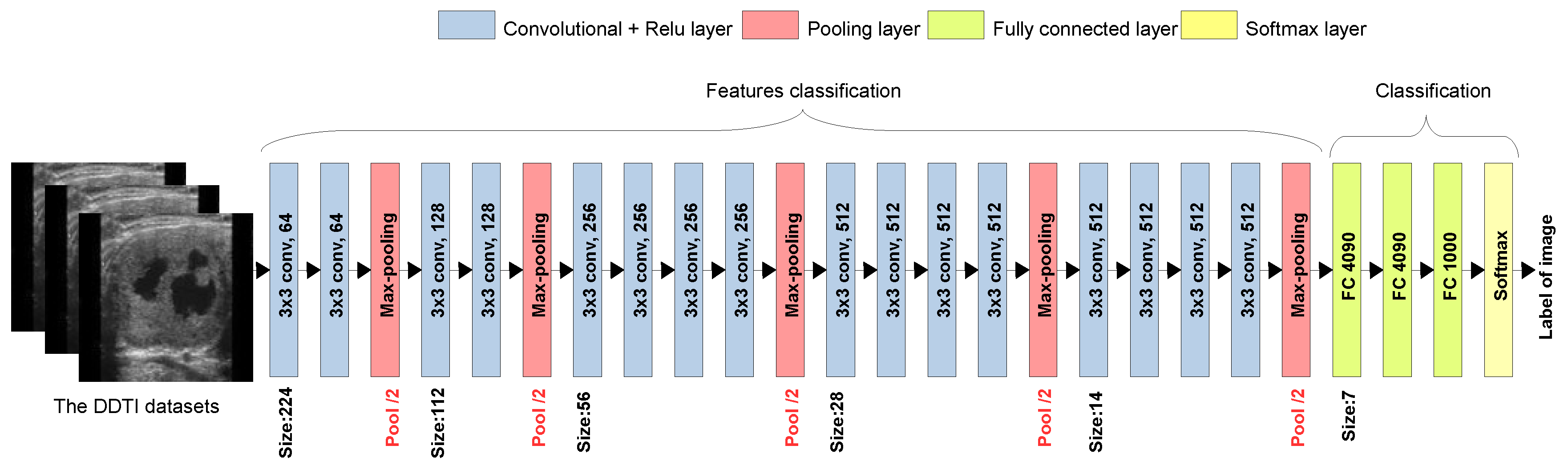
The VGG19 architecture, illustrated in
Figure 4, is employed in this study as a pre-trained model for TL. It consists of a structured DL model used for feature extraction and classification. It begins with an input image (from the DDTI dataset) and processes it through multiple convolutional layers with 3×3 filters and ReLU activation, which capture hierarchical features such as edges, textures, and patterns. After every two or four convolutional layers, a max pooling layer (red) is applied to reduce spatial dimensions while preserving essential features. The model progressively increases the number of filters from 64 to 512, allowing deeper layers to extract more complex representations. Once feature extraction is complete, the fully connected layers flatten the extracted features and pass them through two dense layers with 4096 neurons, followed by a Softmax layer that classifies the image into 1000 possible categories. In TL, the convolutional base is typically frozen, and the fully connected layers are modified or fine-tuned for new classification tasks, making VGG19 highly effective for our calassification of TC images [
39,
40]. By leveraging TL, these models could retain valuable pre-learned representations, minimizing the data and computational resources required for training, a crucial advantage in medical imaging applications where data scarcity is a challenge. In TL, modifying layers between a pre-trained model and a proposed model involves carefully adapting the network architecture to balance feature reuse and task-specific learning. First, the pre-trained model, typically a deep neural network VGG, is loaded, and its architecture is examined. The earlier convolutional layers embedding layers in transformers, which capture fundamental and transferable features, are usually frozen to retain their learned representations. The fully connected (dense) layers or task-specific heads in natural language processing models, which encode high-level, domain-specific features, are removed or replaced with new layers customized for the target task. This replacement often involves adding new dense layers, batch normalization, dropout (to prevent overfitting), incorporating an output layer that uses task-specific activation functions, like softmax for multi-class classification and sigmoid for binary cases. Fine-tuning can be applied to some middle layers if the new dataset is sufficiently large, gradually unfreezing layers while using a lower learning rate to prevent catastrophic forgetting of previously learned features. Additionally, TL methods such as feature extraction (where only new layers are trained) or full fine-tuning (where pre-trained weights are adjusted) are selected based on dataset size, computational power, and the similarity between the source and target domains. The modified model is then compiled and trained, leveraging techniques like learning rate scheduling and data augmentation to improve adaptation while ensuring that the knowledge from the pre-trained model enhances the performance of the new task.
The details of VGG19 layers are provided in
Table 4 as an example.
4.5. Performance Metrics
Common metrics used for evaluating ultrasound-based TC classification. These metrics offer a thorough assessment of the model’s performance in differentiating between benign and malignant thyroid nodules. The
Accuracy (Acc) measures the overall correctness of predictions. As a harmonic mean of precision and recall, the F1-score serves as a comprehensive metric for evaluating classification performance, especially in imbalanced datasets:
Sensitivity (Sen) quantifies the proportion of actual malignant cases correctly identified,
Specificity (Spe) measures the proportion of benign cases correctly classified, and
Precision (P) indicates the ratio of predicted malignancies that are confirmed as actual malignant instances:
These metrics collectively ensure a reliable and multidimensional assessment of classification performance. The abbreviations TP, TN, FP, and FN refer to true positives, true negatives, false positives, and false negatives in that order.
5. Results and Discussion
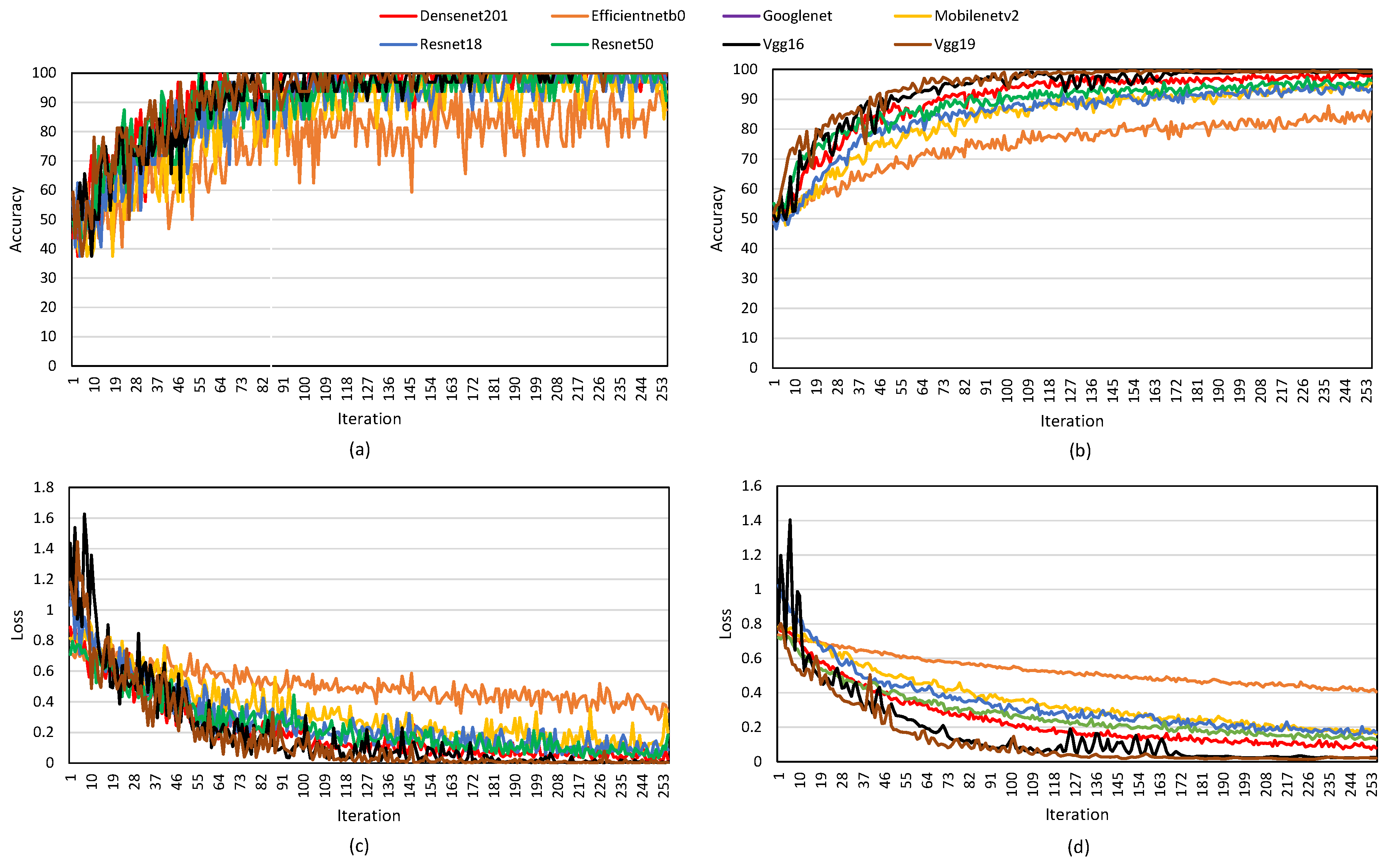
The obtained training and validation accuracy, along with their corresponding losses for different DL models, are illustrated in
Figure 5. To justify the adoption of VGG19 in this study, multiple pretrained models were assessed and compared. The obtained results demonstrate that VGG19 consistently outperforms other models in TC classification using the BT+TL approach. The training accuracy curve (
Figure 5 (a)) shows that VGG19, ResNet50, and DenseNet201 achieve rapid convergence, surpassing 90% accuracy early in the training process, while MobileNetV2, EfficientNetB0, and GoogLeNet exhibit slower improvements and lower final accuracy values. Similarly, the validation accuracy graph (
Figure 5 (b)) confirms that VGG19 maintains the highest accuracy, followed by ResNet50 and DenseNet201, indicating strong generalization to unseen data. In contrast, MobileNetV2 and GoogLeNet display lower validation accuracy, suggesting difficulties in capturing complex patterns in TC images. The training loss curve (
Figure 5 (c)) illustrates a steady decrease across all models, with VGG19 and ResNet50 achieving the lowest final loss values, reflecting their ability to minimize classification errors efficiently. However, GoogLeNet and MobileNetV2 maintain relatively higher loss values, implying weaker learning performance. The validation loss curve (
Figure 5 (d)) further supports these findings, as VGG19, ResNet50, and DenseNet201 exhibit smooth, consistently decreasing validation loss, whereas MobileNetV2 and GoogLeNet show more fluctuations, suggesting overfitting or instability during validation. These results highlight the superior performance of VGG19, which not only achieves the highest accuracy but also demonstrates better stability, faster convergence, and lower loss values, making it the most effective model for TC classification in this study.
In
Table 5, the performance of the WT and is compared under varying quadtree decomposition thresholds (
T) across four evaluation metrics: accuracy, sensitivity, specificity, and F1 score. The WT serves as a strong baseline, achieving an accuracy of 0.9430, sensitivity of 0.9302, specificity of 0.9433, and an F1 score of 0.9584. When the is applied, performance improves at lower thresholds (
), with enhanced accuracy (0.9511), sensitivity (0.9622), specificity (0.9774), and F1 score (0.9807) compared to the WT. As the threshold increases to
, both accuracy (0.9635) and F1 score (0.9645) continue to rise; however, sensitivity decreases to 0.9108, indicating a reduced ability to identify true positives. The optimal performance for the is observed at
, yielding the highest accuracy (0.9891), sensitivity (0.9811), and F1 score (0.9889), while maintaining a strong specificity of 0.9731.
In
Table 6, the performance of the proposed model is compared with several previous approaches. The model introduced by Bakht et al. [
22] reports a lower accuracy of 0.9305 and sensitivity of 0.929, with an F1 score of 0.928, indicating relatively weaker performance. The approach by Ma et al. [
21] achieves an accuracy of 0.9257 and a high sensitivity of 0.9869, but lacks specificity and only reports an F1 score of 0.9596. Vasile et al. [
41] presents a good specificity of 0.9843 and an accuracy of 0.9735; however, the F1 score is not provided. The proposed approach +TL (VGG19) model, achieves the highest performance, with an accuracy of 0.9891, sensitivity of 0.9811, specificity of 0.9731, and an F1 score of 0.9889. In summary, the proposed BT+TL (VGG19) model outperforms all other methods, particularly in terms of accuracy, sensitivity, and F1 score, highlighting its superiority in classification tasks.
6. Conclusion
In this paper, we introduced a novel thyroid nodule TN classification framework that tightly integrates geometric feature encoding via the Bandelet Transform with Transfer Learning (TL) on a pre-trained VGG19 backbone. The core technical innovation lies in employing a quadtree-driven BT to locally adapt basis functions to the intrinsic flow of nodule contours, yielding a sparse set of directional coefficients that encode anisotropic texture and edge continuity more faithfully than standard wavelets. By systematically varying the quadtree subdivision threshold, we demonstrated that finer-scale geometric adaptivity optimises the trade-off between coefficient sparsity and reconstruction fidelity, directly translating into a marked improvement in classification metrics (98.91% accuracy, 98.11% sensitivity, 97.31% specificity, 98.89% F1-score).
From a methodological standpoint, our ablation study confirms three key findings:
Geometric versus separable bases: Replacing classical DWT with BT yields a consistent 2–4% gain across all metrics, underscoring the value of geometry-aware multiscale analysis in ultrasound images.
Quadtree threshold optimisation: Intermediate thresholds strike the best balance, whereas overly coarse (T < 10) or overly fine (T > 50) partitions either underfit large-scale structures or overfit noise, respectively.
Transfer Learning strategy: Leveraging VGG19’s mid-level feature maps, frozen through the initial training epochs and then selectively unfrozen, accelerates convergence, mitigates overfitting on the limited DDTI dataset, and reduces training time by approximately 40% compared to end-to-end learning.
Our contributions extend beyond mere performance gains. To the best of our knowledge, this is the first study to fuse a bandelet-based preprocessing pipeline with TL for thyroid cancer detection. This fusion not only enhances geometric feature fidelity but also streamlines the integration with deep convolutional architectures, offering a generalizable blueprint for other anisotropic tissue classification tasks (e.g., liver lesion or breast mass detection). Moreover, the quadtree-Bandelet module itself is lightweight—requiring only O(N Log N) operations for an N-pixel image—and can be plugged into any CNN or transformer-based classifier with minimal modification.
Looking forward, several avenues warrant exploration. First, extending the BT stage to incorporate adaptive Lagrangian rate-distortion optimisation could further refine coefficient selection under constrained bit budgets, facilitating on-device inference for portable ultrasound systems. Second, integrating vision Transformers as the downstream classifier may capitalise on the global attention mechanism to exploit further the long-range dependencies encoded by bandelet features. Finally, exploring semi-supervised or self-supervised pre-training on large, unlabeled ultrasound repositories—augmented by synthetic data generated via generative adversarial networks—could alleviate the label scarcity that continues to challenge medical imaging applications.
In summary, the proposed BT+TL framework not only achieves state-of-the-art classification accuracy on the DDTI dataset but also establishes a novel paradigm for coupling geometry-adaptive transforms with modern deep learning, charting a clear path toward more interpretable, efficient, and broadly applicable medical image analysis pipelines.
Author Contributions
Conceptualization, Y.H., H.K., M.C.G. and J.H.; Methodology, Y.H. and H.K.; Data curation, H.K.; Resources, Y.H. and H.K.; Investigation, Y.H., H.K., M.C.G. and J.H.; Visualization, H.K., M.C.G. and J.H.; Writing—original draft, Y.H., H.K., M.C.G. and J.H.; Writing—review and editing, Y.H., H.K., M.C.G. and J.H.
Funding
The APC and Open Access fees for this work are funded by the University of Liverpool.
Institutional Review Board Statement
This research was deemed not to require ethical approval.
Informed Consent Statement
Not Applicable.
Data Availability Statement
Data will be made available on request.
Conflicts of Interest
The authors declare that they have no known competing interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- Cao, C.L.; Li, Q.L.; Tong, J.; Shi, L.N.; Li, W.X.; Xu, Y.; Cheng, J.; Du, T.T.; Li, J.; Cui, X.W. Artificial intelligence in thyroid ultrasound. Frontiers in Oncology 2023, 13, 1060702. [Google Scholar] [CrossRef]
- Habchi, Y.; Himeur, Y.; Kheddar, H.; Boukabou, A.; Atalla, S.; Chouchane, A.; Ouamane, A.; Mansoor, W. AI in thyroid cancer diagnosis: Techniques, trends, and future directions. Systems 2023, 11, 519. [Google Scholar] [CrossRef]
- Yu, X.; Wang, H.; Ma, L. Detection of thyroid nodules with ultrasound images based on deep learning. Current Medical Imaging 2020, 16, 174–180. [Google Scholar] [CrossRef]
- Mazari, A.C.; Kheddar, H. Deep learning-and transfer learning-based models for covid-19 detection using radiography images. In Proceedings of the 2023 International Conference on Advances in Electronics, Control and Communication Systems (ICAECCS), 2023; IEEE; pp. 1–4. [Google Scholar]
- Habchi, Y.; Kheddar, H.; Himeur, Y.; Ghanem, M.C. Machine learning and transformers for thyroid carcinoma diagnosis. Journal of Visual Communication and Image Representation 2026, 115, 104668. [Google Scholar] [CrossRef]
- Pavithra, S.; Vanithamani, R.; Judith, J. Classification of stages of thyroid nodules in ultrasound images using transfer learning methods. In Proceedings of the Proc. 2nd Int. Conf. Image Process. Capsule Netw. (ICIPCN 2021), 2022; Springer; pp. 241–253. [Google Scholar]
- Sureshkumar, V.; Jaganathan, D.; Ravi, V.; Velleangiri, V.; Ravi, P. A Comparative Study on Thyroid Nodule Classification using Transfer Learning Methods. The Open Bioinformatics Journal 2024, 17. [Google Scholar] [CrossRef]
- Fang, Y.; Liu, J.; Li, J.; Cheng, J.; Hu, J.; Yi, D.; Xiao, X.; Bhatti, U.A. Robust zero-watermarking algorithm for medical images based on SIFT and Bandelet-DCT. Multimedia Tools and Applications 2022, 81, 16863–16879. [Google Scholar] [CrossRef]
- Beladgham, M.; Habchi, Y.; Taleb-Ahmed, A.; et al. Medical video compression using bandelet based on lifting scheme and SPIHT coding: In search of high visual quality. Informatics in Medicine Unlocked 2019, 17, 100244. [Google Scholar] [CrossRef]
- Vahdati, S.; Khosravi, B.; Robinson, K.A.; Rouzrokh, P.; Moassefi, M.; Akkus, Z.; Erickson, B.J. A multi-view deep learning model for thyroid nodules detection and characterization in ultrasound imaging. Bioengineering 2024, 11, 648. [Google Scholar] [CrossRef]
- Wang, X.; Zhang, H.; Fan, H.; Yang, X.; Fan, J.; Wu, P.; Ni, Y.; Hu, S. Multimodal MRI Deep Learning for Predicting Central Lymph Node Metastasis in Papillary Thyroid Cancer. Cancers 2024, 16, 4042. [Google Scholar] [CrossRef]
- Gummalla, K.; Ganesan, S.; Pokhrel, S.; Somasiri, N. Enhanced early detection of thyroid abnormalities using a hybrid deep learning model. Journal of Innovative Image Processing 2024, 6, 244–261. [Google Scholar] [CrossRef]
- Chandana, K.H.; Prasan, U. Thyroid disease detection using CNN techniques. THYROID 2023, 55. [Google Scholar]
- Wang, Z.; Qu, L.; Chen, Q.; Zhou, Y.; Duan, H.; Li, B.; Weng, Y.; Su, J.; Yi, W. Deep learning-based multifeature integration robustly predicts central lymph node metastasis in papillary thyroid cancer. BMC cancer 2023, 23, 128. [Google Scholar] [CrossRef] [PubMed]
- Qi, Q.; Huang, X.; Zhang, Y.; Cai, S.; Liu, Z.; Qiu, T.; Cui, Z.; Zhou, A.; Yuan, X.; Zhu, W.; et al. Ultrasound image-based deep learning to assist in diagnosing gross extrathyroidal extension thyroid cancer: a retrospective multicenter study. EClinicalMedicine 2023, 58. [Google Scholar] [CrossRef]
- Zhang, F.; Sun, Y.; Wu, X.; Meng, C.; Xiang, M.; Huang, T.; Duan, W.; Wang, F.; Sun, Z. Analysis of the application value of ultrasound imaging diagnosis in the clinical staging of thyroid cancer. Journal of Oncology 2022, 2022, 8030262. [Google Scholar] [CrossRef]
- Shah, A.A.; Daud, A.; Bukhari, A.; Alshemaimri, B.; Ahsan, M.; Younis, R. DEL-Thyroid: deep ensemble learning framework for detection of thyroid cancer progression through genomic mutation. BMC Medical Informatics and Decision Making 2024, 24, 198. [Google Scholar] [CrossRef]
- Zhang, X.; Lee, V.C. Deep Learning Empowered Decision Support Systems for Thyroid Cancer Detection and Management. Procedia Computer Science 2024, 237, 945–954. [Google Scholar] [CrossRef]
- Zhang, X.; Liu, F.; Lee, V.C.S.; Jassal, K.; Di Muzio, B.; Lee, J.C. SSRN 4944552; Dynamic Ensemble Transfer Learning with Multi-View Ultrasonography for Improving Thyroid Cancer Diagnostic Reliability. Available at.
- Chen, W.; Gu, Z.; Liu, Z.; Fu, Y.; Ye, Z.; Zhang, X.; Xiao, L. A new classification method in ultrasound images of benign and malignant thyroid nodules based on transfer learning and deep convolutional neural network. Complexity 2021, 2021, 6296811. [Google Scholar] [CrossRef]
- Ma, J.; Bao, L.; Lou, Q.; Kong, D. Transfer learning for automatic joint segmentation of thyroid and breast lesions from ultrasound images. International journal of computer assisted radiology and surgery 2022, 17, 363–372. [Google Scholar] [CrossRef]
- A. B. Bakht, S.J.; Dina, R.; Almarzouqi, H.; Khandoker, A.; Werghi, N. Thyroid nodule cell classification in cytology images using transfer learning approach. In Proceedings of the Proc. 12th Int. Conf. Soft Comput. Pattern Recognit. (SoCPaR 2020), 2021; Springer; pp. 539–549. [Google Scholar]
- Lu, H.; Wang, H.; Zhang, Q.; Won, D.; Yoon, S.W. A dual-tree complex wavelet transform based convolutional neural network for human thyroid medical image segmentation. In Proceedings of the 2018 IEEE international conference on healthcare informatics (ICHI), 2018; IEEE; pp. 191–198. [Google Scholar]
- Wang, C.W.; Lin, K.Y.; Lin, Y.J.; Khalil, M.A.; Chu, K.L.; Chao, T.K. A soft label deep learning to assist breast cancer target therapy and thyroid cancer diagnosis. Cancers 2022, 14, 5312. [Google Scholar] [CrossRef]
- Sharma, R.; Mahanti, G.K.; Chakraborty, C.; Panda, G.; Rath, A. An IoT and deep learning-based smart healthcare framework for thyroid cancer detection. ACM Transactions on Internet Technology; 2023. [Google Scholar]
- Chao, Z.; Duan, X.; Jia, S.; Guo, X.; Liu, H.; Jia, F. Medical image fusion via discrete stationary wavelet transform and an enhanced radial basis function neural network. Applied Soft Computing 2022, 118, 108542. [Google Scholar] [CrossRef]
- Habchi, Y.; Beladgham, M.; Taleb-Ahmed, A. RGB Medical Video Compression Using Geometric Wavelet and SPIHT Coding. International Journal of Electrical and Computer Engineering 2016, 6, 1627–1636. [Google Scholar] [CrossRef]
- Le Pennec, E.; Mallat, S. Sparse geometric image representations with bandelets. IEEE transactions on image processing 2005, 14, 423–438. [Google Scholar] [CrossRef]
- Deo, B.S.; Pal, M.; Panigrahi, P.K.; Pradhan, A. An ensemble deep learning model with empirical wavelet transform feature for oral cancer histopathological image classification. International Journal of Data Science and Analytics 2024, 1–18. [Google Scholar] [CrossRef]
- Habchi, Y.; Kheddar, H.; Himeur, Y.; Boukabou, A.; Atalla, S.; Mansoor, W.; Al-Ahmad, H. Deep transfer learning for kidney cancer diagnosis. arXiv 2024. arXiv:2408.04318. [CrossRef]
- DDTI. Available online: http://cimalab.unal.edu.co/?lang=en&mod=project&id=31 (accessed on 2023-03-01).
- Li, X.; Fu, C.; Xu, S.; Sham, C.W. Thyroid Ultrasound Image Database and Marker Mask Inpainting Method for Research and Development. Ultrasound in Medicine & Biology 2024, 50, 509–519. [Google Scholar] [CrossRef]
- Wang, R.; Zhou, H.; Fu, P.; Shen, H.; Bai, Y. A multiscale attentional unet model for automatic segmentation in medical ultrasound images. Ultrasonic Imaging 2023, 45, 159–174. [Google Scholar] [CrossRef]
- Goodman, J.; Sarkani, S.; Mazzuchi, T. Distance-based probabilistic data augmentation for synthetic minority oversampling. ACM/IMS Transactions on Data Science (TDS) 2022, 2, 1–18. [Google Scholar] [CrossRef]
- Khan, A.A.; Chaudhari, O.; Chandra, R. A review of ensemble learning and data augmentation models for class imbalanced problems: Combination, implementation and evaluation. Expert Systems with Applications 2024, 244, 122778. [Google Scholar] [CrossRef]
- Wang, X.; Chen, W.; Gao, J.; Wang, C. Hybrid image denoising method based on non-subsampled contourlet transform and bandelet transform. IET Image Processing 2018, 12, 778–784. [Google Scholar] [CrossRef]
- Raja, S. Secured medical image compression using DES encryption technique in Bandelet multiscale transform. International Journal of Wavelets, Multiresolution and Information Processing 2018, 16, 1850028. [Google Scholar] [CrossRef]
- Kheddar, H.; Himeur, Y.; Al-Maadeed, S.; Amira, A.; Bensaali, F. Deep transfer learning for automatic speech recognition: Towards better generalization. Knowledge-Based Systems 2023, 277, 110851. [Google Scholar] [CrossRef]
- Alzubaidi, L.; Fadhel, M.A.; Al-Shamma, O.; Zhang, J.; Santamaría, J.; Duan, Y.; Oleiwi, R.S. Towards a better understanding of transfer learning for medical imaging: a case study. Applied Sciences 2020, 10, 4523. [Google Scholar] [CrossRef]
- Salehi, A.W.; Khan, S.; Gupta, G.; Alabduallah, B.I.; Almjally, A.; Alsolai, H.; Siddiqui, T.; Mellit, A. A study of CNN and transfer learning in medical imaging: Advantages, challenges, future scope. Sustainability 2023, 15, 5930. [Google Scholar] [CrossRef]
- Vasile, C.M.; Udrîştoiu, A.L.; Ghenea, A.E.; Popescu, M.; Gheonea, C.; Niculescu, C.E.; Ungureanu, A.M.; Udrîştoiu, Ş.; Drocaş, A.I.; Gruionu, L.G. Intelligent diagnosis of thyroid ultrasound imaging using an ensemble of deep learning methods. Medicina 2021, 57, 395. [Google Scholar] [CrossRef] [PubMed]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).