Submitted:
15 December 2025
Posted:
18 December 2025
You are already at the latest version
Abstract
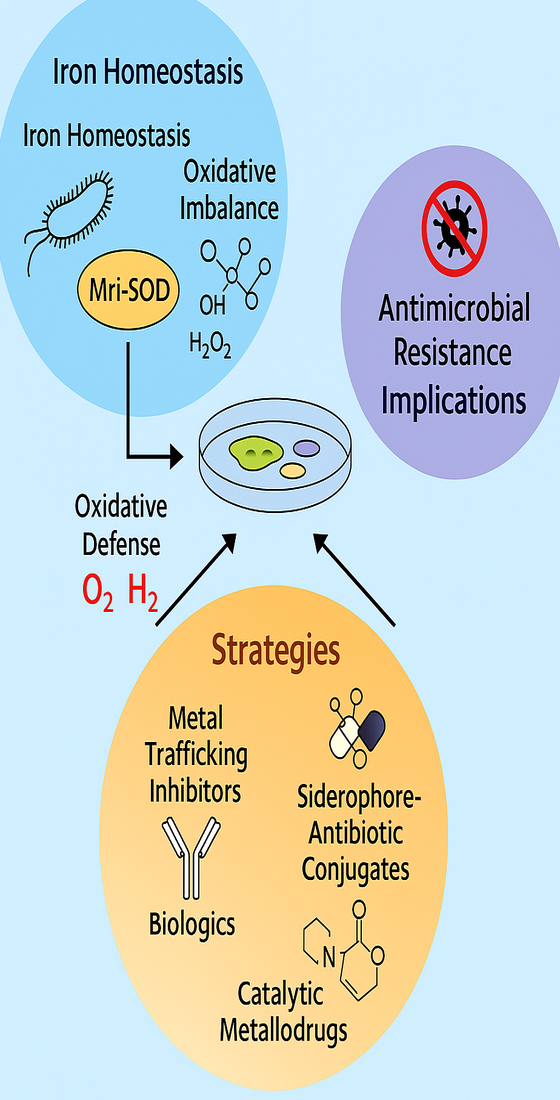
Keywords:
1. Introduction
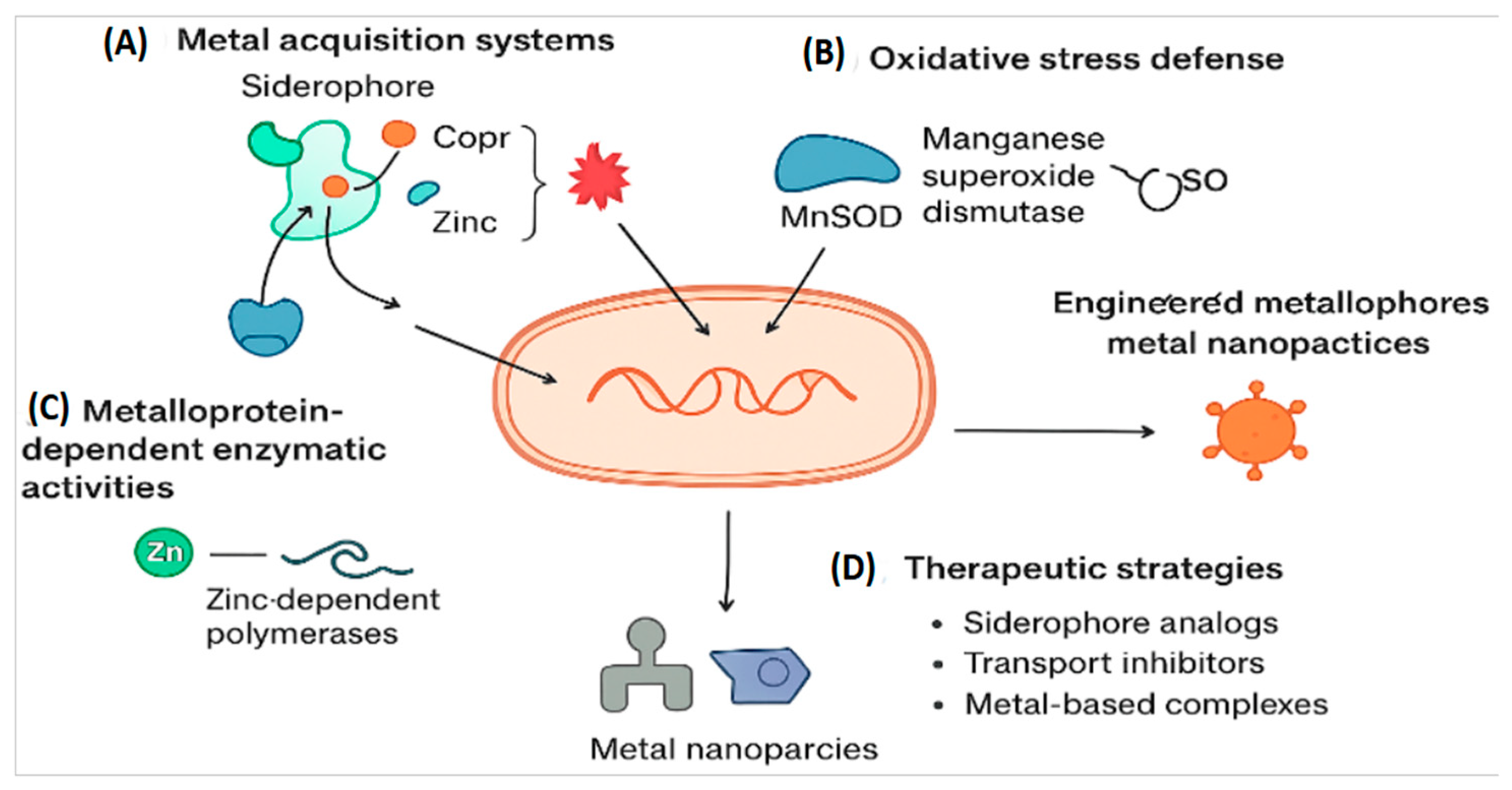
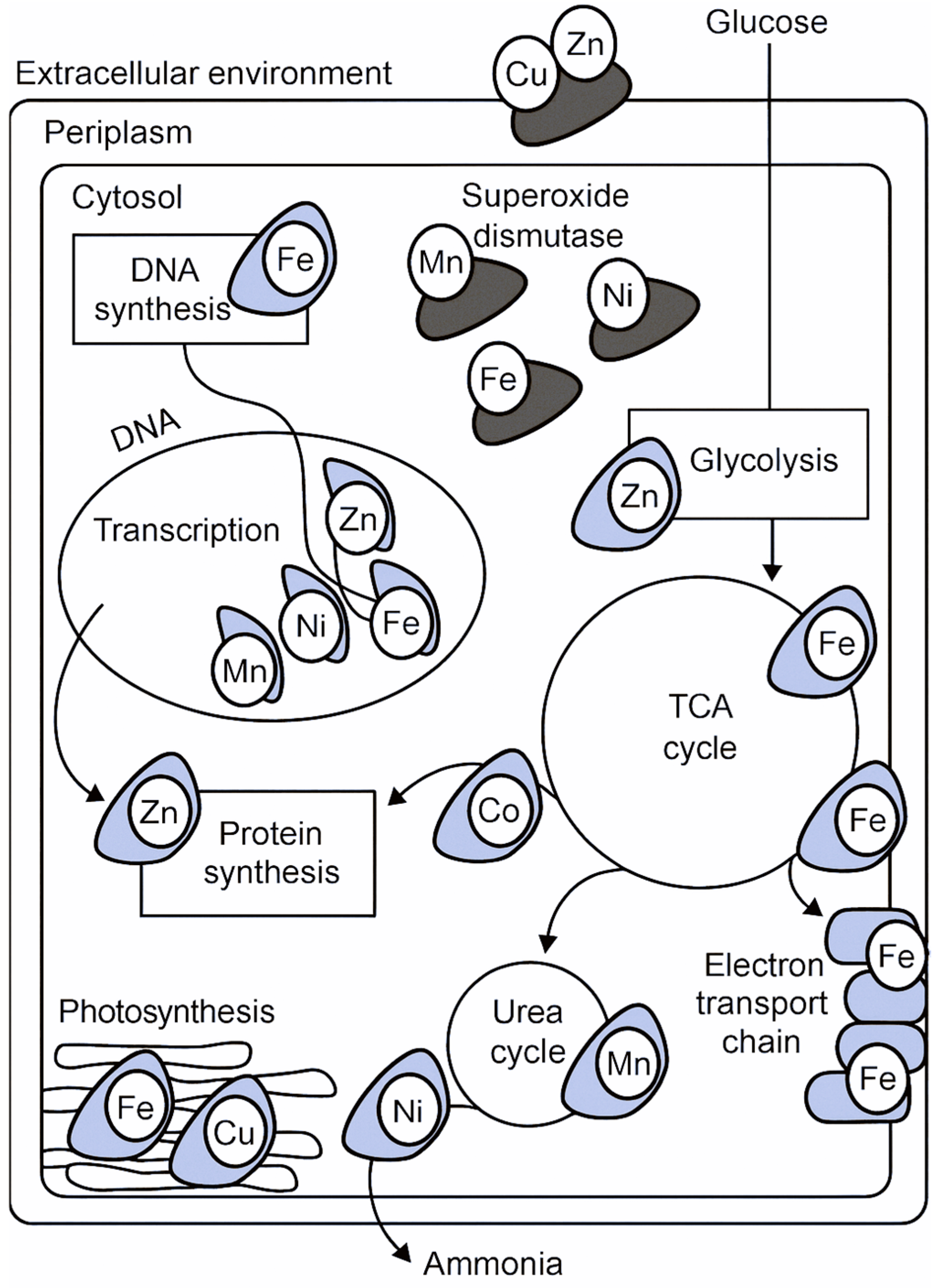
1.1. Mechanisms of Metalloprotein Targeting in Bacterial Pathogenesis
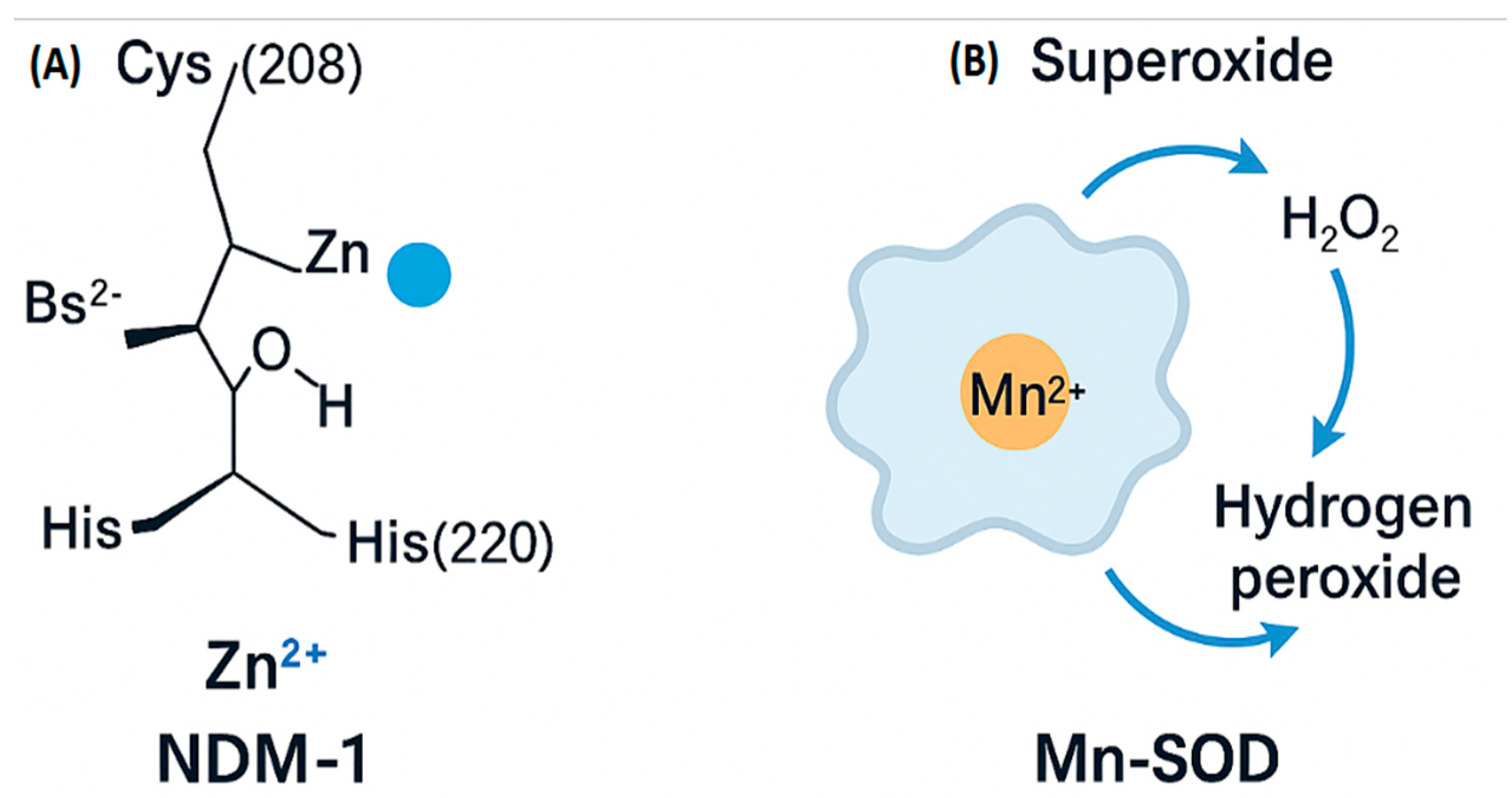
1.1.1. Oxidative Stress Response:
1.1.2. Enzymatic Activity:
1.1.3. Metal Acquisition Systems:
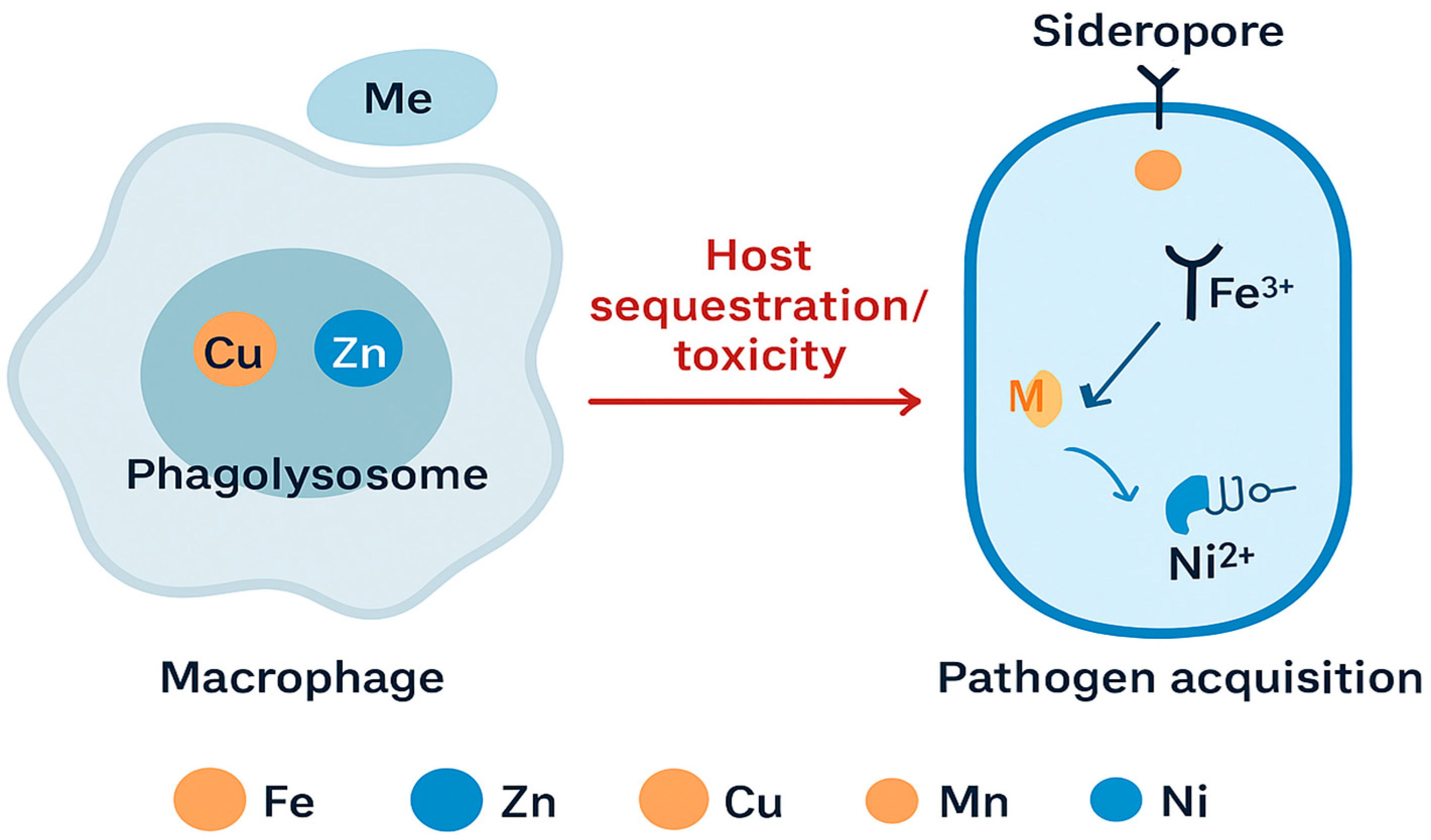
1.1.4. Host–Pathogen Metal Competition:
1.2. Functional and Therapeutic Implications
1.3. From Mechanisms to Strategies: Integrating Metalloprotein Insights into Antibacterial Development
2. Targeting Metal Homeostasis: Experimental Insights
2.1. Mechanisms of Action
2.2. Importance in Bacterial Survival
2.3. Potential for Drug Development
- Extracellular disruption: Charged species and ionic metals destabilize the bacterial envelope and proton motive force.
- Intracellular targeting: Metal pharmacophores penetrate cells and undergo organometallic transformations triggered by bacterial reductants.
3. Innovative Approaches to Target Metalloproteins: Toward Mechanism-Informed Antibacterial Design.
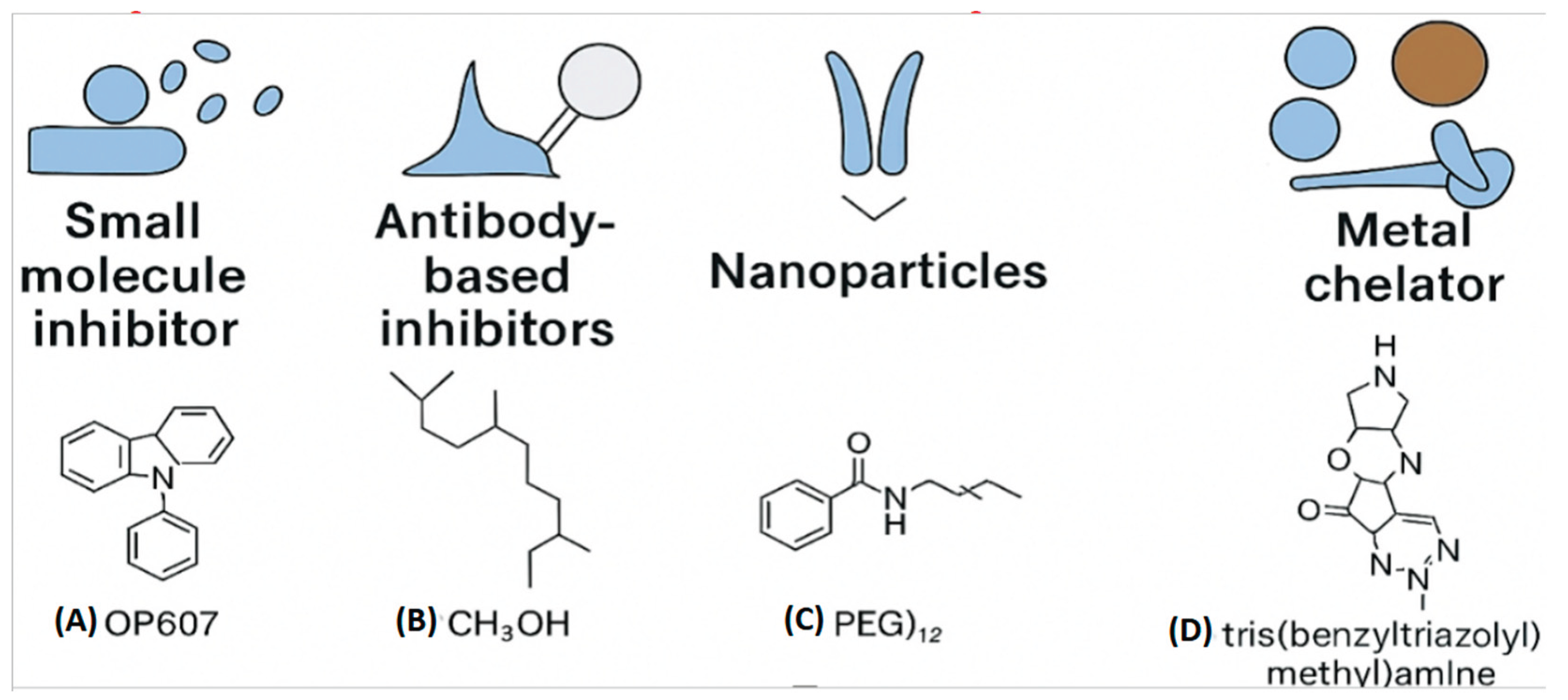
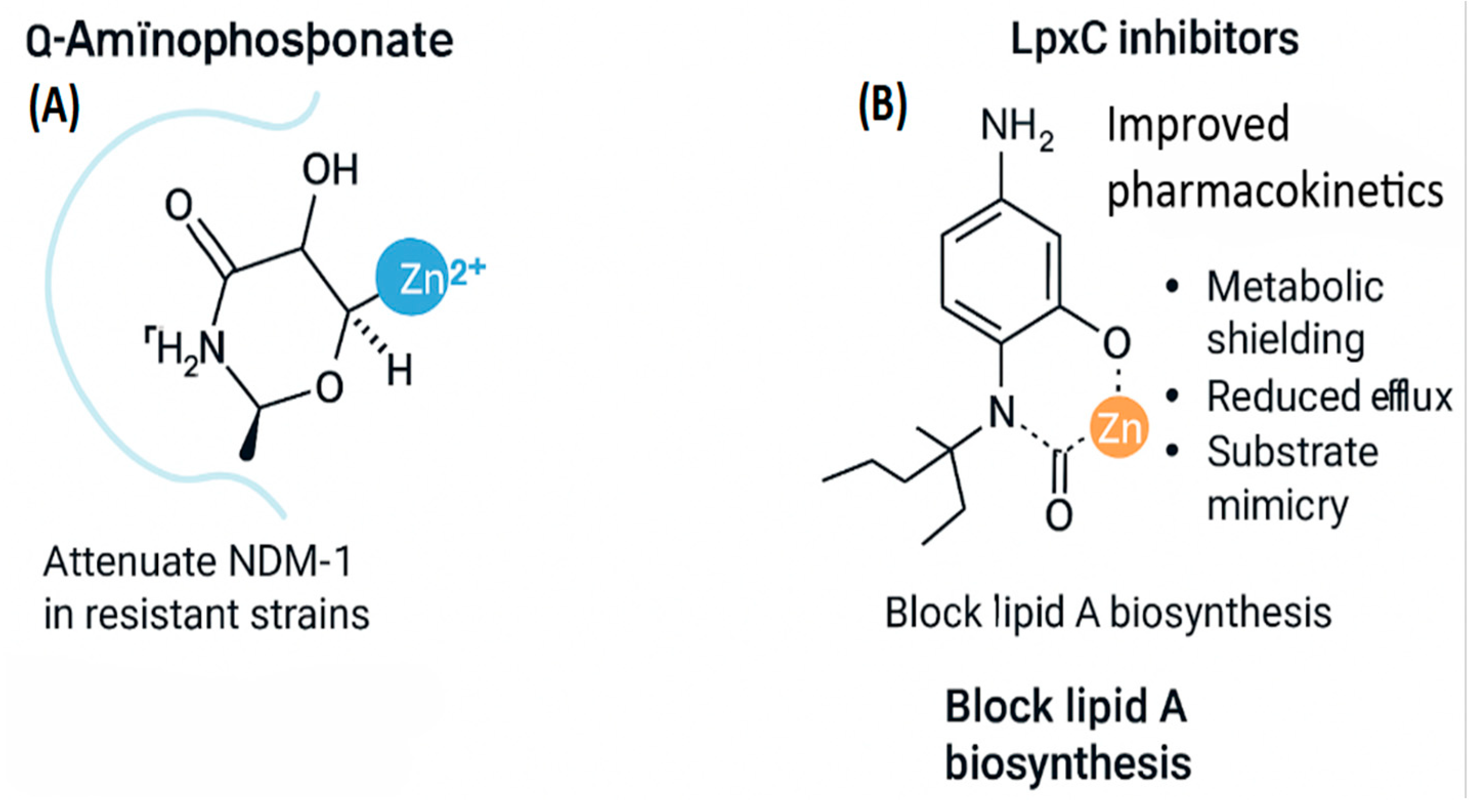
3.1. Small Molecule Inhibitors: Beyond Chelation
- A benzene ring substituted with amino (–NH₂) and hydroxyl (–OH) groups to enhance solubility and hydrogen bonding.
- A central amine linker that positions the phosphonate for optimal interaction.
- A bidentate phosphonic acid group (–PO(OH)₂) that stabilizes the inhibitor–enzyme complex without permanently displacing Zn²⁺.
- Steric shielding of the hydroxamate group, preventing metabolic degradation.
- Polarity tuning, which reduces susceptibility to efflux via RND transporters.
- Scaffold rigidification, enhancing binding specificity and minimizing off-target interactions.
3.2. Monoclonal Antibodies and Biologics: Precision Targeting
- Limited tissue penetration arises from large molecular size (~150 kDa for IgG) and poor diffusion across epithelial barriers. Glycosylation patterns and charge distribution further restrict movement through dense extracellular matrices.
- Binding-site barriers result in peripheral sequestration, where high-affinity binding near vasculature prevents deeper tissue access.
- Immunogenicity risks are elevated in chronic infections, where repeated exposure to recombinant proteins may trigger host immune responses.
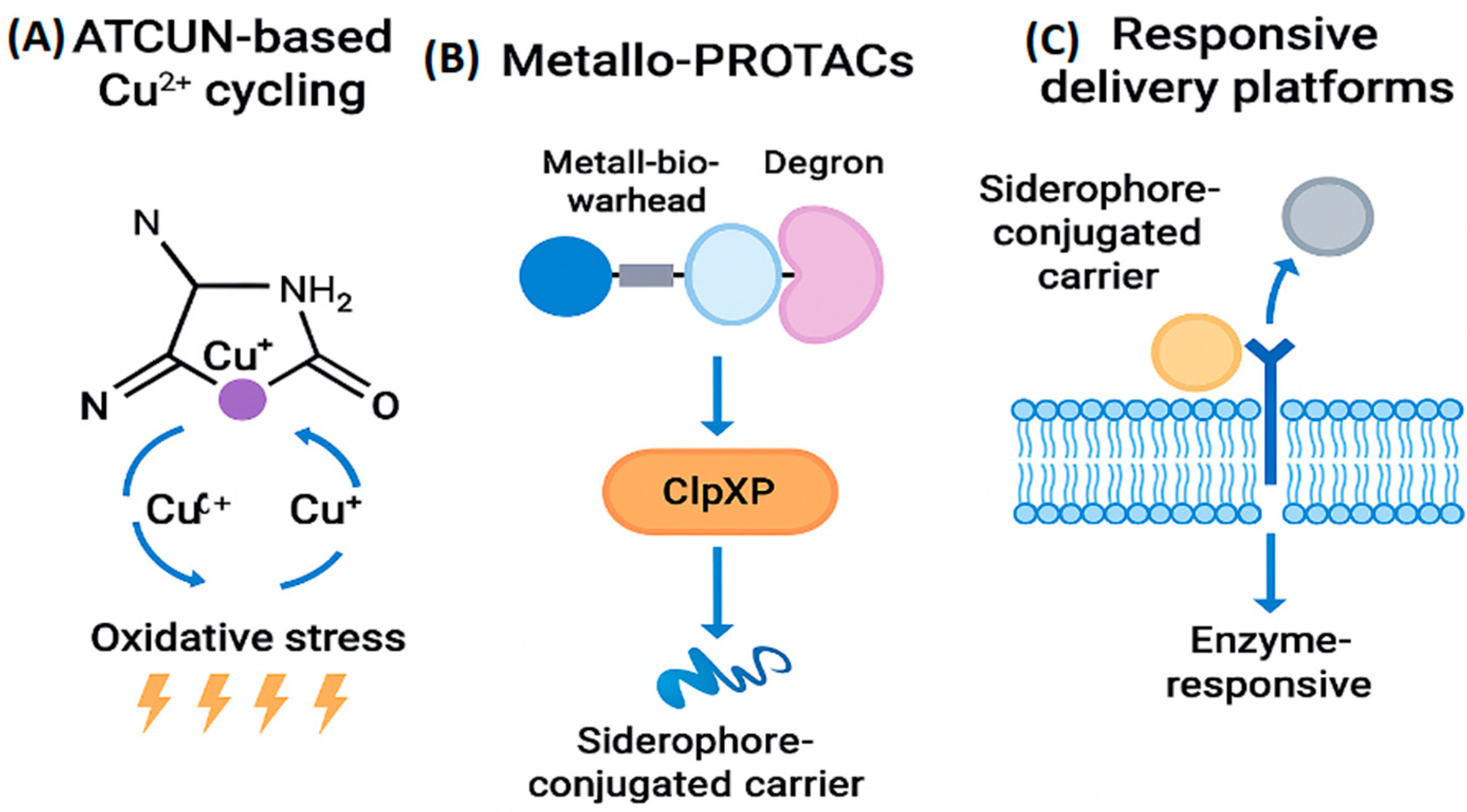
3.3. Catalytic Metallodrugs and Metallo-PROTACs: Mechanistic Innovation
- A metal-binding warhead that engages the target metalloprotein.
- A linker domain optimized for bacterial permeability and stability.
- A degron motif recognized by bacterial proteases, triggering selective degradation.
3.4. Future Directions and Expert Perspective
4. Critical Appraisal and Innovation Pathways
4.1. Isoform Selectivity and Structural Precision
4.2. Delivery Platform Optimization
4.3. Multi-Target Resistance Mitigation
4.4. Interdisciplinary Integration for Clinical Translation
-
Mapping Metalloprotein Networks:Conduct metagenomic profiling across diverse pathogens to delineate conserved and strain-specific vulnerabilities[53]. This includes mining environmental and clinical datasets to identify underexplored metalloprotein families.
-
Designing Hybrid Therapeutics:Develop dual-function agents that inhibit microbial growth while attenuating virulence. Examples include molecules that combine enzymatic inhibition with immunomodulatory or quorum-sensing disruption effects [54].
-
Refining Host–Pathogen Metal Dynamics:Investigate metal competition at infection sites to define therapeutic windows and minimize collateral effects on host metalloproteins. Leveraging host metal sequestration mechanisms may offer adjunctive therapeutic benefits [55].
-
Standardizing In Vivo Models:Create infection models that accurately recapitulate microenvironments such as biofilms, abscesses, and intracellular niches. These should incorporate immune modulation, metal availability, and pharmacokinetic parameters to predict clinical outcomes [56]
5. Case Studies of Metalloprotein-Targeted Antibacterial Strategies
6. Strategic Challenges and Translational Pathways in Metalloprotein Targeting
7. Concluding Perspectives
- Genomic profiling to identify pathogen-specific vulnerabilities.
- Structure-guided drug design to create non-chelating inhibitors and selective biologics.
- Responsive delivery systems tailored to infection microenvironments.
- Multi-target strategies that disrupt metal trafficking and virulence simultaneously.
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest Statement
References
- F., J.; Staats, C. C.; Pontes, M. H. Editorial: Metal Homeostasis in Microbial Physiology and Virulence. Front. Cell. Infect. Microbiol. 2023, 13, 1183137. [Google Scholar] [CrossRef]
- Begg, S. L. The Role of Metal Ions in the Virulence and Viability of Bacterial Pathogens. Biochem. Soc. Trans. 2019, 47(1), 77–87. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.; Koh, J.; Cha, S.; Roe, J. Activation of Zinc Uptake Regulator by Zinc Binding to Three Regulatory Sites. Nucleic Acids Res. 2024, 52(8), 4185–4197. [Google Scholar] [CrossRef]
- Chandrangsu, P.; Rensing, C.; Helmann, J. D. Metal Homeostasis and Resistance in Bacteria. Nat. Rev. Microbiol. 2017, 15(6), 338–350. [Google Scholar] [CrossRef]
- Karnwal, A.; Jassim, A. Y.; Mohammed, A. A.; Mohammad Said, A. R.; Selvaraj, M.; Malik, T. Addressing the global challenge of bacterial drug resistance: Insights, strategies, and future directions. Front. Microbiol. 2025, 16, 1517772. [Google Scholar] [CrossRef]
- Vitali, V.; Zineddu, S.; Messori, L. Metal Compounds as Antimicrobial Agents: Smart Approaches for Discovering New Effective Treatments. RSC Adv. 2025, 15, 748. [Google Scholar] [CrossRef] [PubMed]
- Simmons, K. J.; Chopra, I.; Fishwick, C. W. G. Structure-Based Discovery of Antibacterial Drugs. Nat. Rev. Microbiol. 2010, 8(4), 278–289. [Google Scholar] [CrossRef]
- Kehl-Fie, T. E.; Skaar, E. P. Nutritional Immunity beyond Iron: A Role for Manganese and Zinc. Curr. Opin. Chem. Biol. 2010, 14(2), 218–224. [Google Scholar] [CrossRef]
- Shirakawa, K. T.; Sala, F. A.; Miyachiro, M. M.; Job, V.; Trindade, D. M.; Dessen, A. Architecture and Genomic Arrangement of the MurE–MurF Bacterial Cell Wall Biosynthesis Complex. Proc. Natl. Acad. Sci. U.S.A. 2023, 120(21), e2219540120. [Google Scholar] [CrossRef]
- Rossolini, G. M. Extensively Drug-Resistant Carbapenemase-Producing Enterobacteriaceae: An Emerging Challenge for Clinicians and Healthcare Systems. J. Intern. Med. 2015, 277(5), 528–531. [Google Scholar] [CrossRef]
- King, A. M.; Reid-Yu, S. A.; Wang, W.; King, D. T.; De Pascale, G.; Strynadka, N. C.; Walsh, T. R.; Coombes, B. K.; Wright, G. D. AMA Overcomes Antibiotic Resistance by NDM and VIM Metallo-β-Lactamases. Nature 2014, 510(7506), 503–506. [Google Scholar] [CrossRef]
- Palica, K.; Deufel, F.; Skagseth, S.; Di Santo Metzler, G. P.; Thoma, J.; Andersson Rasmussen, A.; Valkonen, A.; Sunnerhagen, P.; Leiros, H.-K.; Andersson, H.; Erdelyi, M. α-Aminophosphonate Inhibitors of Metallo-β-Lactamases NDM-1 and VIM-2. RSC Med. Chem. 2023, 14(11), 2277–2300. [Google Scholar] [CrossRef]
- Hijazi, A. K.; El-Khateeb, M.; Taha, Z. A.; et al. Anti-Bacterial and Anti-Fungal Properties of a Set of Transition Metal Complexes Bearing a Pyridine Moiety and [B(C₆F₅)₄]₂ as a Counter Anion. Molecules 2025, 30(15), 3121. [Google Scholar] [CrossRef] [PubMed]
- Will, V.; Frey, C.; Normant, V.; et al. The Role of FoxA, FiuA, and FpvB in Iron Acquisition via Hydroxamate-Type Siderophores in Pseudomonas aeruginosa. Sci. Rep. 2024, 14, 18795. [Google Scholar] [CrossRef]
- Beck, A.; Goetsch, L.; Dumontet, C.; Corvaïa, N. Strategies and Challenges for the Next Generation of Antibody–Drug Conjugates. Nat. Rev. Drug Discov. 2017, 16(5), 315–337. [Google Scholar] [CrossRef] [PubMed]
- Hood, M. I.; Skaar, E. P. Nutritional Immunity: Transition Metals at the Pathogen–Host Interface. Nat. Rev. Microbiol. 2012, 10(8), 525–537. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Su, Z.; Chen, P.; Tian, X.; Wu, L.; Tang, D.-J.; Li, P.; Deng, H.; Ding, P.; Fu, Q.; Tang, J.-L.; Ming, Z. Structural Basis for Zinc-Induced Activation of a Zinc Uptake Transcriptional Regulator. Nucleic Acids Res. 2021, 49(11), 6511–6528. [Google Scholar] [CrossRef]
- Šink, R.; Kotnik, M.; Zega, A.; Barreteau, H.; Gobec, S.; Blanot, D.; et al. Crystallographic Study of Peptidoglycan Biosynthesis Enzyme MurD: Domain Movement Revisited. PLoS ONE 2016, 11(3), e0152075. [Google Scholar] [CrossRef]
- Murray, C. J. L.; et al. Global Burden of Bacterial Antimicrobial Resistance. Lancet 2022, 399(10325), 629–655. [Google Scholar] [CrossRef]
- Huang, Z.; Bian, X.; Li, Y.; et al. In Vitro Pharmacokinetics/Pharmacodynamics of FL058 (a Novel β-Lactamase Inhibitor) Combined with Meropenem against Carbapenemase-Producing Enterobacterales. Front. Pharmacol. 2024, 15, 1282480. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, S.; Chen, W.; et al. CRISPR-Cas9 and CRISPR-Assisted Cytidine Deaminase Enable Precise and Efficient Genome Editing in Klebsiella pneumoniae. Appl. Environ. Microbiol. 2018, 84, e01834-18. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.; Wu, X.; Li, Z.; et al. Molecular Mechanism of Siderophore Regulation by the Pseudomonas aeruginosa BfmRS Two-Component System in Response to Osmotic Stress. Commun. Biol. 2024, 7, 295. [Google Scholar] [CrossRef]
- Colaço, H. G.; Santo, P. E.; Matias, P. M.; Bandeiras, T. M.; Vicente, J. B. Roles of Escherichia coli ZinT in Cobalt, Mercury and Cadmium Resistance and Structural Insights into the Metal Binding Mechanism. Metallomics 2016, 8(3), 327–336. [Google Scholar] [CrossRef]
- Capdevila, D. A.; Wang, J.; Giedroc, D. P. Bacterial Strategies to Maintain Zinc Metallostasis at the Host–Pathogen Interface. J. Biol. Chem. 2016, 291(40), 20858–20868. [Google Scholar] [CrossRef]
- Djoko, K. Y.; Ong, C. L.; Walker, M. J.; McEwan, A. G. The Role of Copper and Zinc Toxicity in Innate Immune Defense against Bacterial Pathogens. J. Biol. Chem. 2015, 290(31), 18954–18961. [Google Scholar] [CrossRef]
- Poole, K. At the Nexus of Antibiotics and Metals: The Impact of Cu and Zn on Antibiotic Activity and Resistance. Trends Microbiol. 2017, 25(10), 820–832. [Google Scholar] [CrossRef] [PubMed]
- Palmer, L. D.; Skaar, E. P. Transition Metals and Virulence in Bacteria. Annu. Rev. Genet. 2016, 50, 67–91. [Google Scholar] [CrossRef]
- King, A.; Reid-Yu, S.; Wang, W.; et al. Aspergillomarasmine A Overcomes Metallo-β-Lactamase Antibiotic Resistance. Nature 2014, 510, 503–506. [Google Scholar] [CrossRef]
- Garau, G.; Di Guilmi, A. M.; Hall, B. G. Structure-Based Phylogeny of the Metallo-β-Lactamases. Antimicrob. Agents Chemother. 2005, 49(7), 2778–2784. [Google Scholar] [CrossRef] [PubMed]
- Palica, K.; Deufel, F.; Skagseth, S.; et al. α-Aminophosphonate Inhibitors of Metallo-β-Lactamases NDM-1 and VIM-2. RSC Med. Chem. 2023, 14(11), 2277–2300. [Google Scholar] [CrossRef]
- Kehl-Fie, T. E.; Skaar, E. P. Nutritional Immunity Beyond Iron: A Role for Manganese and Zinc. Curr. Opin. Chem. Biol. 2010, 14(2), 218–224. [Google Scholar] [CrossRef] [PubMed]
- Lemire, J.; Harrison, J.; Turner, R. Antimicrobial Activity of Metals: Mechanisms, Molecular Targets and Applications. Nat. Rev. Microbiol. 2013, 11, 371–384. [Google Scholar] [CrossRef]
- Jiang, D.; Ye, Z.; Hsieh, C. Y.; et al. MetalProGNet: A Structure-Based Deep Graph Model for Metalloprotein–Ligand Interaction Predictions. Chem. Sci. 2023, 14, 2054–2069. [Google Scholar] [CrossRef]
- Kumar, L.; Bisen, M.; Harjai, K.; et al. Advances in Nanotechnology for Biofilm Inhibition. ACS Omega 2023, 8, 21391–21409. [Google Scholar] [CrossRef]
- Jumper, J.; Evans, R.; Pritzel, A.; et al. Highly Accurate Protein Structure Prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y. Single-Particle Cryo-EM at Crystallographic Resolution. Cell 2015, 161, 450–457. [Google Scholar] [CrossRef]
- Li, G.; Hu, Y.; Xu, F.; et al. Chelation Strategies in Antimicrobial Design. Bioorg. Med. Chem. 2019, 27, 115–123. [Google Scholar] [CrossRef]
- Zhang, H.; Hao, Q. Crystal Structure of NDM-1 Reveals Zinc-Binding Site. PLoS ONE 2011, 6(9), e24621. [Google Scholar] [CrossRef]
- Costa, S.; Ragusa, M. A.; Lo Buglio, G.; Scilabra, S. D.; Nicosia, A. The Repertoire of Tissue Inhibitors of Metalloproteases: Evolution, Regulation of Extracellular Matrix Proteolysis, Engineering and Therapeutic Challenges. Life 2022, 12(8), 1145. [Google Scholar] [CrossRef]
- Jain, R. K.; Stylianopoulos, T. Delivering Nanomedicine to Solid Tumors. Nat. Rev. Clin. Oncol. 2010, 7, 653–664. [Google Scholar] [CrossRef]
- Békés, M.; Langley, D. R.; Crews, C. M. PROTAC Targeted Protein Degraders. Nat. Rev. Drug Discov. 2022, 21, 181–200. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Chen, J.; et al. Siderophore-Conjugated Nanoparticles. ACS Nano 2021, 15, 1234–1245. [Google Scholar] [CrossRef]
- Su, J.; Li, Z.; Liao, B.; et al. Microcalorimetric Study of the Effect of Manganese on the Growth and Metabolism in a Heterogeneously Expressing Manganese-Dependent Superoxide Dismutase (Mn-SOD) Strain. J. Therm. Anal. Calorim. 2017, 130, 1407–1416. [Google Scholar] [CrossRef]
- Gasparrini, A. J.; Wang, B.; Sun, X.; et al. Persistent Metagenomic Signatures of Early-Life Hospitalization and Antibiotic Treatment in the Infant Gut Microbiota and Resistome. Nat. Microbiol. 2019, 4(12), 2285–2297. [Google Scholar] [CrossRef]
- Becker, K. W.; Skaar, E. P. Metal Limitation and Toxicity at the Interface between Host and Pathogen. FEMS Microbiol. Rev. 2014, 38(6), 1235–1249. [Google Scholar] [CrossRef]
- Krokidis, M. G.; Koumadorakis, D. E.; Lazaros, K.; Ivantsik, O.; Exarchos, T. P.; Vrahatis, A. G.; Kotsiantis, S.; Vlamos, P. AlphaFold3: An Overview of Applications and Performance Insights. Int. J. Mol. Sci. 2024, 26(8), 3671. [Google Scholar] [CrossRef]
- Haloi, N.; Howard, R. J.; Lindahl, E. Cryo-EM Ligand Building Using AlphaFold3-like Model and Molecular Dynamics. PLoS Comput. Biol. 2025, 21(8), e1013367. [Google Scholar] [CrossRef]
- Lysenko, V.; Gao, M.-L.; Sterk, F. A. C.; Innocenti, P.; Slingerland, C. J.; Martin, N. I. ACS Infect. Dis. 2025, 11(8), 2301–2309. [CrossRef]
- Sherry, N. L.; Lee, J. Y. H.; Giulieri, S. G.; Connor, C. H.; Horan, K.; Lacey, J. A.; Lane, C. R.; Carter, G. P.; Seemann, T.; Egli, A.; Stinear, T. P.; Howden, B. P. Genomics for Antimicrobial Resistance—Progress and Future Directions. Antimicrob. Agents Chemother. 2025, 69, e01082-24. [Google Scholar] [CrossRef] [PubMed]
- Kasmanas, J. C.; Magnúsdóttir, S.; Zhang, J.; Smalla, K.; Schloter, M.; Stadler, P. F.; Ponce, A. C.; Rocha, U. Integrating Comparative Genomics and Risk Classification by Assessing Virulence, Antimicrobial Resistance, and Plasmid Spread in Microbial Communities with gSpreadComp. GigaScience 2025, 14, giaf072. [Google Scholar] [CrossRef]
- Yeung, A.; Hale, C.; Clare, S.; Palmer, S.; Bartholdson Scott, J.; Baker, S.; Dougan, G. Using a Systems Biology Approach To Study Host–Pathogen Interactions. Microbiol. Spectr. 2019, 7(2), BAI-0021-2019. [Google Scholar] [CrossRef]
- Johnson, D. E. In Computational Systems Pharmacology and Toxicology; Richardson, R. J., Johnson, D. E., Eds.; The Royal Society of Chemistry: Cambridge, 2017; pp 1–18.
- Francine, P. Systems Biology: New Insight into Antibiotic Resistance. Microorganisms 2022, 10(12), 2362. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Xu, Y.; Xu, X.; Chen, X.; Chen, H.; Zhang, J.; Ma, J.; Zhang, W.; Zhang, R.; Chen, J. Metagenomic Identification of Pathogens and Antimicrobial-Resistant Genes in Bacterial Positive Blood Cultures by Nanopore Sequencing. Front. Cell. Infect. Microbiol. 2023, 13, 1283094. [Google Scholar] [CrossRef]
- Olsen, N. S.; Riber, L. Metagenomics as a Transformative Tool for Antibiotic Resistance Surveillance: Highlighting the Impact of Mobile Genetic Elements with a Focus on the Complex Role of Phages. Antibiotics 2025, 14(3), 296. [Google Scholar] [CrossRef]
- Gräff, Á. T.; Barry, S. M. Siderophores as Tools and Treatments. npj Antimicrob. Resist. 2024, 2, 47. [Google Scholar] [CrossRef]
- Keskey, R. C.; Xiao, J.; Hyoju, S.; et al. Enterobactin Inhibits Microbiota-Dependent Activation of AhR to Promote Bacterial Sepsis in Mice. Nat. Microbiol. 2025, 10, 388–404. [Google Scholar] [CrossRef]
- Kayode, M.; Ayodeji, O. E.; Yetunde, B. B.; et al. Synthesis, Characterization and Antimicrobial Activity of Mixed Antibiotic–Vitamin Metal Complexes Involving Metronidazole and Vitamin B1 (Thiamine). Asian J. Chem. Sci. 2024, 14(6), 46–59. [Google Scholar] [CrossRef]
- Nahar, L.; Hagiya, H.; Gotoh, K.; et al. New Delhi Metallo-β-Lactamase Inhibitors: A Systematic Scoping Review. J. Clin. Med. 2024, 13(14), 4199. [Google Scholar] [CrossRef] [PubMed]
- Ogasawara, Y.; Shimizu, Y.; Sato, Y.; et al. Identification of Actinomycin D as a Specific Inhibitor of the Alternative Pathway of Peptidoglycan Biosynthesis. J. Antibiot. 2020, 73, 125–127. [Google Scholar] [CrossRef]
- Huang, Z.; Bian, X.; Li, Y.; et al. In Vitro Pharmacokinetics/Pharmacodynamics of FL058 (a Novel β-Lactamase Inhibitor) Combined with Meropenem against Carbapenemase-Producing Enterobacterales. Front. Pharmacol. 2024, 15, 1282480. [Google Scholar] [CrossRef] [PubMed]
- Schalk, I. J. Bacterial siderophores: Diversity, uptake pathways and applications. Nature Reviews Microbiology 2024, 23(1), 24–40. [Google Scholar] [CrossRef]
- Katsube, T.; Echols, R.; Arjona Ferreira, J. C.; Krenz, H. K.; Berg, J. K.; Galloway, C. Cefiderocol, a Siderophore Cephalosporin for Gram-Negative Bacterial Infections: Pharmacokinetics and Safety in Subjects with Renal Impairment. J. Clin. Pharmacol. 2016, 57(5), 584–592. [Google Scholar] [CrossRef] [PubMed]
- Sheldon, J. R.; Skaar, E. P. Acinetobacter baumannii Can Use Multiple Siderophores for Iron Acquisition, but Only Acinetobactin Is Required for Virulence. PLoS Pathog. 2020, 16(10), e1008995. [Google Scholar] [CrossRef] [PubMed]
- Serebnitskiy, Z.; Orban, K.; Finkel, S. E. A Role for Dps Ferritin Activity in Long-Term Survival of Escherichia coli. Microbiol. Spectr. 2025, 13, e01837-24. [Google Scholar] [CrossRef]
- Clemente, C. M.; Prieto, J. M.; Martí, M. Precision Docking for Metalloproteins. J. Chem. Inf. Model. 2024, 64(5), 1581–1592. [Google Scholar] [CrossRef]
- Sheng, Y.; Li, H.; Liu, M.; et al. A Manganese-Superoxide Dismutase from Thermus thermophilus HB27 Suppresses Inflammatory Responses and Alleviates Experimentally Induced Colitis. Inflamm. Bowel Dis. 2019, 25(10), 1644–1655. [Google Scholar] [CrossRef]
- Almeida, M. R.; Videira, M. A.; Lima, J. C.; Saraiva, L. Mechanistic Insights into Staphylococcus aureus IsdG-Ferrochelatase Interactions: A Key to Understanding Haem Homeostasis in Pathogens. J. Inorg. Biochem. 2025, 269, 112878. [Google Scholar] [CrossRef] [PubMed]

| Clinical Stage | Toxicity Profile | Selectivity Index | Delivery Method | Target Mechanism | Agent |
| Preclinical | Potential host metal depletion | Moderate | Systemic (small molecule) | Chelates Fe³⁺, disrupts metalloenzymes | 2,2′-Bipyridyl [1,14] |
| Experimental (in vivo) | Low (host-independent) | High | Endogenous bacterial secretion | Siderophore-mediated Fe³⁺ sequestration | Enterobactin (overproduced) [1,4] |
| Phase I/II clinical trials | Minimal off-target effects | High | Intravenous or oral | Zinc-binding at β-lactamase active site | NDM-1 Inhibitors [10,11] |
| Preclinical | Low toxicity, favorable pharmacokinetics | High | Topical/systemic nanoparticle | Chelates Fe³⁺, inhibits biofilm formation | OP607 [12,13] |
| Early-stage development | Low systemic toxicity | Very high | Intravenous (biologic) | Surface metalloprotein neutralization | Antibody-based inhibitors [15,16] |
| Limitations | Advantages | Evidence | Therapeutic Strategy | Mechanism | Pathogens | Target Class |
| Selectivity challenges | Unique bacterial targets | In vitro inhibition | Small molecule inhibitors | Copper uptake & oxidative stress | P. aeruginosa | Copper Transport Proteins [26] |
| Resistance via transporter mutations | Pathogen-specific | Mouse peritonitis, UTI models | Siderophore analogs, chelators | High-affinity Fe³⁺ acquisition | E. coli, | Iron-Siderophore Systems [1,4] |
| S. aureus | ||||||
| Risk of off-target ROS imbalance | Direct virulence target | Genetic knockout & virulence models | Enzyme inhibitors | ROS detoxification | S. aureus | Mn-Superoxide Dismutase [2] |
| Host enzyme similarity | Essential enzymes | Structural biology + inhibitor screens | Zn-binding site inhibitors | DNA replication/repair | N. gonorrhoeae | Zn-dependent Enzymes [3] |
| Redundancy across strains | Virulence disruption | In vitro & tissue models | Protease inhibitors | Host tissue invasion | H. pylori | Metalloprotein-ases [27] |
| Representative Example | Disadvantages | Advantages | Strategy Type |
| Broad-spectrum MMP inhibitors (failed trials) | Poor selectivity; systemic toxicity (e.g., musculoskeletal pain) | High potency; simple design | Zn-chelating small molecules [5,26] |
| S1′ pocket-targeted MMP-7 inhibitors | Requires detailed structural data; complex SAR | Isoform selectivity; avoids metal stripping | Non-chelating pocket binders [7,8] |
| ADAMTS-selective peptide inhibitors | Proteolytic instability; delivery challenges | High affinity; modular design | Peptides / Peptidomimetics [15] |
| Engineered TIMPs | Costly; limited tissue penetration; immunogenicity | High specificity; long half-life | Biologics (TIMPs, antibodies) [24,27] |
| ATCUN motif-based metallodrugs | Metal lability; systemic toxicity | Novel mechanisms; prodrug activation | Catalytic metallodrugs [18,26] |
| Pt-PROTACs degrading Trx1/TrxR1 | Large size; permeability and E3 ligase constraints | Permanent target removal | Targeted degraders (metallo-PROTACs) [26] |
| MMP-responsive nanoparticles | Complex formulation; variable enzyme expression | Spatial control; reduced systemic exposure | Responsive delivery systems [13] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




