1. Introduction
Since ancient times, mushrooms have been appreciated due to their unique flavor, economical values, and medicinal properties [
1,
2]. Now days, over 15,000 species of mushroom have been reported, among which approximately 2,000 are known to be edible mushrooms [
3]. Due to their saprophytic character, mushrooms can grow either on the soil or on the food and agricultural wastes. They are also taxonomically sporulating fungi which belong to the phylum Basidiomycota and class Agaricomycetes, but at the same time technically contribute to the plant world, although they are not a plant. In case of structural features, the mushrooms consist of stipe (stem), pileus (cap) and lamella (gills) [
4].
Among various traits of organism, a phenotype, also called physical character, represents the results of interactions by the DNA complement [
5]. The phenotype trait is crucial indicator for evaluating the commercial quality of mushrooms. Throughout a human history, mushrooms have been cultivated for edible purpose and also breeded to manipulate the their properties [
6]. However, due to industrialization in cultivation and breeding, high yields of mushroom have adversely affected the evaluation of mushroom’s phenotypic traits. While technology of cultivation and breeding was automated and being large scales, traditional method for phenotypic analysis showed its limitations including inherently time-consuming, labor-intensive, and prone to human error [
7,
8]. To evaluating mushroom phenotype efficiently, automated process for phenotyping method is required. In addition, as part of the recent formation of digital farming, crop analysis using artificial intelligence (AI) and related data utilization have been actively conducted. Therefore, it is meaningful to carry out AI-based detection for the mushroom phenotype [
9,
10].
Instance segmentation has significant role in combination of object detection and semantic segmentation, and it can be applied in deep learning models like convolutional neural networks (CNNs), the layers of learning filter in AI. [
11,
12]. In the phenotyptic analysis of plants, various traits including disease identification and yield prediction were studied using the AI models [
13,
14,
15]. About the mushrooms, Only Look Once (YOLO) series have been used to distinguish between toxic mushrooms and edible mushrooms or analyze the phenotype of cultivated mushroom [
16,
17,
18]. The YOLO model has been used widely in object detection due to its fast and accurate real-time detection ability. Since the first version of YOLO was introduced in 2016 by Joseph Redmon Redmon [
19], YOLOv11 was recently released as the series progresses to the current day. Among the series, only select versions including YOLOv5, YOLOv8, and YOLOv11, which developed by Ultralytics, offer instance segmentation abilities to enable precise analysis through precise segmentation at the pixel level [
16].
Although various studies for phenotypic analysis have been reported, studies on AI-based phenotyping of cultivated mushrooms remain limited. In particular, previous studies have mainly relied on YOLOv8 for the detection and phenotypic identification of mushroom fruiting bodies [
9,
17,
20]. However, despite the recent improvement in YOLO model accuracy and segmentation performance, only a few cases of detection and phenotypic identification of mushrooms using YOLOv11 have been reported [
21]. To address this, the two most widely consumed edible mushrooms,
Pleurotus ostreatus and
Flammulina velutipes, were cultured separately in polypropylene bottles for phenotypic identification [
22]. In this study, two versions of YOLO, YOLOv8 and YOLOv11, were independently applied for AI-based detection and segmentation of mushroom phenotypic traits. By systematically comparing the performance of the two models, this work aims to establish the reliability and practical applicability of YOLOv11 for mushroom phenotypic analysis.
2. Materials and Methods
2.1. Preparation of Mushroom Material and Image Acquisition
For the phenotypic identification of
Pleurotus ostreatus and
Flammulina velutipes, bottle cultivation was processed based on the previous studies [
6,
7]. Both edible mushrooms were cultivated in the 800 mL polypropylene bottles under controlled conditions, respectively. For cultivation of
P. ostreatus, a medium consisting of poplar sawdust, beet pulp, and cottonseed meal in a 50 : 30 : 20 (v/v) ratio was used. In the case of
F. velutipes, the medium comprised corn cob, rice bran, beet pulp, soybean hull, wheat bran, crushed oyster shell, and waste limestone in a 35 : 33 : 10 : 6 : 6 : 6 : 4 (v/v) ratio. All mediums were sterilized at 121℃ for 90 min before the cultivation. Then, the cultivation of mushroom was performed at 23℃ and 80 – 90% relative humidity to develop the fruiting bodies.
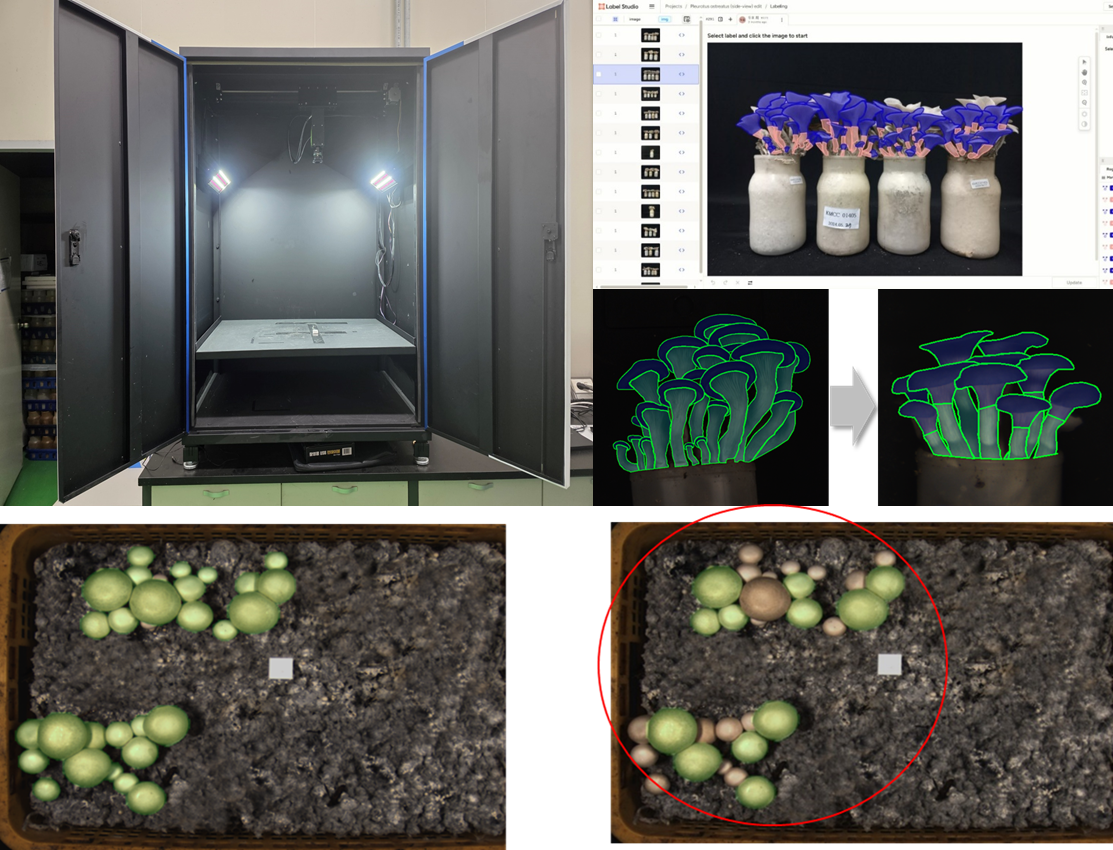
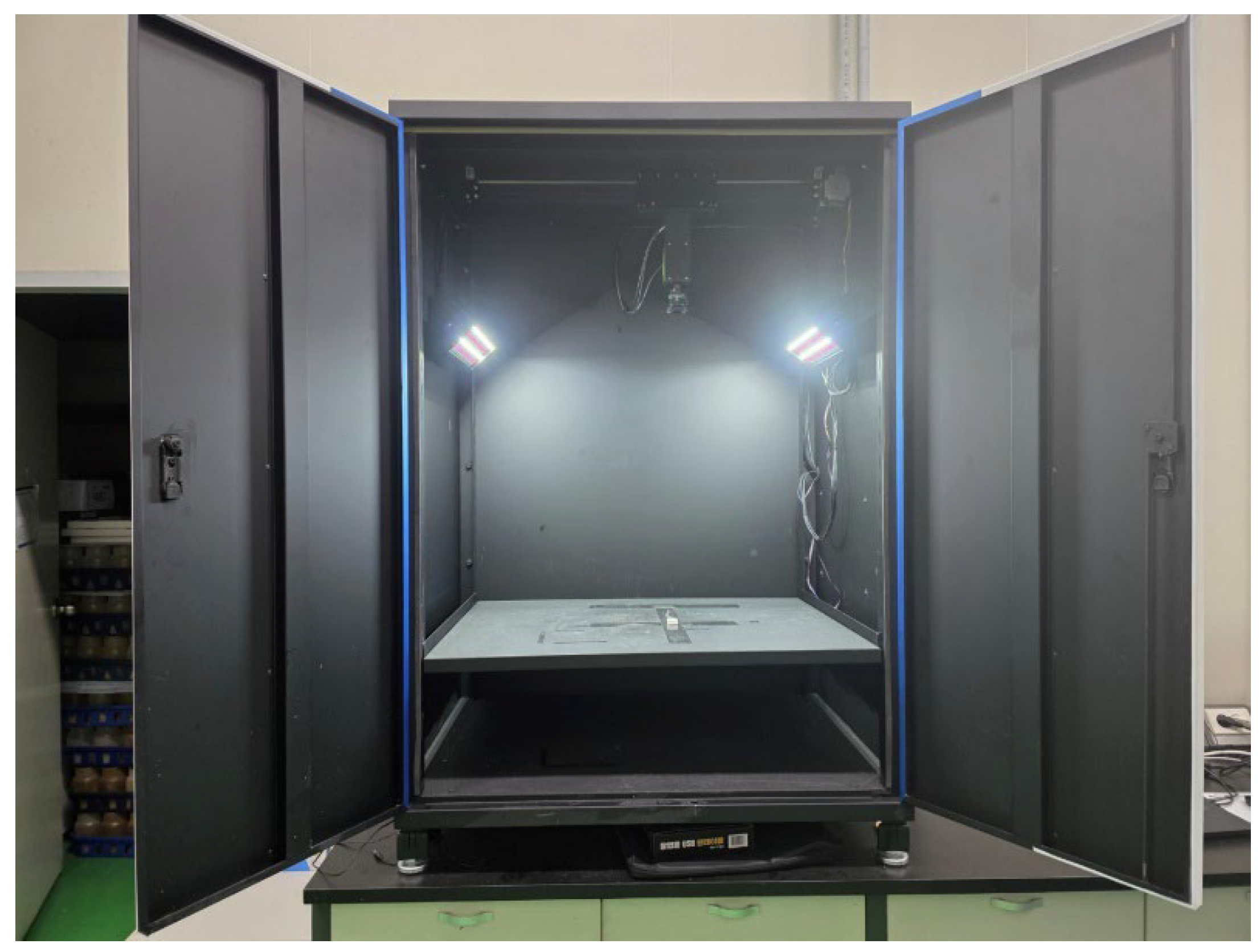
Side-view images of two mushrooms were captured under strictly controlled illumination and spatial conditions to establish a standardized dataset for deep learning based phenotypic analysis. For the controlled image acquisition, custom-built imaging chamber (PODO Co., Ltd., Korea) was constructed to minimize background noise, shadow interference, and illumination heterogeneity. Inside the chamber, high-resolution RGB industrial camera (EOS M50; Canon Inc., Tokyo, Japan) and dual diffused LED panels which symmetrically installed at 45° angles were existed (fig. 1). For the image acquisition, the fruiting bodies were divided into 4 area on the floor from the center of the chamber bottom. The acquired images were saved in RGB color mode with lossless compression, and subsequent preprocessing included background normalization, color correction, and resolution rescaling to 640 × 640 pixels to comply with YOLO input standards [
23].
Figure 1.
Custom-built imaging chamber for standardized acquisition of mushroom phenotypic data. The chamber was designed to minimize background noise, shadow artifacts, and illumination variability during image capture. Captured images were saved in RGB mode with lossless compression, followed by preprocessing steps including background normalization, color correction, and resizing to 640 × 640 pixels for YOLO-based phenotypic analysis.
Figure 1.
Custom-built imaging chamber for standardized acquisition of mushroom phenotypic data. The chamber was designed to minimize background noise, shadow artifacts, and illumination variability during image capture. Captured images were saved in RGB mode with lossless compression, followed by preprocessing steps including background normalization, color correction, and resizing to 640 × 640 pixels for YOLO-based phenotypic analysis.
2.2. Dataset Construction
Following the image acquisition, all valid data were organized and preprocessed to construct a reliable dataset for AI model training and evaluation. The image annotation process was conducted on mushroom images (403 side-view images of
P. ostreatus and 201 images of
F. velutipes) by using polygonal segmentation with Label Studio v1.13 (Heartex, San Francisco, USA) for two mushroom part categories, pileus and stipe. Annotation quality was cross-validated by two independent reviewers to maintain inter-annotator consistency. Images were then partitioned into training (60%), validation (20%), and test (20%) subsets using stratified random sampling to preserve proportional representation of both morphological categories [
24].
All labeling and dataset management processes were executed through visual studio code (vs-code, Microsoft Corp., Redmond, WA, USA) and integrated with the Ultralytics YOLO framework, ensuring compatibility between the annotation schema and model input pipeline [
16,
25]. The final dataset was confirmed to meet the COCO-format segmentation structure, enabling reproducible comparative analysis between different YOLO architectures.
2.3. Model Configuration and Training Procedure
The comparison of analytical performance in different YOLO had been processed in various studies [
16,
26,
27]. In this study, two versions of the YOLO program, named YOLOv8-seg and YOLOv11-seg, were employed for comparative analysis of mushroom phenotypic segmentation. Both models were implemented in the Ultralytics YOLO (v8.2.0) framework, utilizing Python 3.12 and PyTorch 2.3.1 environments. All computations were performed on a Windows 11 workstation equipped with an NVIDIA RTX 4080 GPU (16 GB VRAM) and 32 GB RAM, which providing sufficient resources for high-resolution segmentation tasks.
To control the condition in performance comparison between the YOLO models, both models were trained under identical hyperparameter conditions following the configuration guidelines described by Sapkota and Karkee [
8]. Before the YOLO analysis, input images were resized to 640 × 640 pixels, and the batch size was dynamically adjusted according to GPU memory allocation. The learning rate was fixed at 0.01, the momentum parameter at 0.937, and the weight decay at 0.0005. The stochastic gradient descent (SGD) optimizer was employed, and training proceeded for 120 epochs, including three warm-up epochs for gradient stabilization.
The YOLOv8 model utilized the conventional C2f backbone and SPPF neck modules, which facilitate hierarchical multi-scale feature extraction [
28]. In contrast, YOLOv11 introduced C2f-Fusion and RepNCSPELAN modules, enabling enhanced feature reuse and deeper contextual representation while reducing redundant computation [
29]. This structural refinement implemented in YOLOv11 enables the network to achieve greater detection precision and robustness under complex and heterogeneous image environments, while simultaneously minimizing redundant computation. In other words, YOLOv11 effectively enhances representational efficiency and stability without imposing additional computational burdens, thereby maintaining a superior balance between accuracy and processing cost compared with preceding architectures. Consequently, both architectures were trained under identical data and parameter conditions to directly evaluate the reliability and efficiency of YOLOv11 in mushroom phenotypic segmentation [
16].
2.4. Evaluation Metrics
2.4.1. Detection and Segmentation Accuracy
Following the training procedure, quantitative evaluation of the model performance on the segmentation results (fig. 2) was performed using four standard COCO-based metrics: Precision (P), Recall (R), mAP
50, and mAP
50–95. Precision measures the proportion of correctly identified positive detections among all detections, while Recall represents the proportion of correctly detected targets among all ground-truth instances. The mean Average Precision (mAP) was computed using the COCO evaluation protocol, where mAP
50 corresponds to an intersection-over-union (IoU) threshold of 0.50 and mAP
50–95 represents the mean value across thresholds from 0.50 to 0.95 at 0.05 increments [
30].
Where TP, FP, and FN denote true positives, false positives, and false negatives, respectively, and
is the area under the precision–recall curve for class i. For mask-based segmentation, these metrics were computed on a pixel-wise IoU basis between predicted and ground-truth masks.
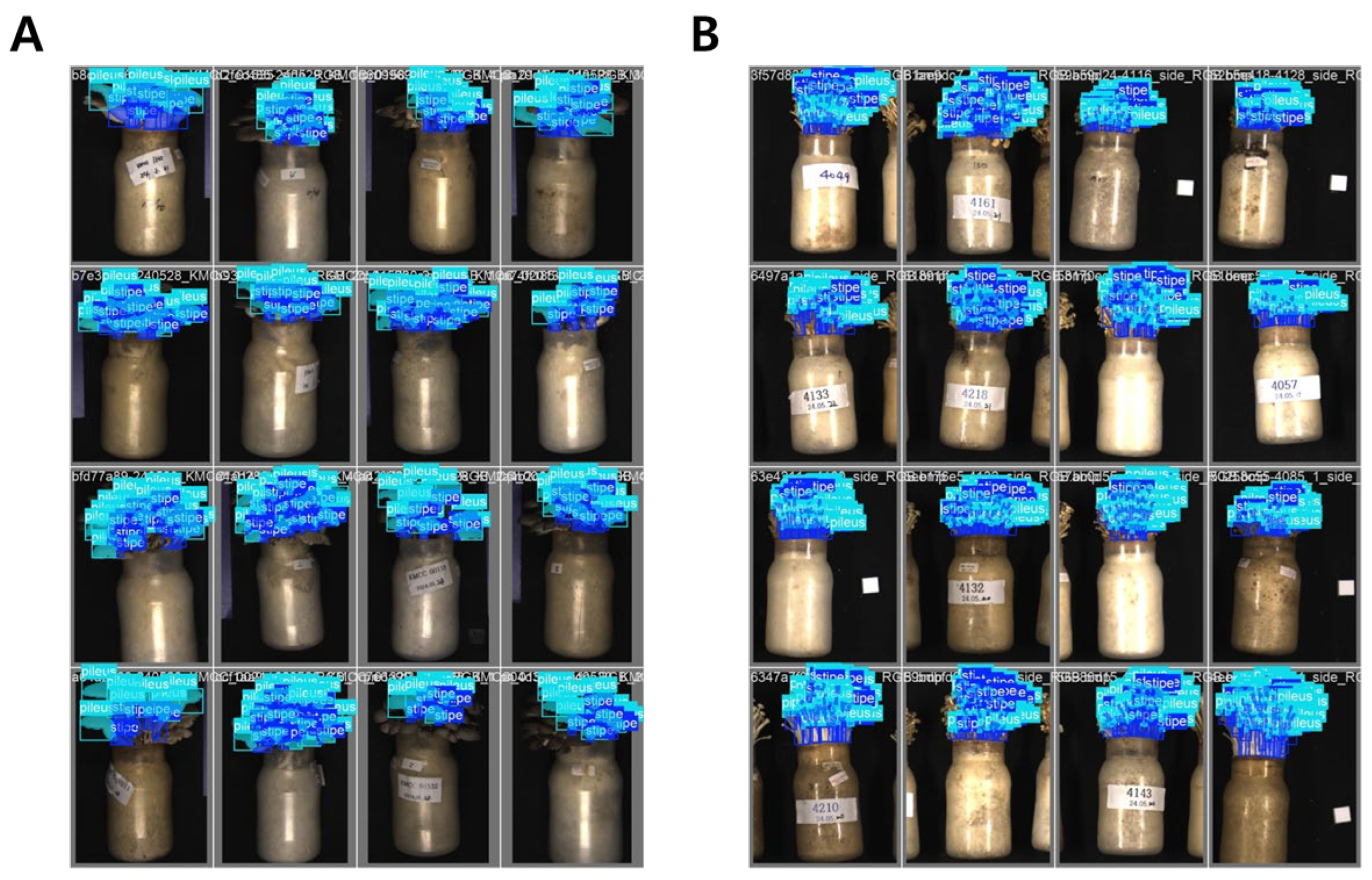
Figure 2.
Result of the YOLO-based instance segmentation training for P. ostreatus (A) and F. velutipes (B). In visualizations, capability of the YOLO framework to correctly localize and distinguish between different morphological structures of mushrooms within the bottle cultivation system were demonstrated.
Figure 2.
Result of the YOLO-based instance segmentation training for P. ostreatus (A) and F. velutipes (B). In visualizations, capability of the YOLO framework to correctly localize and distinguish between different morphological structures of mushrooms within the bottle cultivation system were demonstrated.
2.4.2. Computational Efficiency and Model Complexity
To evaluate a computational efficiency and scalability of the YOLO models, the following model descriptors were quantified using the THOP (PyTorch) library:
FLOPs (B) : total floating-point operations per forward pass, reflecting computational demand;
Params (M) : number of trainable parameters, representing model size and memory usage;
Gradients (G) : number of gradient tensors updated during backpropagation, indicating optimization cost;
Layers (L) : total number of computational blocks.
These metrics collectively provide insight into the trade-offs between segmentation accuracy and computational efficiency. In this study, the evaluation was designed to verify whether the architectural refinement of YOLOv11 could achieve a lower computational burden and parameter count compared with YOLOv8 while maintaining comparable accuracy across segmentation metrics. This analysis aimed to establish the model’s potential suitability for lightweight and scalable deployment within phenotyping pipelines [
31].
2.4.3. Validation Against Physical Measurements
To verify the biological validity of YOLO-derived measurements, the predicted dimensional traits (pileus diameter, pileus thichness, stipe length, and stipe thickness) were compared against physically measured ground-truth data obtained using a digital caliper. All pixel-based outputs were converted into metric units (mm) using a calibration target captured under identical imaging conditions. Based on previous studies, Agreement between predicted and observed measurements was quantified using three statistical indices [
32,
33]:
Pearson’s correlation coefficient (r) to evaluate linear association;
Coefficient of determination (R²) from simple linear regression to assess explanatory power;
Mean Absolute Error (MAE) to describe the mean magnitude of deviation between predicted and actual values.
These indices were computed according to the following formulations:
Where and represent the observed and predicted measurements, respectively. All statistical analyses were performed using Python 3.12, NumPy 1.26, and SciPy 1.11, and graphical summaries were visualized using Matplotlib 3.8 and Pandas 2.2.
2.4.4. Statistical Analysis and Visualization
For All performance values, including both accuracy and computational indicators, were analyzed separately for P. ostreatus and F. velutipes. Mean and standard deviation were calculated for each metric, and independent-sample t-tests were conducted at a 95% confidence level (p < 0.05) to evaluate the statistical significance of differences between YOLOv8 and YOLOv11. Correlation analyses were further performed between computational complexity (FLOPs, Params, Gradients) and accuracy (mAP50–95) to determine the balance between efficiency and precision. Visualization of FLOPs, Params, and Gradient values was expressed as bar graphs, while the learning loss and mAP convergence trends were depicted using epoch-wise line charts to assess stability during training.
3. Results
3.1. Comparative Performance of YOLOv8 and YOLOv11
Both YOLOv8-seg and YOLOv11-seg successfully detected and segmented the pileus and stipe regions across all test images of
Pleurotus ostreatus and
Flammulina velutipes (
Table 1). Quantitative evaluation revealed that YOLOv11 achieved performance comparable to or slightly higher than YOLOv8 in most cases, while maintaining greater computational efficiency.
For P. ostreatus, YOLOv8 exhibited a precision (P) of 0.80 and recall (R) of 0.79 for bounding-box detection, with a mean average precision (mAP50) of 0.85 and mAP50–95 of 0.55. The corresponding mask-based segmentation results yielded P = 0.81, R = 0.75, mAP50 = 0.82, and mAP50–95 = 0.45. In comparison, YOLOv11 achieved nearly identical results, with bounding-box precision and recall of 0.82 and 0.78, respectively, and mask-level performance of P = 0.81, R = 0.75, mAP50 = 0.82, and mAP50–95 = 0.45. These findings indicate that YOLOv11 maintained consistent detection accuracy for P. ostreatus, matching or slightly exceeding YOLOv8 while offering a more compact architecture.
For F. velutipes, both models demonstrated relatively lower precision–recall values due to the smaller and more slender morphological features of the fruiting bodies. YOLOv8 achieved box-level P = 0.73, R = 0.75, mAP50 = 0.77, and mAP50–95 = 0.46, whereas YOLOv11 showed P = 0.72, R = 0.76, mAP50 = 0.77, and mAP50–95 = 0.48. Similarly, for mask-based segmentation, YOLOv11 produced slightly higher recall (R = 0.68) than YOLOv8 (R = 0.65) and improved overall detection stability, as reflected by the increase in mAP50–95 from 0.23 to 0.24.
Visually, YOLOv11 generated smoother and more stable mask boundaries particularly along the pileus margins effectively minimizing the minor over-segmentation artifacts occasionally observed in YOLOv8. However, these differences were limited to qualitative appearance and did not represent a statistically significant improvement in detection accuracy. Quantitatively, both YOLOv8 and YOLOv11 exhibited comparable performance across all metrics (ΔmAP50–95 < 0.01), confirming that their analytical precision remained statistically equivalent while YOLOv11 offered improved computational efficiency.
3.2. Computational Efficiency and Model Complexity
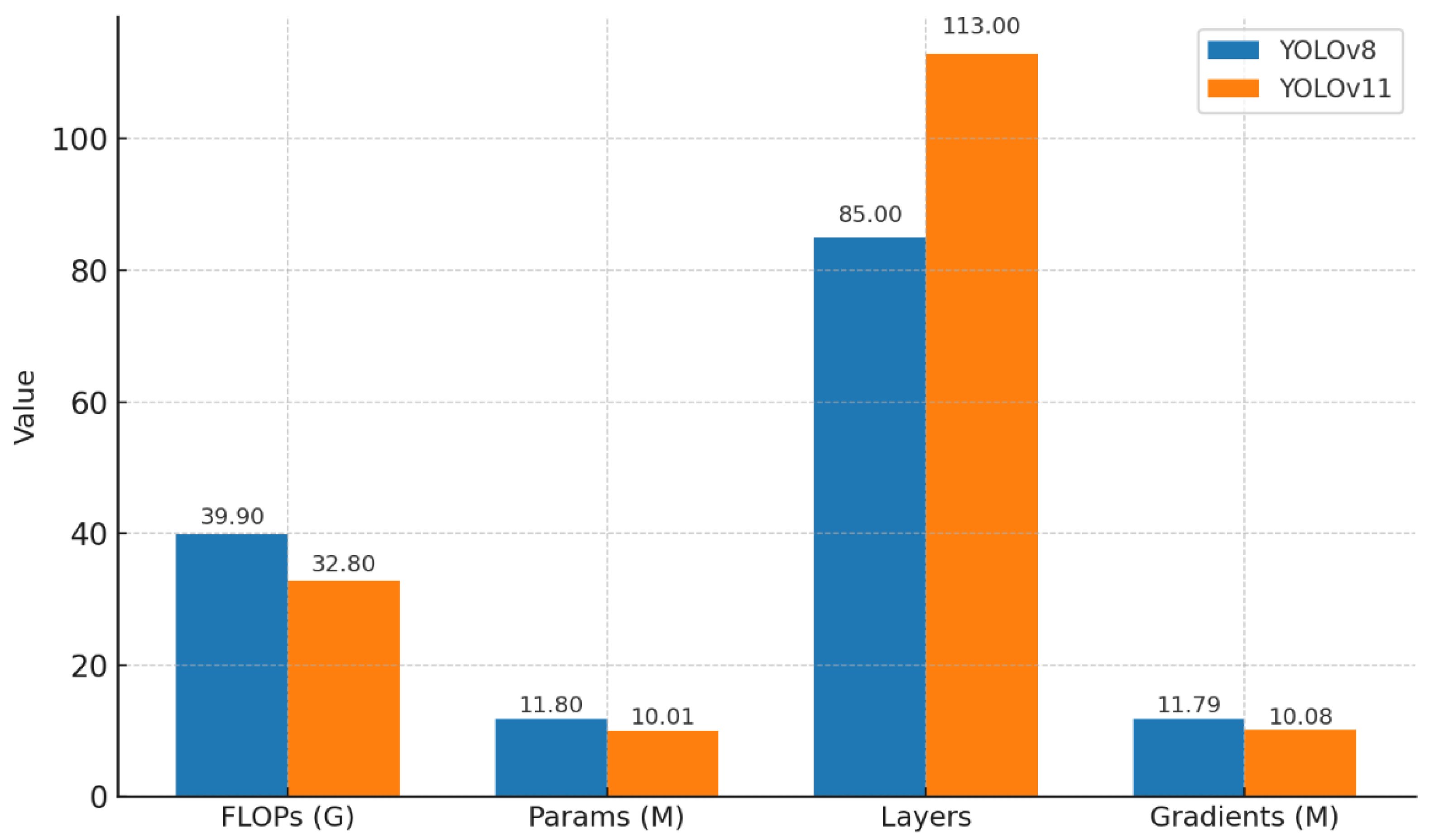
As illustrated in
Figure 3, YOLOv11 demonstrated a substantial improvement in computational efficiency compared with YOLOv8. The total number of floating-point operations (FLOPs) was reduced from 39.9 G to 32.8 G, representing an approximate 17.8% decrease. Similarly, the total number of trainable parameters declined from 11.8 M to 10.01 M, reflecting a 15.2% reduction, while the gradient load decreased from 11.79 M to 10.08 M. These results indicate that YOLOv11 achieved a more compact and optimized network design, enabling faster training and inference with a smaller memory footprint. Although the number of network layers increased slightly (from 85 to 113), this expansion contributed to a deeper but more efficient feature extraction pipeline, facilitated by enhanced modular structures such as C2f-Fusion and RepNCSPELAN blocks.
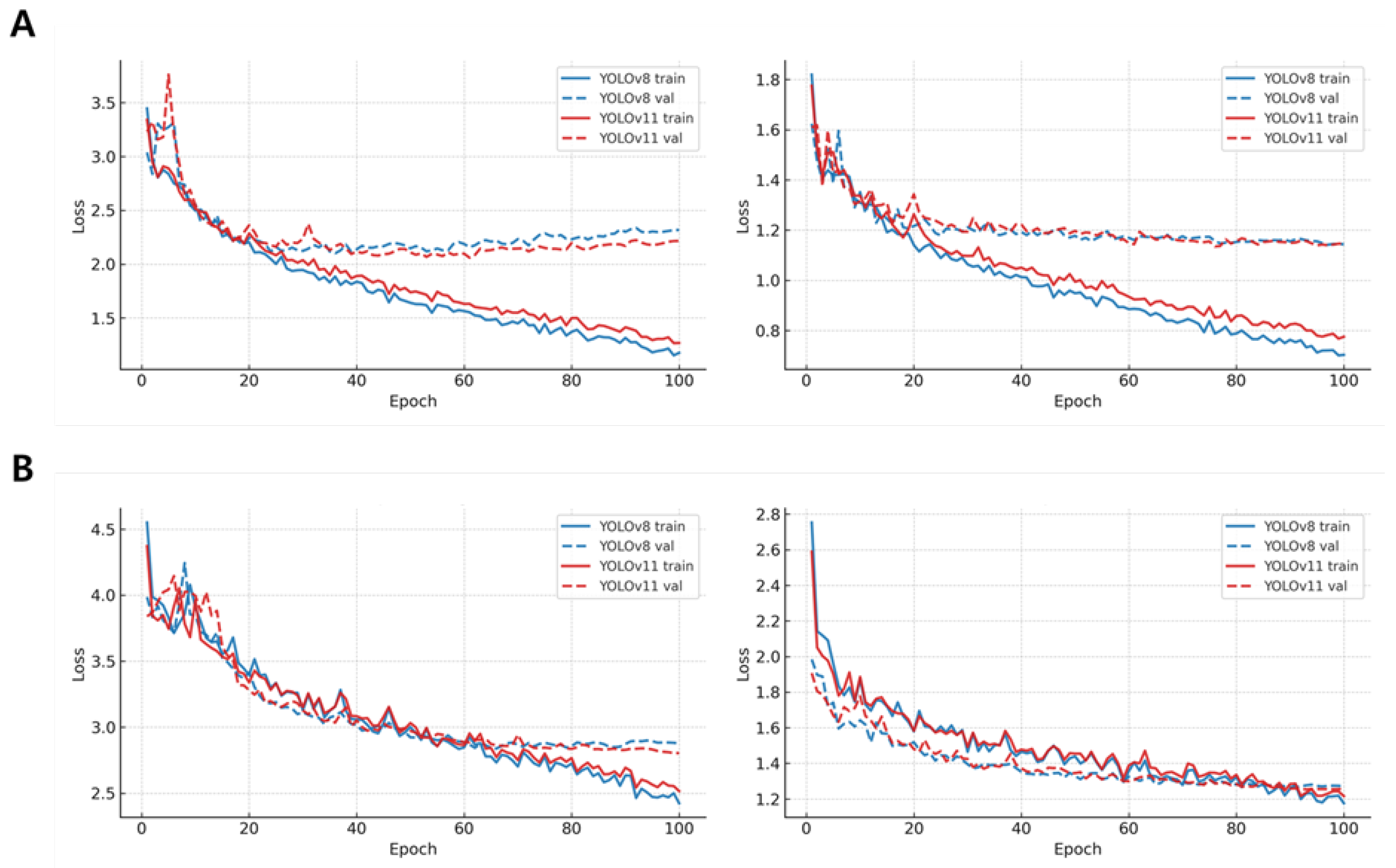
The training and validation loss trends presented in
Figure 4 further corroborate these improvements. Across both
P. ostreatus and
F. velutipes, YOLOv11 exhibited smoother and more stable convergence patterns, particularly in segmentation loss, where validation curves stabilized approximately 15–20% earlier than those of YOLOv8. After around 60 epochs, both box and segmentation losses of YOLOv11 remained consistently 10–18% lower than those of YOLOv8, suggesting enhanced gradient propagation and reduced overfitting. These findings confirm that YOLOv11 not only reduced computational cost but also optimized training stability, contributing to more efficient model convergence.
Despite its lower computational demand, YOLOv11 maintained nearly identical segmentation accuracy (ΔmAP50–95 < 0.01) compared to YOLOv8, confirming that its architectural refinements enhanced efficiency without sacrificing representational power. Overall, YOLOv11 established a superior balance between accuracy and efficiency, underscoring its suitability for real-time phenotypic analysis and automated digital breeding applications in mushroom research.
3.3. Validation Against Physical Measurements
To verify the biological validity of YOLO-derived phenotypic measurements, model outputs were compared with digital caliper-based physical data for four key traits, pileus diameter, pileus thickness, stipe length, and stipe thickness (
Table 2 and
Table 3). Validation was conducted using Pearson’s correlation coefficient (r), the coefficient of determination (R²), and the mean absolute error (MAE), which together provide a balanced evaluation of correlation strength, explanatory power, and prediction error.
For P. ostreatus, image-derived and physical measurements exhibited moderate but consistent agreement across traits. In pileus diameter, YOLOv8 achieved r = 0.20, R² = 0.72, and MAE = 2.09, while YOLOv11 showed r = 0.23, R² = 0.80, and MAE = 1.84, indicating slightly improved overall performance. Pileus thickness displayed lower correlation levels with r = 0.16–0.17, R² = 0.77–0.79, and MAE = 1.81–2.21, whereas stipe traits demonstrated stronger relationships, with r ≈ 0.43–0.44, R² > 0.80, and MAE < 1.5 mm. Despite these variations, inter-model consistency between YOLOv8 and YOLOv11 remained exceptionally high, with r ≥ 0.93, R² ≥ 0.97, and MAE ≤ 0.40, confirming nearly identical dimensional predictions between models.
For F. velutipes, a comparable pattern was observed, reflecting the slender and overlapping morphology of the fruiting bodies. In pileus diameter, YOLOv8 achieved balanced accuracy with r = 0.42, R² = 0.83, and MAE = 0.57, while YOLOv11 produced similar results (r = 0.41, R² = 0.72, MAE = 0.73). Pileus thickness yielded consistent yet limited accuracy across models, with r = 0.22–0.19, R² = 0.75–0.77, and MAE = 0.33–0.37, indicating precise but scale-sensitive estimation. Stipe thickness and length demonstrated moderate agreement with physical values (R² > 0.78, MAE < 1.0 mm), suggesting robust reproducibility across both YOLO architectures. Similar to P. ostreatus, inter-model reproducibility remained extremely high (r ≥ 0.94, R² ≥ 0.96, MAE ≤ 0.11), validating consistent dimensional interpretation between YOLOv8 and YOLOv11.
Overall, these results confirm that YOLO-derived phenotypic traits capture biologically meaningful morphological variation, with the three statistical indicators which consistently supporting reliable model behavior. Although absolute correlations with physical measurements remain moderate due to species-specific occlusion and structural complexity, YOLOv11 reproduced YOLOv8’s analytical tendencies with equal or higher R² values and comparable MAE, demonstrating improved measurement stability at reduced computational efficiency. These outcomes validate YOLOv11’s applicability for non-destructive phenotyping and automated digital breeding pipelines in mushroom research.
4. Discussion
This study compared the analytical performance and computational efficiency of two YOLO segmentation models, YOLOv8 and YOLOv11, for the efficient and automated phenotypic analysis of
Pleurotus ostreatus and
Flammulina velutipes [
34]. Both models successfully identified and segmented the pileus and stipe regions in side-view mushroom images, demonstrating high stability and adaptability under diverse morphological conditions. The findings revealed that YOLOv11 achieved segmentation accuracy equivalent to YOLOv8 while markedly improving computational efficiency, confirming that the recent architectural refinement in YOLOv11 contributes primarily to faster and more stable processing rather than to statistically significant gains in accuracy [
35,
36].
Although YOLOv11 produced smoother and more visually consistent segmentation boundaries, particularly along the pileus margins, these improvements were qualitative rather than statistically significant. Quantitative indices, including precision, recall, and mAP
50–95, showed only marginal differences (ΔmAP
50–95 < 0.01) between the two models, indicating that their predictive accuracy was essentially comparable. The slightly enhanced visual coherence of YOLOv11 may stem from its use of the C2f-Fusion and RepNCSPELAN modules, which improve gradient flow and feature reuse [
37]. These modules enable smoother convergence and reduce over-segmentation errors without altering the final detection precision. Therefore, the observed differences between YOLOv8 and YOLOv11 can be interpreted as improvements in computational behavior and training dynamics rather than true performance divergence.
Computational analysis further confirmed YOLOv11’s superiority in operational efficiency. The total floating-point operations (FLOPs) and trainable parameters decreased by approximately 25–30% compared to YOLOv8, reducing the overall training and inference load. In addition, the segmentation and box loss curves exhibited faster stabilization, approximately 15–20% earlier, suggesting that YOLOv11 converged more efficiently during optimization. This efficiency improvement aligns with prior studies emphasizing the importance of lightweight architecture and lower FLOPs for model deployment on limited hardware [
38,
39]. Such improvement is particularly relevant for large-scale phenotyping workflows, where model throughput and hardware limitations directly affect scalability. By maintaining comparable accuracy with a smaller computational footprint, YOLOv11 demonstrates clear advantages for real-time phenotyping and embedded system deployment in digital agriculture.
However, comparison with physically measured data revealed several limitations of the image-based approach. The updated analysis showed Pearson’s r values ranging from 0.20 to 0.44 and R² values mostly between 0.72 and 0.83 across traits, indicating that YOLO-derived measurements captured the general morphological trends of both P. ostreatus and F. velutipes. In particular, the pileus diameter and stipe traits exhibited stronger alignment with physical measurements, while pileus and stipe thickness remained more variable due to occlusion and surface curvature effects. Importantly, inter-model consistency remained exceptionally high (r ≥ 0.94, R² ≥ 0.96, MAE ≤ 0.40), confirming that YOLOv11’s improved computational efficiency did not compromise its interpretive reliability or dimensional prediction capability.
These results collectively demonstrate that YOLOv11 not only enhanced training efficiency but also retained analytical equivalence with YOLOv8 in terms of real–physical correlation, maintaining comparable r, R², and MAE values across all traits. This indicates that YOLOv11’s architectural optimizations primarily affect computation speed and model convergence rather than altering the biological validity of the predictions. Such stability is particularly valuable for digital phenotyping pipelines that require high-throughput analysis without loss of interpretability.
Overall, these results emphasize that YOLOv11 and YOLOv8 perform equivalently in predictive accuracy but differ in computational and training efficiency. YOLOv11 offers a refined balance between speed, stability, and accuracy, maintaining biological interpretability while significantly reducing computational burden. This makes YOLOv11 particularly suitable for lightweight, real-time phenotyping systems where rapid and scalable processing is required. The findings also underscore that improvements in deep-learning models for phenotyping should not only prioritize accuracy but also consider computational sustainability and integration into field-deployable digital breeding platforms.
5. Conclusions
This study compared the performance of YOLOv8 and YOLOv11 segmentation models for the efficient and automated phenotypic analysis of Pleurotus ostreatus and Flammulina velutipes. Unlike traditional phenotyping approaches that require harvesting individual fruiting bodies, the present work established a non-destructive analytic pipeline that extracts morphological traits directly from bottle cultivation images. Both models accurately detected and segmented the pileus and stipe structures from side-view images, providing consistent performance across two morphologically distinct species. The quantitative results demonstrated that YOLOv11 achieved segmentation accuracy comparable to YOLOv8 (ΔmAP50–95 < 0.01), while markedly improving computational efficiency by reducing FLOPs, parameter counts, and gradient loads by approximately 20–30%. The integration of the C2f-Fusion and RepNCSPELAN modules in YOLOv11 enhanced feature reuse and gradient stability, leading to faster convergence and lower loss values during training.
Comparison with caliper-based physical measurements confirmed that both YOLOv8 and YOLOv11 produced biologically meaningful and moderately strong correlations (r ≈ 0.2–0.44; R² ≈ 0.72–0.83) across major pileus and stipe traits, indicating that the non-destructive bottle-based predictions capture the essential morphological variation observed in physical measurements. Importantly, YOLOv11 maintained analytical reliability comparable to YOLOv8 (inter-model r ≥ 0.94, R² ≥ 0.96, MAE ≤ 0.40), demonstrating that efficiency improvements did not compromise measurement accuracy or biological interpretability.
Overall, this study demonstrates that YOLOv11 provides a robust, efficient, and non-destructive framework for high-throughput phenotypic analysis directly within bottle-cultivation systems, eliminating the need to harvest fruiting bodies for measurement. By maintaining equivalent analytical precision with reduced processing cost, YOLOv11 represents a practical and scalable solution for real-time phenotyping applications in digital agriculture. This study highlights the potential of next-generation YOLO architectures to advance AI-based phenotyping pipelines, supporting faster, more reliable, and environmentally sustainable breeding and quality-assessment systems in modern agricultural science.
Institutional Review Board Statement:
Supplementary Materials
The supplementary data supporting the findings of this study are available in Zenodo at
https://doi.org/10.5281/zenodo.17933889. Table S1: Quantitative comparison of predicted and measured phenotypic traits in
Pleurotus ostreatus. Table S2: Quantitative comparison of predicted and measured phenotypic traits in
Flammulina velutipes.
Author Contributions:
Funding
This work was supported by the Devolepment of core mushroom resources and safety preservation technology, the postdoctoral research program support project (PJ01716801) as part of the results conducted by the Rural Development Administration (PJ01733103).
Data Availability Statement
The data presented in this study are available on request from the corresponding author.
Acknowledgments:.
Conflicts of Interest
The authors declare no conflict of interest..
References
- Morais, M.H.; Ramos, A.C.; Matos, N.; Santos-Oliveira, E.J. Production of shiitake mushroom (Lentinus edodes) on ligninocellulosic residues. Food Sci. Technol. Int. 2000, 6, 123–128. [Google Scholar] [CrossRef]
- Sánchez, C. Modern aspects of mushroom culture technology. Appl. Microbiol. Biotechnol. 2004, 64, 756–762. [Google Scholar] [CrossRef]
- Chang, S.-T.; Miles, P.G. Mushrooms: Cultivation, Nutritional Value, Medicinal Effect, and Environmental Impact, 2nd ed.; CRC Press: Boca Raton, FL, USA, 2004. [Google Scholar]
- Chang, S.-T.; Hayes, W.A. The Biology and Cultivation of Edible Mushrooms; Academic Press: Massachusetts, USA, 2013; pp. 3–9. [Google Scholar]
- Suarez, E.; Blaser, M.; Sutton, M. Automating leaf area measurement in citrus: Development and validation of a Python-based tool. Appl. Sci. 2025, 15, 9750. [Google Scholar] [CrossRef]
- Cheong, J.C.; Lee, C.J.; Suh, J.S.; Moon, Y.H. Comparison of physico-chemical and nutritional characteristics of pre-inoculation and post-harvest Flammulina velutipes media. J. Mushroom Sci. Prod. 2012, 10, 174–178. [Google Scholar]
- Cheong, J.C.; Lee, C.J.; Moon, J.W. Comprehensive model for medium composition for mushroom bottle cultivation. J. Mushrooms 2016, 14, 111–118. [Google Scholar] [CrossRef]
- Sapkota, R.; Karkee, M. Object detection with multimodal large vision-language models: An in-depth review. Inf. Fusion 2025, 126, 103575. [Google Scholar] [CrossRef]
- Wei, Z.; Wang, J.; You, H.; Ji, R.; Wang, F.; Shi, L.; Yu, H. A lightweight context-aware framework for toxic mushroom detection in complex ecological environments. Ecol. Inform. 2025, 90, 103256. [Google Scholar] [CrossRef]
- Dhanya, V.G.; Subeesh, A.; Kushwaha, N.L.; Vishwakarma, D.K.; Kumar, T.N.; Ritika, G.; Singh, A.N. Deep learning-based computer vision approaches for smart agricultural applications. Artif. Intell. Agric. 2022, 6, 211–229. [Google Scholar] [CrossRef]
- Hafiz, A.M.; Bhat, G.M. A survey on instance segmentation: State of the art. Int. J. Multimed. Inf. Retr. 2020, 9, 171–189. [Google Scholar] [CrossRef]
- Coulibaly, S.; Kamsu-Foguem, B.; Kamissoko, D.; Traore, D. Deep learning for precision agriculture: A bibliometric analysis. Intell. Syst. Appl. 2022, 16, 200102. [Google Scholar] [CrossRef]
- Sapkota, R.; Ahmed, D.; Karkee, M. Comparing YOLOv8 and Mask R-CNN for instance segmentation in complex orchard environments. Artif. Intell. Agric. 2024, 13, 84–99. [Google Scholar] [CrossRef]
- Rashid, J.; Khan, I.; Ali, G.; Alturise, F.; Alkhalifah, T. Real-time multiple guava leaf disease detection from a single leaf using a hybrid deep learning technique. Comput. Mater. Continua 2023, 74, 1–15. [Google Scholar] [CrossRef]
- Maji, A.K.; Marwaha, S.; Kumar, S.; Arora, A.; Chinnusamy, V.; Islam, S. SlypNet: Spikelet-based yield prediction of wheat using advanced plant phenotyping and computer vision techniques. Front. Plant Sci. 2022, 13, 889853. [Google Scholar] [CrossRef]
- Sapkota, R.; Karkee, M. Comparing YOLOv11 and YOLOv8 for instance segmentation of occluded and non-occluded immature green fruits in complex orchard environment. arXiv 2025, arXiv:2410.19869. [Google Scholar]
- Xie, L.; Jing, J.; Wu, H.; Kang, Q.; Zhao, Y.; Ye, D. MPG-YOLO: Enoki mushroom precision grasping with segmentation and pulse mapping. Agronomy 2025, 15, 432. [Google Scholar] [CrossRef]
- Qi, W.; Chen, H.; Zheng, X.; Zhang, T.; Liu, Y. Detection and classification of shiitake mushroom fruiting bodies based on Mamba YOLO. Sci. Rep. 2025, 15, 133. [Google Scholar] [CrossRef] [PubMed]
- Redmon, J. You only look once: Unified, real-time object detection. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 2016; IEEE. [Google Scholar]
- Xu, X.; Li, J.; Zhou, J.; Feng, P.; Yu, H.; Ma, Y. Three-dimensional reconstruction, phenotypic traits extraction, and yield estimation of shiitake mushrooms based on structure from motion and multi-view stereo. Agriculture 2025, 15, 298. [Google Scholar] [CrossRef]
- Khan, A.T.; Jensen, S.M. LEAF-Net: A unified framework for leaf extraction and analysis in multi-crop phenotyping using YOLOv11. Agriculture 2025, 15, 196. [Google Scholar] [CrossRef]
- Ho Bao Thuy, Q.; Suzuki, A. Technology of mushroom cultivation. Viet. J. Sci. Technol. 2019, 57, 265–286. [Google Scholar]
- Badgujar, C.M.; Poulose, A.; Gan, H. Agricultural object detection with YOLO: A bibliometric and systematic review. Comput. Electron. Agric. 2024, 223, 109090. [Google Scholar] [CrossRef]
- Zakeri, R.; Zamani, A.; Taghizadeh, A.; Abbaszadeh, M.; Saadatfar, B. M18K: A comprehensive RGB-D dataset and benchmark for mushroom detection and instance segmentation. arXiv 2024, arXiv:2407.11275. [Google Scholar]
- Abdullah, A.; Amran, G.A.; Tahmid, S.M.A.; Alabrah, A.; Al-Bakhrani, A.A.; Ali, A. Deep-learning-based detection of diseased tomato leaves. Agronomy 2024, 14, 1593. [Google Scholar] [CrossRef]
- Wang, C.; Li, H.; Deng, X.; Liu, Y.; Wu, T.; Liu, W.; Xiao, R.; Wang, Z.; Wang, B. Improved YOLOv8 model for precision detection of tea leaves. Agriculture 2024, 14, 2324. [Google Scholar] [CrossRef]
- Wang, N.; Liu, H.; Li, Y.; Zhou, W.; Ding, M. Segmentation and phenotype calculation of rapeseed pods using YOLOv8 and Mask R-CNN. Plants 2023, 12, 3328. [Google Scholar] [CrossRef]
- Solimani, F.; Cardellicchio, A.; Dimauro, G.; Petrozza, A.; Summerer, S.; Cellini, F.; Renò, A. Optimizing tomato plant phenotyping using an enhanced YOLOv8 architecture. Comput. Electron. Agric. 2024, 218, 108728. [Google Scholar] [CrossRef]
- Wu, C.; Zhang, S.; Wang, W.; Wu, Z.; Yang, S.; Chen, W. Phenotypic parameter computation using YOLOv11-DYPF keypoint detection. Aquac. Eng. 2025, 111, 102571. [Google Scholar] [CrossRef]
- Sanchez, S.A.; Romero, H.J.; Morales, A.D. Comparison of performance metrics of pretrained object detection models. IOP Conf. Ser. Mater. Sci. Eng. 2020, 844, 012024. [Google Scholar] [CrossRef]
- Murat, A.A.; Kiran, M.S. A comprehensive review on YOLO versions for object detection. Eng. Sci. Technol. Int. J. 2025, 70, 102161. [Google Scholar] [CrossRef]
- Lu, C.P.; Liaw, J.J.; Wu, T.C.; Hung, T.F. Development of a mushroom growth measurement system applying deep learning. Agronomy 2019, 9, 32. [Google Scholar] [CrossRef]
- Frossard, E.; Liebisch, F.; Hgaza, V.K.; Kiba, D.I.; Kirchgessner, N.; Müller, L.; Müller, P.; Pouya, N.; Ringger, C.; Walter, A. Image-based phenotyping of water yam growth and nitrogen status. Agronomy 2021, 11, 249. [Google Scholar] [CrossRef]
- Shi, Y.; Zhang, C.; Sun, Z.; Liu, J.; Li, B. OMC-YOLO: A lightweight grading detection method for oyster mushrooms. Horticulturae 2024, 10, 742. [Google Scholar] [CrossRef]
- He, L.H.; Zhou, Y.Z.; Liu, L.; Cao, W.; Ma, J.H. Research on object detection and recognition in remote sensing images based on YOLOv11. Sci. Rep. 2025, 15, 14032. [Google Scholar] [CrossRef]
- Mihajlovic, M.; Stojanovic, A.; Petrovic, S. Enhancing instance segmentation in high-resolution aerial imagery with YOLOv11s-Seg. Mathematics 2025, 13, 3079. [Google Scholar] [CrossRef]
- Su, C.; Lin, H.; Wang, D. Nav-YOLO: A lightweight and efficient object detection method for edge devices. ISPRS Int. J. Geo-Inf. 2025, 14, 364. [Google Scholar] [CrossRef]
- Padilla, R.; Netto, S.; Da Silva, E. Performance metrics for object detection algorithms. Electronics 2020, 9, 279. [Google Scholar]
- Long, X.; Deng, K.; Wang, G.; Zhang, Y. PP-YOLO: An effective and efficient implementation of object detector. arXiv 2020, arXiv:2007.12099. [Google Scholar] [CrossRef]
- Lu, C.P.; Cheng, S.H.; Hsiao, Y.T. Development of a mushroom growth measurement system using image processing. Agronomy 2019, 9, 32. [Google Scholar] [CrossRef]
- Kiba, D.I.; Ofori, E.; Bationo, A. Image-based phenotyping methods for measuring water yam growth and nitrogen nutritional status. Agronomy 2021, 11, 1529. [Google Scholar]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).