Submitted:
15 December 2025
Posted:
16 December 2025
You are already at the latest version
Abstract
Keywords:
Introduction
Genomics Capacity
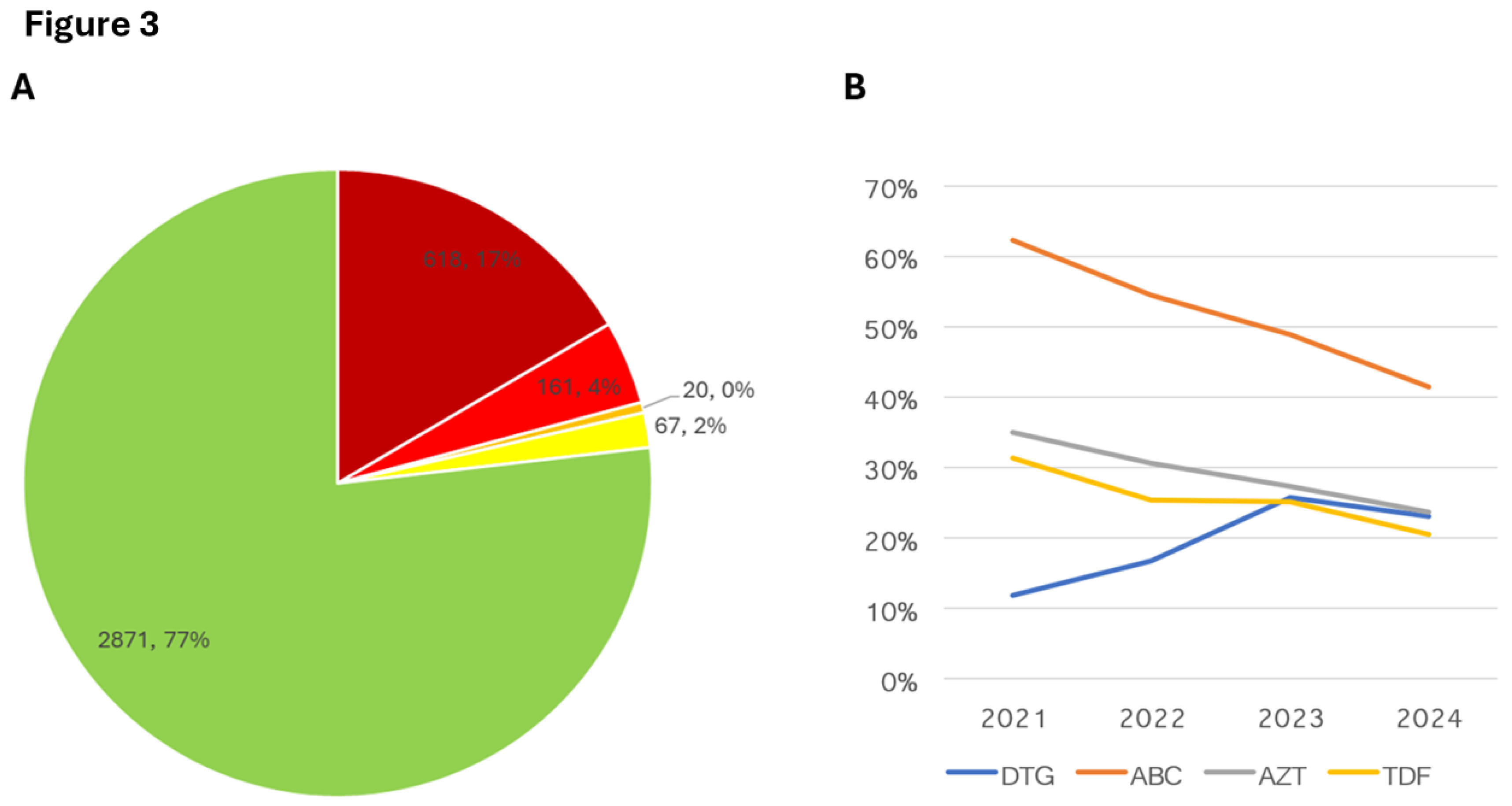
SARS-CoV-2 Variant Surveillance
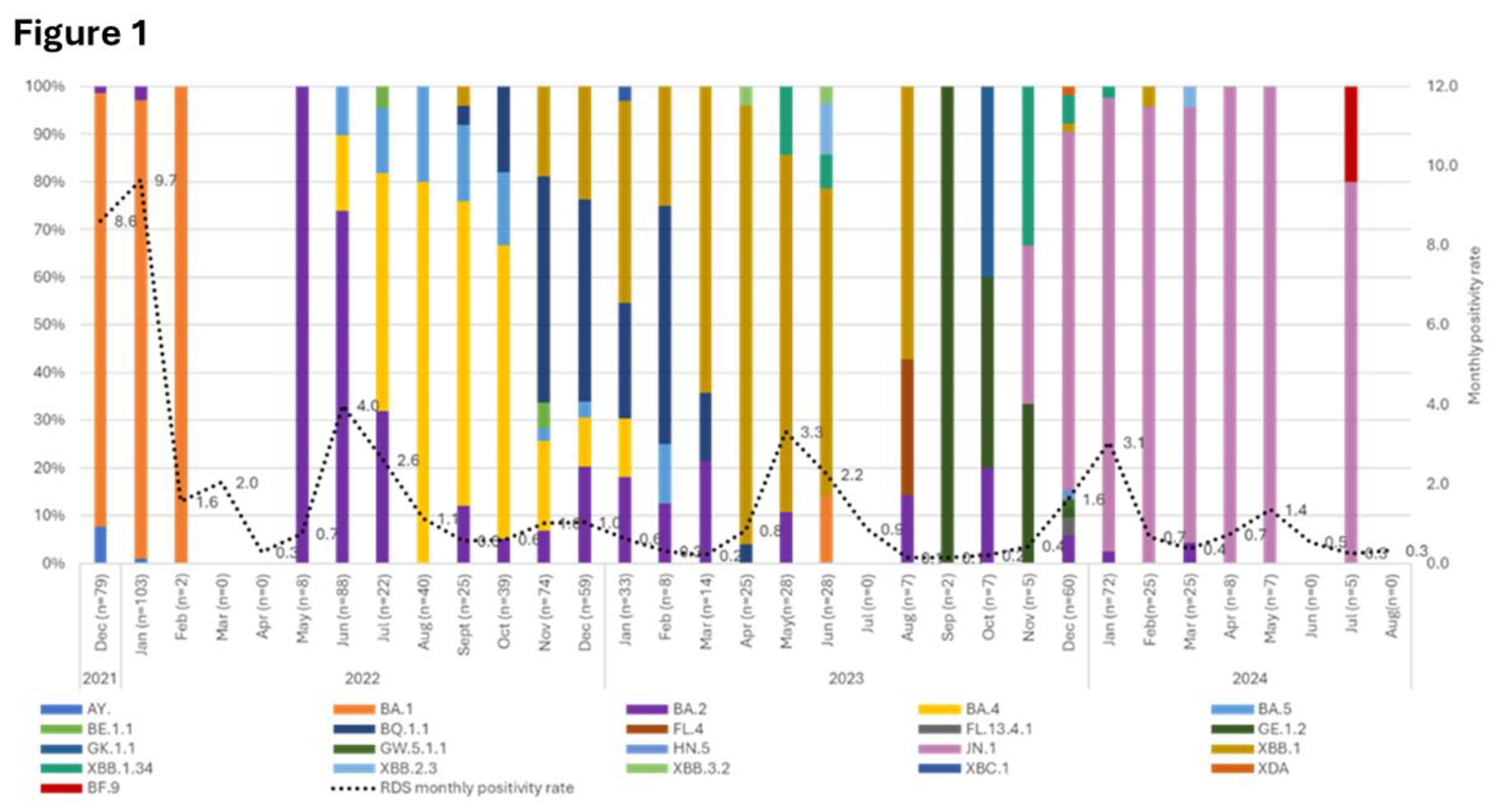
Genomics in Wastewater-Based Surveillance
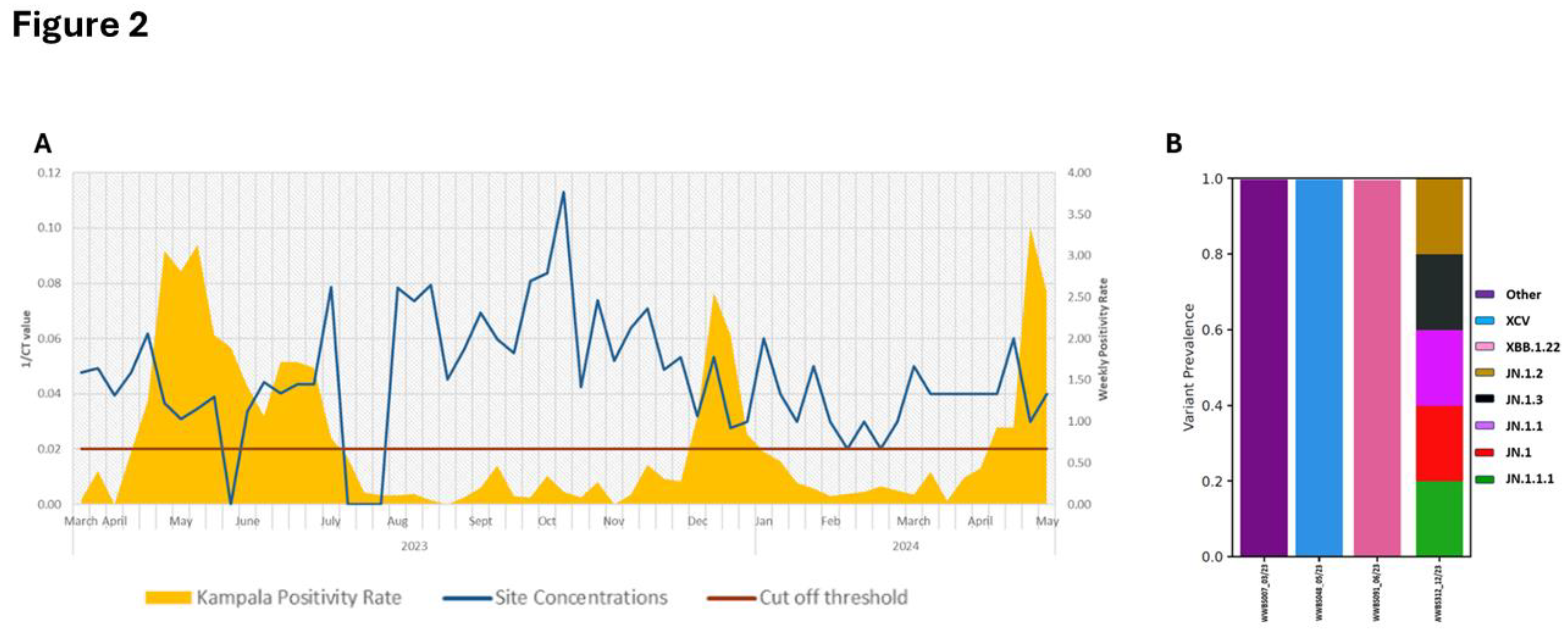
Ebola Sudan Outbreak Investigation
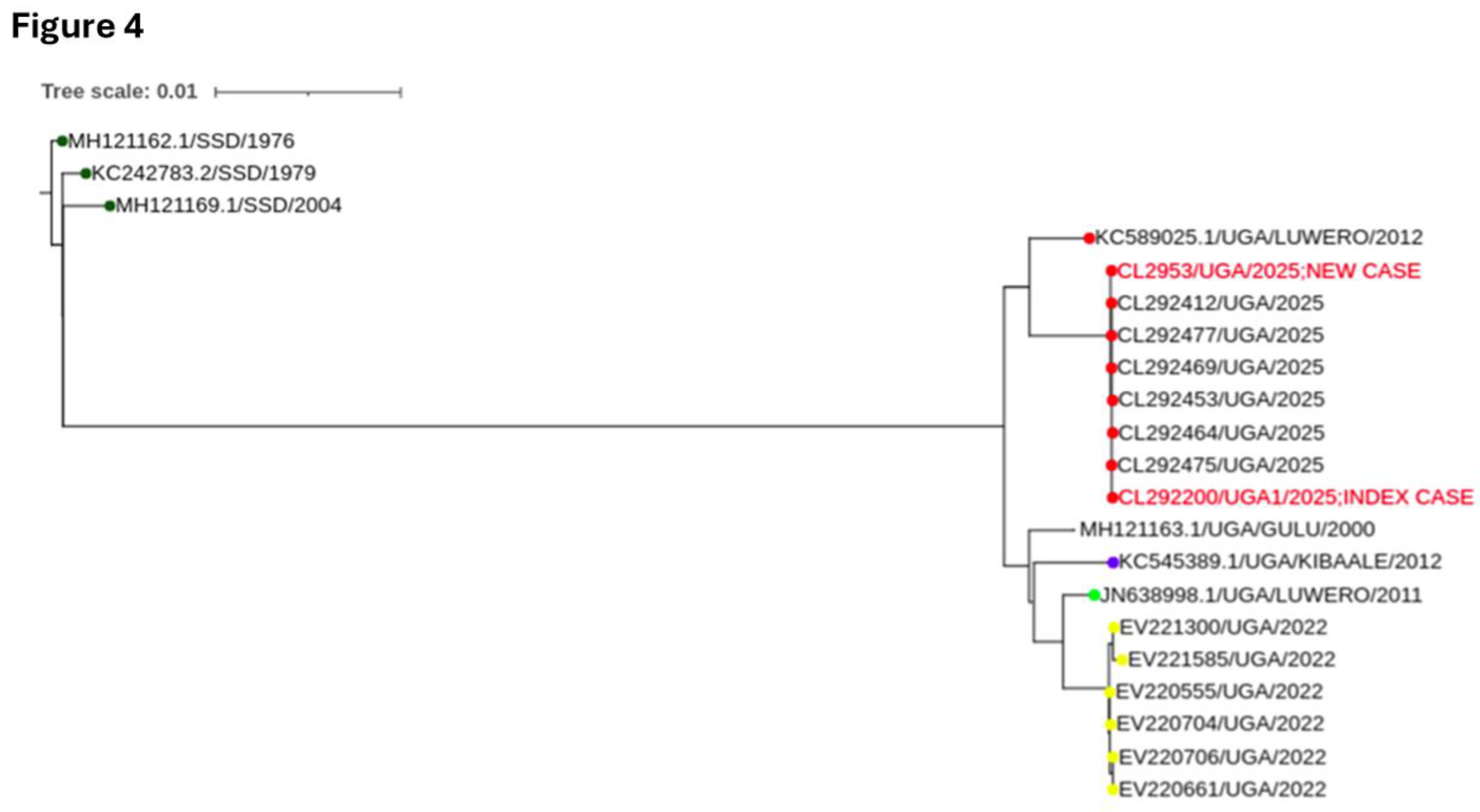
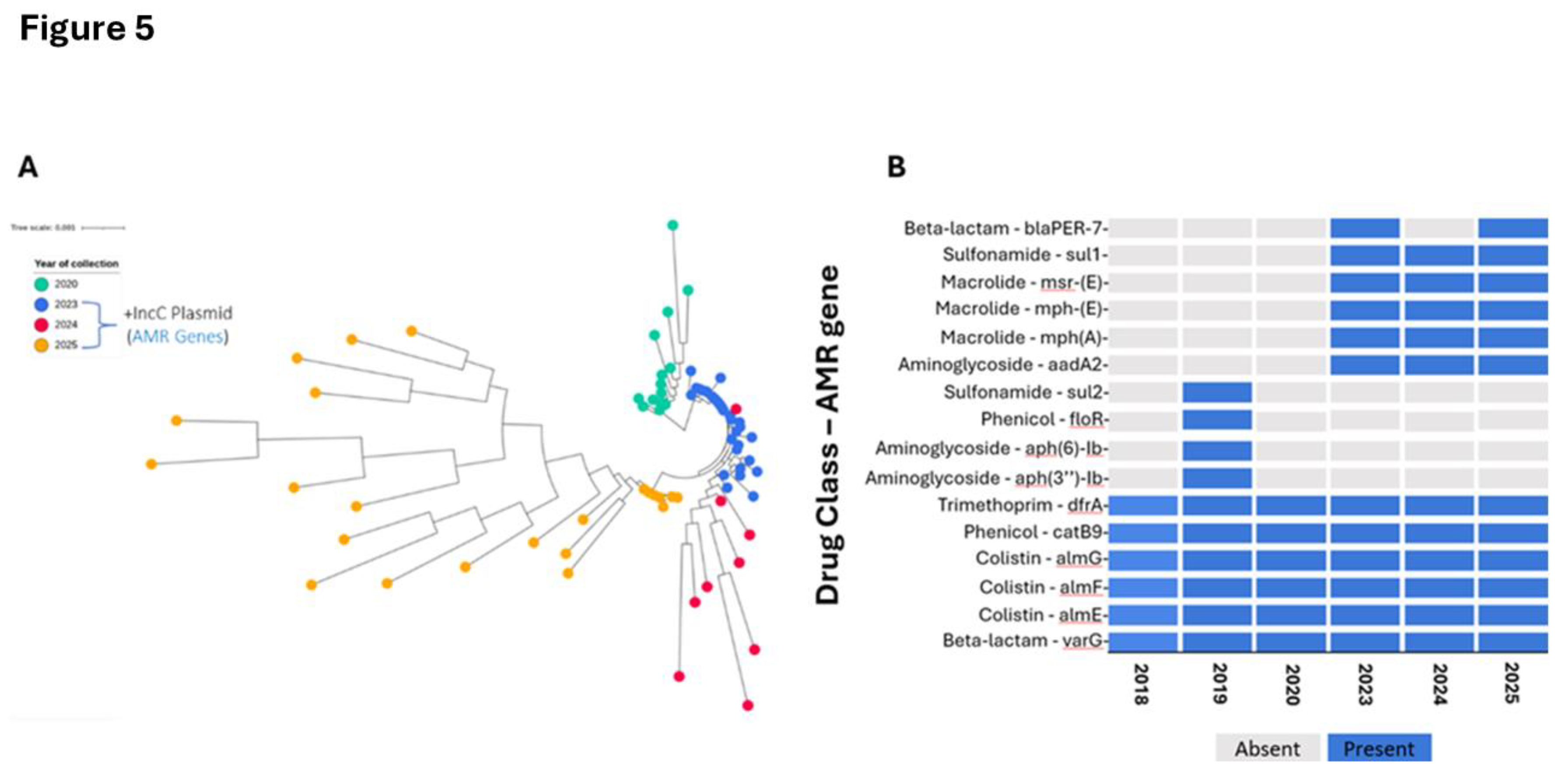
Cholera Outbreak Investigation
Genomics in Outbreak Response of Unknown Aetiologies
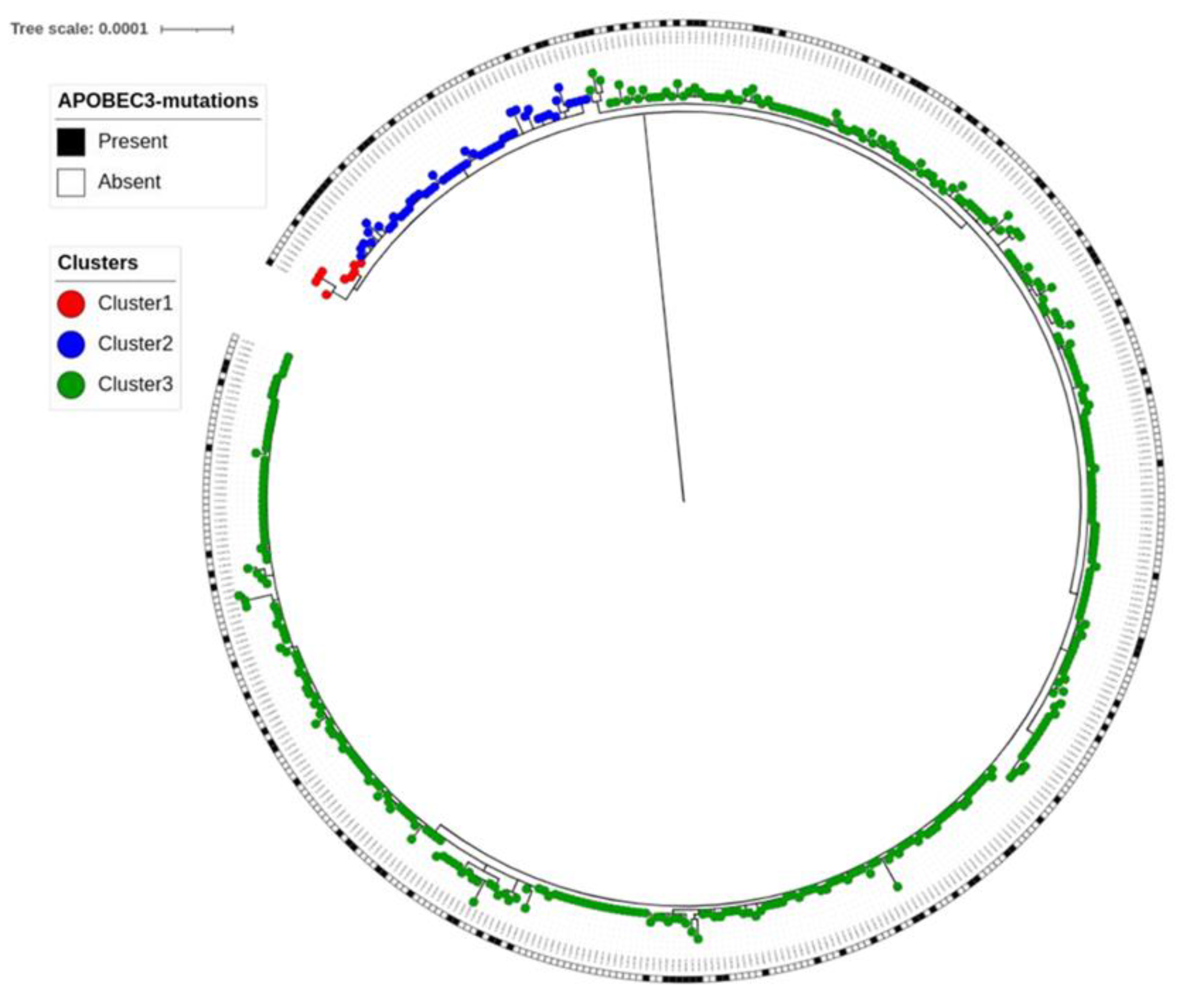
Genomics in Real-Time Mpox Outbreak Management
Genomics in Malaria Surveillance
Genomics for Liquid Biopsy in Oncology Diagnosis
Discussion and Conclusion
Ethical Considerations
Acknowledgments
Author Contributions
Data Availability
References
- Murray CJL. The Global Burden of Disease Study at 30 years. Nat Med (2022) 28:2019–2026. [CrossRef]
- Max Roser, Hannah Ritchie, Fiona Spooner. Burden of Disease. Our World in Data (2021).
- Mremi IR, George J, Rumisha SF, Sindato C, Kimera SI, Mboera LEG. Twenty years of integrated disease surveillance and response in Sub-Saharan Africa: challenges and opportunities for effective management of infectious disease epidemics. One Health Outlook (2021) 3:22. [CrossRef]
- Grad YH, Lipsitch M. Epidemiologic data and pathogen genome sequences: a powerful synergy for public health. Genome Biol (2014) 15:538. [CrossRef]
- Gardy JL, Loman NJ. Towards a genomics-informed, real-time, global pathogen surveillance system. Nat Rev Genet (2018) 19:9–20. 10.1038/nrg.2017.88Armstrong GL, Maccannell DR, Taylor J, Carleton HA, Neuhaus EB, Bradbury RS, Posey JE, Gwinn M, Summary MPH. Pathogen Genomics in Public Health. N Engl J Med (2019) 381:2569–2580. [CrossRef]
- Armstrong GL, Maccannell DR, Taylor J, Carleton HA, Neuhaus EB, Bradbury RS, Posey JE, Gwinn M, Summary MPH. Pathogen Genomics in Public Health. N Engl J Med (2019) 381:2569–2580. [CrossRef]
- Baker KS, Jauneikaite E, Hopkins KL, Lo SW, Sánchez-Busó L, Getino M, Howden BP, Holt KE, Musila LA, Hendriksen RS, et al. Genomics for public health and international surveillance of antimicrobial resistance. Lancet Microbe (2023) 4:e1047–e1055. [CrossRef]
- Inzaule SC, Tessema SK, Kebede Y, Ogwell Ouma AE, Nkengasong JN. Genomic-informed pathogen surveillance in Africa: opportunities and challenges. Lancet Infect Dis (2021) 21: e281–e289. 10.1016/S1473-3099(20)30939-7Onywera H, Ondoa P, Nfii F, Ogwell A, Kebede Y, Christoffels A, Tessema SK. Boosting pathogen genomics and bioinformatics workforce in Africa. Lancet Infect Dis (2024) 24: e106–e112. [CrossRef]
- Onywera H, Ondoa P, Nfii F, Ogwell A, Kebede Y, Christoffels A, Tessema SK. Boosting pathogen genomics and bioinformatics workforce in Africa. Lancet Infect Dis (2024) 24:e106–e112. [CrossRef]
- Kanyerezi S, Ayitewala A, Ssemaganda A, Makoha C, Kabahita JM, Murungi M, Tusabe GW, Tenywa W, Oundo HR, Sseruyange J, et al. Near Real-Time Genomic Characterization of the 2025 Sudan Ebolavirus Outbreak in Uganda’s Index Case: Insights into Evolutionary Origins. https://virological.org/t/near-real-time-genomic-characterization-of-the-2025-sudan-ebolavirus-outbreak-in-uganda-s-index-case-insights-into-evolutionary-origins/990 (2025).
- Kanyerezi S, Sserwadda I, Ssemaganda A, Seruyange J, Ayitewala A, Oundo HR, Tenywa W, Kagurusi BA, Tusabe G, Were S, et al. HIV-DRIVES: HIV drug resistance identification, variant evaluation, and surveillance pipeline. Access Microbiol (2024) 6:000815. v3. [CrossRef]
- Africa Centres for Disease Control. Integrated Genomic Surveillance for Outbreak Detection (DETECT). The Africa Pathogen Genomics Initative (2024).
- World Health Organization. WHO chief declares end to COVID-19 as a global health emergency. https://news.un.org/en/story/2023/05/1136367 (2023).
- World Health Organization. Tracking SARS-CoV-2 variants. https://www.who.int/activities/tracking-SARS-CoV-2-variants/ (2024).
- World Health Organization. JN.1 Initial Risk Evaluation. https://www.who.int/docs/default-source/coronaviruse/18122023_jn1_ire_clean.pdf?sfvrsn=6103754a_3 (2023).
- Naveed Siddiqui A, Musharaf I, Gulumbe BH. The JN.1 variant of COVID-19: immune evasion, transmissibility, and implications for global health. Ther Adv Infect Dis (2025) 12:. [CrossRef]
- Manyana S, Gounder L, Pillay M, Manasa J, Naidoo K, Chimukangara B. HIV-1 Drug Resistance Genotyping in Resource Limited Settings: Current and Future Perspectives in Sequencing Technologies. Viruses (2021) 13:. [CrossRef]
- Aboulker J-P. Drug-resistant genotyping in HIV-1 therapy. The Lancet (1999) 354:1120. [CrossRef]
- World Health Organization. Updated recommendations on first line and second-line antiretroviral regimens and post-exposure prophylaxis and recommendations on early infant diagnosis of HIV: interim guidelines: supplement to the 2016 consolidated guidelines on the use of antiretroviral drugs for treating and preventing HIV infection. (2018).
- Phillips AN, Venter F, Havlir D, Pozniak A, Kuritzkes D, Wensing A, Lundgren JD, De Luca A, Pillay D, Mellors J, et al. Risks and benefits of dolutegravir-based antiretroviral drug regimens in sub-Saharan Africa: a modelling study. Lancet HIV (2019) 6: e116–e127. [CrossRef]
- Vitoria M, Hill A, Ford N, Doherty M, Clayden P, Venter F, Ripin D, Flexner C, Domanico PL. The transition to dolutegravir and other new antiretrovirals in low-income and middle-income countries: What are the issues? AIDS (2018) 32:1551–1561. [CrossRef]
- Ayitewala A, Ssewanyana I, Kiyaga C. Next generation sequencing based in-house HIV genotyping method: validation report. AIDS Res Ther (2021) 18:. [CrossRef]
- Parkin NT, Avila-Rios S, Bibby DF, Brumme CJ, Eshleman SH, Harrigan PR, Howison M, Hunt G, Ji H, Kantor R, et al. Multi-Laboratory Comparison of Next-Generation to Sanger-Based Sequencing for HIV-1 Drug Resistance Genotyping. Viruses (2020) 12:. [CrossRef]
- Ministry of Health. Consolidated Guidelines for the Prevention and Treatment of HIV and AIDS in Uganda. (2022).
- Abdullah W, W BP, Kenneth K, Kayiira M, Hellen A-T, Daniel K, T SM, D KJ, Peter W, R BR, et al. Sudan Virus Disease among Health Care Workers, Uganda, 2022. New England Journal of Medicine (2024) 391:285–287. [CrossRef]
- Kyobe Bosa Henry, Wayengera Misaki, Nabadda Suzan, Olaro Charles, Bahatungire Rony, Kalungi Samuel, Bakamutumaho Barnabas, Muruta Allan, Kagirita Atek, Muwonge Haruna, et al. Sudan Ebola virus disease outbreak in Uganda a role for cryptic transmission. Nat Med (2025).
- Tanui CK, Tessema SK, Tegegne MA, Tebeje YK, Kaseya J. Unlocking the power of molecular and genomics tools to enhance cholera surveillance in Africa. Nat Med (2023) 29:2387–2388. [CrossRef]
- Africa Centres for Disease Control and Prevention. Scientists Call for Broader and Stronger Network for Genomic Surveillance of Cholera in Africa. https://africacdc.org/news-item/scientists-call-for-broader-and-stronger-network-for-genomic-surveillance-of-cholera-in-africa/# (2024).
- De R. Mobile Genetic Elements of Vibrio cholerae and the Evolution of Its Antimicrobial Resistance. Frontiers in Tropical Diseases (2021) 2:. [CrossRef]
- Mboowa G, Matteson NL, Tanui CK, Kasonde M, Kamwiziku GK, Akanbi OA, Chitio JJE, Kagoli M, Essomba RG, Ayitewala A, et al. Multicountry genomic analysis underscores regional cholera spread in Africa. (2024). [CrossRef]
- ARTIC Network Project. artic-mpxv-illumina-nf. (2025).
- Jahankir MJB, Sounderrajan V, Rao SS, Thangam T, Kamariah N, Kurumbati A, Harshavardhan S, Ashwin A, Jeyaraj S, Parthasarathy K. Accelerated mutation by host protein APOBEC in Monkeypox virus. Gene Rep (2024) 34:101878. [CrossRef]
- Suspène R, Raymond KA, Boutin L, Guillier S, Lemoine F, Ferraris O, Tournier JN, Iseni F, Simon-Lorière E, Vartanian JP. APOBEC3F Is a Mutational Driver of the Human Monkeypox Virus Identified in the 2022 Outbreak. Journal of Infectious Diseases (2023) 228:1421–1429. [CrossRef]
- World Health Organization. WHO guidelines for malaria. WHO Global Malaria Programme. (2021).
- National Malaria Control Division Ministry of Health Uganda. Malaria reduction and elimination strategic plan: 2021-2025. (2021).
- Agaba BB, Travis J, Smith D, Rugera SP, Zalwango MG, Opigo J, Katureebe C, Mpirirwe R, Bakary D, Antonio M, et al. Emerging threat of artemisinin partial resistance markers (pfk13 mutations) in Plasmodium falciparum parasite populations in multiple geographical locations in high transmission regions of Uganda. Malar J (2024) 23:330. [CrossRef]
- Conrad MD, Asua V, Garg S, Giesbrecht D, Niaré K, Smith S, Namuganga JF, Katairo T, Legac J, Crudale RM, et al. Evolution of Partial Resistance to Artemisinins in Malaria Parasites in Uganda. New England Journal of Medicine (2023) 389:722–732. [CrossRef]
- Kiyaga, S., Mbabazi, M., Katairo, T., Kabbale, K.D., Asua, V., Nsengimaana, B., Wiringilimaana, I., Semakuba, F.Ddumba., Mwubaha, C., Nakasaanya, J., et al. (2025). Accuracy of Plasmodium falciparum genetic data for estimating parasite prevalence and malaria incidence in Uganda. Preprint. [CrossRef]
- Tukwasibwe S, Garg S, Katairo T, Asua V, Kagurusi BA, Mboowa G, Crudale R, Tumusiime G, Businge J, Alula D. Varied Prevalence of antimalarial drug resistance markers in different populations of newly arrived refugees in Uganda. J Infect Dis (2024) 230:497–504.
- Aranda-Díaz A, Neubauer Vickers E, Murie K, Palmer B, Hathaway N, Gerlovina I, Boene S, García-Ulloa M, Cisteró P, Katairo T, et al. Sensitive and modular amplicon sequencing of Plasmodium falciparum diversity and resistance for research and public health. Sci Rep (2025) 15:10737. [CrossRef]
- Aydemir O, Janko M, Hathaway NJ, Verity R, Mwandagalirwa MK, Tshefu AK, Tessema SK, Marsh PW, Tran A, Reimonn T, et al. Drug-Resistance and Population Structure of Plasmodium falciparum Across the Democratic Republic of Congo Using High-Throughput Molecular Inversion Probes. J Infect Dis (2018) 218:946–955. [CrossRef]
- Katairo T, Asua V, Nsengimaana B, Tukwasibwe S, Semakuba FD, Wiringilimaana I, Kagurusi BA, Mwubaha C, Nakasaanya J, Garg S. Performance of molecular inversion probe DR23K and Paragon MAD4HatTeR Amplicon sequencing panels for detection of Plasmodium falciparum mutations associated with antimalarial drug resistance. Malar J (2025) 24:188.
- Agaba BB, Anderson K, Gresty K, Prosser C, Smith D, Nankabirwa JI, Nsobya S, Yeka A, Opigo J, Gonahasa S, et al. Molecular surveillance reveals the presence of pfhrp2 and pfhrp3 gene deletions in Plasmodium falciparum parasite populations in Uganda, 2017–2019. Malar J (2020) 19:300. [CrossRef]
- Agaba, Bosco B, Nankabirwa JI, Yeka A, Nsobya S, Gresty K, Anderson K, Mbaka P, Prosser C, Smith D, et al. Limitations of rapid diagnostic tests in malaria surveys in areas with varied transmission intensity in Uganda 2017-2019: Implications for selection and use of HRP2 RDTs. PLoS One (2021) 15: e0244457-. [CrossRef]
- Agaba BB, Smith D, Travis J, Pasay C, Nabatanzi M, Arinaitwe E, Ssewanyana I, Nabadda S, Cunningham J, Kamya MR, et al. Limited threat of Plasmodium falciparum pfhrp2 and pfhrp3 gene deletion to the utility of HRP2-based malaria RDTs in Northern Uganda. Malar J (2024) 23:3. [CrossRef]
- Kabbale KD, Nsengimaana B, Semakuba FD, Kagurusi BA, Mwubaha C, Wiringilimaana I, Katairo T, Kiyaga S, Mbabazi M, Gonahasa S. Field evaluation of the Bioline Malaria Ag Pf/Pan rapid diagnostic test causes of microscopy discordance and performance in Uganda. Malar J (2025) 24:138.
- Ogwang MD, Zhao W, Ayers LW, Mbulaiteye SM. Accuracy of Burkitt Lymphoma Diagnosis in Constrained Pathology Settings: Importance to Epidemiology. Arch Pathol Lab Med (2011) 135:445–450. [CrossRef]
- Mawalla WF, Morrell L, Chirande L, Achola C, Mwamtemi H, Sandi G, Mahawi S, Kahakwa A, Ntemi P, Hadija N, et al. Treatment delays in children and young adults with lymphoma: a report from an East Africa lymphoma cohort study. Blood Adv (2023) 7:4962–4965. [CrossRef]
- Chamba CC, Howard K, Mawalla WF, Christopher H, Seruyange J, Josephat E, Legason ID, Ogwang M, Ayitewala A, Mremi A, et al. Performance of a Liquid Biopsy Diagnostic Prediction Model for EBV-Positive Burkitt Lymphoma in Sub-Saharan Africa. Blood (2023) 142:4364. [CrossRef]
| Lab ID | Pathogen Detected |
|---|---|
| RPGS-222 | Rotavirus A; Enterovirus (Coxsackieviruses) |
| RPGS-223 | Rotavirus A; Enterovirus (Coxsackieviruses) |
| RPGS-224 | Rotavirus A; Enterovirus (Coxsackieviruses) |
| RPGS-225 | Rotavirus A; Enterovirus (Coxsackieviruses) |
| RPGS-226 | Enterovirus (Coxsackieviruses) |
| RPGS-227 | Rotavirus A; Enterovirus (Coxsackieviruses) |
| RPGS-228 | Enterovirus (Coxsackieviruses) |
| RPGS-230 | Enterovirus (Coxsackieviruses) |
| RPGS-231 | Rotavirus A; Enterovirus (Coxsackieviruses) |
| RPGS-232 | Enterovirus (Coxsackieviruses) |
| Exhibit material | Sample id | Culture | Genomics |
|---|---|---|---|
| Cooked posho | D | NIL | Bacillus cereus |
| C | NIL | Bacillus cereus | |
| Cooked beans | A | Bacillus cereus | Bacillus cereus |
| Uncooked Posho flour | N3 | Bacillus cereus | NIL |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).