Submitted:
24 January 2026
Posted:
27 January 2026
You are already at the latest version
Abstract
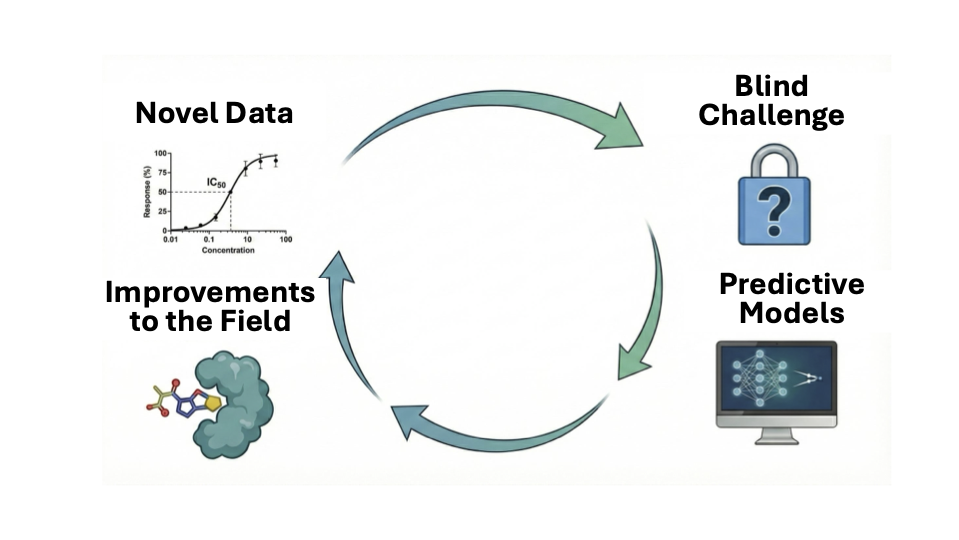
Keywords:
Drug Discovery Is Costly
Computer-Aided Drug Discovery (CADD) Holds Enormous Potential to Accelerate Progress
ADMET Properties and Anti-Target as a Focus for Blind Challenges
Blind Challenges Require Evergreen Data Generation Efforts
Funding
References
- Scannell, J. W.; Blanckley, A.; Boldon, H.; Warrington, B. Diagnosing the Decline in Pharmaceutical R&D Efficiency. Nat. Rev. Drug Discov. 2012, 11(3), 191–200. [Google Scholar] [CrossRef] [PubMed]
- Ringel, M. S.; Scannell, J. W.; Baedeker, M.; Schulze, U. Breaking Eroom’s Law. Nat. Rev. Drug Discov. 2020, 19(12), 833–834. [Google Scholar] [CrossRef] [PubMed]
- Erooms_law: Eroom’s Law; Github.
- Brown, F. K.; Sherer, E. C.; Johnson, S. A.; Holloway, M. K.; Sherborne, B. S. The Evolution of Drug Design at Merck Research Laboratories. J. Comput. Aided Mol. Des. 2017, 31(3), 255–266. [Google Scholar] [CrossRef] [PubMed]
- Shirts, M. R.; Mobley, D. L.; Brown, S. P. Free-Energy Calculations in Structure-Based Drug Design. Drug Design 2010, 1, 61–86. [Google Scholar]
- Retchin, M.; Wang, Y.; Takaba, K.; Chodera, J. D. DrugGym: A Testbed for the Economics of Autonomous Drug Discovery. bioRxiv 2024, 2024.05.28.596296. [Google Scholar] [CrossRef] [PubMed]
- Ahdritz, G.; Bouatta, N.; Floristean, C.; Kadyan, S.; Xia, Q.; Gerecke, W.; O’Donnell, T. J.; Berenberg, D.; Fisk, I.; Zanichelli, N.; Zhang, B.; Nowaczynski, A.; Wang, B.; Stepniewska-Dziubinska, M. M.; Zhang, S.; Ojewole, A.; Guney, M. E.; Biderman, S.; Watkins, A. M.; Ra, S.; Lorenzo, P. R.; Nivon, L.; Weitzner, B.; Ban, Y.-E. A.; Chen, S.; Zhang, M.; Li, C.; Song, S. L.; He, Y.; Sorger, P. K.; Mostaque, E.; Zhang, Z.; Bonneau, R.; AlQuraishi, M. OpenFold: Retraining AlphaFold2 Yields New Insights into Its Learning Mechanisms and Capacity for Generalization. Nat. Methods 2024, 21(8), 1514–1524. [Google Scholar] [CrossRef] [PubMed]
- Graber, D.; Stockinger, P.; Meyer, F.; Mishra, S.; Horn, C.; Buller, R. Resolving Data Bias Improves Generalization in Binding Affinity Prediction. Nat. Mach. Intell. 2025, 7(10), 1713–1725. [Google Scholar] [CrossRef] [PubMed]
- Bernett, J.; Blumenthal, D. B.; Grimm, D. G.; Haselbeck, F.; Joeres, R.; Kalinina, O. V.; List, M. Guiding Questions to Avoid Data Leakage in Biological Machine Learning Applications. Nat. Methods 2024, 21(8), 1444–1453. [Google Scholar] [CrossRef] [PubMed]
- Yin, J.; Henriksen, N. M.; Slochower, D. R.; Shirts, M. R.; Chiu, M. W.; Mobley, D. L.; Gilson, M. K. Overview of the SAMPL5 Host-Guest Challenge: Are We Doing Better? J. Comput. Aided Mol. Des. 2017, 31(1), 1–19. [Google Scholar] [CrossRef] [PubMed]
- Gaieb, Z.; Liu, S.; Gathiaka, S.; Chiu, M.; Yang, H.; Shao, C.; Feher, V. A.; Walters, W. P.; Kuhn, B.; Rudolph, M. G.; Burley, S. K.; Gilson, M. K.; Amaro, R. E. D3R Grand Challenge 2: Blind Prediction of Protein-Ligand Poses, Affinity Rankings, and Relative Binding Free Energies. J. Comput. Aided Mol. Des. 2018, 32(1), 1–20. [Google Scholar] [CrossRef] [PubMed]
- Wagner, J. R.; Churas, C. P.; Liu, S.; Swift, R. V.; Chiu, M.; Shao, C.; Feher, V. A.; Burley, S. K.; Gilson, M. K.; Amaro, R. E. Continuous Evaluation of Ligand Protein Predictions: A Weekly Community Challenge for Drug Docking. Structure 2019, 27(8), 1326–1335.e4. [Google Scholar] [CrossRef] [PubMed]
- Ackloo, S.; Al-Awar, R.; Amaro, R. E.; Arrowsmith, C. H.; Azevedo, H.; Batey, R. A.; Bengio, Y.; Betz, U. A. K.; Bologa, C. G.; Chodera, J. D.; Cornell, W. D.; Dunham, I.; Ecker, G. F.; Edfeldt, K.; Edwards, A. M.; Gilson, M. K.; Gordijo, C. R.; Hessler, G.; Hillisch, A.; Hogner, A.; Irwin, J. J.; Jansen, J. M.; Kuhn, D.; Leach, A. R.; Lee, A. A.; Lessel, U.; Morgan, M. R.; Moult, J.; Muegge, I.; Oprea, T. I.; Perry, B. G.; Riley, P.; Rousseaux, S. A. L.; Saikatendu, K. S.; Santhakumar, V.; Schapira, M.; Scholten, C.; Todd, M. H.; Vedadi, M.; Volkamer, A.; Willson, T. M. CACHE (Critical Assessment of Computational Hit-Finding Experiments): A Public-Private Partnership Benchmarking Initiative to Enable the Development of Computational Methods for Hit-Finding. Nat Rev Chem 2022, 6(4), 287–295. [Google Scholar] [CrossRef] [PubMed]
- Feynman, R. P. Cargo Cult Science. Engineering and Science 1974, 37(7), 10–13. [Google Scholar]
- Garrido, A.; Lepailleur, A.; Mignani, S. M.; Dallemagne, P.; Rochais, C. HERG Toxicity Assessment: Useful Guidelines for Drug Design. Eur. J. Med. Chem. 2020, 195(112290), 112290. [Google Scholar] [CrossRef] [PubMed]
- Bowes, J.; Brown, A. J.; Hamon, J.; Jarolimek, W.; Sridhar, A.; Waldron, G.; Whitebread, S. Reducing Safety-Related Drug Attrition: The Use of in Vitro Pharmacological Profiling. Nat. Rev. Drug Discov. 2012, 11(12), 909–922. [Google Scholar] [CrossRef] [PubMed]
- Denisov, I. G.; Makris, T. M.; Sligar, S. G.; Schlichting, I. Structure and Chemistry of Cytochrome P450. Chem. Rev. 2005, 105(6), 2253–2277. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



