Submitted:
11 December 2025
Posted:
12 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Cell Lines and Induction of Replication Stress
2.2. DNA Damage Induction with Illudin S
2.3. Cell Viability MTS Assay
2.4. DNA Damage and Replication Stress Markers
2.5. BrdU IF Assay
2.6. Cell Viability and Apoptosis
2.7. Propidium Iodide-Based Flow Cytometric Analysis of Cell-Cycle Phase and Sub G1 Population
2.8. Senescence-Associated β-Galactosidase (SA-β-Gal) Staining
2.9. Proximity Ligation Assay (PLA)
2.10. Perfil Transcriptómico (RNA-seq)
2.11. Whole-Exome Sequencing (WES)
2.12. Statistical Analysis
2.13. Declaration of Generative AI and AI-Assisted Technologies in the Writing Process
3. Results
3.1. Hydroxyurea-Induced Replication Stress in a TC-NER–Deficient Model
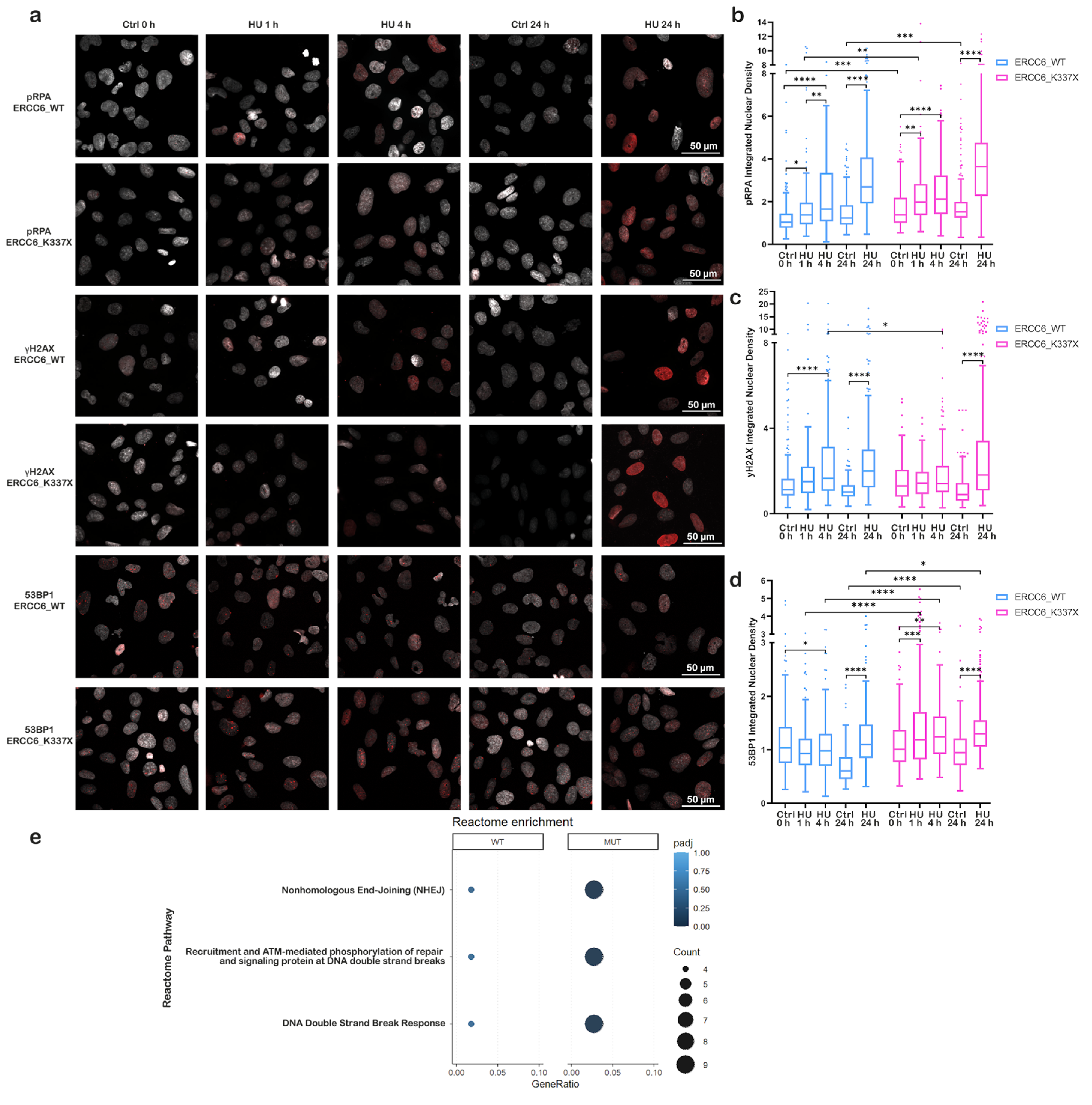
3.2. ERCC6 Deficiency Compromises Transcription–Replication Conflict (TRC) Resolution and Promotes Early NHEJ Activation Under Replication Stress
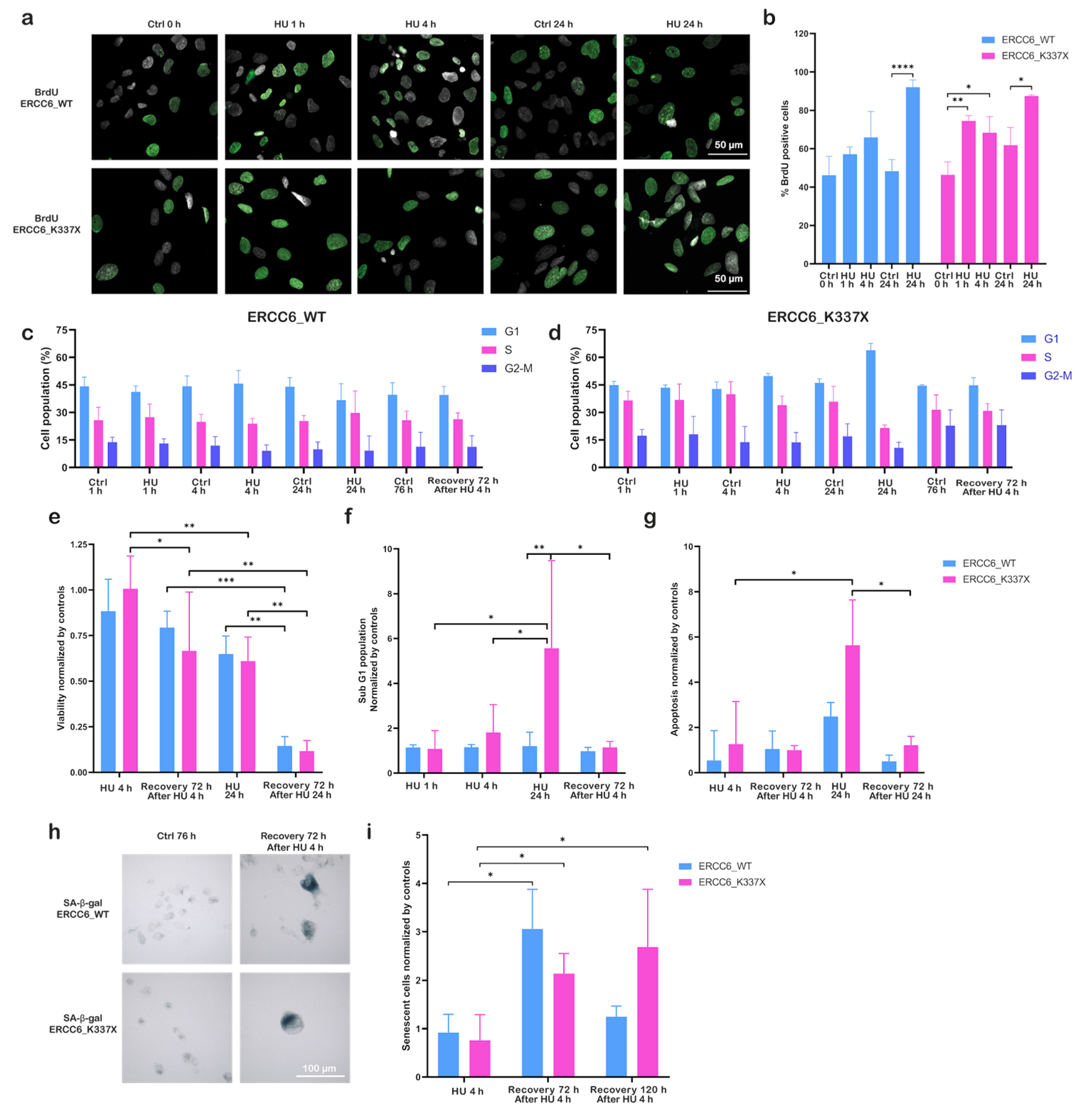
3.3. ERCC6 Deficiency Impairs Replication Dynamics and Promotes Senescence Under Replication Stress
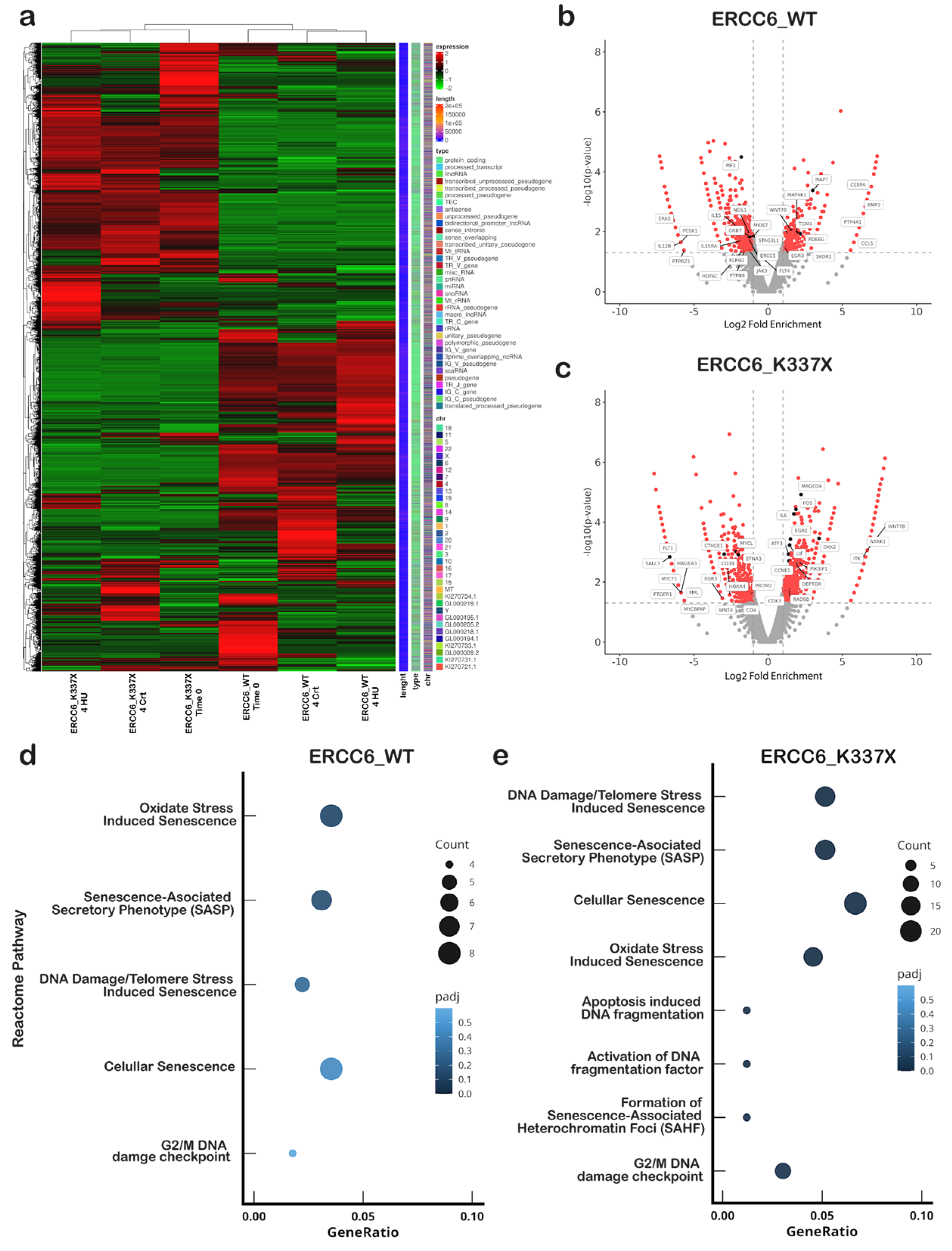
3.4. Divergent Transcriptional Programs Emerge Under Replication Stress Depending on ERCC6 Status
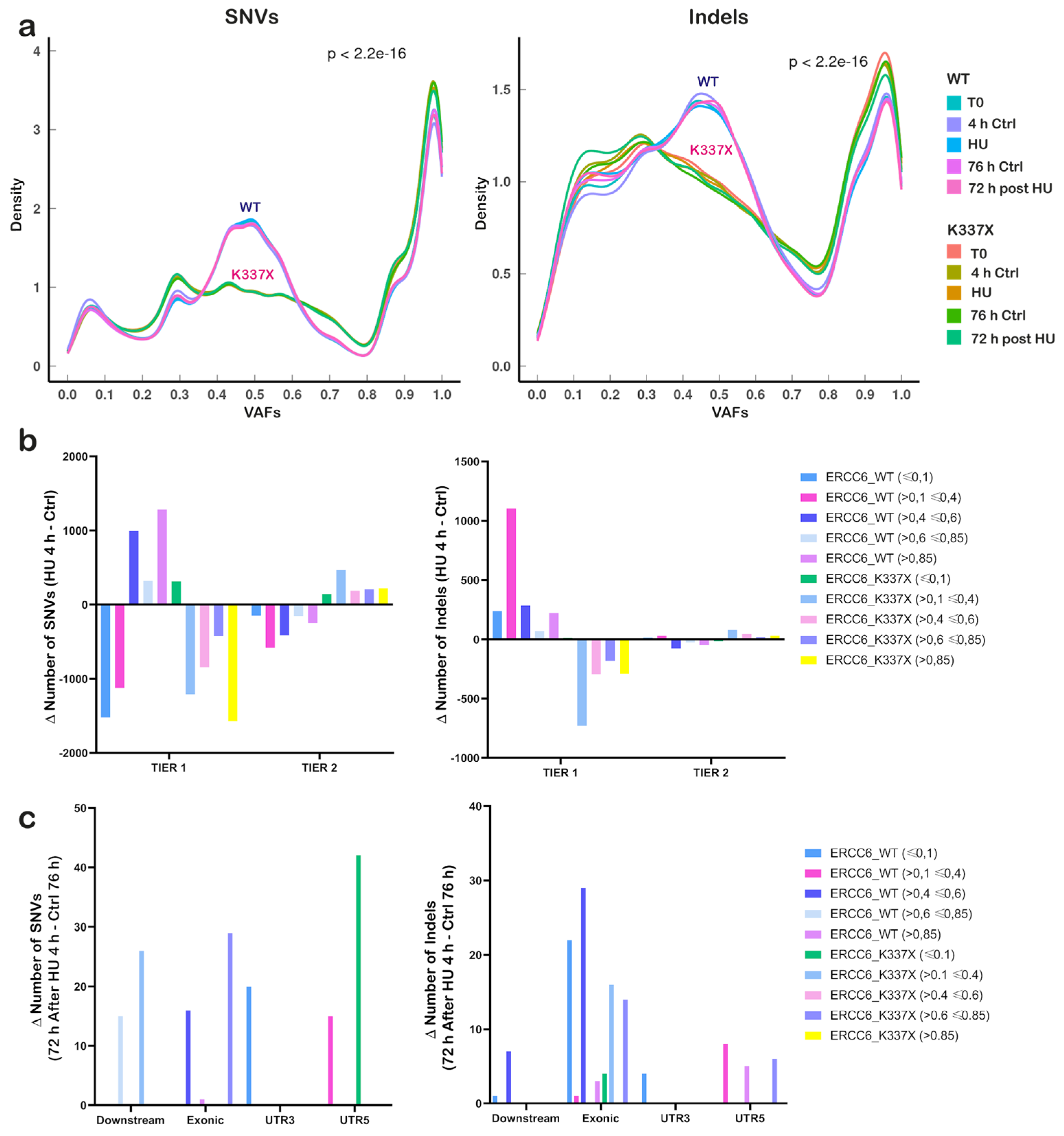
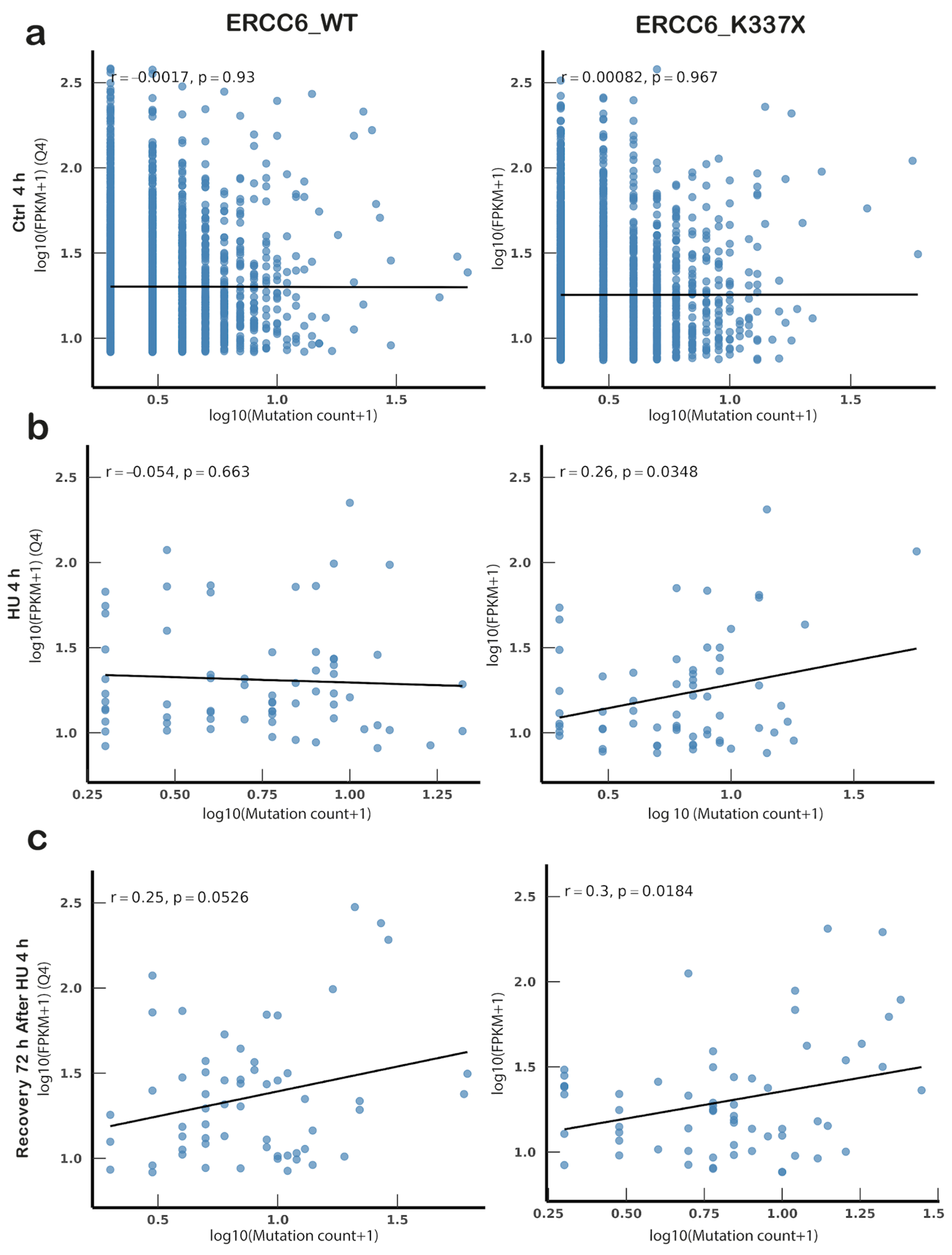
3.5. ERCC6 Status Determines Distinct Mutational Patterns and Expression–Mutation Coupling Under Replication Stress
4. Discussion
5. Limitations of the Study
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| Abbreviation | Definition |
| ERCC6 | Excision Repair Cross-Complementation Group 6 |
| CSB | Cockayne Syndrome Group B protein |
| CS | Cockayne Syndrome |
| RS | Replication Stress |
| TC-NER | Transcription-Coupled Nucleotide Excision Repair |
| TRCs | Transcription–Replication Conflicts |
| DSBs | DNA Double-Strand Breaks |
| HR | Homologous Recombination |
| NHEJ | Non-Homologous End Joining |
| Alt-EJ | Alternative End Joining |
| BIR | Break-Induced Replication |
| GG-NER | Global Genome Nucleotide Excision Repair |
| ATM | Ataxia Telangiectasia Mutated kinase |
| CDK2 | Cyclin-Dependent Kinase 2 |
| PARP1 | Poly(ADP-ribose) Polymerase 1 |
| PARP2 | Poly(ADP-ribose) Polymerase 2 |
| RPA | Replication Protein A |
| pRPA | Phosphorylated Replication Protein A |
| γH2AX | Phosphorylated Histone H2AX (Ser139) |
| 53BP1 | p53-Binding Protein 1 |
| RNAPII | RNA Polymerase II |
| PCNA | Proliferating Cell Nuclear Antigen |
| PLA | Proximity Ligation Assay |
| HU | Hydroxyurea |
| BrdU | 5-bromo-2′-deoxyuridin |
References
- Saxena, S.; Zou, L. Hallmarks of DNA Replication Stress. Mol. Cell 2022, 82, 2298–2314. [Google Scholar] [CrossRef] [PubMed]
- Wendel, B.M.; Hollingsworth, S.; Courcelle, C.T.; Courcelle, J. UV-induced DNA Damage Disrupts the Coordination between Replication Initiation, Elongation and Completion. Genes Cells 2021, 26, 94–108. [Google Scholar] [CrossRef] [PubMed]
- In Het Panhuis, W.; Tsaalbi-Shtylik, A.; Schönke, M.; Van Harmelen, V.; Pronk, A.C.M.; Streefland, T.C.M.; Sips, H.C.M.; Afkir, S.; Willems Van Dijk, K.; Rensen, P.C.N.; et al. Rev1 Deficiency Induces Replication Stress to Cause Metabolic Dysfunction Differently in Males and Females. Am. J. Physiol.-Endocrinol. Metab. 2022, 322, E319–E329. [Google Scholar] [CrossRef] [PubMed]
- Jegadesan, N.K.; Branzei, D. DDX11 Loss Causes Replication Stress and Pharmacologically Exploitable DNA Repair Defects. Proc. Natl. Acad. Sci. 2021, 118, e2024258118. [Google Scholar] [CrossRef]
- Ito, S.S.; Nakagawa, Y.; Matsubayashi, M.; Sakaguchi, Y.M.; Kobashigawa, S.; Matsui, T.K.; Nanaura, H.; Nakanishi, M.; Kitayoshi, F.; Kikuchi, S.; et al. Inhibition of the ATR Kinase Enhances 5-FU Sensitivity Independently of Nonhomologous End-Joining and Homologous Recombination Repair Pathways. J. Biol. Chem. 2020, 295, 12946–12961. [Google Scholar] [CrossRef]
- Musiałek, M.W.; Rybaczek, D. Hydroxyurea—The Good, the Bad and the Ugly. Genes 2021, 12, 1096. [Google Scholar] [CrossRef]
- Hills, S.A.; Diffley, J.F.X. DNA Replication and Oncogene-Induced Replicative Stress. Curr. Biol. 2014, 24, R435–R444. [Google Scholar] [CrossRef]
- Niehrs, C.; Luke, B. Regulatory R-Loops as Facilitators of Gene Expression and Genome Stability. Nat. Rev. Mol. Cell Biol. 2020, 21, 167–178. [Google Scholar] [CrossRef]
- Lalonde, M.; Trauner, M.; Werner, M.; Hamperl, S. Consequences and Resolution of Transcription–Replication Conflicts. Life 2021, 11, 637. [Google Scholar] [CrossRef]
- Kumar, C.; Remus, D. Looping out of Control: R-Loops in Transcription-Replication Conflict. Chromosoma 2024, 133, 37–56. [Google Scholar] [CrossRef]
- Rinaldi, C.; Pizzul, P.; Longhese, M.P.; Bonetti, D. Sensing R-Loop-Associated DNA Damage to Safeguard Genome Stability. Front. Cell Dev. Biol. 2021, 8, 618157. [Google Scholar] [CrossRef] [PubMed]
- Sollier, J.; Stork, C.T.; García-Rubio, M.L.; Paulsen, R.D.; Aguilera, A.; Cimprich, K.A. Transcription-Coupled Nucleotide Excision Repair Factors Promote R-Loop-Induced Genome Instability. Mol. Cell 2014, 56, 777–785. [Google Scholar] [CrossRef] [PubMed]
- Batenburg, N.L.; Thompson, E.L.; Hendrickson, E.A.; Zhu, X. Cockayne Syndrome Group B Protein Regulates DNA Double-Strand Break Repair and Checkpoint Activation. EMBO J. 2015, 34, 1399–1416. [Google Scholar] [CrossRef]
- Batenburg, N.L.; Walker, J.R.; Noordermeer, S.M.; Moatti, N.; Durocher, D.; Zhu, X.-D. ATM and CDK2 Control Chromatin Remodeler CSB to Inhibit RIF1 in DSB Repair Pathway Choice. Nat. Commun. 2017, 8, 1921. [Google Scholar] [CrossRef] [PubMed]
- Oka, Y.; Nakazawa, Y.; Shimada, M.; Ogi, T. Endogenous Aldehyde-Induced DNA–Protein Crosslinks Are Resolved by Transcription-Coupled Repair. Nat. Cell Biol. 2024, 26, 784–796. [Google Scholar] [CrossRef]
- Van Sluis, M.; Yu, Q.; Van Der Woude, M.; Gonzalo-Hansen, C.; Dealy, S.C.; Janssens, R.C.; Somsen, H.B.; Ramadhin, A.R.; Dekkers, D.H.W.; Wienecke, H.L.; et al. Transcription-Coupled DNA–Protein Crosslink Repair by CSB and CRL4CSA-Mediated Degradation. Nat. Cell Biol. 2024, 26, 770–783. [Google Scholar] [CrossRef]
- Carnie, C.J.; Acampora, A.C.; Bader, A.S.; Erdenebat, C.; Zhao, S.; Bitensky, E.; Van Den Heuvel, D.; Parnas, A.; Gupta, V.; D’Alessandro, G.; et al. Transcription-Coupled Repair of DNA–Protein Cross-Links Depends on CSA and CSB. Nat. Cell Biol. 2024, 26, 797–810. [Google Scholar] [CrossRef]
- Tuo, J.; Jaruga, P.; Rodriguez, H.; Bohr, V.A.; Dizdaroglu, M. Primary Fibroblasts of Cockayne Syndrome Patients Are Defective in Cellular Repair of 8-hydroxyguanine and 8-hydroxyadenine Resulting from Oxidative Stress. FASEB J. 2003, 17, 668–674. [Google Scholar] [CrossRef]
- Dianov, G. Repair of 8-Oxoguanine in DNA Is Deficient in Cockayne Syndrome Group B Cells. Nucleic Acids Res. 1999, 27, 1365–1368. [Google Scholar] [CrossRef]
- Ranes, M.; Boeing, S.; Wang, Y.; Wienholz, F.; Menoni, H.; Walker, J.; Encheva, V.; Chakravarty, P.; Mari, P.-O.; Stewart, A.; et al. A Ubiquitylation Site in Cockayne Syndrome B Required for Repair of Oxidative DNA Damage, but Not for Transcription-Coupled Nucleotide Excision Repair. Nucleic Acids Res. 2016, 44, 5246–5255. [Google Scholar] [CrossRef]
- Menoni, H.; Wienholz, F.; Theil, A.F.; Janssens, R.C.; Lans, H.; Campalans, A.; Radicella, J.P.; Marteijn, J.A.; Vermeulen, W. The Transcription-Coupled DNA Repair-Initiating Protein CSB Promotes XRCC1 Recruitment to Oxidative DNA Damage. Nucleic Acids Res. 2018, 46, 7747–7756. [Google Scholar] [CrossRef] [PubMed]
- Kumar, N.; Raja, S.; Van Houten, B. The Involvement of Nucleotide Excision Repair Proteins in the Removal of Oxidative DNA Damage. Nucleic Acids Res. 2020, 48, 11227–11243. [Google Scholar] [CrossRef] [PubMed]
- Thorslund, T.; Von Kobbe, C.; Harrigan, J.A.; Indig, F.E.; Christiansen, M.; Stevnsner, T.; Bohr, V.A. Cooperation of the Cockayne Syndrome Group B Protein and Poly(ADP-Ribose) Polymerase 1 in the Response to Oxidative Stress. Mol. Cell. Biol. 2005, 25, 7625–7636. [Google Scholar] [CrossRef]
- Boetefuer, E.L.; Lake, R.J.; Dreval, K.; Fan, H.-Y. Poly(ADP-Ribose) Polymerase 1 (PARP1) Promotes Oxidative Stress–Induced Association of Cockayne Syndrome Group B Protein with Chromatin. J. Biol. Chem. 2018, 293, 17863–17874. [Google Scholar] [CrossRef] [PubMed]
- Bilkis, R.; Lake, R.J.; Cooper, K.L.; Tomkinson, A.; Fan, H.-Y. The CSB Chromatin Remodeler Regulates PARP1- and PARP2-Mediated Single-Strand Break Repair at Actively Transcribed DNA Regions. Nucleic Acids Res. 2023, 51, 7342–7356. [Google Scholar] [CrossRef]
- Deem, A.; Keszthelyi, A.; Blackgrove, T.; Vayl, A.; Coffey, B.; Mathur, R.; Chabes, A.; Malkova, A. Break-Induced Replication Is Highly Inaccurate. PLoS Biol. 2011, 9, e1000594. [Google Scholar] [CrossRef]
- Batenburg, N.L.; Mersaoui, S.Y.; Walker, J.R.; Coulombe, Y.; Hammond-Martel, I.; Wurtele, H.; Masson, J.-Y.; Zhu, X.-D. Cockayne Syndrome Group B Protein Regulates Fork Restart, Fork Progression and MRE11-Dependent Fork Degradation in BRCA1/2-Deficient Cells. Nucleic Acids Res. 2021, 49, 12836–12854. [Google Scholar] [CrossRef]
- Batenburg, N.L.; Walker, J.R.; Zhu, X.-D. CSB Regulates Pathway Choice in Response to DNA Replication Stress Induced by Camptothecin. Int. J. Mol. Sci. 2023, 24, 12419. [Google Scholar] [CrossRef]
- Batenburg, N.L.; Sowa, D.J.; Walker, J.R.; Andres, S.N.; Zhu, X.-D. CSB and SMARCAL1 Compete for RPA32 at Stalled Forks and Differentially Control the Fate of Stalled Forks in BRCA2-Deficient Cells. Nucleic Acids Res. 2024, 52, 5067–5087. [Google Scholar] [CrossRef]
- Martin, H.A.; Pedraza-Reyes, M.; Yasbin, R.E.; Robleto, E.A. Transcriptional De-Repression and Mfd Are Mutagenic in Stressed Bacillus Subtilis Cells. J. Mol. Microbiol. Biotechnol. 2011, 21, 45–58. [Google Scholar] [CrossRef]
- Million-Weaver, S.; Samadpour, A.N.; Moreno-Habel, D.A.; Nugent, P.; Brittnacher, M.J.; Weiss, E.; Hayden, H.S.; Miller, S.I.; Liachko, I.; Merrikh, H. An Underlying Mechanism for the Increased Mutagenesis of Lagging-Strand Genes in Bacillus Subtilis. Proc. Natl. Acad. Sci. 2015, 112. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Sahin, O.; Barton, Y.-W.; Zhang, Q. Key Role of Mfd in the Development of Fluoroquinolone Resistance in Campylobacter Jejuni. PLoS Pathog. 2008, 4, e1000083. [Google Scholar] [CrossRef] [PubMed]
- Ragheb, M.N.; Thomason, M.K.; Hsu, C.; Nugent, P.; Gage, J.; Samadpour, A.N.; Kariisa, A.; Merrikh, C.N.; Miller, S.I.; Sherman, D.R.; et al. Inhibiting the Evolution of Antibiotic Resistance. Mol. Cell 2019, 73, 157–165.e5. [Google Scholar] [CrossRef] [PubMed]
- Galhardo, R.S.; Hastings, P.J.; Rosenberg, S.M. Mutation as a Stress Response and the Regulation of Evolvability. Crit. Rev. Biochem. Mol. Biol. 2007, 42, 399–435. [Google Scholar] [CrossRef]
- Portman, J.R.; Brouwer, G.M.; Bollins, J.; Savery, N.J.; Strick, T.R. Cotranscriptional R-Loop Formation by Mfd Involves Topological Partitioning of DNA. Proc. Natl. Acad. Sci. U. S. A 2021, 118, e2019630118. [Google Scholar] [CrossRef]
- Sarmini, L.; Meabed, M.; Emmanouil, E.; Atsaves, G.; Robeska, E.; Karwowski, B.T.; Campalans, A.; Gimisis, T.; Khobta, A. Requirement of Transcription-Coupled Nucleotide Excision Repair for the Removal of a Specific Type of Oxidatively Induced DNA Damage. Nucleic Acids Res. 2023, 51, 4982–4994. [Google Scholar] [CrossRef]
- Khobta, A.; Sarmini, L. Transcription-Coupled Nucleotide Excision Repair: A Faster Solution or the Only Option? Biomolecules 2025, 15, 1026. [Google Scholar] [CrossRef]
- Sarmini, L.; Kitsera, N.; Meabed, M.; Khobta, A. Transcription Blocking Properties and Transcription-Coupled Repair of N2-Alkylguanine Adducts as a Model for Aldehyde-Induced DNA Damage. J. Biol. Chem. 2025, 301, 108459. [Google Scholar] [CrossRef]
- Vogelstein, B.; Papadopoulos, N.; Velculescu, V.E.; Zhou, S.; Diaz, L.A.; Kinzler, K.W. Cancer Genome Landscapes. Science 2013, 339, 1546–1558. [Google Scholar] [CrossRef]
- McGranahan, N.; Swanton, C. Biological and Therapeutic Impact of Intratumor Heterogeneity in Cancer Evolution. Cancer Cell 2015, 27, 15–26. [Google Scholar] [CrossRef]
- Parsels, L.A.; Karnak, D.; Parsels, J.D.; Zhang, Q.; Vélez-Padilla, J.; Reichert, Z.R.; Wahl, D.R.; Maybaum, J.; O’Connor, M.J.; Lawrence, T.S.; et al. PARP1 Trapping and DNA Replication Stress Enhance Radiosensitization with Combined WEE1 and PARP Inhibitors. Mol. Cancer Res. 2018, 16, 222–232. [Google Scholar] [CrossRef] [PubMed]
- Vesela, E.; Chroma, K.; Turi, Z.; Mistrik, M. Common Chemical Inductors of Replication Stress: Focus on Cell-Based Studies. Biomolecules 2017, 7, 19. [Google Scholar] [CrossRef] [PubMed]
- Colicchia, V.; Petroni, M.; Guarguaglini, G.; Sardina, F.; Sahún-Roncero, M.; Carbonari, M.; Ricci, B.; Heil, C.; Capalbo, C.; Belardinilli, F.; et al. PARP Inhibitors Enhance Replication Stress and Cause Mitotic Catastrophe in MYCN-Dependent Neuroblastoma. Oncogene 2017, 36, 4682–4691. [Google Scholar] [CrossRef] [PubMed]
- Xu, R.; Zhu, S.; Zhang, W.; Xu, H.; Tu, C.; Wang, H.; Wang, L.; He, N.; Liu, T.; Guo, X.; et al. A Dual Approach with Organoid and CRISPR Screening Reveals ERCC6 as a Determinant of Cisplatin Resistance in Osteosarcoma. Adv. Sci. 2025, 12, 2500632. [Google Scholar] [CrossRef]
- Dimri, G.P.; Lee, X.; Basile, G.; Acosta, M.; Scott, G.; Roskelley, C.; Medrano, E.E.; Linskens, M.; Rubelj, I.; Pereira-Smith, O. A Biomarker That Identifies Senescent Human Cells in Culture and in Aging Skin in Vivo. Proc. Natl. Acad. Sci. 1995, 92, 9363–9367. [Google Scholar] [CrossRef]
- Panier, S.; Boulton, S.J. Double-Strand Break Repair: 53BP1 Comes into Focus. Nat. Rev. Mol. Cell Biol. 2014, 15, 7–18. [Google Scholar] [CrossRef]
- Chowdhury, D.; Xu, X.; Zhong, X.; Ahmed, F.; Zhong, J.; Liao, J.; Dykxhoorn, D.M.; Weinstock, D.M.; Pfeifer, G.P.; Lieberman, J. A PP4-Phosphatase Complex Dephosphorylates γ-H2AX Generated during DNA Replication. Mol. Cell 2008, 31, 33–46. [Google Scholar] [CrossRef]
- Mentani, A.; Maresca, M.; Shiriaeva, A. Prime Editing: Mechanistic Insights and DNA Repair Modulation. Cells 2025, 14, 277. [Google Scholar] [CrossRef]
- Jeon, J.; Kang, T.-H. Transcription-Coupled Repair and R-Loop Crosstalk in Genome Stability. Int. J. Mol. Sci. 2025, 26, 3744. [Google Scholar] [CrossRef]
- Deaconescu, A.M. Mfd – at the Crossroads of Bacterial DNA Repair, Transcriptional Regulation and Molecular Evolvability. Transcription 2021, 12, 156–170. [Google Scholar] [CrossRef]
- Low, G.K.M.; Ng, G.Y.-Q.; Zeegers, D.; Ting, A.; Gopalakrishnan, K.; Khaw, A.K.; Jayapal, M.; Hande, M.P. Protective Role of Cockayne Syndrome B (CSB) Protein in Maintaining Genome Integrity in Human Cells under Oxidative Stress. Mutat. Res. - Genet. Toxicol. Environ. Mutagen. 2025, 907, 503887. [Google Scholar] [CrossRef]
- Herr, L.M.; Schaffer, E.D.; Fuchs, K.F.; Datta, A.; Brosh, R.M. Replication Stress as a Driver of Cellular Senescence and Aging. Commun. Biol. 2024, 7, 616. [Google Scholar] [CrossRef]
- Reid-Bayliss, K.S.; Arron, S.T.; Loeb, L.A.; Bezrookove, V.; Cleaver, J.E. Why Cockayne Syndrome Patients Do Not Get Cancer despite Their DNA Repair Deficiency. Proc. Natl. Acad. Sci. 2016, 113, 10151–10156. [Google Scholar] [CrossRef]
- Bilkis, R.; Lake, R.J.; Fan, H.-Y. ATP-Dependent Chromatin Remodeler CSB Couples DNA Repair Pathways to Transcription with Implications for Cockayne Syndrome and Cancer Therapy. Cells 2025, 14, 239. [Google Scholar] [CrossRef]
- Kotsantis, P.; Silva, L.M.; Irmscher, S.; Jones, R.M.; Folkes, L.; Gromak, N.; Petermann, E. Increased Global Transcription Activity as a Mechanism of Replication Stress in Cancer. Nat. Commun. 2016, 7, 13087. [Google Scholar] [CrossRef]
- Hidmi, O.; Shatleh, D.; Flayshman, S.O.; Monin, J.; Aqeilan, R.I. Super-Enhancers Shape the Landscape and Repair Dynamics of Transcription-Associated DNA Breaks in Cancer 2025.
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).



