Submitted:
09 December 2025
Posted:
11 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Methods - Data Synthesis and Analysis
2.1. Pan-Cancer Analysis from TCGA
2.2. Survival Analysis
2.3. Gene Expression Analysis from TCGA
3. Mechanistic Principles of Oncofetal Reprogramming
3.1. Colorectal Cancer (CRC)
3.2. Breast Cancer
3.3. Lung Cancer (NSCLC)
3.4. Liver Cancer (HCC)
5. Conclusion
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| YAP1 | Yes Associated Protein 1 |
| AP-1 | Activator Protein 1 |
| CSC | cancer stem cell |
| CRC | colorectal cancer |
| EMT | epithelial-to-mesenchymal transition |
| revSCs | revival stem cells |
| NSCLC | non-small cell lung cancer |
| HCC | hepatocellular carcinoma |
| AFP | α- Fetoprotein |
| GPC3 | Glypican-3 |
| RBP | RNA-binding protein |
| LUAD | lung adenocarcinoma |
| MSF | migration-stimulating factor |
| piRNAs | PIWI-interacting RNA |
| TPM | Transcript per million |
| KM | Kaplan-Meier |
References
- Sharma; Blériot, C.; Currenti, J.; Ginhoux, F. Oncofetal reprogramming in tumour development and progression. Nat Rev Cancer 2022, 22(10), 593–602. [Google Scholar] [CrossRef]
- Zaidi, S. K.; et al. Bivalent Epigenetic Control of Oncofetal Gene Expression in Cancer. Mol Cell Biol 2017, 37(23). [Google Scholar] [CrossRef]
- Huang, J. F.; et al. Genome-wide screening identifies oncofetal lncRNA Ptn-dt promoting the proliferation of hepatocellular carcinoma cells by regulating the Ptn receptor. Oncogene 2019, 38(18), 3428–3445. [Google Scholar] [CrossRef] [PubMed]
- West, R. C.; Bouma, G. J.; Winger, Q. A. Shifting perspectives from ‘oncogenic’ to oncofetal proteins; How these factors drive placental development. Reproductive Biology and Endocrinology 2018, 16(1). [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; et al. Oncofetal TRIM71 drives liver cancer carcinogenesis through remodeling CEBPA-mediated serine/glycine metabolism. Theranostics 2024, 14(13), 4948–4966. [Google Scholar] [CrossRef] [PubMed]
- Cao, J.; et al. Oncofetal reprogramming in tumor development and progression: novel insights into cancer therapy; John Wiley and Sons Inc, 01 Dec 2023. [Google Scholar] [CrossRef]
- Currenti, J.; Mishra, A.; Wallace, M.; George, J.; Sharma, A. Immunosuppressive mechanisms of oncofetal reprogramming in the tumor microenvironment: implications in immunotherapy response; Portland Press Ltd, 01 Apr 2023. [Google Scholar] [CrossRef]
- Oo, H. Z.; et al. Oncofetal chondroitin sulfate is a highly expressed therapeutic target in non-small cell lung cancer. Cancers (Basel) 2021, 13(17). [Google Scholar] [CrossRef]
- Mzoughi, S.; et al. Oncofetal reprogramming drives phenotypic plasticity in WNT-dependent colorectal cancer. Nat Genet 2025, 57(2), 402–412. [Google Scholar] [CrossRef]
- Liang; et al. Advancements and applications of single-cell multi-omics techniques in cancer research: Unveiling heterogeneity and paving the way for precision therapeutics; Elsevier B.V, 01 Mar 2024. [Google Scholar] [CrossRef]
- Zou, X.; Wang, Y.; Luan, M.; Zhang, Y. Multi-omics and single-cell approaches reveal molecular subtypes and key cell interactions in hepatocellular carcinoma. Front Pharmacol 2025, 16. [Google Scholar] [CrossRef]
- Kim, M.; et al. SALL4 Is Required for YAP1-Dependent Malignant and Regenerative Hepatocyte-to-Cholangiocyte Reprogramming. Cancer research communications 2025, 5(9), 1714–1727. [Google Scholar] [CrossRef]
- Hu, S.; et al. NOTCH-YAP1/TEAD-DNMT1 Axis Drives Hepatocyte Reprogramming Into Intrahepatic Cholangiocarcinoma. Gastroenterology 2022, 163(2), 449–465. [Google Scholar] [CrossRef]
- Hsieh, M. H.; et al. Liver cancer initiation requires translational activation by an oncofetal regulon involving LIN28 proteins. Journal of Clinical Investigation 2024, 134(11). [Google Scholar] [CrossRef] [PubMed]
- Faraji, F.; et al. YAP-driven malignant reprogramming of oral epithelial stem cells at single cell resolution. Nature Communications 2025, 16(1). [Google Scholar] [CrossRef]
- Schott; et al. The IGF2BP1 oncogene is a druggable m6A-dependent enhancer of YAP1-driven gene expression in ovarian cancer. NAR Cancer 2025, 7(1). [Google Scholar] [CrossRef]
- Bala, R.; et al. Targeting the Hippo/YAP Pathway: A Promising Approach for Cancer Therapy and Beyond; John Wiley and Sons Inc, 01 Sep 2025. [Google Scholar] [CrossRef]
- Watt, K. I.; et al. The Hippo pathway effector YAP is a critical regulator of skeletal muscle fibre size. Nat Commun 2015, 6. [Google Scholar] [CrossRef]
- Wu, Z.; Nicoll, M.; Ingham, R. J. AP-1 family transcription factors: a diverse family of proteins that regulate varied cellular activities in classical hodgkin lymphoma and ALK+ ALCL; BioMed Central Ltd, 01 Dec 2021. [Google Scholar] [CrossRef]
- He, L.; Pratt, H.; Gao, M.; Wei, F.; Weng, Z.; Struhl, K. YAP and TAZ are transcriptional co-activators of AP-1 proteins and STAT3 during breast cellular transformation. 2021, 10. [Google Scholar] [CrossRef]
- Zhuo; et al. Integrative epigenetic analysis reveals AP-1 promotes activation of tumor-infiltrating regulatory T cells in HCC. Cellular and Molecular Life Sciences 2023, 80(4). [Google Scholar] [CrossRef]
- Szulzewsky, F.; Holland, E. C.; Vasioukhin, V. YAP1 and its fusion proteins in cancer initiation, progression and therapeutic resistance. Dev Biol 2021, 475, 205–221. [Google Scholar] [CrossRef]
- Frost, T. C.; et al. YAP1 and WWTR1 expression inversely correlates with neuroendocrine markers in Merkel cell carcinoma. Journal of Clinical Investigation 2023, 133(5). [Google Scholar] [CrossRef]
- Zhang, H.; et al. Metabolic reprogramming and immune evasion: the interplay in the tumor microenvironment; BioMed Central Ltd, 01 Dec 2024. [Google Scholar] [CrossRef]
- Li, Z.; Pai, R.; Gupta, S.; Currenti, J.; Gui, W.; Di Bartolomeo, A. Presence of onco-fetal neighborhoods in hepatocellular carcinoma is associated with relapse and response to immunotherapy. Nat Cancer 2024, 5(1), 167–186. [Google Scholar] [CrossRef] [PubMed]
- Yui, S.; et al. YAP/TAZ-Dependent Reprogramming of Colonic Epithelium Links ECM Remodeling to Tissue Regeneration. Cell Stem Cell 2018, 22(1), 35–49.e7. [Google Scholar] [CrossRef] [PubMed]
- Ayyaz; et al. Single-cell transcriptomes of the regenerating intestine reveal a revival stem cell. Nature 2019, 569(7754), 121–125. [Google Scholar] [CrossRef]
- Roulis, M.; et al. Paracrine orchestration of intestinal tumorigenesis by a mesenchymal niche. Nature 2020, 580(7804), 524–529. [Google Scholar] [CrossRef]
- Jacquemin, G.; et al. Paracrine signalling between intestinal epithelial and tumour cells induces a regenerative programme. Elife 2022, 11. [Google Scholar] [CrossRef]
- Han, T.; et al. Lineage reversion drives wnt independence in intestinal cancer. Cancer Discov 2020, 10(10), 1590–1609. [Google Scholar] [CrossRef] [PubMed]
- Cheung, P.; et al. Regenerative Reprogramming of the Intestinal Stem Cell State via Hippo Signaling Suppresses Metastatic Colorectal Cancer. Cell Stem Cell 2020, 27(4), 590–604.e9. [Google Scholar] [CrossRef] [PubMed]
- Vasquez, E. G.; et al. Dynamic and adaptive cancer stem cell population admixture in colorectal neoplasia. Cell Stem Cell 2022, 29(8), 1213–1228.e8. [Google Scholar] [CrossRef] [PubMed]
- Qin, X.; Cardoso Rodriguez, F.; Sufi, J.; Vlckova, P.; Claus, J.; Tape, C. J. An oncogenic phenoscape of colonic stem cell polarization. Cell 2023, 186(25), 5554–5568.e18. [Google Scholar] [CrossRef]
- Verhagen, M. P.; et al. Non-stem cell lineages as an alternative origin of intestinal tumorigenesis in the context of inflammation. Nat Genet 2024, 56(7), 1456–1467. [Google Scholar] [CrossRef]
- Kobayashi, S.; et al. Collagen type I-mediated mechanotransduction controls epithelial cell fate conversion during intestinal inflammation. Inflamm Regen 2022, 42(1). [Google Scholar] [CrossRef]
- Heinz, M. C.; et al. Liver Colonization by Colorectal Cancer Metastases Requires YAP-Controlled Plasticity at the Micrometastatic Stage. Cancer Res 2022, 82(10), 1953–1968. [Google Scholar] [CrossRef]
- Moorman; et al. Progressive plasticity during colorectal cancer metastasis. Nature 2025, 637(8047), 947–954. [Google Scholar] [CrossRef] [PubMed]
- Bobbitt, J. R.; Seachrist, D. D.; Keri, R. A. Chromatin Organization and Transcriptional Programming of Breast Cancer Cell Identity; Endocrine Society, 01 Aug 2023. [Google Scholar] [CrossRef]
- Iglesia, M. D.; et al. Differential chromatin accessibility and transcriptional dynamics define breast cancer subtypes and their lineages. Nat Cancer 2024, 5(11), 1713–1736. [Google Scholar] [CrossRef] [PubMed]
- Fredlund, E.; Staaf, J.; Rantala, J. K.; Kallioniemi, O.; Borg, Å.; Ringnér, M. The gene expression landscape of breast cancer is shaped by tumor protein p53 status and epithelial-mesenchymal transition. Breast Cancer Research 2012, 14(4). [Google Scholar] [CrossRef] [PubMed]
- Franco, H. L.; et al. Enhancer transcription reveals subtype-specific gene expression programs controlling breast cancer pathogenesis. Genome Res 2018, 28(2), 159–170. [Google Scholar] [CrossRef]
- Shi, W.; Zhong, B.; Dong, J.; Hu, X.; Li, L. Super enhancer-driven core transcriptional regulatory circuitry crosstalk with cancer plasticity and patient mortality in triple-negative breast cancer. Front Genet 2023, 14. [Google Scholar] [CrossRef]
- Spike, T.; Engle, D. D.; Lin, J. C.; Cheung, S. K.; La, J.; Wahl, G. M. A mammary stem cell population identified and characterized in late embryogenesis reveals similarities to human breast cancer. Cell Stem Cell 2012, 10(2), 183–197. [Google Scholar] [CrossRef]
- Zvelebil, M.; et al. Embryonic mammary signature subsets are activated in Brca1-/-and basal-like breast cancers. Breast Cancer Research 2013, 15(2). [Google Scholar] [CrossRef]
- Pfefferle, D.; Spike, B. T.; Wahl, G. M.; Perou, C. M. Luminal progenitor and fetal mammary stem cell expression features predict breast tumor response to neoadjuvant chemotherapy. Breast Cancer Res Treat 2015, 149(2), 425–437. [Google Scholar] [CrossRef]
- Soady, K. J.; et al. Mouse mammary stem cells express prognostic markers for triple-negative breast cancer. Breast Cancer Research 2015, 17(1). [Google Scholar] [CrossRef]
- McMullen, J. R. W.; Soto, U. Newly identified breast luminal progenitor and gestational stem cell populations likely give rise to HER2-overexpressing and basal-like breast cancers. Discover Oncology 2022, 13(1). [Google Scholar] [CrossRef]
- Matouk, J.; et al. Oncofetal H19 RNA promotes tumor metastasis. Biochim Biophys Acta Mol Cell Res 2014, 1843(7), 1414–1426. [Google Scholar] [CrossRef]
- Huang, X.; Zhang, H.; Guo, X.; Zhu, Z.; Cai, H.; Kong, X. Insulin-like growth factor 2 mRNA-binding protein 1 (IGF2BP1) in cancer; BioMed Central Ltd, 28 Jun 2018. [Google Scholar] [CrossRef]
- Li, S.; Jiang, M. Elevated insulin-like growth factor 2 mRNA binding protein 1 levels predict a poor prognosis in patients with breast carcinoma using an integrated multi-omics data analysis. Front Genet 2022, 13. [Google Scholar] [CrossRef] [PubMed]
- Yan, Q.; Fang, X.; Li, C.; Lan, P.; Guan, X. Oncofetal proteins and cancer stem cells; Portland Press Ltd, 01 Sep 2022. [Google Scholar] [CrossRef]
- Paço, S. A. de B.; Garcia; Castro, J. L.; Costa-Pinto, A. R.; Freitas, R. Roles of the hox proteins in cancer invasion and metastasis; MDPI AG, 01 Jan 2021. [Google Scholar] [CrossRef]
- Ishii, H.; Afify, S. M.; Hassan, G.; Salomon, D. S.; Seno, M. Cripto-1 as a potential target of cancer stem cells for immunotherapy; MDPI AG, 02 May 2021. [Google Scholar] [CrossRef]
- de Bessa Garcia, S. A.; Araújo, M.; Pereira, T.; Mouta, J.; Freitas, R. HOX genes function in Breast Cancer development; Elsevier B.V., 01 Apr 2020. [Google Scholar] [CrossRef]
- Matsuura, H.; Hakomori, S.-I. The oncofetal domain of fibronectin defined by monoclonal antibody FDC-6: Its presence in fibronectins from fetal and tumor tissues and its absence in those from normal adult tissues and plasma (hepatoma/COOH-terminal region). 1985. Available online: https://www.pnas.org.
- Guller, S.; et al. Release of oncofetal fibronectin from human placenta. Placenta 2003, 24(8–9), 843–850. [Google Scholar] [CrossRef]
- Kaczmarek; Castellani, P.; Nicolo, G.; Spina, B.; Allemanni, G.; Zardi, L. Distribution of oncofetal fibronectin isoforms in normal, hyperplastic and neoplastic human breast tissues. Int J Cancer 1994, 59(1), 11–16. [Google Scholar] [CrossRef]
- Geiger, P.; Mayer, B.; Wiest, I.; Schulze, S.; Jeschke, U.; Weissenbacher, T. Binding of galectin-1 to breast cancer cells MCF7 induces apoptosis and inhibition of proliferation in vitro in a 2D- and 3D- cell culture model. BMC Cancer 2016, 16(1). [Google Scholar] [CrossRef]
- Sapra, P.; et al. Long-term tumor regression induced by an antibody-drug conjugate that targets 5T4, an oncofetal antigen expressed on tumor-initiating cells. Mol Cancer Ther 2013, 12(1), 38–47. [Google Scholar] [CrossRef]
- Tapaneeyakorn, S.; Chantima, W.; Thepthai, C.; Dharakul, T. Production, characterization, and in vitro effects of a novel monoclonal antibody against Mig-7. Biochem Biophys Res Commun 2016, 475(2), 149–153. [Google Scholar] [CrossRef]
- An, Y.; Hu, Y.; Li, X.; Li, Z.; Duan, J.; Da Yang, X. Selection of a novel DNA aptamer against OFA/iLRP for targeted delivery of doxorubicin to AML cells. Sci Rep 2019, 9(1). [Google Scholar] [CrossRef] [PubMed]
- Bhatlekar, S.; Fields, J. Z.; Boman, B. M. Role of HOX genes in stem cell differentiation and cancer; Hindawi Limited, 2018. [Google Scholar] [CrossRef]
- Donlic; Zafferani, M.; Padroni, G.; Puri, M.; Hargrove, A. E. Regulation of MALAT1 triple helix stability and in vitro degradation by diphenylfurans. Nucleic Acids Res 2020, 48(14), 7653–7664. [Google Scholar] [CrossRef]
- Laughney, M.; et al. Regenerative lineages and immune-mediated pruning in lung cancer metastasis. Nat Med 2020, 26(2), 259–269. [Google Scholar] [CrossRef] [PubMed]
- Ben-Porath; et al. An embryonic stem cell-like gene expression signature in poorly differentiated aggressive human tumors. Nat Genet 2008, 40(5), 499–507. [Google Scholar] [CrossRef] [PubMed]
- Hassan, A.; Guoan, C.; Kalemkerian, G. P.; Wicha, M. S.; Beer, D. G. An embryonic stem cell-like signature identifies poorly differentiated lung adenocarcinoma but not squamous cell carcinoma. Clinical Cancer Research 2009, 15(20), 6386–6390. [Google Scholar] [CrossRef]
- Wang; et al. Multi-omics analyses reveal biological and clinical insights in recurrent stage I non-small cell lung cancer. Nature Communications 2025, 16(1). [Google Scholar] [CrossRef]
- Wang, Y.; Dong, Q.; Zhang, Q.; Li, Z.; Wang, E.; Qiu, X. Overexpression of yes-associated protein contributes to progression and poor prognosis of non-small-cell lung cancer. Cancer Sci 2010, 101(5), 1279–1285. [Google Scholar] [CrossRef] [PubMed]
- Damelin, M.; et al. Delineation of a cellular hierarchy in lung cancer reveals an oncofetal antigen expressed on tumor-initiating cells. Cancer Res 2011, 71(12), 4236–4246. [Google Scholar] [CrossRef]
- Stern, P. L.; Harrop, R. 5T4 oncofoetal antigen: an attractive target for immune intervention in cancer; Springer Science and Business Media Deutschland GmbH, 01 Apr 2017. [Google Scholar] [CrossRef]
- Ahmad; et al. Emerging roles of long noncoding RNA H19 in human lung cancer; John Wiley and Sons Ltd, 01 Jun 2024. [Google Scholar] [CrossRef]
- Cohn, E.; et al. Reactivation of multiple fetal mirnas in lung adenocarcinoma. Cancers (Basel) 2021, 13(11). [Google Scholar] [CrossRef] [PubMed]
- Fujiwara, Y.; et al. Oncofetal IGF2BP3-mediated control of microRNA structural diversity in the malignancy of early-stage lung adenocarcinoma. Proc Natl Acad Sci U S A 2024, 121(36). [Google Scholar] [CrossRef]
- Pewarchuk, M. E.; et al. Identification of oncofetal PIWI-interacting RNAs as potential prognostic biomarkers in non-small cell lung cancer. Front Genet 2025, 16. [Google Scholar] [CrossRef]
- Schor, S. L.; et al. “Migration-Stimulating Factor: A Genetically Truncated Onco-Fetal Fibronectin Isoform Expressed by Carcinoma and Tumor-Associated Stromal Cells,” 2003. Available online: http://aacrjournals.org/cancerres/article-pdf/63/24/8827/2511244/zch02403008827.pdf.
- De Zuani, M.; et al. Single-cell and spatial transcriptomics analysis of non-small cell lung cancer. Nat Commun 2024, 15(1). [Google Scholar] [CrossRef]
- Wu, H.; et al. The role of YAP1 in liver cancer stem cells: proven and potential mechanisms; BioMed Central Ltd, 01 Dec 2022. [Google Scholar] [CrossRef]
- Liu; et al. “Immunotherapy, targeted therapy, and their cross talks in hepatocellular carcinoma,” 2023; Frontiers Media SA. [CrossRef]
- Han, S. X.; et al. Expression and clinical significance of YAP, TAZ, and AREG in hepatocellular carcinoma. J Immunol Res 2014, 2014. [CrossRef]
- Biagioni, F.; et al. Decoding YAP dependent transcription in the liver. Nucleic Acids Res 2022, 50(14), 7959–7971. [Google Scholar] [CrossRef] [PubMed]
- Simile, M. M.; et al. Post-translational deregulation of YAP1 is genetically controlled in rat liver cancer and determines the fate and stem-like behavior of the human disease. Oncotarget 2016, 7(31), 49194–49216. [Google Scholar] [CrossRef]
- Li, H.; et al. Deregulation of Hippo kinase signalling in Human hepatic malignancies. Liver International 2012, 32(1), 38–47. [Google Scholar] [CrossRef] [PubMed]
- Patel, S. H.; Camargo, F. D.; Yimlamai, D. Hippo Signaling in the Liver Regulates Organ Size, Cell Fate, and Carcinogenesis; W.B. Saunders, 01 Feb 2017. [Google Scholar] [CrossRef]
- Hyun; Oh, S. H.; Premont, R. T.; Guy, C. D.; Berg, C. L.; Diehl, A. M. Dysregulated activation of fetal liver programme in acute liver failure. Gut 2019, 68(6), 1076–1087. [Google Scholar] [CrossRef]
- Fitamant; et al. YAP Inhibition Restores Hepatocyte Differentiation in Advanced HCC, Leading to Tumor Regression. Cell Rep 2015, 10(10), 1692–1707. [Google Scholar] [CrossRef] [PubMed]
- Cubero, F. J.; Martinez-Chantar, M. L. Plasticity of adult hepatocytes and readjustment of cell fate: A novel dogma in liver disease; BMJ Publishing Group, 01 Jun 2019. [Google Scholar] [CrossRef]
- Devan, R.; et al. The role of glypican-3 in hepatocellular carcinoma: Insights into diagnosis and therapeutic potential. 2024. [Google Scholar] [CrossRef]
- Cheng, S. W.; et al. Lin28B is an oncofetal circulating cancer stem cell-like marker associated with recurrence of hepatocellular carcinoma. PLoS One 2013, 8(11). [Google Scholar] [CrossRef] [PubMed]
- Yong, J.; et al. Oncofetal Gene SALL4 in Aggressive Hepatocellular Carcinoma. New England Journal of Medicine 2013, 368(24), 2266–2276. [Google Scholar] [CrossRef]
- Yin, F.; Han, X.; Yao, S. K.; Wang, X. L.; Yang, H. C. Importance of SALL4 in the development and prognosis of hepatocellular carcinoma. World J Gastroenterol 2016, 22(9), 2837–2843. [Google Scholar] [CrossRef]
- Han, S. X.; et al. Serum SALL4 is a novel prognosis biomarker with tumor recurrence and poor survival of patients in hepatocellular carcinoma. J Immunol Res 2014, 2014. [CrossRef]
- Tietze; Kessler, S. M. The Good, the Bad, the Question–H19 in Hepatocellular Carcinoma. Cancers (Basel) 2020, 5(12). [Google Scholar] [CrossRef]
- Wang, S.; Wu, X.; Liu, Y.; Yuan, J.; Yang, F. Long noncoding RNA H19 inhibits the proliferation of fetal liver cells and the Wnt signaling. FEBS Lett 2016, 4(590), 559–570. [Google Scholar] [CrossRef]
- Ding; et al. Long non-coding RNA PVT1 is associated with tumor progression and predicts recurrence in hepatocellular carcinoma patients. Oncol Lett 2015, 9(2), 955–963. [Google Scholar] [CrossRef]
- Sharma; et al. Onco-fetal Reprogramming of Endothelial Cells Drives Immunosuppressive Macrophages in Hepatocellular Carcinoma. Cell 2020, 183(2), 377–394.e21. [Google Scholar] [CrossRef]
- Nanki; Fujita, J.; Yang, Y.; Hojo, S.; Bandoh, S. Expression of Oncofetal Fibronectin and Syndecan-1 mRNA in 18 Human Lung Cancer Cell Lines. Tumor Biology 2001, no. 22, 390–396. [Google Scholar] [CrossRef] [PubMed]
- Samanta, S.; et al. IMP3 promotes stem-like properties in triple-negative breast cancer by regulating SLUG. Oncogene 2016, 35(9), 1111–1121. [Google Scholar] [CrossRef] [PubMed]
- dos Reis, S.; et al. Increased expression of the pathological O-glycosylated form of oncofetal fibronectin in the multidrug resistance phenotype of cancer cells. Matrix Biology 2023, 118, 47–68. [Google Scholar] [CrossRef] [PubMed]
- Yong, J.; et al. Targeting SALL4 by entinostat in lung cancer. Oncotarget. 2016, 7, pp. 75425–75440. Available online: www.impactjournals.com/oncotarget.
- Glaß; Hüttelmaier, S. IGF2BP1—An Oncofetal RNA-Binding Protein Fuels Tumor Virus Propagation; Multidisciplinary Digital Publishing Institute (MDPI), 01 Jul 2023. [Google Scholar] [CrossRef]
- Xiang, M.; et al. Oncofetal HLF transactivates c-Jun to promote hepatocellular carcinoma development and sorafenib resistance. Gut 2019, 68(10), 1858–1871. [Google Scholar] [CrossRef]
- Pandey; et al. ROR1 potentiates FGFR signaling in basal-like breast cancer. Cancers (Basel) 2019, 11(5). [Google Scholar] [CrossRef]
- Nicolb; Salvi, S.; Oliveri, G.; Borsi, L.; Castellani, P.; Zardi, L. Expression of tenascin and of the ED-B containing oncofetal fibronectin isoform in human cancer. 1990. [Google Scholar]
- Clausen, T. M.; et al. Oncofetal chondroitin sulfate glycosaminoglycans are key players in integrin signaling and tumor cell motility. Molecular Cancer Research 2016, 14(12), 1288–1299. [Google Scholar] [CrossRef] [PubMed]
- Pickholtz; et al. “Cooperation between BRCA1 and vitamin D is critical for histone acetylation of the p21waf1 promoter and for growth inhibition of breast cancer cells and cancer stem-like cells,” 2014. Available online: www.impactjournals.com/oncotarget/.
- Rokavec, H. Li; Jiang, L.; Hermeking, H. The p53/miR-34 axis in development and disease; Oxford University Press, 2014. [Google Scholar] [CrossRef]
- Agarwal, A; Bahadur, RP. Modular architecture and functional annotation of human RNA-binding proteins containing RNA recognition motif. Biochimie;Epub 2023, 209, 116–130. [Google Scholar] [CrossRef] [PubMed]
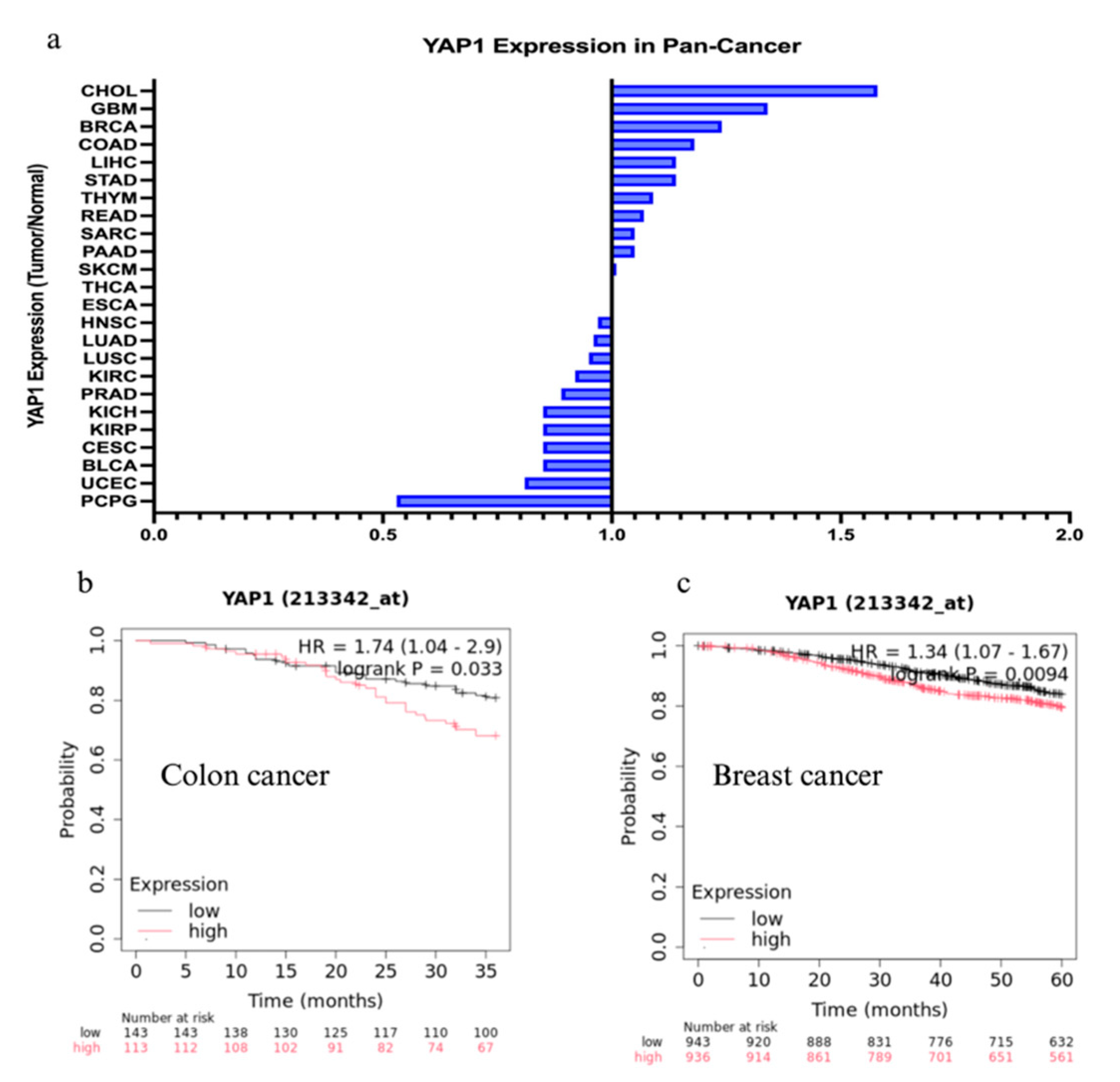
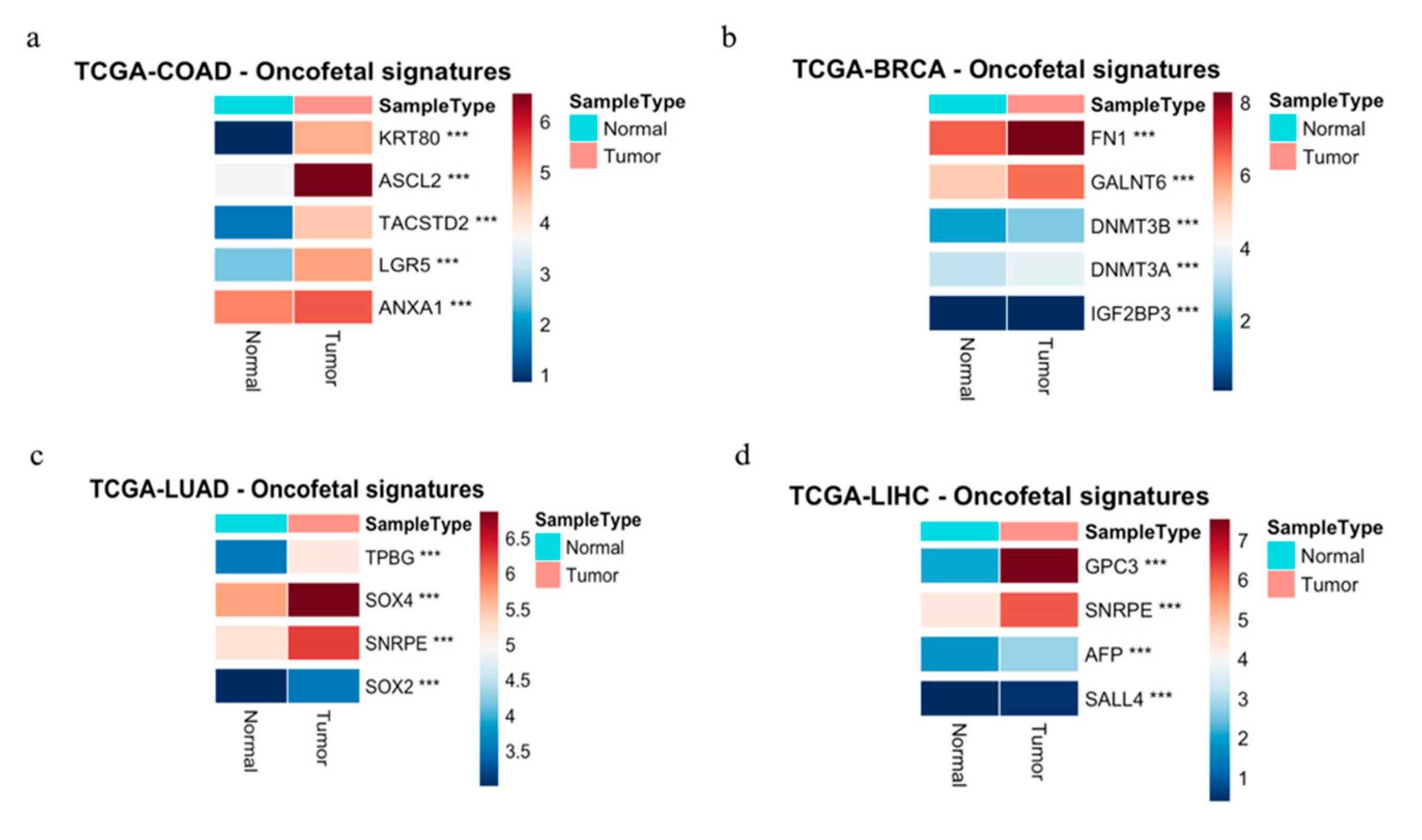
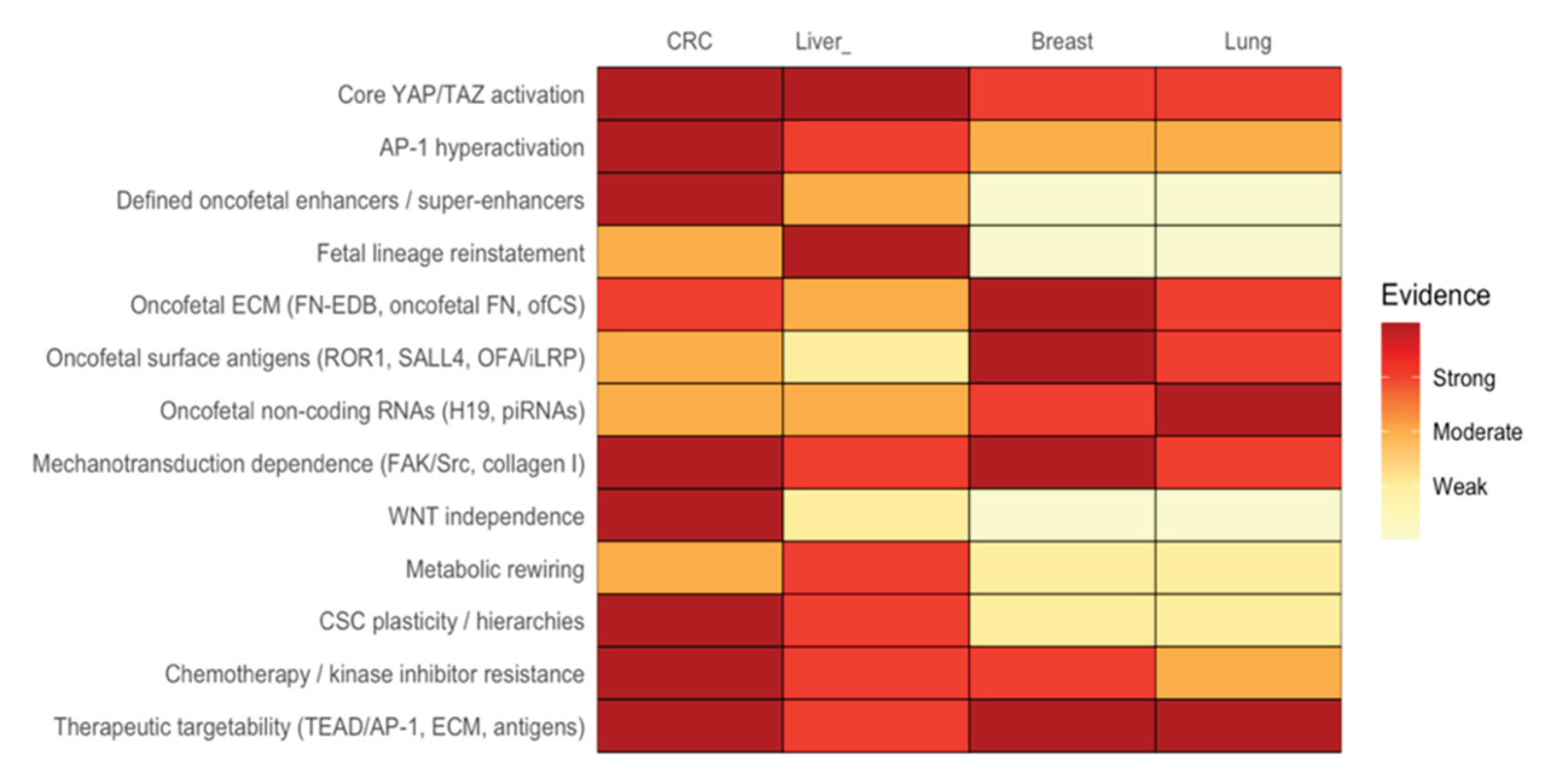
| Cancer type | Oncofetal signatures | References |
|---|---|---|
| Colorectal cancer (CRC) | TACSTD2 (TROP2), ANXA1, KRT80, THBS1, CTGF, CYR61, WWTR1 (TAZ), TEAD4, FOSL1, RUNX2 | [9,26,27,28,29,30,32,35,37] |
| Breast cancer | FN1 (oncofetal FN isoform), GALNT6, IGF2BP3 (IMP3), IGF2BP1 | [97,98] |
| NSCLC | TPBG (5T4), FN1 (oncofetal FN), SDC1, SALL4 | [59,96,99] |
| Liver cancer (HCC) | AFP, GPC3, SALL4, SNRPE | [89] |
| Other solid tumors | IGF2BP1, IGF2BP3, SALL4, TPBG (5T4), oncofetal chondroitin sulfate–modified proteoglycans (ofCS) | [100] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




