Submitted:
09 December 2025
Posted:
10 December 2025
You are already at the latest version
Abstract
RNA chaperones play a crucial role in the biogenesis and function of various RNAs in bacteria. They facilitate the interaction of small regulatory trans-encoded sRNAs with mRNAs, thereby significantly altering the pattern of gene expression in cells. This allows bacteria responses quickly to changing environmental conditions, such as stress or adaptation to host organisms. Despite the identification of a large number of sRNAs in mycobacteria, none of the most common RNA chaperones has been found in their genomes. We characterized the cold shock protein CspB from Mycobacterium tuberculosis as a potential RNA chaperone. It forms a dimer due to its elongated C-terminal region, which is a hairpin composed of two α-helices. We demonstrated that CspB from M. tuberculosis exhibits high affinity for the two studied sRNAs from the same organism and, unlike the single-domain CspA, could be considered as a potential RNA chaperone in mycobacteria. Thus, it may be involved in the regulation of bacterial pathogenesis via interactions with sRNAs.
Keywords:
1. Introduction
2. Results
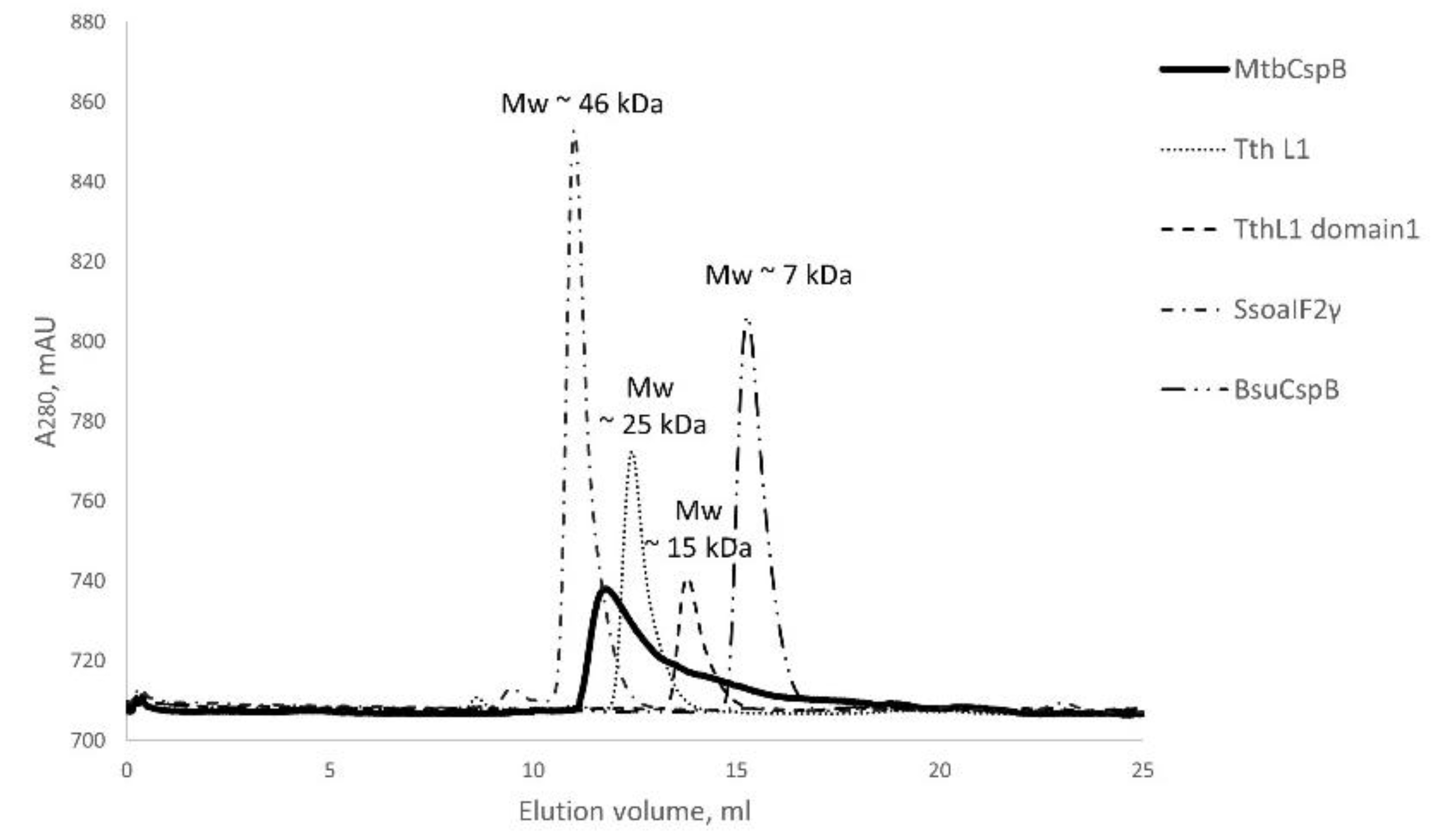
2.1. MtbCspB Forms Dimers
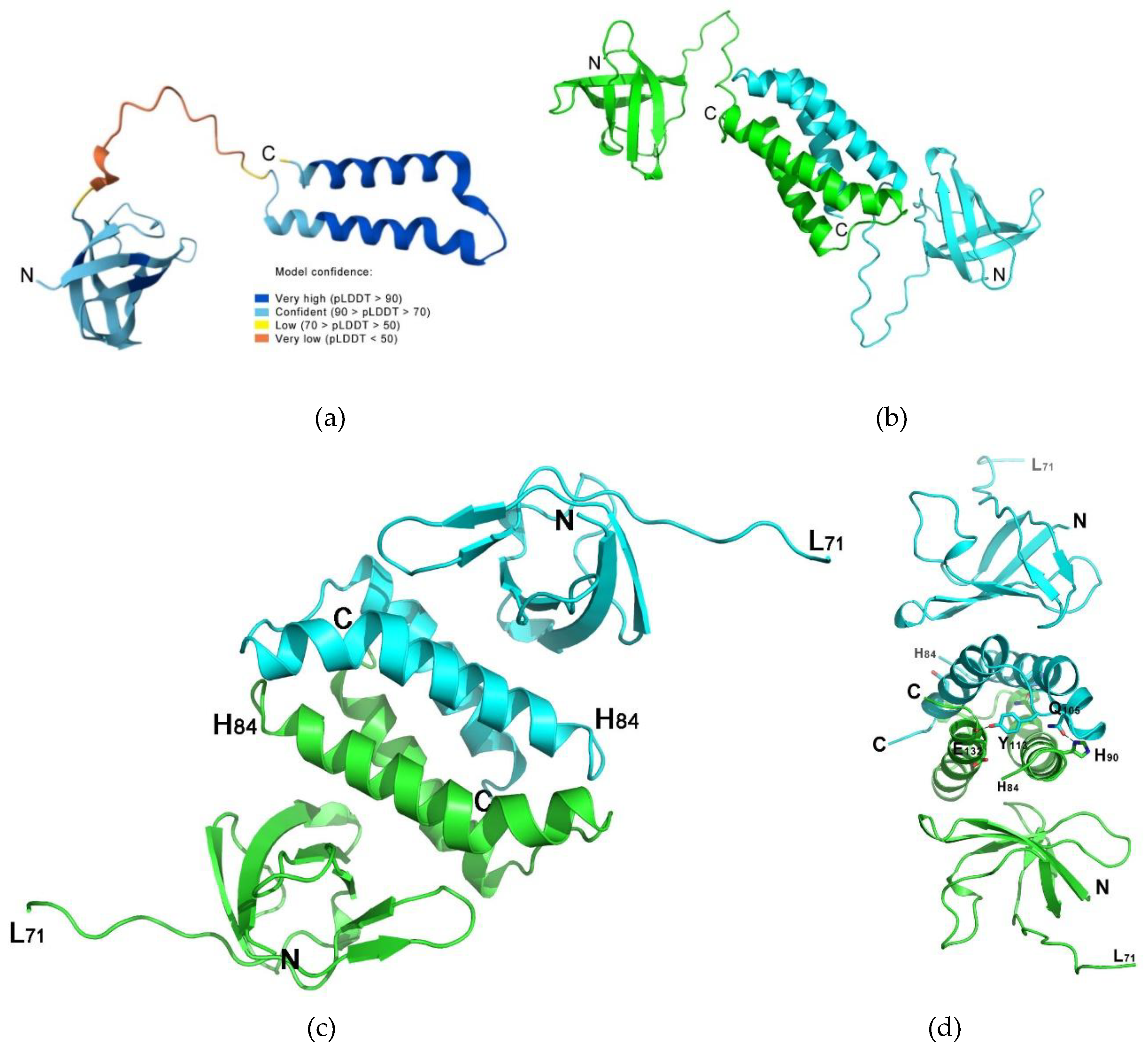
2.2. The MtbCspB Crystal Structure
2.3. MtbCspB Is Able to Bind sRNAs MTS0997 and MTS1338
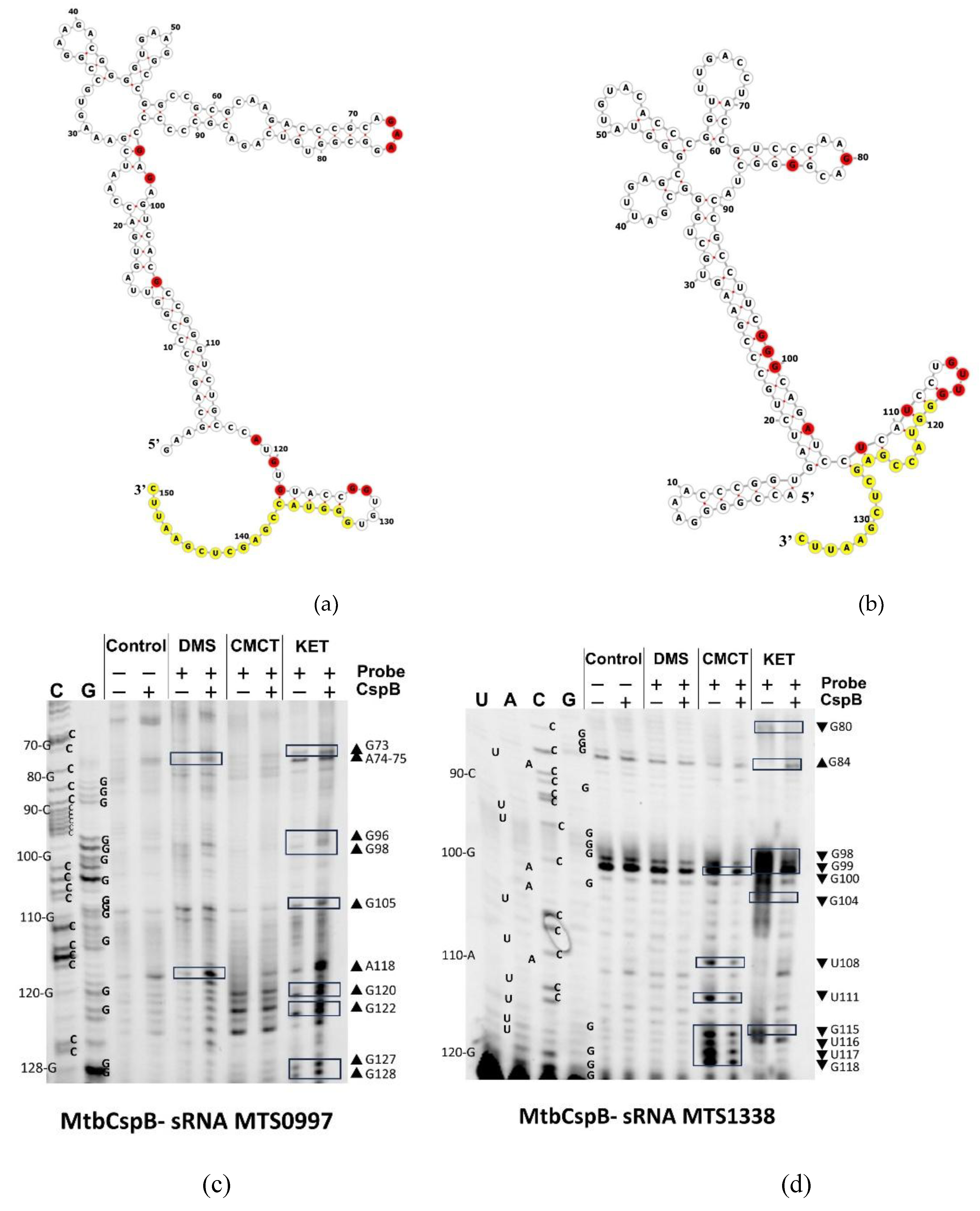
2.4. MtbCspB Modulates the Structure of MTS0997 and MTS1338 sRNAs Differently
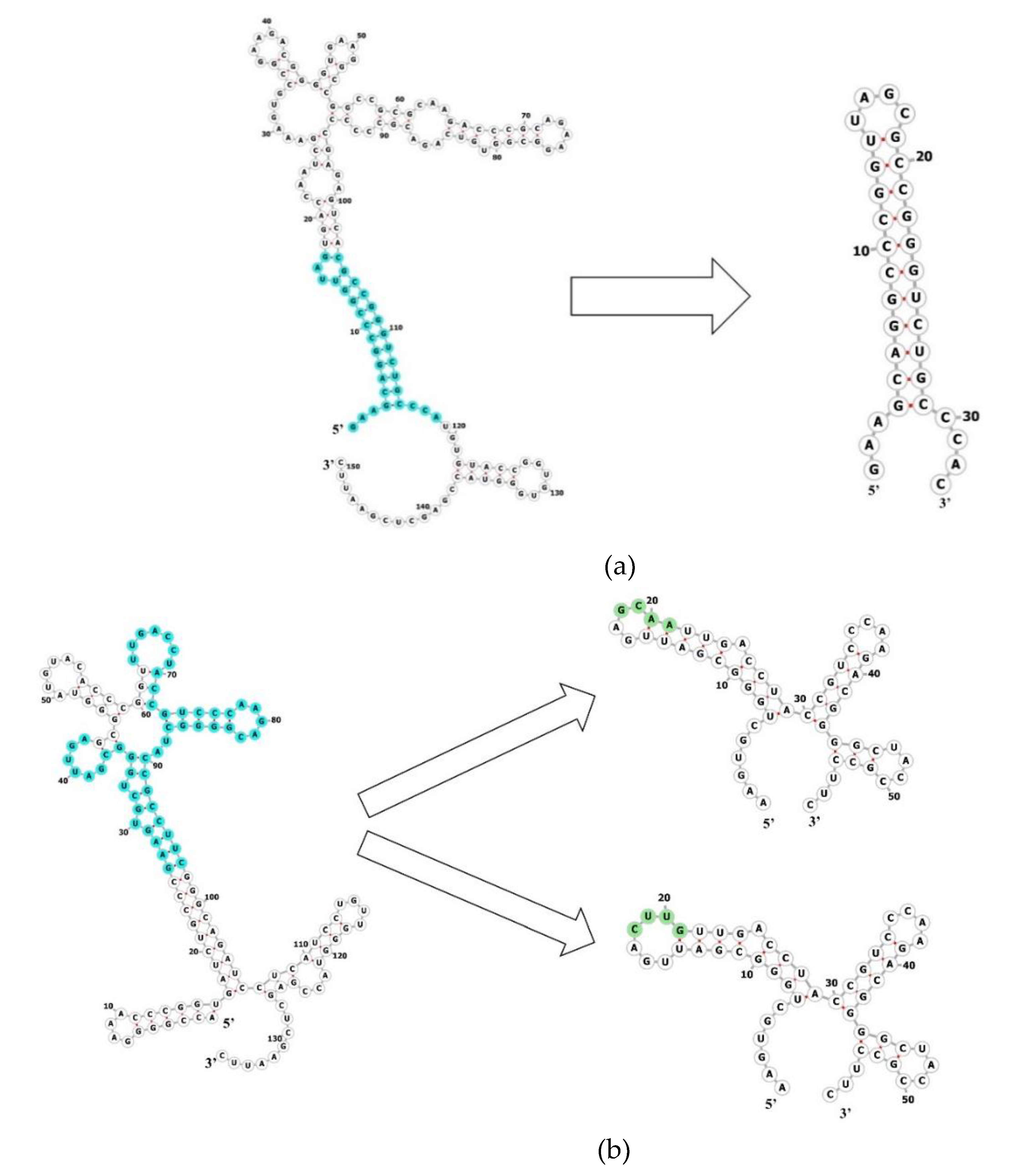
2.5. MtbCspB Can Bind to Short Variants of the RNAs
3. Discussion
4. Materials and Methods
4.1. Gene Cloning, Expression and Purification of MtbCspB
4.2. Analysis of Protein Particle Size by Gel Filtration
4.3. Protein Crystallization
4.4. X-Ray Diffraction Data Collection and Processing

4.5. Cloning and Purification of RNAs
4.6. RNA Chemical Probing
4.6.1. Chemical Modifications of RNA
4.6.2. Fluorescent Primer Extension
4.7. Analysis of MtbCspB-RNA Interaction
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Dutta, T.; Srivastava, S. Small RNA-Mediated Regulation in Bacteria: A Growing Palette of Diverse Mechanisms. Gene 2018, 656, 60–72. [Google Scholar] [CrossRef] [PubMed]
- Djapgne, L.; Oglesby, A.G. Impacts of Small RNAs and Their Chaperones on Bacterial Pathogenicity. Front. Cell. Infect. Microbiol. 2021, 11, 561. [Google Scholar] [CrossRef] [PubMed]
- Martini, B.A.; Grigorov, A.S.; Skvortsova, Y.V.; Bychenko, O.S.; Salina, E.G.; Azhikina, T.L. Small RNA MTS1338 Configures a Stress Resistance Signature in Mycobacterium Tuberculosis. Int. J. Mol. Sci. 2023, 24, 7928. [Google Scholar] [CrossRef] [PubMed]
- Azhikina, T.L.; Ignatov, D.V.; Salina, E.G.; Fursov, M.V.; Kaprelyants, A.S. Role of Small Noncoding RNAs in Bacterial Metabolism. Biochemistry (Moscow) 2015, 80, 1633–1646. [Google Scholar] [CrossRef]
- Arnvig, K.B.; Young, D.B. Non-Coding RNA and Its Potential Role in Mycobacterium Tuberculosis Pathogenesis. RNA Biol. 2012, 9, 427–436. [Google Scholar] [CrossRef]
- Ignatov, D.V.; Salina, E.G.; Fursov, M.V.; Skvortsov, T.A.; Azhikina, T.L.; Kaprelyants, A.S. Dormant Non-Culturable Mycobacterium Tuberculosis Retains Stable Low-Abundant MRNA. BMC Genomics 2015, 16, 954. [Google Scholar] [CrossRef]
- Moores, A.; Riesco, A.B.; Schwenk, S.; Arnvig, K.B. Expression, Maturation and Turnover of DrrS, an Unusually Stable, DosR Regulated Small RNA in Mycobacterium Tuberculosis. PLoS One 2017, 12, 1–27. [Google Scholar] [CrossRef]
- Pelly, S.; Bishai, W.R.; Lamichhane, G. A Screen for Non-Coding RNA in Mycobacterium Tuberculosis Reveals a CAMP-Responsive RNA That Is Expressed during Infection. Gene 2012, 500, 85–92. [Google Scholar] [CrossRef]
- Katsuya-Gaviria, K.; Paris, G.; Dendooven, T.; Bandyra, K.J. Bacterial RNA Chaperones and Chaperone-like Riboregulators: Behind the Scenes of RNA-Mediated Regulation of Cellular Metabolism. RNA Biol. 2022, 19, 419–436. [Google Scholar] [CrossRef]
- Sahoo, B.R.; Bardwell, J.C. Protein and RNA Chaperones. Mol. Aspects Med. 2025, 104, 101384. [Google Scholar] [CrossRef]
- Coskun, F.S.; Płociński, P.; van Oers, N.S.C. Small RNAs Asserting Big Roles in Mycobacteria. Non-Coding RNA 2021, 7, 69. [Google Scholar] [CrossRef] [PubMed]
- Taneja, S.; Dutta, T. On a Stake-out: Mycobacterial Small RNA Identification and Regulation. Non-coding RNA Res. 2019, 4, 86–95. [Google Scholar] [CrossRef] [PubMed]
- Czapski, T.R.; Trun, N. Expression of Csp Genes in E. Coli K-12 in Defined Rich and Defined Minimal Media during Normal Growth, and after Cold-Shock. Gene 2014, 547, 91–97. [Google Scholar] [CrossRef] [PubMed]
- Yamanaka, K.; Fang, L.; Inouye, M. The CspA Family in Escherichia Coli: Multiple Gene Duplication for Stress Adaptation. Mol. Microbiol. 1998, 27, 247–255. [Google Scholar] [CrossRef]
- Bae, W.; Xia, B.; Inouye, M.; Severinov, K. Escherichia Coli CspA-Family RNA Chaperones Are Transcription Antiterminators. Proc. Natl. Acad. Sci. U. S. A 2000, 97, 7784–7789. [Google Scholar] [CrossRef]
- Giuliodori, A.M.; Di Pietro, F.; Marzi, S.; Masquida, B.; Wagner, R.; Romby, P.; Gualerzi, C.O.; Pon, C.L. The CspA MRNA Is a Thermosensor That Modulates Translation of the Cold-Shock Protein CspA. Mol. Cell 2010, 37, 21–33. [Google Scholar] [CrossRef]
- Rennella, E.; Sára, T.; Juen, M.; Wunderlich, C.; Imbert, L.; Solyom, Z.; Favier, A.; Ayala, I.; Weinhäupl, K.; Schanda, P.; et al. RNA Binding and Chaperone Activity of the E. Coli Cold-Shock Protein CspA. Nucleic Acids Res. 2017, 45, 4255–4268. [Google Scholar] [CrossRef]
- Jiang, W.; Hou, Y.; Inouye, M. CspA, the Major Cold-Shock Protein of Escherichia Coli, Is an RNA Chaperone. J. Biol. Chem. 1997, 272, 196–202. [Google Scholar] [CrossRef]
- Zhang, Y.; Burkhardt, D.H.; Rouskin, S.; Li, G.; Weissman, J.S.; Gross, C.A. A Stress Response That Monitors and Regulates MRNA Structure Is Central to Cold Shock Adaptation. Mol. Cell 2018, 70, 274–286.e7. [Google Scholar] [CrossRef]
- Pankratova, P.Y.; Lekontseva, N.V.; Nikulin, A.D. Characterization of DNA/RNA Binding Properties of CspA Protein from Mycobacterium Tuberculosis. Silico Res. Biomed. 2025, 1, 100033. [Google Scholar] [CrossRef]
- Lekontseva, N.; Mikhailina, A.; Fando, M.; Kravchenko, O.; Balobanov, V.; Tishchenko, S.; Nikulin, A. Crystal Structures and RNA-Binding Properties of Lsm Proteins from Archaea Sulfolobus Acidocaldarius and Methanococcus Vannielii: Similarity and Difference of the U-Binding Mode. Biochimie 2020, 175, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Fando, M.S.; Mikhaylina, A.O.; Lekontseva, N.V.; Tishchenko, S.V.; Nikulin, A.D. Structure and RNA-Binding Properties of Lsm Protein from Halobacterium Salinarum. Biochemistry (Moscow) 2021, 86, 833–842. [Google Scholar] [CrossRef] [PubMed]
- Jørgensen, M.G.; Pettersen, J.S.; Kallipolitis, B.H. SRNA-Mediated Control in Bacteria: An Increasing Diversity of Regulatory Mechanisms. Biochim. Biophys. Acta - Gene Regul. Mech. 2020, 1863, 194504. [Google Scholar] [CrossRef] [PubMed]
- Schwenk, S.; Arnvig, K.B. Regulatory RNA in Mycobacterium Tuberculosis, Back to Basics. Pathog. Dis. 2018, 76, 1–12. [Google Scholar] [CrossRef]
- McCoy, A.J.; Grosse-Kunstleve, R.W.; Adams, P.D.; Winn, M.D.; Storoni, L.C.; Read, R.J. Phaser Crystallographic Software. J. Appl. Crystallogr. 2007, 40, 658–674. [Google Scholar] [CrossRef]
- Murshudov, G.N.; Vagin, A.A.; Dodson, E.J. Refinement of Macromolecular Structures by the Maximum-Likelihood Method. Acta Crystallogr. D. Biol. Crystallogr. 1997, 53, 240–255. [Google Scholar] [CrossRef]
- Winn, M.D.; Ballard, C.C.; Cowtan, K.D.; Dodson, E.J.; Emsley, P.; Evans, P.R.; Keegan, R.M.; Krissinel, E.B.; Leslie, A.G.W.; McCoy, A.; et al. Overview of the CCP4 Suite and Current Developments. Acta Crystallogr. D. Biol. Crystallogr. 2011, 67, 235–242. [Google Scholar] [CrossRef]
- Emsley, P.; Cowtan, K. Coot : Model-Building Tools for Molecular Graphics. Acta Crystallogr. Sect. D Biol. Crystallogr. 2004, 60, 2126–2132. [Google Scholar] [CrossRef]
- Nevskaya, N. Ribosomal Protein L1 Recognizes the Same Specific Structural Motif in Its Target Sites on the Autoregulatory MRNA and 23S RRNA. Nucleic Acids Res. 2005, 33, 478–485. [Google Scholar] [CrossRef]
- Stern, S.; Moazed, D.; Noller, H.F. Structural Analysis of RNA Using Chemical and Enzymatic Probing Monitored by Primer Extension. In Brenner’s Encyclopedia of Genetics; Elsevier, 1988; Vol. 164, pp. 481–489. ISBN 9780080961569. [Google Scholar]
- Katsamba, P.S.; Park, S.; Laird-Offringa, I.A. Kinetic Studies of RNA–Protein Interactions Using Surface Plasmon Resonance. Methods 2002, 26, 95–104. [Google Scholar] [CrossRef]
| RNA | ka (M-1*s-1) | kd (s-1) | KD (nM) |
|---|---|---|---|
| MTS0997 sRNA | 2.33E+06 | 2.62E-04 | 0.112±0.0056 |
| MTS1338 sRNA | 4.22E+06 | 1.25E-03 | 0.296± 0.0148 |
| oligoU RNA1 | 1.18E+04 | 2.23E-04 | 18.8±0.94 |
| oligoC RNA2 | not detected | not detected | not detected |
| RNA | ka (M-1*s-1) | kd (s-1) | KD (nM) |
|---|---|---|---|
| shortMTS0997 | 2.59E+04 8.31E+03* |
1.59E-03 1.22E-03* |
61.3±3.06 147±7.4* |
| shortMTS1338-GCAA | 2.19E+04 8.72E+03* |
2.93E-04 5.11E-04* |
13.3±0.67 58.6±2.93* |
| shortMTS1338-CUUG | 4.61E+04 8.64E+03* |
1.82E-03 1.11E-03* |
39.5±1.98 128±6.4* |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).




