Submitted:
05 December 2025
Posted:
05 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Literature-driven biomarker selection
2.1.1. Eligibility criteria
2.1.2. Information sources and search strategy
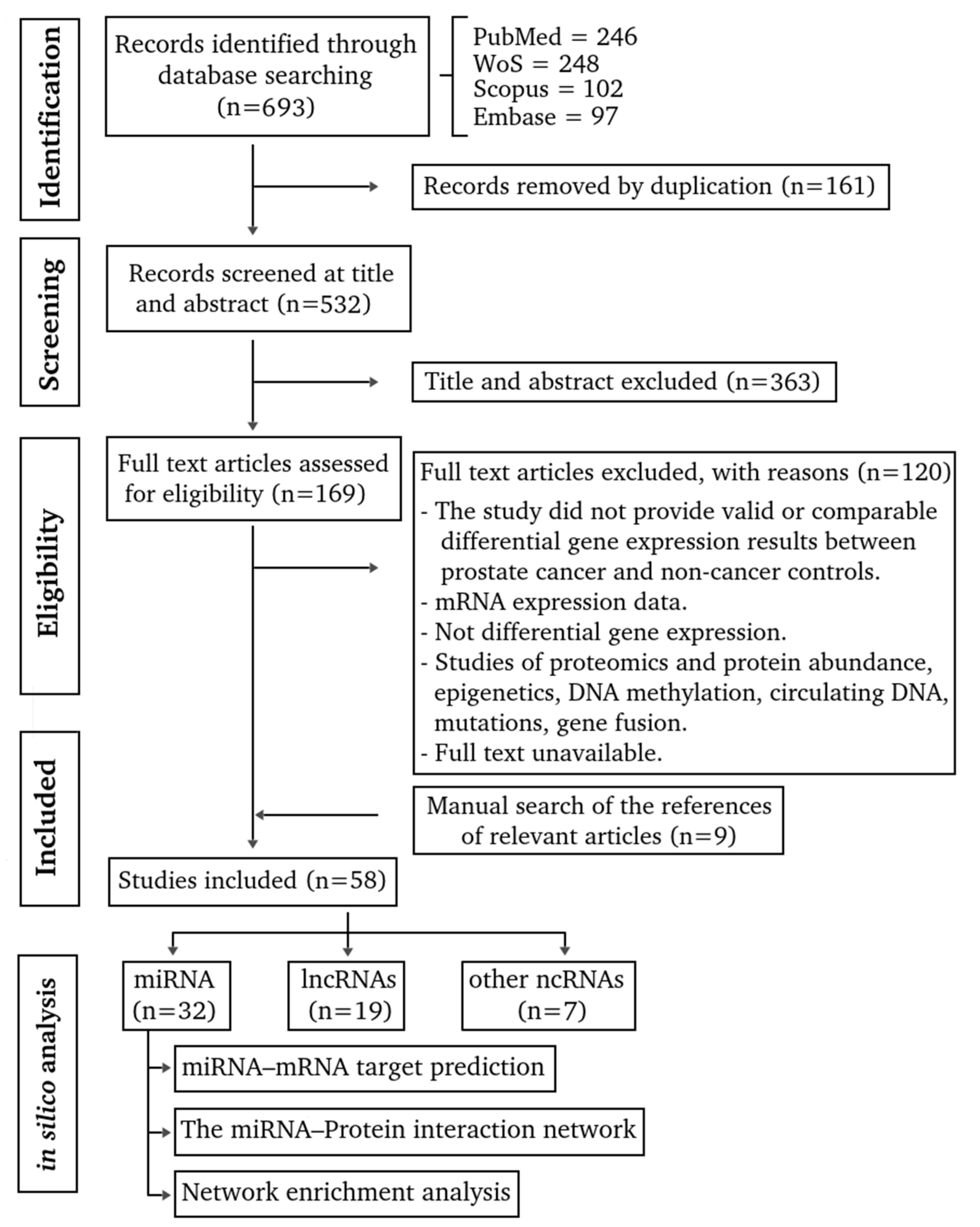
2.1.3. Selection of articles
2.1.4. Data collection process
2.2. In silico analysis
2.2.2. The miRNA–mRNA target interaction network analysis.
2.2.3. Network enrichment analysis.
3. Results
3.1. Evidence Mapping
3.2. The miRNA-based molecular diagnostic biomarkers
3.3. The lncRNA-based molecular diagnostic biomarkers
3.4. Other ncRNA-based molecular diagnostic biomarkers
3.5. miRNA–mRNA target prediction
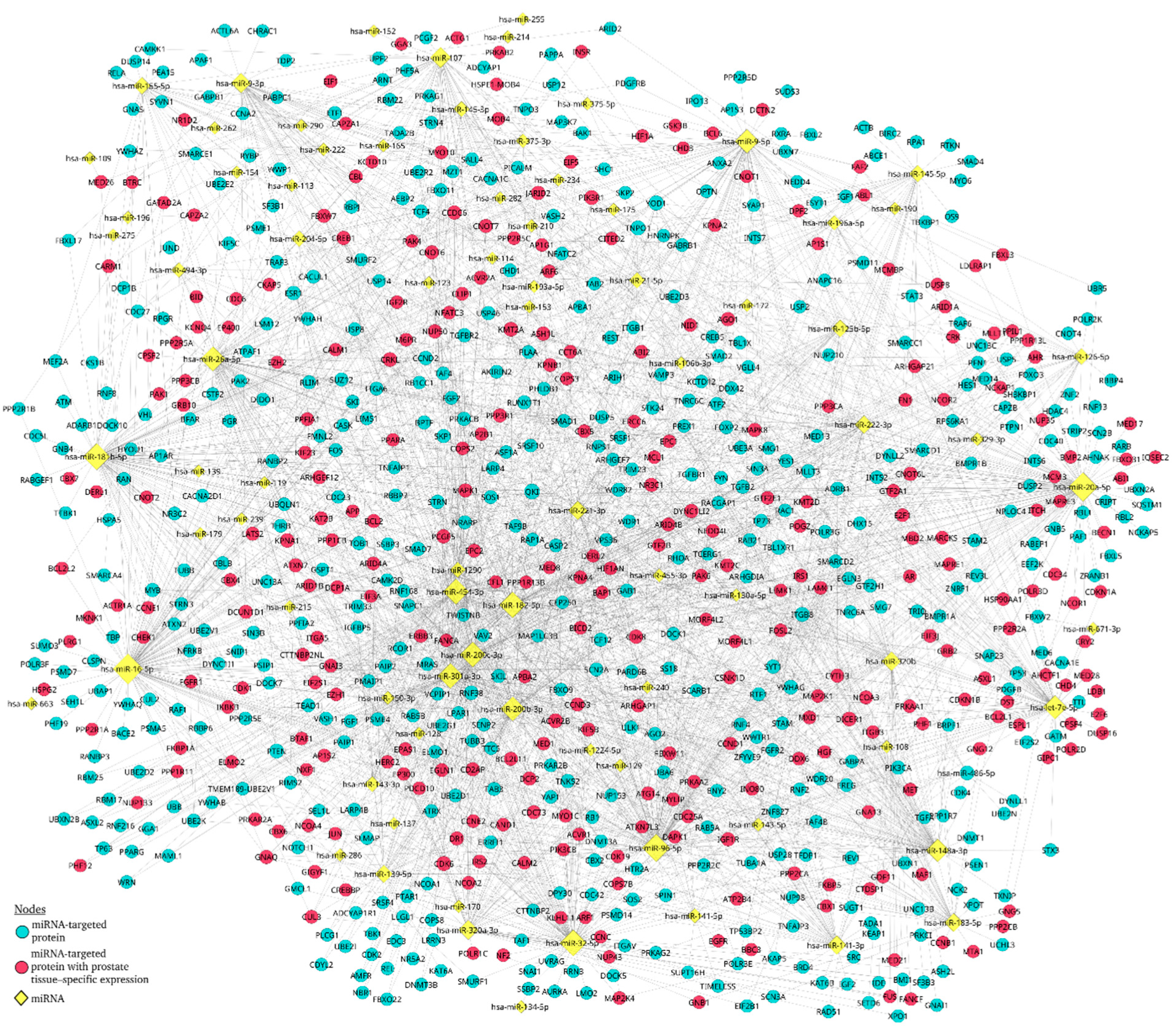
3.6. The miRNA–Protein interaction network
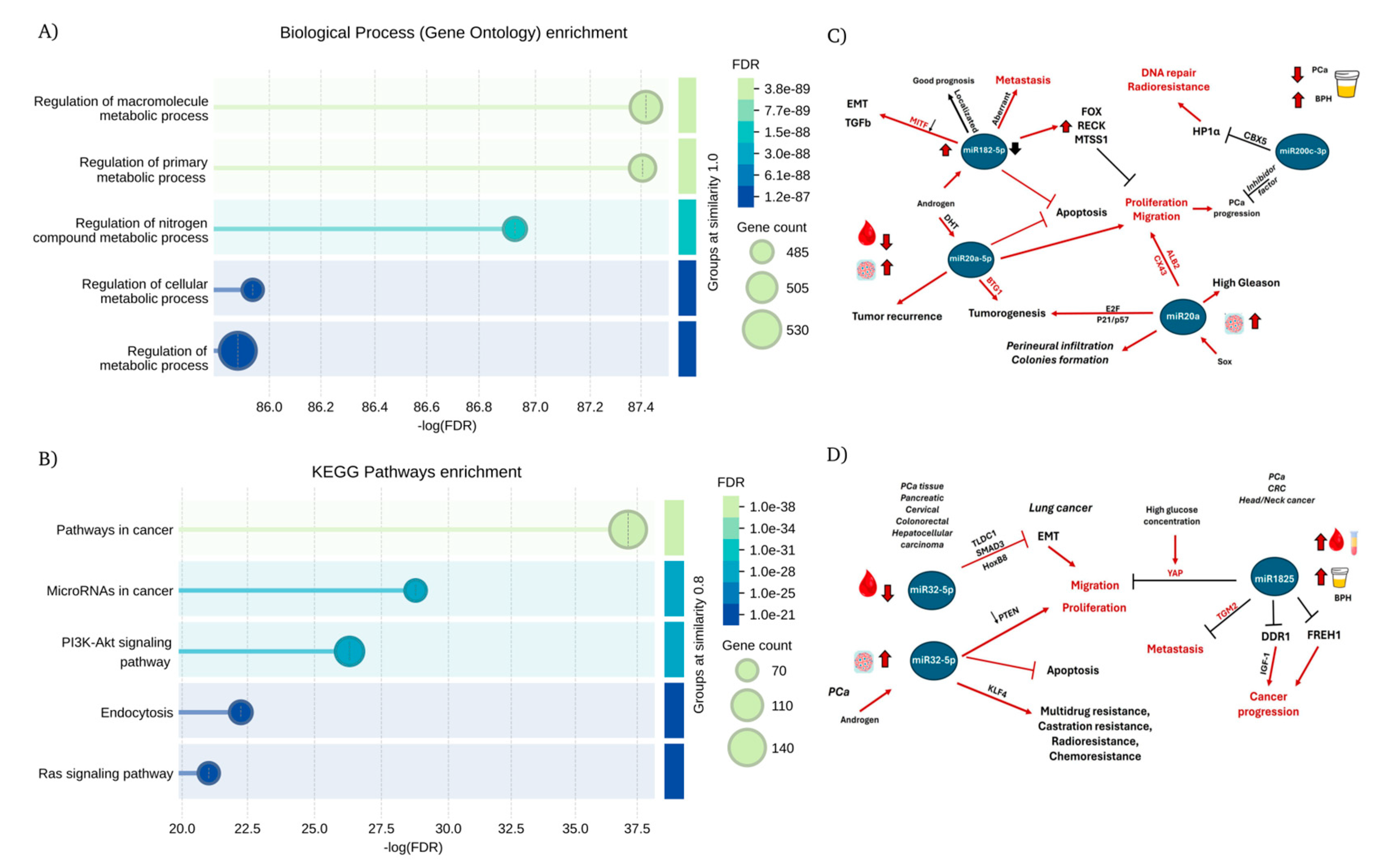
3.7. Network enrichment analysis.
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ACVR2B | Activin A receptor type IIB |
| ANOVA | Analysis of Variance |
| AUC | Area Under the Curve |
| BPH | Benign Prostatic Hyperplasia |
| CDK6 | Cyclin-Dependent Kinase 6 |
| circRNA | Circular RNA |
| DDR1 | Discoidin Domain Receptor 1 |
| EMT | Epithelial–Mesenchymal Transition |
| FG | Fold Change |
| FDR | False Discovery Rate |
| FOX | Forkhead Box Transcription Factors |
| GO | Gene Ontology |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| lncRNA | Long Non-Coding RNA |
| miRNA | microRNA |
| ncRNA | Non-Coding RNA |
| PCa | Prostate Cancer |
| PCA3 | Prostate Cancer Antigen 3 |
| PI3K | Phosphoinositide 3-Kinase |
| piRNA | PIWI-Interacting RNA |
| PRISMA | Preferred Reporting Items for Systematic Reviews and Meta-Analyses |
| PSA | Prostate-Specific Antigen |
| PTEN | Phosphatase and Tensin Homolog |
| qPCR | Quantitative Polymerase Chain Reaction |
| RT-qPCR | Reverse Transcription Quantitative Polymerase Chain Reaction |
| QKI | Quaking Homolog |
| RNU1A-1 | RNA, U1 Small Nuclear 1A |
| ROC | Receiver Operating Characteristic |
| snoRNA | Small Nucleolar RNA |
| snRNA | Small Nuclear RNA |
| STRING | Search Tool for the Retrieval of Interacting Genes/Proteins |
| TBL1XR1 | Transducin Beta-Like 1 X-Linked Receptor 1 |
| UCA1 | Urothelial Cancer Associated 1 |
References
- Bray, F.; Laversanne, M.; Sung, H.; Ferlay, J.; Siegel, R. L.; Soerjomataram, I.; amp; Jemal, A. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: a cancer journal for clinicians 2024, 74, 229–263. [Google Scholar] [CrossRef]
- International Agency for Research on Cancer. GLOBOCAN 2022: Estimated Cancer Incidence, Mortality and Prevalence Worldwide in 2022; IARC: Lyon, France, 2022; Available online: https://gco.iarc.who.int (accessed on 25 October 2025).
- Bergengren, O.; Pekala, K.R.; Matsoukas, K.; Fainberg, J.; Mungovan, S.F.; Bratt, O.; Bray, F.; Brawley, O.; Luckenbaugh, A.N.; Mucci, L.; Morgan, T.M.; Carlsson, S.V. 2022 Update on Prostate Cancer Epidemiology and Risk Factors-A Systematic Review. European urology 2023, 84, 191–206. [Google Scholar] [CrossRef]
- Raychaudhuri, R.; Lin, D.W.; Montgomery, R.B. Prostate Cancer: A Review. JAMA 2025, 333, 1433–1446. [Google Scholar] [CrossRef]
- Abdelmonem, H.B.; Kamal, L.T.; Wardy, L.W.; Ragheb, M.; Hanna, M.M.; Elsharkawy, M.; Abdelnaser, A. Non-Coding RNAs: Emerging Biomarkers and Therapeutic Targets in Cancer and Inflammatory Diseases. Front. Oncol. 2025, 15, 1534862. [Google Scholar] [CrossRef]
- Ahamed, Y.; Hossain, M.; Baral, S.; Al-Raiyan, A.U.; Ashraf, S.B.; Sun, W. The Research Progress on Diagnostic Indicators Related to Prostate-Specific Antigen Gray-Zone Prostate Cancer. BMC Cancer 2025, 25, 1264. [Google Scholar] [CrossRef]
- U.S. Food and Drug Administration. P100033: PROGENSA PCA3 Assay—Premarket Approval. Available online: https://www.accessdata.fda.gov/cdrh_docs/pdf10/P100033A.pdf (accessed on 10 June 2025).
- Bijnsdorp, I.V.; van Royen, M.E.; Verhaegh, G.W.; Martens-Uzunova, E.S. The Non-Coding Transcriptome of Prostate Cancer: Implications for Clinical Practice. Mol. Diagn. Ther. 2017, 21, 385–400. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Tang, X.; Wang, B.; Chen, M.; Zheng, J.; Chang, K. Current Landscape of Exosomal Non-Coding RNAs in Prostate Cancer: Modulators and Biomarkers. Noncoding RNA Res. 2024, 9, 1351–1362. [Google Scholar] [CrossRef] [PubMed]
- Page, M.J.; McKenzie, J.E.; Bossuyt, P.M.; Boutron, I.; Hoffmann, T.C.; Mulrow, C.D.; Shamseer, L.; Tetzlaff, J.M.; Akl, E.A.; Brennan, S.E.; et al. The PRISMA 2020 Statement: An Updated Guideline for Reporting Systematic Reviews. BMJ 2021, 372, n71. [Google Scholar] [CrossRef] [PubMed]
- Albarracín-Navas, L.; Almonte-Becerril, M.; Guerrero, E.; Rivadeneira, J.; Telechea-Fernández, M.; Guzmán, E.; Calderón, F.; Hernández-Leal, M.J.; Otzen, T.; Manterola, C.; Duque, G.; Riffo-Campos, Á.L. Differential Protein-Coding Gene Expression Profile in Patients with Prostate Cancer. Biomedicines 2024, 12, 2509. [Google Scholar] [CrossRef]
- Gene Database. Available online: https://www.ncbi.nlm.nih.gov/gene/ (accessed on 3 November 2025).
- Chen, Y.; Wang, X. miRDB: An Online Database for Prediction of Functional microRNA Targets. Nucleic Acids Res. 2020, 48, D127–D131. [Google Scholar] [CrossRef]
- Agarwal, V.; Bell, G.W.; Nam, J.W.; Bartel, D.P. Predicting Effective microRNA Target Sites in Mammalian mRNAs. eLife 2015, 4, e05005. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.Y.; Lin, Y.C.; Li, J.; Huang, K.Y.; Shrestha, S.; Hong, H.C.; Tang, Y.; Chen, Y.G.; Jin, C.N.; Yu, Y.; et al. miRTarBase 2020: Updates to the Experimentally Validated microRNA–Target Interaction Database. Nucleic Acids Res. 2020, 48, D148–D154. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Kirsch, R.; Koutrouli, M.; Nastou, K.; Mehryary, F.; Hachilif, R.; Gable, A.L.; Fang, T.; Doncheva, N.T.; Pyysalo, S.; et al. The STRING Database in 2023: Protein–Protein Association Networks and Functional Enrichment Analyses for Any Sequenced Genome of Interest. Nucleic Acids Res. 2023, 51, D638–D646. [Google Scholar] [CrossRef] [PubMed]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- The Gene Ontology Consortium. Expansion of the Gene Ontology Knowledgebase and Resources. Nucleic Acids Res. 2017, 45, D331–D338. [Google Scholar] [CrossRef]
- Kanehisa, M.; Furumichi, M.; Sato, Y.; Matsuura, Y.; Ishiguro-Watanabe, M. KEGG: Biological Systems Database as a Model of the Real World. Nucleic Acids Res. 2025, 53, D672–D677. [Google Scholar] [CrossRef]
- Kabassum, R.; Nath, A.; Preininger, M.; Gibson, G. Geographical, Environmental and Pathophysiological Influences on the Human Blood Transcriptome. Curr. Genet. Med. Rep. 2013, 1, 203–211. [Google Scholar] [CrossRef]
- Temilola, D.O.; Wium, M.; Paccez, J.; Salukazana, A.S.; Otu, H.H.; Carbone, G.M.; Kaestner, L.; Cacciatore, S.; Zerbini, L.F. Potential of miRNAs in Plasma Extracellular Vesicle for the Stratification of Prostate Cancer in a South African Population. Cancers 2023, 15, 3968. [Google Scholar] [CrossRef]
- Nonn, L.; Vaishnav, A.; Gallagher, L.; Gann, P.H. mRNA and micro-RNA Expression Analysis in Laser-Capture Microdissected Prostate Biopsies: Valuable Tool for Risk Assessment and Prevention Trials. Exp. Mol. Pathol. 2010, 88, 45–51. [Google Scholar] [CrossRef]
- Mahn, R.; Heukamp, L.C.; Rogenhofer, S.; von Ruecker, A.; Müller, S.C.; Ellinger, J. Circulating microRNAs (miRNA) in Serum of Patients with Prostate Cancer. Urology 2011, 77, 1265.e9–1265.e16. [Google Scholar] [CrossRef]
- Lange, T.; Stracke, S.; Rettig, R.; Lendeckel, U.; Kuhn, J.; Schlüter, R.; Rippe, V.; Endlich, K.; Endlich, N. Identification of miR-16 as an Endogenous Reference Gene for the Normalization of Urinary Exosomal miRNA Expression Data from CKD Patients. PLoS ONE 2017, 12, e0183435. [Google Scholar] [CrossRef] [PubMed]
- Rezaei, S.; Jafari Najaf Abadi, M.H.; Bazyari, M.J.; Jalili, A.; Kazemi Oskuee, R.; Aghaee-Bakhtiari, S.H. Dysregulated microRNAs in Prostate Cancer: In Silico Prediction and In Vitro Validation. Iran J. Basic Med. Sci. 2024, 27, 611–620. [Google Scholar] [CrossRef] [PubMed]
- Daniel, R.; Wu, Q.; Williams, V.; Clark, G.; Guruli, G.; Zehner, Z. A Panel of MicroRNAs as Diagnostic Biomarkers for the Identification of Prostate Cancer. Int. J. Mol. Sci. 2017, 18, 1281. [Google Scholar] [CrossRef] [PubMed]
- Mohammadi Torbati, P.; Asadi, F.; Fard-Esfahani, P. Circulating miR-20a and miR-26a as Biomarkers in Prostate Cancer. Asian Pac. J. Cancer Prev. 2019, 20, 1453–1456. [Google Scholar] [CrossRef]
- Pełka, K.; Klicka, K.; Grzywa, T.M.; Gondek, A.; Marczewska, J.M.; Garbicz, F.; Szczepaniak, K.; Paskal, W.; Włodarski, P.K. miR-96-5p, miR-134-5p, miR-181b-5p and miR-200b-3p Heterogeneous Expression in Sites of Prostate Cancer versus Benign Prostate Hyperplasia—Archival Samples Study. Histochem. Cell Biol. 2021, 155, 423–433. [Google Scholar] [CrossRef]
- Goel, S.; Bergholz, J.S.; Zhao, J.J. Targeting CDK4 and CDK6 in Cancer. Nat. Rev. Cancer 2022, 22, 356–372. [Google Scholar] [CrossRef]
- He, Y.; Xu, W.; Xiao, Y.T.; et al. Targeting Signaling Pathways in Prostate Cancer: Mechanisms and Clinical Trials. Signal Transduct. Target. Ther. 2022, 7, 198. [Google Scholar] [CrossRef]
- Stoen, M.J.; Andersen, S.; Rakaee, M.; Pedersen, M.I.; Ingebriktsen, L.M.; Donnem, T.; Lombardi, A.P.G.; Kilvaer, T.K.; Busund, L.R.; Richardsen, E. Overexpression of miR-20a-5p in Tumor Epithelium Is an Independent Negative Prognostic Indicator in Prostate Cancer—A Multi-Institutional Study. Cancers 2021, 13, 4096. [Google Scholar] [CrossRef]
- Kolokotronis, T.; Majchrzak-Stiller, B.; Buchholz, M.; Mense, V.; Strotmann, J.; Peters, I.; Skrzypczyk, L.; Liffers, S.T.; Menkene, L.M.; Wagner, M.; Glanemann, M.; Betsou, F.; Ammerlaan, W.; Schmidt, R.; Schröder, C.; Uhl, W.; Braumann, C.; Höhn, P. Differential miRNA and Protein Expression Reveals miR-1285, Its Targets TGM2 and CDH-1, as Well as CD166 and S100A13 as Potential New Biomarkers in Patients with Diabetes Mellitus and Pancreatic Adenocarcinoma. Cancers 2024, 16, 2726. [Google Scholar] [CrossRef]
- Guo, X.; Han, T.; Hu, P.; et al. Five MicroRNAs in Serum as Potential Biomarkers for Prostate Cancer Risk Assessment and Therapeutic Intervention. Int. Urol. Nephrol. 2018, 50, 2193–2200. [Google Scholar] [CrossRef]
- Haj-Ahmad, T.A.; Abdalla, M.A.; Haj-Ahmad, Y. Potential Urinary miRNA Biomarker Candidates for the Accurate Detection of Prostate Cancer among Benign Prostatic Hyperplasia Patients. J. Cancer 2014, 5, 182–191. [Google Scholar] [CrossRef]
- Capik, O.; Gundogdu, B.; Tatar, A.; Sahin, A.; Chen, F.; Creighton, C.J.; Karatas, O.F. Oncogenic miR-1825 Promotes Head and Neck Carcinogenesis via Targeting FREM1. J. Cell. Biochem. 2023, 124, 1628–1645. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Luo, J.; Liu, J.; Wu, H.; Li, Y.; Xu, Y.; Liu, L.; Liu, X.; Zhang, Q. Cancer-Secreted Exosomal miR-1825 Induces Angiogenesis to Promote Colorectal Cancer Metastasis. Cancer Cell Int. 2025, 25, 63. [Google Scholar] [CrossRef] [PubMed]
- Baumann, B.; Acosta, A.M.; Richards, Z.; Deaton, R.; Sapatynska, A.; Murphy, A.; Kajdacsy-Balla, A.; Gann, P.H.; Nonn, L. Association of High miR-182 Levels with Low-Risk Prostate Cancer. Am. J. Pathol. 2019, 189, 911–923. [Google Scholar] [CrossRef] [PubMed]
- Stafford, M.Y.C.; McKenna, D.J. miR-182 Is Upregulated in Prostate Cancer and Contributes to Tumor Progression by Targeting MITF. Int. J. Mol. Sci. 2023, 24, 1824. [Google Scholar] [CrossRef]
- Danarto, R.; Astuti, I.; Umbas, R.; Haryana, S.M. Urine miR-21-5p and miR-200c-3p as Potential Non-Invasive Biomarkers in Patients with Prostate Cancer. Turk. J. Urol. 2019, 46, 26–30. [Google Scholar] [CrossRef]
- Labbé, M.; Chang, M.; Saintpierre, B.; Letourneur, F.; de Beaurepaire, L.; Véziers, J.; Deshayes, S.; Cotinat, M.; Fonteneau, J.F.; Blanquart, C.; Potiron, V.; Supiot, S.; Fradin, D. Loss of miR-200c-3p Promotes Resistance to Radiation Therapy via the DNA Repair Pathway in Prostate Cancer. Cell Death Dis. 2024, 15, 751. [Google Scholar] [CrossRef]
- Zhang, J.; Zhang, H.; Qin, Y.; Chen, C.; Yang, J.; Song, N.; Gu, M. MicroRNA-200c-3p/ZEB2 Loop Plays a Crucial Role in the Tumor Progression of Prostate Carcinoma. Ann. Transl. Med. 2019, 7, 141. [Google Scholar] [CrossRef]
- Zhang, J.X.; Yang, W.; Wu, J.Z.; Zhou, C.; Liu, S.; Shi, H.B.; Zhou, W.Z. MicroRNA-32-5p Inhibits Epithelial–Mesenchymal Transition and Metastasis in Lung Adenocarcinoma by Targeting SMAD Family 3. J. Cancer 2021, 12, 2258–2267. [Google Scholar] [CrossRef]
- Yuan, P.; Tang, C.; Chen, B.; Lei, P.; Song, J.; Xin, G.; Wang, Z.; Hui, Y.; Yao, W.; Wang, G.; Zhao, G. miR-32-5p Suppresses the Proliferation and Migration of Pancreatic Adenocarcinoma Cells by Targeting TLDC1. Mol. Med. Rep. 2021, 24, 752. [Google Scholar] [CrossRef]
- Ambrozkiewicz, F.; Karczmarski, J.; Kulecka, M.; Paziewska, A.; Cybulska, M.; Szymanski, M.; Dobruch, J.; Antoniewicz, A.; Mikula, M.; Ostrowski, J. Challenges in Cancer Biomarker Discovery Exemplified by the Identification of Diagnostic MicroRNAs in Prostate Tissues. Biomed Res. Int. 2020, 2020, 9086829. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




