Submitted:
04 December 2025
Posted:
05 December 2025
You are already at the latest version
Abstract
Keywords:
Before You Begin
- Prepare the sucrose solutions needed for polysome gradient formation.
- 2.
- Grow yeast cultures to mid-log phase.
- 3.
- Culture mammalian cells to 70%–80% confluence.
Innovation
Preparation of 5x PGB
- Prepare the 5x Polysome Gradient Buffer (yPGB or mPGB) solution as described in the Materials and Equipment section.
- Filter through a 0.22mm syringe filter.
[Optional] Designing Primers for qPCR-Based Quantification
- 3.
- Design forward and reverse qPCR primers against 25S rRNA.
- 4.
- Identify genes of interest as a set of at least 15 mRNAs that are highly expressed under both standard and stress conditions.
- 5.
- Design forward and reverse qPCR primers specific to the mRNAs of interest in such a way that the forward primer contains the start codon on the mRNA.
Preparation of Sucrose Gradients
- 6.
- Prepare light (eg. 10% w/v) and heavy (eg. 50% w/v) solutions of sucrose by mixing 80% (w/v) sucrose with 5x PGB (mPGB or yPGB depending on cells used, final concentration 1x) and diluting with an appropriate volume of water and mix for 2-3h at 4 °C with gentle rocking.
- 7.
- Degas the solutions by centrifugation at 1500g for 10min.
- 8.
- Mark the half-way point on the 13.2mL ultracentrifuge tube with the help of the marker block
- 9.
- Pour the light solution till the mark.
- 10.
- With a broad-gauge syringe, layer the heavy solution at the bottom of the tube up to the halfway mark. Follow STAR Protocol #4182 for a detailed video [3].
- 11.
- Then place the appropriate caps on the tubes and remove excess liquid that may have collected in the cap.
- 12.
- Use the Gradient Master part of the Gradient Station Machine to make the gradient. Use the appropriate programs corresponding to the gradient used.
- 13.
- Keep at 4 °C overnight until it is time to load the samples.
Key Resources Table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
| Chemicals, peptides, and recombinant proteins | ||
| Cycloheximide | Thermo Scientific | Cat#357420050 |
| GlycoBlue | Thermo Scientific | Cat#AM9516 |
| RNasIN Plus | Promega | Cat#N2615 |
| SuperScript IV | Thermo Scientific | Cat#18090200 |
| Nuclease Free Water | Thermo Scientific | Cat#AM9932 |
| Random hexamers | Promega | Cat#C1181 |
| oligodT15 | Promega | Cat#C1101 |
| KAPA SYBR Fast | Sigma Aldrich | Cat#KK4611 |
| Tris-HCl pH 8.0 | Thermo Scientific | Cat#AM9856 |
| MgCl2 | Thermo Scientific | Cat#AM9530G |
| DTT | Thermo Scientific | Cat#707265ML |
| KCl | Thermo Scientific | Cat#AM9640G |
| Cycloheximide (RNA grade) | Thermo Scientific | Cat#J66004-XF |
| cOmplete EDTA free Protease Inhibitor | Sigma Aldrich | Cat#COEDTAF-RO |
| Sucrose | Thermo Scientific | Cat#036508-30 |
| Phenol (pH 4.5) | Sigma Aldrich | Cat#P4682-400ML |
| Chloroform-Isoamyl Alcohol (24:1) | Sigma Aldrich | Cat#25666-500ML |
| Absolute Ethanol | Sigma Aldrich | Cat#1009831000 |
| Triton X-100 | Sigma Aldrich | Cat#T9284 |
| Phosphate buffered saline (PBS) | Sigma Aldrich | Cat#D8537 |
| Critical commercial assays | ||
| RNeasy MinElute Cleanup Kit | Qiagen | Cat#74204 |
| RNA Clean and Concentrator | Zymo | Cat#R1013 |
| Qiaseq Fastselect Yeast Kit | Qiagen | Cat#334217 |
| SMART-Seq Stranded | TaKaRa | Cat#634444 |
| TranscriptAid T7 High Yield Transcription Kit | Thermo Scientific | Cat#K0441 |
| TruSeq Stranded kit | Illumina | Cat#20020599 |
| Deposited data | ||
| Polysome profiling of murine macrophages without rRNA depletion, at 37° C (Control) and 42° C (Heat-shock) | This paper | GEO: GSE311393 |
| Experimental models: Cell lines | ||
| Mouse: J774A.1 Murine macrophages | Laboratory of Prof. Dr. Jean Pieters | ATCC: TIB-67 |
| Experimental models: Organisms/strains | ||
| S. cerevisiae: Strain background: BY4743 | EUROSCARF | Y20000 |
| Oligonucleotides | ||
| Sc_RDN25-1_qPCR_ForwardPrimer | AATCTCGCATTTCACTGGGC | 9608 |
| Sc_RDN25-1_qPCR_ReversePrimer | TTGACTTACGTCGCAGTCCT | 9614 |
| Mm_RN28S1_qPCR_ForwardPrimer | GGGTTTAGACCGTCGTGAGA | 10133 |
| Mm_RN28S1_qPCR_ReversePrimer | CTCAGCCAAGCACATACACC | 10139 |
| Other | ||
| Zirconia/Silica beads | Biospec | Cat#11079105z |
| Ultracentrifuge tubes 13.2mL | Beckman Coulter | Cat#C14293 |
| Caps for Ultracentrifuge tubes | BioComp | Cat#105-414-1 |
| SW41-Ti Ultracentrifuge Rotor | Beckman Coulter | Cat#331362 |
| Lysing Matrix Y | MPBio | Cat#116960050-CF |
| Teenprep™ Lysing Matrix Y tubes | MPBio | Cat#116975050 |
| Lightcycler 480 | Roche | RRID:SCR_018626 |
| Gradient Station | BioComp | Cat#153 |
| Marker Block (for SW41-Ti Rotor) | BioComp | Cat#105-614A |
| 23G syringe | B Braun | Cat#4665635 |
| 5mL microfuge tube (DNA Lobind) | Eppendorf | Cat#0030108310s |
| 2mL microfuge tube (DNA Lobind) | Eppendorf | Cat#0030108078 |
Materials and Equipment
| Reagent | Final concentration |
| Tris-HCl pH 8.0 | 100mM |
| MgCl2 | 50mM |
| KCl | 250mM |
| Dithiothreitol (DTT) | 5mM |
| Cycloheximide (RNA grade) | 1mg/mL |
| cOmplete EDTA free Protease Inhibitor | 1 tab/10mL |
| RNasIN RNase Inhibitor | 80U/mL |
| Nuclease Free Water | n/a |
| Reagent | Final concentration |
| Tris-HCl pH 8.0 | 100mM |
| MgCl2 | 50mM |
| Dithiothreitol (DTT) | 5mM |
| Cycloheximide (RNA grade) | 1mg/mL |
| cOmplete EDTA free Protease Inhibitor | 1 tab/10mL |
| RNasIN RNase Inhibitor | 40U/mL |
| Nuclease Free Water | n/a |
| Reagent | Final concentration |
| 5x PGB (mPGB or yPGB) | 1x |
| Triton X-100 | 1% |
| Nuclease Free Water | N/A |
| Reagent | Final concentration |
| Cycloheximide | 300µg/mL |
| Yeast culturing media (eg. CSM Complete) | N/A |
| Reagent | Final concentration |
| Cycloheximide | 200µg/mL |
| 1x PBS | N/A |
Step-by-Step Method Details
Collection and Lysis of Cells
-
Collect the cells as follows:
-
Option 1: Yeast cells (use at least 2x108 cells):
-
Add 1x volume of yeast culture (e.g. 30mL) to 2x volume of Yeast cell collection media (e.g. 60mL). Immediately mix by inversion and cool in a 50% Ethanol bath at -20 °C for 2 minutes.CRITICAL: Use of ice-cold collection media and cooling in a -20 °C Ethanol bath helps stop translation process immediately by dropping the temperature in the tube to <10 °C, as opposed to stopping it in a gradual manner, which would potentially lead to ribosomal run-off [4].
- Centrifuge at 1500g for 10min in a pre-cooled centrifuge at 4 °C.
- Completely remove supernatant and proceed to lysis.
-
-
Option 2: Mammalian Cells (use at least 1x107 cells):
- Remove media by aspiration and immediately place the culture dish on ice
- Add Mammalian cell collection media cooled to 4 °C. to the culture dish and scrape the cells.
- Collect the cells in an appropriate tube and centrifuge at 1500g for 10min in a pre-cooled centrifuge at 4 °C.
- Completely remove supernatant and proceed to lysis.
-
-
Lyse the cells as follows:
-
Option 1: Yeast cells:
- Re-suspend the cells in 600µL of 1x yPLB on ice.
- Add the mixture to 2mL microfuge tubes containing 500µL Zirconia/Silica beads
- Lyse at 4 °C in a Thermomixer by intermittent vortexing at 2000rpm, 30s on 30s off, 20 cycles.
-
Option 2: Mammalian cells:
- Re-suspend the cells in 600µL of 1x mPLB on ice.
- Pass the mixture 10 times through a sterile 23G needle.
- Carry out passive lysis by incubating on ice for 10min.
- Pass the mixture 10 times through a sterile 23G needle.
- Centrifuge at 4 °C, 10000g for 10min and collect the supernatant.
-
Measure A260 using a spectrophotometer.Note: Due to the presence of RNase Inhibitors, Protease Inhibitors and detergents in PLB, all of which absorb in the UV range, it will be necessary to dilute the sample at least 10-fold in water for A260 measurement.Pause point: Samples may be immediately used for polysome profiling or flash frozen in liquid nitrogen and stored at -80 °C for several months. Thaw slowly on ice or a metal block and resuspend before polysome profiling.
-
Ultracentrifugation and Polysome Profiling
- 3.
- Pre-cool the ultracentrifuge to 4 °C.
- 4.
- Layer 2-5 A260 units of the lysate on the top of the sucrose gradients in the ultracentrifuge tube.
- 5.
- Weigh the ultracentrifuge tubes along with the buckets and make sure they are balanced
- 6.
- Load the buckets containing the tubes into the rotor and carry out the ultracentrifugation at 275000g (equivalent to 40,000 rpm in an SW41Ti rotor) for 2h.
- 7.
- When the ultracentrifugation is about to finish (<10min left), switch on the Gradient Station machine and carry out the cleaning, blanking and general quality control steps as instructed in the TRIAX program in the computer attached to the Gradient Station.
- 8.
- Load 2mL tubes in the fractionation rack (code 19) as instructed and place it in the Gilson collector unit.
- 9.
- Carefully take out the rotor from the ultracentrifuge and keep the buckets with the tubes at 4 °C without disturbing the gradients.
- 10.
- Carefully take out the ultracentrifuge tube from the bucket and place it in the tube holder from the Gradient station machine under the piston.
- 11.
- Fractionate the gradient, collect the fractionated samples and save the polysome profile trace generated by the TRIAX program.
- 12.
- Keep the fractions in ice if used immediately or store at -80 °C.
RNA Extraction
- 13.
- Prepare Phenol:Chloroform:Isoamyl Alcohol = 25:24:1 mix by mixing equal volumes of acid phenol (pH = 4.5) and chloroform:isoamyl alcohol (24:1) and keep at 37 °C.
- 14.
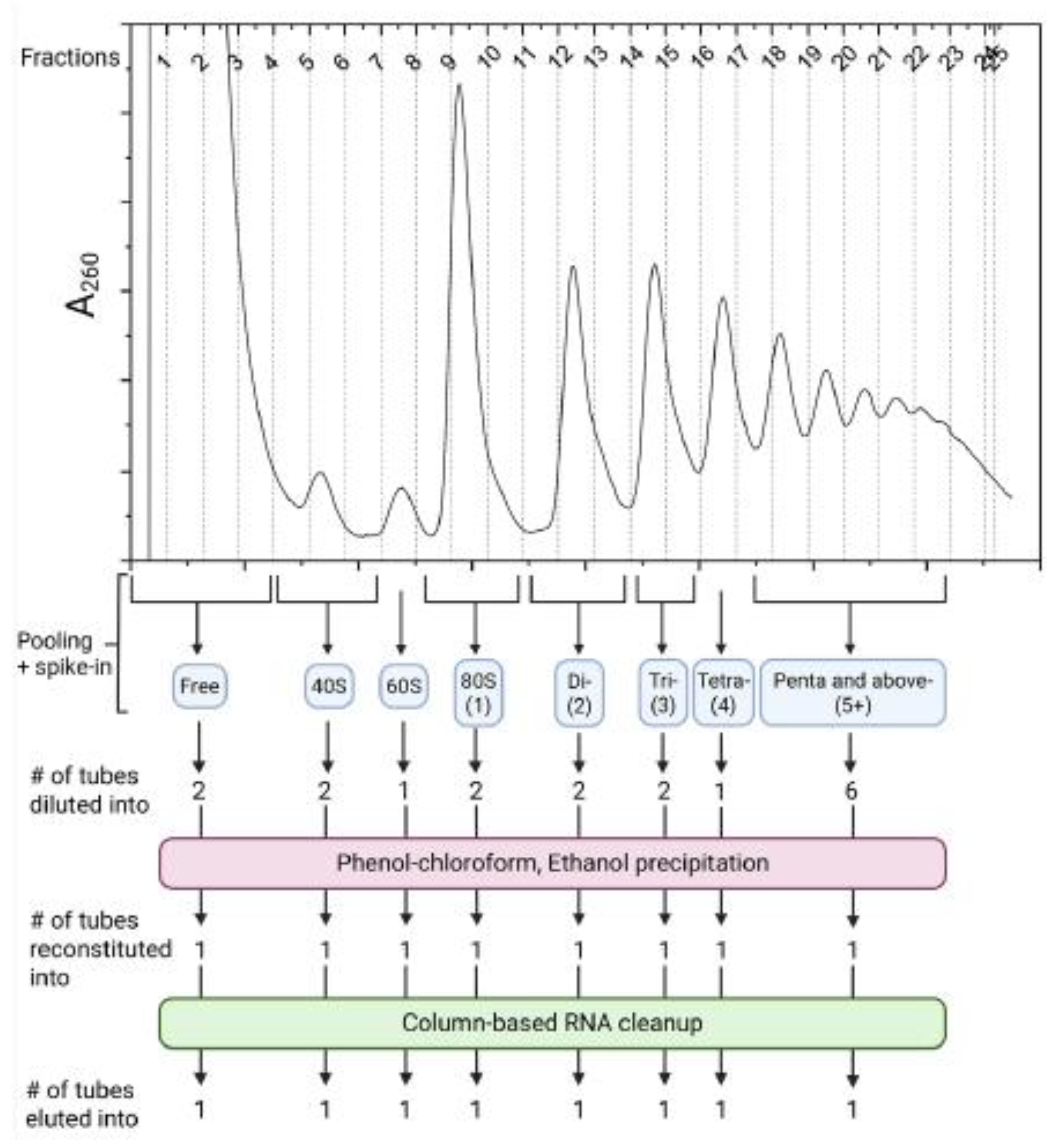
- From the polysome profile trace, identify the fractions (collected in step 11) corresponding to a particular mRNA-ribosome complexation state (e. Free, 40S, 60S, Monosome (80S), Disome, etc.)
- 15.
- Pool the fractions corresponding to each particular mRNA-ribosome complexation state together in a 5mL microfuge tube and add equal volumes of spike-in RNAs to each pool.
- 16.
- If any pool exceeds 1200 µL, divide it into multiple 5-mL microfuge tubes as needed. Add nuclease free water to a final volume of 1200µL.
- 17.
- If any tube contains a pooled sample with a sucrose concentration >30%, split into multiple 5mL microfuge tubes as needed. Add nuclease free water to a final volume of 1200µL.
- 18.
- Add 1200µL of the hot phenol-chloroform-isoamyl alcohol mix (prepared in step 13) to each 5mL microfuge tube containing 1200µL of sample. Mix by vortexing and let stand for 1min.
- 19.
- Centrifuge the 5mL microfuge tubes at 40 °C for 10min at a speed of 16000g.
- 20.
- Pipet out 1mL of the upper (aqueous) phase into a fresh 5mL microfuge tube.
- 21.
- Add 100µL of Sodium acetate (pH = 5.0), 2µL of glycoblue and 3.3mL of absolute ethanol.
- 22.
-
Vortex the tubes and keep at -80 °C overnight for precipitation.Pause point: The tubes can be stored at -80 °C for up to a year.
- 23.
- Prepare 75% ethanol and keep chilled in ice.
- 24.
- Centrifuge the tubes at 4 °C for 1h at a speed of 20000g.
- 25.
- Discard the supernatant while taking care to not disturb the pellet (blue)
- 26.
- Add 2mL of 75% ethanol and vortex to wash the pellet.
- 27.
- Centrifuge at 4 °C for 10min at a speed of 20000g.
- 28.
- Completely remove any traces of ethanol and let dry for 1min on the bench.
- 29.
- Reconstitute the pellets from each pool in warm nuclease free water (at 50 °C). For pools which have been separated into multiple tubes, dissolve one pellet first and then use the resulting solution to dissolve the next one from the same pool.
- 30.
- Further purify this RNA using column-based cleanup (e.g. RNeasy Minelute) including the on-column DNase digestion step. Elute in 30µL of nuclease free water.
Quantification
- 31.
-
Measure the RNA intensities as follows:
-
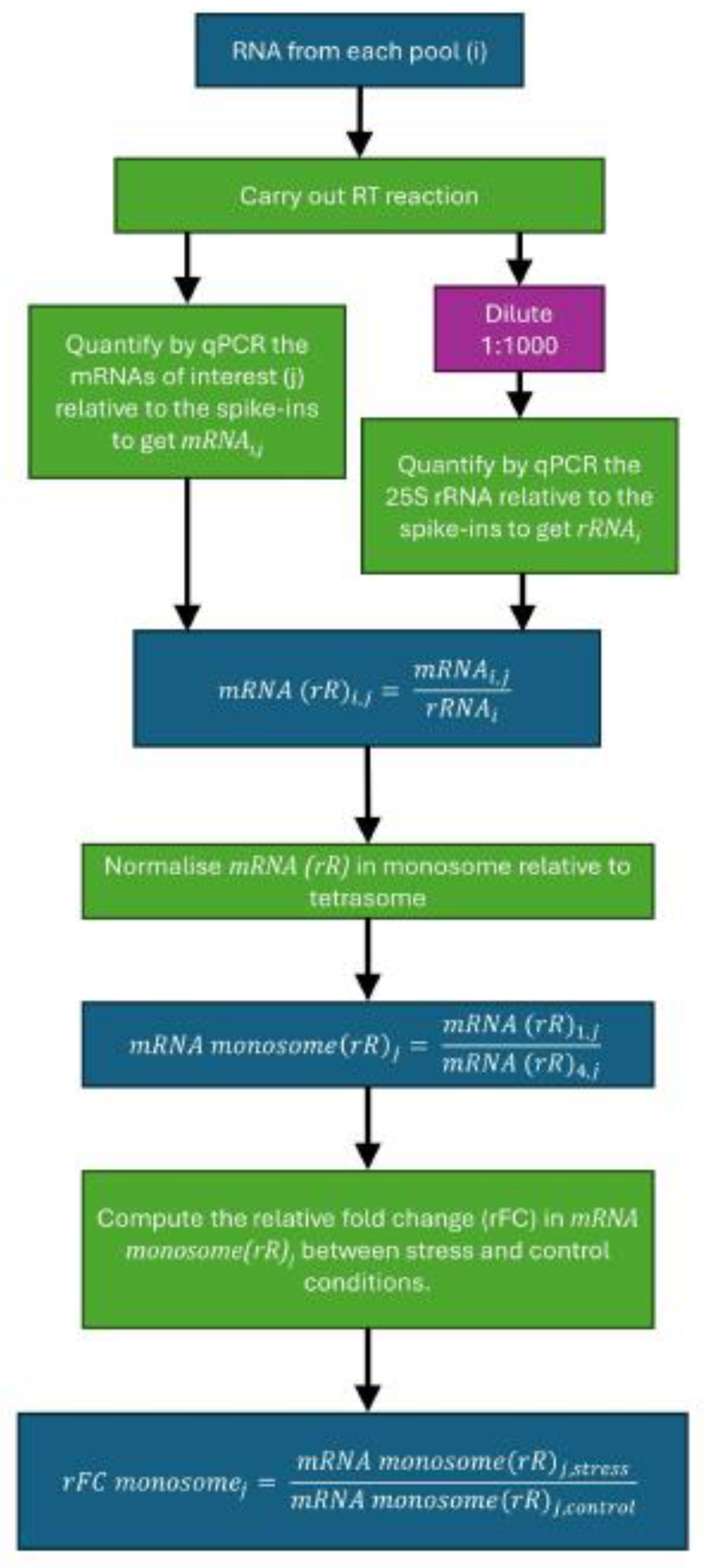
Option 1: qPCR: Scheme in Figure 2.
- Set up a reverse transcription reaction with 5µL of the eluted RNA using SuperScript IV. Use oligo(dT)15, 25S rRNA reverse primer as well as the reverse primers for the spike-in RNAs as the RT primers.
- Carry out reverse transcription at 50 °C for 1h.
- Probe the cDNA generated in step i. for the genes of interest by qPCR using primers flanking the start codon as described in preparation step “Designing primers for qPCR-based quantification,” while also measuring the spike-in RNAs as control.
-
Dilute the cDNA generated in step i. 1:1000 and probe for 25S rRNA by qPCR, while also measuring the spike-in RNAs as control.Note: 25S rRNA (present in the large ribosomal subunit) rather than the 18S rRNA (present in the small subunit) was used in order to prevent signal from the scanning 40S ribosomal subunits that could be associated with mRNAs in each peak of the polysome profile (Troubleshooting 4).CRITICAL: If this heavy dilution is not carried out, the rRNA Ct values will be too low to be measured correctly.
- Normalize both the mRNAs of interest (j) and the 25S rRNA to the spike-in RNA intensities for each pool (i) to get mRNAi,j and rRNAj respectively.
- Calculate mRNA relative to rRNA for each mRNA of interest in each pool as follows:
-
For each mRNA, normalize the ratio of mRNA to rRNA in monosome relative to tetrasome, as follows:Note: Here 1 refers to the monosome, 4 to the tetrasome (4 ribosomes) and j is the species of mRNA.
-
Compute the relative fold change (rFC) in monosomal mRNA/rRNA ratios between stress (e.g., heat shock) and control conditions for each mRNA species j:Note: A lower rFC monosome value indicates a higher prevalence of silent ribosomes.
-
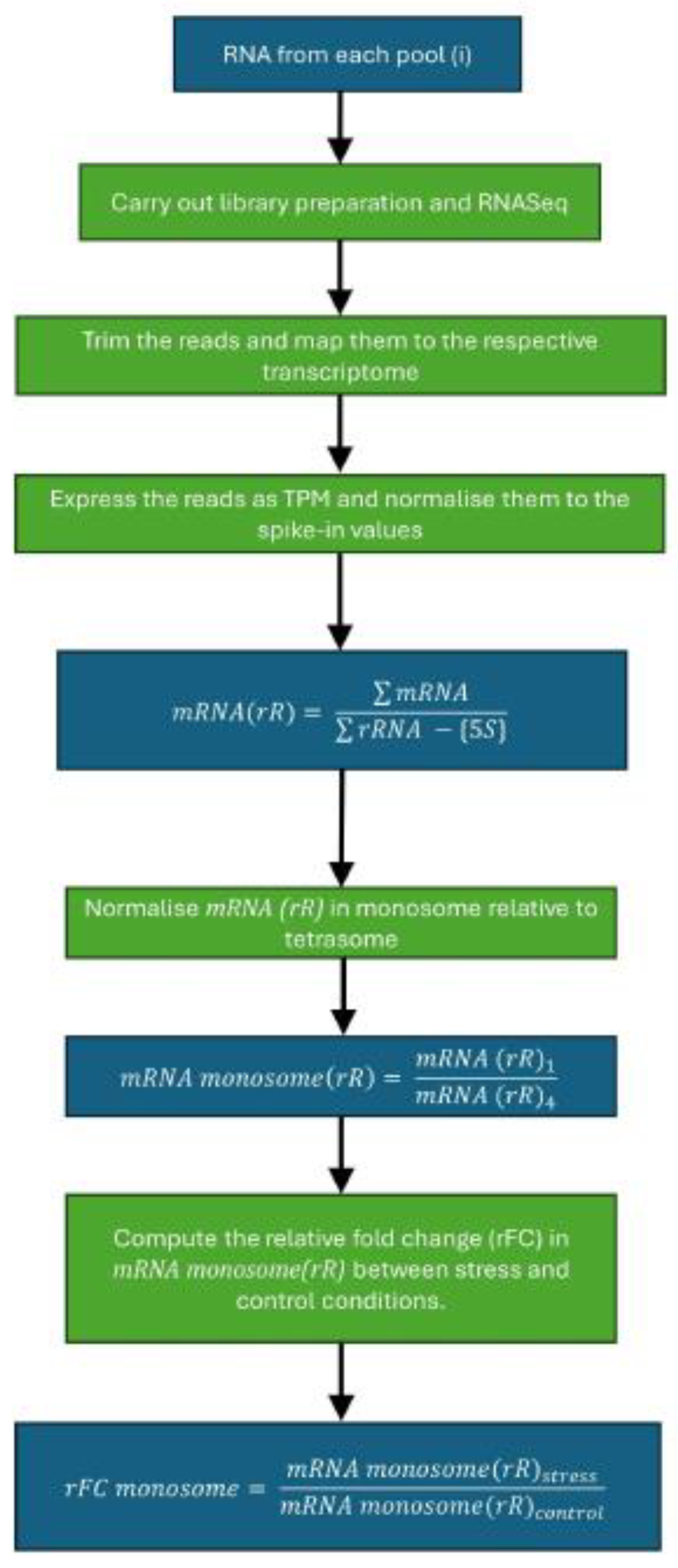
Option 2: RNA-Seq: Scheme in Figure 3.
- Quantify the eluted RNA from step 30 using Qubit or any other fluorescence-based measurement technique. Troubleshooting 5.
-
Carry out library preparation on the RNA using SMART-Seq Stranded kit if using yeast samples and TruSeq Stranded kit if using mammalian samples. Follow the kit instructions for library preparation and downstream quality control.CRITICAL: Since here we are using the rRNA levels as a readout for ribosomes and using this to calculate silent ribosomes, it is imperative that no mRNA enrichment or rRNA depletion step is carried out.
- Carry out RNA-Seq on the prepared libraries.
- Trim the reads from the RNA-Seq using TrimGalore and map them onto the respective transcriptome (along with the spike-ins) using Salmon [5].
- Express the reads as transcripts-per-million (TPM) and normalize them to the spike-in values.
- For each pool, calculate mRNA relative to rRNA as follows:Note: 5S rRNA has been seen to display anomalous amplification which may be linked to its small size. As a result, it is not used as a readout for rRNA levels [1].
- Normalize the ratio of mRNA to rRNA in monosome (1) relative to tetrasome (4) as follows:
- Compute the relative fold change (rFC) in monosomal mRNA/rRNA ratios between stress (e.g., heat shock) and control conditions:
-
Expected Outcomes
Limitations
Troubleshooting
Problem 1: Peaks Are Not Well Defined.
Problem 2: Peaks Are Too Congested for Fractionation.
Potential Solution:
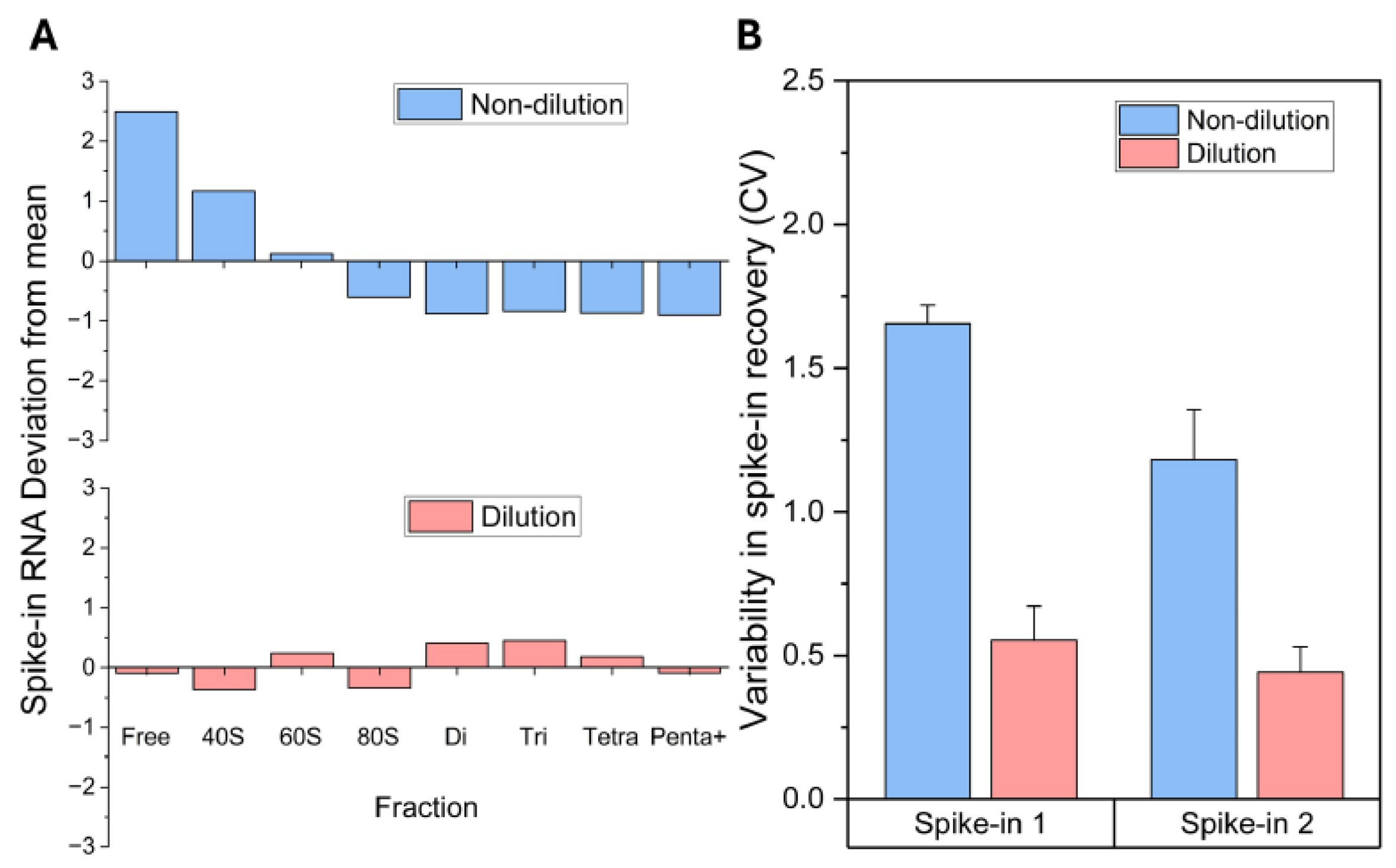
Problem 3: Spike in Levels Show a Large Deviation Across Pools.
Potential Solution:
Problem 4: Low Ct Value of 25S rRNA
Potential Solution:
Problem 5: Low RNA Yield
Potential Solution:
Resource Availability
Lead Contact
Technical Contact
Materials Availability
Data and Code Availability
Author Contributions
Acknowledgments
Declaration of Interests
References
- Rahaman, S., Schiffelholz, N., Mittal, N., Frohlich, K.E., Zavolan, M., and Becskei, A. (2025). Heat shock induces silent ribosomes and reorganizes mRNA turnover. Cell Rep 44, 116447. [CrossRef]
- Delaney, C.E., Becskei, A. (2025). Detection and Function of the Eukaryotic Vacant Ribosome. Preprints. [CrossRef]
- De Siqueira, M.K., Nouhi, Z., Zhao, Y., Wang, S., Xiao, X., Yang, X., Hulea, L., and Villanueva, C.J. (2025). Protocol to perform polysome profiling in primary differentiating murine adipocytes. STAR Protoc 6, 103799. [CrossRef]
- Masek, T., Valasek, L., and Pospisek, M. (2011). Polysome analysis and RNA purification from sucrose gradients. Methods Mol Biol 703, 293–309. [CrossRef]
- Patro, R., Duggal, G., Love, M.I., Irizarry, R.A., and Kingsford, C. (2017). Salmon provides fast and bias-aware quantification of transcript expression. Nat Methods 14, 417–419. [CrossRef]
- Rahaman, S., Faravelli, S., Voegeli, S., and Becskei, A. (2023). Polysome propensity and tunable thresholds in coding sequence length enable differential mRNA stability. Sci Adv 9, eadh9545. [CrossRef]
- Jaquet, V., Wallerich, S., Voegeli, S., Turos, D., Viloria, E.C., and Becskei, A. (2022). Determinants of the temperature adaptation of mRNA degradation. Nucleic Acids Res 50, 1092–1110. [CrossRef]
- Shalgi, R., Hurt, J.A., Krykbaeva, I., Taipale, M., Lindquist, S., and Burge, C.B. (2013). Widespread Regulation of Translation by Elongation Pausing in Heat Shock. Molecular Cell 49, 439–452. [CrossRef]
- Liu, B., Han, Y., and Qian, S.B. (2013). Cotranslational response to proteotoxic stress by elongation pausing of ribosomes. Mol Cell 49, 453–463. [CrossRef]
- Liu, B., and Qian, S.B. (2016). Characterizing inactive ribosomes in translational profiling. Translation (Austin) 4, e1138018. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



