1. Introduction
Seeds play a vital role in plant survival, enabling species to persist through unfavorable conditions, disperse across environments and protect embryos from stresses such as desiccation [
1]. Seed security is an essential yet often overlooked component of global food security [
2]. As the foundation of agriculture, seeds serve as the primary reproductive unit of plants and the vehicle for delivering genetic potential that supports crop productivity, diversity and resilience in changing environments [
3]. The use of high-quality seed ensures uniform germination, vigorous seedling growth and higher yields, whereas poor-quality or contaminated seed can result in weak establishment and significant yield losses [
4]. Moreover, seed health encompassing purity, germination capacity and free from pathogens and pests directly influences crop performance and long-term agricultural sustainability [
5].
Seed health testing continues to play a critical role, even with rapid advances in diagnostics and seed technology. The international trade of seeds has increased substantially with globalization, expanding both commercial opportunities and phytosanitary risks, which underscores the continuing importance of effective seed health testing [
6]. A single contaminated lot may introduce exotic pathogens into new regions, potentially disrupting production or triggering regulatory action, consistent with known risks of contaminants in traded commodities [
7]. Seedborne pathogens can significantly reduce crop yields and seed quality, leading to economic losses for growers and potential disruptions in market supply chains [
8,
9,
10]. Beyond these direct impacts, ensuring seed health is also a crucial component of food safety, as it mitigates the introduction of toxin-producing and human-pathogenic microorganisms into the food supply. The implementation of rigorous seed sanitation and treatment protocols reduces the dependence on post-emergence disease management practices, contributing to improved crop health and minimizing the risk of contaminant introduction into the food production system [
11].
Because seed health affects many aspects of crop production and trade, it is important to clearly define what it entails. In this context, the term refers to the overall physiological and pathological condition of the seed, encompassing both its physical quality and free from infectious agents [
12]. Within this framework, seedborne, seed-associated, and surface contamination are distinct but related terms. Seedborne pathogens are internally present within the seed tissues and can be transmitted to the seedling [
13,
14]. Seed-associated microorganisms may occur on or within the seed without necessarily being transmitted or causing disease [
15]. Surface contamination refers to the presence of pathogens adhering to the outer seed surface, which can often be managed through disinfestation or seed treatment. Seed disinfestation targets spores and other propagules on the seed surface, whereas disinfection aims to eliminate pathogens that have penetrated the internal seed tissues [
16].
Despite improvements in technology, several challenges continue to limit seed health testing. Testing seeds for plant pathogens can be challenging, as infested seeds are often asymptomatic, pathogen populations occur at very low levels, and contamination may be unevenly distributed within a seed lot. These factors make detection difficult and may require large sample sizes or highly sensitive assays to achieve reliable results [
17]. The seed industry ensures the delivery of healthy seeds while complying with national and international phytosanitary regulations. Seed health testing combines direct methods, which confirm pathogen viability and pathogenicity, with indirect methods such as ELISA, PCR, or high-throughput sequencing for rapid prescreening [
12]. While molecular assays are sensitive, they may detect noninfectious fragments and require confirmation by classical tests [
12]. The integration of both approaches allows accurate risk assessment, supporting safe international seed movement and global food security [
18,
19]. This review highlights recent developments and ongoing challenges in detecting and managing seedborne fungi, bacteria, and viruses/viroids, with brief notes on oomycetes and nematodes that can occasionally be associated with seed movement. Emphasis is given on diagnostic advances, harmonization efforts, and regulatory perspectives, illustrating why seed health testing remains indispensable in the modern era of global agriculture.
2. Regulatory & Standards Landscape
Advances in molecular diagnostics, high resolution imaging, and emerging artificial intelligence (AI) assisted detection tools are increasingly shaping seed health testing, making regulatory harmonization essential to ensure that new technologies are validated, recognized, and consistently applied across trading systems [
12,
13,
20]. The regulation of seed health is supported by a network of international, regional, and national organizations that develop testing standards, diagnostic protocols, and certification systems. Together, these frameworks ensure that seeds moving through domestic and international trade channels are tested, certified, and managed consistently to minimize phytosanitary risk while enabling market access [
18]. At the global level, several key bodies play distinct yet complementary roles. The International Seed Testing Association (ISTA) establishes standardized seed testing methods, including molecular assays that are increasingly incorporated into routine diagnostics, and issues certificates widely recognized in international trade [
21]. In the United States, the Association of Official Seed Analysts (AOSA) and the Society of Commercial Seed Technologists (SCST) maintain and standardize seed testing standards for both regulatory and commercial purposes, with ongoing discussions regarding integration of qPCR, LAMP, and other molecular technologies into official rulebooks [
22]. Within Europe, the European and Mediterranean Plant Protection Organization (EPPO) develops diagnostic protocols and plant health standards, including detailed molecular and serological methods for detecting seedborne pathogens [
23]. For the vegetable seed sector, the International Seed Health Initiative for Vegetable Crops (ISHI Veg) focuses on standardized diagnostic methods and validation data and is currently evaluating next generation sequencing (NGS) approaches and digital imaging tools to establish pathways for future standardization [
18]. Broader phytosanitary oversight is provided by the International Plant Protection Convention (IPPC), which develops International Standards for Phytosanitary Measures (ISPMs) that guide how countries regulate the movement of plants and seeds to prevent the spread of quarantine pests [
24,
25]. Complementing these, the OECD Seed Schemes provide an internationally recognized framework for varietal certification and facilitate movement of seed among participating countries under standardized labeling and quality systems [
26].
Accreditation and validation form the technical foundation that ensures confidence in test results and comparability between laboratories [
22]. Many seed testing laboratories operate under ISO/IEC 17,025 accreditation, which establishes requirements for technical competence, quality management, and method reliability [
27]. New diagnostic methods, particularly qPCR, LAMP, NGS, high throughput imaging, and emerging AI assisted classification tools, undergo formal validation pathways that assess parameters such as analytical sensitivity, specificity, repeatability, and reproducibility [
28]. Collaborative ring trials and proficiency testing are essential components of this process, allowing multiple laboratories to evaluate the same method under comparable conditions [
29]. These exercises help identify variability, refine protocols, and demonstrate that a test can perform reliably across different operators and environments, ultimately supporting broader acceptance by regulators and trading partners [
30]. However, while validation frameworks for molecular assays are well established, comparable regulatory pathways for AI based image analysis and digital diagnostics are still under development, creating uncertainty around how such tools will be incorporated into official standards.
Despite these advances, achieving full standardization across laboratories and regulatory systems remains challenging. Discrepancies persist in the choice of target pathogens, sample sizes and sampling strategies, and acceptable thresholds for detection or tolerance [
30]. Some standards emphasize presence or absence criteria, while others set quantitative thresholds based on inoculum levels or disease risk [
22,
31]. Rapid molecular tests may detect non-viable or residual pathogen DNA, raising questions about the biological relevance of positive results [
32]. The introduction of high throughput sequencing and AI assisted imaging further complicates interpretation, as these technologies can generate highly sensitive outputs whose regulatory significance is not yet fully defined. Additionally, differences in regulatory interpretation and resource capacity among countries contribute to inconsistent implementation. For example, studies examining seed sector regulation in Sub Saharan Africa have found that, despite regional standardization efforts, variations in national legal frameworks and enforcement capacities continue to create gaps between written law and practical application [
33]. Because of these differences, a seed lot that meets the standards in one system may not comply with another, causing uncertainty for exporters, labs, and regulators.
The ongoing evolution of seed trade, combined with rapid innovation in diagnostic technologies, underscores the need for greater international coordination. Building consensus on validation frameworks, data sharing, performance criteria, and risk-based decision making will be key to aligning scientific advances, particularly molecular, digital, and AI assisted tools, with regulatory trust [
33]. Only through such standardization can the global seed industry maintain both the efficiency of commerce and the integrity of plant health protection.
3. Sampling Theory & Study Design
Accurate detection of pathogens in seed lots depends on well-designed sampling strategies. Because seed lots are large and contamination is often rare and uneven, traditional sampling rules are not always sufficient. This section introduces modern approaches such as risk-based and Bayesian designs that improve detection while managing cost and uncertainty. It also covers pooling methods, models for heterogeneity, and operational factors like sample handling and transport. Together, these elements support reliable, efficient, and scientifically grounded sampling plans.
3.1. Lot Size and Statistical Sampling Plans
Seed lots often contain extremely large numbers of individual units, and pathogen presence is typically rare and nonuniform. The classical “square-root rule” (e.g. sample size ∝ √N, where N = lot size) can sometimes serve as a rough heuristic [
34] but does not account for heterogeneity or prior information. In modern frameworks, two important enhancements are:
Risk-based sampling designs: these allocate more sampling effort to lots with higher prior risk (due to origin, field history, or previous surveillance results). The idea is that not all lots are equal in their prior probability of contamination.
Bayesian sampling designs: these explicitly incorporate prior probabilities of infection (or contamination) and allow calculation of posterior probability of presence/absence after sampling [
35].
In a Bayesian framework, one might specify a prior distribution on the proportion of infected seeds in the lot (e.g. Beta distribution) and then update that with observed negatives/positives to compute a posterior credible interval for the infection prevalence or a posterior probability of “freedom from infection.” This is often more informative than frequentist fixed-size designs when pathogen prevalence is very low [
36].
When designing a sampling plan, one can calculate, for any proposed sample size n, the probability of detecting at least one infected seed given a true underlying prevalence p, using:
P(detect at least one infected seed)=1−(1−p)n
Where:
Assumptions:
For example:
This means a 95% chance of detecting at least one infected seed.
One also often defines a required confidence level (e.g. 95% or 99%) that the lot is “free” of infection at or below a threshold prevalence (the “damage threshold”) if no positives are found. [
34] outlines this in the context of seed health testing. In regulatory settings, such probabilistic designs support transparent decision thresholds (i.e. lot acceptance or rejection) and can be aligned with phytosanitary risk tolerance.
3.2. Composite/Pooling Strategies
Pooling (or compositing) is a practical strategy to reduce the number of assays needed, by mixing subsamples from multiple units (or seeds) and testing them together. If a pooled test is negative, one infers none of the constituent units are positive; if positive, further deconvolution or individual testing may follow.
Key factors in pooling design:
Pool size dilution effect: if the pathogen load per infected seed is low, pooling too many seeds may dilute the target nucleic acid below the assay’s limit of detection (LOD).
Homogenization and mixing uniformity: to reduce sampling bias, the pool must be well mixed so that each aliquot is representative.
Adaptive or algorithmic pooling: modern approaches use Bayesian or information-theoretic optimization (e.g. maximizing mutual information) to define which pools to test, how large, and how to split in follow-ups [
37].
D-optimal pooling design: for a known prior infection probability and test error rates, one can optimize pool groupings to maximize the information gained per assay [
38] though originally applied to SARS-CoV-2, the general approach is transferable to seed pathogen screening.
Regulatory frameworks such as ISTA (International Seed Testing Association) and EPPO (European and Mediterranean Plant Protection Organization) set specific limits on pooling size for official testing. These limits are designed to ensure assay sensitivity and minimize false negatives due to dilution. Compliance with such standards is essential for results to be accepted in international trade and phytosanitary certification.
Thus, pooling strategies must balance cost efficiencies against increased false negative risk due to dilution or sample heterogeneity.
3.3. Heterogeneity Models and Implications for Detection Limits
Real seed lots frequently exhibit aggregation or clustering of infected seeds rather than a uniform random distribution. A simple binomial model (constant p) often underestimates variability. Two alternative models are:
Negative binomial distribution: to model over dispersed counts of infected seeds (i.e. variance > mean). Under this model one can derive the probability that a sample of size n will include at least one infected seed, accounting for clustering.
Hierarchical occupancy-detectability models: for example, when each sub-unit (e.g. packet, bag, sublot) has its own probability of being infected, and detection within is probabilistic [
39].
Beta-binomial model: this approach assumes that the infection probability itself varies across subsamples, following a beta distribution. It captures extra-binomial variation and allows for more realistic estimation of detection probabilities when infection rates are not constant.
These models help to better estimate the false negative rate at given sample sizes, particularly under patchy infection. For example, even with an assay LOD of one infected seed per 1000, if infected seeds cluster in only certain sublots, one might miss them if sampling is not spatially stratified.
In practice, heterogeneity modeling may draw from spatial sampling data, prior outbreak maps, or controlled spiking experiments. Simulation (Monte Carlo) is often used to evaluate different sampling designs under assumed clustering parameters.
3.4. Chain of Custody, Transport, Storage, and Pre-Analytical Variables
Ensuring that sampling integrity is preserved from sampling to analysis is as critical as the statistical design. Key considerations include:
Chain of custody/traceability: each subsample must be uniquely barcoded or labeled with parent lot identifier, sampler ID, timestamp, and handling record (e.g. in a laboratory information management system). This is essential for auditability and data integrity under international standards (ISO/IEC 17025).
Transport and storage conditions: fluctuations in temperature, humidity, or physical agitation can degrade pathogen viability or nucleic acid integrity. For molecular assays, samples should often be stored at ≤ 4 °C or frozen and preserved with desiccants to limit nucleic acid degradation.
Pre-treatment and residual inhibitors: seed surface treatments (e.g. fungicides, coatings, seed treatments) or decontamination agents may inhibit downstream assays (e.g. PCR inhibition). Standardized wash steps or inhibitor removal protocols are necessary.
Homogenization before subsampling: to counter segregation (e.g. by size, density, or seed damage) during handling, bulk mixing before splitting into subsamples reduces bias.
Temporal stability and decay: for pathogen viability or nucleic acid detection, decay during storage is a known factor; stability studies may be needed to quantify how storage intervals affect detectable load (i.e. degradation rate).
Digitalization plays a growing role in managing these variables. Automated sample tracking, sensor-based condition monitoring, and AI-assisted anomaly detection can help ensure sample integrity and flag deviations in real time. Linking metadata to assay results enables retrospective analysis and supports adaptive sampling strategies. These technologies not only improve operational efficiency but also enhance the reliability of statistical inferences drawn from the sampling process. Poor control of pre-analytical steps may inflate false negatives or variability and confound statistical assumptions from the sampling model
3.5. Integration with Diagnostic Workflows
To close the loop, sampling metadata (lot ID, subsample IDs, spatial/prior risk variables) should be electronically linked to assay results (positive/negative, Ct values, confirmatory tests) via a LIMS. This allows:
Retrospective evaluation of sampling efficacy,
Recalibration of risk models,
Estimation of residual risk for lots with negative results.
Furthermore, iterative or adaptive sampling can be implemented: early negative results may reduce further sampling intensity, but borderline results might trigger additional subsamples. Bayesian updates allow recalculation of posterior risk and dynamic decision thresholds.
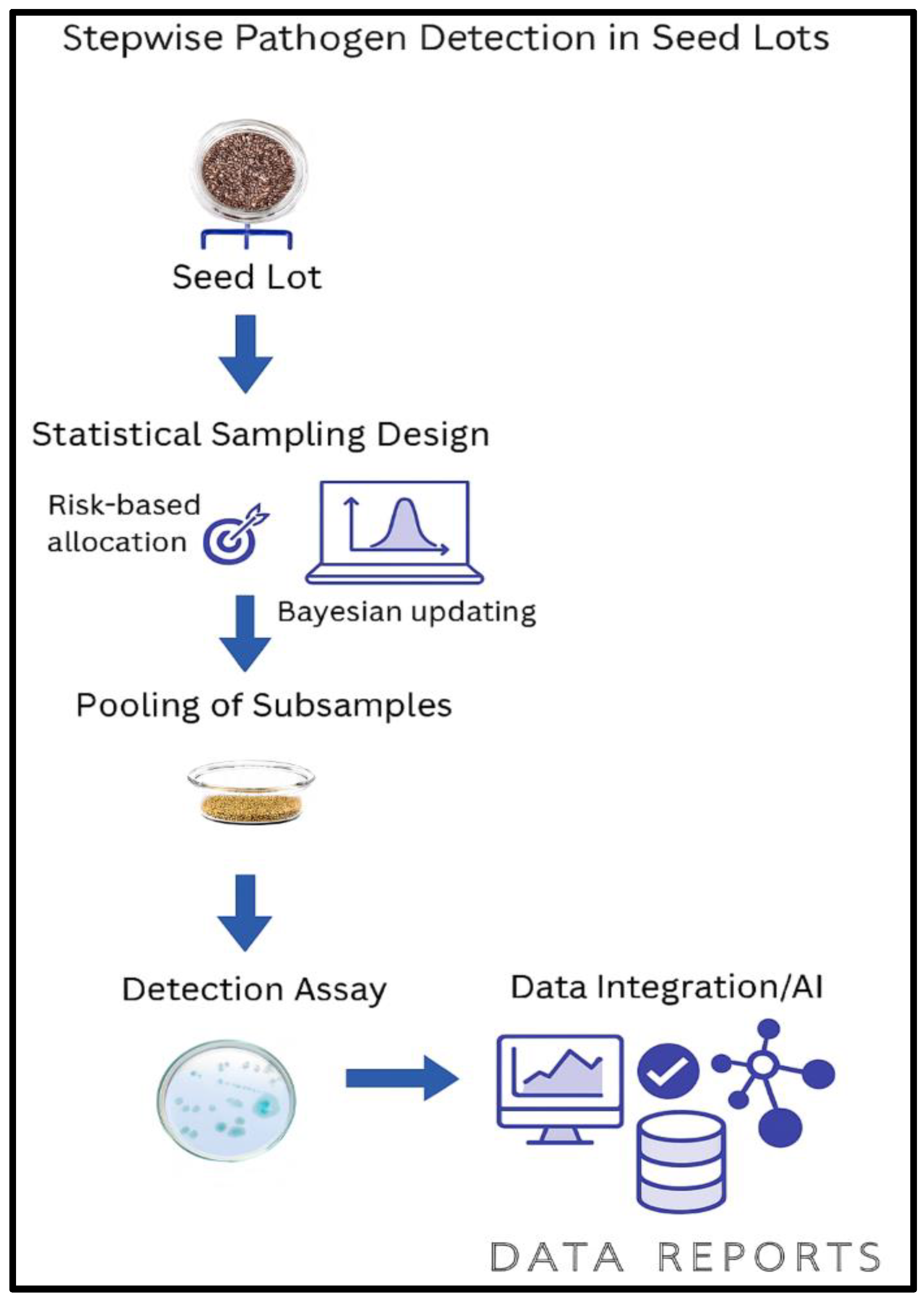
This integration also lays the groundwork for machine learning and AI-based risk prediction. By aggregating historical sampling data, assay outcomes, and contextual metadata, predictive models can be trained to identify patterns associated with contamination risk. These models can then inform future sampling strategies, prioritize high-risk lots, and optimize resource allocation. As shown in
Figure 3.1, the workflow integrates sampling metadata, pooling design, detection assays, and data systems to support adaptive diagnostics and machine learning–driven risk prediction.
4. Conventional Diagnostics (Baseline)
4.1. Overview
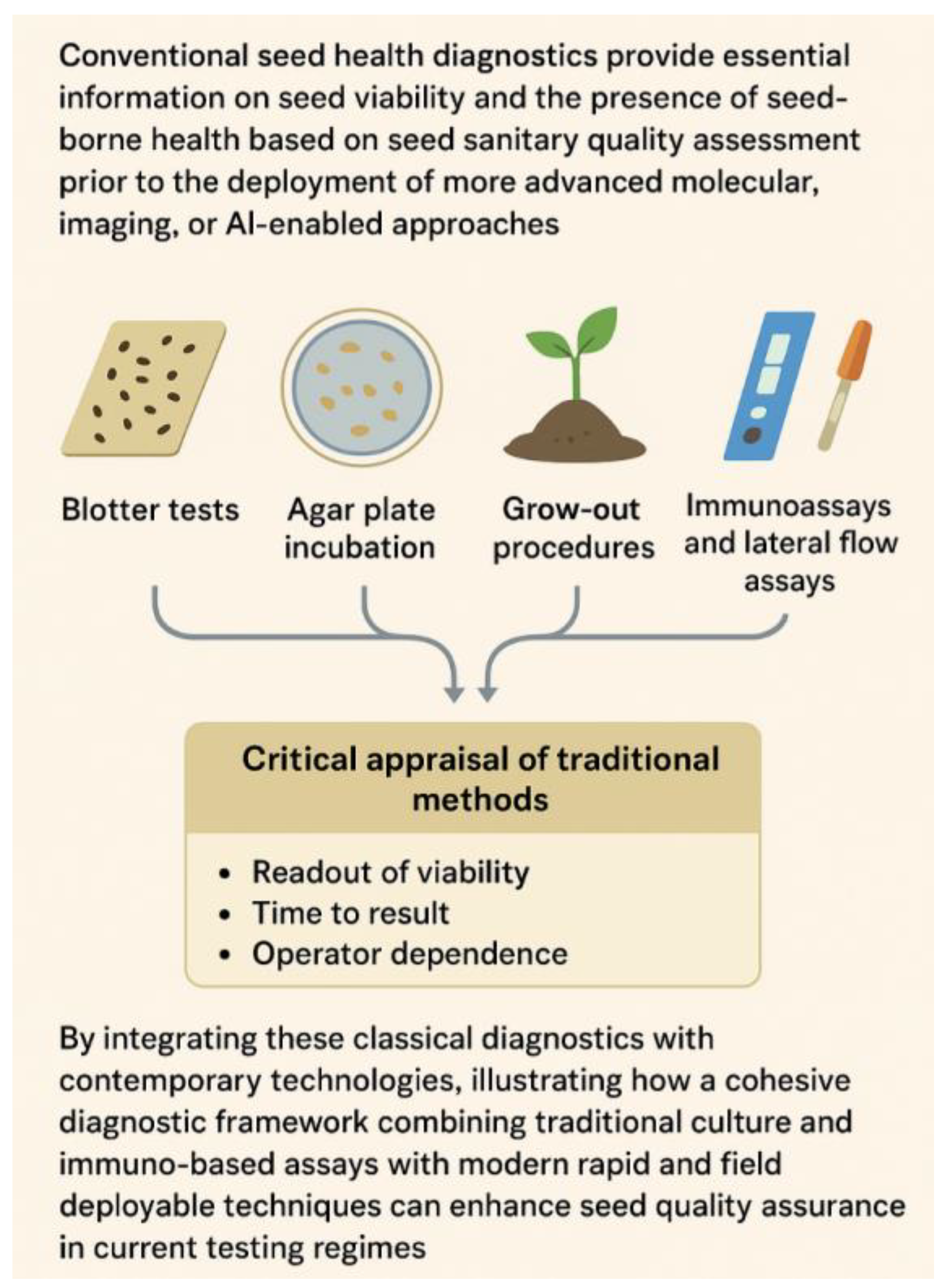
Conventional seed health diagnostics forms the foundation of seed sanitary quality assessment, providing essential information on seed viability and the presence of seedborne pathogens, prior to advanced molecular, imaging, or AI-enabled approaches. The established approaches such as blotter tests, agar plate incubations, grow-out procedures, and seedling assays, ELISA and lateral flow immuno-assays remain integral to seed health certification. A critical appraisal of these traditional methods is presented with particular emphasis on key performance dimensions readout of viability, time to result, and operator dependence that influence decision making in seed-health management and phytosanitary certification [
40,
41]. By integrating these classical diagnostics with contemporary technologies, how a cohesive diagnostic framework combining culture and immuno-based assays with rapid and field deployable techniques can enhance seed quality assurance [
42]. While conventional methods offer direct viability readouts and relatively cost-effective implementation, they may entail longer turnaround times and require trained personnel, considerations which motivate the ongoing development and integration of rapid adjuncts and automated readouts in contemporary seed-health programs [
43,
42]. The conventional diagnostics put traditional seed-health assays as indispensable anchors for quality assurance, while clarifying their limitations and the value of synergistic integration with modern diagnostic modalities [
44,
45].
4.2. Blotter, Agar Plate, Grow-Out, Incubation and Seedling Assays
Traditional seed health diagnostics comprise a family of complementary, culture-based methods used to assess seed viability and detect seed-borne pathogens. Blotter tests, agar plate (and grow-out) assays, incubation followed by seedling evaluations, and related seedling assays collectively provide rapid screening, culturable pathogen recovery, and functional readouts of seed health [
44]. When integrated with immunoassays and newer rapid modalities, these conventional assays remain foundational anchors for seed health certification, regulatory screening, and quality assurance programs. Across the literature, these methods are described as rapid, relatively cost-effective, and standardizable (e.g., ISTA guidelines), yet they depend on culturable organisms, can be observer-subjective, and may entail variable turnaround times for culture recovery and pathogen identification. These themes emerge consistently across contemporary reviews and primary studies and are explicitly linked to their roles in baseline diagnostics, sanitary status assessment, and informing sanitation interventions [
46].
The interrelationships among blotter, agar plate/grow-out, incubation, and seedling assays are best understood as a spectrum: blotter and seedling-based viability readouts provide rapid, functional signals of seed health; agar plate and grow-out methods enable pathogen recovery, identification, and pathogenicity testing; incubation/seedling assays translate infections into measurable growth impacts, yielding composite health risk metrics [
47]. This synthesis aligns with the framing of conventional diagnostics as “baselines” that can be complemented by molecular, imaging, and AI-based approaches to form cohesive seed health frameworks [
48,
49].
4.2.1. Blotter Tests and Seedling Germination Assays.
Blotter tests are a rapid, controlled-condition screening method to detect seed-borne pathogens and to evaluate germination performance and seed vigor. They yield direct viability readouts and can reveal pathogenic impairment via abnormal seedling development or germination failure, informing sanitary status and vigor assessments [
46,
50]. Standard vigor assessments often use early germination counts as proxies for seed lot vigor, acknowledging that seed deterioration slows germination and reduces vigor indices [
51]. Blotter test observations are qualitative but can be supported by laboratory confirmation for pathogen identity and viability [
52]. Blotter-based screening informs seed lot decisions and sanitary status, serving as a rapid first-pass filter before more resource-intensive assays or downstream management interventions [
52].
Agar plate and grow-out assays enable recovery and identification of culturable seed-borne fungi and bacteria, providing essential information on the spectrum of seed-transmitted pathogens and supporting phenotypic characterization. This phenotypic data supports decisions regarding seed treatment, sanitation, and intervention strategies; culture-based methods align with ISTA guidelines and have underpinned health surveillance across multiple crops [
53,
54]. These culture-based approaches support pathogenicity testing when needed for quarantine and certification decisions, reinforcing the role of traditional methods in regulatory contexts. However, they rely on culturable organisms; some pathogens may be non-culturable or slow growing, potentially underestimating disease burden [
47].
Incubation evaluations followed by seedling assays quantify disease transmission potential and seed health risk by observing the functional consequences of seed-borne infections on early growth. Seedling-based metrics capture both viability and pathogen impact, providing a robust baseline for sanitary quality where rapid screening alone cannot differentiate latent infections [
55,
56]. These assays reflect the real-world consequences of seed infections on establishment and yield, particularly for crops where seedling vigor is closely tied to field performance [
57]. Like blotter tests, seedling assessments can be subject to environmental variation and observer interpretation; standardization of conditions and scoring is essential to ensure comparability across laboratories and programs [
58,
59]. Blotter and seedling assays provide rapid, cost-effective insights into seed vigor and infection potential. When supported by confirmatory culture or molecular testing, they serve as reliable first-line indicators of seed lot health.
4.2.2. Synthesis of Baseline Methods
Collectively, blotter tests, agar plate/grow-out, incubation, and seedling assays form a diagnostic continuum for rapid viability screening to pathogen recovery assessment. They are widely used, standardized (including ISTA guidelines), and foundational for seed health certification, screening during production, and regulatory programs. The blended use of these methods provides a practical pathway from rapid viability signals (blotter/seedling germination) to detailed pathogen recovery and characterization (agar plate/grow-out), with incubation-seedling stages offering functional assessments of disease transmission risk [
60]. A central caveat is reliance on culturable organisms and potential subjectivity in observations; integrating these traditional assays with molecular, immunoassay-based, and imaging approaches can enhance decision-making, turning traditional anchors into part of a cohesive, multi-modal diagnostic framework [
61]. This integrated perspective is consistent with recommendations to couple conventional diagnostics with rapid adjuncts and automated readouts to improve seed-health programs [
61,
62].
4.3. ELISA and Lateral Flow for Priority Pathogens
4.3.1. ELISA and Seed-Health Applications
ELISA (enzyme-linked immunosorbent assay) remains a core high-throughput method for screening seed lots for priority seed-borne pathogens, especially viruses and, to a lesser extent, bacteria and fungi [
46]. It detects pathogen antigens, such as viral coat proteins or bacterial/fungal determinants through antibody–antigen binding on microplates with enzyme-linked colorimetric, fluorescent, or chemiluminescent detection. DAS-ELISA, indirect ELISA, and competitive ELISA offer different balances of sensitivity, antibody requirements, and ease of standardization [
63]. Reliable performance depends heavily on optimized extraction because seed matrices rich in oils, polyphenols, and storage proteins can inhibit antigen capture; ISTA protocols emphasize bulked seed extracts, homogenization, clarification, and spiking approaches for validation. ELISA remains widely used for seed-borne viruses such as potyviruses and tobamoviruses, and commercial kits support routine screening in certification, quarantine, and surveillance programs [
64]. Although adaptations exist for bacterial pathogens (e.g., Xanthomonas spp.) and some fungal antigens, their deployment is more limited due to challenges in antibody development and antigen extraction [
65,
66].
4.3.2. LFIA Principles, Advantages, and Performance Considerations
Lateral-flow immunoassays (LFIA) apply the same antibody–antigen recognition to membrane strips, generating visible test lines as extracts migrate by capillarity. Sandwich and competitive designs enable rapid, field-ready detection within minutes, and recent advances integrate nanoparticles, engineered reporters, and portable readers for improved sensitivity and quantitation [
67]. LFIA is increasingly used for on-site screening of seed-borne viruses and select bacterial/fungal targets, though careful validation remains essential [
66]. Compared with nucleic-acid assays, both ELISA and LFIA offer speed and low cost but generally lower analytical sensitivity than PCR/qPCR/RT-qPCR; molecular assays detect lower titers while immunoassays may tolerate some inhibitors yet risk cross-reactivity [
68]. Because immunoassays detect antigen irrespective of viability, positive results require confirmation through culture, grow-out, or PCR for regulatory decisions [
69]. Best practice, therefore, positions ELISA/LFIA as front-end screens within integrated workflows endorsed by ISTA and proficiency-testing programs [
70], with strong user acceptance reported for LFIA in on-farm and quarantine contexts [
71]. The main conventional diagnostic methods—blotter tests, agar plate incubation, grow-out procedures, and immunoassays—are summarized in
Figure 4.1
.
4.4. Strengths/Limitations: Viability Readout, Time-To-Result, Operator Dependence
A key limitation shared by most conventional diagnostic assays—including blotter, agar plate, incubation, grow-out, and seedling tests, as well as immunoassays such as ELISA and lateral-flow immunoassays (LFIAs) is their variable capacity to infer pathogen viability. Culture-based methods (e.g., agar plate and blotter assays) provide a direct indication of viability, since detection depends on the successful germination, growth, and sporulation of the pathogen from the seed; for this reason, they remain the gold standard for viability confirmation and are routinely required for regulatory seed health certification [
72]. In contrast, immunoassays detect antigenic proteins or epitopes that may persist even in non-viable or inactivated cells, meaning a positive ELISA or LFIA result verifies the presence of pathogen-derived antigenic material but not infectivity or transmission potential [
73]. This limitation poses challenges for quarantine and phytosanitary decisions, where confirmation of “live” pathogens is critical. The need to discriminate between viable and non-viable propagules has therefore stimulated the development of viability-PCR (vPCR) and other advanced nucleic-acid–based techniques that incorporate intercalating dyes (e.g., PMA or EMA) to selectively suppress amplification of DNA from dead cells, thereby improving the biological relevance of molecular detection, although these remain outside the scope of most routine or conventional seed diagnostic workflows [
74].
Immunoassays such as ELISA and lateral flow immunoassays (LFIA) provide rapid turnaround (ELISA in a few hours, LFIA in minutes) combined with relatively low cost per sample when assays are batched, making them attractive for moderately high throughput screening (e.g. as in seed health testing protocols). LFIA requires minimal laboratory equipment and modest operator training, which enables field deployment or use in resource-limited settings. Where well-characterized antibodies and standardized extraction protocols are used, the assays often yield good reproducibility; and ELISA formats lend themselves to automation (e.g. via plate washers and microplate readers) to further scale throughput. Because immunoassays detect antigenic molecules and not viability, positive detections do not guarantee that the pathogen is alive or infectious, thus confirmation via culture, bioassay, or grow-out may be required to satisfy regulatory phytosanitary or seed health endpoints (as noted in seed health best practices) [
75]. The sensitivity of ELISA and LFIA is typically lower than that of molecular (PCR/qPCR) assays, and low-titer infections may escape detection (ELISA and LFIA are often less sensitive than nucleic acid methods) [
46]. In addition, antibody cross-reactivity and interference from complex seed matrices can lead to false positives or negatives unless validation and rigorous controls are in place (a known limitation in lateral flow immunoassays) [
76]. Particularly for LFIA, the visual readout may be subjective; using quantitative strip readers can mitigate subjectivity but introduces additional cost and logistical burden (e.g. device calibration, power needs, transport) [
77].
International and national seed testing organizations (e.g., ISTA, national seed health committees) provide guidelines and proficiency testing schemes that include ELISA for virus testing and recommend validation steps (limit of detection, specificity, pooled sample strategies). Proficiency testing documents describe how to prepare spiked seed material for ELISA/LFIA validation and performance monitoring. These frameworks are important for harmonizing results across labs and for demonstrating the suitability of immunoassays in certification programs [
78]. On the sensitivity enhancements, nanoparticle labels, signal amplification, and reader-based LFIA are improving LFIA sensitivity toward ELISA/qPCR levels in some applications [
79], and the multiplexing attempts to multiplex ELISA panels and LFIA strips (multi-line membranes, microarray ELISAs) aim to screen for multiple priority pathogens in a single run attractive for seed companies and quarantine services [
80]. The workflow integration consists of hybrid pipelines that combine LFIA/ELISA screening with reflex molecular testing (qPCR) and culture/grow-out confirmation are becoming the operational norm in many advanced seed laboratories, while digital capture & AI: smartphone readers for LFIA and image-analysis of ELISA plates are improving objectivity, traceability, and remote data aggregation [
81].
In practice, immunoassays (ELISA and LFIA) are best deployed as screening tools, especially for large seed lots or at point-of-entry inspections, with any positive results then subjected to confirmatory viability or culture-based tests to satisfy regulatory phytosanitary or seed health requirements (e.g. as recommended in seed health chapters). It is critical to validate extraction protocols and pooling strategies for each seed species and target pathogen, to minimize matrix effects or dilution losses that could impair assay sensitivity or specificity (as highlighted in ISTA/ISHI best practice documents). Participating in proficiency testing or inter-laboratory comparisons helps ensure consistency and comparability across laboratories and supports regulatory confidence in the assay results (a key element of ISTA’s accreditation and quality assurance programs). Finally, when feasible, using reader-assisted LFIA devices or plate readers for ELISA reduces subjectivity in interpretation and enables quantitative or semi-quantitative data capture, which is valuable for trend analysis, quality control, and performance monitoring (as discussed in immunoassay review literature) [
82].
The comparison in
Table 4.1 highlights that conventional assays remain essential for seed health testing, particularly when viability or cultural confirmation is required. However, their limitations in speed, standardization, and operator dependence have driven the adoption of molecular, imaging, and AI-based diagnostics discussed in later sections. ELISA and lateral-flow immunoassays serve as rapid, affordable screening tools for priority pathogens—mainly viruses—but their inability to confirm viability and lower sensitivity compared to molecular methods means they should not be used as sole evidence for regulatory decisions. Current literature supports integrated workflows where immunoassays provide initial screening, followed by confirmatory molecular or culture-based tests. Technical improvements such as nanoparticle labels, multiplexing, and reader devices are narrowing sensitivity gaps and improving objectivity [
83,
84].
5. Molecular Diagnostics- Core Methods
Molecular diagnostics provide essential tools for seed health testing by enabling precise detection, quantification, and differentiation of plant pathogens. Techniques such as polymerase chain reaction (PCR), real-time quantitative PCR (qPCR), and digital PCR (dPCR) offer high specificity and sensitivity, especially when extraction methods are optimized for challenging matrices. In regulated seed testing workflows, end-point PCR remains valuable for rapid exclusion and identity verification, while advanced formats like nested PCR address
5.1. End-Point PCR and Nested PCR
Classical Polymerase chain reaction (PCR) is pivotal due to its robust performance in routine diagnostics, handling scenarios that demand single target checks or when only limited information is necessary [
85]. Classical end-point PCR read on agarose gels or capillaries, remains useful for single-target identity checks and rapid exclusion screening in routine workflows [
86]. Established uses include detection of
Fusarium oxysporum f. sp.
lactucae in lettuce seeds [
87] and
Clavibacter michiganensis subsp
. michiganensis (Cmm) in tomato seed-lots [
88], which helped set early molecular benchmarks for regulated pathogens.
Nested PCR increases detection sensitivity by introducing an internal primer set, thereby enhancing the probability of identifying trace pathogens subjected to strong sanitization or environmental challenges. To mitigate contamination, physical separation of setup areas, filtered pipette tips, and enzymatic approaches to control amplification carryover are recommended. Nested PCR has been applied to detect
Xanthomonas axonopodis pv.
allii from onion seeds [
89],
Alternaria carthami in safflower seeds,
Candidatus Phytoplasma asteris’ associated with Phyllody and Witches’ Broom in Pea (Singh et al., 2024) and 16 SrI and 16 SrVI phytoplasma from carrot seed [
90]. Seed extracts often contain inhibitors such as polysaccharides, polyphenols, and treatment residues; mitigation includes matrix-adapted extraction, dilution-to-extinction, BSA/PVP additives, or bead/silica clean-ups, plus an internal amplification control (IAC) to detect residual inhibition [
91].
5.2. Quantitative and Digital PCR
Quantitative real-time PCR (qPCR) extends PCR from presence/absence to calibrated measurement and high-throughput surveillance. Minimum Information for Publication of Quantitative Real-Time PCR Experiments (MIQE) compliant protocols, emphasizing calibration range, efficiency, and reproducibility, have led to significant improvements in transparency and inter-laboratory reliability [
92]. In seed testing, practical limit of detection (LOD)/limit of quantification (LOQ) depend more on extraction recovery and inhibitors than on instrument optics; include process controls (e. g., exogenous spikes or plant targets) and define guard-banded decision thresholds for lot release. qPCR assay is developed qPCR for detection and quantification of Cmm in tomato seeds [
93],
Xanthomonas translucens pv.
undulosa in wheat seeds [
94],
Pseudomonas syringae pv.
tomato (Pst) [
95] and
Fusarium oxysporum f. sp.
phaseoli in Common bean seeds. The multiplex qPCR for the detection of Cmm, Pst and pathogenic
Xanthomonas species in tomato [
96] and simultaneous detection of
Colletotrichum truncatum,
Corynespora cassiicola, and
Sclerotinia sclerotiorum in soybean seeds [
97].
When measurements are robust at very low copy number or in inhibitor-rich matrices, digital PCR (dPCR) partitions reactions into droplets/nanowells for poisson-based absolute counting, avoiding standard curves and improving precision near action thresholds. In seeds, dPCR has improved quantitation for Cmm in tomato seeds [
98] and
Stagonosporopsis cucurbitacearum in seeds of
Cucurbita maxima [
99]. Notably, dPCR allows for absolute quantitation at very low copy numbers, supporting reliable decision-making even near threshold levels. Following the dMIQE framework is crucial for achieving reproducible and comparable results across laboratories as well as for publication. [
100].
5.3. Isothermal Amplification: LAMP, RPA
Isothermal amplification techniques like loop-mediated isothermal amplification (LAMP) and recombinase polymerase amplification (RPA) provide fast, equipment-minimal alternatives suitable for large-scale screening and field applications. These approaches use multiple primers and strand-displacing polymerases, allowing for rapid results with minimal preparation.
Isothermal amplification offers fast results and minimal equipment, which suits border points and large screening programs. Loop-mediated isothermal amplification (LAMP, ~60 – 65 °C) uses 4-6 primers with a strand-displacing polymerase and yields results in ≤30 min with turbidity, fluorescence, or color readouts that tolerate partially purified extracts [
101]. LAMP assays are developed for detection of various seed borne pathogens such as
Pseudomonas syringae pv.
actinidiae in kiwifruit seed [
102],
Fusarium fujikuroi and
Magnaporthe oryzae in rice seed [
103], and
Cercospora sojina in soyabean seeds [
104].
Recombinase polymerase amplification (RPA) technology employs a coordinated enzymatic system consisting of recombinase, single-stranded DNA binding proteins, and strand-displacing polymerase to facilitate specific recognition and amplification of the target sequence [
105]. This process occurs under isothermal conditions, typically within a temperature range of 25 °C to 43 °C. The detection of amplified products is achieved through lateral flow dipstick (LFD) analysis using appropriately designed sequence-specific probes. RPA is also integrating with CRISPR/Cas12 which enhance sensitivity, specificity, and rapidness of test from crude sap [
106]. RPA assays are established for detection of toxigenic
Fusarium verticillioides in maize seeds [
107],
Pantoea stewartii subsp.
stewartii in maize seeds and seedlings [
108] and
Xanthomonas oryzae pv.
oryzae in rice seeds [
109]. Both methods generate high amplicon loads; use closed-tube formats, UDG/dUTP carryover control, pre-aliquoted mastermixes, and strict separation of setup and analysis zones. Positive isothermal screens are commonly confirmed by qPCR/dPCR for quantitation or by amplicon sequencing for identity, combining speed with regulatory-grade specificity.
5.4. Reverse-Transcription Assays for RNA Viruses/Viroids
RT-PCR/RT-qPCR are the methods of choice for seedborne RNA viruses and viroids. One-step RT-qPCR combines reverse transcription and amplification in a single closed tube, reducing handling and lowering contamination risk while allowing multiplex internal controls [
110]. RT-qPCR protocols are validated for detection and quantification of Tomato brown rugose fruit virus (ToBRFV) in tomato/pepper seeds [
111], Pepino mosaic virus in tomato seeds [
112], Potato spindle tuber viroid (PSTVd) and Tomato chlorotic dwarf viroid (TCDVd) in tomato seeds [
113] and Cucumber green mottle mosaic virus in zucchini seeds [
114]. Because RNA is inherently unstable, RNA handling must involve RNase-free procedures, stabilization agents (e.g., guanidinium thiocyanate), and robust extraction/RT controls such as armored RNA or transcript-based references. Replicate definitions, Cq acceptance thresholds, and reflex criteria for ambiguous amplification signals should be clearly specified.
5.5. Viability-Linked PCR
Assessing seed viability after pathogen sanitation is challenging with DNA-based diagnostics alone, as nucleic acids from non-viable cells can persist and inaccurately reflect actual risks. Viability-linked PCR employs dye intercalators like propidium monoazide (PMA) or ethidium monoazide (EMA) to distinguish living cells by preventing amplification of compromised genetic material. PMA-based approaches are favored for their specificity and limited impact on viable cells, while complementary RNA-focused techniques offer further validation [
115]. PMA-qPCR has been developed to differentiate viable
Acidovorax citrulli in watermelon seeds [
116], viable
Xanthomonas euvesicatoria, X.
gardneri, X.
perforans, and X.
vesicatoria in tomato seeds [
117] and viable Cmm in tomato seeds [
118]. Optimization of dye concentration, light exposure parameters, and quenching conditions for the specific matrix is critical for the reliability of viability PCR assays. RNA centered approaches, including RNase pretreatment or the interrogation of labile mRNA species, serve as complementary lines of evidence and may be effectively integrated with RT-qPCR for the detection of bacterial and fungal viability. In accordance with best practices, the simultaneous reporting of total DNA and viability PCR-adjusted values, alongside conventional culture-based or grow-out methodologies when applicable, provides a robust framework for defensible lot-level assessments.
Seed-lot heterogeneity at low prevalence requires risk-based sampling and validated pooling schemes, with seed counts per extract so that results can be expressed as copies per seed or per gram and translated into prevalence estimates. Layer controls: IACs to flag inhibition; extraction/process controls to track recovery; inclusivity panels covering target diversity; and exclusivity panels to challenge near-neighbors. Guard-band decision thresholds such as Cq cutoffs in qPCR, positivity rules in isothermal assays, and copy-number gates in dPCR, also repeatability/reproducibility studies under inhibitor challenge, and verify in inter-laboratory settings. Follow MIQE/dMIQE with explicit LoB/LoD/LoQ, acceptance criteria, and reflex logic for harmonization and regulatory acceptance [
92,
99].
6. Next-Gen & Meta-Omics
Recent advances in high-throughput sequencing and meta-omics are transforming seed health diagnostics by enabling broad-spectrum screening of microbial communities and detection of low-titer seedborne pathogens. This section reviews three major workflows: amplicon metabarcoding, shotgun metagenomics or targeted capture, and portable sequencing with adaptive sampling. It concludes with a discussion of the bioinformatics, reference databases, and false-discovery management strategies that are essential for reliable implementation.
6.1. Amplicon Metabarcoding for Survey Screens
Amplicon metabarcoding uses universal or semi-universal primers targeting conserved genetic loci such as bacterial 16S rRNA, fungal ITS, or animal COI to survey microbial communities in seed materials [
118]. The workflow typically involves bulk DNA extraction from seed lots, amplification of barcoding loci, deep sequencing, and bioinformatic clustering into operational taxonomic units (OTUs) or amplicon sequence variants (ASVs). Compared with single-target assays, metabarcoding allows parallel detection of diverse microbial taxa and provides a cost-efficient approach for broad surveillance [
119].
However, several sources of bias can affect accuracy, including primer mismatches, uneven amplification efficiencies, tag switching, and incomplete reference databases [
120]. To mitigate these limitations, best practices recommend inclusion of mock-community controls, dual indexing, rigorous demultiplexing thresholds, and transparent reporting of clustering and filtering parameters [
118].
6.2. Shotgun Metagenomics and Targeted Capture for Low-Titer Pathogens
When pathogen loads are low or when strain-level resolution is required, shotgun metagenomic sequencing provides an untargeted approach by sequencing total DNA from both host and microbes [
121]. In seed health testing, metagenomics has been applied to reconstruct microbial assemblages associated with seeds and seedlings [
122].
For very low-titer pathogens, targeted capture or enrichment techniques such as hybridization-based bait libraries or PCR-based enrichment can increase detection sensitivity and reduce sequencing waste [
123]. Despite their analytical power, these methods are still challenged by host DNA background, the need for deep sequencing to detect rare taxa, and potential false positives when taxonomic assignment is uncertain [
124]. Reliable workflows therefore include statistical thresholds, genome completeness criteria, and independent confirmation of candidate detections.
6.3. Portable Sequencing and Adaptive Sampling
Portable sequencing technologies, particularly those from Oxford Nanopore Technologies (ONT), now make it possible to perform sequencing on site in seed testing laboratories. Adaptive sampling allows selective enrichment of pathogen reads during sequencing, which improves sensitivity without additional sample enrichment. These platforms can provide same-day confirmation of priority pathogens in quarantine or trade inspection contexts. Although field deployable, portable sequencing still requires rigorous sample preparation, quality control, and access to either local or cloud-based bioinformatics pipelines.
6.4. Bioinformatics Pipelines, Reference Databases, and False-Discovery Management
Across metabarcoding, metagenomics, and portable sequencing workflows, result reliability depends on robust bioinformatics pipelines, curated reference databases, and effective false-discovery management. Standard pipelines include quality filtering, host-read removal, taxonomic assignment, abundance normalization, and statistical validation. The limited availability of curated, high-quality reference genomes for many seedborne pathogens remains a major challenge [
119].
To minimize false positives and negatives, laboratories should apply transparent parameter settings, include negative and mock controls, set minimum read-support thresholds, and confirm rare taxa using orthogonal methods such as qPCR or culture [
124]. As computational tools evolve, machine learning classifiers, federated databases, and standardized reporting formats are expected to further streamline multi-pathogen seed diagnostics and strengthen data interoperability.
7. CRISPR-Based Diagnostics
Agriculture diagnostics refers to scientific interpretation, detection and monitoring of diseases, pests, nutrition deficiencies and genetic traits in crops, seeds and soil that enable timely measures for input optimization and the development of healthy and high-yielding plants. Diagnostic tools vary across several categories like Serological techniques, including ELISA (Enzyme-Linked Immunosorbent Assay), Immunostrip Tests/Lateral Flow Assays and Western Blotting are confirming the presence of specific pathogens or toxins in plant tissues based on antigen–antibody interactions [
125]. Similarly, biochemical testing involves the study of metabolites, enzymes and biomarkers that reflect plant physiological responses to environmental changes, stress, or nutrient deficiencies. Enzyme assays, isozyme assay and metabolite profiling are common techniques implemented under biochemical testing [
126,
127] Non-destructive imaging based diagnostic technique is evolving rapidly to obtain visual images or spectral signatures using sensors that detect plant stress or damage based on color, temperature or reflectance pattern. X-ray imaging, Thermal Imaging, Hyperspectral imaging and others enable real-time, high-throughput monitoring and can be integrated with AI or ML for precision agriculture application [
128,
129]. Molecular techniques represent the most sensitive, specific and precise diagnostic techniques as they utilize specific DNA or RNA sequence to determine traits in plants and seeds. These include techniques including Polymerase Chain Reaction (PCR) for amplifying target DNA to detect plant pathogens, DNA Barcoding and Sequencing to Identify unknown species or genetic variations and advance techniques like CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats) based Diagnostics (SHERLOCK, DETECTR), Loop-Mediated Isothermal Amplification (LAMP) that enable early disease detection, genetic purity testing and pathogen surveillance [
130,
131]. In the present scenario, integrated diagnostic approaches are implemented and crop diagnostics are employed for seed and crop health certification, purity testing and pathogen-free propagation material is making it an indispensable part of sustainable crop production, biosecurity and global food safety systems [
132].
7.1. Evolution of CRISPR-Cas Systems for Agricultural Diagnostics
CRISPR Cas system originated in the early 2010 as an adoptive immune mechanism in bacteria and archaea was providing defense invading bacteriophages (virus) by remembering or recognizing the foreign genome and cleaving the genetic material upon reinfection. Same principle was used to understand the distinctness of Cas family in precise nucleic acid detection and trans-cleavage activity and discovery of diagnostics like DETECTR (Cas12 system used in detection of specific DNA targets) and SHERLOCK (Cas13 system used to detect both DNA and RNA) platforms were developed. These systems produce visual or fluorescent signals in successful recognition of target sequences without need for complex laboratory equipment [
133,
134,
135].
Recent breakthroughs in CRISPR Cas technology have made it simpler, precise and versatile in application and recent study demonstrates use of amplification free CRISPR–Cas12a assay with multiplexed detection designed for Candidatus Phytoplasma detection and this platform used an engineered LbCas12a-Ultra variant alongside optimized 7nt stem-loop reporters could detect phytoplasmas at an accuracy of 99.4% and was further adapted for instrument-free lateral flow assays suitable for field diagnostics [
136,
137]. Current trends in CRISPR diagnostic systems show integration of microfluidic chips, AI-assisted signal quantification and portable smartphone interfaces for real-time agricultural surveillance can lead to breakthroughs aligned with precision agriculture is allowing proactive management of disease, pest and genetic purity of crop species [
138]. CRISPR-Cas9 system functions using three major components like the Cas enzyme, guide RNA (gRNA) and a short DNA sequence known as the Protospacer Adjacent Motif (PAM). Each have fixed role with gRNA (fusion of two natural RNAs-crRNA (CRISPR RNA) and tracrRNA (trans-activating CRISPR RNA)) that is complementary to target DNA, binds to targeting sequence (crRNA) and tracrRNA (scaffolding sequence) directs Cas protein to target DNA to form ribonucleoprotein complex. PAM sequence is located adjacent to targeted DNA sequence and helps Cas enzyme to recognize and bind to the target DNA with high specificity. Cas protein (Cas9, Cas12, Cas13) acts as a molecular scissor creating double stranded breaks at targeted DNA sequences allowing genome editing [
139,
140]. The steps involved in CRISPR/Cas9-based genome editing are summarized in
Figure 7.1 [
141].
Figure 7.
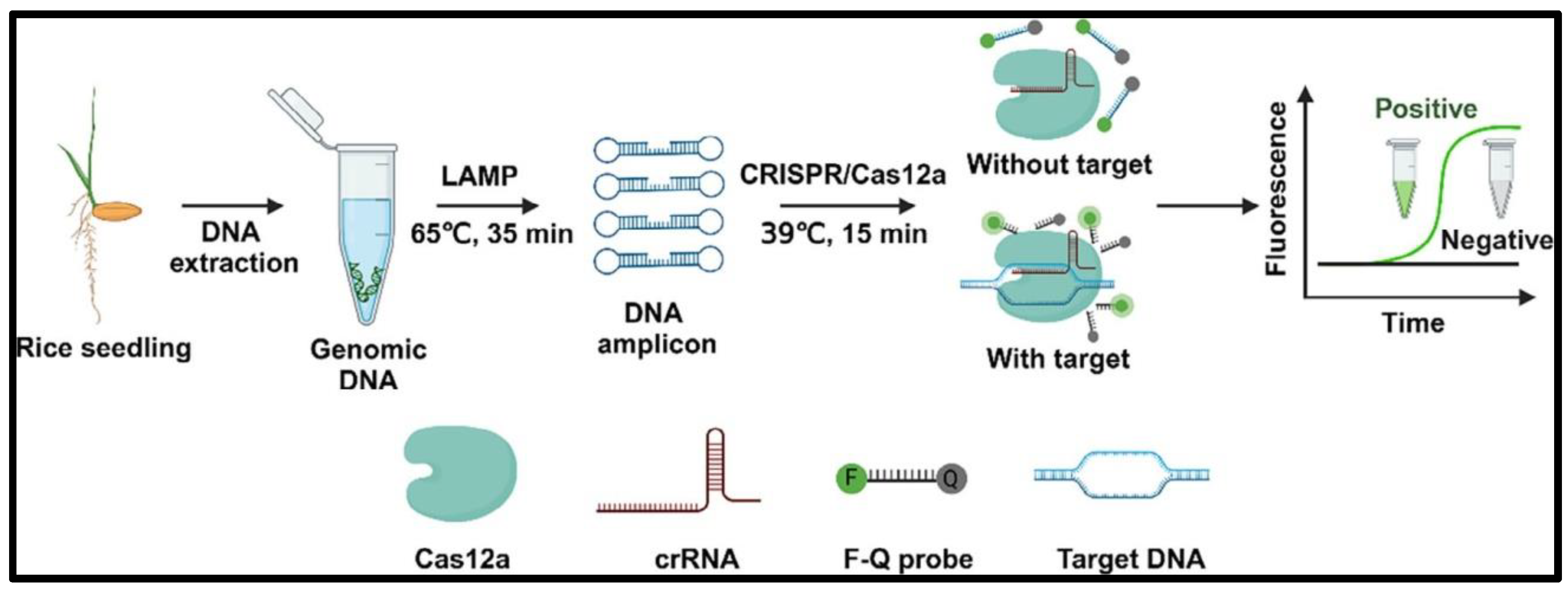
2. Flow chart of CRISPR/LbCas12a-LAMP biosensor used for on-site diagnosis of Rice Bakanae Disease [
153].
Figure 7.
2. Flow chart of CRISPR/LbCas12a-LAMP biosensor used for on-site diagnosis of Rice Bakanae Disease [
153].
7.2. Application of CRISPR Gene Editing on Seed Health Diagnostics
The advent of the CRISPR-Cas system has transformed molecular biology by facilitating accurate and sequence-specific DNA targeting. It employs guide RNA (gRNA) molecules to steer the Cas protein to specific DNA regions for targeted cleavage. This elevated specificity reduces nonspecific amplification and improves the precision of molecular detection and diagnostic procedures. The CRISPR-Cas system functions as an adaptable, programmable immunity mechanism that safeguards organisms against invading nucleic acids. The CRISPR-Cas system has emerged as a potent technique in nucleic acid detection and diagnostics due to its precision and adaptability, potentially transforming the identification and management of plant diseases. Among the several CRISPR-associated enzymes, Cas12a (a type V, class 2 endonuclease) is distinguished by its distinctive RNA-guided DNA cleavage capability. Upon binding of its guide RNA to the target double-stranded DNA, Cas12a is activated, initiating trans-cleavage activity that cleaves a fluorophore/quencher-labeled single-stranded DNA reporter, resulting in a visible fluorescence signal [
142,
143]. When integrated with isothermal amplification methods, CRISPR-Cas12a facilitates the detection of low-abundance nucleic acid targets by specifically recognizing the amplified DNA sequences [
144,
145,
146,
147]. Despite its effective application in other research domains, the promise of this technique for identifying Rice Bakanae Disease (RBD) remains under investigation and established an on-site diagnostic platform for RBD by combining the trans-cleavage capability of LbCas12a with LAMP, a swift DNA amplification technique facilitates the early identification and enhancing the management and regulation of the disease in the field. Nucleic acids are good biomarkers in molecular diagnostics because of their stability, dependable amplification and compatibility with several reporter systems [
148,
149]. While PCR is the benchmark for nucleic acid detection, it possesses significant drawbacks, such being time-consuming, necessitating advanced laboratory apparatus and relying on proficient workers [
148,
150]. Conversely, isothermal amplification techniques, which function at a constant temperature, provide a more rapid, straightforward and field-compatible alternative. These methodologies are exceptionally appropriate for point-of-care testing (POCT) and point-of-need testing (PONT). Recombinase Polymerase Amplification (RPA) and LAMP have demonstrated remarkable potential for field applications and the integration of isothermal amplification with CRISPR/Cas-based systems markedly improves detection specificity and facilitates visual readouts via lateral flow assays (LFA) or colorimetric detection. These biosensing technologies are particularly advantageous in resource-constrained environments, providing economical, highly sensitive and user-friendly diagnostic solutions appropriate for extensive screening [
148,
150,
151]. Numerous CRISPR-based biosensing systems have been established, including DETECTR (which integrates RT-LAMP and Cas12 for LFA-based detection) [
152], AIOD-CRISPR (a one-pot Cas12a-based fluorescent system) (Chao zhan et al., 2024), iSCAN (a two-pot RT-LAMP–CRISPR-Cas12a platform), iSCAN-V2 (which combines RT-RPA and CRISPR/Cas12b for SARS-CoV-2 detection) [
150,
154], Vigilant (utilizing a dCas9–VirD2 fusion with ssDNA reporters) and Bio-SCAN (a biotin-linked CRISPR-based detection system) [
155].
Recent LFAs have garnered heightened interest in point-of-care diagnostics owing to its rapidity, simplicity and minimal equipment prerequisites. They are especially proficient in swift diagnosis, food authenticity verification, and environmental surveillance in resource-constrained settings. Two principal domains of current LFA research encompass: (1) the integration of LFAs with isothermal amplification techniques (such as LAMP or RPA) to facilitate ultra-sensitive detection independent of laboratory facilities, and (2) the innovation of novel labelling materials that improve sensitivity and permit quantitative detection utilizing portable devices. Future developments in lateral flow nucleic acid testing are anticipated to emphasize microfluidic integration and the amalgamation of LFA with CRISPR-based technologies, facilitating swift, precise, and portable molecular diagnostics. This study illustrates the efficacy of a CRISPR/LbCas12a-LAMP biosensor for the on-site detection of Rice Bakanae Disease, facilitating practical and accessible crop disease control measures (
Figure 7.2).
Figure 7.
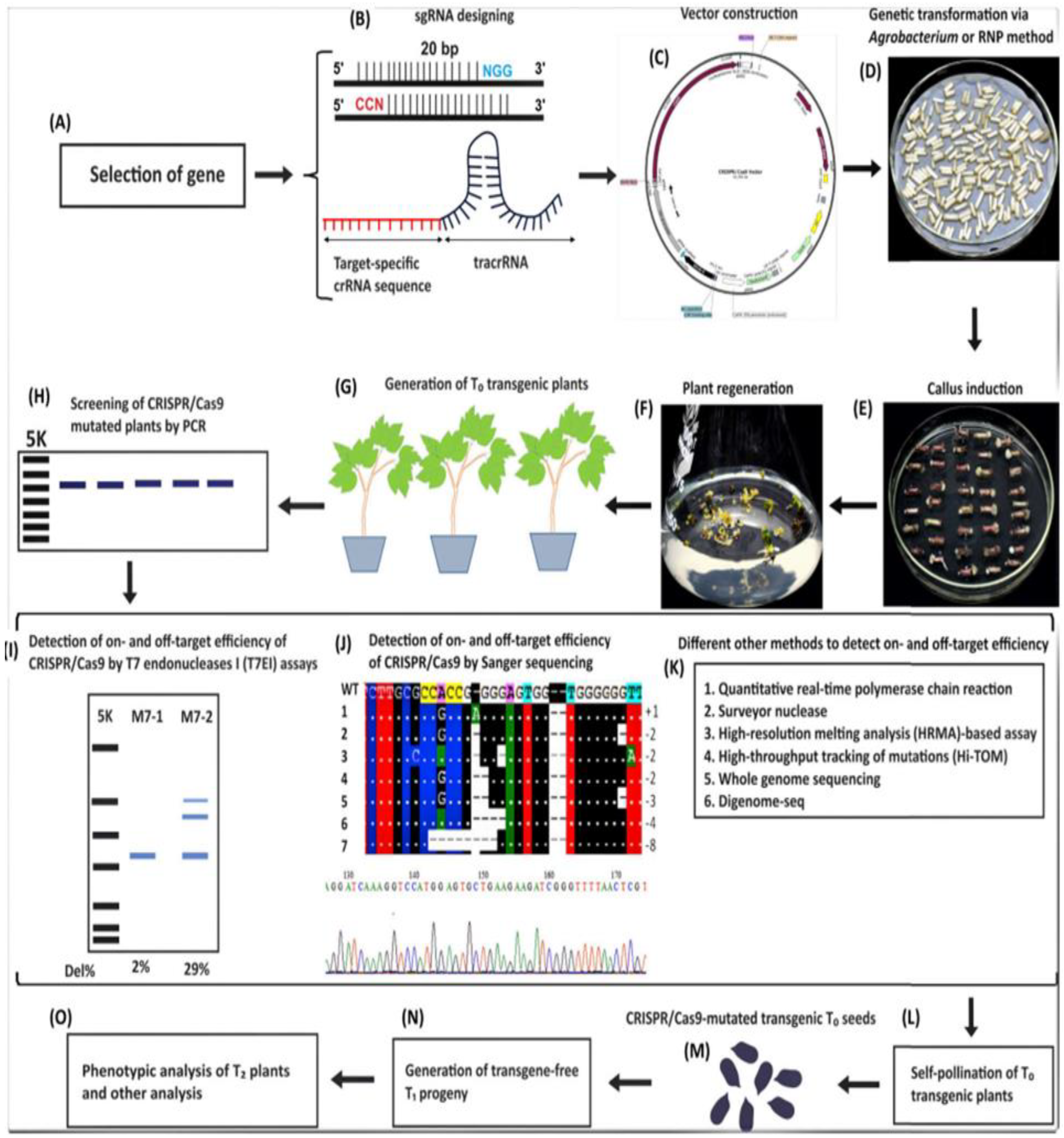
1. Flow chart of CRISPR/Cas9-based genome editing steps:(A) Target gene selection; (B) sgRNA design; (C) Vector construction; (D) Delivery via Agrobacterium or ribonucleoprotein (RNP); (E) Callus induction; (F) Plant regeneration; (G) Generation of T0 transgenic plants; (H) PCR screening; (I–K) On/off-target detection by T7E1 and sequencing; (L) Self-pollination for homozygous T1 plants; (M) CRISPR-mutated T0 seeds; (N) Transgene-free T1 progeny; (O) Phenotypic analysis.
Figure 7.
1. Flow chart of CRISPR/Cas9-based genome editing steps:(A) Target gene selection; (B) sgRNA design; (C) Vector construction; (D) Delivery via Agrobacterium or ribonucleoprotein (RNP); (E) Callus induction; (F) Plant regeneration; (G) Generation of T0 transgenic plants; (H) PCR screening; (I–K) On/off-target detection by T7E1 and sequencing; (L) Self-pollination for homozygous T1 plants; (M) CRISPR-mutated T0 seeds; (N) Transgene-free T1 progeny; (O) Phenotypic analysis.
A variety of CRISPR-based biosensing systems have been created to improve the accuracy and rapidity of nucleic acid detection. DETECTR combines RT-LAMP with Cas12 and employs a reporter molecule to generate a lateral flow assay (LFA) readout [
153]. AIOD-CRISPR is a singular detection method utilizing Cas12a facilitating real-time fluorescence visualization of target nucleic acids [
156]. Vigilant utilizes a chimeric fusion of nuclease-dead Cas9 (dCas9) and VirD2, along with a single-stranded DNA (ssDNA) reporter to create a sensitive detection complex [
157]. Bio-SCAN (biotin-coupled selective CRISPR-based assay for nucleic acid detection) is a promising platform that provides excellent specificity and adaptability for many diagnostic applications. The development of the diagnostic model was influenced by several critical factors like rapid prediction time, capacity to handle extensive image datasets and a compact model size compatible with the smartphone hardware typically accessible to farmers. This work presents an innovative and pragmatic method for detecting crop diseases utilizing common Android smartphones enhancing accessibility for rural people and agricultural consultants in the field. The above suggested technique possesses both theoretical significance in enhancing precision agriculture and practical advantages for disease categorization in field environments can be further developed to detect micronutrient deficiency signs in crops, a frequently neglected yet vital component of plant health assessment [
158].
8. Imaging & Non-Destructive Phenotyping
The application of non-invasive and imaging techniques with minimal human involvement is highly significant in the agricultural sector and crop development. Contemporary imaging technologies facilitate the automated visualization of multiple parameters for the characterization of biological specimens, thereby diminishing subjectivity and enhancing the analytical process. Additionally, the integration of two or more imaging modalities has played a crucial role in the identification of novel physicochemical tools and the real-time interpretation of datasets. Below are some key imaging and non-destructive methods as outlined [
159].
8.1. Radiation Imaging (X-Ray, CT and MRI)
Ionizing radiation (IR) is a form of high-energy radiation that possesses sufficient energy to displace an electron (a negatively charged particle) from an atom or molecule, resulting in ionization. This type of radiation serves as a significant asset in agricultural sciences and seed technology, frequently employed to tackle issues related to food microbiological safety and seed storage. Gamma (γ) radiation, a high-energy variant of IR, can penetrate and interact with living tissues [
160]. Traditional methods for insect detection, including grain flotation, the Berlese funnel technique, probes and traps, as well as measurements of carbon dioxide and uric acid, exhibit various limitations; they can be subjective, destructive, inaccurate, time-consuming, and ineffective in identifying internal insect infestations [
161]. Different insect detection methods vary in specificity: ELISA and PCR target unique insect proteins or genes, while IR detects general chemical groups. Acoustic, electrical conductance, and electronic nose methods rely on insects’ physical activity or emitted chemicals, and acid hydrolysis detects insect waste through chemical reactions. [
162].
This review paper presents the fundamental principles of imaging techniques and provides a summary of their use in detecting insects and fungi in stored products in real time. Among emerging non-destructive techniques, X-ray imaging, magnetic resonance imaging (MRI), thermal imaging and NIR hyperspectroscopy have been evaluated for rapid and accurate assessment of infestation. Pioneer studies on Nuclear magnetic resonance (NMR) have been shown to effectively detect concealed infestations of wheat caused by the granary weevil (
Sitophilus granarius), However, the sensitivity achieved was quite low and there have been no significant attempts to employ MRI for detecting stored product pests since that time. The other two advanced methods near-infrared (NIR) hyper-spectroscopy and X-ray imaging have shown promise for real-time applications [
163].
The impact of X-rays on living organisms remains incompletely understood. Within the spectrum of infrared sources, X-rays possess wavelengths ranging from 0.01 to 10 nm, which correspond to frequencies between 30 and 30,000 PHz (where 1 Petahertz equals 10^15 Hertz) and energies spanning from 120 eV to 120 keV. Soft X-rays, characterized by energies between 0.12 and 12 keV, are particularly effective for agricultural applications due to their limited penetration ability and capacity to reveal internal density variations. Very few studies have been published regarding the effects of X-ray exposure on seed performance since the 1960s [
164]. The existing understanding of X-ray effects on plants is still limited, primarily addressing a few physiological aspects and requires further exploration. New insights into the molecular and physiological mechanisms that confer resistance in plants are needed. The application of this imaging technique on hard red winter wheat samples infested with
S. oryzae pupae was examined by [
165]. The samples were placed in a plastic tube containing either 0, 50 or 100 infested kernels per kilogram of wheat. To enhance the contrast between the voids within the kernels and those between them, the inter-kernel spaces were filled with corn oil. The average detection accuracy for samples containing five infested kernels per 100 g was 94.4 ± 7.3%, while for ten infested kernels per 100 g, it was 87.3 ± 7.9%. The slightly lower detection accuracy for ten infested kernels can be attributed to overlapping kernels or the presence of air bubbles in the oil. Therefore, CT imaging could serve as a viable alternative for detecting pests in stored products. The advent of advanced technology has broadened the possibilities for non-destructive assessment of seed quality. Techniques such as X-ray imaging, Computed Tomography (CT), Magnetic Resonance Imaging (MRI) and ultrasound have been investigated for the non-destructive evaluation of indicators that are not visible on the surface of a variety of agricultural products [
166].
8.2. Near-Infrared (NIR)
Near-Infrared light penetrates the seed and emits a signal associated with its internal components, which are indicative of seed health. Consequently, NIR can distinguish between low vigor seeds, hidden insects and dead seeds based on the specific type of NIR signal emitted. Furthermore, seeds infected with fungal pathogens produce a distinct NIR signal, allowing for the identification of seeds with fungal contamination. NIR signals have also been utilized to differentiate hybrid watermelon seeds from inbred seeds, as well as to assess the duration required for seed priming to achieve optimal outcomes. While NIR technology is commercially available and used in the seed sector for seed quality assessment, its adoption is still limited due to cost, calibration needs and variability across crops [
167,
168].
NIR spectroscopy has developed into a rapid, dependable, precise and cost-effective method for the compositional analysis of seed health [
169]. This method is applicable for both qualitative and quantitative assessments. The NIR technique yields insights based on the reflectance characteristics of various substances found in a product. It operates on the principle of electromagnetic wavelength absorption within the range of 780-2500 nm. Classical absorption spectroscopy can be employed to ascertain the concentrations of components such as water, protein, fat and carbohydrates. NIR system as the most effective approach for identifying individual wheat kernels that harbor live or deceased internal rice weevils at different life stages. Machine vision systems are utilized to support grading, cleaning and the diagnosis of diseases and insects within food grain handling operations [
170]. At present, there is an increasing application of computer vision systems in seed health for quality assurance, encompassing a spectrum from standard inspections to advanced vision-guided robotic control [
171].
8.3. Thermal Imaging
Infrared thermal imaging is widely used in assessing seed quality, including evaluations of germination performance, viability, and vigor. It is also employed for estimating morphological characteristics, detecting diseases and insect infestations, and monitoring seed quality during storage. [
172]. Depending on the resolution, sensor sensitivity and the range of temporal data captured by thermal cameras, this technique is applied in various contexts. The infrared radiation that thermal cameras capture consists of long-wavelength radiation from the electromagnetic spectrum, which ranges from 0.78 µm to 1000 µm. This radiation is further categorized into near-infrared (0.75 - 3 µm), mid-infrared (3-6 µm), far infrared (6-15 µm) and extreme far infrared (15-100 µm) radiation. Typically, thermal cameras that operate within the far infrared radiation range are employed for seed quality evaluation and they record surface temperatures accordingly. The choice between active or passive thermal imaging systems is determined by the emissivity, absorptivity, transmissivity and reflectivity of the infrared radiation emitted by the sample [
173].
Thermal imaging may serve a pivotal function in this trend and could provide an alternative approach for identifying insect infestations, as the respiration of pests generates heat that exceeds that of the seeds [
174]. Lately, thermal imaging has recently emerged as a vital tool for curing diseases and detecting insect infestations in seeds, since the deterioration of seed tissues due to disease progression is typically linked to variations in the surface temperatures of the affected seed areas. An active thermal imaging system was developed [
175] could be used to detect and categorize diseases based on distinct differences in the temperature profiles of infected and healthy seeds. In both the heating and cooling phases, infected seeds exhibited higher surface temperatures than healthy seeds, highlighting the effectiveness of infrared thermal imaging for detecting fungal infections through changes in thermal behavior.
8.4. Multi Spectral and Hyper Spectral Imaging:
Multispectral and Hyperspectral imaging have been investigated as potential analytical tools for the non-destructive analysis and assessment of seed quality and safety. Hyperspectral imaging, which can acquire spectral and spatial information simultaneously, combines the advantages of spectroscopic and imaging techniques. In other words, it simultaneously obtains the chemical information and the spatial distribution of chemical components in heterogeneous samples [
176,
177,
178]. The identification of the wheat grain moth (
Sitotroga cerealella) has been evaluated through X-ray and multispectral imaging (MSI) as reported by [
179]. The study highlights the potential of multispectral imaging for detecting insect eggs located on seed surfaces. Recent research focusing on soybean, maize and sweet corn showed that seeds that are damaged are more likely to develop into abnormal seedlings. Additionally, studies conducted by [
180,
181,
182] demonstrated the effectiveness of MSI in categorizing various types of damage without the requirement for further analytical evaluations. A classification model based on surface features obtained from MSI and multivariate data processing achieved an overall accuracy of 82% in distinguishing between damage classes. As the processes involved in seed health testing are time-consuming and necessitate considerable training for the characterization of pathogenic fungi on seeds, a multispectral imaging system (395-970 nm) was utilized by [
183] to identify the surface properties associated with different fungal infections in spinach seeds. The system successfully distinguished healthy seeds from those infected with multiple fungal pathogens, including
Verticillium spp., Fusarium spp., Stemphylium botryosum, Cladosporium spp., and
Alternaria alternata. They implemented canonical discriminant analysis (CDA) to separate image pixels based on their mean intensity and employed Jefferies-Matusita (JM) distance for the classification and modelling of spectral data, at high accuracy between uninfected seeds and those infected seeds. Furthermore [
184], utilized a multispectral imaging system (375–970 nm) with 19 wavelength bands to differentiate 27 varieties of winter wheat (
Triticum aestivum L.) and nine varieties of triticale (
Triticosecale Wittm. and
Camus) in relation to their resistance to fungal infections. A comparative summary of non-destructive seed health methods, including hyperspectral imaging, X-ray, machine vision, and thermal imaging, is provided in
Table 8.1.
8.5. Chlorophyll Fluorescence Imaging (CF)
Chlorophyll fluorescence operates on the principle that chlorophyll molecules, whenstimulated by light of a specific wavelength (typically within the blue or red spectrum), emit light of a longer wavelength (fluorescence) as they revert to their ground state [
195]. This fluorescence signal can be quantified to evaluate various physiological maturity states of the seed, especially those associated with the functionality of the photosynthetic apparatus. The commercial application of this principle allows for the assessment of chlorophyll fluorescence in the seed coat or embryo, aiding in decision-making regarding harvest timing, maturity and seed viability. The quantity of chlorophyll was correlated with the germination performance and health status of the seeds [
196].
9. Artificial Intelligence (AI) and Machine Learning (ML) In Seed Diagnostics
Artificial intelligence and machine learning have emerged as transformative tools in seed health diagnostics, enabling rapid, non-destructive, and highly scalable analysis of complex datasets. These technologies leverage advanced algorithms to interpret visual, spectral, and molecular data, reducing reliance on manual assessments and improving diagnostic accuracy under low pathogen prevalence. By automating feature extraction and pattern recognition, AI systems can detect subtle indicators of infection, classify seed quality traits, and predict contamination risks with minimal operator bias. Among the various approaches, deep learning models particularly convolutional neural networks have shown exceptional performance in processing high-dimensional imaging and hyperspectral data, laying the foundation for next-generation diagnostic workflows.
9.1. Convolutional Neural Networks (CNN) for Image and Spectrum Classification
Convolutional Neural Networks (CNNs) have become the go-to deep learning workhorses for seed image analysis and pathogen detection across both image and spectral datasets. Convolutional Neural Networks (CNNs) excel at automatically extracting hierarchical features from seed images, enabling identification of morphological traits linked to seed health or pathogen presence without manual feature engineering. Recent work has reported CNN accuracies exceeding 99% in seed quality classification tasks [
197]. CNN architectures have also been adapted for seed health diagnostics. For example, hyperspectral imaging of rice seeds (874.41–1734.91 nm) combined with CNNs enabled detection of Fusarium spp., achieving accuracies above 98% [
198]. The study demonstrated that CNNs consistently outperformed traditional machine-learning models—including Partial Least Squares Discriminant Analysis (PLS-DA) and Support Vector Machines (SVM)—which typically achieved accuracies above 90%. This performance gap reflects CNNs’ ability to learn complex, nonlinear patterns in spectral data without handcrafted features.
Transfer-learning strategies using pre-trained CNNs have been particularly effective when seed-health datasets are limited. Advanced architectures such as EfficientNet variants have been tailored for seed disease detection. MobileNetV2, enhanced with layers such as average pooling, flattening, dense units, dropout, and softmax activation, reached ~96% accuracy across four corn seed disease classes (Broken, Discolored, Silk cut, and Pure) using 21,662 images. Performance gains were further supported by data augmentation, adaptive learning-rate schedules, model checkpointing, and dropout regularization. MobileNet with transfer learning has also achieved 99.55% accuracy in rice seed classification; freezing the first twelve convolutional layers and fine-tuning only the final layer allowed effective learning from a 10,000-image dataset spanning five seed classes [
199].
Hyperspectral imaging in the 400–1000 nm range has enabled identification of Aspergillus flavus in naturally infected peanut seeds. Integrating CNNs with near-infrared hyperspectral imaging (935–1700 nm) and analytical techniques such as Wavelet Packet Analysis (WPA) and Principal Component Analysis (PCA) has further strengthened non-destructive seed health assessment pipelines. SVM-based hyperspectral approaches allow 100% seed testing without destroying samples, offering a major advantage over traditional microbiological assays. These methods have achieved 100% accuracy in detecting Aspergillus spp. in naturally infected corn kernels. Ensemble boosting algorithms—including CatBoost, Gradient Boosting Decision Trees (GBDT), and XGBoost—have also demonstrated robust performance, each exceeding 97.42% accuracy under diverse contamination levels and seed conditions [
200,
201].
Transformers and Classical Machine Learning for Image & Spectrum Classification
Transformer-based models, originally developed for natural language processing, have been effectively adapted for computer vision tasks in agriculture, improving the identification of disease indicators in seed health assessment [
202]. Vision Transformers (ViTs) use self-attention mechanisms to capture global contextual information in images, offering advantages over traditional CNNs in applications where long-range feature relationships matter. Implementations such as Swin Transformers achieve computational efficiency through reduced parameter counts while preserving high accuracy by using hierarchical feature representations built through shifted-window attention. These models have demonstrated strong performance in plant disease detection, with some studies reporting accuracies of up to 95.5% and outperforming architectures like DenseNet-121, VGG-16, and standard CNNs [
203,
204]. The inherent attention mechanisms in transformer architectures also enable interpretable visualizations that highlight image regions influencing classification decisions.
Hybrid architectures that integrate CNN-based local feature extraction with transformer-based global context modeling are emerging as particularly effective for complex diagnostic tasks requiring both fine-scale detail recognition and broader pattern assessment. Advances in deep learning—especially CNNs and transformers—have enabled non-destructive, high-throughput seed analysis with accuracy rates often exceeding 95% for detecting pathogens, assessing germination potential, and identifying physical defects. Additionally, hyperspectral imaging combined with machine learning continues to show exceptional capability in detecting seed-borne pathogens across multiple crop species, with certain pathogen–seed combinations achieving 100% classification accuracy, as summarized in
Table 9.1.
9.2. Classical Machine Learning for Image and Spectrum Classification
Traditional machine learning remains a steady workhorse in seed health testing, especially for spectral analysis, feature-based image classification, and situations where training data are scarce. Classical models such as Support Vector Machines (SVMs), Random Forests (RF), and ensemble methods offer interpretable decision boundaries and often outperform deep learning when datasets are small or when explainability is essential. These approaches have been successfully used to classify seed and seedling quality using descriptors generated through interactive machine-learning workflows, even with limited sample sizes [
218].
SVMs are widely applied for seed discrimination and classification based on morphological, color, and spectral features. For example, SVM models have achieved over 94% accuracy in identifying peanut seeds contaminated with various fungal species using hyperspectral data spanning 967–2499 nm, demonstrating strong performance in high-dimensional spectral spaces. The kernel trick allows SVMs to manage nonlinear decision boundaries effectively. With appropriate kernel functions—such as radial basis function, polynomial, and sigmoid—SVMs consistently perform well across seed species and pathogen types. Least Squares SVM (LS-SVM) models have also been used to detect Cucumber green mottle mosaic virus in watermelon seeds, achieving 92% accuracy with hyperspectral imaging between 950–2500 nm [
219,
220].
Random Forest classifiers and ensemble methods are equally valuable due to their capacity to aggregate multiple decision trees, improving predictive stability and reducing overfitting—common challenges in high-dimensional biological datasets. RF models deliver robust predictions across diverse seed conditions (Yablokova et al., 2024). In comparative studies, RF has demonstrated superior performance over SVM, C4.5, and AdaBoost, and in dry bean classification for seed certification, RF combined with PCA achieved 95.5% accuracy, outperforming k-Nearest Neighbors (95%) and SVM (93.5%) [
221,
222].
Classical discriminant methods such as Linear Discriminant Analysis (LDA) also remain effective for spectral classification in seed health assessments. LDA and PLS-DA have achieved accuracies above 92% when identifying Fusarium-damaged hard wheat kernels using near-infrared hyperspectral imaging (938–1654 nm). Other studies applying LDA in combination with Quadratic Discriminant Analysis (QDA) or Multiple Discriminant Analysis (MDA) have reported over 90% accuracy for detecting Aspergillus glaucus in canola seeds using hyperspectral data between 1000–1600 nm [
223,
224].
9.4. Dataset Curation and Labeling Strategies for Seed Diagnostics
Effective AI or ML in seed health testing depends on well-curated datasets that capture the full range of seed conditions and pathogen interactions, since dataset quality ultimately shapes model performance and generalizability. Dataset development for seed health applications requires systematic collection across multiple varieties, growing seasons, and storage conditions [
225]. Sample sizes vary widely, from fewer than 100 seeds to more than 47,000 samples, with larger datasets generally supporting stronger model training. However, targeted augmentation methods can partially offset limited sample sizes [
226].
A key consideration during dataset creation is how infected seed samples are generated. Naturally infected seeds (54.5 percent) better reflect real field conditions, while artificially inoculated seeds (40.9 percent) allow controlled experimentation with specific pathogens. Although natural infections still require rigorous validation, laboratory-inoculated seeds have been shown to reliably mimic field-infected kernels [
227].
Agricultural datasets are inherently rich and multimodal, incorporating imaging, spectral, and genomic data. Handling these diverse data types demands careful standardization to maintain consistency and interoperability across sources [
225]. Centralized and standardized dataset frameworks also support efficient deep learning workflows, especially when models are fine-tuned from general non-agricultural datasets [
227]. Strengthening standardizations is particularly important because existing public agricultural datasets remain limited in size and structure, making it difficult to fully leverage higher-capacity models and modern computational power for seed health applications [
229].
9.5. Labeling Strategies for Seed Diagnostics
Accurate labeling and validation rely on combining multiple confirmation methods. This requires close attention to pathogen-specific traits while drawing on molecular and spectroscopy tools such as DNA sequencing, qPCR, LAMP, MultispeQ, and hyperspectral imaging. These approaches strengthen model development and improve disease detection by boosting both specificity and sensitivity. They also support disease management, monitoring, and germplasm screening by enabling discrimination between closely related strains before symptoms appear.
Advanced imaging—particularly hyperspectral and infrared thermal technologies—further enables early detection of diseases that were previously invisible in the field (230,231,232,233).
Seed health studies commonly rely on visual confirmation (54.5%), molecular diagnostics such as PCR (13.6%), and culture-based methods (22.7%). Many studies omit confirmatory validation, even though molecular methods provide the highest specificity, albeit at higher cost and with increased sample destruction [
234].
Robust quality control includes inter-rater reliability assessments and alignment with established diagnostic standards. Integrating plant pathology expertise into AI development ensures biological relevance and technical accuracy. Hierarchical labeling frameworks enable both high-level categories (e.g., healthy vs. infected) and fine-grained, pathogen-specific identification [
235]. Finally, combining omics data with advanced imaging provides deeper insight into plant–pathogen interactions. These integrated datasets help reveal new intervention targets for disease management [
236].
9.6. Dataset Augmentation and Class Imbalance for Seed Diagnostics
Data augmentation is essential for mitigating class imbalance, a recurring issue in seed diagnostics where healthy seeds vastly outnumber pathogen-infected ones. Standard augmentation workflows rely on geometric transformations (e.g., rotation, scaling, translation, flipping) and photometric adjustments (e.g., brightness, contrast, color balance) to expand dataset diversity without altering biological relevance.
Generative Adversarial Networks (GANs) further strengthen diagnostic performance by producing synthetic seed images that mimic real pathogen-induced characteristics. Studies applying Deep Convolutional GANs (DCGANs) and CycleGANs to agricultural disease datasets report notable gains in accuracy and recall, particularly for detecting cotton Fusarium wilt, where GAN-generated images effectively captured the underlying distribution of the training data and alleviated limited-sample constraints [
236,
237].
To address class imbalance more directly, the Synthetic Minority Over-Sampling Technique (SMOTE) remains the most widely adopted strategy. SMOTE generates new minority-class samples by interpolating feature space vectors between existing examples and their nearest neighbors [
238,
239]. Advanced variants—including Borderline-SMOTE, Distance-Based SMOTE (D-SMOTE), and Bi-Phasic SMOTE—focus on regions near decision boundaries, where misclassification risk is highest, thereby improving model sensitivity to subtle pathogen signatures.
In applied seed pathology, SMOTE-integrated pipelines have demonstrated strong diagnostic performance. For example, SMOTE-siPLS-stacking models achieved >99% accuracy in identifying
Penicillium decumbens in Choy Sum seeds, underscoring the value of synthetic sample generation for low-prevalence pathogens [
240,
241].
9.7. Model Explainability and Performance Metrics for Seed Diagnostics
Model explainability is essential in seed health diagnostics, where regulatory compliance and diagnostic accuracy require transparent and traceable decision-making. Techniques such as Class Activation Mapping (CAM) and Gradient-weighted Class Activation Mapping (Grad-CAM) identify which regions of a seed image contribute most to a model’s prediction. Grad-CAM generates heatmaps by computing the gradients of a predicted class score with respect to the activations of the final convolutional layer, allowing clear visualization of the spatial features that influence classification outcomes. In agricultural imaging, Grad-CAM and Grad-CAM++ have effectively highlighted pathogen-associated regions in plant and seed images, enabling interpretable assessments of model behavior and facilitating the detection of potential biases or image artifacts that may compromise diagnostic reliability [
242].
Integrating explainability into seed diagnostic workflows strengthens confidence in AI-assisted decisions by ensuring alignment with expert knowledge and established diagnostic criteria. Visual explanations support verification that models rely on biologically meaningful features rather than non-informative correlations. Recent studies increasingly combine multiple interpretability methods—LIME, SHAP, and Grad-CAM—to provide complementary perspectives on model reasoning, capturing both global feature contributions and localized spatial attributions [
243,
244,
245].
Performance metrics for AI/ML models in seed health assessment become especially tricky under imbalanced datasets and very low pathogen prevalence. Even highly specific assays can produce a low probability of true infection among positive results, underscoring the need for a practical interpretive framework when pathogen incidence in seed lots falls below 1% [
246]. Models often appear highly accurate because healthy seeds dominate; however, they typically struggle with rare positive cases. As a result, they may show high sensitivity but low positive predictive value (PPV), leading to elevated false-positive rates and costly, unnecessary interventions.
Receiver Operating Characteristic (ROC) curves visualize the trade-off between true-positive and false-positive rates across thresholds, with ROC-AUC providing a threshold-independent score. Yet for strongly imbalanced datasets—a constant reality in seed health—Precision-Recall (PR) curves provide a more meaningful evaluation [
247]. PR curves directly compare precision (PPV) and recall (sensitivity), making them better suited to situations where true positives are extremely rare. Empirical work confirms that PR-AUC outperforms ROC-AUC as an indicator of model quality under such conditions [
248,
249].
A thorough assessment should include accuracy, precision, recall, and F1-scores to fully interpret model behavior in low-prevalence scenarios. Merged or composite scores can further clarify performance under class imbalance. Considering both PR-AUC and F1-score is critical for seed health diagnostics because these metrics help identify rare, infected seeds while limiting false positives that drive economic losses. Differentiating healthy from infected high-value vegetable seed lots requires careful evaluation across multiple thresholds, as threshold choice directly influences precision, recall, and downstream commercial viability [
250,
251].
In commercial seed lots—typically showing pathogen incidences near 1%—PR curves remain sensitive to genuine performance shifts, while ROC curves may show inflated AUC values that do not translate to practical usefulness. Understanding PPV and Negative Predictive Value (NPV) is essential in this context. Low disease prevalence depresses PPV even when specificity exceeds 99%, resulting in many false positives, whereas NPV remains high. This imbalance directly affects decisions regarding seed-lot acceptance (as illustrated in
Figure 9.1). Under low prevalence, any positive screen requires confirmatory testing to avoid unnecessary rejections. Meanwhile, high NPV supports using AI-based screening to efficiently identify clean lots needing minimal follow-up.
Effective AI-driven workflows must therefore account for both the true detection of infected seeds and the misclassification of healthy seeds, since each outcome affects economic feasibility. Hypothetical model studies show that optimizing precision and recall is necessary to meet eligibility thresholds across seed-sorting pipelines involving pathogens with different prevalence levels. High average precision reflects strong identification of true positives, especially in datasets that weight the positive class. Although the F1-score balances precision and recall, it does not capture behavior across all thresholds the way PR-AUC does—an important limitation when evaluating the reliability of positive classifications [
252,
253].
Adjusting model thresholds substantially shifts the balance between reducing false positives (higher precision) and reducing false negatives (higher recall), and these trade-offs directly influence economic outcomes in seed testing programs [
254]. By calibrating precision and recall appropriately, seed companies can generate sorted fractions that meet specific quantitative and qualitative requirements [
255].
9.8. Edge Deployment vs. Cloud, MLOps in Accredited Labs, Reproducibility and Versioning
Edge-to-cloud decisions shape the performance, cost, and accessibility of AI-driven seed health systems. Choosing between local (edge) and cloud-based processing always comes with trade-offs in compute power, latency, data privacy, and infrastructure needs.
Edge systems that process data directly inside seed testing labs cut latency, strengthen data security, and keep diagnostics running even when connectivity is spotty. But the flip side is real: updating or retraining models on these devices is harder because of limited compute and irregular internet access [256. For that reason, cloud platforms usually handle the heavy lifting—initial model training, large-scale analytics, and fine-tuning on high-performance, AI-optimized hardware. Models are then adapted and deployed onto edge devices for fast, on-site inference.
This hybrid setup blends the strengths of both worlds. The cloud supports computationally demanding development and large-dataset analysis, while the edge enables real-time decision-making inside agricultural environments [
257]. Together, they address the core challenges of applying AI in agriculture by pairing deep analytical power with immediate, field-level insights [
257,
258].
More advanced hybrid architectures—especially those using federated learning—push this further. By keeping sensitive agricultural data closer to where it originates, they improve privacy and reduce reliance on continuous connectivity. At the same time, they still benefit from shared model improvements across sites [
257]. This approach is designed to strengthen accuracy, reproducibility, and version control in accredited seed labs, ensuring consistent performance even across distributed testing environments.
Federated and distributed AI systems are redefining how plant disease detection is carried out, offering stronger generalization across agricultural contexts while reducing communication overhead and training costs. These efficiencies make earlier disease identification far more attainable [
258]. Decentralized learning also enables the formation of grower federations that can share model insights securely and transparently, strengthening collective forecasting and production decision-making [
259]. Advanced neural architectures—including convolutional neural networks and vision transformers—continue to drive progress in detecting leaf diseases and other crop anomalies, mitigating the substantial economic losses and quality reductions associated with plant health issues [
260,
261].
Federated learning directly addresses data heterogeneity and privacy constraints across farms by supporting joint model training without raw data exchange, producing robust disease-classification models from diverse datasets [
262]. By allowing insights gained on one farm to inform others, federated systems nurture a collaborative, resilient ecosystem for disease surveillance and management [
263]. Demonstrated successes in agriculture highlight strong performance of federated approaches using convolutional architectures and attention mechanisms trained on geographically and biologically diverse image sets [
260].
Tiny Machine Learning (TinyML) further shifts diagnostics toward true edge intelligence, enabling efficient deployment of compact models on low-power embedded devices. Models under 140 KB can now achieve ~96% accuracy while consuming only 2.63 mW/h, making continuous, battery-powered plant-health monitoring feasible [Heydari et al., 2025]. Boards such as the Arduino Nano and ESP32 support real-time inference for inline seed testing. Compression methods—quantization, pruning, and knowledge distillation—preserve model accuracy while reducing computational load by 4–10×, making sophisticated inference viable directly on seed-testing hardware.
Cloud infrastructures complement these edge systems by providing the computational depth needed to train large-scale models and process high test volumes. High-performance GPUs, distributed training environments, and the ability to pool diverse seed datasets enable ongoing model refinement and generalizability. Centralized storage allows rapid incorporation of new data, producing stronger models over time. However, cloud dependence introduces latency concerns for fast-turnaround diagnostics and raises privacy considerations in competitive seed markets. Hybrid cloud–edge architectures therefore provide an optimal compromise: edge devices handle initial screenings and transmit flagged samples or summary features to the cloud for deeper analysis, model updates, and cross-location learning.
In accredited seed-testing laboratories, integrating machine learning operations (MLOps) resolves the practical challenges of deploying and maintaining AI models under strict regulatory and quality assurance expectations. Robust CI/CD frameworks support continuous training, validation, and deployment with full auditability—critical for labs operating under ISTA or ISO/IEC 17,025 accreditation [
264,
265,
266]. Automated testing monitors model accuracy against defined benchmarks and alerts personnel when performance drifts.
Seamless integration with laboratory information systems preserves established QA workflows while enabling efficient AI-supported diagnostics. Version control ensures complete traceability for datasets, model parameters, and configuration changes. Continuous monitoring tracks key operational indicators—including prediction accuracy, throughput, and system reliability—to maintain both diagnostic validity and regulatory compliance [
268]. Drift-detection algorithms identify shifts in seed varieties, imaging conditions, or storage environments that could degrade performance [
269].
Quality assurance for AI-based seed-health analysis relies on routine validation against reference benchmarks, inter-laboratory comparison studies, and proficiency testing programs [
267]. These oversight mechanisms ensure that deployed models remain trustworthy and aligned with accreditation criteria over time [
269]. Emerging audit-based certification systems may soon automate evaluation of MLOps lifecycle artifacts using standards such as ISO 25012, enabling rapid issuance or revocation of compliance certificates as models evolve [
267].
10. Automation and High-Throughput Workflows
Automation and high-throughput workflows are redefining the operational landscape of seed health diagnostics, enabling unprecedented speed, precision, and scalability in testing programs. Traditional manual processes—often labor-intensive and prone to variability—are being replaced by integrated systems that combine robotic liquid handling, microfluidic lab-on-chip platforms, and advanced data management tools. These innovations minimize human error, streamline sample preparation, and support multiplexed molecular assays, dramatically reducing turnaround times while maintaining ISO/IEC 17,025 compliance. By coupling automation with digital traceability frameworks such as barcoding and Laboratory Information Management Systems (LIMS), modern seed-testing laboratories can achieve end-to-end process control, ensuring reproducibility and auditability across global phytosanitary networks. This convergence of robotics, microfluidics, and informatics marks a pivotal shift toward compact, modular, and field-deployable diagnostic stations capable of meeting the growing demands of international seed trade and biosecurity.
10.1. Robotics for Sample Handling and Microfluidic/Lab-on-Chip Systems for Nucleic Acids
Automation in diagnostic workflows reduces human error, increases throughput, and enhances reproducibility. Two major technological pillars are robotic liquid-handling systems and microfluidic lab-on-chip modules for nucleic acid extraction, amplification, and detection.
Automated liquid-handling platforms such as those from Hamilton, Tecan, and Beckman Coulter enable high-precision pipetting, plate transfers, mixing, and dilution under tight software control. When paired with automated seed-processing modules—seed grinding, weighing, and bead-beating homogenizers—the workflow tightens even further, cutting down the hands-on chaos and keeping throughput steady and clean. Integrated with plate stackers, incubators, and thermal cyclers, these systems push truly end-to-end molecular pipelines. [
270] showed that fully automated PCR master-mix prep can match the accuracy and precision of a seasoned tech, even in diagnostic settings. All of this reduces operator variability and helps keep labs in line with ISO/IEC 17,025 accreditation expectations.
Microfluidic chips miniaturize extraction, purification, amplification, and detection processes within a compact footprint, decreasing reagent use and assay time while improving reproducibility. [
271,
272] reviewed integrated lab-on-chip platforms for automated nucleic acid preparation and amplification, highlighting their potential for decentralized diagnostics.
A particularly versatile configuration, digital microfluidics (DMF), manipulates discrete droplets through electrostatic actuation, enabling programmable mixing, splitting, routing, incubation, and detection on a planar chip [
273]. DMF platforms are being increasingly applied to nucleic-acid amplification tests because they permit multiplexing and automation without pumps or valves.
Recent progress has produced fully integrated microfluidic systems that combine sample preparation and amplification on the same device. [
274] introduced a platform coupling magnetic-bead RNA extraction with RT-LAMP, reaching limits of detection of 10–100 RNA copies within about one hour. [
275] reported a DMF chip equipped with dual-mode thermal control that automatically performs nucleic-acid amplification and detection. Likewise, [
276] demonstrated a centrifugal “sample-in–result-out” microfluidic system for multiplexed molecular diagnostics.
Integration of robotics and microfluidics now allows the design of modular “plug-and-play” diagnostic stations. Upstream seed homogenization and lysis can be seamlessly linked to nucleic-acid purification and downstream amplification modules, all coordinated by unified automation software. This convergence is paving the way for compact, high-throughput, and field-deployable seed-testing platforms.
10.2. Barcoding, LIMS Integration, and Electronic Chain of Custody
Traceability and data integrity are critical in automated laboratories. Digital barcoding combined with laboratory information management systems (LIMS) underpins a secure electronic chain of custody.
Each sample vessel—tube, microplate, or cartridge—is labeled with a unique identifier such as a 1D barcode, 2D QR code, or RFID tag. [
277] described digital barcode strategies for high-throughput screening that allow pooled assays and accurate sample retrieval through barcode decoding. For pooled or multiplexed testing, combinatorial barcode schemes can encode sample identity even after mixing.
A robust LIMS records sample origin, barcode, timestamps, processing steps, operator IDs, quality checks, and analytical results. Integration between LIMS and robotic instruments ensures every transfer and operation is automatically logged. Time-stamped digital records of handling, storage, and analysis create a verifiable audit trail compliant with ISO/IEC 17,025 and phytosanitary documentation requirements. Error-correction features such as checksum validation and duplicate-ID alerts further enhance reliability. Collectively, well-implemented barcode-LIMS frameworks provide complete traceability from seed lot to final diagnostic result and reduce sample mix-ups in high-volume laboratories.
10.3. Pooled-Testing Algorithms and Deconvolution Strategies
Pooled testing enables laboratories to screen many subsamples with fewer assays, boosting cost-effectiveness when pathogen prevalence is low. However, demonstrating reproducibility and true throughput gains increasingly requires quantitative comparisons—such as limits of detection (LOD), false-negative/false-positive rates, and reaction-savings metrics—across pooling strategies.
Adaptive or hierarchical pooling tests large pools first and retests only positive ones in smaller sub-pools. [
278] developed an adaptive RT-qPCR deconvolution algorithm that minimizes total reactions while maintaining sensitivity, a performance that can be further validated through LOD and error-rate benchmarking. Combinatorial or matrix pooling assigns each sample to overlapping pools so that intersecting positive pools identify likely positives, allowing computation of diagnostic accuracy under different prevalence scenarios.
Bayesian inference methods refine detection by integrating prior probabilities (e.g., lot-level risk) to estimate sample-specific posterior probabilities. Error-tolerant designs mitigate occasional false positives or negatives through structured retesting rules, making quantitative error-rate evaluation especially important. Throughput-optimized frameworks like the D-Optimal Pooling Experimental (DOPE) design [
279] mathematically balance pool size, expected prevalence, and assay sensitivity, and are strengthened when accompanied by empirical LOD and reproducibility comparisons
Software-based pooling managers now automate these computations, import qPCR results, and output per-sample probabilities. In seed diagnostics, pooling protocols must balance efficiency against the risk of target dilution or heterogeneous infection distribution. Because low-prevalence pathogens rarely play fair, over-pooling can easily bury a weak signal—so every strategy needs to be validated under realistic, messy infection patterns to avoid false negatives. Monte Carlo simulations are recommended to validate pooling parameters across realistic prevalence ranges before adoption in accredited testing programs.
11. Interferences, Pitfalls, and QC
Seed health testing is inherently sensitive to a range of technical interferences and operational pitfalls, which can jeopardize its diagnostic performance if left unaddressed [
280]. With rapid adoption of high-throughput technologies to improve speed, sensitivity, cost, and to detect latent infection, the need for well-defined guardrails has become even more critical for quality assurance [
281]. However, the reliability of seed health assays is constantly challenged by seed matrix inhibition, chemical residue from treatments, and the persistent risk of cross-contamination or amplicon carryover [
280,
282,
283]. Addressing these vulnerabilities is critical to ensure consistent interpretation of diagnostic methods across laboratories and reliable decision making in seed health testing.
Emerging diagnostic platforms are expanding the frontiers of seed health testing, as the integration of high-throughput platforms offers a paradigm shift in phytopathology and plant protection strategies worldwide. While these tools are powerful and, in many cases, nondestructive in nature, they can amplify risks if control strategies are overlooked [
284]. Accordingly, appropriate controls are the cornerstone of robust diagnostic workflows to mitigate false positive and false negative results [
285]. Controls are broadly categorized into process controls (positive/negative), amplification controls (positive, negative, and non-template), and internal amplification controls (IACs). Process controls verify method sensitivity, specificity, and confirm that the reagents and tools function properly [
286]. Positive process controls are known samples containing the target organism that are processed alongside test samples to verify detection performance, while negative process controls contain no target organism and confirm the absence of contamination or false positives, additional process controls including mock communities or spike-ins, help identify contamination or cross-reactivity during sample handling, extraction, and downstream processes [
282,
287,
288]. Amplification controls monitor the integrity of a workflow from sample processing to analysis. These include non-template controls (DNA or RNA free water) to check for contamination, positive amplification controls (DNA or RNA from reference isolates for each primer, especially in multiplex PCR) to confirm assay performance, and negative amplification controls (DNA or RNA from non-target isolates) to ensure assay specificity [
282,
286,
289,
290]. IACs are non-target nucleic acid sequences co-amplified with target sequences to monitor assay efficiency and guard against detection inhibition [
291,
288,
282]. The necessity of these controls varies with the diagnostic platform and are implemented as either essential or optional measures. Together, these layers of controls provide a comprehensive quality assurance critical for accurate interpretation of results and maintaining reproducibility across technicians, laboratories, and diagnostic platforms (
Table 11.1).
Seeds are composed of complex metabolites, including phenolic compounds, lipids, and polysaccharides, which can co-purify during nucleic acid extraction and suppress target detection [
292]. The inhibition is particularly problematic in molecular-based assays, highlighting the need for careful sample preparation. To mitigate these inhibitory effects, optimizing extraction protocols, such as incorporating magnetic bead-based purification, is essential, while the inclusion of IACs remains indispensable to monitor the sensitivity of the detection assay [
281]. On the other hand, machine-learning models trained on limited or poorly annotated datasets can introduce bias, leading to systematic misclassification of diagnostic results [
293]. In the absence of robust validation controls, such as algorithm audit trails and curated reference datasets, AI systems may generate outputs that are difficult to verify, undermining both trust and regulatory acceptance while risking financial loss and eroding confidence in innovative diagnostic technologies [
284,
294]. Ultimately, the reliability of diagnostic methods depends on robust sample preparation and thorough validation to ensure both scientific rigor and trust in routine applications.
Seeds are exposed to various physical or biological treatments to manage pests, diseases, and safeguard seed quality during storage and production cycles [
295,
296]. Their widespread application adds additional layers of complexity to diagnostic pipelines, as the residue from these treatments can influence the analytical performance of seed health tests. In nucleic-acid extraction-based assays, treatment residue can chemically modify or degrade target DNA or RNA templates, resulting in poor extraction yield and reduced amplification efficiency [
297]. Emerging AI-based diagnostics add yet another layer of vulnerabilities. Algorithms trained on untreated data fail to establish treatment-related spectral or morphological changes, requiring domain-appropriate training data and validation to ensure diagnostic robustness [
298]. To mitigate these effects, the assay must be validated on both treated and untreated seed lots to account for chemical interference by incorporating process controls and reference materials.
High-throughput molecular diagnostic platforms (e.g., PCR, next-gen, meta-omics etc.,) are particularly prone to contamination, where traces of DNA or RNA from previous reactions and carryover amplicons are the biggest cause of false positive results [
299]. This contamination can aerosolize during sample preparation, pipetting, or other downstream operational processes, allowing the contaminant template to spread and generate spurious results [
300]. For imaging and AI-based diagnostics, such as fluorescence or hyperspectral imaging, inadequate calibration or poorly defined reference standards can distort classification outcomes [
298]. Similarly, without rigorous negative and positive controls to benchmark infected versus healthy samples, subtle changes in lighting, instrument drift, or seed-surface properties may be misinterpreted as pathogen signals [
301]. This compromises reproducibility across laboratories and reduces confidence in adopting these methods as part of routine seed quality assurance [
302,
303]. In addition to inhibition, contamination remains a critical source of false results. Strengthening laboratory hygiene, workflow separation, routine swab testing, and timely calibration of equipment can help safeguard the precision of diagnostic methods.
Ensuring repeatability and reproducibility is critical to obtain credible seed health testing across all diagnostic platforms. In molecular assays, different nucleic acid extraction protocols, reagents, thermal cycle calibrations, or instrumental sensitivity introduce variation, especially for low-titer pathogen results [
304]. Imaging diagnostic platforms face analogous challenges, were sensor resolution, changes in illumination, or seed orientation can alter spectral assessments, undermining comparability between laboratories [
305]. Algorithm performance depends on training-data diversity and quality, such that a model trained on one seed population or treatment type could yield inconsistent predictions between laboratories [
306]. Reference materials and calibrated standards, such as well-characterized seed panels and standardized imaging databases, can serve as benchmarks between technicians and laboratories [
307]. Establishing standardized processes, robust quality controls, and shared reference frameworks is essential to ensure repeatability, reproducibility, and compatibility in seed health testing across laboratories and regions. As seed testing moves toward integrated molecular, imaging, and AI approaches, unified quality assurance frameworks will be key to building trust, supporting trade, and protecting global food security. Guard-banding decision thresholds is a critical safeguard to reduce uncertainty across diagnostic platforms. In molecular assays, this means retesting borderline Ct values; in imaging, it involves adjusting classification cut-offs; and in AI systems, it requires confidence scoring and human review of ambiguous outputs. Cross-platform harmonization through standardized controls, calibration, curated datasets, and inter-laboratory validation is essential to ensure repeatability and reproducibility. Similarly, the publicly available databases and reference panels will play a key role in promoting consistent and reliable seed health results
12. Performance & Economics
Rapid, reliable, and cost-effective diagnostics are essential to safeguard global agriculture against emerging seed-borne pathogens. Diagnostic tools range from traditional visual checks and culture-based or bioassay methods (direct approaches that confirm viability and pathogenicity) to advanced molecular, imaging, and AI-driven approaches (indirect methods). Each offers unique trade-offs in terms of viability confirmation, turnaround time, cost per sample or lot, accuracy, and environmental impact [
208,
209].
A major challenge in seed health testing is decision-making under low pathogen titer. Although the likelihood of an outbreak is low, the consequences of missing an infection can be severe. Understanding assay sensitivity and specificity is therefore critical for confidence in diagnostic results, guiding interpretation, and defining proportional risk mitigation strategies [
290]. Under low prevalence, the sensitivity–specificity trade-off becomes pronounced. Highly sensitive assays (e.g., molecular or nucleic acid-based tests) reduce false negatives and enable early detection, supporting quarantine and exclusion programs [
290,
310]. However, increased sensitivity can also raise false-positive risks, especially when detecting non-viable or trace nucleic acids, potentially triggering unnecessary interventions, retesting, and trade or financial losses [
311].
To manage these trade-offs, diagnostic guardrails are indispensable. These include assay validation, threshold definition, confirmatory workflows, and the use of control materials across test stages. Traditionally, tiered diagnostic methods have been the gold standard—combining indirect prescreen assays (molecular, serological, or imaging-based) with confirmatory direct assays (bioassay or culture-based tests) to enhance efficiency and reliability [
282]. Recent advances show that molecular approaches can achieve comparable accuracy to conventional bioassays with significantly faster turnaround times [
312]. Moving forward, coordinated efforts among industry, academia, and regulators are needed to refine standards, define acceptable uncertainty thresholds, and institutionalize guardrails for timely, cost-effective decisions under low-titer scenarios.
Beyond accuracy, seed health testing must also be operationally feasible. Conventional methods rely on visualizing pathogen growth on seeds, providing definitive evidence under favorable conditions [
313]. However, these phenotypic methods often lack sensitivity and specificity, making it difficult to distinguish closely related species. For example, culture plating may fail to differentiate morphologically similar species, posing regulatory challenges when one species within a genus is of quarantine importance [
314]. While direct methods are inexpensive, they are time-consuming (days to weeks) and have low throughput. In contrast, molecular diagnostics deliver faster results—often within hours—with improved sensitivity and specificity. They can detect pathogens that are difficult to culture and differentiate species with high genetic similarity [
290]. Automation and multiplexing further enhance throughput but increase operational costs for reagents, instruments, and skilled labor. Imaging-based diagnostics offer non-destructive, high-throughput detection with rapid turnaround, though initial equipment costs are high; per-sample costs decrease significantly at scale.
Integration of AI and ML-driven diagnostics into molecular and imaging workflows is transforming turnaround time and throughput. These technologies enable automated pattern recognition, anomaly detection, and predictive analysis for real-time decision-making in seed health programs [
315,
316]. While AI requires substantial upfront investment for model development and training, the cost per lot decreases once systems are optimized. Future research should focus on standard validation workflows, cost-effective automation, and robust data pipelines to ensure scalability and operational convenience. Bridging next-generation technologies with regulatory guardrails will be key to achieving diagnostic precision and operational sustainability.
Successful implementation also depends on alignment with international regulatory frameworks and seed movement logistics. Seeds often cross multiple borders during production, creating risks of accidental or deliberate pathogen introduction [
311,
317]. Harmonized, science-based phytosanitary requirements are critical for resilient and transparent global seed movement [
318]. Variability in diagnostic protocols and interpretation among national plant protection organizations (NPPOs) can lead to inconsistent risk assessments and trade delays, underscoring the need for global alignment in tools, reference materials, and standards [
309].
Recent advances enable data-driven, risk-based decision frameworks to guide actions such as quarantine, eradication, or commercial release of seed lots. Multiple checkpoints throughout production—from site selection to post-harvest conditioning, treatment, storage, and distribution—create integrated surveillance for early detection and proactive risk management [
319]. A harmonized diagnostic framework enhances transparency, reduces trade disruptions, and strengthens global commitment to sustainable seed production. Building shared data platforms, reference libraries, and joint validation programs among NPPOs, research institutions, and industry will support a trusted diagnostic ecosystem anchored by harmonized guardrails.
Finally, sustainability is emerging as a critical dimension. Both direct and indirect testing methods consume water and energy and generate chemical and plastic waste, contributing to environmental impact. Green lab initiatives—such as energy-efficient equipment, workflow optimization, and recycling programs—are increasingly adopted to mitigate these effects [
320,
321]. Integrating diagnostics with green practices supports institutional sustainability goals and reduces resource use and costs. Environmental guardrails, including carbon accounting, eco-friendly consumables, and waste-minimizing benchmarks, can further embed sustainability without compromising analytical quality [
322]. These measures strengthen compliance with emerging environmental regulations and stakeholder confidence in sustainable seed systems [
320,
321,
322].
In summary, seed health testing must balance operational efficiency, diagnostic performance, regulatory compliance, and environmental responsibility. Accurate and timely detection requires harmonized standards, scalable automation, and AI-enabled approaches, integrated with sustainability practices. This convergence will define next-generation seed health testing as scientifically rigorous, economically viable, and environmentally responsible—capable of supporting global trade in an evolving agricultural landscape.
13. Case Studies in Seed Health Diagnostics
13.1. Viral Pathogens: Tomato Brown Rugose Fruit Virus (ToBRFV) in Tomato and Pepper
Tomato brown rugose fruit virus (ToBRFV) has become a model for understanding seedborne viral transmission in solanaceous crops. [
332] quantified ToBRFV distribution within infected tomato seeds and evaluated both seed-to-seedling transmission and the effectiveness of several disinfection treatments. The virus was localized mainly on the seed coat and occasionally within the endosperm but was absent from the embryo. Transmission occurred at low frequencies—approximately 2.8% in cotyledons and 1.8% in the third true leaf. Although most disinfection treatments inactivated viral infectivity, RT-qPCR still detected viral RNA in six of seven treatments showing that molecular detection does not necessarily equate to infectivity.
Subsequent localization work confirmed that ToBRFV contamination is predominantly external and mechanical. Seeds from infected fruit were almost all contaminated on the surface but true biological transmission rates were extremely low (about 0.08%) [
333]. These findings guided current seed-testing protocols that emphasize large composite samples and external surface testing rather than embryo assays.
13.2. Bacterial Pathogens: Xanthomonas Campestris on Brassica Seeds
For bacterial pathogens such as black rot caused by
Xanthomonas campestris pv.
campestris (Xcc), seedborne inoculum is a major concern. [
334] developed a multiplex real-time PCR assay that could detect one infected seed among 10 000 tested and incorporated an internal plant DNA control to identify false negatives from PCR inhibition has set the analytical sensitivity benchmark for modern seed-lot testing.
Infection localization studies by [
335] used GFP-tagged Xcc and confocal microscopy to track colonization in
Brassica oleracea following flower-cluster spray inoculation. They demonstrated both superficial contamination and under high inoculum pressure, limited penetration into internal tissues such as the endosperm and embryo. These observations explain occasional treatment failures when infections become deep-seated.
At an operational scale, [
336] evaluated physical and chemical disinfection methods for naturally contaminated
B. oleracea seeds. A 3% hydrogen-peroxide treatment for 30 min eliminated detectable Xcc without reducing germination (~95%). The authors emphasized that temperature, drying, and seed-lot size critically influence scalability and reliability in industry settings.
13.3. Fungal Pathogens: Fusarium Species in Cereals and Vegetables
DNA from non-viable
Fusarium propagules can persist after treatment, complicating interpretation of molecular results. [
337] addressed this limitation using a propidium-monoazide (PMA) qPCR method that differentiates viable from dead
Fusarium cells in soil and plant material. Their approach achieved detection limits near 82 spores mL
−1 in suspension and ≈ 91 spores g
−1 in soil offering a more realistic measure of infection risk.
Earlier, [
338] established EF-1α-targeted real-time PCR assays for eleven
Fusarium species in cereals and found strong correlations between quantified DNA and mycotoxin content validating the assays for disease surveillance.
For harvested onions, [
339] introduced a pooled-sample PCR protocol that detects latent
Fusarium oxysporum infection. Subsamples from 50 bulbs were combined and dual markers—species identification and pathogenicity genes were amplified from FTA-card or kit-extracted DNA. This workflow increased throughput for screening symptomless bulbs in commercial storage.
13.4. Lessons Learned and Industry Adoption
These examples illustrate key principles for implementing scalable, regulatory-ready diagnostics:
Pathogen localization drives sampling and disinfection strategy. ToBRFV remains largely external on seed coats; therefore, surface assays and disinfection protocols targeting the testa are most effective [
332,
333].
Detection does not always equal infectivity. RT-qPCR or PCR can amplify residual nucleic acids from inactivated pathogens causing risk assessment must incorporate viability indicators [
337,
340].
Analytical sensitivity and real-world validation are critical. Methods must detect extremely low prevalence i.e.
, one infected seed in 10 000 for Xcc—but also remain robust across large seed lots and diverse matrices [
334,
336].
Integrating molecular, imaging, and viability assays yields deeper insight. Confocal imaging verified infection depth for Xcc [
335], while PMA-qPCR demonstrated functional viability for
Fusarium [
337].
Together these studies have informed modern seed-testing frameworks, emphasizing statistically grounded sampling, viability assessment, and clear interpretation criteria that distinguish detection from transmission risk.
14. Data Standards & Traceability
14.1. Introduction
Modern seed health testing increasingly depends on digital infrastructure for managing diagnostic data, certification records and phytosanitary documentation. As diagnostic technologies evolve from conventional assays to high-throughput molecular and imaging systems, data volumes have expanded exponentially. Without standardized frameworks, these datasets become siloed, reducing their value for regulatory and commercial traceability [
340].To ensure interoperability and long-term usability, the
FAIR data principles—that all data should be
Findable, Accessible, Interoperable and
Reusable—have become the cornerstone of digital seed testing systems [
341]. These principles, when applied to seed health diagnostics, promote consistency across laboratories, facilitate international data exchange, and enhance transparency throughout the seed value chain [
342].
In addition, traceability mechanisms—whether audit trails, version-controlled records or digital certificates are now mandatory under phytosanitary frameworks such as the International Plant Protection Convention (IPPC) and the Organization for Economic Co-operation and Development (OECD) Seed Schemes [
343]. These frameworks aim to establish verifiable links between diagnostic results, seed lot identity and export certification, thereby strengthening biosecurity and market confidence.
14.3. Audit Trails, Data Integrity and Compliance
14.3.1. Digital Audit Trails and Chain-of-Custody
A robust audit trail underpins traceable diagnostic workflows. Every data transaction—from seed sampling to report generation—should be time-stamped, digitally signed and linked to authenticated user credentials. This digital chain-of-custody ensures data authenticity and accountability [
348].
In seed-testing laboratories, audit trails are usually managed through laboratory information management systems (LIMS) that log sample accession, reagent batches, instrument calibration, and analyst actions. Such systems facilitate compliance with
ISO/IEC 17,025 accreditation, which requires demonstrable data traceability for all analytical results [
347].
In high-throughput environments, an integrated LIMS can automatically capture image-based pathogen-detection data, store raw files on secure servers and generate immutable PDF/A reports for submission to regulatory authorities. Cryptographic hashing or checksum validation methods are often applied to prevent post-hoc data modification [
349].
14.3.2. Regulatory Compliance and Standardization
International seed trade depends on compliance with phytosanitary frameworks that mandate data traceability. The
International Plant Protection Convention (IPPC) requires phytosanitary certificates to include verifiable details about commodity origin, treatments, and diagnostic history [
344]. Similarly, the
OECD Seed Schemes link each certified lot to the testing laboratory and competent authority (OECD, 2022). Modern compliance audits emphasize electronic document management and metadata traceability. Laboratories adopting
ISO/IEC 17025-compliant LIMS systems report reduced transcription errors and faster audit readiness (Goble et al., 2020).
14.4. Linking Diagnostics to Certificates and Phytosanitary Documentation
Integration of diagnostic data with certification platforms allows verifiable linkage between analytical results and trade documentation. The
IPPC ePhyto Solution provides a global system for National Plant Protection Organizations (NPPOs) to issue and exchange digital phytosanitary certificates [
344].
Within the European Union, the
TRACES NT (Trade Control and Expert System – Next Generation) platform records electronic documentation for movement of plants and plant products, including seed consignments. Authorized users can attach laboratory test reports directly to shipment records [
350].
In the United States, the
Phytosanitary Certificate Issuance and Tracking System (PCIT) managed by USDA APHIS links laboratory test results to issued export certificates, ensuring verifiable traceability to original seed lots [
351]. Collectively, these systems illustrate a global shift toward electronic phytosanitary certification that enhances transparency and reduces fraud.
14.4.1. Blockchain and Distributed Ledger Options
Blockchain has been explored to ensure immutability of certification records. A blockchain ledger can store hashed identifiers of diagnostic results, offering tamper-proof verification without revealing confidential data [
352].
However, large-scale adoption faces challenges in governance, scalability, and regulatory acceptance [
353]. Most authorities currently favor hybrid architectures in which traditional databases manage day-to-day operations while distributed ledgers store integrity-verification tokens [
349].
In practice, digitally signed XML certificates combined with secure API-based validation achieve similar integrity benefits without the complexity of full blockchain integration.
14.5. Pragmatic Approaches for Global Harmonization
Data traceability initiatives must balance technological sophistication with practical feasibility. For many seed-testing laboratories, especially in developing regions, infrastructure constraints hinder rapid adoption of advanced blockchain or AI-integrated systems. Gradual digital transformation strategies emphasizing open standards and incremental interoperability are therefore more sustainable [
342].
Key priorities for harmonization include:
Common metadata standards – adopting MIAPPE and Darwin Core vocabularies across diagnostic databases.
API-based integration – enabling secure data exchange between LIMS, ePhyto, TRACES, and national systems.
Persistent identifiers – assigning DOIs or UUIDs to seed lots, diagnostic assays, and certificates.
Capacity building – training laboratory data managers and phytosanitary officers in FAIR implementation.
Phased blockchain adoption – piloting distributed-ledger systems before national deployment.
Aligning diagnostic data standards with global certification systems will create a transparent, auditable and efficient pathway from laboratory testing to international trade.
15. Ethics, IP, and Adoption
The rapid adoption of molecular, imaging, and artificial intelligence (AI)–based diagnostics in seed health testing has raised complex ethical and legal questions about data ownership, intellectual property (IP), and equitable access. Traditional seed testing relied on open, standardized methods documented by organizations such as the International Seed Testing Association (ISTA). However, newer diagnostic technologies increasingly depend on proprietary datasets, algorithms, and patented reagents, creating tension between innovation and transparency [
354].
This section examines the ethical and regulatory landscape governing modern seed diagnostic technologies, focusing on proprietary AI training data, licensing of molecular assays, and the balance between open science and commercial protection. It also addresses the importance of external validation, biosafety, and bio-surveillance frameworks that ensure responsible deployment in global agriculture.
15.1. Proprietary Datasets and AI Model Governance
AI-based seed health diagnostics such as deep-learning models for hyperspectral imaging or pathogen classification are trained on large, labeled datasets of seed and pathogen images. The quality and bias of these datasets directly affect diagnostic performance. Yet many such datasets are proprietary, owned by commercial developers or private laboratories, limiting reproducibility and independent benchmarking [
355].
From an ethical perspective, data opacity undermines scientific scrutiny and can perpetuate model bias toward certain crops, regions, or pathogens [
356]. In regulated diagnostics, transparency is essential for external validation and accreditation. The OECD Principles on Artificial Intelligence and the FAIR AI guidelines emphasize that algorithms influencing phytosanitary decisions must be explainable and auditable [
357,
358].
To address this, several initiatives advocate open-access AI repositories for agricultural diagnostics, analogous to genomic databases like GenBank. These efforts encourage data standardization (e.g., using MIAPPE metadata), model interpretability, and independent performance verification before regulatory acceptance [
359].
15.2. Ethical Use of Proprietary Data
Ethical governance requires balancing innovation incentives with the public interest. Private companies often invest substantially in curating image libraries, sequencing data, and model architectures. However, withholding such resources can slow scientific progress and widen gaps between high- and low-income testing systems.
A pragmatic approach involves tiered access models, where anonymized training data and performance metrics are shared publicly, while proprietary parameters remain protected. This mirrors practices in human health AI under the EU’s General Data Protection Regulation (GDPR) and the U.S. National Artificial Intelligence Research Resource (NAIRR) framework [
358].
15.3. Licensing of Molecular Assays: Primers, Probes, and Patents
The licensing of molecular diagnostics especially primers, probes, and isothermal amplification systems presents growing IP challenges in the seed health domain. Many primer or probe sequences are patented under biotechnology or diagnostic-use claims [
360].
For instance, proprietary loop-mediated isothermal amplification (LAMP) assays and probe-based qPCR kits for
Clavibacter michiganensis or
Acidovorax citrulli may be covered by exclusive use agreements. This can restrict laboratories from reproducing or modifying assays without licensing, even when the underlying genomic sequences are public [
361].
While patent protection incentivizes innovation, it may also hinder assay harmonization and method validation across regions. The World Intellectual Property Organization (WIPO) has encouraged open licensing frameworks for agricultural diagnostics, particularly in contexts affecting global food security.
15.4. Balancing Innovation and Accessibility
Molecular diagnostics often rely on shared sequence databases such as GenBank and EMBL, where pathogen genomes are publicly available. Ethical conflicts arise when these public resources underpin privately patented assays, leading to “knowledge privatization” [
362]. One proposed solution is the dual licensing model, where diagnostic assays for non-commercial research are open, but commercial applications require licensing fees. This model, used in plant breeding and veterinary diagnostics, could be adapted for seed health testing [
361]. Additionally, collaborative validation under ISTA or EPPO frameworks can help ensure that licensed assays are independently verified, protecting users from over-reliance on vendor claims [
363].
15.4. Transparency, Validation, and External Oversight
A recurring challenge in diagnostic regulation is balancing commercial confidentiality with scientific transparency. Developers of diagnostic assays or AI models may withhold key design details such as algorithmic parameters or reagent compositions to protect trade secrets. However, regulators and accreditation bodies require sufficient information to evaluate reliability and reproducibility [
364].
Best practice now emphasizes tiered transparency, where developers disclose validation data, accuracy metrics, and reference panels while retaining proprietary components. This approach aligns with the European Commission’s AI Act and ISO/IEC 23894:2023 (AI Risk Management), which require explainability without mandating full disclosure of intellectual property [
365].
15.5. Training, Safety, and Biosurveillance Implications
The adoption of advanced diagnostic systems requires specialized training in bioinformatics, data stewardship, and biosafety. Laboratories transitioning from conventional to AI-assisted or molecular diagnostics must ensure personnel are competent in both analytical and ethical dimensions of technology use [
367].
Capacity building programs under the FAO International Plant Protection Convention (IPPC) and the OECD Seed Schemes have emphasized competency-based training to ensure standardized implementation across diverse laboratory infrastructures [
357,
364].
15.6. Biosafety and Dual-Use Risks
Advanced diagnostic systems, particularly those involving synthetic biology or genomic data, carry dual-use potential; the risk that legitimate research may be repurposed for harmful applications [
368]. Seed health laboratories handling exotic or regulated pathogens must therefore implement stringent biosafety and biosecurity measures consistent with Biosafety Level 2 or 3 (BSL-2/3) protocols.
AI-driven pathogen prediction models also raise biosurveillance concerns, as predictive algorithms could inadvertently reveal vulnerabilities in agricultural systems [
365]. Ethical deployment requires clear data governance policies, ensuring diagnostic data are not misused for non-agricultural surveillance or trade manipulation.
Integrating diagnostic data into biosurveillance networks—such as the Global Plant Health Information System (GPHIS) can strengthen early warning systems for transboundary pest and pathogen threats [
364]. However, such integration must respect data sovereignty and privacy, especially when AI is used to predict disease spread from proprietary datasets. Transparent data-sharing agreements and adherence to FAIR and CARE (Collective Benefit, Authority to Control, Responsibility, Ethics) principles are recommended [
369].
15.7. Summary and Outlook
Ethical and IP considerations represent the next frontier in seed diagnostic innovation. As the industry embraces AI and molecular platforms, balancing openness, security, and commercial interests will determine adoption success. Regulatory frameworks must evolve to address these dynamics, supporting ethical innovation ecosystems where proprietary rights coexist with scientific integrity, biosafety, and global accessibility.
Future progress will rely on transparent validation pipelines, fair licensing of molecular reagents, open AI training datasets, and harmonized biosurveillance governance. These measures will ensure that technological advancement in seed diagnostics continues to enhance and not compromise global agricultural resilience and biosecurity.
16. Conclusions & Practitioners
The landscape of seed health diagnostics has shifted profoundly in recent years. What was once a largely manual, culture-based discipline has become a technologically integrated field linking molecular biology, imaging, automation, and artificial intelligence. Laboratories now operate within data-rich ecosystems where accuracy, reproducibility, and traceability are as essential as pathogen detection itself. This review has shown that the future of seed health testing will rely on the intelligent combination of classical and emerging methods, not on the displacement of one by another. Practitioners must therefore balance scientific rigor, regulatory compliance, and technological agility to ensure that diagnostic innovation serves the broader goals of food security, seed trade integrity, and global biosecurity (370; 366).
16.1. Synthesis and Outlook
Seed health testing is entering a period of diagnostic convergence that unites traditional microbiological methods with molecular, imaging, and AI-based approaches (Van der Wolf et al., 2020; Leung et al., 2021). This shift is driven by the need for speed, scalability, and confidence in results across diverse laboratory infrastructures. Integration remains the key theme: combining rapid molecular screens, viability confirmation, and robust data management within harmonized quality frameworks (347; 371).
Standardization will define credibility in the coming decade. International alignment through ISTA, EPPO, and IPPC is advancing validation pathways and promoting digital traceability that ensures test results can be verified across borders (363,366). Artificial intelligence tools must also meet regulatory expectations for transparency and interpretability, with clear documentation of training data and algorithmic performance (356,358). The challenge is to maintain biological realism while embracing computational precision.
Seed diagnostic systems that successfully blend molecular and classical methods will form the foundation of future phytosanitary programs. They offer a path toward reliable, high-throughput, and ethically governed diagnostics that can support both commercial and public-sector goals (372)
16.2. Method Selection Framework by Pathogen Class and Objective
Objective –based Application
Surveillance: Rapid LAMP or LFIA field screening
Quarantine or Certification: ISO 17025-validated qPCR or grow-out confirmation (347)
Research or Discovery: Metagenomics and AI-assisted phenotyping (98)
Routine Industry Testing: Multiplex qPCR combined with imaging for reproducibility and throughput (335)
16.3. Implementation Roadmap for Diagnostic Laboratories
Best practices for transition
Begin with pilot targets, expand gradually, retain legacy assays during evaluation, maintain comprehensive records under FAIR data principles (340), and continuously train personnel in molecular and bioinformatics competencies (366).
16.4. Practitioner Checklist
✅ Method Selection: Align assays with pathogen biology, seed matrix, and national regulations.
✅ Validation: Satisfy ISO 17,025 criteria for sensitivity, specificity, repeatability, and reproducibility (347).
✅ Controls: Incorporate internal and external controls for each batch
✅ Traceability: Use LIMS systems with full audit trails (348).
✅ Data Management: Apply FAIR principles and standardized metadata such as MIAPPE or Darwin Core (340,346).
✅ Ethics and IP: Verify reagent and software licensing; maintain AI transparency (358,354).
✅ Training: Record personnel competency across all diagnostic platforms (343).
✅ Sustainability: Optimize protocols for minimal waste and energy efficiency (342).
16.5. Closing Perspective
Modern seed diagnostics are evolving into integrated systems that unite biology, data science, and regulatory discipline. Laboratories that combine validated molecular and imaging platforms with transparent data governance will define the next generation of seed health assurance. The future of global seed movement depends on such networks of interoperable, ethical, and scientifically credible diagnostics that protect both agricultural productivity and biodiversity.
Author Contributions
Conceptualization, CB; methodology, CB; validation, CB, HMS, and TJR; formal analysis, CB; investigation, CB; resources, all authors; data curation, CB; writing—original draft preparation, CB; writing—review and editing, CB, HMS, and TJR; visualization, CB; supervision, CB; project administration, CB. Section contributions: Introduction, SM; Regulatory & Standards Landscape, SM; Sampling Theory & Study Design, CB; Conventional Diagnostics, ANT; Molecular Diagnostics—Core Methods, JS; Next-Gen & Meta-Omics, HMS; CRISPR-Based Diagnostics, HMS; Imaging & Non-Destructive Phenotyping, HMS; AI/ML for Seed Diagnostics, TJR; Automation & High-Throughput Workflows, SK; Interferences, Pitfalls, and QC, SK; Performance & Economics, SK; Case Studies, CB; Data Standards & Traceability, CB; Ethics, IP, and Adoption, CB; Future Directions, CB; Conclusions & Practitioner Checklist, CB. HMS and TJR also contributed to overall review and organization during the final stages of the manuscript. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
No new data were created or analyzed in this study. Data sharing is not applicable to this article.
Acknowledgments
The authors gratefully acknowledge the extensive body of research and innovation in seed health diagnostics that informed this review. While it was not possible to include every significant contribution, we recognize and appreciate the efforts of scientists, institutions, and industry partners whose work continues to advance molecular, imaging, and AI-based approaches for seed quality assurance. Their collective contributions have laid the foundation for the progress discussed in this manuscript. The authors have reviewed and edited all outputs and take full responsibility for the final content of this publication.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Abdelfattah, A.; Tack, A. J.; Lobato, C.; Wassermann, B.; Berg, G. From seed to seed: The role of microbial inheritance in the assembly of the plant microbiome. Trends in Microbiology 2023, 31(4), 346–355. [Google Scholar] [CrossRef] [PubMed]
- Barret, M.; Guimbaud, J. F.; Darrasse, A.; Jacques, M. A. Plant microbiota affects seed transmission of phytopathogenic microorganisms. Molecular Plant Pathology 2016, 17(6), 791. [Google Scholar] [CrossRef]
- Brunel, S.; Horn, N. M.; Unger, J. G.; Arnitis, R. Implementation of international standards for phytosanitary measures No. 7 phytosanitary certification system and No. 12 phytosanitary certificates. EPPO Bulletin 2013, 43(2), 309–315. [Google Scholar] [CrossRef]
- Buddenhagen, C. E.; Rubenstein, J. M.; Hampton, J. G.; Rolston, M. P. The phytosanitary risks posed by seeds for sowing trade networks. PLOS ONE 2021, 16(11), e0259912. [Google Scholar] [CrossRef] [PubMed]
- Choudhury, R. A.; Garrett, K. A.; Klosterman, S. J.; Subbarao, K. V.; McRoberts, N. A framework for optimizing phytosanitary thresholds in seed systems. Phytopathology 2017, 107(10), 1219–1228. [Google Scholar] [CrossRef]
- Cortes, J. Overview of the regulatory framework in seed trade. In Proceedings of the Second World Seed Conference, Rome, 2009, September; FAO. [Google Scholar]
- Hulme, P. E. Unwelcome exchange: International trade as a direct and indirect driver of biological invasions worldwide. One Earth 2021, 4(5), 666–679. [Google Scholar] [CrossRef]
- Dwivedi, S. L.; Spillane, C.; Lopez, F.; Ayele, B. T.; Ortiz, R. First the seed: Genomic advances in seed science for improved crop productivity and food security. Crop Science 2021, 61(3), 1501–1526. [Google Scholar] [CrossRef]
- Gebeyaw, M. Review on: Impact of seed-borne pathogens on seed quality. American Journal of Plant Biology 2020, 5, 77–81. [Google Scholar] [CrossRef]
- Gitaitis, R.; Walcott, R. The epidemiology and management of seedborne bacterial diseases. Annual Review of Phytopathology 2007, 45(1), 371–397. [Google Scholar] [CrossRef]
- Haider, M. W.; Mishra, M. K.; Fayaz, F.; Ahmad, D.; Mehmood, A. Evaluating the impact of seed-borne mycoflora on seed quality and health of various Oryza sativa varieties. International Journal of Microbiology 2024, 2024(1), 1263308. [Google Scholar] [CrossRef]
- Hiddink, G. A.; Willmann, R.; Woudenberg, J. H.; Souza-Richards, R. Seed health testing: Doing things right. PhytoFrontiers™ 2023, 3(1), 71–74. [Google Scholar] [CrossRef]
- Jones, M.; Marengo, S. Laboratory validation, verification, and accreditation of molecular methods. In Molecular microbial diagnostic methods; Academic Press, 2016; pp. 107–133. [Google Scholar] [CrossRef]
- Kachhap, B.; Pandey, S.; Banerjee, M.; Pandey, A. K.; Kumari, N. Comprehensive review of seed-borne pathogens: Challenges and control in crop production. Current Agriculture Research Journal 2025, 13(2), Article 3. [Google Scholar] [CrossRef]
- Keshavulu, K. International treaties for seed quality assurance: An Indian perspective. In Indian seed sector: Evolution, technology, trade and impact; Springer Nature Singapore, 2025; pp. 343–352. [Google Scholar] [CrossRef]
- Klaedtke, S. M.; Rey, F.; Groot, S. P. Designing a seed health strategy for organic cropping systems, based on a dynamic perspective on seed and plant health. Sustainability 2022, 14(17), 10903. [Google Scholar] [CrossRef]
- Kuhlmann, K.; Nalinya, A. N.; Francis, T.; Spielman, D. J. A comparative study of the legal and regulatory dimension of seed sector development in Sub-Saharan Africa. Agricultural Systems 2025, 228, 104351. [Google Scholar] [CrossRef]
- Lamichhane, J. R.; Venturi, V. Synergisms between microbial pathogens in plant disease complexes: A growing trend. Frontiers in Plant Science 2015, 6, 385. [Google Scholar] [CrossRef]
- Mancini, V.; Romanazzi, G. Seed treatments to control seedborne fungal pathogens of vegetable crops. Pest Management Science 2014, 70(6), 860–868. [Google Scholar] [CrossRef]
- Singh, S. K.; Jha, R.; Pandey, S.; Mohan, C.; Ghosh, S.; Kumari, S.; Singh, A. Artificial intelligence-based tools for next-generation seed quality analysis. In Crop Design; 2025; p. 100094. [Google Scholar] [CrossRef]
- Martín, I.; Gálvez, L.; Guasch, L.; Palmero, D. Fungal pathogens and seed storage in the dry state. Plants 2022, 11(22), 3167. [Google Scholar] [CrossRef]
- McGuire, S.; Sperling, L. The links between food security and seed security: Facts and fiction that guide response. In Global food-price shocks and poor people; Routledge, 2014; pp. 39–54. [Google Scholar] [CrossRef]
- Milivojević, M.; Ripka, Z.; Petrović, T. ISTA rules changes in seed germination testing at the beginning of the 21st century. Journal on Processing and Energy in Agriculture 2018, 22(1), 40–45. [Google Scholar] [CrossRef]
- Mishra, U.; Goswani, S. P.; Chauhan, S.; Kumar, M. P. Principles of quality seed production. In Recent Advances in Seed Science and Technology 1; 2023; pp. 118–134. [Google Scholar] [CrossRef]
- Misra, M. K.; Harries, A.; Dadlani, M. Role of seed certification in quality assurance. In Seed science and technology; Springer, 2023; pp. 267–298. Available online: https://www.researchgate.net/publication/376396735_Role_of_Seed_Certification_in_Quality_Assurance.
- Nallathambi, P.; Umamaheswari, C.; Lal, S. K.; Manjunatha, C.; Berliner, J. Mechanism of seed transmission and seed infection in major agricultural crops in India. In Seed-borne diseases of agricultural crops: Detection, diagnosis & management; Springer Singapore, 2020; pp. 749–791. [Google Scholar] [CrossRef]
- Okezue, M. A.; Adeyeye, M. C.; Byrn, S. J.; Abiola, V. O.; Clase, K. L. Impact of ISO/IEC 17025 laboratory accreditation in sub-Saharan Africa: A case study. BMC Health Services Research 2020, 20(1), 1065. [Google Scholar] [CrossRef]
- Verma, N. V.; Shukla, M.; Kulkarni, R.; Srivastava, K.; Claudic, B.; Savara, J.; Mathew, M. J.; Maurya, R.; Bhattacharjee, G.; Singh, V.; Pandya, A. Emerging extraction and diagnostic tools for detection of plant pathogens: Recent trends, challenges, and future scope. ACS Agricultural Science & Technology 2022, 2(5), 858–881. [Google Scholar] [CrossRef]
- Elias, S. Seed quality testing. In Handbook of seed science and technology; CRC Press, 2024; pp. 561–601. [Google Scholar] [CrossRef]
- Petter, F.; Suffert, M.; McMullen, M.; Griessinger, D.; Roy, A. S. Seed-borne pests and phytosanitary issues: The role of EPPO. In Global perspectives on the health of seeds and plant propagation material; Springer Netherlands, 2014; pp. 29–46. [Google Scholar] [CrossRef]
- Vishunavat, K.; Prabakar, K.; Anand, T. Seed health: Testing and management. In Seed health; Dadlani, M., Ed.; Springer, 2023; p. p. 335. [Google Scholar]
- Walcott, R. R. Detection of seedborne pathogens. HortTechnology 2003, 13(1), 40–47. [Google Scholar] [CrossRef]
- Mahant, V.; Pal, P. Seeding the future: A review of artificial intelligence’s pivotal role in seed technology. In Journal of Crop Science and Biotechnology; 2025; pp. 1–14. [Google Scholar] [CrossRef]
- Morrison, R. H. Sampling in seed health testing. Phytopathology 1999, 89(11), 1084–1087. [Google Scholar] [CrossRef]
- Jennelle, C. S.; Runge, M. C.; MacKay, N.; Converse, K. A. Applying a Bayesian weighted surveillance approach to detect disease presence. Journal of Applied Ecology 2018, 55, 2470–2479. [Google Scholar] [CrossRef]
- Grant, E. H. C.; Mummah, R. O.; Mosher, B. A.; Evans, J.; DiRenzo, G. V. Inferring pathogen presence when sample misclassification and partial observation occur. Methods in Ecology and Evolution 2023, 14(6), 1323–1337. [Google Scholar] [CrossRef]
- Cuturi, M.; Teboul, O.; Berthet, Q.; Doucet, A.; Vert, J.-P. Noisy adaptive group testing using Bayesian sequential experimental design. arXiv. 2020. Available online: https://arxiv.org/abs/2004.12508.
- Daon, Y.; Huppert, A.; Obolski, U. DOPE: D-Optimal pooling experimental design with application for SARS-CoV-2 screening. Journal of the American Medical Informatics Association 2021, 28(12), 2562–2570. [Google Scholar] [CrossRef] [PubMed]
- Grider, J. F.; Udell, B. J.; Reichert, B. E.; Foster, J. T.; Kendall, W. L.; Cheng, T. L.; Frick, W. F. A novel method for estimating pathogen presence, prevalence, load, and dynamics at multiple scales. Scientific Reports 15 2025, 9423. [Google Scholar] [CrossRef]
- Sunani, S. K.; Bashyal, B. M.; Rawat, K.; Manjunatha, C.; Sharma, S.; Prakash, G.; Aggarwal, R. Development of PCR and loop-mediated isothermal amplification assay for the detection of bakanae pathogen Fusarium fujikuroi. European Journal of Plant Pathology 2019, 154(3), 715–717. [Google Scholar] [CrossRef]
- Madhusaisri, S.; B, R.; Pradeep, M.; D, B. Seed Health Techniques for Detection and Management of Alternaria Spp. in Sesame. Journal of Advances in Microbiology 2025, 25(3), 51–58. [Google Scholar] [CrossRef]
- Sundareswaran, S.; Choudhury, P. R.; Vanitha, C.; Yadava, D. K. Seed quality: Variety development to planting—An overview. In Seed Science and Technology; Springer Nature Singapore; 2023; pp. 1–16. [Google Scholar] [CrossRef]
- Buddenhagen, C. E.; Andrade-Piedra, J.; Forbes, G. A.; Kromann, P.; Navarrete, I.; Thomas-Sharma, S.; Xing, Y.; Choudhury, R. A.; Andersen, K. F.; Schulte-Geldermann, E.; Garrett, K. A. Management performance mapping and the value of information for regional prioritization of management interventions. bioRxiv 2018. [Google Scholar] [CrossRef]
- Batzer, J. C.; Shirazi, A.; Gill, D.; Platner-Heidt, E.; Nicolaus, K.; Bray, A.; Mohan, K.; Mathew, F. M.; Mueller, D. S. Seedborne Fungal Detection Differs with Seed Assay Method, and Fungal Diversity and Abundance Are Impacted by Fungicide Treatment, Harvest Timing, and Storage Environment. PhytoFrontiers™ 2024, 4(4), 767–780. [Google Scholar] [CrossRef]
- Kumar, P. L.; Cuervo, M.; Kreuze, J. F.; Muller, G.; Kulkarni, G.; Kumari, S. G.; Massart, S.; Mezzalama, M.; Alakonya, A.; Muchugi, A. Phytosanitary interventions for safe global germplasm exchange and the prevention of transboundary pest spread: the role of CGIAR germplasm health units. Plants 2021, 10(2), 328. [Google Scholar] [CrossRef]
- Santhy, V.; Sandra, N.; Ravishankar, K. V.; Chidambara, B. Molecular Techniques for Testing Genetic Purity and Seed Health. In Seed Science and Technology: Biology, Production, Quality; Dadlani, M., Yadava, D. K., Eds.; Springer Nature Singapore; 2023; pp. 365–389. [Google Scholar] [CrossRef]
- Ishimwe, R.; Abutaleb, K.; Ahmed, F. Applications of thermal imaging in agriculture—A review. Advances in Remote Sensing 2014, 3(3), 128–140. [Google Scholar] [CrossRef]
- Lundström, C.; Lindvall, M. Mapping the landscape of care providers’ quality assurance approaches for AI in diagnostic imaging. Journal of digital imaging 2023, 36(2), 379–387. [Google Scholar] [CrossRef]
- Naseri, S.; Shukla, S.; Hiwale, K.; Jagtap, M. M.; Gadkari, P.; Gupta, K.; Deshmukh, M.; Sagar, S. From pixels to prognosis: A narrative review on artificial intelligence’s pioneering role in colorectal carcinoma histopathology. Cureus 2024, 16(4), e59171. [Google Scholar] [CrossRef]
- Yan, L.; Li, Q.; Fu, K.; Zhou, X.; Zhang, K. Progress in the application of artificial intelligence in ultrasound-assisted medical diagnosis. Bioengineering 2025, 12(3), 288. [Google Scholar] [CrossRef]
- Mangena, P. Analysis of correlation between seed vigour, germination and multiple shoot induction in soybean (Glycine max L. Merr.). Heliyon 2021, 7(9), e07913. [Google Scholar] [CrossRef]
- Goswami, S. K.; Manzar, N.; Kashyap, A. S.; Kumar, R. Contribution of individuals and organizations in the development of seed pathology. In Seed-Borne Diseases of Agricultural Crops: Detection, Diagnosis & Management; Springer, 2020; pp. 65–80. [Google Scholar] [CrossRef]
- Bhandari, A. Revolutionizing Radiology With Artificial Intelligence. Cureus 2024, 16(10), e72646. [Google Scholar] [CrossRef] [PubMed]
- Sindhu, A.; Jadhav, U.; Ghewade, B.; Bhanushali, J.; Yadav, P.; JADHAV, U. Revolutionizing pulmonary diagnostics: a narrative review of artificial intelligence applications in lung imaging. Cureus 2024, 16(4). [Google Scholar] [CrossRef]
- Frischie, S.; Miller, A. L.; Pedrini, S.; Kildisheva, O. A. Ensuring seed quality in ecological restoration: native seed cleaning and testing. Restoration Ecology 2020, 28, S239–S248. [Google Scholar] [CrossRef]
- Xing, M.; Long, Y.; Wang, Q.; Tian, X.; Fan, S.; Zhang, C.; Huang, W. Physiological alterations and nondestructive test methods of crop seed vigor: A comprehensive review. Agriculture 2023, 13(3), 527. [Google Scholar] [CrossRef]
- Şahbaz, S.; Munkvold, G.; Arias, S.; Tok, F. M. Interaction of seedling-pathogens with physiological seed quality affecting soybean emergence and seedling growth. International Journal of Innovative Approaches in Agricultural Research 2023, 7(4), 630–637. [Google Scholar] [CrossRef]
- Ejeta, M.; Haile, M.; Bayisa, E. Assessment of seed quality and seed-borne pathogens of barley (Hordeum vulgare L.) seed collected from formal and informal seed sources. Journal of Plant Pathology & Microbiology 2022, 13(6), 614. [Google Scholar] [CrossRef]
- Franić, I.; Cleary, M.; Aday Kaya, A. G.; Bragança, H.; Brodal, G.; Cech, T. L.; Chandelier, A.; Doğmuş-Lehtijärvi, T.; Eschen, R.; Lehtijärvi, A.; Ormsby, M.; Prospero, S.; Schwanda, K.; Sikora, K.; Szmidla, H.; Talgø, V.; Tkaczyk, M.; Vettraino, A. M.; Perez-Sierra, A. The Biosecurity Risks of International Forest Tree Seed Movements. Current Forestry Reports 2024, 10(2), 89–102. [Google Scholar] [CrossRef]
- Klaedtke, S. M.; Rey, F.; Groot, S. P. Designing a seed health strategy for organic cropping systems, based on a dynamic perspective on seed and plant health. Sustainability 2022, 14(17), 10903. [Google Scholar] [CrossRef]
- Ali, M.; Irfan, M.; Ali, T.; Wei, C. R.; Akilimali, A. Artificial intelligence in dental radiology: a narrative review. Annals of medicine and surgery (2012) 2025, 87(4), 2212–2217. [Google Scholar] [CrossRef] [PubMed]
- Parvin, N.; Joo, S. W.; Jung, J. H.; Mandal, T. K. Multimodal AI in Biomedicine: Pioneering the Future of Biomaterials, Diagnostics, and Personalized Healthcare. Nanomaterials 2025, 15(12), 895. [Google Scholar] [CrossRef]
- Boshra, M. H.; El-Housseiny, G. S.; Farag, M. M. S.; Aboshanab, K. M. Evaluation of ELISA and immunoaffinity fluorometric analytical tools of four mycotoxins in various food categories. AMB Express 2023, 13(1), 123. [Google Scholar] [CrossRef]
- Bhat, A. I.; Aman, R.; Mahfouz, M. Onsite detection of plant viruses using isothermal amplification assays. Plant Biotechnology Journal 2022, 20(10), 1859–1873. [Google Scholar] [CrossRef] [PubMed]
- Ishino, Y.; Shinagawa, H.; Makino, K.; Amemura, M.; Nakata, A. Nucleotide sequence of the IAP gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. Journal of Bacteriology 1987, 169(12), 5429–5433. [Google Scholar] [CrossRef] [PubMed]
- Venbrux, M.; Crauwels, S.; Rediers, H. Current and emerging trends in techniques for plant pathogen detection. Front Plant Sci 2023, 14, 1120968. [Google Scholar] [CrossRef]
- Omidfar, K.; Riahi, F.; Kashanian, S. Lateral Flow Assay: A Summary of Recent Progress for Improving Assay Performance. Biosensors 2023, 13(9), 837. Available online: https://www.mdpi.com/2079-6374/13/9/837. [CrossRef] [PubMed]
- Xinying, Y.; Xin, L.; Lili, Y.; Qiuyue, Z.; Yongzhe, P.; Jijuan, C. Detection of Cucumber green mottle mosaic virus in low-concentration virus-infected seeds by improved one-step pre-amplification RT-qPCR. Plant Methods 2022, 18(1), 70. [Google Scholar] [CrossRef]
- Moghadam, B. Y.; Connelly, K. T.; Posner, J. D. Two orders of magnitude improvement in detection limit of lateral flow assays using isotachophoresis. Analytical chemistry 2015, 87(2), 1009–1017. [Google Scholar] [CrossRef]
- Munyaneza, V.; Chen, D.; Hu, X. Detection of seed vigour differences in Festuca sinensis seed lots. Seed Science and Technology 2022, 50(1), 61–75. [Google Scholar] [CrossRef]
- Molin, C.; Ribeiro, N. R.; Matusomoto, M. N.; Lütkemeyer, A. J.; Bordignon, K. B.; Ferreira, M. L. B.; Barbieri, M.; Deuner, C. C.; Huzar-Novakowiski, J. Seed treatment for controlling damping-off caused by Globisporangium irregulare and Globisporangium ultimum var. sporangiiferum in soybean from southern Brazil. Crop Protection 2021, 149, 105782. [Google Scholar] [CrossRef]
- Mathur, S. B.; Kongsdal, O. Common laboratory seed health testing methods for detecting fungi; International Seed Testing Association (ISTA): Bassersdorf/Copenhagen, 2003; p. 425 pp. ISBN 3-906549-35-6/978-3906549354. [Google Scholar]
- Boonham, N.; Kreuze, J.; Winter, S.; van der Vlugt, R.; Bergervoet, J.; Tomlinson, J.; Mumford, R. Methods in virus diagnostics: From ELISA to next generation sequencing. Virus Research 186 2014, 20–31. [Google Scholar] [CrossRef]
- Martinelli, F.; Scalenghe, R.; Davino, S.; Panno, S.; Scuderi, G.; Ruisi, P.; Villa, P.; Stroppiana, D.; Boschetti, M.; Goulart, L. R. Advanced methods of plant disease detection: A review. Agronomy for Sustainable Development 2015, 35(1), 1–25. [Google Scholar] [CrossRef]
- Jung, Y. J.; Lee, H. J.; Bae, S.; Kim, J. H.; Kim, D. H.; Kim, H. K.; Nam, K. H.; Nogoy, F. M.; Cho, Y. G.; Kang, K. K. Acquisition of seed dormancy breaking in rice (Oryza sativa L.) via CRISPR/Cas9-targeted mutagenesis of OsVP1 gene. Plant Biotechnology Reports 13 2019, 511–520. [Google Scholar] [CrossRef]
- Shahjahan, T.; Javed, B.; Sharma, V.; Tian, F. Overview of Various Components of Lateral-Flow Immunochromatography Assay for the Monitoring of Aflatoxin and Limit of Detection in Food Products: A Systematic Review. Chemosensors 2023, 11(10), 520. [Google Scholar] [CrossRef]
- Hepp, J.; Shiraishi, M.; Tran, M.; Henson, E.; Ananthanarayanan, M.; Soangra, R. Exploring Teslasuit’s Potential in Detecting Sequential Slip-Induced Kinematic Changes among Healthy Young Adults. Sensors 2023, 23(14), 6258. [Google Scholar] [CrossRef]
- Grimault, V.; Olivier, V.; Rolland, M.; Darrasse, A.; Jacques, M. Chapter 7: Validated seed health testing methods. international rules for seed testing, effective 1 january 2024, 7 21: Detection of Xanthomonas axonopodis pv. phaseoli and Xanthomonas axonopodis pv. phaseoli var. fuscans in Phaseolus vulgaris (bean) seed. Technical report. The International Seed Testing Association (ISTA), Richtiarkade 18, CH-8304 Wallisellen, Switzerland, 2024. [Google Scholar]
- Mirica, A. C.; Stan, D.; Chelcea, I. C.; Mihailescu, C. M.; Ofiteru, A.; Bocancia-Mateescu, L. A. Latest Trends in Lateral Flow Immunoassay (LFIA) Detection Labels and Conjugation Process. Front Bioeng Biotechnol 2022, 10, 922772. [Google Scholar] [CrossRef]
- Anfossi, L.; Di Nardo, F.; Cavalera, S.; Giovannoli, C.; Baggiani, C. Multiplex Lateral Flow Immunoassay: An Overview of Strategies towards High-throughput Point-of-Need Testing. Biosensors 2019, 9(1), 2. Available online: https://www.mdpi.com/2079-6374/9/1/2. [CrossRef]
- Kim, J.; Shin, M.-S.; Shin, J.; Kim, H.-M.; Pham, X.-H.; Park, S.-M.; Kim, D.-E.; Kim, Y. J.; Jun, B.-H. Recent trends in lateral flow immunoassays with optical nanoparticles. International Journal of Molecular Sciences 2023, 24(11), 9600. [Google Scholar] [CrossRef]
- Park, J. Lateral Flow Immunoassay Reader Technologies for Quantitative Point-of-Care Testing. Sensors 2022, 22(19), 7398. Available online: https://www.mdpi.com/1424-8220/22/19/7398. [CrossRef] [PubMed]
- Anfossi, L.; Di Nardo, F.; Cavalera, S.; Giovannoli, C.; Baggiani, C. Multiplex lateral flow immunoassay: An overview of strategies towards high-throughput point-of-need testing. Biosensors 2018, 9(1), 2. [Google Scholar] [CrossRef]
- Jones, A.; Dhanapala, L.; Kankanamage, R. N. T.; Kumar, C. V.; Rusling, J. F. Multiplexed Immunosensors and Immunoarrays. Analytical chemistry 2020, 92(1), 345–362. [Google Scholar] [CrossRef] [PubMed]
- Vincelli, P.; Tisserat, N. Nucleic acid–based pathogen detection in applied plant pathology. Plant Disease 2008, 92(5), 660–669. [Google Scholar] [CrossRef] [PubMed]
- Schrader, C.; Schielke, A.; Ellerbroek, L.; Johne, R. PCR inhibitors: Occurrence, properties and removal. Journal of Applied Microbiology 2012, 113(5), 1014–1026. [Google Scholar] [CrossRef]
- Mbofung, G. C. Y.; Pryor, B. M.; Anand, G.; Kapoor, R.; A PCR-based assay for detection of Fusarium oxysporum f. sp. lactucae in lettuce seed. Nested PCR assay for specific and sensitive detection of Alternaria carthami. Plant Disease;Archives of Microbiology 2010, 94(7) 202(1), 860–866 171–179. [Google Scholar] [CrossRef]
- Hadas, R.; Kritzman, G.; Klietman, F.; Gefen, T.; Manulis, S. Comparison of extraction procedures and determination of the detection threshold for Clavibacter michiganensis ssp. michiganensis in tomato seeds. Plant pathology 2005, 54(5), 643–649. [Google Scholar] [CrossRef]
- Robene-Soustrade, I.; Legrand, D.; Gagnevin, L.; Chiroleu, F.; Laurent, A.; Pruvost, O. Multiplex nested PCR for detection of Xanthomonas axonopodis pv. allii from onion seeds. Applied and Environmental Microbiology 2010, 76(9), 2697–2703. [Google Scholar] [CrossRef]
- Zelyüt, F. R.; Ertunç, F.; Şenal, D. The association of 16SrVI and 16SrI phytoplasma groups with carrot seeds and weeds in Ankara and Konya provinces in Turkey. Plant Protection Bulletin 2022, 62(1), 24–33. [Google Scholar] [CrossRef]
- Sanna, M.; Spadaro, D.; Gullino, M. L.; Mezzalama, M. Optimization of a Loop-Mediated Isothermal Amplification Assay for On-Site Detection of Fusarium fujikuroi in Rice Seed. Agronomy 2021, 11(8), 1580. [Google Scholar] [CrossRef]
- Bustin, S. A.; Benes, V.; Garson, J. A.; Hellemans, J.; Huggett, J.; Kubista, M.; Wittwer, C. T. The MIQE guidelines: Minimum information for publication of quantitative real-time PCR experiments. Clinical Chemistry 2009, 55(4), 611–622. [Google Scholar] [CrossRef]
- Brochu, A. S.; Dumonceaux, T. J.; Valenzuela, M.; Bélanger, R.; Pérez-López, E. A New Multiplex TaqMan qPCR for Precise Detection and Quantification of Clavibacter michiganensis in Seeds and Plant Tissue. Plant Disease 2024, 108(8), 2272–2282. [Google Scholar] [CrossRef]
- Sarkes, A.; Yang, Y.; Dijanovic, S.; Fu, H.; Zahr, K.; Harding, M. W.; Feng, J. Detection of Xanthomonas translucens pv. undulosa, pv. translucens, and pv. secalis by quantitative PCR. Plant Disease 2022, 106(11), 2876–2883. [Google Scholar] [CrossRef]
- Fanelli, V.; Cariddi, C.; Finetti-Sialer, M. Selective detection of Pseudomonas syringae pv. tomato using dot blot hybridization and real-time PCR. Plant Pathology 2007, 56(4), 683–691. [Google Scholar] [CrossRef]
- Peňázová, E.; Dvořák, M.; Ragasová, L.; Kiss, T.; Pečenka, J.; Čechová, J.; Eichmeier, A. Multiplex real-time PCR for the detection of Clavibacter michiganensis subsp. michiganensis, Pseudomonas syringae pv. tomato and pathogenic Xanthomonas species on tomato plants. PLoS ONE 2020, 15(1), e0227559. [Google Scholar] [CrossRef]
- Ciampi-Guillardi, M.; Ramiro, J.; Moraes, M. H. D. D.; Barbieri, M. C. G.; Massola, N. S., Jr. Multiplex qPCR assay for direct detection and quantification of Colletotrichum truncatum, Corynespora cassiicola, and Sclerotinia sclerotiorum in soybean seeds. Plant Disease 2020, 104(11), 3002–3009. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Tian, Q.; Zhou, P.; Zhao, W.; Sun, X. Evaluation of droplet digital PCR for the detection of black canker disease in tomato. Plant Disease 2022, 106(2), 395–405. [Google Scholar] [CrossRef] [PubMed]
- Murolo, S.; Moumni, M.; Mancini, V.; Allagui, M. B.; Landi, L.; Romanazzi, G. Detection and quantification of Stagonosporopsis cucurbitacearum in seeds of Cucurbita maxima using droplet digital polymerase chain reaction. Frontiers in Microbiology 2022, 12, 764447. [Google Scholar] [CrossRef] [PubMed]
- Huggett, J. F. The digital MIQE guidelines update: minimum information for publication of quantitative digital PCR experiments for 2020. Clinical chemistry 2020, 66(8), 1012–1029. [Google Scholar] [CrossRef]
- Notomi, T.; Okayama, H.; Masubuchi, H.; Yonekawa, T.; Watanabe, K.; Amino, N.; Hase, T. Loop-mediated isothermal amplification of DNA. Nucleic Acids Research 2000, 28(12), e63. [Google Scholar] [CrossRef]
- Suzaki, K.; Sawada, H.; Kisaki, G. Loop-mediated isothermal amplification of bacterial effector genes to detect Pseudomonas syringae pv. actinidiae biovars 1 and 3. Journal of General Plant Pathology 2022, 88(1), 2–9. [Google Scholar] [CrossRef]
- Ortega, S. F.; Tomlinson, J.; Hodgetts, J.; Spadaro, D.; Gullino, M. L.; Boonham, N. Development of loop-mediated isothermal amplification assays for the detection of seedborne fungal pathogens Fusarium fujikuroi and Magnaporthe oryzae in rice seed. Plant Disease 2018, 102(8), 1549–1558. [Google Scholar] [CrossRef] [PubMed]
- Tan, X.; Liu, M.; Deng, J.; Gong, S.; Gao, Y.; Li, D.; Zhang, J.; Ruan, C.; Sun, W.; Hu, Y.; Peng, Z. Rapid detection of Frogeye leaf spot pathogen in seeds by LAMP assays to protect soybean production. Plant Disease 2025, 109(10), Article PDIS-10-24-2156-SR. [Google Scholar] [CrossRef]
- Zhang, S.; Duan, M.; Li, S.; Hou, J.; Qin, T.; Teng, Z.; Hu, J.; Zhang, H.; Xia, X. Current status of recombinase polymerase amplification technologies for the detection of pathogenic microorganisms. Diagnostic Microbiology and Infectious Disease 108 2023, 116097. [Google Scholar] [CrossRef]
- Anbazhagan, P.; Parameswari, B.; Anitha, K.; Chaitra, G. V.; Bajaru, B.; Rajashree, A.; Mangrauthia, S. K.; Yousuf, F.; Chalam, V. C.; Singh, G. P. Advances in plant pathogen detection: Integrating recombinase polymerase amplification with CRISPR/Cas systems. 3 Biotech 2024, 14(9), 214. [Google Scholar] [CrossRef]
- Liang, X.; Zhang, X.; Xi, K.; Liu, Y.; Jijakli, M. H.; Guo, W. Development of an RPA-based CRISPR/Cas12a assay in combination with a lateral flow strip for rapid detection of toxigenic Fusarium verticillioides in maize. Food Control 157 2024, 110172. [Google Scholar] [CrossRef]
- Cai, L.; Tian, Q.; Meng, Q.; Bao, X.; Xu, P.; Liu, J.; Wang, H. Recombinase polymerase amplification assay for rapid field diagnosis of Stewart’s wilt of corn pathogen Pantoea stewartii subsp. stewartii. Agriculture 2023, 13(10). [Google Scholar] [CrossRef]
- Capote, N.; Bertolini, E.; Cambra, M. Using RT-PCR for plant RNA virus detection. CABI Reviews 2009, 1–17. [Google Scholar] [CrossRef]
- Wang, J.; Li, A.; Mao, C.; Li, X.; Zhang, C. Recombinase polymerase amplification-lateral flow dipstick for rapid detection of Xanthomonas oryzae pv. oryzae in rice. Crop Protection 191 2025, 107153. [Google Scholar] [CrossRef]
- Tiberini, A.; Manglli, A.; Taglienti, A.; Vučurović, A.; Brodarič, J.; Ferretti, L.; Luigi, M.; Gentili, A.; Mehle, N. Development and Validation of a One-Step Reverse Transcription Real-Time PCR Assay for Simultaneous Detection and Identification of Tomato Mottle Mosaic Virus and Tomato Brown Rugose Fruit Virus. Plants 2022, 11(4), 489. [Google Scholar] [CrossRef]
- Ling, K. S.; Wechter, W. P.; Jordan, R. Development of a one-step immunocapture real-time TaqMan RT-PCR assay for the broad-spectrum detection of Pepino mosaic virus. Journal of Virological Methods 2007, 144(1–2), 65–72. [Google Scholar] [CrossRef] [PubMed]
- Bakker, D.; Bruinsma, M.; Dekter, R. W.; Toonen, M. A. J.; Verhoeven, J. T. J.; Koenraadt, H. M. S. Detection of PSTVd and TCDVd in seeds of tomato using real-time RT-PCR. EPPO Bulletin 2015, 45(1), 14–21. [Google Scholar] [CrossRef]
- Xinying, Y.; Xin, L.; Lili, Y.; Qiuyue, Z.; Yongzhe, P.; Jijuan, C. Detection of cucumber green mottle mosaic virus in low-concentration virus-infected seeds by improved one-step pre-amplification RT-qPCR. Plant Methods 2022, 18(1), 70. [Google Scholar] [CrossRef]
- Nocker, A.; Richter-Heitmann, T.; Montijn, R.; Schuren, F.; Kort, R. Discrimination between live and dead cells in bacterial communities from environmental water samples analyzed by 454 pyrosequencing. International Microbiology 2010, 13(2), 59–65. [Google Scholar] [CrossRef]
- Tian, Q.; Feng, J. J.; Hu, J.; Zhao, W. J. Selective detection of viable seed-borne Acidovorax citrulli by real-time PCR with propidium monoazide. Scientific Reports 2016, 6(1), 35457. [Google Scholar] [CrossRef]
- Wang, H.; Wagnon, R.; Moreno, D.; Timilsina, S.; Jones, J.; Vallad, G.; Turechek, W. W. A long-amplicon viability-qPCR test for quantifying living pathogens that cause bacterial spot in tomato seed. Plant Disease 2022, 106(5), 1474–1485. [Google Scholar] [CrossRef]
- Tedersoo, L.; Anslan, S.; Bahram, M.; Kõljalg, U.; Abarenkov, K. Best practices in metabarcoding of fungi: From experimental design to publication in mycological ecology. Molecular Ecology 2022, 31(20), 4885–4910. [Google Scholar] [CrossRef]
- Piombo, E.; Abdelfattah, A.; Droby, S.; Cocolin, L. Metagenomics approaches for the detection and characterisation of plant pathogenic fungi and bacteria in high-throughput sequencing diagnostics. Microorganisms 2021, 9(1), 188. [Google Scholar] [CrossRef]
- Guenay-Greunke, Y.; Droege, G.; Hohlfeld, M.; Gemeinholzer, B. Handling of targeted amplicon sequencing data focusing on index hopping, primer bias, and read depth variation. Scientific Reports 11 2021, 19803. [Google Scholar] [CrossRef]
- Sharpton, T. J. An introduction to the analysis of shotgun metagenomic data. Frontiers in Plant Science 5 2014, 209. [Google Scholar] [CrossRef]
- Torres-Cortés, G.; Garcia, B. J.; Nerva, L.; Marasco, R.; Venturi, V. Functional microbial features driving community assembly in seed and seedling microbiota. Frontiers in Plant Science 9 2018, 902. [Google Scholar] [CrossRef]
- Buytaers, F. E.; Fraiture, M. A.; Berbers, B.; Vandermassen, E.; Hoffman, S.; Papazova, N.; Vanneste, K.; Marchal, K.; Roosens, N. H. C.; De Keersmaecker, S. C. J. A shotgun metagenomics approach to detect and characterize unauthorized genetically modified microorganisms in microbial fermentation products. Food Chemistry: Molecular Sciences 2021, 2, 100023. [Google Scholar] [CrossRef]
- Pust, M.-M.; Tümmler, B. Identification of core and rare species in metagenome samples based on shotgun metagenomic sequencing, Fourier transforms, and spectral comparisons. ISME Communications 2021, 1(1), 2. [Google Scholar] [CrossRef] [PubMed]
- Tsedaley, B. Review on seed health tests and detection methods of seedborne diseases. Journal of Biology, Agriculture and Healthcare 2015, 5(5). Available online: http://www.iiste.org/Journals/index.php/JBAH/article/view/20641/21576.
- McDonald, M. B. Seed quality assessment. Seed Science Research 1998, 8(2), 265–276. [Google Scholar] [CrossRef]
- Elias, S. Seed quality testing. In Handbook of Seed Science and Technology; CRC Press, 2024; pp. 561–601. [Google Scholar]
- Arnal Barbedo, J. G. Digital image processing techniques for detecting, quantifying and classifying plant diseases. SpringerPlus 2013, 2(1), 660. [Google Scholar] [CrossRef] [PubMed]
- Bock, C. H.; Poole, G. H.; Parker, P. E.; Gottwald, T. R. Plant disease severity estimated visually, by digital photography and image analysis, and by hyperspectral imaging. Critical Reviews in Plant Sciences 2010, 29(2), 59–107. [Google Scholar] [CrossRef]
- Sahoo, J. P.; Samal, K. C. Molecular diagnostic techniques in seed purity assessment. Biotica Research Today 2020, 2(6), 503–505. [Google Scholar] [CrossRef]
- Sendekie, Y. Review on seed genetic purity for quality seed production. International Journal of Scientific Engineering and Science 2020, 4(10), 1–7. [Google Scholar]
- Akhtar, J.; Kandan, A.; Singh, B.; Kumar, P.; Khan, Z.; Gawade, B. H.; Kumar, S.; Dubey, S. C. Diagnostics of seed-borne plant pathogens for safe introduction and healthy conservation of plant genetic resources. In Current Trends in Plant Disease Diagnostics and Management Practices; Springer International Publishing, 2016; pp. 429–440. [Google Scholar] [CrossRef]
- Koonin, E. V.; Makarova, K. S. Origins and evolution of CRISPR-Cas systems. Philosophical Transactions of the Royal Society B 2019, 374(1772), 20180087. [Google Scholar] [CrossRef]
- Kaminski, M. M.; Abudayyeh, O. O.; Gootenberg, J. S.; Zhang, F.; Collins, J. J. CRISPR-based diagnostics. Nature Biomedical Engineering 2021, 5(7), 643–656. [Google Scholar] [CrossRef] [PubMed]
- Del Giovane, S.; Bagheri, N.; Di Pede, A. C.; Chamorro, A.; Ranallo, S.; Migliorelli, D.; Burr, L.; Paoletti, S.; Altug, H.; Porchetta, A. Challenges and perspectives of CRISPR-based technology for diagnostic applications. TrAC Trends in Analytical Chemistry 2024, 172, 117594. [Google Scholar] [CrossRef]
- Lagner, J. R.; Newberry, E. A.; Rivera, Y.; Zhang, L.; Vakulskas, C. A.; Qi, Y. Amplification-free detection of plant pathogens by improved CRISPR-Cas12a systems: A case study on phytoplasma. Frontiers in Plant Science 2025, 16, 1544513. [Google Scholar] [CrossRef]
- Jaybhaye, S. G.; Chavhan, R. L.; Hinge, V. R.; Deshmukh, A. S.; Kadam, U. S. CRISPR-Cas assisted diagnostics of plant viruses and challenges. Virology 2024, 597, 110160. [Google Scholar] [CrossRef] [PubMed]
- Liu, G. Advancing CRISPR/Cas biosensing with integrated devices. ACS Sensors 2025, 10(2), 575–576. [Google Scholar] [CrossRef]
- Wang, M.; Zhang, R.; Li, J. CRISPR/Cas systems redefine nucleic acid detection: Principles and methods. Biosensors and Bioelectronics 2020, 165, 112430. [Google Scholar] [CrossRef]
- Hille, F.; Charpentier, E. CRISPR-Cas: Biology, mechanisms and relevance. Philosophical Transactions of the Royal Society B: Biological Sciences 2016, 371(1707), 20150496. [Google Scholar] [CrossRef]
- Manghwar, H.; Lindsey, K.; Zhang, X.; Jin, S. CRISPR/Cas system: Recent advances and future prospects for genome editing. Trends in Plant Science 2019, 24(12), 1102–1125. [Google Scholar] [CrossRef]
- Li, S.-Y.; Cheng, Q.-X.; Liu, J.-K.; Nie, X.-Q.; Zhao, G.-P.; Wang, J. CRISPR-Cas12a has both cis- and trans-cleavage activities on single-stranded DNA. Cell Research 2018, 28(4), 491–493. [Google Scholar] [CrossRef] [PubMed]
- Fonfara, I.; Richter, H.; Bratovic, M.; Le Rhun, A.; Charpentier, E. The CRISPR-associated DNA-cleaving enzyme Cpf1 also processes precursor CRISPR RNA. Nature 2016, 532(7599), 517–521. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.-Y.; Oh, S.-W. Filtration-based LAMP-CRISPR/Cas12a system for the rapid, sensitive and visualized detection of Escherichia coli O157:H7. Talanta 241 2022, 123186. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Cao, X.; Meng, Y.; Richards, D.; Wu, J.; Ye, Z. Drop CRISPR: A LAMP-Cas12a based digital method for ultrasensitive detection of nucleic acid. Biosensors and Bioelectronics 211 2022, 114377. [Google Scholar] [CrossRef]
- Liu, H.; Wang, J.; Zeng, H.; Liu, X.; Jiang, W.; Wang, Y. RPA-Cas12a-FS: A frontline nucleic acid rapid detection system for food safety based on CRISPR-Cas12a combined with recombinase polymerase amplification. Food Chemistry 334 2021, 127608. [Google Scholar] [CrossRef]
- Xu, J.; Ma, J.; Li, Y.; Kang, L.; Yuan, B.; Li, S. General RPA-CRISPR/Cas12a sensing platform for Brucella spp. detection in blood and milk samples. Sensors and Actuators B: Chemical 364 2022, 131864. [Google Scholar] [CrossRef]
- Zheng, C.; Wang, K.; Zheng, W.; Cheng, Y.; Li, T.; Cao, B.; Jin, Q. Rapid developments in lateral flow immunoassay for nucleic acid detection. Analyst 2021, 146(5), 1514–1528. [Google Scholar] [CrossRef]
- Zhu, H.; Zhang, H.; Xu, Y.; Lassakova, S.; Korabecna, M.; Neuzil, P. PCR past, present and future. Biotechniques 2020, 69(4), 317–325. [Google Scholar] [CrossRef]
- Aman, R.; Mahas, A.; Mahfouz, M. Nucleic acid detection using CRISPR/Cas biosensing technologies. ACS Synthetic Biology 2020, 9(5), 1226–1233. [Google Scholar] [CrossRef]
- Anders, C.; Niewoehner, O.; Duerst, A.; Jinek, M. Structural basis of PAM-dependent target DNA recognition by the Cas9 endonuclease. Nature 2014, 513(7519), 569–573. [Google Scholar] [CrossRef] [PubMed]
- Broughton, J. P.; Deng, X.; Yu, G.; Fasching, C. L.; Servellita, V.; Singh, J.; Miao, X. CRISPR–Cas12-based detection of SARS-CoV-2. Nature Biotechnology 2020, 38(7), 870–874. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Zhao, T.; Li, J.; Liu, X.; Wang, W.; Huang, X.; Liu, Y.; Sun, Y.; Wei, P. Ultrasensitive and on-site diagnosis of rice bakanae disease based on CRISPR-LbCas12a coupled with LAMP. Pest Management Science 2024. [Google Scholar] [CrossRef]
- Ali, Z.; Sanchez, E.; Tehseen, M.; Mahas, A.; Marsic, T.; Aman, R.; Sivakrishna Rao, G. Bio-Scan: A CRISPR/dCas9-based lateral flow assay for rapid, specific, and sensitive detection of SARS-CoV-2. ACS Synthetic Biology 2022, 11(2), 406–419. [Google Scholar] [CrossRef]
- Ali, Z.; Aman, R.; Mahas, A.; Rao, G.S.; Tehseen, M.; Marsic, T.; Salunke, R. Iscan: An Rt-lamp-coupled CRISPR-Cas12 module for rapid, sensitive detection of Sars-CoV-2. Virus Res. 2020, 288, 198129. [Google Scholar] [CrossRef]
- Ding, X.; Yin, K.; Li, Z.; Lalla, R. V.; Ballesteros, E.; Sfeir, M. M.; Liu, C. Ultrasensitive and visual detection of SARS-CoV-2 using all-in-one dual CRISPR-Cas12a assay. Nature Communications 11 2020, 4711. [Google Scholar] [CrossRef]
- Marsic, T.; Ali, Z.; Tehseen, M.; Mahas, A.; Hamdan, S.; Mahfouz, M. Vigilant: An engineered VirD2-Cas9 complex for lateral flow assay-based detection of SARS-CoV-2. Nano Letters 2021, 21(8), 3596–3603. [Google Scholar] [CrossRef]
- Gupta, A.; Srivastava, M.; Mahanta. Offline handwritten character recognition using neural network. IEEE International Conference on Computer Applications and Industrial Electronics (ICCAIE), 2011; IEEE; pp. 102–107. [Google Scholar] [CrossRef]
- Kaliramesh, S.; Chelladurai, V.; Jayas, D. S.; Alagusundaram, K.; White, N. D. G.; Fields, P. G. Detection of infestation by Callosobruchus maculatus in mung bean using near-infrared hyperspectral imaging. Journal of Stored Products Research 52 2013, 107–111. [Google Scholar] [CrossRef]
- Neethirajan, S.; Karunakaran, C.; Jayas, D. S.; White, N. D. G. Detection techniques for stored-product insects in grain. Food Control 2007, 18(2), 157–166. [Google Scholar] [CrossRef]
- Singh, C. B.; Jayas, D. S.; Paliwal, J.; White, N. D. G. Detection of insect-damaged wheat kernels using near-infrared hyperspectral imaging. Journal of Stored Products Research 2009, 45(2), 151–155. [Google Scholar] [CrossRef]
- Chambers, J; McKevitt, N J; Stubbs, M R. Nuclear magnetic resonance spectroscopy for studying the development and detection of the grain weevil, Sitophilus granarius (L.) (Cole-optera: Curculionidae), within wheat kernels. Bull Entom Res. 1984, 74, 707–724. [Google Scholar] [CrossRef]
- Maghirang, E. B.; Dowell, F. E.; Baker, J. E.; Throne, J. E. Automated detection of single wheat kernels containing live or dead insects using near-infrared reflectance spectroscopy. Transactions of the ASAE 2003, 46(4), 1277–1282. [Google Scholar] [CrossRef]
- Toews, M. D.; Pearson, T. C.; Campbell, J. F. Imaging and automated detection of Sitophilus oryzae L. (Coleoptera: Curculionidae) pupae in hard red winter wheat. Journal of Economic Entomology 2006, 99(2), 583–592. [Google Scholar] [CrossRef]
- Gunasekaran, S.; Paulsen, M. R.; Shove, G. C. Optical methods for non-destructive quality evaluation of agricultural and biological materials. Journal of Agricultural Engineering Research 32 1985, 209–241. [Google Scholar] [CrossRef]
- Huang, M.; Wang, Q. G.; Zhu, Q. B. Review of seed quality and safety tests using optical sensing technologies. Seed Science and Technology 2015, 43(3), 337–366. [Google Scholar] [CrossRef]
- Nicolai, B. M.; Beullens, K.; Bobelyn, E. Non-destructive measurement of fruit and vegetable quality by means of NIR spectroscopy: A review. Postharvest Biology and Technology 2007, 46(2), 99–118. [Google Scholar] [CrossRef]
- Kim, S. S.; Phyu, M. R.; Kim, J. M.; Lee, S. H. Authentication of rice using near infrared reflectance spectroscopy. Cereal Chemistry 2003, 80(3), 346–349. [Google Scholar] [CrossRef]
- Dowell, F. E.; Throne, J. E.; Baker, J. E. Automated non-destructive detection of internal insect infestation of wheat kernels by using near-infrared reflectance spectroscopy. Journal of Economic Entomology 1998, 91(4), 899–904. [Google Scholar] [CrossRef]
- Zayas, I.; Pomeranz, Y.; Lai, F. S. Discrimination of wheat and nonwheat components in grain samples by image analysis. Cereal Chemistry 1989, 66(3), 233–237. Available online: https://www.cerealsgrains.org/publications/cc/backissues/1989/Documents/66_233.pdf.
- El Masry, G.; Mandour, N.; Al-Rejaie, S.; Belin, E.; Rousseau, D. Recent applications of multispectral imaging in seed phenotyping and quality monitoring: An overview. Sensors 2019, 19(5), 1090. [Google Scholar] [CrossRef]
- Ring, E. F. J. The historical development of temperature measurement in medicine. Infrared Physics & Technology 2007, 49(3), 297–301. [Google Scholar] [CrossRef]
- Damcevski, K.; Annis, P.; Waterford, C. Effect of grain on apparent respiration of adult stored-product Coleoptera in an airtight system: Implications for fumigant testing. Journal of Stored Products Research 1998, 34(4), 331–339. [Google Scholar] [CrossRef]
- Chelladurai, V.; Jayas, D.; White, N. Thermal imaging for detecting fungal infection in stored wheat. Journal of Stored Products Research 2010, 46(3), 174–179. [Google Scholar] [CrossRef]
- França-Silva, F.; Rego, C. H. Q.; Gomes-Junior, F. G.; Brancaglioni, V. A.; Hirai, W. Y.; Rodrigues, D. B.; Almeida, A. D. S.; Martins, A. B. N.; Tunes, L. V. M. D. Determination of Sitotroga cerealella infestation in wheat seeds by radiographic and multispectral images. Agronomy Journal 2020, 112(6), 3695–3703. [Google Scholar] [CrossRef]
- Ambrose, A.; Kandpal, L. M.; Kim, M. S.; Lee, W.-H.; Cho, B.-K. High speed measurement of corn seed viability using hyperspectral imaging. Infrared Physics & Technology 75 2016, 173–179. [Google Scholar] [CrossRef]
- Zhao, Y.; Zhang, C.; Zhu, S.; Gao, P.; Feng, L.; He, Y. Non-destructive and rapid variety discrimination and visualization of single grape seed using near-infrared hyperspectral imaging technique and multivariate analysis. Molecules 2018, 23(6), 1352. [Google Scholar] [CrossRef] [PubMed]
- Kandpal, L. M.; Lohumi, S.; Kim, M. S.; Kang, J.-S.; Cho, B.-K. Near-infrared hyperspectral imaging system coupled with multivariate methods to predict viability and vigor in muskmelon seeds. Sensors and Actuators B: Chemical 229 2016, 534–544. [Google Scholar] [CrossRef]
- Dale, L. M.; Thewis, A.; Boudry, C.; Rotar, I.; Dardenne, P.; Baeten, V.; Pierna, J. A. F. Hyperspectral imaging applications in agriculture and agro-food product quality and safety control: A review. Applied Spectroscopy Reviews 2013, 48(2), 142–159. [Google Scholar] [CrossRef]
- Chomontowski, C.; Podlaski, S. Impact of sugar beet seed priming using the SMP method on the properties of the pericarp. BMC Plant Biology 2020, 20(1), 1–9. [Google Scholar] [CrossRef]
- Salimi, Z.; Boelt, B. Classification of processing damage in sugar beet (Beta vulgaris) seeds by multispectral image analysis. Sensors 2019, 19(10), 2360. [Google Scholar] [CrossRef]
- Olesen, M. H.; Carstensen, J. M.; Boelt, B. Multispectral imaging as a potential tool for seed health testing of spinach (Spinacia oleracea L.). Seed Science and Technology 2011, 39(1), 140–150. [Google Scholar] [CrossRef]
- Vresak, M.; Olesen, M. H.; Gislum, R.; Bavec, F.; Jørgensen, J. R. The use of image-spectroscopy technology as a diagnostic method for seed health testing and variety identification. PLoS ONE 2016, 11(3), e0152011. [Google Scholar] [CrossRef]
- Jalink, H; van der Schoor, R; Birnbaum, Y E; Bino, R J. Seed chlorophyll content as an indicator for seed maturity and seed quality. Acta Hortic. 1999, 504, 219–228. [Google Scholar] [CrossRef]
- Tomazello-Filho, M.; Brazolin, S.; Chagas, M.P.; Oliveira, J.T.S.; Ballarin, A.W. Application of X-ray technique in nondestructive evaluation of eucalypt wood. Maderas. Ciencia y Tecnologia 2008, 10, 139–149. [Google Scholar] [CrossRef]
- Ahmed, M. R.; Yasmin, J.; Park, E.; Kim, G.; Kim, M. S.; Wakholi, C.; Mo, C.; Cho, B. K. Classification of watermelon seeds using morphological patterns of X-ray imaging: A comparison of conventional machine learning and deep learning. Sensors 2020, 20(23), 6753. [Google Scholar] [CrossRef] [PubMed]
- Huang, M.; Wang, Q. G.; Zhu, Q. B.; Qin, J. W.; Huang, G. Review of seed quality and safety tests using optical sensing technologies. Seed Science and Technology 2015, 43(3), 337–366. [Google Scholar] [CrossRef]
- Wu, Y.; Chen, B.; Zhao, X.; Kang, Y.; Ding, Y. Review of weed detection methods based on computer vision. Sensors 2021, 21(11), 3647. [Google Scholar] [CrossRef]
- Zhao, S.; Haque, R.; Wang, S. Automated seed identification with computer vision: Challenges and opportunities. Seed Science and Technology 2022, 50(1), 75–102. [Google Scholar] [CrossRef]
- Farokhzad, S.; Modaress Motlagh, A.; Ahmadi Moghadam, P.; Jalali Honarmand, S.; Kheiralipour, K. Application of infrared thermal imaging technique and discriminant analysis methods for non-destructive identification of fungal infection of potato tubers. Journal of Food Measurement and Characterization 2020, 14(1), 88–94. [Google Scholar] [CrossRef]
- Kranner, I.; Kastberger, G.; Hartbauer, M.; Pritchard, H. W. Noninvasive diagnosis of seed viability using infrared thermography. Proceedings of the National Academy of Sciences 2010, 107(8), 3912–3917. [Google Scholar] [CrossRef]
- Baranowski, P.; Mazurek, W.; Walczak, R. T. The use of thermography for pre-sowing evaluation of seed germination capacity. International Conference on Quality in Chains: An Integrated View on Fruit and Vegetable Quality (Acta Horticulturae, 2003; Vol. 604, pp. 459–465. [Google Scholar] [CrossRef]
- Chinnasamy, S.; Sundareswaran, K. S.; Subramaniyan, K.; Raja, P. R.; Renganayaki, S.; Marimuthu. Volatile organic compound analysis as advanced technology to detect seed quality in groundnut. Journal of Applied and Natural Science 2022, 14(3), 885–894. [Google Scholar] [CrossRef]
- Furizal, A.; Ma’arif, A.; Azzawagama Firdaus, W.; Rahmaniar. Future potential of E-nose technology: A review. International Journal of Robotics and Control Systems 2023, 3(3), 449–469. [Google Scholar] [CrossRef]
- Galletti, P. A.; Carvalho, M. E. A.; Hirai, W. Y.; Brancaglioni, V. A.; Arthur, V.; Barboza da Silva, C. Integrating optical imaging tools for rapid and non-invasive characterization of seed quality: Tomato (Solanum lycopersicum L.) and carrot (Daucus carota L.) as study cases. Frontiers in Plant Science 11 2020, 577851. [Google Scholar] [CrossRef]
- Konstantinova, P.; van den Bulk, R.; Jalink, H. Chlorophyll fluorescence sorting as a method for improvement of barley (Hordeum vulgare L.) seed health and germination. Seed Science and Technology 2002, 30(2), 519–529. [Google Scholar] [CrossRef]
- Xie, B.; Chen, B.; Ma, J.; Chen, J.; Zhou, Y.; Han, X.; Xiong, Z.; Yu, Z.; Huang, F. Rapid identification of Choy sum seeds infected with Penicillium decumbens based on hyperspectral imaging and stacking ensemble learning. Food Analytical Methods 2024, 17(2), 416–425. [Google Scholar] [CrossRef]
- Zhang, H.; Huang, L.; Huang, W.; Dong, Y.; Weng, S.; Zhao, J.; Ma, H.; Liu, L. Detection of wheat Fusarium head blight using UAV-based spectral and image feature fusion. Frontiers in Plant Science 13 2022, 1004427. [Google Scholar] [CrossRef] [PubMed]
- Agustiono, W.; Safitri, F. A.; Setiawan, W.; Chan, C. Transfer learning using MobileNet for rice seed image classification. Communications in Mathematical Biology and Neuroscience 2023, 2023, 132. [Google Scholar] [CrossRef]
- Wu, Q.; Xu, L.; Zou, Z.; Wang, J.; Zeng, Q.; Wang, Q.; Zhen, J.; Wang, Y.; Zhao, Y.; Zhou, M. Rapid nondestructive detection of peanut varieties and peanut mildew based on hyperspectral imaging and stacked machine learning models. Frontiers in Plant Science 13 2022, 1047479. [Google Scholar] [CrossRef]
- Chu, X.; Wang, W.; Ni, X.; Li, C.; Li, Y. Classifying maize kernels naturally infected by fungi using near-infrared hyperspectral imaging. Infrared Physics & Technology 2020, 105, 103242. [Google Scholar] [CrossRef]
- Kabala, D. M.; Hafiane, A.; Bobelin, L.; Canals, R. Image-based crop disease detection with federated learning. Scientific Reports 2023, 13(1). [Google Scholar] [CrossRef]
- Mehdipour, S.; Mirroshandel Mgomezulu, W. R.; Chitete, M.; Maonga, B. B.; Phiri, M. Machine learning approaches for grain seed quality assessment: a comparative study of maize seed samples in Malawi. Deleted Journal 2025, 7(6). [Google Scholar] [CrossRef]
- Mehreen, S.; Goëau, H.; Bonnet, P.; Chau, S.; Champ, J.; Joly, A. Estimating Compositions and Nutritional Values of Seed Mixes Based on Vision Transformers. Plant Phenomics 2023, 5. [Google Scholar] [CrossRef]
- Femenias Llaneras, A.; Llorens, E.; Ramos, A.; Marín, S. Near-infrared hyperspectral imaging evaluation of Fusarium damage and DON in single wheat kernels. Food Control 142 2022, 109239. [Google Scholar] [CrossRef]
- Xu, P.; Sun, W.; Xu, K.; Zhang, Y.; Tan, Q.; Qing, Y.; Yang, R. Identification of defective maize seeds using hyperspectral imaging combined with deep learning. Foods 2022, 12(1), 144. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zhai, M.; Zheng, L.; Zhou, L.; Xie, X.; Zhao, W.; Zhang, W. Efficient residual network using hyperspectral images for corn variety identification. Frontiers in Plant Science 15 2024, 1376915. [Google Scholar] [CrossRef]
- Jin, B.; Zhang, C.; Jia, L.; Tang, Q.; Gao, L.; Zhao, G.; Qi, H. Identification of rice seed varieties based on near-infrared hyperspectral imaging technology combined with deep learning. ACS Omega 2022, 7(1), 1021–1032. [Google Scholar] [CrossRef]
- Sudki, J. M.; Fonseca de Oliveira, G. R.; de Medeiros, A. D.; Mastrangelo, T.; Arthur, V.; Amaral da Silva, E. A.; Mastrangelo, C. B. Fungal identification in peanuts seeds through multispectral images: Technological advances to enhance sanitary quality. Frontiers in Plant Science 14 2023, 1112916. [Google Scholar] [CrossRef]
- De Silva, M.; Brown, D. Multispectral Plant Disease Detection with Vision Transformer-Convolutional Neural Network Hybrid Approaches. Sensors (Basel, Switzerland) 2023, 23(20), 8531. [Google Scholar] [CrossRef]
- Jung, D.-H.; Kim, J. D.; Kim, H.-Y.; Lee, T. S.; Kim, H. S.; Park, S. H. A hyperspectral data 3D convolutional neural network classification model for diagnosis of gray mold disease in strawberry leaves. Frontiers in Plant Science 13 2022, 837020. [Google Scholar] [CrossRef]
- Guo, X.; Feng, Q.; Guo, F. CMTNet: a hybrid CNN-transformer network for UAV-based hyperspectral crop classification in precision agriculture. Sci Rep 2025, 15, 12383. [Google Scholar] [CrossRef]
- Kabir, M. A.; Lee, I.; Lee, S.-H. Deep Learning-Based Detection of Aflatoxin B1 Contamination in Almonds Using Hyperspectral Imaging: A Focus on Optimized 3D Inception–ResNet Model. Toxins 2025, 17(4), 156. [Google Scholar] [CrossRef]
- Wu, N.; Liu, F.; Meng, F.; Li, M.; Zhang, C.; He, Y. Rapid and Accurate Varieties Classification of Different Crop Seeds Under Sample-Limited Condition Based on Hyperspectral Imaging and Deep Transfer Learning. Frontiers in bioengineering and biotechnology 2021, 9, 696292. [Google Scholar] [CrossRef]
- Han, Z.; Gao, J. Pixel-level aflatoxin detecting based on deep learning and hyperspectral imaging. Computers and Electronics in Agriculture 164 2019, 104888. [Google Scholar] [CrossRef]
- Li, G.; Ye, M. Hyperspectral image classification via transformer-based spectral-spatial attention decoupling and adaptive gating. arXiv 2025, arXiv:2506.08324. [Google Scholar] [CrossRef]
- de Medeiros, A. D.; da Silva, L. J.; Ribeiro, J. P. O.; Ferreira, K. C.; Rosas, J. T. F.; Santos, A. A.; da Silva, C. B. Machine Learning for Seed Quality Classification: An Advanced Approach Using Merger Data from FT-NIR Spectroscopy and X-ray Imaging. In Sensors; Multidisciplinary Digital Publishing Institute; 2020; Volume 20, Issue 15. [Google Scholar] [CrossRef]
- Qiao, X.; Jiang, J.; Qi, X.; Guo, H.; Yuan, D. Utilization of spectral-spatial characteristics in shortwave infrared hyperspectral images to classify and identify fungi-contaminated peanuts. Food Analytical Methods 2017, 10, 3503–3514. [Google Scholar] [CrossRef] [PubMed]
- Seo, Y.; Kim, G.; Lim, J.; Lee, A.; Kim, B.; Jang, J.; Mo, C.; Kim, M. S. Non-destructive detection software for cucumber green mottle mosaic virus in watermelon seeds using a near-infrared hyperspectral imaging system. Biosystems Engineering 2019, 181, 83–92. [Google Scholar]
- Yablokova, A.; Koвaлeв, И. B.; Kovalev, D.; Podoplelova, V.; Kobilova, A. Computer vision methods and algorithms for automatic detection and classification of objects in decision support systems in agriculture. E3S Web of Conferences 2024, 548, 3023. [Google Scholar] [CrossRef]
- Oliveira, M. M.; Cerqueira, B. V.; Barbon, S.; Barbin, D. F. Classification of fermented cocoa beans (cut test) using computer vision. Journal of Food Composition and Analysis 2020, 97, 103771. [Google Scholar] [CrossRef]
- Breiman, L. Random Forests. Machine Learning 2001, 45, 5–32.222. [Google Scholar] [CrossRef]
- Delwiche, S. R.; Rodriguez, I. T.; Rausch, S. R.; Graybosch, R. A. Estimating percentages of Fusarium-damaged kernels in hard wheat by near-infrared hyperspectral imaging Data version control. Journal of Cereal Science 2019, 87, 18–24. Available online: https://dvc.org. [CrossRef]
- Senthilkumar, T.; Jayas, D. S.; White, N. D. G.; Fields, P. G.; Gräfenhan, T. Detection of fungal infection in canola using near-infrared hyperspectral imaging. Journal of Stored Products Research 2015, 63, 80–88. [Google Scholar] [CrossRef]
- Williamson, H. F.; Brettschneider, J.; Cáccamo, M.; Davey, R.; Goble, C.; Kersey, P.; May, S.; Morris, R. J.; Ostler, R.; Pridmore, T.; Rawlings, C.; Studholme, D. J.; Tsaftaris, S. A.; Leonelli, S. Data management challenges for artificial intelligence in plant and agricultural research. F1000Research 2021, 10, 324. [Google Scholar] [CrossRef] [PubMed]
- Dong, J.; Yin, H.; Zheng, R.; Yoo, S. J.; Gu, Y. H. PlantInfoCMS: Scalable Plant Disease Information Collection and Management System for Training AI Models. Sensors 2023, 23(11), 5032. [Google Scholar] [CrossRef]
- Joshi, A.; Guevara, D.; Earles, M. Standardizing and Centralizing Datasets for Efficient Training of Agricultural Deep Learning Models. Plant Phenomics 2023, 5. [Google Scholar] [CrossRef] [PubMed]
- Waltz, L.; Katari, S.; Hong, C.; Anup, A.; Colbert, J.; Potlapally, A.; Dill, T. D.; Porter, C.; Engle, J. D.; Stewart, C.; Subramoni, H.; Shearer, S. A.; Machiraju, R.; Ortez, O. A.; Lindsey, L. E.; Nandi, A.; Khanal, S. Cyberinfrastructure for machine learning applications in agriculture: experiences, analysis, and vision. Frontiers in Artificial Intelligence 2025, 7. [Google Scholar] [CrossRef]
- Sykes, J. R.; Denby, K.; Franks, D. W. Computer vision for plant pathology: A review with examples from cocoa agriculture [Review of Computer vision for plant pathology: A review with examples from cocoa agriculture]. In Applications in Plant Sciences; Botanical Society of America, 2023; Volume 12, 2. [Google Scholar] [CrossRef]
- Geiser, D. M.; Martin, F. N.; Espíndola, A. S.; Brown, J. K.; Bell, T. H.; Yang, Y.; Kang, S. Knowledge Gaps, Research Needs, and Opportunities in Plant Disease Diagnostic Assay Development and Validation. PhytoFrontiersTM 2023, 3(1), 51. [Google Scholar] [CrossRef]
- Lahlali, R.; Gachara, G.; Özer, G.; Hussain, T. Editorial: Perspective challenges for applied research in potato pathogens: From molecular biology to bioinformatics. Frontiers in Microbiology 2023, 14. [Google Scholar] [CrossRef]
- Li, A.; Marković, M.; Edwards, P.; Leontidis, G. Model pruning enables localized and efficient federated learning for yield forecasting and data sharing. Expert Systems with Applications 2023, 242, 122847. [Google Scholar] [CrossRef]
- Ferreira, L. C.; Carvalho, I. C. B.; Jorge, L. A. C.; Quezado-Duval, A. M.; Rossato, M. Hyperspectral imaging for the detection of plant pathogens in seeds: Recent developments and challenges. Frontiers in Plant Science 2024, 15, 1387925. [Google Scholar] [CrossRef]
- Shoaib, M.; Shah, B.; Sayed, N.; Ali, F.; Ullah, R.; Hussain, I. Deep learning for plant bioinformatics: an explainable gradient-based approach for disease detection. Frontiers in Plant Science 2023, 14. [Google Scholar] [CrossRef] [PubMed]
- Nizamani, M. M.; Zhang, Q.; Muhae-Ud-Din, G.; Wang, Y. High-throughput sequencing in plant disease management: a comprehensive review of benefits, challenges, and future perspectives [Review of High-throughput sequencing in plant disease management: a comprehensive review of benefits, challenges, and future perspectives]. In Phytopathology Research; BioMed Central, 2023; Volume 5, 1. [Google Scholar] [CrossRef]
- Zhang, Z.; Ma, L.; Wei, C.; Yang, M.; Qin, S.; Lv, X.; Zhang, Z. Cotton Fusarium wilt diagnosis based on generative adversarial networks in small samples. Frontiers in Plant Science 14 2023, 1290774. [Google Scholar] [CrossRef]
- Alshammari, K.; et al. An improved pear disease classification approach using GAN-based image synthesisand deep learning. Scientific Reports 2024, 14, Article 5714. [Google Scholar] [CrossRef] [PubMed]
- Chawla, N. V.; Bowyer, K. W.; Hall, L. O.; Kegelmeyer, W. P. SMOTE: Synthetic minority over-sampling technique. Journal of Artificial Intelligence Research 2002, 16, 321–357. [Google Scholar] [CrossRef]
- Fernández, A.; García, S.; Galar, M.; Prati, R. C.; Krawczyk, B.; Herrera, F. Learning from imbalanced data sets. In Springer; 2018. [Google Scholar] [CrossRef]
- Sowjanya, A. M.; Rao, G. S.; Singamaneni, K. K. Effective treatment of imbalanced datasets in health care using modified SMOTE coupled with stacked deep learning algorithms. BMC Medical Informatics and Decision Making 2022, 22, 38. [Google Scholar] [CrossRef] [PubMed]
- Kawakura, S.; Hirafuji, M.; Ninomiya, S.; Shibasaki, R. Visual analysis of agricultural workers using explainable artificial intelligence (XAI) on class activation map (CAM) with characteristic point data output from OpenCV-based analysis. European Journal of Artificial Intelligence and Machine Learning 2023, 2(1), 1–8. [Google Scholar] [CrossRef]
- Khan, H. Advancements in Crop Analysis through Deep Learning and Explainable. AI 2025, 242. [Google Scholar] [CrossRef]
- Alhammad, S. M.; Khafaga, D. S.; Elhady, W. M.; Samy, F. M.; Hosny, K. M. Deep learning and explainable AI for classification of potato leaf diseases. Frontiers in Artificial Intelligence 2025, 7. [Google Scholar] [CrossRef]
- Agrawal, R.; Gupta, T.; Gupta, S.; Chauhan, S. S.; Patel, P.; Hamdare, S. Fostering trust and interpretability: integrating explainable AI (XAI) with machine learning for enhanced disease prediction and decision transparency. Diagnostic Pathology 2025, 20(1). [Google Scholar] [CrossRef] [PubMed]
- Vasileiou, M.; Kyrgiakos, L. S.; Kleisiari, C.; Kleftodimos, G.; Vlontzos, G.; Belhouchette, H.; Pardalos, P. M. Transforming weed management in sustainable agriculture with artificial intelligence: A systematic literature review towards weed identification and deep learning. Crop Protection 2023, 176, 106522. [Google Scholar] [CrossRef]
- Saito, T.; Rehmsmeier, M. The precision-recall plot is more informative than the ROC plot when evaluating binary classifiers on imbalanced datasets. PLoS ONE 2015, 10(3), e0118432. [Google Scholar] [CrossRef]
- Nehoshtan, Y.; Carmon, E.; Yaniv, O.; Sharon, A.; Rotem, O. Robust seed germination prediction using deep learning and RGB image data. Scientific Reports 2021, 11(1). [Google Scholar] [CrossRef] [PubMed]
- MacNeil, L.; Missan, S.; Luo, J.; Trappenberg, T.; LaRoche, J. Plankton classification with high-throughput submersible holographic microscopy and transfer learning. BMC Ecology and Evolution 2021, 21(1). [Google Scholar] [CrossRef] [PubMed]
- Matsuda, O.; Ohara, Y. Last-percent improvement in eligibility rates of crop seeds based on quality evaluation using near-infrared imaging spectrometry. PLoS ONE 2023, 18(9). [Google Scholar] [CrossRef]
- Ford, A. T.; Lan, J.; Ng, K. Detection of multiple cardiac abnormalities using Convolution, Positional Encoder and Transformer on 12-lead ECG recordings. In medRxiv; Cold Spring Harbor Laboratory, 2024. [Google Scholar] [CrossRef]
- Ahmed, A. M. A. Exploring MLOps Dynamics: An Experimental Analysis in a Real-World Machine Learning Project. In arXiv; Cornell University, 2023. [Google Scholar] [CrossRef]
- Hemalatha, S.; Jayachandran, J. J. B. A Multitask Learning-Based Vision Transformer for Plant Disease Localization and Classification. International Journal of Computational Intelligence Systems 2024, 17(1). [Google Scholar] [CrossRef]
- Folino, F.; Folino, G.; Pisani, F. S.; Pontieri, L.; Sabatino, P. Efficiently approaching vertical federated learning by combining data reduction and conditional computation techniques. Journal Of Big Data 2024, 11(1). [Google Scholar] [CrossRef]
- Hu, K.; Wang, Z.; Coleman, G.; Bender, A.; Yao, T.; Zeng, S.; Song, D.; Schumann, A. W.; Walsh, M. Deep learning techniques for in-crop weed recognition in large-scale grain production systems: a review [Review of Deep learning techniques for in-crop weed recognition in large-scale grain production systems: a review]. In Precision Agriculture; Springer Science+Business Media, 2023; Volume 25, 1, p. 1. [Google Scholar] [CrossRef]
- Batistatos, M. C.; de Cola, T.; Kourtis, M.; Apostolopoulou, V.; Xilouris, G.; Sagias, N. C. AGRARIAN: A Hybrid AI-Driven Architecture for Smart Agriculture 2025. [CrossRef]
- Pintus, M.; Colucci, F.; Maggio, F. Emerging Developments in Real-Time Edge AIoT for Agricultural Image Classification. IoT 2025, 6(1), 13. [Google Scholar] [CrossRef]
- Chelotti, J. O.; Martinez-Rau, L. S.; Ferrero, M.; Vignolo, L. D.; Galli, J. R.; Planisich, A. M.; Rufiner, H. L.; Giovanini, L. L. Livestock feeding behaviour: A review on automated systems for ruminant monitoring [Review of Livestock feeding behaviour: A review on automated systems for ruminant monitoring]. In Biosystems Engineering; Elsevier BV, 2024; Volume 246, p. 150. [Google Scholar] [CrossRef]
- Puppala, S.; Sinha, K. Towards Secure and Efficient Farming using Self-Regulating Heterogeneous Federated Learning in Dynamic Network Conditions. 2025. [Google Scholar] [CrossRef]
- Mugisha, S.; Kisitu, R.; Tushabe, F. Hybrid Knowledge Transfer through Attention and Logit Distillation for On-Device Vision Systems in Agricultural IoT. 2025. [Google Scholar] [CrossRef]
- Chorney, W.; Rahman, A.; Wang, Y.; Wang, H.; Peng, Z. Federated Learning for Heterogeneous Multi-Site Crop Disease Diagnosis 2025. [CrossRef]
- Heydari, S.; Mahmoud, Q. H. Tiny Machine Learning and On-Device Inference: A Survey of Applications, Challenges, and Future Directions. Sensors 2025, 25(10), 3191. [Google Scholar] [CrossRef]
- Singla, A. Machine Learning Operations (MLOps): Challenges and Strategies. Journal of Knowledge Learning and Science Technology ISSN 2959-6386 (Online) 2023, 2(3), 333. [Google Scholar] [CrossRef]
- Ahmed, S.; Revolinski, S.; Maughan, P. J.; Savic, M.; Kalin, J. E. R.; Burke, I. C. Deep learning–based detection and quantification of weed seed mixtures. Weed Science 2024, 1. [Google Scholar] [CrossRef]
- Knoblauch, D.; Großmann, J. Towards a Risk-Based Continuous Auditing-Based Certification for Machine Learning. The Review of Socionetwork Strategies 2023, 17(2), 255. [Google Scholar] [CrossRef]
- Garg, S.; Pundir, P.; Rathee, G.; Gupta, P. K.; Garg, S.; Ahlawat, S. On Continuous Integration/Continuous Delivery for Automated Deployment of Machine Learning Models using MLOps. 25 2021. [CrossRef]
- Bade, S. Demystifying Mlops: Bridging The Gap Between Machine Learning And Operations. International Journal Of Information Technology And Management Information Systems 2025, 16(2), 1376. [Google Scholar] [CrossRef]
- Moskalenko, V.; Kharchenko, V. Resilience-aware MLOps for AI-based medical diagnostic system. Frontiers in Public Health 2024, 12. [Google Scholar] [CrossRef]
- Cob-Parro, A. C.; Lalangui, Y.; Lazcano, R. Fostering Agricultural Transformation through AI: An Open-Source AI Architecture Exploiting the MLOps Paradigm. Agronomy 2024, 14(2), 259. [Google Scholar] [CrossRef]
- Lundstrom, C.; Lindvall, M. Mapping the Landscape of Care Providers’ Quality Assurance Approaches for AI in Diagnostic Imaging [Review of Mapping the Landscape of Care Providers’ Quality Assurance Approaches for AI in Diagnostic Imaging]. In Journal of Digital Imaging; Springer Science+Business Media, 2022; Volume 36, 2, p. 379. [Google Scholar] [CrossRef]
- Fedele, G.; Hill, G.; Sweetford, A.; Haines, J.; Smith, R. Development and validation of automated methods for PCR master-mix preparation in diagnostic laboratories. SLAS Technology 2024, 29(5), 100195. [Google Scholar] [CrossRef] [PubMed]
- Yin, H.; Zhang, X.; Zhang, Y.; Chen, X. Integrated microfluidic systems with sample preparation and nucleic acid amplification. Lab on a Chip 2019, 19(24), 4237–4259. [Google Scholar] [CrossRef]
- Obino, D.; Vassalli, M.; Franceschi, A.; Alessandrini, A.; Facci, P.; Viti, F. An overview on microfluidic systems for nucleic-acids extraction from human raw samples. Sensors 2021, 21(9), 3058. [Google Scholar] [CrossRef]
- Thai, D. A.; Liu, Y. Nucleic acid amplification tests in digital microfluidics: The promise of next-generation point-of-care diagnostics. Microsystems & Nanoengineering 11 2025, 155. [Google Scholar] [CrossRef]
- Sun, A.; Liu, X.; Vopařilová, P.; Zhou, S.; Chen, R. An integrated microfluidic platform for nucleic-acid testing. Microsystems & Nanoengineering 10 2024, 66. [Google Scholar] [CrossRef]
- Liao, Q.; Zhou, L.; Zhang, F.; Hu, X. A digital microfluidic chip with dual-mode thermal control for automated nucleic-acid testing. Analytica Chimica Acta 1279 2025, 348473. [Google Scholar] [CrossRef]
- Xiao, Y.; Zhou, M.; Liu, C.; Li, H.; Tang, X. Fully integrated and automated centrifugal microfluidic chip for point-of-care multiplexed molecular diagnostics. Biosensors and Bioelectronics 255 2024, 116240. [Google Scholar] [CrossRef]
- Yang, Z.; Zhao, L.; Han, J. Digital barcodes for high-throughput screening. In ACS Chemical & Biomedical Engineering.; 2024. [Google Scholar] [CrossRef]
- Crone, M. A.; Priestman, M.; Ciechonska, M.; Alcock, R.; Reeve, R.; Hughes, G. L.; Freeman, T. M. A simple, efficient, and scalable approach to pooled RT-qPCR testing for SARS-CoV-2 detection based on adaptive deconvolution algorithms. BMC Bioinformatics 22 2021, 515. [Google Scholar] [CrossRef]
- Daon, Y.; Huppert, A.; Obolski, U. DOPE: D-optimal pooling experimental design with application for SARS-CoV-2 screening. arXiv 2021, arXiv:2103.03706. [Google Scholar] [CrossRef] [PubMed]
- Emerson, J. B.; Adams, R. I.; Román, C. M. B.; Brooks, B.; Coil, D. A.; Dahlhausen, K.; Rothschild, L. J. Schrödinger’s microbes: Tools for distinguishing the living from the dead in microbial ecosystems. Microbiome 2017, 5(1), 86. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R.; Kumar Lal, M.; Prasad, P.; Tiwari, R. K. Editorial: Current advancements in real-time plant pathogen diagnostics: From lab assays to in-field detection. Frontiers in Plant Science 14 2023. [Google Scholar] [CrossRef]
- International Seed Health Initiative (ISHI); International Seed Federation (ISF). Best practices in PCR assays in seed health tests (Version 5); International Seed Federation: Nyon, Switzerland, 2025; Available online: 4_Best-Practices_PCR_v5.pdf. [Google Scholar]
- Newberry, E. A.; Srivastava, S.; Nunziata, S. O.; Mathew, R.; Mark, N.; Rívera, Y. Evaluation of metabarcoding methods for plant disease surveillance. PhytoFrontiers™ 3 2023, 785–794. [Google Scholar] [CrossRef]
- González-Rodríguez, V. E.; Izquierdo-Bueno, I.; Cantoral, J. M.; Carbú, M.; Garrido, C. Artificial intelligence: A promising tool for application in phytopathology. Horticulturae 10 2024, 197. [Google Scholar] [CrossRef]
- Martin, F. N.; Batuman, O.; Luster, D. G.; Miles, T. D.; Rivera, Y.; Sharma, P.; Geiser, D.; Cardwell, K. How controls improve diagnostic assay performance: Hitchhiker’s guide to diagnostic assay controls. PhytoFrontiers™ 5 2025, 127–137. [Google Scholar] [CrossRef]
- International Seed Health Initiative (ISHI); International Seed Health Initiative. Best practices for molecular techniques in seed health tests. Nyon, Switzerland, 2016. Available online: https://worldseed.org/wp-content/uploads/2016/01/ISHI-Veg_BestPractices_MolecularTechniques_2014.pdf.
- National Seed Health Systems (ISHI). Seed Health Testing Method. 2025. Available online: https://seedhealth.org/seed-health-testing-methods/.
- Galla, G.; Praeg, N.; Colla, F.; Rzehak, T.; Illmer, P.; Seeber, J.; Hauffe, H. C. Mock community as an in situ positive control for amplicon sequencing of microbiotas from the same ecosystem. Scientific Reports 13 2023, 4056. [Google Scholar] [CrossRef]
- Sharma, P.; Luster, D. G. State of the field of plant pathogen diagnostic assay development and validation. PhytoFrontiers™ 3 2023, 18–22. [Google Scholar] [CrossRef]
- Cardwell, K.; Dennis, G.; Flannery, A. R.; Fletcher, J.; Luster, D.; Nakhla, M.; Rice, A.; Shiel, P.; Stack, J.; Walsh, C.; et al. Principles of diagnostic assay validation for plant pathogens: A basic review of concepts. Plant Health Progress 19 2018, 272–278. [Google Scholar] [CrossRef]
- Hoorfar, J.; Malorny, B.; Abdulmawjood, A.; Cook, N.; Wagner, M.; Fach, P. Practical considerations in design of internal amplification controls for diagnostic PCR assays. Journal of Clinical Microbiology 2004, 42(5), 1863–1868. [Google Scholar] [CrossRef] [PubMed]
- Abdel-Latif, A.; Osman, G. Comparison of three genomic DNA extraction methods to obtain high DNA quality from maize. Plant Methods 2017, 13(1), 1. [Google Scholar] [CrossRef] [PubMed]
- Miftahushudur, T.; Sahin, H. M.; Grieve, B.; Yin, H. A survey of methods for addressing imbalance data problems in agriculture applications. Remote Sensing 17 2025, 454. [Google Scholar] [CrossRef]
- Jhaveri, M.; Chirputkar, A.; Ashok, P. The efficacy of artificial intelligence in making best marketing decisions. In Proceedings of the 2023 International Conference on Innovative Data Communication Technologies and Application (ICIDCA), 2023; pp. 225–229. [Google Scholar]
- Paulikienė, S.; Benesevičius, D.; Benesevičienė, K.; Ūksas, T. Review—Seed treatment: Importance, application, impact, and opportunities for increasing sustainability. Agronomy 15 2025, 1689. [Google Scholar] [CrossRef]
- Mulvaney, M. J.; Verhulst, N.; Herrera, J. M.; Mezzalama, M.; Govaerts, B. Improved wheat performance with seed treatments under dry sowing on permanent raised beds. Field Crops Research 164 2014, 189–198. [Google Scholar] [CrossRef]
- Munkvold, G. P. Seed pathology progress in academia and industry. Annual Review of Phytopathology 47 2009, 285–311. [Google Scholar] [CrossRef]
- Ferreira, L. da C.; Carvalho, I. C. B.; Jorge, L. A. de C.; Quezado-Duval, A. M.; Rossato, M. Hyperspectral imaging for the detection of plant pathogens in seeds: Recent developments and challenges. Frontiers in Plant Science 15 2024. [Google Scholar] [CrossRef]
- De Sousa, M. V.; Machado, J. D. C.; Simmons, H. E.; Munkvold, G. P. Real-time quantitative PCR assays for the rapid detection and quantification of Fusarium oxysporum f. sp. phaseoli in Phaseolus vulgaris (common bean) seeds. Plant Pathology 2015, 64(2), 478–488. [Google Scholar] [CrossRef]
- Das, P. K.; Ganguly, S. B.; Mandal, B. Mitigating polymerase chain reaction/amplicon contamination in a high-risk high-burden mycobacterial reference laboratory in a resource-limited setting. International Journal of Mycobacteriology 7 2018, 332–337. [Google Scholar] [CrossRef] [PubMed]
- Khalifa, M.; Albadawy, M. AI in diagnostic imaging: Revolutionising accuracy and efficiency. Computer Methods and Programs in Biomedicine Update 5 2024, 100146. [Google Scholar] [CrossRef]
- Barbedo, J. G. A. A review on the main challenges in automatic plant disease identification based on visible range images. Biosystems Engineering 144 2016, 52–60. [Google Scholar] [CrossRef]
- Arnal Barbedo, J. G. Plant disease identification from individual lesions and spots using deep learning. Biosystems Engineering 180 2019, 96–107. [Google Scholar] [CrossRef]
- Tan, S. C.; Yiap, B. C. DNA, RNA, and protein extraction: The past and the present. Journal of Biomedical Biotechnology 2009 2009, 574398. [Google Scholar] [CrossRef] [PubMed]
- Nansen, C.; Imtiaz, M. S.; Mesgaran, M. B.; Lee, H. Experimental data manipulations to assess performance of hyperspectral classification models of crop seeds and other objects. Plant Methods 18 2022, 74. [Google Scholar] [CrossRef] [PubMed]
- Hong, K.; Zhou, Y.; Han, H. The pipelines of deep learning-based plant image processing. Quantitative Plant Biology 6 2025, e23. [Google Scholar] [CrossRef]
- Grimault, V.; Laffont, J.-L.; Buimer, M.; Serandat, I. Organizing and analyzing results of the seed health proficiency tests, version 3.0; International Seed Testing Association (ISTA): Bassersdorf, Switzerland, 2021; Available online: https://www.seedtest.org/api/rm/647V48YB6FBPKVV/pt-p-03-organizingandanalysingshptsv3-0.pd.
- International Seed Federation (ISF). ISF viewpoint on indirect seed health tests; International Seed Federation: Athens, Greece, 2013; Available online: https://worldseed.org/wp-content/uploads/2015/10/Indirect_Seed_Health_Tests_2013.pdf.
- Hiddink, G.; Baldwin, T.; Guerra, L. de L.; Spiekerman, M.; Hogan, C. The use of direct and indirect methods in seed health testing. APSNET. 2 August 2018. Available online: https://www.apsnet.org.
- Baltes, N. J.; Gil-Humanes, J.; Cermak, T.; Atkins, P. A.; Voytas, D. F. DNA replicons for plant genome engineering. Plant Cell 2014, 26(1), 151–163. [Google Scholar] [CrossRef] [PubMed]
- Schaad, N. W.; Mortensen, C. N.; Li, J.; Feng, J.; Luo, L.; Mazzaglia, A.; Balestra, G. M. Technical challenges for specific, sensitive detection of seed-borne bacterial pathogens. In Global perspectives on the health of seeds and plant propagation material; Gullino, M. L., Munkvold, G., Eds.; Springer Netherlands: Dordrecht, 2014; pp. 59–66. ISBN 978-94-017-9389-6. [Google Scholar]
- Al Rwahnih, M.; Klaassen, V.; Erickson, T.; Alabi, O. J.; Stevens, K.; Hwang, M. S.; Port, L. A new era in federal quarantine and state certification diagnostics at clean plant centers in the United States. Plant Disease 2025, 109(6), 1392–1403. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhang, F.; Li, X.; Baller, A. A.; Qi, Y.; Starker, C. G. Transcription activator-like effector nucleases enable efficient plant genome engineering. Plant Physiology 2013, 161(1), 20–27. [Google Scholar] [CrossRef]
- Chai, A. L.; Ben, H. Y.; Guo, W. T.; Shi, Y. X.; Xie, X. W.; Li, L.; Li, B. J. Quantification of viable cells of Pseudomonas syringae pv. tomato in tomato seed using propidium monoazide and a real-time PCR assay. Plant Disease 2020, 104(8), 2225–2232. [Google Scholar] [CrossRef] [PubMed]
- Islam, M.; Azad, A.; Arman, S. E.; Alyami, S. A.; Hasan, M. M. PlantCareNet: An advanced system to recognize plant diseases with dual-mode recommendations for prevention. Plant Methods 21 2025, 52. [Google Scholar] [CrossRef]
- Sujatha, R.; Krishnan, S.; Chatterjee, J. M.; Gandomi, A. H. Advancing plant leaf disease detection integrating machine learning and deep learning. Scientific Reports 15 2025, 11552. [Google Scholar] [CrossRef]
- Munkvold, G.; du Toit, L.; Dunkle, R. Seed pathology: Challenges and advances in ensuring a safe global seed supply. Annual Review of Phytopathology 63 2025, 43–62. [Google Scholar] [CrossRef]
- Choudhury, R. A.; Garrett, K. A.; Klosterman, S. J.; Subbarao, K. V.; McRoberts, N. A framework for optimizing phytosanitary thresholds in seed systems. Phytopathology® 2017, 107(10), 1219–1228. [Google Scholar] [CrossRef]
- U.S. Department of Agriculture; Animal and Plant Health Inspection Service (APHIS). ReFreSH: Regulatory framework for seed health accreditation standard. U.S. Department of Agriculture, Animal and Plant Health Inspection Service: Washington, DC, 2023. Available online: https://www.seedtest.org/api/rm/647V48YB6FBPKVV/pt-p-03-organizingandanalysingshptsv3-0.pd.
- Freese, T.; Elzinga, N.; Heinemann, M.; Lerch, M. M.; Feringa, B. L. The relevance of sustainable laboratory practices. RSC Sustainability 2 2024, 1300–1336. [Google Scholar] [CrossRef]
- Rai, S.; Sriram, N.; Alva, P.; Ashraf, A. A.; Kumar, S.; Nayak, S. Advancing green laboratory practices: A review of sustainability in healthcare. International Journal of Medical Biochemistry 2024, 7(3), 201–207. [Google Scholar] [CrossRef]
- Durgan, J.; Rodríguez-Martínez, M.; Rouse, B. Green labs: A guide to developing sustainable science in your organization. Immunology & Cell Biology 2023, 101(4), 289–301. [Google Scholar] [CrossRef]
- Massart, S.; Adams, I.; Al Rwahnih, M.; Baeyen, S.; Bilodeau, G. J.; Blouin, A. G.; Boonham, N.; Candresse, T.; Chandellier, A.; De Jonghe, K.; Fox, A.; Gaafar, Y. Z. A.; Gentit, P.; Haegeman, A.; Ho, W.; Hurtado-Gonzales, O.; Jonkers, W.; Kreuze, J.; Kutjnak, D.; Landa, B. B.; Liu, M.; Maclot, F.; Malapi-Wight, M.; Maree, H. J.; Martoni, F.; Mehle, N.; Minafra, A.; Mollov, D.; Moreira, A. G.; Nakhla, M.; Petter, F.; Piper, A. M.; Ponchart, J. P.; Rae, R.; Remenant, B.; Rivera, Y.; Rodoni, B.; Botermans, M.; Roenhorst, J. W.; Rollin, J.; Saldarelli, P.; Santala, J.; Souza-Richards, R.; Spadaro, D.; Studholme, D. J.; Sultmanis, S.; van der Vlugt, R.; Tamisier, L.; Trontin, C.; Vazquez-Iglesias, I.; Vicente, C. S. L.; van de Vossenberg, B. T. L. H.; Westenberg, M.; Wetzel, T.; Ziebell, H.; Lebas, B. S. M. Guidelines for the reliable use of high throughput sequencing technologies to detect plant pathogens and pests. Peer Community Journal 2 2022, e62. [Google Scholar] [CrossRef]
- Gauthier, N. P. G.; Chorlton, S. D.; Krajden, M.; Manges, A. R. Agnostic sequencing for detection of viral pathogens. Clinical Microbiology Reviews 36 2023, e0011922. [Google Scholar] [CrossRef] [PubMed]
- Détain, A.; Bhowmik, P.; Leborgne-Castel, N.; Ochatt, S. Latest biotechnology tools and targets for improving abiotic stress tolerance in protein legumes. Environmental and Experimental Botany 2022, 104824. [Google Scholar] [CrossRef]
- Zhu, Y.; Yu, X.; Wu, J. CRISPR/Cas: A toolkit for plant disease diagnostics. Trends in Plant Science 2025, 30(3), 245–248. [Google Scholar] [CrossRef] [PubMed]
- Cermak, T.; Doyle, E. L.; Christian, M.; Wang, L.; Zhang, Y.; Schmidt, C. Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Research 39 2011, e82. [Google Scholar] [CrossRef]
- Mahlein, A.-K. Plant disease detection by imaging sensors – Parallels and specific demands for precision agriculture and plant phenotyping. Plant Disease 2016, 100(2), 241–251. [Google Scholar] [CrossRef]
- Ang, M. C.-Y.; Lew, T. T. S. Non-destructive technologies for plant health diagnosis. Frontiers in Plant Science 13 2022, 884454. [Google Scholar] [CrossRef]
- Singh, C.; Randhawa, G. S.; Farooque, A. A.; Gill, Y. S.; Fraser, A.; Kumar, L. K. M.; Barrett, R.; Al-Mughrabi, K. AgriScout: AI-powered robot for precise detection of PVY-infected potato plants. Computers and Electronics in Agriculture 238 2025, 110781. [Google Scholar] [CrossRef]
- Zarbakhsh, S.; Fakhrzad, F.; Rajkovic, D.; Piekutowska, M.; Niedbała, G. Approaches and challenges in machine learning for monitoring agricultural products and predicting plant physiological responses to biotic and abiotic stresses. Current Plant Biology 2025, 100535. [Google Scholar] [CrossRef]
- Davino, S.; Caruso, A. G.; Bertacca, S.; Barone, S.; Panno, S. Tomato brown rugose fruit virus: Seed transmission rate and efficacy of different seed disinfection treatments. Plants 2020, 9(11), 1615. [Google Scholar] [CrossRef]
- Salem, N. M.; Sulaiman, A. M.; Samarah, N.; Turina, M. Localization and mechanical transmission of Tomato brown rugose fruit virus in tomato seeds. Plant Disease 2022, 106(1), 122–130. [Google Scholar] [CrossRef]
- Berg, T.; Tesoriero, L.; Hailstones, D. L. A multiplex real-time PCR assay for detection of Xanthomonas campestris from brassicas. Letters in Applied Microbiology 2006, 42(6), 624–630. [Google Scholar] [CrossRef]
- Van der Wolf, J. M.; van Beek, T. A.; van den Bulk, R. W.; van Doorn, J. Colonization of siliques and seeds of Brassica oleracea by Xanthomonas campestris pv. campestris after spray-inoculation of flower clusters. European Journal of Plant Pathology 154 2019, 445–461. [Google Scholar] [CrossRef]
- Sanna, M.; Gilardi, G.; Gullino, M. L.; Mezzalama, M. Evaluation of physical and chemical disinfection methods of Brassica oleracea seeds naturally contaminated with Xanthomonas campestris pv. campestris. Journal of Plant Diseases and Protection 129 2022, 1145–1152. [Google Scholar] [CrossRef]
- Chen, L.; Li, L.; Xie, X.; Chai, A.; Shi, Y.; Fan, T.; Xie, J.; Li, B. An improved method for quantification of viable Fusarium cells in infected soil products by propidium monoazide coupled with real-time PCR. Microorganisms 2022, 10(5), 1037. [Google Scholar] [CrossRef]
- Nicolaisen, M.; Supronienė, S.; Kærgaard Nielsen, L.; Lazzaro, I.; Spliid, N. H.; Justesen, A. F. Real-time PCR for quantification of eleven individual Fusarium species in cereals. Journal of Microbiological Methods 2009, 76(3), 234–240. [Google Scholar] [CrossRef]
- Latvala, S.; Haapalainen, M.; Kivijärvi, P.; Suojala-Ahlfors, T.; Iivonen, S.; Hannukkala, A. Sampling and PCR method for detecting pathogenic Fusarium oxysporum strains in onion harvest. Letters in Applied Microbiology 2020, 70(3), 210–220. [Google Scholar] [CrossRef] [PubMed]
- Wilkinson, M. D.; Dumontier, M.; Aalbersberg, I. J.; Appleton, G.; Axton, M.; Baak, A.; Blomberg, N.; Boiten, J.-W.; Bonino da Silva Santos, L.; Bourne, P. E.; Bouwman, J.; Brookes, A. J.; Clark, T.; Crosas, M.; Dillo, I.; Dumon, O.; Edmunds, S.; Evelo, C. T.; Finkers, R.; Gonzalez-Beltran, A.; Gray, A. J. G.; Groth, P.; Goble, C.; Grethe, J. S.; Heringa, J.; t Hoen, P. A. C.; Hooft, R.; Kuhn, T.; Kok, R.; Kok, J.; Lusher, S. J.; Martone, M. E.; Mons, A.; Packer, A. L.; Persson, B.; Rocca-Serra, P.; Roos, M.; van Schaik, R.; Sansone, S.-A.; Schultes, E.; Sengstag, T.; Slater, T.; Strawn, G.; Swertz, M. A.; Thompson, M.; van der Lei, J.; van Mulligen, E.; Velterop, J.; Waagmeester, A.; Wittenburg, P.; Wolstencroft, K.; Zhao, J.; Mons, B. The FAIR guiding principles for scientific data management and stewardship. Scientific Data 3 2016, 160018. [Google Scholar] [CrossRef] [PubMed]
- Mons, B.; Neylon, C.; Velterop, J.; Dumontier, M.; Bonino da Silva Santos, L. O.; Wilkinson, M. D. Cloudy, increasingly FAIR; revisiting the FAIR Data guiding principles for the European Open Science Cloud. Information Services & Use 2017, 37(1), 49–56. [Google Scholar] [CrossRef]
- Qi, X.; Zhang, C.; Zhu, J.; Liu, C.; Huang, C.; Li, X.; Xie, C. Genome editing enables next-generation hybrid seed production technology. Molecular Plant 2020, 13(1), 1–8. [Google Scholar] [CrossRef]
- OECD. OECD Seed Schemes: Rules and Directions for the International Certification of Seed Moving in International Trade; OECD Publishing: Paris, 2025; Available online: https://www.oecd.org/content/dam/oecd/en/topics/policy-sub-issues/seeds/rules-and-regulation-eng.pdf.
- FAO. ePhyto Solution: Electronic Phytosanitary Certification. Food and Agriculture Organization of the United Nations. 2023. Available online: https://www.ippc.int.
- Papoutsoglou, E. A.; Faria, D.; Arend, D.; Arnaud, E.; Athanasiadis, I. N.; Chaves, I.; Coppens, F.; Cornut, G.; Costa, B. V.; Ćwiek-Kupczyńska, H.; Droesbeke, B.; Finkers, R.; Gruden, K.; Junker, A.; King, G. J.; Krajewski, P.; Lange, M.; Laporte, M.-A.; Michotey, C.; Oppermann, M.; Ostler, R.; Poorter, H.; Ramírez-Gonzalez, R.; Ramšak, Ž.; Reif, J. C.; Rocca-Serra, P.; Sansone, S.-A.; Scholz, U.; Tardieu, F.; Uauy, C.; Usadel, B.; Visser, R. G. F.; Weise, S.; Kersey, P. J.; Miguel, C. M.; Adam-Blondon, A.-F. Enabling reusability of plant phenomic datasets with MIAPPE 1.1. New Phytologist 2020, 227(1), 260–273. [Google Scholar] [CrossRef]
- Wieczorek, J.; Bloom, D.; Guralnick, R.; Blum, S.; Döring, M.; Robertson, T.; Vieglais, D. Darwin Core: An evolving community-developed biodiversity data standard. PLoS ONE 2012, 7(1), e29715. [Google Scholar] [CrossRef]
- ISO. International Organization for Standardization. 2017. Available online: https://www.iso.org/standard/66912.html.
- Hughes, L. D.; Tsueng, G.; DiGiovanna, J.; Horvath, T. D.; Rasmussen, L. V.; Savidge, T. C.; Stoeger, T.; Turkarslan, S.; Wu, Q.; Wu, C.; Su, A. I.; Pache, L.; NIAID Systems Biology Data Dissemination Working Group. Addressing barriers in FAIR data practices for biomedical data. Scientific data 2023, 10(1), 98. [Google Scholar] [CrossRef]
- Raimundo, R.; Rosário, A. Blockchain System in the Higher Education. European Journal of Investigation in Health, Psychology and Education 2021, 11(1), 276–293. [Google Scholar] [CrossRef] [PubMed]
- European Commission. TRACES NT: Trade Control and Expert System – Next Generation; Brussels, Belgium, 2022; Available online: https://food.ec.europa.eu/horizontal-topics/traces_en.
- USDA. Phytosanitary Certificate Issuance and Tracking System (PCIT). United States Department of Agriculture – Animal and Plant Health Inspection Service. 2023. Available online: https://pcit.aphis.usda.gov/pcit/faces/signIn.jsf.
- Casino, F.; Dasaklis, T. K.; Patsakis, C. A systematic literature review of blockchain-based applications: Current status, classification and open issues. Telematics and Informatics 36 2019, 55–81. [Google Scholar] [CrossRef]
- Tripoli, M.; Schmidhuber, J. Emerging Opportunities for the Application of Blockchain in the Agri-food Industry. Revised version. In Rome and Geneva, FAO and ICTSD; 2020; Available online: https://openknowledge.fao.org/handle/20.500.14283/ca9934en.
- Oforleta, C. Reassessing trade secret protections in the era of AI: A comparative perspective on legal and ethical challenges. SSRN Electronic Journal 2025. [Google Scholar] [CrossRef]
- Barbedo, J. G. A. A review on the main challenges in automatic plant disease identification based on visible range images. Biosystems Engineering 144 2016, 52–60. [Google Scholar] [CrossRef]
- Gebru, T.; Morgenstern, J.; Vecchione, B.; Vaughan, J. W.; Wallach, H.; Daumé, H., III; Crawford, K. Datasheets for datasets. Communications of the ACM 2021, 64(12), 86–92. [Google Scholar] [CrossRef]
- OECD. Tools for Trustworthy AI Retrieved. 2021. Available online: https://www.oecd.org/content/dam/oecd/en/publications/reports/2021/06/tools-for-trustworthy-ai_0e36bb08/008232ec-en.pdf.
- Research ethics and integrity in the artificial intelligence era. In Frontiers in Research Metrics and Analytics; Phiri Chigwada, J., Ngulube, P., Eds.; 2023; Available online: https://www.frontiersin.org/research-topics/62732/research-ethics-and-integrity-in-the-artificial-intelligence-era.
- Lamprecht, A.-L.; Garcia, L.; Kuzak, M.; Martinez, C.; Arcila, R.; Del Pico, E. Towards FAIR principles for research software. Data Science 2021, 4(1), 17–33. [Google Scholar] [CrossRef]
- Brougher, J. T. Intellectual property and health technologies: Balancing innovation and the public’s health; Springer, 2014. [Google Scholar] [CrossRef]
- Ensuring innovation in diagnostics for bacterial infection: Implications for policy. In Observatory Studies Series; Morel, C., McClure, L., Edwards, S., et al., Eds.; European Observatory on Health Systems and Policies: Copenhagen, Denmark, 2016; Volume No. 44, Available online: https://www.ncbi.nlm.nih.gov/books/NBK447306/.
- Cohen-Sasson, O.; Tur-Sinai, O. Facilitating open science without sacrificing IP rights: A novel tool for improving replicability of published research: A novel tool for improving replicability of published research. EMBO reports 2022, 23(9), e55841. [Google Scholar] [CrossRef]
- EPPO. PM 7/98 (5): Specific requirements for laboratories preparing plant pest diagnostic protocols. EPPO Bulletin 2023, 52(2), 198–215. [Google Scholar] [CrossRef]
- Haibe-Kains, B.; Adam, G. A.; Hosny, A.; Khodakarami, F.; Waldron, L.; Wang, B.; McIntosh, C.; Goldenberg, A.; Kundaje, A.; Greene, C. S.; Broderick, T.; Hoffman, M. M.; Leek, J. T.; Korthauer, K.; Huber, W.; Brazma, A.; Pineau, J.; Tibshirani, R.; Hastie, T.; Aerts, H. J. W. L.; Massive Analysis Quality Control (MAQC) Society Board of Directors. Transparency and reproducibility in artificial intelligence. Nature 2020, 586(7829), E14–E16. [Google Scholar] [CrossRef]
- European Commission. Artificial Intelligence Act – Regulation of the European Parliament and Council on AI Systems; EU Publications: Brussels, 2024; Available online: https://eur-lex.europa.eu/legal-content/EN/TXT/PDF/?uri=OJ:L_202401689.
- FAO. International Plant Protection Convention: Capacity Development and Global Plant Health Information System. Food and Agriculture Organization of the United Nations. 2023. Available online: https://www.ippc.int.
- National Research Council (US) Committee on a New Government-University Partnership for Science and Security. Science and Security in a Post 9/11 World: A Report Based on Regional Discussions Between the Science and Security Communities IV, Biosecurity and Dual-Use Research in the Life Sciences. National Academies Press (US): Washington (DC), 2007. Available online: https://www.ncbi.nlm.nih.gov/books/NBK11496/.
- Dubov, A. The concept of governance in dual-use research. Medicine, health care, and philosophy 2014, 17(3), 447–457. [Google Scholar] [CrossRef] [PubMed]
- Carroll, S. R.; Garba, I.; Figueroa-Rodríguez, O. L.; Holbrook, J.; Lovett, R.; Materechera, S.; Hudson, M. The CARE principles for Indigenous data governance. Data Science Journal 2020, 19(43), 1–12. [Google Scholar] [CrossRef]
- ISTA. International Rules for Seed Testing.; International Seed Testing Association: Bassersdorf, 2023. [Google Scholar]
- ISTA. Accreditation and Proficiency Testing Program Manual.; International Seed Testing Association: Bassersdorf, 2021. [Google Scholar]
- Gitaitis, R.; Walcott, R. Seedborne bacterial pathogens in vegetables. Annual Review of Phytopathology 2012, 50, 223–246. [Google Scholar] [CrossRef]
- Eads, A.; Groth-Helms, D.; Davenport, B.; Cha, X.; Li, R.; Walsh, C.; Schuetz, K. The commercial validation of three tomato brown rugose fruit virus assays. PhytoFrontiers™ 2023, 3(3), 233–240. [Google Scholar] [CrossRef]
- Li, Y.; Tan, G.; Lan, P.; Zhang, A.; Liu, Y.; Li, R.; Li, F. Detection of tobamoviruses by RT-PCR using a novel pair of degenerate primers. Journal of virological methods 2018, 259, 122–128. [Google Scholar] [CrossRef] [PubMed]
- Clark, M. F.; Adams, A. N. Characteristics of the microplate method of ELISA. Journal of General Virology 1977, 34, 475–483. [Google Scholar] [CrossRef]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |