1. Introduction
Diet strongly influences the human gastrointestinal microbiome, shaping both community composition and metabolic function. Among dietary factors, fibre is particularly important. Fermentable fibres are metabolised by gut microbes to short-chain fatty acids such as acetate, propionate, and butyrate, which support epithelial energy supply, reinforce intestinal barrier integrity, and modulate inflammatory pathways (David et al., 2022; Hou et al., 2021). These metabolites are central to gut health and exemplify the bidirectional communication between host physiology and microbial ecology.
Despite broad consensus on the benefits of dietary fibre, translating microbiome science into real-world interventions remains challenging. Small participant numbers, high inter-individual variability, and the complex multivariate nature of microbiome data often limit statistical power and interpretability in human nutrition studies (Hughes et al., 2019). Baseline diet quality and habitual fibre intake are increasingly recognised as important determinants of microbial responsiveness (David et al., 2022; Hughes et al., 2019).
At the taxonomic level, fibre interventions often increase the relative abundance of butyrate-producing taxa, including members of the Lachnospiraceae such as Agathobacter, while reducing opportunistic or inflammation-linked genera such as Escherichia-Shigella (Karim et al., 2024). These microbial responses align with improvements in diet quality and ecological resilience. Recent pilot work conducted at CSIRO and La Trobe University (Melbourne, VIC, Australia) demonstrated that consumption of a fibre-enriched complex carbohydrate delivery vehicle induced measurable modulation of both oral and gastrointestinal microbiota in an autistic cohort, including reductions in opportunistic taxa and increases in beneficial fermenters (Shindler et al., submitted).
Building on this foundation, the present study introduces a Hybrid Intelligence Framework (HIF), a structured human-AI collaborative workflow designed to integrate domain expertise with generative-AI reasoning. The framework enables exploratory interpretation of complex, small-sample datasets in which traditional statistical tools alone may be insufficient. To demonstrate its potential, we applied HIF to the same two-week intervention (Shindler et al., submitted) in which twelve participants consumed a fibre-enriched food daily while providing oral and faecal microbiome samples and daily enjoyment scores. The HIF analysis was complemented by an independent statistical evaluation by the La Trobe University microbiome team using established tools such as ANCOM-BC, PERMANOVA, DESeq2, and MaAsLin2.
Together, these analyses illustrate how human-AI collaboration can enhance microbiome interpretation by combining transparent computational reasoning with disciplinary expertise. This work represents the first application of a structured HIF in food and sensory science, highlighting its promise for complex, multivariate biological research.
2. Methods
This section outlines the study design, data generation procedures, the independent subject-matter-expert (SME) statistical pipeline, and the HIF used for exploratory analysis.
2.1. Study Design and Participants
This pilot intervention enrolled twelve autistic adults aged 18 to 59 years, who consumed a fibre-enriched savoury nugget portion (6 nuggets) once daily for two weeks. Microbiome samples were collected from oral and faecal sites at three timepoints: baseline, mid-intervention, and end-intervention.
At study onset, participants completed a baseline dietary intake questionnaire covering specified fibre-relevant food groups and reported consumption frequency. No dietary grouping was defined at this stage. For the Hybrid Intelligence analysis, these raw responses were later combined into a continuous fibre-intake score for each participant, which was used to derive a two-bin split (low versus high fibre intake) for exploratory comparison. Participants also recorded daily enjoyment of the product during the intervention on a five-point scale (1 = not at all enjoyable; 5 = very enjoyable). For the independent SME analysis, dietary grouping was conducted separately using a Healthy Diet Index (HDI).
Eligible participants were required to have a history of, or current, feeding challenges characterised by selective eating patterns, restricted dietary preferences, or difficulty achieving adequate nutritional intake. Proficiency in reading and understanding English was required to ensure that participants could follow written study materials, including consent forms and food preparation instructions.
Exclusion criteria were established to minimise risk and ensure participant safety. Individuals with a documented history of severe allergic reactions to any ingredient in the study product were excluded. The test product contained soy and chicken protein; therefore, participants with known allergies to these ingredients were deemed ineligible. Individuals with medical conditions that could impair safe swallowing were also excluded. In addition, participants with impaired taste or smell perception were excluded, as such sensory disruptions could interfere with product consumption or adherence to the intervention protocol. Pregnant or breastfeeding individuals were excluded due to potential concerns regarding product safety during pregnancy and lactation.
The trial was registered at the Australian New Zealand Clinical Trials Registry (ANZCTR) (ACTRN12624000361505) and reviewed by the Human Research Ethics Committee of La Trobe University (HEC23487) and by the CSIRO Health and Medical Human Research Ethics Committee (2024_014_RR). All participants provided written informed consent before the start of the trial.
2.2. Fibre-Enriched Food Product
The intervention used a fibre-enriched savoury nugget developed by CSIRO as a food-based vehicle for increasing daily fibre intake. Participants received the products frozen and were instructed to consume one standard serving of six nuggets per day for two weeks. More information about the product is provided in the companion paper (Shindler et al., submitted).
2.3. Data Generation and SME Pipeline
Oral and faecal microbiome profiling and the primary statistical analyses were conducted by the La Trobe University microbiome team using an established procedure covering community-level comparisons and differential-abundance modelling. Full wet-lab procedures, sequencing workflows, bioinformatics steps, and statistical model specifications are provided in the companion paper (Shindler et al., submitted) and were applied here without substantive modification. This SME methodology served as the independent reference analysis against which the Hybrid Intelligence workflow was compared.
2.4. Problem Framing
The analytical goal was to interpret a small, multivariate dataset linking a fibre-enriched food intervention with microbiome responsiveness and sensory behaviour. Four guiding questions structured the hybrid intelligence approach:
- (i)
Which participants showed the largest microbiome shifts across the three timepoints?
- (ii)
To what extent did baseline diet quality or fibre intake relate to responsiveness?
- (iii)
Which taxa most plausibly contributed to observed changes?
- (iv)
Was daily enjoyment associated with patterns of microbial change?
Given the limited sample size and short study duration, the HIF workflow prioritised direction of effect, internal consistency across timepoints, and coherence with the independent SME statistical analysis rather than significance testing alone. The aim was to surface interpretable patterns while avoiding over-interpretation of a constrained dataset.
2.5. Data Structuring
The dataset was organised to support transparent HIF analysis while remaining compatible with the independent SME statistical dataset. All inputs were harmonised into a tidy, table-oriented scheme keyed by participant ID, sample site (oral or faecal), and timepoint (baseline, mid-intervention, end-intervention).
2.5.1. Inputs
- (i)
Genus-level relative abundances and diversity summaries exported directly from the SME dataset, retaining the same filtering, taxonomic annotations, and sample identifiers.
- (ii)
Behaviour: Daily enjoyment ratings (five-point scale) were averaged within each participant to provide a behavioural indicator. One low-adherence participant (AFIC011) was retained for completeness and noted during sensitivity checks. Daily gastrointestinal symptom logs were collected alongside enjoyment ratings but were not part of the primary HIF analysis and are not reported further.
- (iii)
Diet context: Raw baseline dietary questionnaire responses for fibre-relevant food groups and consumption frequencies, along with the HDI groupings generated independently in the SME analysis.
- (iv)
All samples followed the AFIC### convention and carried explicit timepoint and site labels (Pre, Mid, Final; Oral, Faecal), enabling complete longitudinal tracking for each participant.
2.5.2. Harmonisation
All datasets were brought into a single participant–timepoint table to ensure consistency between the HIF inputs and the SME statistical dataset. Identifiers, timepoints, and site labels were checked for alignment, and column names were standardised across sources. Enjoyment ratings were averaged per participant, and no additional feature engineering was applied. To stabilise exploratory summaries of microbial change, very low-prevalence taxa were removed before calculating directional fold changes.
2.5.3. Derived Variables (HIF-Specific)
Baseline dietary questionnaire responses were combined into a simple continuous fibre-intake score representing habitual consumption of fibre-rich foods. The score was calculated by summing frequencies for five food groups (legumes, whole-grain cereal, salad, fruit, and cereal), each rated on a one-to-five scale, giving a total range of five to twenty-five. This provided an approximate index of baseline dietary fibre exposure.
For exploratory comparison, participants were grouped into low-fibre (≤20) or high-fibre (>20) categories. This grouping is independent of the Healthy Diet Index categories used in the SME analysis. Additional variables included the average daily enjoyment score. The HDI category was not part of the dataset used for the HIF workflow and was introduced only through the SME analysis; it is therefore used in this manuscript solely for cross-comparison with SME findings, not as an input to the HIF analysis.
2.5.4. Quality Checks
Numeric fields were checked for sensible ranges and categorical fields were verified against expected labels. Missing values were left as missing; no imputation was applied. All merges and derived variables were reviewed to ensure that the HIF inputs matched the underlying raw data and the SME dataset structure.
2.6. Hybrid Exploration (Conversation-Centred HIF)
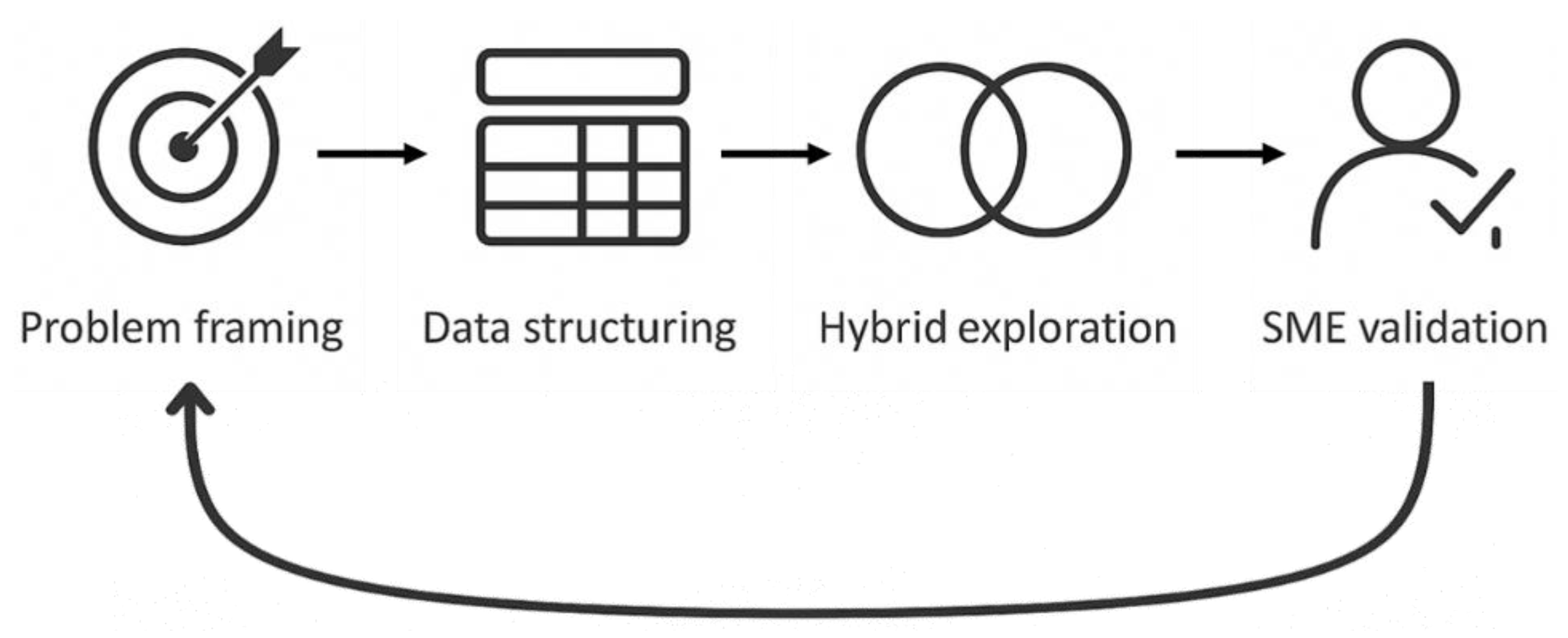
HIF treats analysis as an iterative human-AI dialogue, with an emphasis on transparency, explicit validation and refinement. Related principles for effective human–AI collaboration and traceability have been discussed previously (Amershi et al., 2019; Vössing et al., 2022), but here they are operationalised for small-sample, multivariate food-microbiome studies. The AI collaborator surfaces potential patterns, inconsistencies, and alternative explanations, and these are accepted, redirected, or rejected based on biological plausibility and alignment with the independent SME assessment. The approach follows four core principles: human-in-the-loop decision control, transparency, explicit validation, and iterative refinement (
Figure 1).
To operationalise these principles for this small-sample, multivariate microbiome dataset, analysis proceeded through a structured sequence of conversational passes aligned with problem framing, data structuring, hybrid exploration, SME validation, and iteration (
Figure 2). Each pass focused on a narrow, well-defined question, and outputs were logged to maintain traceability.
Although illustrated as a loop, the workflow is inherently non-linear; any stage can lead forward or backward depending on the outcome of the human-AI dialogue, including returning to problem framing if required.
2.6.1. Session Setup
Inputs: harmonised tables from
Section 2.5 (microbiome relative abundances, derived diet variables, enjoyment ratings).
Scope: one or two concrete questions per pass, such as whether low-fibre participants showed larger shifts over time or whether enjoyment aligned with responsiveness.
Guardrails: statistical or causal interpretations were not accepted unless they were consistent with the independent SME analysis (ANCOM-BC, PERMANOVA, DESeq2, MaAsLin2). Uncertainty was explicitly surfaced at each step.
2.6.2. AI Pattern Surfacing
The model proposed candidate patterns linking changes over time, dietary context, sample site, and enjoyment. Each proposal included a brief explanation of the underlying evidence (e.g., directional fold-change trends or participant-level consistency) and an uncertainty note describing alternative interpretations or potential confounders.
2.6.3. Human Curation
The analyst evaluated each pattern for biological plausibility, internal consistency, and coherence with the study design. Weak or unsupported proposals were discarded. Ambiguous cases were reframed for a later pass, while promising patterns were annotated with contextual reasoning drawn from microbiome domain expertise.
2.6.4. Synthesis (Human + AI)
Promising elements were consolidated into short, testable narratives. For orientation, simple exploratory checks (e.g., non-parametric group comparisons or paired difference summaries) were used to verify direction of effect only. All inferential claims relied on the SME statistical outputs. In cases where HIF and SME findings diverged, the discrepancy was flagged and revisited in subsequent passes. Simple quantitative summaries and exploratory visualisations (e.g., line plots, scatterplots and paired comparisons) were also generated within the HIF workflow using session-based Python scripts to support pattern checking. These did not replace the formal statistical outputs of the SME analysis.
2.6.5. Decision Criteria
Narratives were retained when they met all the following:
- (i)
Internal coherence across the three timepoints,
- (ii)
Contextual plausibility within the dietary and sensory setting, and
- (iii)
Consistency with SME statistical results.
Patterns not meeting all criteria were labelled exploratory and carried forward transparently.
2.6.6. Iteration
The conversation cycled until patterns stabilised and no new explanatory value emerged. Iteration typically required three to five passes per question, with later passes focused on resolving discrepancies between HIF and SME findings.
2.6.7. Audit Trail and Transparency
Prompts, intermediate notes, keep/discard decisions, and consolidated narratives were logged. This ensured a full audit trail linking raw inputs, HIF reasoning steps, and final outputs.
2.6.8. Outputs
The HIF workflow generated:
Participant-level responsiveness profiles (per site and timepoint);
Group-level narratives relating baseline diet context (fibre score, low/high categories, HDI grouping) to microbiome shifts;
Shortlists of taxa contributing to observed changes, each tagged as retained or exploratory;
Areas of convergence and divergence relative to the SME analyses.
Analyses were performed at both participant and group levels to accommodate substantial inter-individual variability.
3. Results
3.1. Overview
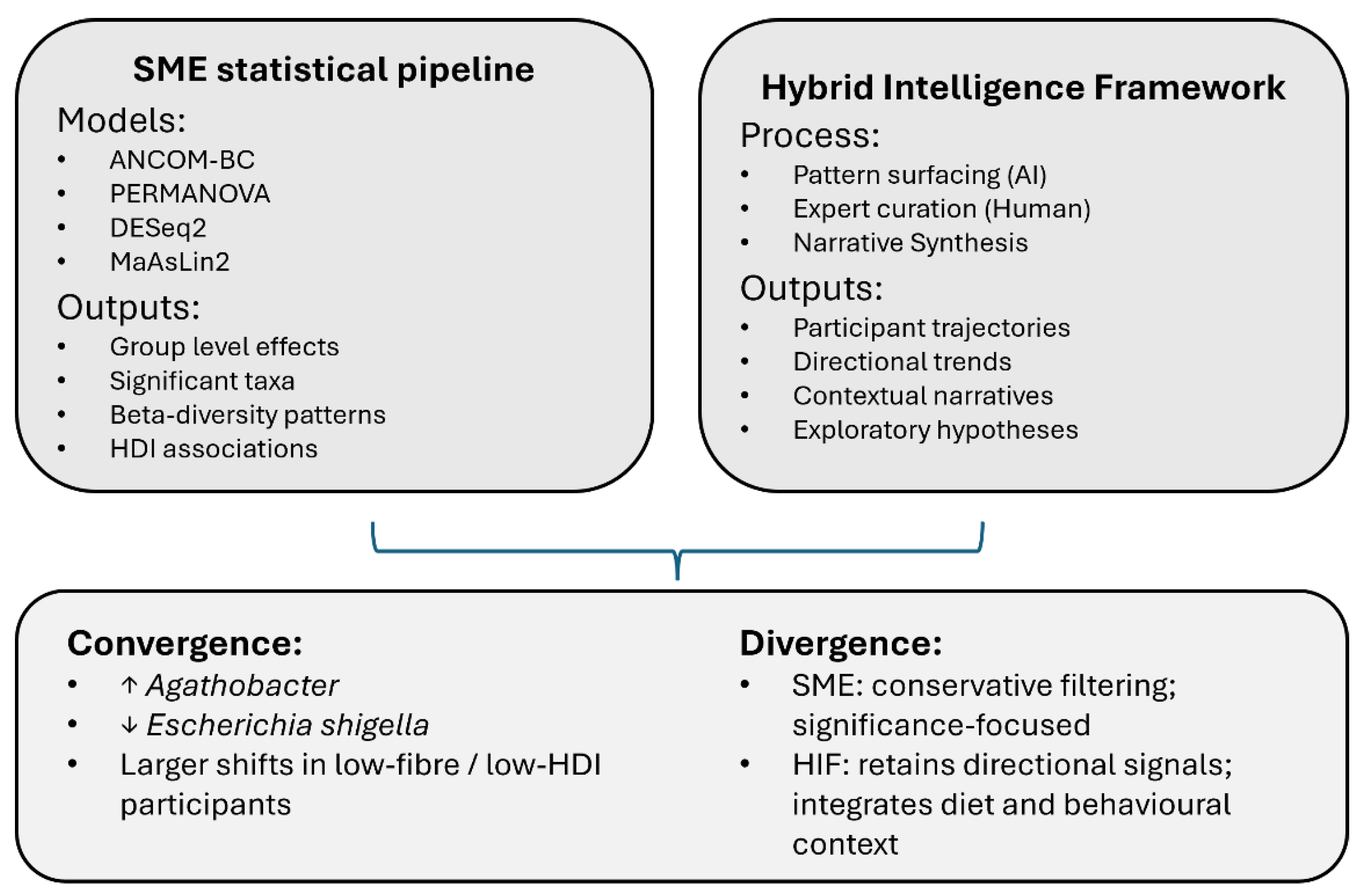
The HIF and the independent SME statistical analysis were applied to the same microbiome dataset. Because the manuscript focuses on methodology rather than full biological interpretation, this section presents the main points where HIF and SME analyses converged or diverged, plus minimal orientation needed for the reader. Detailed statistical outputs remain in Shindler et al. (submitted). A schematic summary of the comparison is provided in
Figure 3.
3.2. Baseline Diet Quality and Diversity Patterns
Both HIF and SME analyses identified baseline diet quality as an important contextual factor. Using the continuous baseline fibre-intake score and the derived low/high split, HIF identified a consistent directional trend: participants with lower baseline fibre tended to show larger compositional shifts across the intervention. This pattern appeared in both oral and faecal profiles and was visible in several participants from baseline to week 1 and maintained to week 2 of the study.
The SME team reached the same overall conclusion using the HDI. Ordination trajectories based on UniFrac distances showed greater displacement among low-HDI participants, and PERMANOVA on gastrointestinal beta diversity indicated significant differences between HDI groups. Alpha-diversity measures (richness, Shannon, Simpson) varied between individuals and across timepoints but showed no clear group-level shifts attributable to fibre intake or HDI in either analysis.
Although exact participant allocation differed slightly between the fibre-score split (HIF) and the HDI split (SME), the same individuals generally fell into the lower diet-quality grouping in both methods.
3.3. Taxa Contributing to Observed Changes
Through iterative passes, HIF highlighted a concise set of genera that shifted with the intervention across multiple participants. Increases in the butyrate-associated genus Agathobacter were seen in several of the more responsive cases, while opportunistic or inflammation-linked genera, including Escherichia-Shigella, Fusobacterium and Pseudomonas, tended to decrease. These patterns appeared across participants and were internally coherent across the three timepoints.
The SME differential-abundance approach identified the same genera as key contributors to change. Agathobacter was detected as a positive signal, and statistically significant reductions in Fusobacterium and Pseudomonas were reported in the gastrointestinal and oral microbiota, alongside decreases in Escherichia-Shigella for several participants.
Both analytical approaches therefore converged on the same high-level microbial interpretation: modest but directionally favourable shifts towards increased fermenters and reduced opportunistic taxa in a subset of participants, particularly those with lower baseline diet quality.
3.4. Divergence Between HIF and SME
The main convergence between approaches lies in the narrative they support. Both HIF and SME analyses indicate that participants with lower baseline diet quality or lower habitual fibre intake showed larger microbiome shifts over the two-week intervention, and that these shifts often involved increases in Agathobacter and decreases in genera such as Escherichia-Shigella, Fusobacterium and Pseudomonas. No contradictions between HIF and SME outputs were identified; rather, HIF summaries reflected patterns detected by the established statistical methods.
Divergence primarily reflects analytical emphasis rather than conflicting results. The SME analysis uses conservative filtering and significance-based thresholds, reporting only effects that meet model-specific criteria and emphasising group-level and genus-level findings with formal P-values. In contrast, HIF retains directional trends that are biologically plausible and contextually meaningful even when sample size limits statistical power. HIF integrates dietary and behavioural context directly into interpretation and focuses on participant-level trajectories, using SME outputs as an anchor rather than a replacement for narrative reasoning.
Taken together, both approaches land on the same core story; the divergence lies in scope and interpretive flexibility rather than in the underlying signals.
3.5. Behavioural Enjoyment
Behavioural enjoyment was explored within HIF as an interpretive axis, for example by considering whether participants with higher average enjoyment scores followed different trajectories. SME PERMANOVA models also examined enjoyment rankings for gastrointestinal beta diversity and detected some compositional differences, but these did not translate into robust genus-level associations and were not a central focus of the SME interpretation. Given the small sample size and the exploratory nature of these observations, enjoyment-related patterns are treated qualitatively in the discussion section rather than as primary outcomes in the Results.
4. Discussion
Because the HIF is implemented as a dialogue between human expertise and AI-enabled pattern surfacing, the interpretation offered here emerges from that joint process rather than from either contributor alone.
4.1. Principal Findings
Both HIF and the independent SME analyses identified a consistent pattern: participants with lower baseline fibre intake or lower HDI showed larger microbiome shifts across the two-week intervention. This was visible in both oral and faecal sites and was sustained from week 1 to week 2 for several individuals. At the genus level, Agathobacter increased in multiple participants and Escherichia-Shigella decreased, matching the SME findings.
These convergent outcomes indicate that HIF can surface interpretable hypotheses that align with conventional statistics in small-sample settings.
4.2. Methodological Contribution: Human-AI Collaboration in Small-Sample, Multivariate Data
HIF operationalises human-AI collaboration as an iterative conversation in which the model proposes candidate patterns and the analyst curates, reframes, and validates them. By focusing on directionality, internal consistency across timepoints, and agreement with the SME assessment, HIF mitigates over-interpretation in small cohorts. The framework is transparent by design, with logged prompts, intermediate summaries, and keep or discard decisions linking each conclusion to its underlying reasoning. This AI-assisted methodology complements established statistical workflows rather than displacing them and provides a practical route to integrate behavioural and dietary context with microbiome summaries where formal hypotheses are difficult to pre-specify.
4.3. Alignment with Conventional Analysis
HIF recovered all major signals detected by the SME analysis of data, including the greater responsiveness of low-fibre and low-HDI participants, increases in Agathobacter, and reductions in Escherichia-Shigella. The close correspondence between the two analyses supports the use of HIF as a complementary interpretive layer. In this setting, the SME statistics provided formal significance testing, while HIF contributed participant-level trajectories, contextual reasoning, and integrative narratives.
4.4. Behavioural Context and Adherence
Enjoyment was explored as an interpretive axis within HIF. Although SME analyses detected minor differences in gastrointestinal beta diversity by enjoyment ranking, these did not translate to stable genus-level effects. Within HIF, participants with higher enjoyment tended to adhere more consistently to the intervention, and clearer microbiome trajectories were observed in those cases.
Accordingly, enjoyment is treated as an exploratory behavioural correlate. The ability to examine such behavioural, dietary, and biological context within a unified reasoning loop illustrates a key strength of HIF relative to conventional statistical analysis.
4.5. Implications for Microbiome and Small-Sample Nutrition Studies
For early-phase nutrition and microbiome research, this case study suggests that HIF is particularly useful when investigators must interpret complex multivariate readouts in small cohorts. In such settings, conventional statistics alone can struggle to provide clear, interpretable narratives, whereas HIF helps to organise exploratory analysis, highlight biologically plausible trajectories, and link these to diet and behavioural context. The dual use of HIF and SME analysis here shows how exploratory, conversation-centred interpretation can sit alongside a pre-specified statistical pipeline without conflict.
For applied work in food innovation and sensory intervention trials, HIF offers a practical way to integrate microbiome, dietary and enjoyment data within a single reasoning loop, without requiring a full multidisciplinary analytical team at every step. The framework can help identify which patterns are most promising for follow-up, which participants or subgroups warrant closer examination, and which hypotheses should be prioritised for testing in larger, more definitive studies. These implications are most relevant for feasibility studies and pilot interventions, where decisions about whether and how to scale a concept often rely on limited but heterogeneous data.
5. Conclusions
This case study demonstrates that a structured HIF can generate coherent, biologically plausible insights from small, multivariate datasets that are traditionally difficult to interpret. By combining human expertise with AI-enabled pattern surfacing in an iterative conversational workflow, HIF enabled a single analyst to integrate diet, behaviour, and microbiome context in a way that would normally require multiple specialist roles. The resulting narratives aligned closely with an independent SME statistical pipeline, indicating that HIF can recover key signals while offering broader contextual interpretation.
The comparison also clarifies the relationship between the two approaches. In this study, the SME analysis served as the pre-specified statistical reference point, while HIF provided integrative, hypothesis-generating interpretation across behavioural, sensory, nutritional and microbial domains. The SME outputs were used to check and refine emerging HIF narratives rather than to constrain what could be explored. This division of labour positions HIF not as a replacement for specialist analysis but as a methodology that expands what individuals can do, improves analytical scalability and accelerates early-phase research.
Although exploratory, the incorporation of enjoyment illustrates HIF’s ability to merge behavioural context with biological readouts. Participants with higher enjoyment tended to show clearer microbial trajectories, consistent with better adherence. These signals should not be over-interpreted, but they demonstrate the value of a framework that can flexibly incorporate multiple data streams without requiring dedicated domain-specific analysts for each.
More broadly, the HIF is intentionally domain-agnostic. The principles applied here can support work in nutrition, product development, sensory science, engineering, and other fields where small datasets, high variability, or complex contextual questions make conventional pipelines hard to deploy. The present study represents the first structured application of HIF to food and sensory science, and it establishes a foundation for future work where problem framing, literature synthesis, experimental design, and data interpretation can be integrated into a single, transparent workflow.
Future applications should evaluate HIF in larger and longer studies, preregister prompts and decision criteria, and formally benchmark performance across different types of data. As research questions become more complex and more interdisciplinary, the capacity of HIF to unify reasoning across domains will become increasingly valuable.
Acknowledgments
This work was conducted using the Hybrid Intelligence Framework (HIF), integrating large language model (“Skippy”, GPT-5) analysis with human expertise in microbiome science and sensory evaluation. We thank the La Trobe University team for conducting independent statistical analyses.
References
- Amershi, S., Weld, D., Vorvoreanu, M., Fourney, A., Nushi, B., Collisson, P., Suh, J., Iqbal, S., Bennett, P.N., Inkpen, K., Teevan, J., Kikin-Gil, R., Horvitz, E. Guidelines for human–AI interaction. In Proceedings of the 2019 CHI Conference on Human Factors in Computing Systems, 2019; pp. 1–13. [CrossRef]
- David, L. A., Maurice, C. F., Carmody, R. N., Gootenberg, D. B., Button, J. E., Wolfe, B. E., Ling, A. V., Devlin, A. S., Varma, Y., Fischbach, M. A., Biddinger, S. B., Dutton, R. J., & Turnbaugh, P. J. Microbiota responses to different prebiotics are conserved within individuals but defined by baseline SCFA levels and habitual fibre intake. Microbiome 2022, 10, 114. [CrossRef]
- Hou, L., Yang, Y., Sun, B., Jing, Y., Deng, W., et al. Dietary fibre, gut microbiota, short-chain fatty acids and host metabolism. American Journal of Life Sciences 2021, 9, 162–172. [CrossRef]
- Hughes, R. L., Marco, M. L., Hughes, J. P., Keim, N. L., & Kable, M. E. The role of the gut microbiome in predicting response to diet and the development of precision nutrition models – Part I: Overview of current methods. Advances in Nutrition 2019, 10, 953–978. [CrossRef]
- Karim, M. R., Iqbal, S., Mohammad, S., Morshed, M. N., Haque, M. A., Mathiyalagan, R., Yang, D. C., Kim, Y. J., & Song, J. H. Butyrate’s (a short-chain fatty acid) microbial synthesis, absorption, and preventive roles against colorectal and lung cancer. Archives of Microbiology 2024, 206, 137. [CrossRef]
- Shindler, A., Mosca, A. C., Appelqvist, Batista, C., Srinivasagam Ramasamy, A., I., Brown, C., Moschonis, G., Belski, R., Cooke, M., Franks, A., Lane, A., Smilveska, S., & St. John, B. (submitted). Ingestion of a novel complex carbohydrate delivery vehicle leads to beneficial modulation of the oral and gastrointestinal microbiota of autistic adults: A pilot study [submitted].
- Vössing, M., Kühl, N., Lind, M., & Satzger, G. Designing transparency for effective human–AI collaboration. Information Systems Frontiers 2022. [CrossRef]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).