3. Results
3.1. Demographic, Systemic, and Oral Clinical Characteristics of the 31 Participants
A total of 31 participants were included in this study. Their demographic, systemic, and oral health characteristics are summarized in
Table 1. The study population consisted of adults with varying systemic conditions, including diabetes and hypertension. The number of remaining teeth and oral hygiene levels also differed among individuals. These data illustrate the heterogeneity of participants’ backgrounds, highlighting the potential influence of both systemic and local oral factors on the microbial composition in the oral cavity.
Systemic conditions (hypertension, hyperlipidemia, diabetes), lifestyle factors (current smoking and drinking), dental status (number of teeth, DMFT, DT, MT, FT), oral hygiene status and periodontitis status are listed. Continuous variables are presented as mean ± standard deviation (SD), and categorical variables as n (%). Abbreviations; DMFT, total number of decayed, missing and filled teeth: DT, decayed teeth; MT, missing teeth; FT, filled teeth. Oral hygiene was categorized as “good” when the plaque control record was <20%. Periodontitis was defined as having periodontal pockets ≥4 mm in ≥20% of teeth. These variables were included as covariates in the analyses of α diversity, β diversity and microbial composition.
3.2. Comparison of α Diversity Among Saliva, Plaque and Tongue Coating
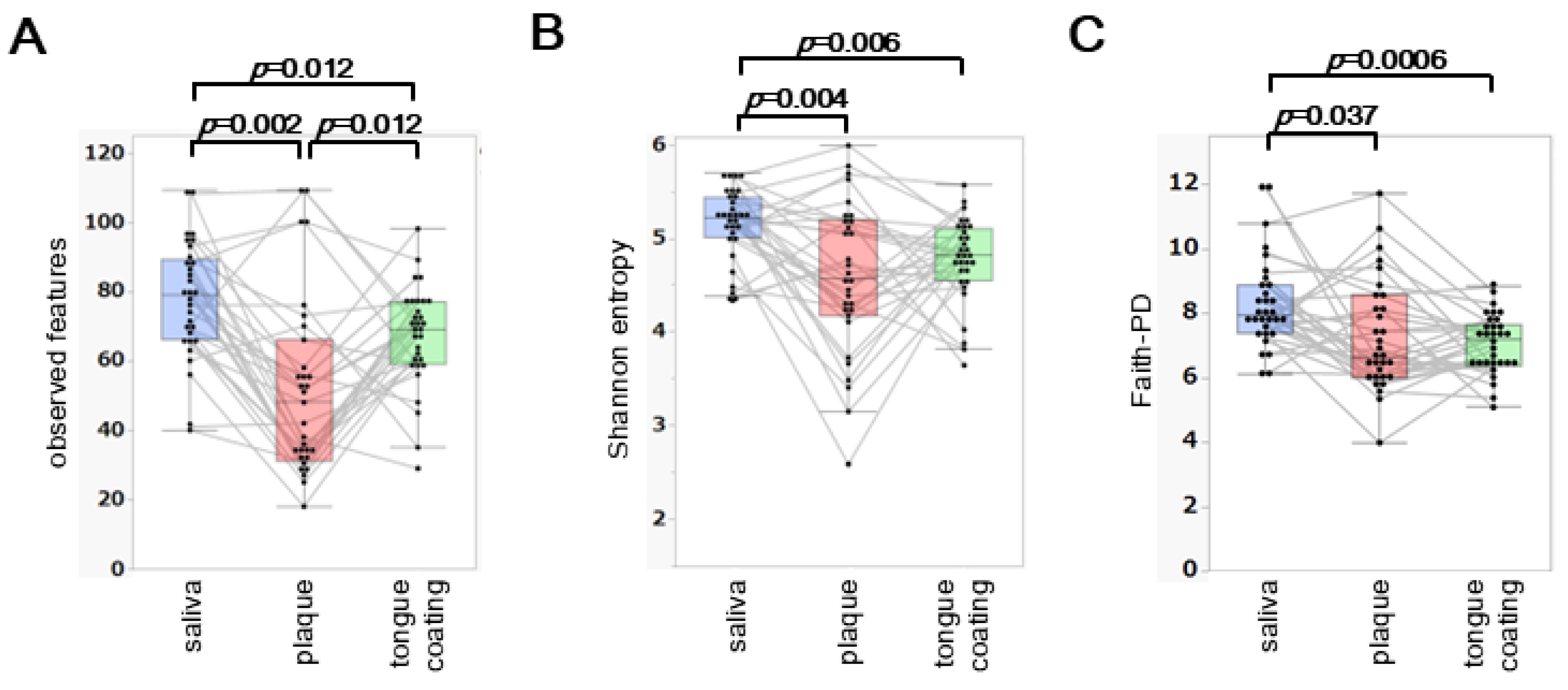
Three α diversity metrics, observed features, Shannon entropy and Faith’s phylogenetic diversity (Faith-PD), were compared among saliva, plaque and tongue coating from the same 31 participants.
Observed features differed significantly across all three sites (saliva > tongue coating > plaque;
p < 0.05) (
Figure 1A). For Shannon entropy, saliva showed the highest diversity, whereas plaque and tongue coating did not differ significantly (
Figure 1B). Faith-PD showed a pattern identical to that of Shannon entropy (
Figure 1C), with saliva exhibiting the highest phylogenetic diversity and no significant difference between tongue coating and plaque.
Overall, saliva consistently displayed the highest α diversity, while plaque and tongue coating showed similar diversity profiles. The lower richness in plaque likely reflects its more specialized, biofilm-associated microbial structure compared with the more dynamic environments of saliva and tongue coating.
(A–C) Boxplots of observed features (A), Shannon entropy (B) and Faith’s phylogenetic diversity (Faith-PD) (C) across saliva, plaque and tongue coating. Each thin line links samples from the same individual to illustrate intra-individual variation across oral sites. Statistical comparisons were performed using the paired Steel–Dwass test and only significant p values (p < 0.05) are shown within the figure.
3.3. Background Factors Influencing α Diversity
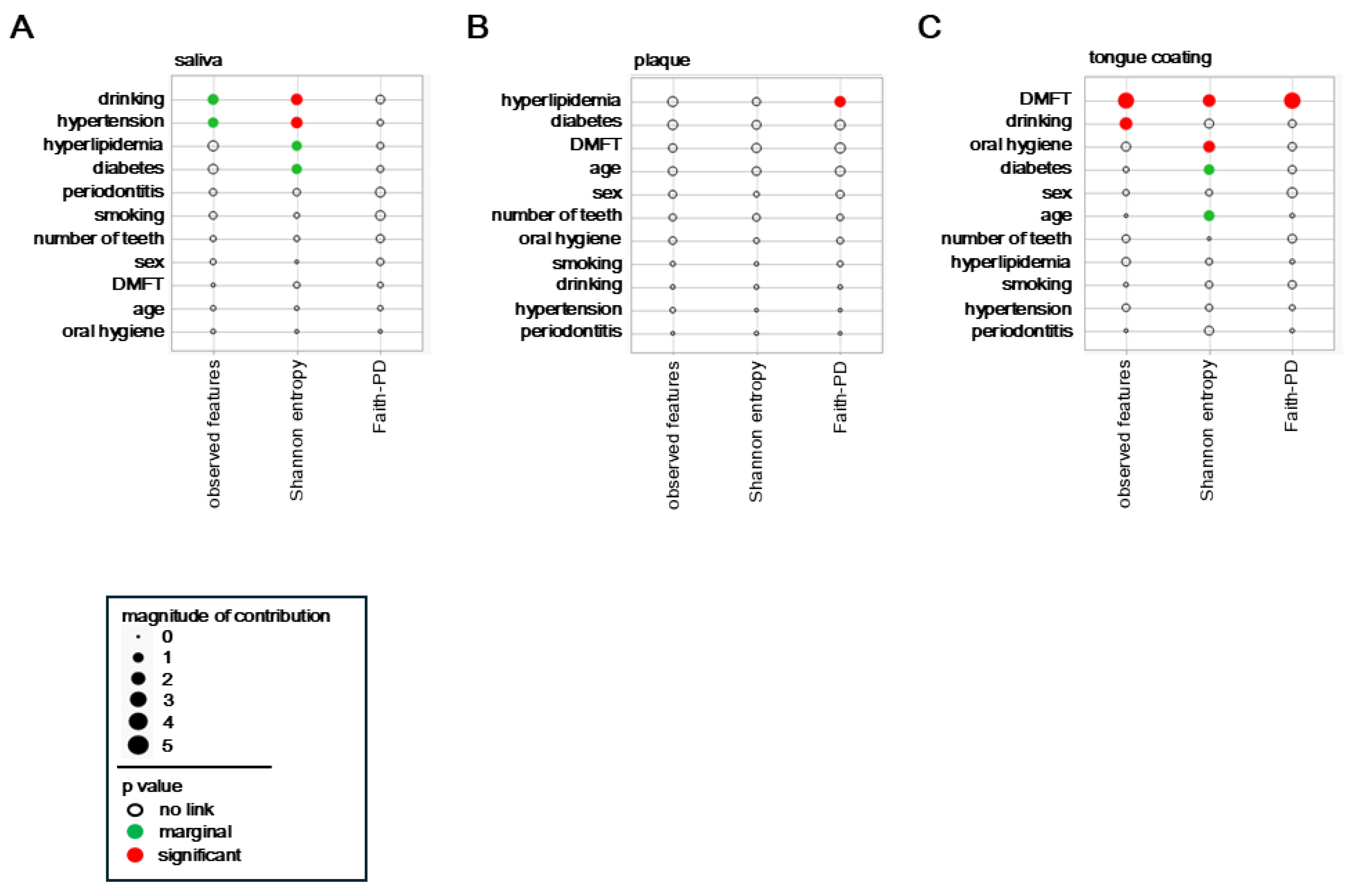
The associations between host and lifestyle factors and α diversity were analyzed separately for saliva, plaque, and tongue coating. Each circle represents one explanatory variable, with its area corresponding to the effect magnitude in the multivariate model. Red circles denote significant associations (p < 0.05), green circles indicate marginal associations (0.05 ≤ p < 0.10), and the position along the axis reflects the direction and relative contribution of each factor.
In saliva, Shannon entropy showed significant associations (red circles) with alcohol consumption and hypertension, indicating that the balance of salivary microbial communities is sensitive to behavioral and vascular factors (
Figure 2A). Moreover, observed features also tended to increase (green circles) with alcohol consumption and hypertension, suggesting that both richness and evenness are modulated by these conditions. Diabetes and hyperlipidemia displayed marginal associations (green circles) with α diversity, implying that systemic metabolic health may subtly influence the salivary ecosystem.
In plaque, only Faith-PD diversity showed a significant positive relationship (red circle) with hyperlipidemia, while other systemic factors exerted minimal effects (
Figure 2B). This indicates that plaque diversity is primarily governed by local biofilm dynamics, with limited responsiveness to systemic conditions.
In tongue coating, the DMFT index exhibited strong and consistent associations (red circles) across all α diversity metrics, observed features, Shannon entropy, and Faith-PD (
Figure 2C), demonstrating that cumulative oral disease experience profoundly shapes tongue surface microbial diversity. Additionally, observed features were marginally associated with alcohol consumption (green circle), and Shannon entropy was significantly influenced by oral hygiene (red circle). Diabetes also showed a marginal association (green circle), suggesting that metabolic disorders may affect the microbial ecology of the tongue dorsum.
Collectively, these findings indicate that systemic and lifestyle-related factors predominantly shape salivary and tongue–coating diversity, whereas plaque diversity remains locally constrained. Thus, saliva and tongue coating microbiota appear to be more strongly influenced by systemic and lifestyle-related factors, while plaque is comparatively stable and locally regulated. This gradient of systemic influence across oral sites underscores the complex interplay between host metabolism, behavior, and site-specific microenvironments in regulating oral microbial diversity.
3.4.β Diversity Analysis of Saliva, Plaque and Tongue Coating Microbiota
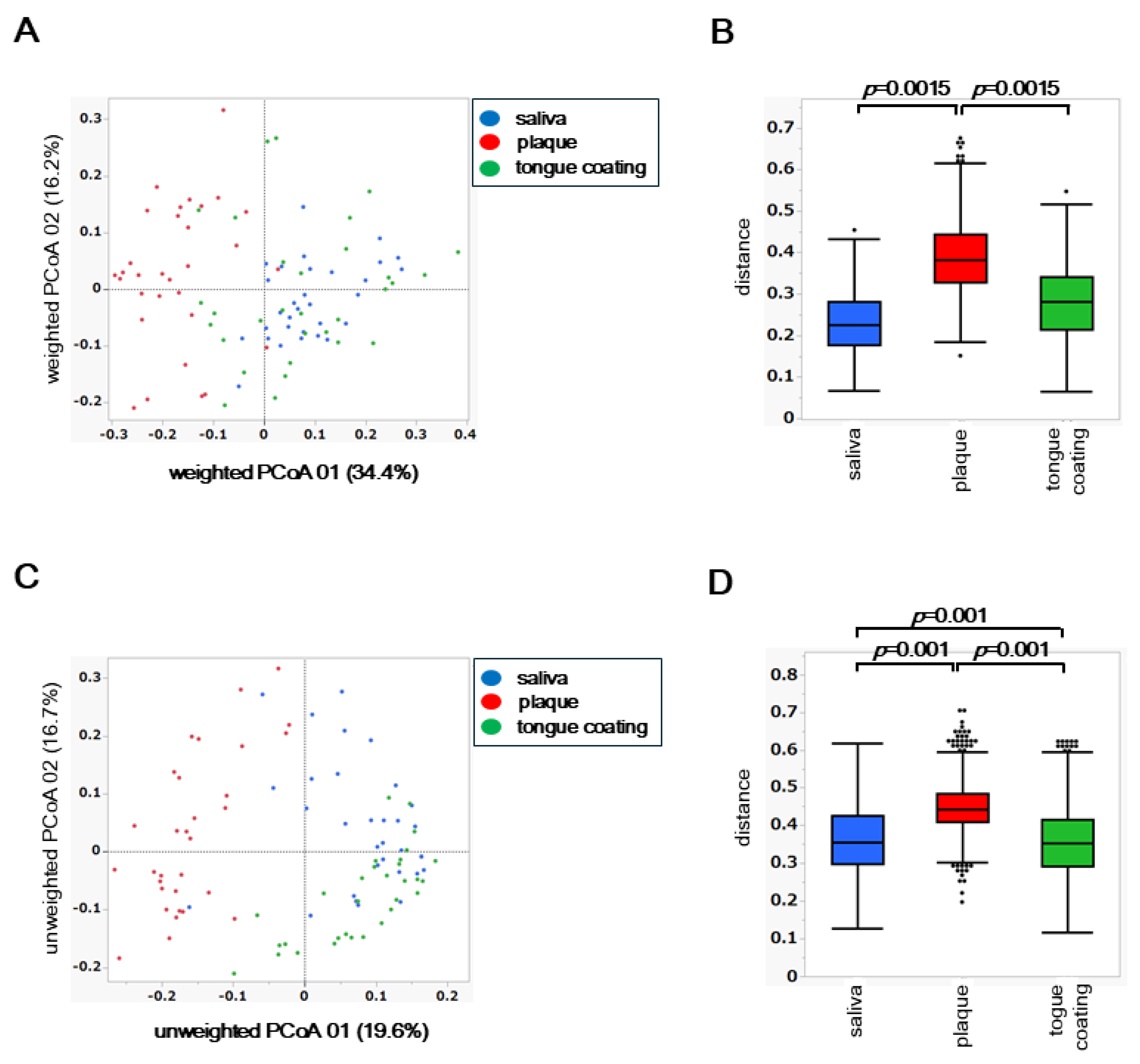
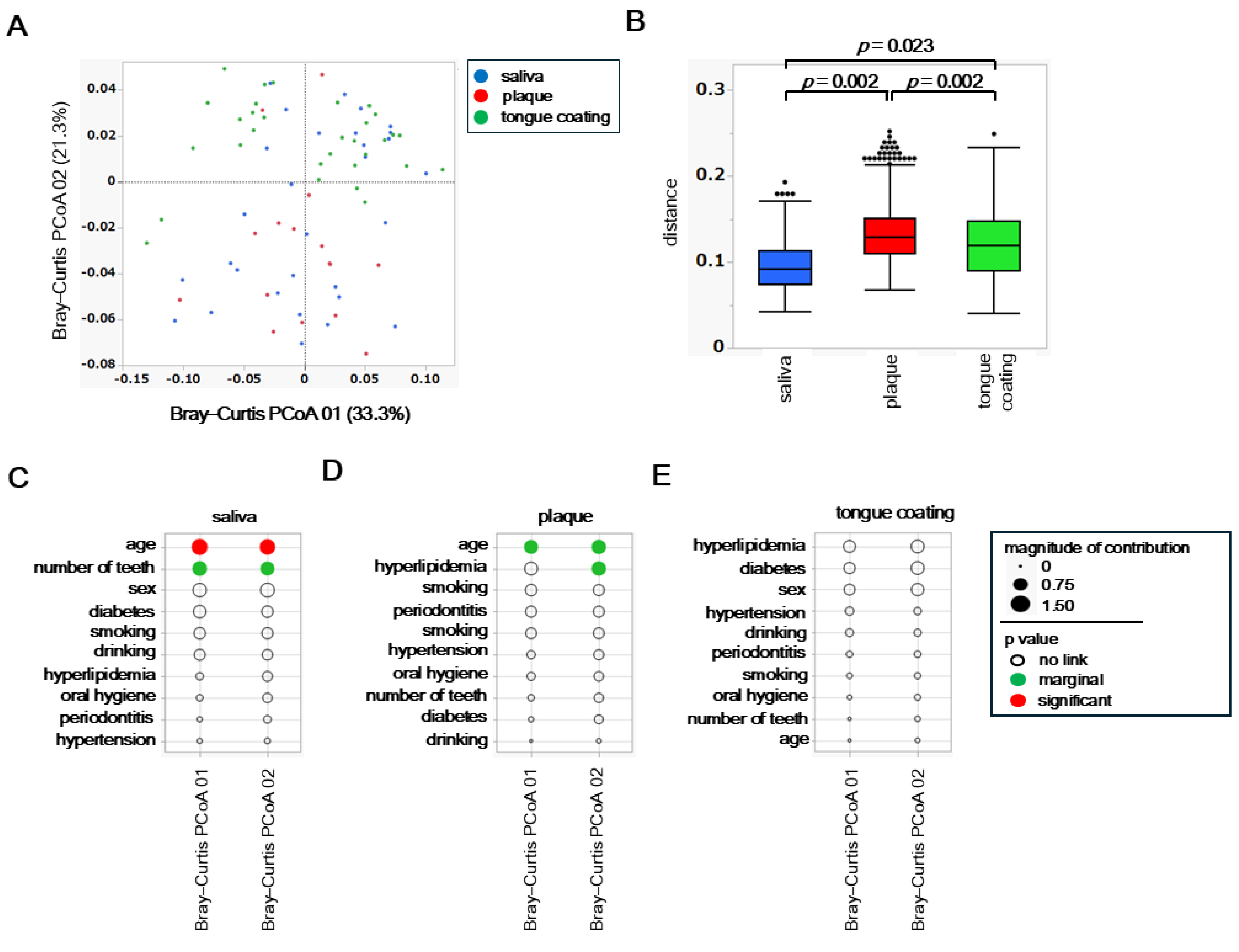
To assess inter-site differences in the overall composition of the oral microbiota, principal coordinate analysis (PCoA) was performed based on weighted and unweighted UniFrac distances. Differences among the three oral sites, saliva, plaque, and tongue coating, were statistically evaluated using permutational multivariate analysis of variance (PERMANOVA).
In the weighted UniFrac PCoA, which reflects differences in the abundance of dominant taxa, the microbial community structure of plaque was clearly separated from those of saliva and tongue coating (
Figure 3A). This finding indicates that plaque harbors a distinct microbiome shaped by its dense and adherent biofilm environment, where abundant taxa dominate in a site-specific manner. PERMANOVA confirmed a significant difference among the three oral sites (
p < 0.05), primarily driven by the compositional divergence of plaque from the other two niches (
Figure 3B).
In contrast, the unweighted UniFrac PCoA, which is based on the presence or absence of taxa including low-abundance species, revealed that saliva, plaque, and tongue coating each formed distinct clusters (
Figure 3C). PERMANOVA of the unweighted distances also showed significant differences among all site pairs (
p < 0.05), suggesting that rare or low-frequency taxa further contribute to the differentiation among oral niches (
Figure 3D).
Together, these findings demonstrate that while the dominant bacterial community of plaque differs most strongly from saliva and tongue coating, the entire microbial assemblage, including rare taxa, differs across all oral sites. This indicates a layered ecological organization in the oral cavity, in which abundant core microbiota are partially shared between saliva and tongue coating, whereas niche-specific and rare species define finer structural distinctions.
3.5. Background Factors Associated with β Diversity
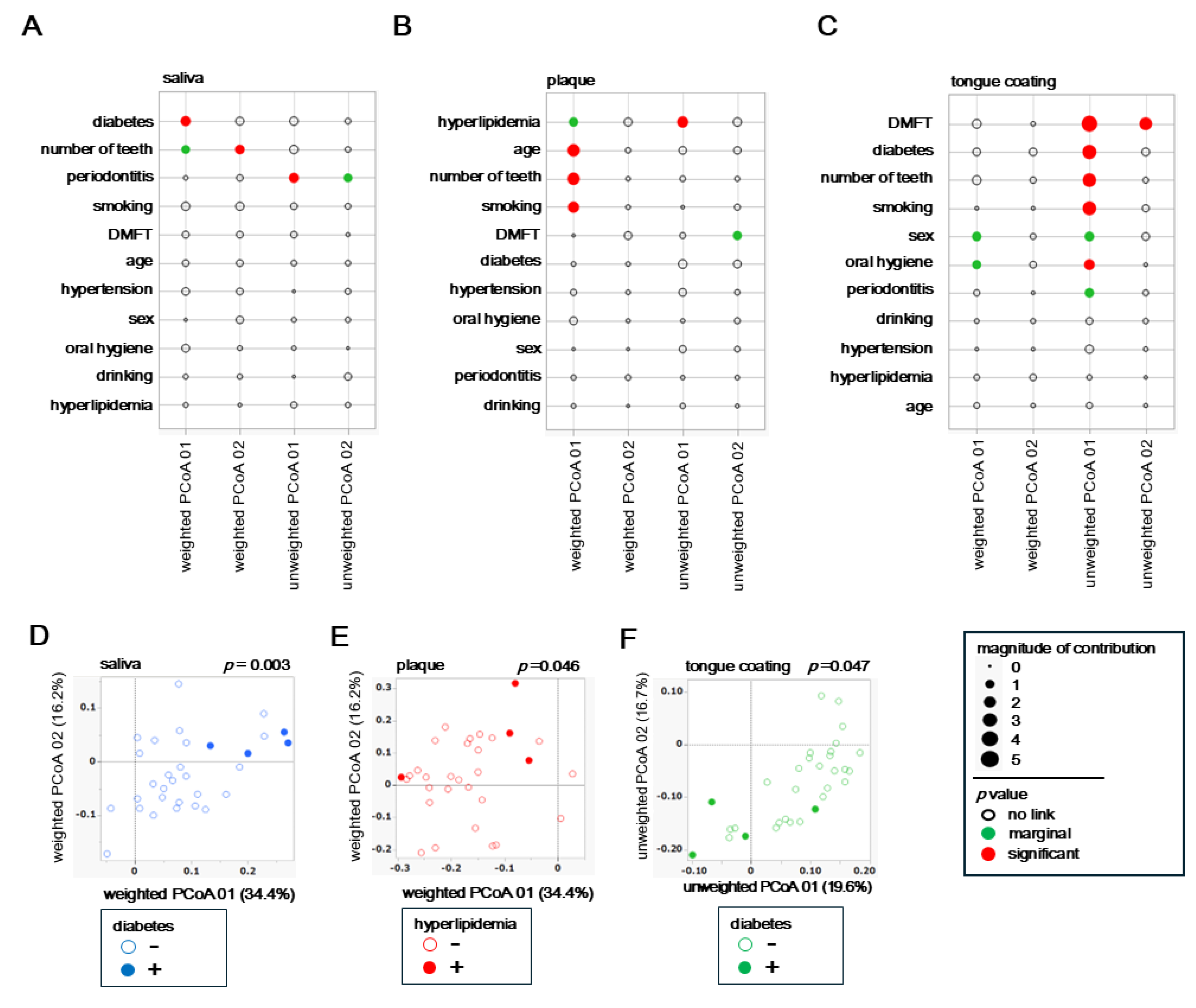
To examine how host background factors shape overall microbial community structure, associations between each variable and β diversity were evaluated using both weighted and unweighted UniFrac PCoA results for saliva, plaque and tongue coating. Weighted analyses represent variations driven by dominant taxa, whereas unweighted analyses highlight differences in rare or minor taxa.
In saliva, the weighted PCoA showed strong associations with diabetes and number of teeth (
Figure 4A), whereas the unweighted PCoA indicated a closer relationship with periodontitis. In plaque, the weighted PCoA revealed involvement of multiple factors such as hyperlipidemia, age, number of teeth and smoking history (
Figure 4B), while the unweighted PCoA identified hyperlipidemia as the sole contributing factor. In tongue coating, the weighted analysis suggested potential influences of sex and oral hygiene (
Figure 4C), whereas the unweighted PCoA indicated broader associations involving diabetes, various oral clinical indices, and smoking. These results indicate that the factors influencing microbial β diversity differed markedly among oral sites, with each niche being affected by distinct systemic or oral conditions. For examples, systemic metabolic conditions predominantly affected the salivary microbiota, both systemic and lifestyle factors interacted within plaque, and complex oral environmental conditions exerted the strongest effects on the tongue coating community.
To further illustrate these associations, PCoA plots were generated for the most influential systemic factors identified in each site. In saliva, individuals with diabetes (blue dots) were clearly separated along the weighted PCoA axis (
Figure 4D), confirming that diabetes significantly alters dominant salivary bacterial composition. In plaque, subjects with hyperlipidemia (red dots) showed partial separation in the weighted PCoA (
Figure 4E), suggesting that lipid metabolism influences biofilm-associated microbiota. In tongue coating, participants with diabetes (green dots) were separated in the unweighted PCoA, indicating compositional differences among low-abundance taxa on the tongue surface.
Together, these results demonstrate a layered pattern of host–microbiome interaction. Systemic conditions primarily reshape the dominant taxa composition in saliva and plaque, while intraoral and metabolic factors modulate the presence of minor taxa in tongue coating. Thus, saliva and tongue coating represent more dynamic and host-responsive niches, whereas plaque remains ecologically stable and locally constrained.
3.6. Correlation of β Diversity Among Oral Sites Based on Mantel Test Analysis
After identifying how systemic, lifestyle, and oral factors influenced β diversity at each oral site (
Figure 4), we next examined how similar the β diversity patterns were among saliva, plaque, and tongue coating within the same individuals using Mantel correlation analysis.
Figure 5A–C show correlations based on weighted UniFrac distances, while
Figure 5D–F depict those based on unweighted UniFrac distances, using samples collected simultaneously from the same individuals.
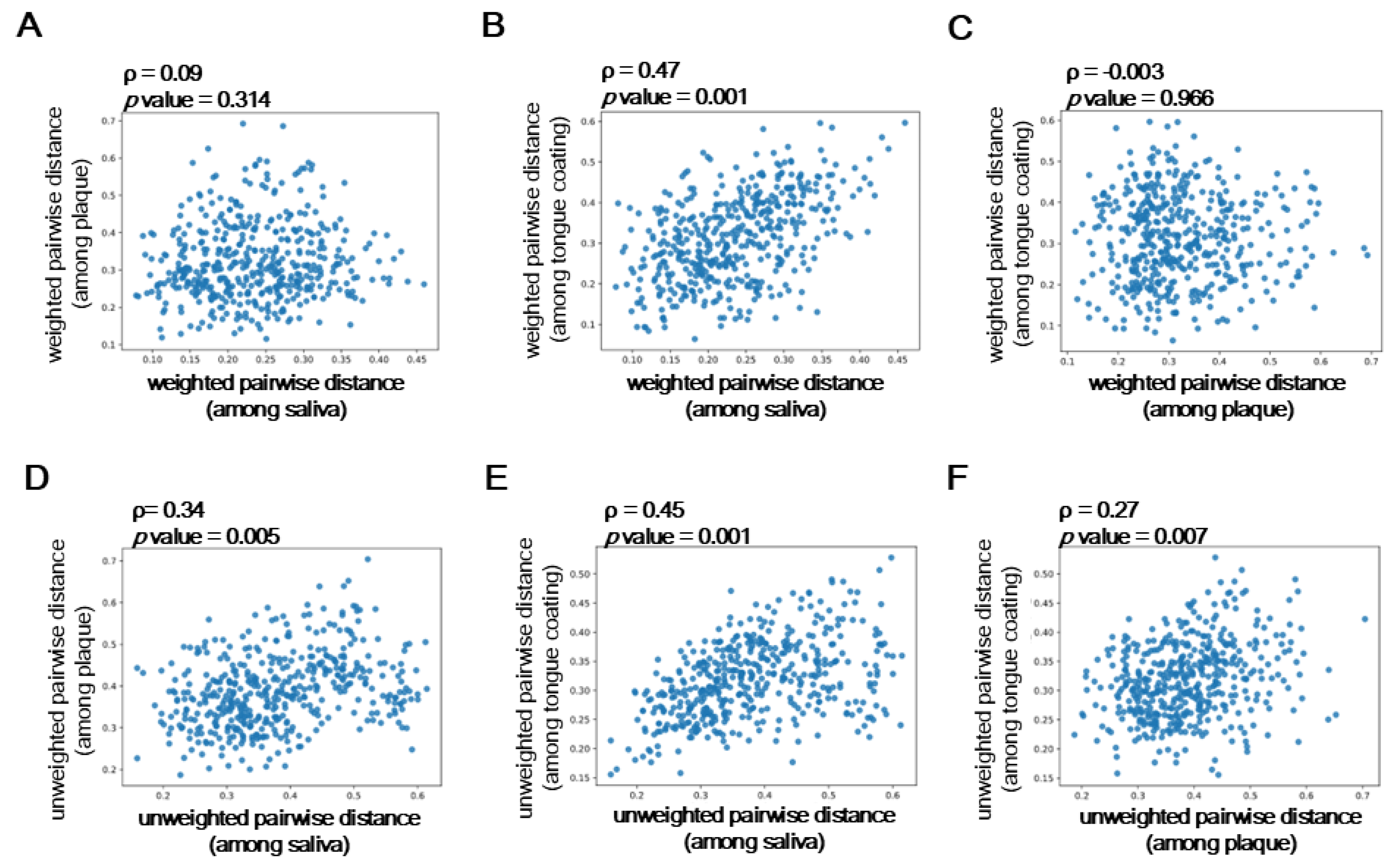
In
Figure 5A, the Spearman’s correlation coefficient (ρ) between saliva and plaque was weak and not statistically significant (ρ = 0.09,
p = 0.314), suggesting that salivary microbiota composition differs considerably from that of the plaque biofilm. In contrast,
Figure 5B shows that saliva and tongue coating exhibited the strongest correlation (ρ = 0.47,
p = 0.001), indicating that these two microbial niches share highly similar community structures.
Figure 5C, representing plaque versus tongue coating, showed almost no correlation (ρ = −0.003,
p = 0.966), reflecting the distinct microbial profiles between the hard biofilm and the mucosal surface.
Similarly, the unweighted UniFrac-based correlations (
Figure 5D–F) showed consistent tendencies.
Figure 5D (saliva vs plaque) showed a moderate but weaker correlation (ρ = 0.34,
p = 0.005), while
Figure 5E (saliva vs tongue coating) again showed the highest correlation (ρ= 0.45,
p = 0.001).
Figure 5F (plaque vs tongue coating) exhibited a lower but still significant association (ρ = 0.27,
p = 0.007), suggesting a limited overlap of minor taxa between these two sites.
Overall, these Mantel test results demonstrate that saliva and tongue coating harbor the most similar microbial community structures, whereas plaque remains compositionally distinct and locally stable. This pattern highlights the ecological heterogeneity within the oral cavity, saliva and tongue coating, reflecting dynamic and interchangeable communities, while plaque represents a more static, site-specific biofilm ecosystem.
3.7. Class-Level Composition and Factors Associated with the Firmicutes/Bacteroidetes (F/B) Ratio
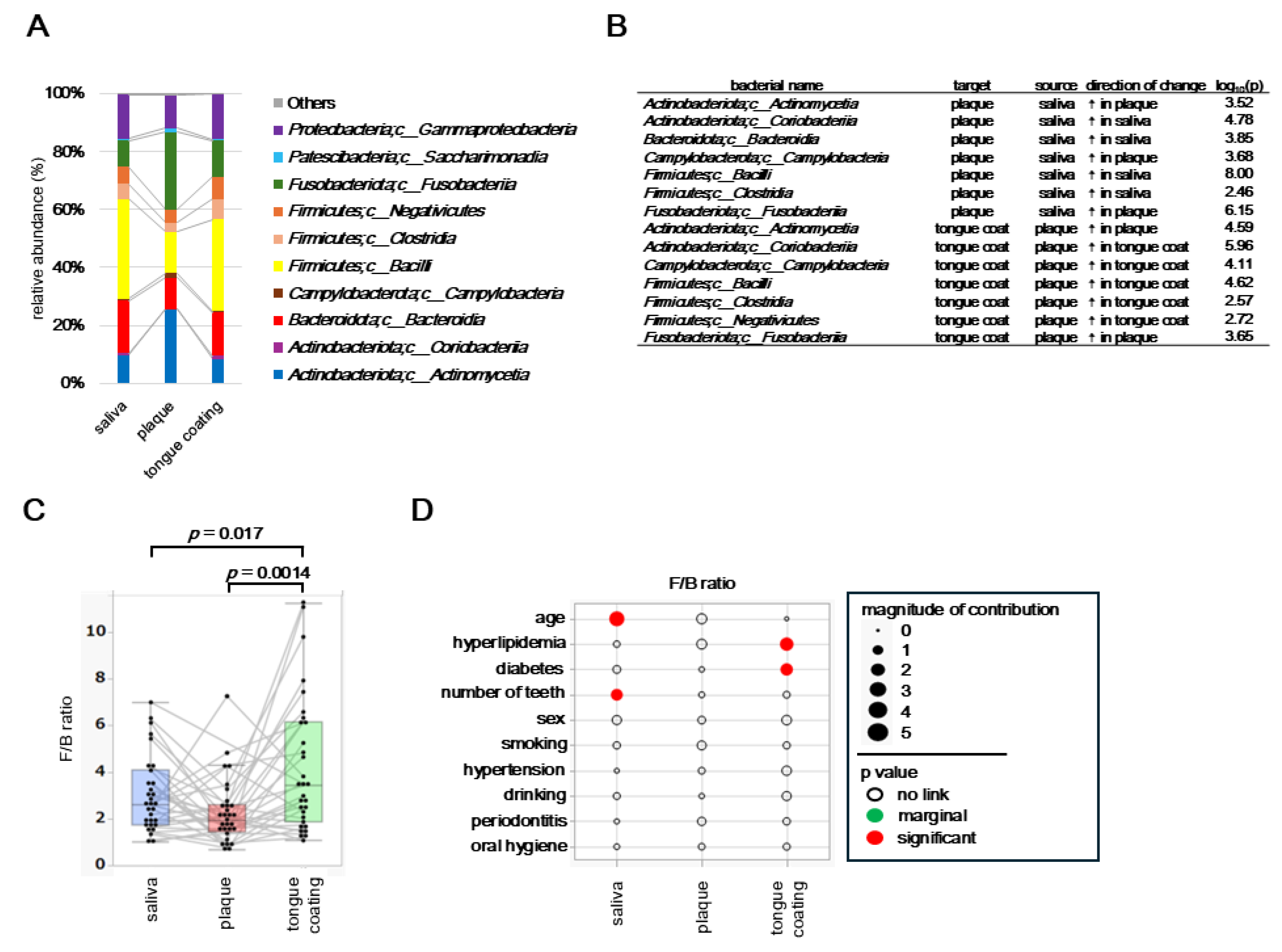
At the class level, microbial compositions differed markedly among saliva, plaque, and tongue coating, with only classes showing a relative abundance ≥ 1% displayed and taxa below this threshold were grouped as “Others”.
Saliva and tongue coating were dominated by
Bacilli and
Bacteroidia, whereas plaque exhibited higher proportions of
Actinobacteria and
Fusobacteria, reflecting the anaerobic and biofilm-enriched nature of this niche (
Figure 6A). Pairwise comparisons using the Steel–Dwass test identified several classes with significant inter-site differences (
Figure 6B).
Bacilli and
Bacteroidia were significantly more abundant in saliva and tongue coating, while
Actinobacteria and
Fusobacteria were enriched in plaque, highlighting ecological differentiation between fluid-associated and surface-associated habitats in the oral cavity.
The
Firmicutes/
Bacteroidetes (F/B) ratio, an indicator of microbial balance, varied significantly among oral sites (
Figure 6C). The ratio was highest in tongue coating, lowest in plaque, and intermediate in saliva (
p = 0.017 for saliva vs plaque and
p = 0.0014 for saliva vs tongue coating, Steel–Dwass test). Each thin line in the boxplot connects samples from the same participant, showing consistent intra-individual patterns across oral sites. Background factors influencing the F/B ratio were examined using multivariate models for each site (
Figure 6D). In saliva, the F/B ratio was significantly associated with age and number of teeth, suggesting that oral aging and tooth loss influence microbial balance in salivary communities. In plaque, no significant factors were detected, indicating that the plaque microbiome is locally regulated and less influenced by systemic background. In tongue coating, the F/B ratio was significantly associated with diabetes and hyperlipidemia, suggesting that microbial balance on the tongue dorsum reflects systemic metabolic status.
These results demonstrate that both microbial composition and the F/B ratio differ substantially among oral sites. While salivary F/B ratios are affected by oral conditions such as age and tooth loss, tongue coating F/B ratios are modulated by systemic metabolic disorders including diabetes and hyperlipidemia. In contrast, the plaque microbiome remains stable and locally maintained. Together, these findings indicate that saliva and tongue coating are sensitive indicators of oral and systemic health, whereas plaque represents a more resilient and localized microbial ecosystem.
3.8. Class-Level Heatmap Analysis of Oral Microbiota
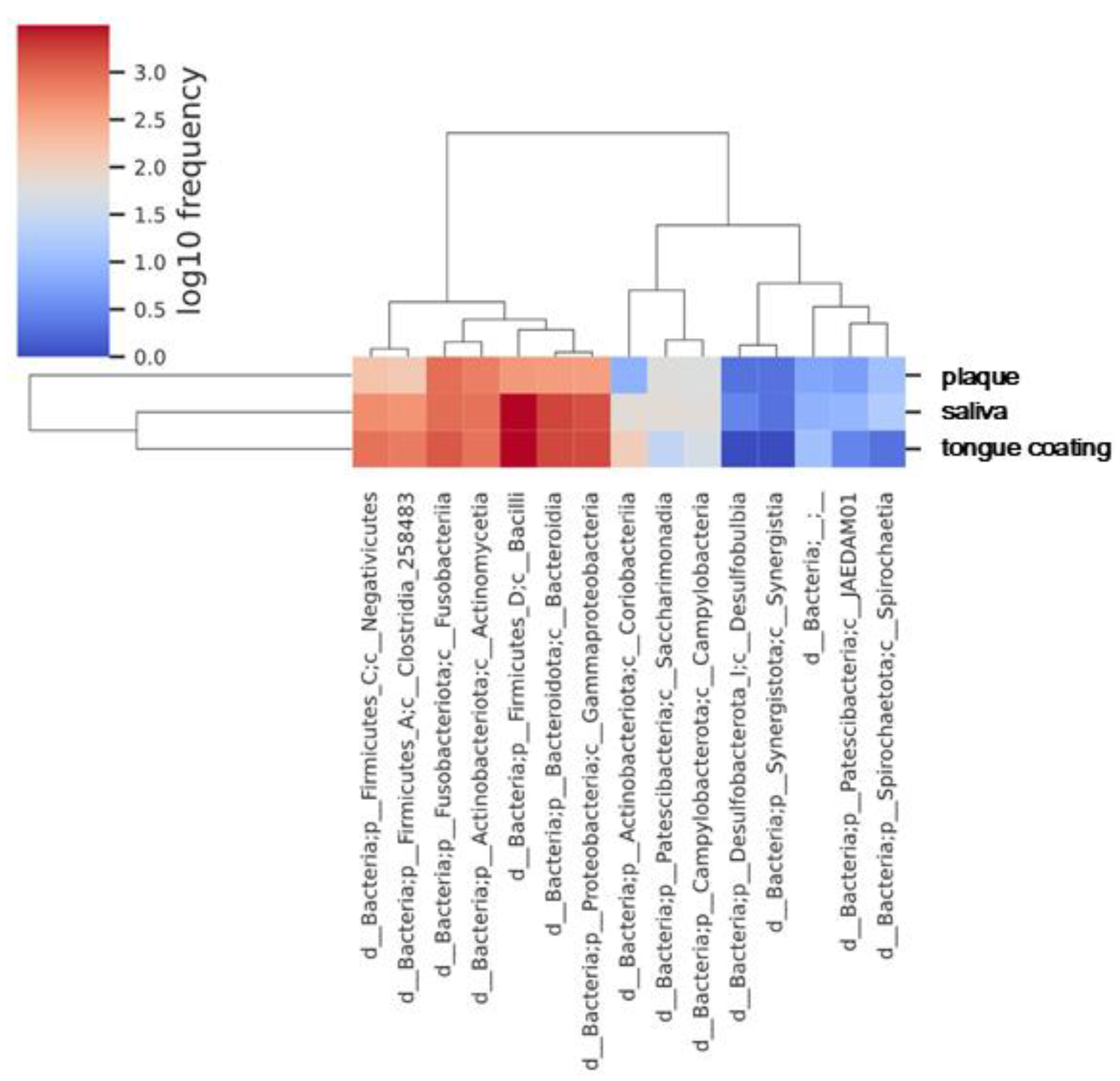
To further compare the microbial community structures among saliva, plaque, and tongue coating, a heatmap based on the relative abundance of bacterial classes was generated. Hierarchical clustering revealed distinct compositional patterns among the three oral sites.
Saliva and tongue coating samples clustered together, indicating that these two habitats share a closely related class-level microbiota (
Figure 7). Both were characterized by the dominance of
Bacilli and
Bacteroidia, representing mucosal-type bacterial communities that are continuously exposed to oxygen. In contrast, plaque samples formed a distinct cluster, enriched in Actinobacteria and Fusobacteria, which are typical of anaerobic, biofilm-associated environments. This clustering pattern is consistent not only with theα diversity findings (
Figure 1) but also with the β diversity analyses (
Figure 3), which showed that saliva and tongue coating have similar overall microbial structures. These results indicate that saliva and tongue coating harbor a related and dynamically connected microbiota, whereas plaque constitutes an ecologically distinct and more stable microbial community.
Heatmap of the log₁₀-transformed frequencies of bacterial classes in saliva, plaque and tongue coating. Values were averaged within each sample type before plotting. Hierarchical clustering was performed using Euclidean distance and Ward’s linkage method to group classes with similar distribution patterns. Sample types are shown on the horizontal axis, and bacterial classes are shown on the vertical axis.
3.9. Correlation Network Analysis of of Class-Level Bacteria Within and Across Oral Sites
To explore intra- and inter-site relationships among bacterial taxa, correlation networks were constructed using Spearman’s correlations (ρ ≥ 0.4, p < 0.05). Each node represents a bacterial class, node size corresponds to its relative abundance, and each edge indicates a significant positive correlation (red) or negative collection (blue).
In saliva, the network showed a simple and linear pathway, connecting a few major taxa such as
Bacilli and
Bacteroidia (
Figure 8A). This structure indicates a relatively stable and streamlined microbial community in which a small number of dominant taxa form a continuous relationship network.
The plaque network was the most complex, forming a dense web of interconnections consistent with the biofilm nature of plaque (
Figure 8B). It was overlaid on the Socransky pyramid, a classic model of subgingival microbial ecology [
30]. Even at the class level, the hierarchical structure resembled this model, where basal taxa (e.g.,
Actinobacteria) appeared to stabilize or suppress upper-layer taxa (e.g.,
Fusobacteria), reflecting ecological control within the plaque biofilm. The tongue coating network was also simple, similar to saliva, and largely followed a single main route of connections among
Bacteroidia and
Negativicutes (
Figure 8C). This linear topology suggests that the tongue surface microbiota forms a stable and less complex community under dynamic but controlled mucosal conditions.
Figure 8D integrates the three oral sites into a cross-site correlation network. No significant edges were detected between identical bacterial classes across different sites. Instead, multiple significant correlations emerged between distinct classes across sites, implying interdependent but functionally non-redundant relationships among oral habitats. These cross-site links may reflect ecological accelerators or brakes that modulate microbial balance and adaptability within the oral ecosystem, a novel finding unique to this intra-individual design.
Overall, saliva and tongue coating shared simple, nearly linear network structures, whereas plaque displayed a complex hierarchical pattern reminiscent of the Socransky pyramid. The integrated network revealed unique inter-class relationships rather than direct overlaps, suggesting that oral microbial communities across different niches are ecologically connected yet functionally distinct within the same individual.
Correlation networks were constructed using Spearman correlations (ρ ≥ 0.4,
p < 0.05). Nodes represent bacterial classes; node size reflects their relative abundance within the corresponding sample type, and node shape denotes the oral site (circle: saliva; triangle: plaque; square: tongue coating). Edge colors indicate the direction of correlation (red: positive; blue: negative), and edge thickness represents correlation strength. Node colors correspond to the class-level color scheme used in
Figure 7. (A) Network of saliva samples. (B) Network of plaque samples displayed over the simplified Socransky complex pyramid. (C) Network of tongue coating samples. (D) Cross-site network integrating saliva, plaque and tongue coating; node size reflects the relative abundance within each respective site.
3.10. ANCOM-BC Analysis of Differential Bacterial Genus Abundance Across Oral Sites
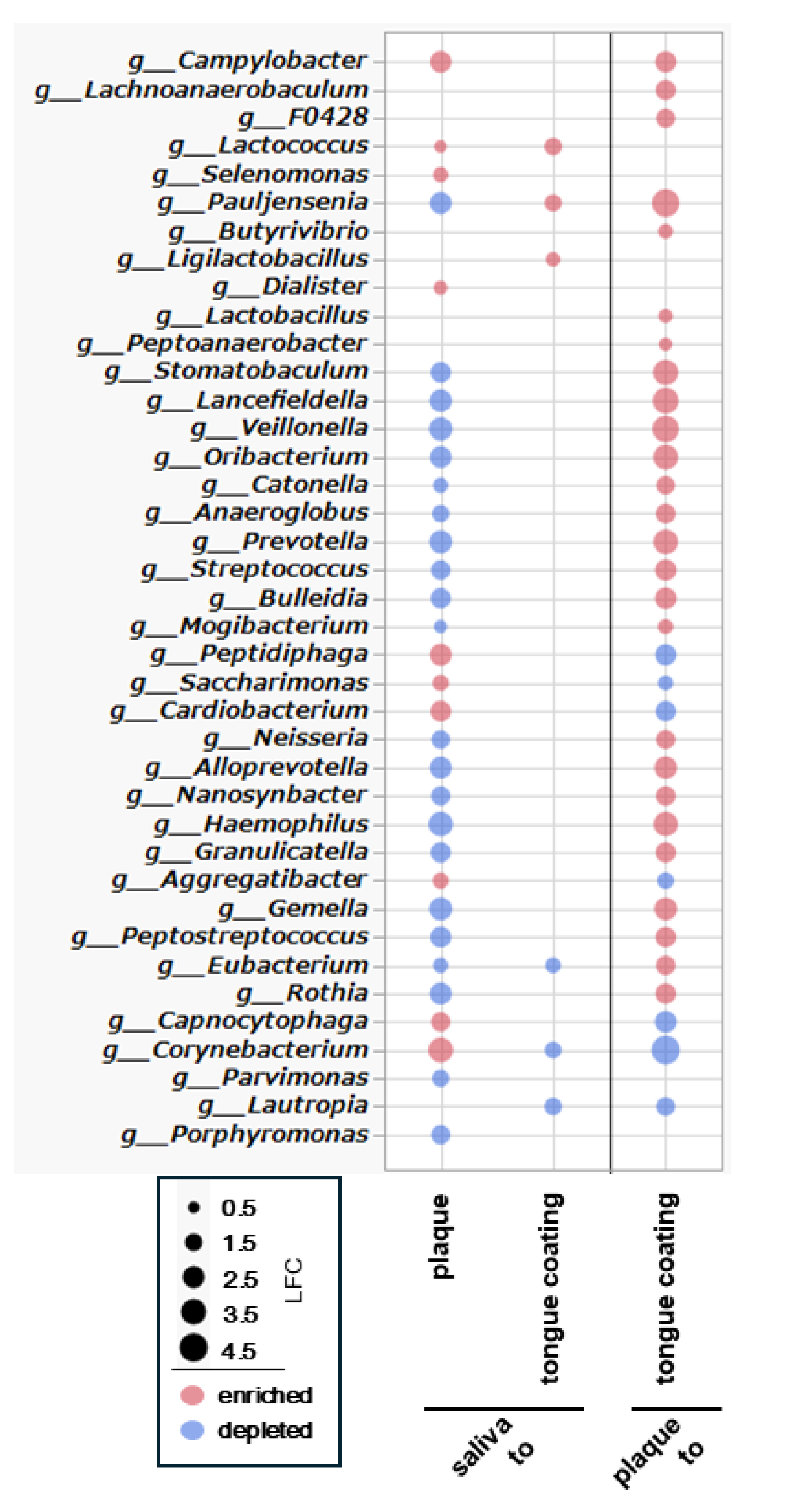
To clarify genus-level differences among oral sites, ANCOM-BC was applied to all pairwise comparisons (saliva–plaque, saliva–tongue coating and plaque–tongue coating). Because 16S rRNA sequencing offers limited species-level resolution, analyses were performed at the genus level and visualized in a single bubble plot summarizing significant log₂ fold changes (
Figure 9).
In the comparison between saliva and plaque, plaque exhibited larger divergence, with 10 genera enriched and 22 depleted relative to saliva. Enriched genera included anaerobic and biofilm-associated taxa such as Campylobacter, Lactococcus, and Selenomonas, whereas genera that are more abundant and commonly found in saliva such as Streptococcus and Neisseria were depleted. These findings highlight the strong ecological separation between fluid-associated (saliva) and biofilm-associated (plaque) communities.
In contrast, the comparison between saliva and tongue coating revealed only modest differences, with 3 genera enriched and 3 depleted. Enriched taxa included mucosa-associated genera such as Lactococcus, reflecting the close ecological relationship between saliva and the tongue coating and their shared exposure to oxygenated mucosal surfaces.
A broader shift emerged in the comparison between plaque and tongue coating, in which tongue coating demonstrated 26 enriched and 7 depleted genera. Mucosal or oxygen-tolerant genera such as Streptococcus, Neisseria, and Haemophilus were enriched, whereas several plaque-associated anaerobic taxa were reduced. This pattern underscores the distinction between the dense anaerobic biofilm of plaque and the more dynamic, oxygen-exposed microbiota of the tongue surface.
In summary, the genus-level ANCOM-BC analysis demonstrates that plaque is the most compositionally distinct niche, while saliva and tongue coating are similar, differing only in a few genera. These findings provide a taxonomic framework that supports the ecological and functional contrasts observed across oral sites in subsequent analyses.
Differentially abundant bacterial genera among saliva, plaque and tongue coating were identified using ANCOM-BC and visualized in a bubble plot. Each bubble represents a genus with significant differences (q < 0.05). Bubble color indicates the direction of log₂ fold change (LFC) (red: enriched; blue: depleted), and bubble size reflects the magnitude of the log₂ fold change. Comparisons are shown for saliva→plaque, saliva→tongue coating and plaque→tongue coating.
3.11. Functional Prediction of Oral Microbiota Using PICRUSt2
Functional prediction of the oral microbiota was performed using PICRUSt2, and Bray–Curtis PCoA was applied to assess differences in predicted metabolic functions among saliva, plaque, and tongue coating.
The overall ordination pattern showed clear separation among the three sites, similar to the unweighted UniFrac PCoA results (
Figure 10A), indicating that differences in minor (low-abundance) taxa substantially contribute to functional divergence across oral habitats. Plaque exhibited the most distinct functional profile (
Figure 10B). This suggests that functional differentiation within the oral microbiota arises largely from variation in low-abundance bacterial members.
Background factor analysis revealed that different host characteristics influenced metabolic pathway variation across sites, the functional profile in saliva was significantly associated with age and number of teeth (
Figure 10C), implying that oral functional diversity reflects both aging and dentition status. In plaque, age and hyperlipidemia were the main factors associated with Bray–Curtis dissimilarities (
Figure 10D), suggesting that biofilm functional capacity is affected by systemic lipid metabolism as well as local aging. In tongue coating, the functional profiles appeared relatively stable (
Figure 10E), showing minimal influence from host background factors, consistent with its role as a mucosal surface maintaining ecological homeostasis.
These findings indicate that functional differences among oral sites are primarily driven by minor taxa, and that host background factors such as age, dentition, and lipid metabolism selectively influence site-specific functional profiles.
Saliva reflects age- and dentition-related changes, plaque functions respond to systemic lipid status, and tongue coating functions remain stable within the oral ecosystem.
3.12. Distribution of Metabolic Pathways Among Oral Sites
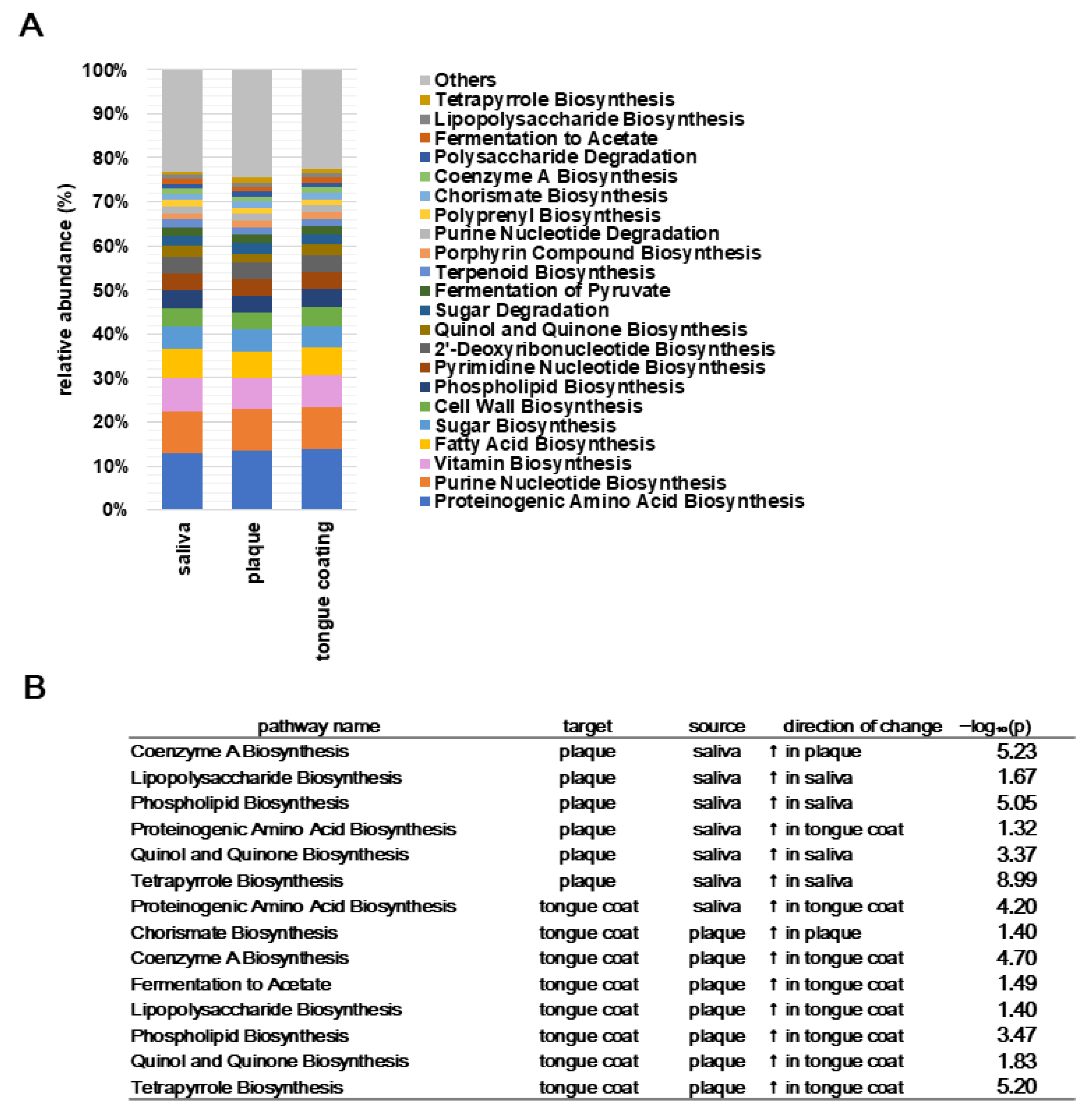
To characterize the predicted metabolic functions across oral habitats, PICRUSt2-based pathway predictions were summarized and statistically compared. In
Figure 11A, distinct compositional patterns were observed across oral sites. Plaque showed enrichment of biosynthetic pathways such as proteinogenic amino acid biosynthesis, phospholipid biosynthesis, and tetrapyrrole biosynthesis, consistent with the anabolic, biofilm-associated nature of this community (
Figure 11B). Saliva exhibited a balanced mixture of biosynthetic and degradative functions, reflecting continuous microbial exchange and environmental fluctuations. In contrast, tongue coating displayed higher proportions of fermentation- and carbohydrate-related pathways (e.g., fermentation to acetate, sugar degradation), consistent with its adaptation to a mucosal surface with variable oxygen tension. As summarized in
Figure 11B, the Steel–Dwass test revealed several pathways with significant differences among the three sites. Plaque was enriched in amino acid, coenzyme A, and phospholipid biosynthesis, while saliva and tongue coating showed increased representation of fermentation and polysaccharide degradation pathways.
These results demonstrate that biosynthetic activity predominates in plaque, while mucosal sites (saliva and tongue coating) exhibit greater fermentative and carbohydrate-degrading potential. The distribution of functional pathways mirrors the structural and physiological differences among oral sites, forming a gradient from anabolic to catabolic metabolism within the oral cavity.
3.13. Differential Analysis of Metabolic Pathways Among Oral Sites
After clarifying the overall distribution of metabolic pathways and identifying significant inter-site differences, we performed ANCOM-BC analysis on PICRUSt2 inferred metabolic pathways (
Figure 12).
In the saliva–plaque comparison, nine pathways were enriched in plaque. LPS biosynthesis was the only increasing biosynthetic pathway, while the other enriched pathways belonged to degradation-related functions. Despite clear genus-level differences, the modest number of pathway shifts suggests partial functional overlap between saliva and plaque.
A greater functional divergence appeared between saliva and tongue coating, where 23 significantly altered pathways were decreased in tongue coating. Many were degradation pathways, including several nitrogen-related degradation pathways. Because these pathways participate in the breakdown of nitrogenous compounds linked to volatile sulfur compound formation, the reductions suggest a distinct metabolic environment on the tongue surface. This functional separation contrasts with the strong taxonomic similarity between saliva and tongue coating.
The plaque–tongue coating comparison showed the widest set of changes, with 37 pathways differing significantly (three enriched and 34 depleted in tongue coating). Many of the depleted pathways were general degradation and nitrogen-related degradation pathways, indicating broad functional down-regulation on the tongue surface. This pattern shows that the tongue coating has a markedly reduced metabolic degradation capacity, making it functionally more distant from plaque as well as from saliva.
Overall, pathway-level ANCOM-BC identifies tongue coating as the most functionally distinct niche, exhibiting the largest number and magnitude of pathway shifts, particularly reductions in degradation and nitrogen-related processes. This overall pattern is consistent with the Bray–Curtis functional separation in
Figure 10A and 10B. However, it clearly contrasts with the genus-level ANCOM-BC results in
Figure 9, where saliva and tongue coating appeared taxonomically similar. These findings indicate that tongue coating can display substantial functional divergence even when its genus-level composition overlaps with that of other oral sites.
Metabolic pathways predicted by PICRUSt2 were compared across saliva, plaque, and tongue coating using ANCOM-BC. Only pathways showing significant differences (q < 0.05) and with an absolute log₂ fold change (|LFC|) ≥ 1.0 are displayed. Each bubble represents one pathway; red indicates enrichment and blue indicates depletion, and bubble size corresponds to the magnitude of the LFC.