Submitted:
30 November 2025
Posted:
02 December 2025
Read the latest preprint version here
Abstract
Keywords:
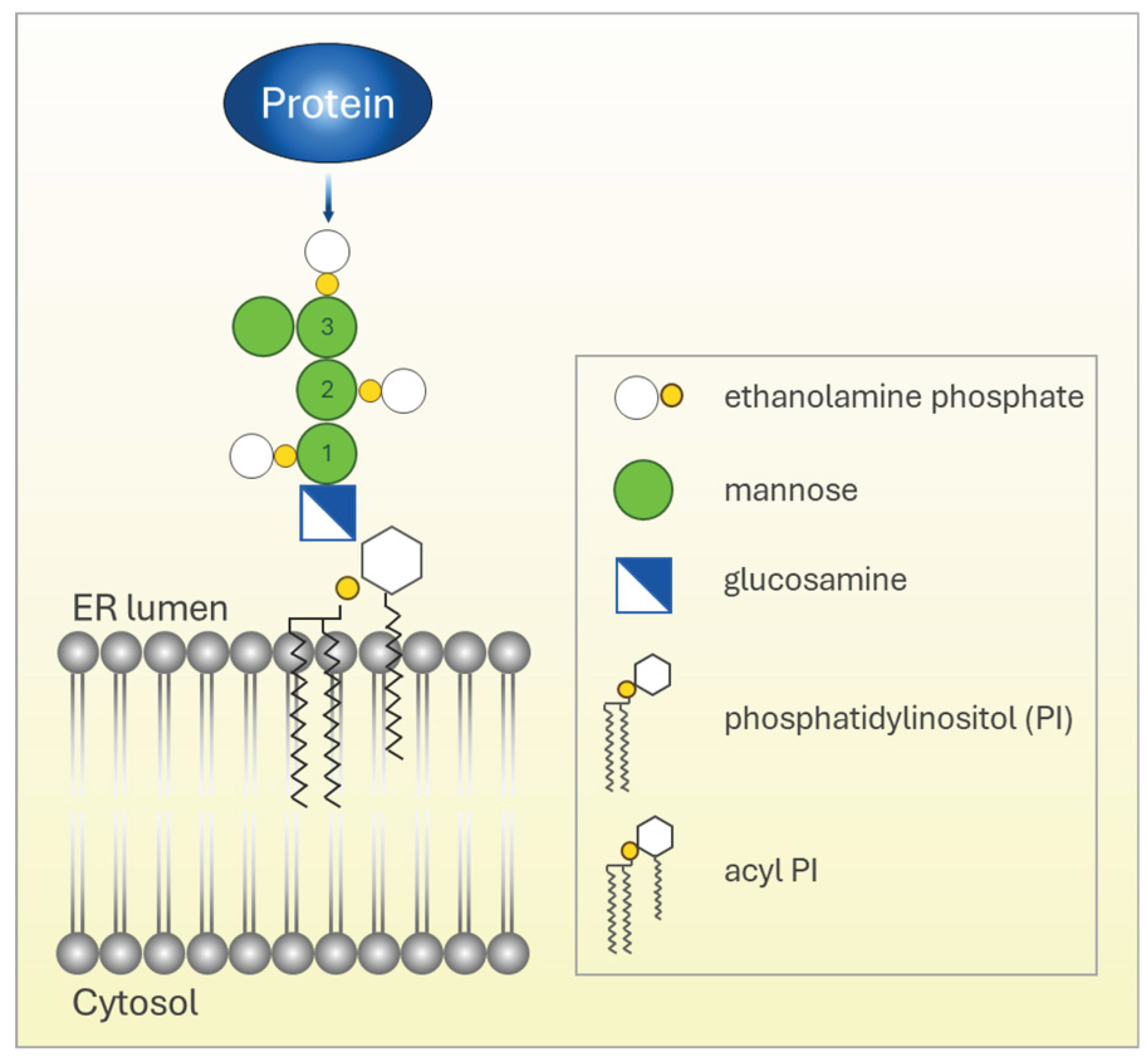
1. Biogenesis of GPI-Anchored Proteins
2. Physiological and Clinical Significance of GPI-Anchored Proteins
3. Protein Quality Control in the Secretory Pathway
4. GPI Anchor Remodeling and Implications for GPI-AP QC
5. Mechanisms of GPI-AP QC in Different Model Systems
5.1. Yeast
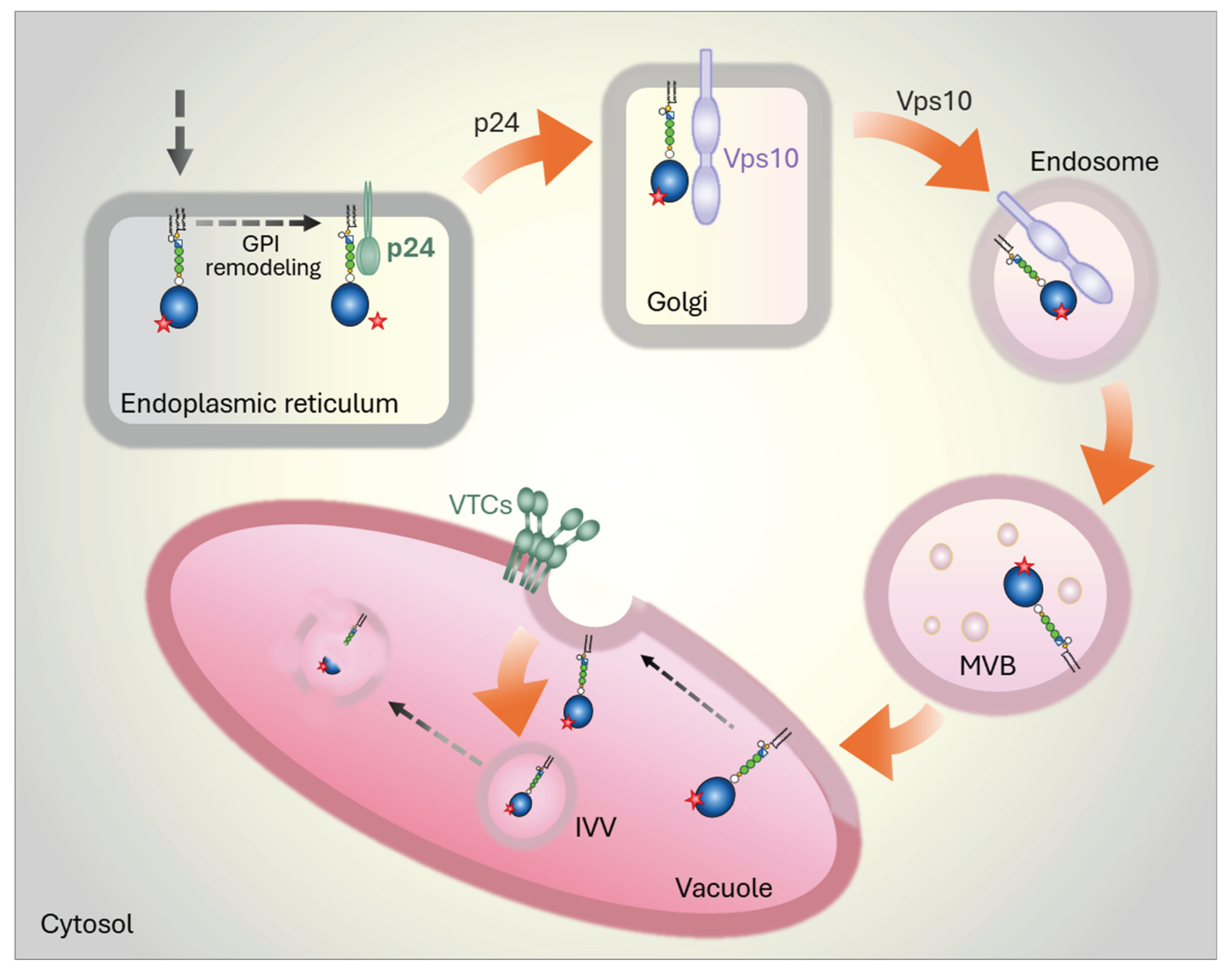
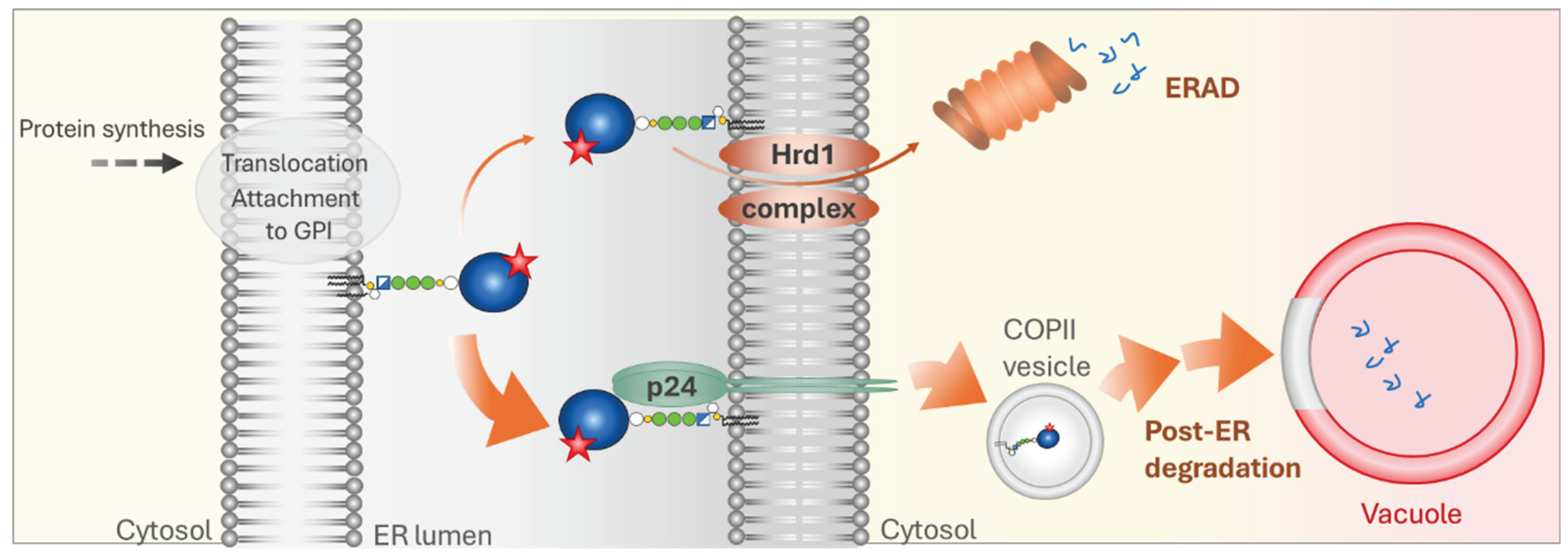
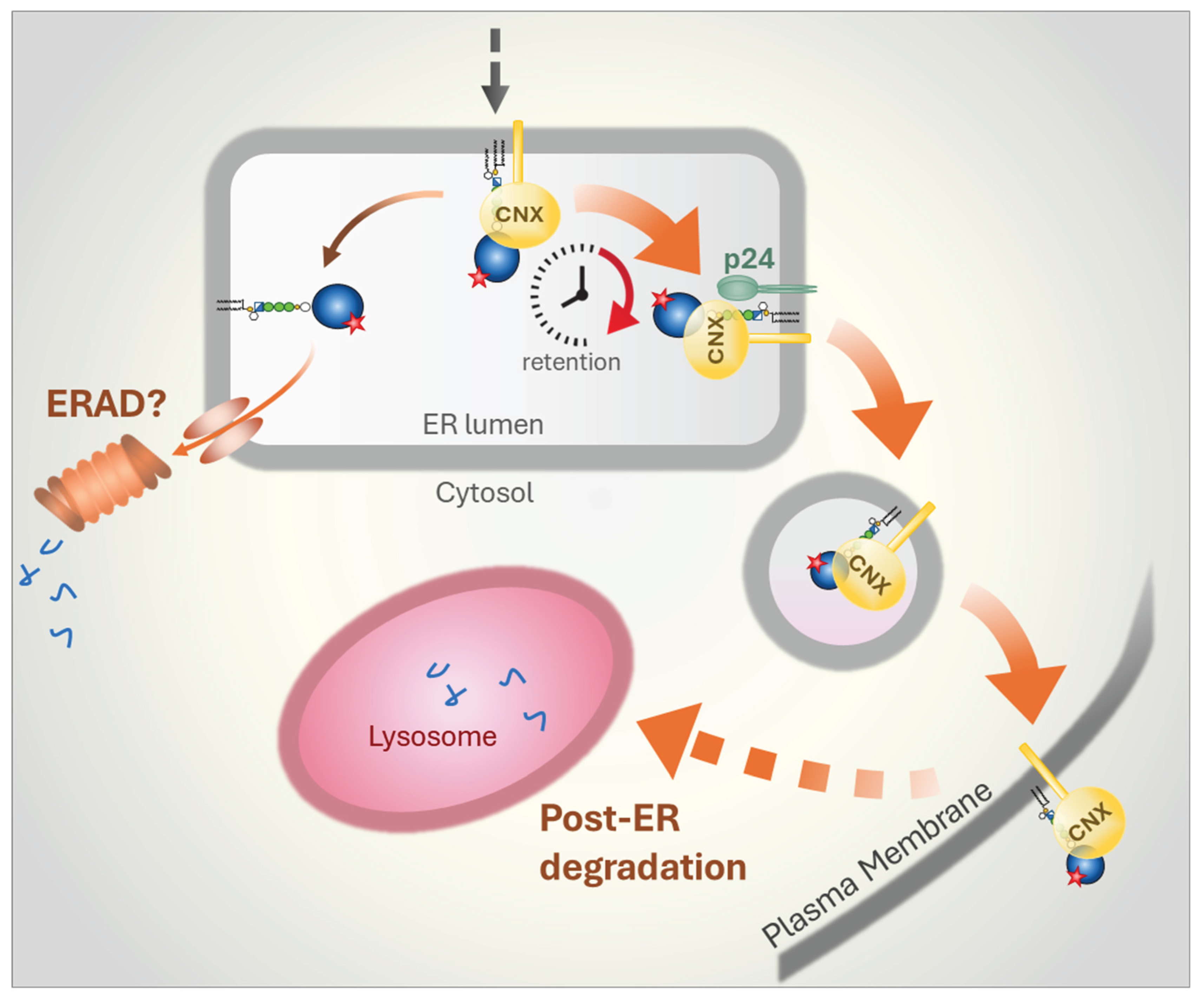
5.2. Mammalian Cells
5.3. Plants
5.4. Protozoa
6. Summary and Outlook
Acknowledgments
Conflicts of Interest
References
- O. Nosjean, A. Briolay, and B. Roux, ‘Mammalian GPI proteins: sorting, membrane residence and functions’, Biochim Biophys Acta, vol. 1331, no. 2, pp. 153–186, Sep. 1997. [CrossRef]
- P. Orlean and A. K. Menon, ‘Thematic review series: lipid posttranslational modifications. GPI anchoring of protein in yeast and mammalian cells, or: how we learned to stop worrying and love glycophospholipids’, J Lipid Res, vol. 48, no. 5, pp. 993–1011, May 2007. [CrossRef]
- T. Kinoshita, ‘Biosynthesis and biology of mammalian GPI-anchored proteins’, vol. 10, no. 3, Mar. 2020. [CrossRef]
- T. Kinoshita, ‘Towards a thorough understanding of mammalian glycosylphosphatidylinositol-anchored protein biosynthesis’, Glycobiology, vol. 34, no. 11, Sep. 2024. [CrossRef]
- R. Watanabe et al., ‘Initial enzyme for glycosylphosphatidylinositol biosynthesis requires PIG-P and is regulated by DPM2’, EMBO J, vol. 19, no. 16, pp. 4402–4411, Aug. 2000. [CrossRef]
- R. Watanabe, K. Ohishi, Y. Maeda, N. Nakamura, and T. Kinoshita, ‘Mammalian PIG-L and its yeast homologue Gpi12p are N-acetylglucosaminylphosphatidylinositol de-N-acetylases essential in glycosylphosphatidylinositol biosynthesis’, Biochem J, vol. 339 ( Pt 1), no. Pt 1, pp. 185–192, Apr. 1999. [CrossRef]
- Y. Wang et al., ‘Genome-wide CRISPR screen reveals CLPTM1L as a lipid scramblase required for efficient glycosylphosphatidylinositol biosynthesis’, Proc Natl Acad Sci U S A, vol. 119, no. 14, Apr. 2022. [CrossRef]
- Y. Murakami et al., ‘PIG-W is critical for inositol acylation but not for flipping of glycosylphosphatidylinositol-anchor’, Mol Biol Cell, vol. 14, no. 10, pp. 4285–4295, Oct. 2003. [CrossRef]
- N. Jiménez-Rojo and H. Riezman, ‘On the road to unraveling the molecular functions of ether lipids’, FEBS Lett, vol. 593, no. 17, pp. 2378–2389, Sep. 2019. [CrossRef]
- Y. Xu et al., ‘Structures of liganded glycosylphosphatidylinositol transamidase illuminate GPI-AP biogenesis’, Nature Communications 2023 14:1, vol. 14, no. 1, pp. 1–17, Sep. 2023. [CrossRef]
- D. Li, ‘Structure and Function of the Glycosylphosphatidylinositol Transamidase, a Transmembrane Complex Catalyzing GPI Anchoring of Proteins’, Subcell Biochem, vol. 104, pp. 425–458, 2024. [CrossRef]
- S. Mayor, A. K. Menon, and G. A. M. Cross, ‘Transfer of glycosyl-phosphatidylinositol membrane anchors to polypeptide acceptors in a cell-free system’, J Cell Biol, vol. 114, no. 1, pp. 61–71, 1991. [CrossRef]
- S. E. Maxwell, S. Ramalingam, L. D. Gerber, L. Brink, and S. Udenfriend, ‘An active carbonyl formed during glycosylphosphatidylinositol addition to a protein is evidence of catalysis by a transamidase’, J Biol Chem, vol. 270, no. 33, pp. 19576–19582, 1995. [CrossRef]
- D. K. Sharma, J. Vidugiriene, J. D. Bangs, and A. K. Menon, ‘A cell-free assay for glycosylphosphatidylinositol anchoring in African trypanosomes. Demonstration of a transamidation reaction mechanism’, J Biol Chem, vol. 274, no. 23, pp. 16479–16486, Jun. 1999. [CrossRef]
- S. J. Grimme, B. A. Westfall, J. M. Wiedman, C. H. Taron, and P. Orlean, ‘The essential Smp3 protein is required for addition of the side-branching fourth mannose during assembly of yeast glycosylphosphatidylinositols’, J Biol Chem, vol. 276, no. 29, pp. 27731–27739, Jul. 2001. [CrossRef]
- N. Stahl, M. A. Baldwin, R. Hecker, K. M. Pan, A. L. Burlingame, and S. B. Prusiner, ‘Glycosylinositol phospholipid anchors of the scrapie and cellular prion proteins contain sialic acid’, Biochemistry, vol. 31, no. 21, pp. 5043–5053, Feb. 1992. [CrossRef]
- M. Ishida et al., ‘Ethanolamine-phosphate on the second mannose is a preferential bridge for some GPI-anchored proteins’, EMBO Rep, vol. 23, no. 7, Jul. 2022. [CrossRef]
- M. D. Resh, ‘Trafficking and signaling by fatty-acylated and prenylated proteins’, Nat Chem Biol, vol. 2, no. 11, pp. 584–590, 2006. [CrossRef]
- M. E. Linder and R. J. Deschenes, ‘Palmitoylation: policing protein stability and traffic’, Nat Rev Mol Cell Biol, vol. 8, no. 1, pp. 74–84, Jan. 2007. [CrossRef]
- K. Simons and D. Toomre, ‘Lipid rafts and signal transduction’, Nat Rev Mol Cell Biol, vol. 1, no. 1, pp. 31–39, 2000. [CrossRef]
- J. B. Helms and C. Zurzolo, ‘Lipids as targeting signals: lipid rafts and intracellular trafficking’, Traffic, vol. 5, no. 4, pp. 247–254, Apr. 2004. [CrossRef]
- S. Sasaki et al., ‘Protein sorting upon exit from the endoplasmic reticulum dominates Golgi biogenesis in budding yeast’, FEBS Lett, vol. 598, no. 5, pp. 548–555, Mar. 2024. [CrossRef]
- M. S. Vogt, G. F. Schmitz, D. V. Silva, H. U. Mösch, and L. O. Essen, ‘Structural base for the transfer of GPI-anchored glycoproteins into fungal cell walls’, Proc Natl Acad Sci U S A, vol. 117, no. 36, pp. 22061–22067, Sep. 2020. [CrossRef]
- A. Schwede and M. Carrington, ‘Bloodstream form Trypanosome plasma membrane proteins: antigenic variation and invariant antigens’, Parasitology, vol. 137, no. 14, pp. 2029–2039, Dec. 2010. [CrossRef]
- M. A. Davitz, D. Hereld, S. Shak, J. Krakow, P. T. Englund, and V. Nussenzweig, ‘A glycan-phosphatidylinositol-specific phospholipase D in human serum’, Science, vol. 238, no. 4823, pp. 81–84, 1987. [CrossRef]
- G. Kondoh et al., ‘Angiotensin-converting enzyme is a GPI-anchored protein releasing factor crucial for fertilization’, Nat Med, vol. 11, no. 2, pp. 160–166, Feb. 2005. [CrossRef]
- K. Watanabe et al., ‘Growth factor induction of Cripto-1 shedding by glycosylphosphatidylinositol-phospholipase D and enhancement of endothelial cell migration’, J Biol Chem, vol. 282, no. 43, pp. 31643–31655, Oct. 2007. [CrossRef]
- D. Corda, M. G. Mosca, N. Ohshima, L. Grauso, N. Yanaka, and S. Mariggiò, ‘The emerging physiological roles of the glycerophosphodiesterase family’, FEBS J, vol. 281, no. 4, pp. 998–1016, Feb. 2014. [CrossRef]
- Y. Fujihara and M. Ikawa, ‘GPI-AP release in cellular, developmental, and reproductive biology’, J Lipid Res, vol. 57, no. 4, pp. 538–545, Apr. 2016. [CrossRef]
- M. A. Wulf, A. Senatore, and A. Aguzzi, ‘The biological function of the cellular prion protein: an update’, BMC Biology 2017 15:1, vol. 15, no. 1, pp. 34-, May 2017. [CrossRef]
- E. De Cecco and G. Legname, ‘The role of the prion protein in the internalization of α-synuclein amyloids’, Prion, vol. 12, no. 1, pp. 23–27, Jan. 2018. [CrossRef]
- A. R. Castle and D. Westaway, ‘Prion Protein Endoproteolysis: Cleavage Sites, Mechanisms and Connections to Prion Disease’, J Neurochem, vol. 169, no. 1, p. e16310, Jan. 2025. [CrossRef]
- B. Caughey et al., ‘Prions and protein aggregates as pathogens, self-propagating structures, biomarkers, and therapeutic targets’, Microbiology and Molecular Biology Reviews, Sep. 2025. [CrossRef]
- M. Zahra, A. Idris, M. Q. Wei, N. A. J. McMillan, and A. L. Munn, ‘Unmasking the tumorigenic potential of cellular prion protein in cancer progression’, Biochim Biophys Acta Mol Basis Dis, vol. 1872, no. 1, Jan. 2026. [CrossRef]
- S. B. Prusiner, ‘Molecular biology of prions causing infectious and genetic encephalopathies of humans as well as scrapie of sheep and BSE of cattle.’, Dev Biol Stand, vol. 75, pp. 55–74, 1991.
- E. Dvorakova et al., ‘Detection of the GPI-anchorless prion protein fragment PrP226* in human brain’, BMC Neurol, vol. 13, Sep. 2013. [CrossRef]
- G. Zanusso et al., ‘Gerstmann-sträussler-scheinker disease and anchorless prion protein mice share prion conformational properties diverging from sporadic creutzfeldt-jakob disease’, Journal of Biological Chemistry, vol. 289, no. 8, pp. 4870–4881, Feb. 2014. [CrossRef]
- P. Shen et al., ‘Characterization of Anchorless Human PrP With Q227X Stop Mutation Linked to Gerstmann-Sträussler-Scheinker Syndrome In Vivo and In Vitro’, Mol Neurobiol, vol. 58, no. 1, pp. 21–33, Jan. 2021. [CrossRef]
- S. Li, Q. Tang, Y. Jiang, and X. Chen, ‘Inherited glycosylphosphatidylinositol deficiency: a review from molecular and clinical perspectives’, Acta Biochim Biophys Sin (Shanghai), vol. 56, no. 8, pp. 1234–1243, Aug. 2024. [CrossRef]
- Y. Murakami et al., ‘Mutations in PIGB Cause an Inherited GPI Biosynthesis Defect with an Axonal Neuropathy and Metabolic Abnormality in Severe Cases’, Am J Hum Genet, vol. 105, no. 2, pp. 384–394, Aug. 2019. [CrossRef]
- Y. Murakami and T. Kinoshita, ‘Inherited GPI deficiencies: A new disease with intellectual disability and epilepsy’, No To Hattatsu, vol. 47, no. 1, pp. 5–13, Jan. 2015.
- K. Bellai-Dussault, T. T. M. Nguyen, N. V. Baratang, D. A. Jimenez-Cruz, and P. M. Campeau, ‘Clinical variability in inherited glycosylphosphatidylinositol deficiency disorders’, Clin Genet, vol. 95, no. 1, pp. 112–121, Jan. 2019. [CrossRef]
- J. Sidpra et al., ‘The clinical and genetic spectrum of inherited glycosylphosphatidylinositol deficiency disorders’, Brain, vol. 147, no. 8, pp. 2775–2790, Aug. 2024. [CrossRef]
- A. Hill, A. E. Dezern, T. Kinoshita, and R. A. Brodsky, ‘Paroxysmal nocturnal haemoglobinuria’, Nat Rev Dis Primers, vol. 3, May 2017. [CrossRef]
- D. G. Gamage and T. L. Hendrickson, ‘Gpi transamidase and gpi anchored proteins: Oncogenes and biomarkers for cancer’, Crit Rev Biochem Mol Biol, vol. 48, no. 5, pp. 446–464, Sep. 2013. [CrossRef]
- P. R. Gilson et al., ‘Identification and stoichiometry of glycosylphosphatidylinositol-anchored membrane proteins of the human malaria parasite Plasmodium falciparum’, Mol Cell Proteomics, vol. 5, no. 7, pp. 1286–1299, 2006. [CrossRef]
- F. Kamena et al., ‘Synthetic GPI array to study antitoxic malaria response’, Nat Chem Biol, vol. 4, no. 4, pp. 238–240, 2008.
- R. Nagar et al., ‘The major surface protein of malaria sporozoites is GPI-anchored to the plasma membrane’, Journal of Biological Chemistry, vol. 300, no. 8, Aug. 2024. [CrossRef]
- M. A. J. Ferguson, ‘The structure, biosynthesis and functions of glycosylphosphatidylinositol anchors, and the contributions of trypanosome research’, J Cell Sci, vol. 112, no. 17, pp. 2799–2809, 1999. [CrossRef]
- J. Lukeš, D. Speijer, A. Zíková, J. D. Alfonzo, H. Hashimi, and M. C. Field, ‘Trypanosomes as a magnifying glass for cell and molecular biology’, Trends Parasitol, vol. 39, no. 11, pp. 902–912, Nov. 2023. [CrossRef]
- L. Lemus, R. S. Hegde, and V. Goder, ‘New frontiers in quality control: the case of GPI-anchored proteins’, Nat Rev Mol Cell Biol, May 2023. [CrossRef]
- L. Ellgaard and A. Helenius, ‘Quality control in the endoplasmic reticulum’, Nat Rev Mol Cell Biol, vol. 4, no. 3, pp. 181–191, 2003.
- R. Strasser, ‘Protein Quality Control in the Endoplasmic Reticulum of Plants’, Annu Rev Plant Biol, vol. 69, pp. 147–172, Apr. 2018. [CrossRef]
- R. L. Wiseman, J. S. Mesgarzadeh, and L. M. Hendershot, ‘Reshaping endoplasmic reticulum quality control through the unfolded protein response’, Mol Cell, vol. 82, no. 8, pp. 1477–1491, Apr. 2022. [CrossRef]
- L. Krshnan, M. L. van de Weijer, and P. Carvalho, ‘Endoplasmic Reticulum-Associated Protein Degradation’, Cold Spring Harb Perspect Biol, vol. 14, no. 12, 2022. [CrossRef]
- J. C. Christianson, E. Jarosch, and T. Sommer, ‘Mechanisms of substrate processing during ER-associated protein degradation’, Nat Rev Mol Cell Biol, vol. 24, no. 11, pp. 777–796, Nov. 2023. [CrossRef]
- I. Fregno and M. Molinari, ‘Proteasomal and lysosomal clearance of faulty secretory proteins: ER-associated degradation (ERAD) and ER-to-lysosome-associated degradation (ERLAD) pathways’, Crit Rev Biochem Mol Biol, vol. 54, no. 2, pp. 153–163, Mar. 2019. [CrossRef]
- S. Schwabl and D. Teis, ‘Protein quality control at the Golgi’, Curr Opin Cell Biol, vol. 75, Apr. 2022. [CrossRef]
- E. Hong, A. R. Davidson, and C. A. Kaiser, ‘A pathway for targeting soluble misfolded proteins to the yeast vacuole’, J Cell Biol, vol. 135, no. 3, pp. 623–633, 1996.
- S. Wang and D. T. Ng, ‘Evasion of endoplasmic reticulum surveillance makes Wsc1p an obligate substrate of Golgi quality control’, Mol Biol Cell, vol. 21, no. 7, pp. 1153–1165, 2010.
- O. Schmidt et al., ‘ Endosome and Golgi-associated degradation ( EGAD ) of membrane proteins regulates sphingolipid metabolism ’, EMBO J, vol. 38, no. 15, Aug. 2019. [CrossRef]
- A. Eisenberg-Lerner et al., ‘Golgi organization is regulated by proteasomal degradation’, Nat Commun, vol. 11, no. 1, Dec. 2020. [CrossRef]
- A. Ruggiano, O. Foresti, and P. Carvalho, ‘ER-associated degradation: Protein quality control and beyond’, 2014. [CrossRef]
- V. Goder, E. Alanis-Dominguez, and M. Bustamante-Sequeiros, ‘Lipids and their (un)known effects on ER-associated protein degradation (ERAD)’, Biochimica et Biophysica Acta (BBA) - Molecular and Cell Biology of Lipids, Jun. 2019. [CrossRef]
- D. Kumari and J. L. Brodsky, ‘The targeting of native proteins to the endoplasmic reticulum-associated degradation (Erad) pathway: An expanding repertoire of regulated substrates’, Biomolecules, vol. 11, no. 8, Aug. 2021. [CrossRef]
- S. Vashist, W. Kim, W. J. Belden, E. D. Spear, C. Barlowe, and D. T. Ng, ‘Distinct retrieval and retention mechanisms are required for the quality control of endoplasmic reticulum protein folding’, J Cell Biol, vol. 155, no. 3, pp. 355–368, 2001.
- D. Hellerschmied, Y. V. Serebrenik, L. Shao, G. M. Burslem, and C. M. Crews, ‘Protein folding state-dependent sorting at the Golgi apparatus’, Mol Biol Cell, vol. 30, no. 17, pp. 2296–2308, Aug. 2019. [CrossRef]
- W. M. Henne, N. J. Buchkovich, and S. D. Emr, ‘The ESCRT pathway’, Dev Cell, vol. 21, no. 1, pp. 77–91, Jul. 2011. [CrossRef]
- M. U. Jørgensen, S. D. Emr, and J. R. Winther, ‘Ligand recognition and domain structure of Vps10p, a vacuolar protein sorting receptor in Saccharomyces cerevisiae’, Eur J Biochem, vol. 260, no. 2, pp. 461–469, Mar. 1999. [CrossRef]
- C. L. Gelling, I. W. Dawes, D. H. Perlmutter, E. A. Fisher, and J. L. Brodsky, ‘The Endosomal protein-sorting receptor sortilin has a role in trafficking α-1 antitrypsin’, Genetics, vol. 192, no. 3, pp. 889–903, Nov. 2012. [CrossRef]
- I. Fitzgerald and B. S. Glick, ‘Secretion of a foreign protein from budding yeasts is enhanced by cotranslational translocation and by suppression of vacuolar targeting’, Microb Cell Fact, vol. 13, no. 1, Aug. 2014. [CrossRef]
- L. Lemus, Z. Matić, L. Gal, A. Fadel, M. Schuldiner, and V. Goder, ‘Post-ER degradation of misfolded GPI-anchored proteins is linked with microautophagy’, Curr Biol, vol. 31, no. 18, pp. 4025-4037.e5, Sep. 2021. [CrossRef]
- C. MacDonald, J. A. Payne, M. Aboian, W. Smith, D. J. Katzmann, and R. C. Piper, ‘A Family of Tetraspans Organizes Cargo for Sorting into Multivesicular Bodies’, Dev Cell, vol. 33, no. 3, pp. 328–342, May 2015. [CrossRef]
- F. Schimmoller, B. Singer-Kruger, S. Schroder, U. Kruger, C. Barlowe, and H. Riezman, ‘The absence of Emp24p, a component of ER-derived COPII-coated vesicles, causes a defect in transport of selected proteins to the Golgi’, Embo J, vol. 14, no. 7, pp. 1329–1339, 1995.
- M. Muniz, C. Nuoffer, H. P. Hauri, and H. Riezman, ‘The Emp24 complex recruits a specific cargo molecule into endoplasmic reticulum-derived vesicles’, J Cell Biol, vol. 148, no. 5, pp. 925–930, 2000.
- S. Takida, Y. Maeda, and T. Kinoshita, ‘Mammalian GPI-anchored proteins require p24 proteins for their efficient transport from the ER to the plasma membrane’, Biochemical Journal, vol. 409, no. 2, pp. 555–562, Jan. 2008. [CrossRef]
- S. Lopez et al., ‘Dual independent roles of the p24 complex in selectivity of secretory cargo export from the endoplasmic reticulum’, Cells, vol. 9, no. 5, May 2020. [CrossRef]
- C. Bernat-Silvestre et al., ‘P24 Family proteins are involved in transport to the plasma membrane of gpi-anchored proteins in plants’, Plant Physiol, vol. 184, no. 3, pp. 1333–1347, Nov. 2020. [CrossRef]
- Z. Ji, R. Nagar, S. M. Duncan, M. L. Sampaio Guther, and M. A. J. Ferguson, ‘Identification of the glycosylphosphatidylinositol-specific phospholipase A2 (GPI-PLA2) that mediates GPI fatty acid remodeling in Trypanosoma brucei’, Journal of Biological Chemistry, vol. 299, no. 8, Aug. 2023. [CrossRef]
- S. Tanaka, Y. Maeda, Y. Tashima, and T. Kinoshita, ‘Inositol deacylation of glycosylphosphatidylinositol-anchored proteins is mediated by mammalian PGAP1 and yeast Bst1p’, J Biol Chem, vol. 279, no. 14, pp. 14256–14263, Apr. 2004. [CrossRef]
- C. Bernat-Silvestre et al., ‘AtPGAP1 functions as a GPI inositol-deacylase required for efficient transport of GPI-anchored proteins’, Plant Physiol, vol. 187, no. 4, pp. 2156–2173, Dec. 2021. [CrossRef]
- W. L. Roberts, J. J. Myher, A. Kuksis, M. G. Low, and T. L. Rosenberry, ‘Lipid analysis of the glycoinositol phospholipid membrane anchor of human erythrocyte acetylcholinesterase. Palmitoylation of inositol results in resistance to phosphatidylinositol-specific phospholipase C’, J Biol Chem, vol. 263, no. 35, pp. 18766–18775, 1988. [CrossRef]
- M. Fujita, M. Umemura, T. Yoko-o, and Y. Jigami, ‘PER1 Is Required for GPI-Phospholipase A(2) Activity and Involved in Lipid Remodeling of GPI-anchored Proteins’, Mol Biol Cell, vol. 17, no. 12, pp. 5253–5264, 2006.
- R. Bosson, M. Jaquenoud, and A. Conzelmann, ‘GUP1 of Saccharomyces cerevisiae Encodes an O-Acyltransferase Involved in Remodeling of the GPI Anchor’, Mol Biol Cell, vol. 17, no. 6, pp. 2636–2645, 2006.
- M. Umemura, M. Fujita, O. T. Yoko, A. Fukamizu, and Y. Jigami, ‘Saccharomyces cerevisiae CWH43 is involved in the remodeling of the lipid moiety of GPI anchors to ceramides’, Mol Biol Cell, vol. 18, no. 11, pp. 4304–4316, 2007.
- T. Kinoshita and M. Fujita, ‘Biosynthesis of GPI-anchored proteins: special emphasis on GPI lipid remodeling’, J Lipid Res, vol. 57, no. 1, pp. 6–24, Jan. 2016. [CrossRef]
- M. Fujita, Y. Maeda, M. Ra, Y. Yamaguchi, R. Taguchi, and T. Kinoshita, ‘GPI glycan remodeling by PGAP5 regulates transport of GPI-anchored proteins from the ER to the Golgi’, Cell, vol. 139, no. 2, pp. 352–365, 2009.
- J. Manzano-Lopez et al., ‘COPII Coat Composition Is Actively Regulated by Luminal Cargo Maturation’, Curr Biol, vol. 25, no. 2, pp. 152–162, 2015.
- M. Fujita et al., ‘Sorting of GPI-anchored proteins into ER exit sites by p24 proteins is dependent on remodeled GPI’, J Cell Biol, vol. 194, no. 1, pp. 61–75, 2011.
- G. A. Castillon et al., ‘The yeast p24 complex regulates GPI-anchored protein transport and quality control by monitoring anchor remodeling’, Mol Biol Cell, vol. 22, no. 16, pp. 2924–2936, 2011.
- S. Rodriguez-Gallardo et al., ‘Ceramide chain length-dependent protein sorting into selective endoplasmic reticulum exit sites.’, Sci Adv, vol. 6, no. 50, pp. 8237–8248, Dec. 2020. [CrossRef]
- S. Rodriguez-Gallardo et al., ‘Quality-controlled ceramide-based GPI-anchored protein sorting into selective ER exit sites’, Cell Rep, vol. 39, no. 5, May 2022. [CrossRef]
- H. Hirayama, M. Fujita, T. Yoko-o, and Y. Jigami, ‘O-mannosylation is required for degradation of the endoplasmic reticulum-associated degradation substrate Gas1*p via the ubiquitin/proteasome pathway in Saccharomyces cerevisiae’, J Biochem, vol. 143, no. 4, pp. 555–567, 2008.
- A. Ashok and R. S. Hegde, ‘Retrotranslocation of prion proteins from the endoplasmic reticulum by preventing GPI signal transamidation’, Mol Biol Cell, vol. 19, no. 8, pp. 3463–3476, 2008.
- P. Satpute-Krishnan, M. Ajinkya, S. Bhat, E. Itakura, R. S. Hegde, and J. Lippincott-Schwartz, ‘ER stress-induced clearance of misfolded GPI-anchored proteins via the secretory pathway’, Cell, vol. 158, no. 3, pp. 522–533, 2014.
- N. Sikorska et al., ‘Limited ER quality control for GPI-anchored proteins’, J Cell Biol, vol. 213, no. 6, pp. 693–704, 2016.
- A. Ashok and R. S. Hegde, ‘Selective Processing and Metabolism of Disease-Causing Mutant Prion Proteins’, PLoS Pathog, vol. 5, no. 6, p. e1000479, Jun. 2009. [CrossRef]
- V. Goder and A. Melero, ‘Protein O-mannosyltransferases participate in ER protein quality control’, J Cell Sci, vol. 124, no. Pt 1, pp. 144–153, 2011.
- M. Fujita, O. T. Yoko, and Y. Jigami, ‘Inositol deacylation by Bst1p is required for the quality control of glycosylphosphatidylinositol-anchored proteins’, Mol Biol Cell, vol. 17, no. 2, pp. 834–850, 2006.
- C. Harty, S. Strahl, and K. Romisch, ‘O-mannosylation protects mutant alpha-factor precursor from endoplasmic reticulum-associated degradation’, Mol Biol Cell, vol. 12, no. 4, pp. 1093–1101, 2001.
- C. Xu, S. Wang, G. Thibault, and D. T. W. Ng, ‘Futile Protein Folding Cycles in the ER Are Terminated by the Unfolded Protein O-Mannosylation Pathway’, Science (1979), vol. 340, no. 6135, pp. 978 LP-- 981, 2013.
- L. Lemus, H. Meyer, A. I. Rodríguez-Rosado, M. Schuldiner, and V. Goder, ‘O-mannosylation of misfolded ER proteins promotes ERAD’, EMBO J, vol. in press, Nov. 2025. [CrossRef]
- A. Uttenweiler, H. Schwarz, H. Neumann, and A. Mayer, ‘The Vacuolar Transporter Chaperone (VTC) Complex Is Required for Microautophagy’, Mol Biol Cell, vol. 18, no. 1, pp. 166–175, Jan. 2007. [CrossRef]
- C. W. Wang, Y. H. Miao, and Y. S. Chang, ‘A sterol-enriched vacuolar microdomain mediates stationary phase lipophagy in budding yeast’, Journal of Cell Biology, vol. 206, no. 3, pp. 357–366, 2014. [CrossRef]
- T. Tsuji et al., ‘Niemann-Pick type C proteins promote microautophagy by expanding raft-like membrane domains in the yeast vacuole’, Elife, vol. 6, Jun. 2017. [CrossRef]
- L. Lemus and V. Goder, ‘Pep4-dependent microautophagy is required for post-ER degradation of GPI-anchored proteins’, Autophagy, vol. 18, no. 1, pp. 223–225, 2022. [CrossRef]
- G. G. Kovács, G. Trabattoni, J. A. Hainfellner, J. W. Ironside, R. S. G. Knight, and H. Budka, ‘Mutations of the prion protein gene phenotypic spectrum’, J Neurol, vol. 249, no. 11, pp. 1567–1582, 2002. [CrossRef]
- A. Ruiz-Argüelles and L. Llorente, ‘The role of complement regulatory proteins (CD55 and CD59) in the pathogenesis of autoimmune hemocytopenias’, Autoimmun Rev, vol. 6, no. 3, pp. 155–161, Jan. 2007. [CrossRef]
- B. Puig, H. Altmeppen, and M. Glatzel, ‘The GPI-anchoring of PrP: implications in sorting and pathogenesis’, Prion, vol. 8, no. 1, pp. 11–18, 2014.
- J. Ma and S. Lindquist, ‘Wild-type PrP and a mutant associated with prion disease are subject to retrograde transport and proteasome degradation’, Proc Natl Acad Sci U S A, vol. 98, no. 26, pp. 14955–14960, 2001.
- Y. Yedidia, L. Horonchik, S. Tzaban, A. Yanai, and A. Taraboulos, ‘Proteasomes and ubiquitin are involved in the turnover of the wild-type prion protein’, Embo J, vol. 20, no. 19, pp. 5383–5391, 2001.
- G. Petris et al., ‘CD4 and BST-2/tetherin proteins retro-translocate from endoplasmic reticulum to cytosol as partially folded and multimeric molecules’, J Biol Chem, vol. 289, no. 1, pp. 1–12, 2014.
- Y. J. Wang et al., ‘The association of the vanin-1 N131S variant with blood pressure is mediated by endoplasmic reticulum-associated degradation and loss of function’, PLoS Genet, vol. 10, no. 9, p. e1004641, 2014.
- B. Drisaldi et al., ‘Mutant PrP is delayed in its exit from the endoplasmic reticulum, but neither wild-type nor mutant PrP undergoes retrotranslocation prior to proteasomal degradation’, J Biol Chem, vol. 278, no. 24, pp. 21732–21743, 2003.
- Y.-S. S. Liu et al., ‘N -Glycan–dependent protein folding and endoplasmic reticulum retention regulate GPI-anchor processing’, J Cell Biol, vol. 217, no. 2, pp. 585–599, Feb. 2018. [CrossRef]
- X. Y. Guo, Y. S. Liu, X. D. Gao, T. Kinoshita, and M. Fujita, ‘Calnexin mediates the maturation of GPI-anchors through ER retention’, vol. 295, no. 48, pp. 16393–16410, Nov. 2020.
- A. M. Cheatham, N. R. Sharma, and P. Satpute-Krishnan, ‘Competition for calnexin binding regulates secretion and turnover of misfolded GPI-anchored proteins’, Journal of Cell Biology, vol. 222, no. 10, Oct. 2023. [CrossRef]
- J. Hong et al., ‘Molecular basis of the inositol deacylase PGAP1 involved in quality control of GPI-AP biogenesis’, Nat Commun, vol. 15, no. 1, Dec. 2024. [CrossRef]
- P. Satpute-Krishnan, M. Ajinkya, S. Bhat, E. Itakura, R. S. Hegde, and J. Lippincott-Schwartz, ‘ER stress-induced clearance of misfolded GPI-anchored proteins via the secretory pathway’, Cell, vol. 158, no. 3, pp. 522–533, 2014, [Online]. Available: http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Citation&list_uids=25083867.
- C. Stordeur, K. Puth, J. P. Saenz, and R. Ernst, ‘Crosstalk of lipid and protein homeostasis to maintain membrane function’, Biol Chem, vol. 395, no. 3, pp. 313–326, 2014.
- Y. Wang et al., ‘Cross-talks of glycosylphosphatidylinositol biosynthesis with glycosphingolipid biosynthesis and ER-associated degradation’, Nat Commun, vol. 11, no. 1, Dec. 2020. [CrossRef]
- C. Hetz and C. Soto, ‘Stressing out the ER: a role of the unfolded protein response in prion-related disorders’, Curr Mol Med, vol. 6, no. 1, pp. 37–43, Feb. 2006. [CrossRef]
- E. Ronzier and P. Satpute-Krishnan, ‘TMED9 coordinates the clearance of misfolded GPI-anchored proteins out of the ER and into the Golgi’, PLoS Biol, vol. 23, no. 4, Apr. 2025. [CrossRef]
- E. Zavodszky and R. S. Hegde, ‘Misfolded GPI-anchored proteins are escorted through the secretory pathway by ER-derived factors’, Elife, vol. 8, May 2019. [CrossRef]
- N. Desnoyer and R. Palanivelu, ‘Bridging the GAPs in plant reproduction: a comparison of plant and animal GPI-anchored proteins’, Plant Reprod, vol. 33, no. 3–4, pp. 129–142, Dec. 2020. [CrossRef]
- Z. Xu et al., ‘Glycosylphosphatidylinositol anchor lipid remodeling directs proteins to the plasma membrane and governs cell wall mechanics’, Aug. 2022. [CrossRef]
- Y. J. Shin, U. Vavra, and R. Strasser, ‘Proper protein folding in the endoplasmic reticulum is required for attachment of a glycosylphosphatidylinositol anchor in plants’, Plant Physiol, vol. 186, no. 4, pp. 1878–1892, Aug. 2021. [CrossRef]
- M. J. Skelly, ‘Dropping anchor: Stringent quality control prevents GPI anchoring of severely misfolded proteins in plants’, Aug. 01, 2021, Plant Physiol. [CrossRef]
- M. A. McDowell, D. M. Ransom, and J. D. Bangs, ‘Glycosylphosphatidylinositol-dependent secretory transport in Trypanosoma brucei’, Biochem J, vol. 335 ( Pt 3), no. Pt 3, pp. 681–689, Nov. 1998. [CrossRef]
- J. D. Bangs, D. M. Ransom, M. A. McDowell, and E. M. Brouch, ‘Expression of bloodstream variant surface glycoproteins in procyclic stage Trypanosoma brucei: role of GPI anchors in secretion’, EMBO J, vol. 16, no. 14, pp. 4285–4294, Jul. 1997. [CrossRef]
- C. Tiengwe, C. M. Koeller, and J. D. Bangs, ‘Endoplasmic reticulum-associated degradation and disposal of misfolded GPI-anchored proteins in Trypanosoma brucei’, Mol Biol Cell, vol. 29, no. 20, pp. 2397–2409, Oct. 2018. [CrossRef]
- F. Paturiaux-Hanocq et al., ‘A role for the dynamic acylation of a cluster of cysteine residues in regulating the activity of the glycosylphosphatidylinositol-specific phospholipase C of Trypanosoma brucei’, J Biol Chem, vol. 275, no. 16, pp. 12147–12155, Apr. 2000. [CrossRef]
- A. J. Szempruch et al., ‘Extracellular Vesicles from Trypanosoma brucei Mediate Virulence Factor Transfer and Cause Host Anemia’, Cell, vol. 164, no. 1–2, pp. 246–257, Jan. 2016. [CrossRef]
- P. Garrison et al., ‘Turnover of Variant Surface Glycoprotein in Trypanosoma brucei Is a Bimodal Process’, mBio, vol. 12, no. 4, pp. 1725–1746, Aug. 2021. [CrossRef]
- J. D. Bangs, ‘Evolution of Antigenic Variation in African Trypanosomes: Variant Surface Glycoprotein Expression, Structure, and Function’, Bioessays, vol. 40, no. 12, Dec. 2018. [CrossRef]
- K. Halbleib et al., ‘Activation of the Unfolded Protein Response by Lipid Bilayer Stress’, Mol Cell, vol. 67, no. 4, pp. 673-684.e8, Aug. 2017. [CrossRef]
- R. Covino, G. Hummer, and R. Ernst, ‘Integrated Functions of Membrane Property Sensors and a Hidden Side of the Unfolded Protein Response’, Mol Cell, vol. 71, no. 3, pp. 458–467, Aug. 2018. [CrossRef]
- M. C. Jonikas et al., ‘Comprehensive characterization of genes required for protein folding in the endoplasmic reticulum’, Science (1979), vol. 323, no. 5922, pp. 1693–1697, 2009.
- W. J. Masterson, J. Raper, T. L. Doering, G. W. Hart, and P. T. Englund, ‘Fatty acid remodeling: a novel reaction sequence in the biosynthesis of trypanosome glycosyl phosphatidylinositol membrane anchors’, Cell, vol. 62, no. 1, pp. 73–80, Jul. 1990. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



