Submitted:
26 November 2025
Posted:
27 November 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
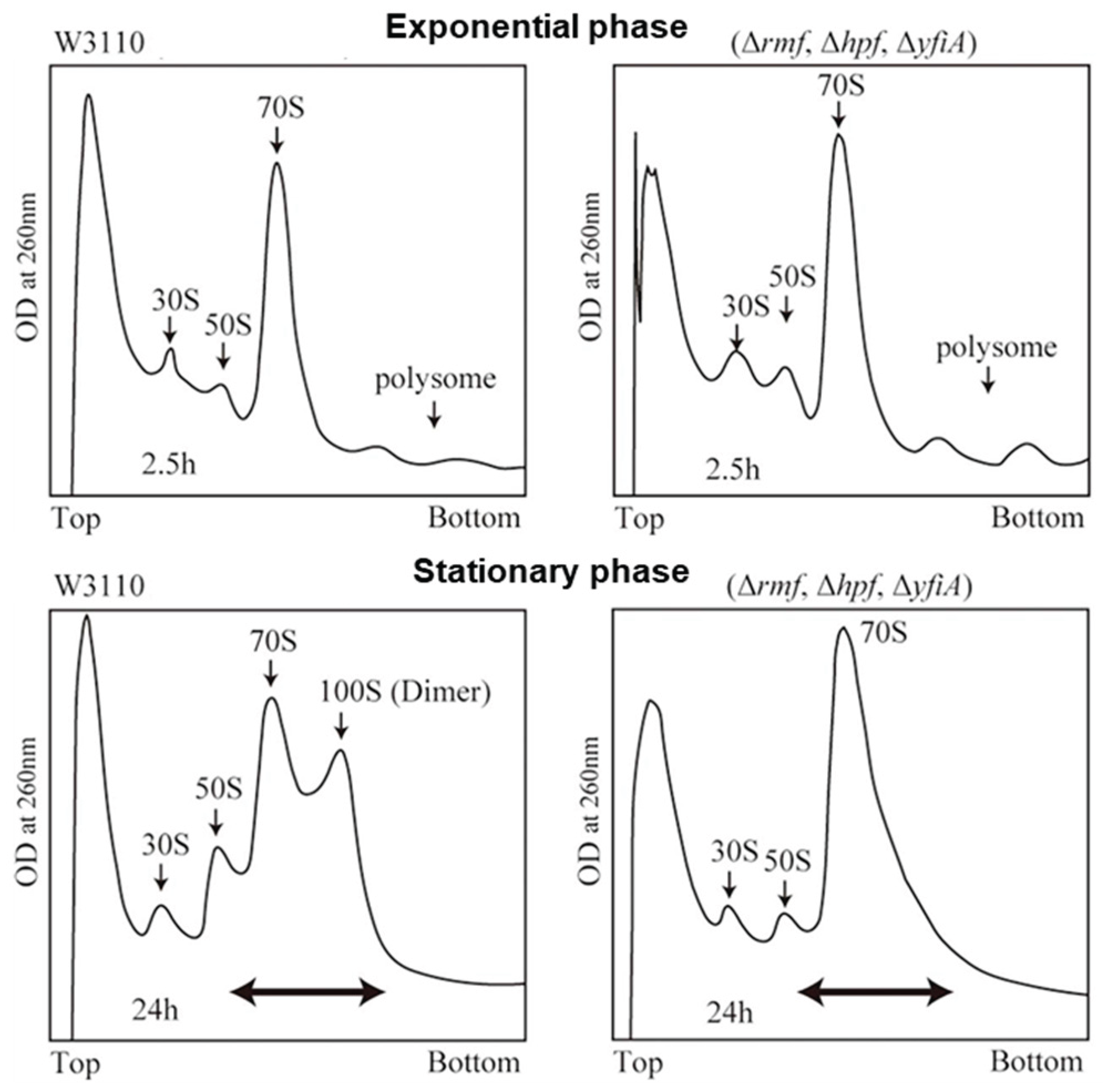
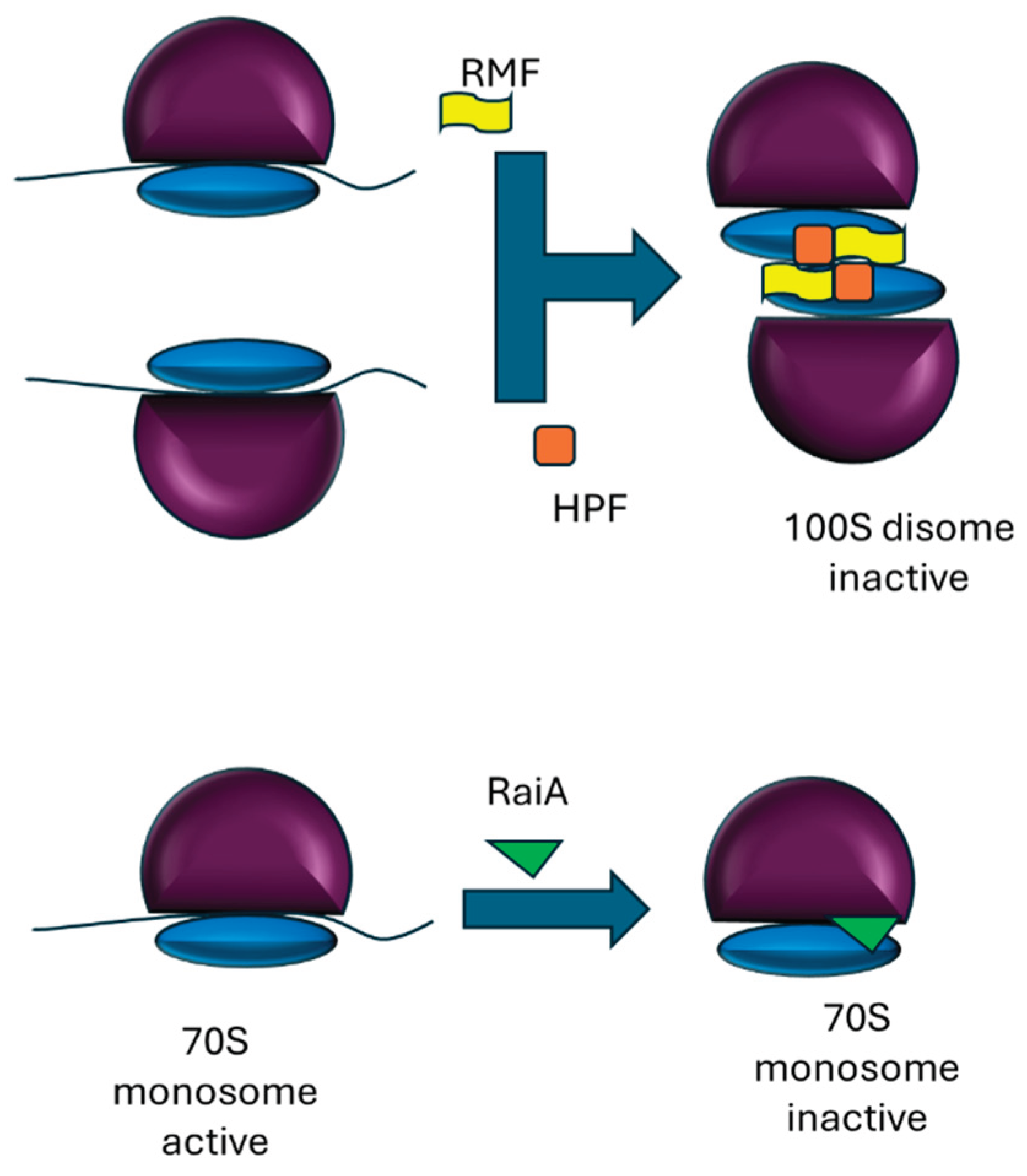
2. Dormant Ribosomes in Prokaryotes and Plant Chloroplasts
3. From Bacteria to Eukaryotes: Challenges to Identifying Vacant Ribosomes
3.1. Protein Interactions with Dormant Ribosomes in Yeasts
3.2. Dimeric Hibernating Ribosomes in Eukaryotes?
3.3. Vacant Monosomes in Metazoans and Mammals
4. Methodological Challenges in Quantifying Processes Related to the Formation of Vacant Ribosomes
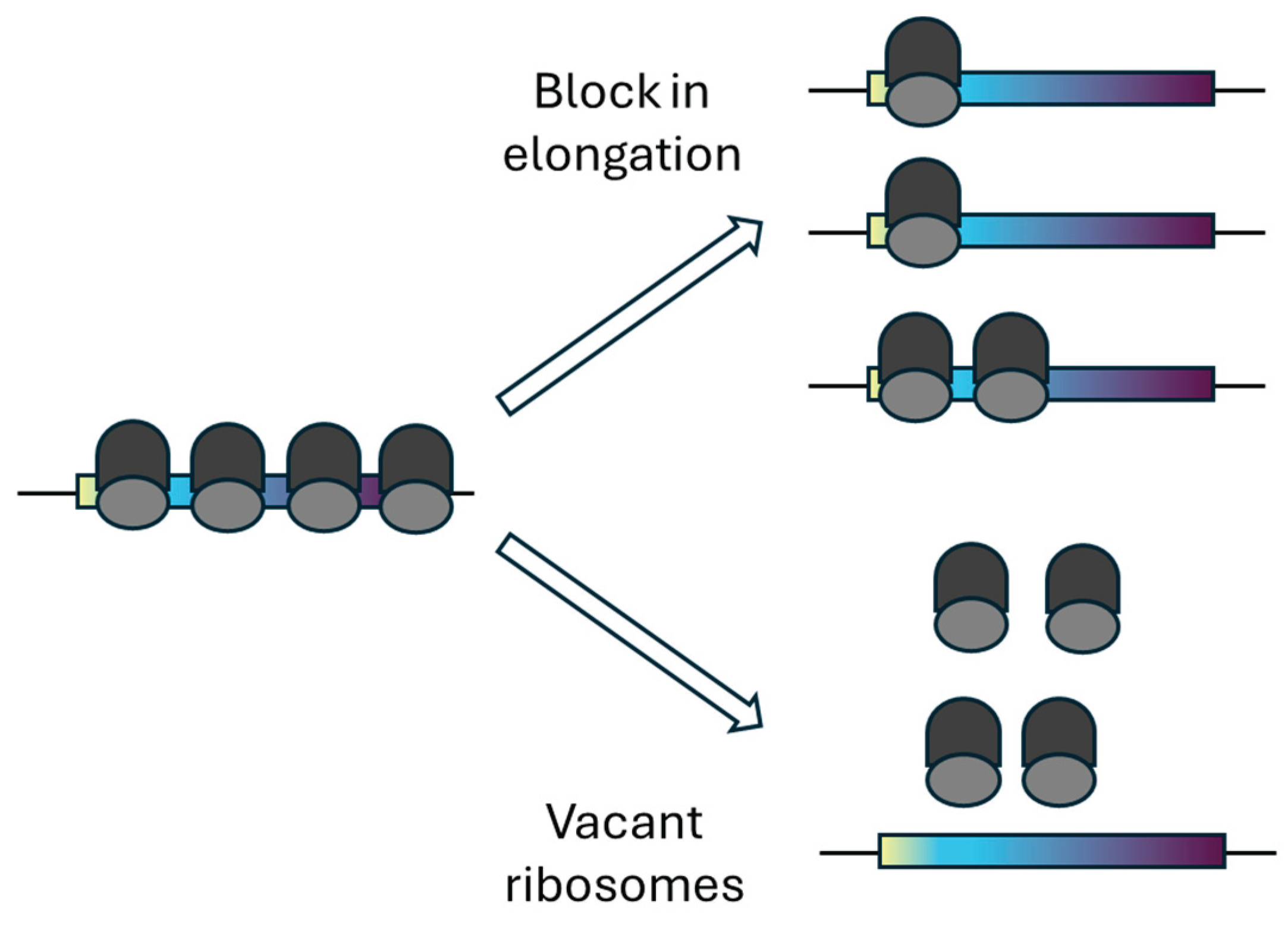
4.1. Distinguishing Vacant Ribosomes from Elongation Blockade
4.2. Quantifying Post-Transcriptional Processes Under Stress
4.3. Stress-Induced Changes in Cytoplasmic Biophysical Properties
5. Biological and Biotechnological Relevance of Dormant and Inactive Ribosomes
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Gregory B, Rahman N, Bommakanti A, Shamsuzzaman M, Thapa M, Lescure A, et al. The small and large ribosomal subunits depend on each other for stability and accumulation. Life Sci Alliance. 2019;2(2).
- Prossliner T, Skovbo Winther K, Sorensen MA, Gerdes K. Ribosome Hibernation. Annu Rev Genet. 2018;52:321–48.
- Prossliner T, Gerdes K, Sorensen MA, Winther KS. Hibernation factors directly block ribonucleases from entering the ribosome in response to starvation. Nucleic Acids Res. 2021;49(4):2226–39.
- Yoshida H, Nakayama H, Maki Y, Ueta M, Wada C, Wada A. Functional Sites of Ribosome Modulation Factor (RMF) Involved in the Formation of 100S Ribosome. Front Mol Biosci. 2021;8:661691.
- Yoshida H, Wada A. The 100S ribosome: ribosomal hibernation induced by stress. Wiley Interdiscip Rev RNA. 2014;5(5):723–32.
- Wada A, Yamazaki Y, Fujita N, Ishihama A. Structure and probable genetic location of a “ribosome modulation factor” associated with 100S ribosomes in stationary-phase Escherichia coli cells. Proc Natl Acad Sci U S A. 1990;87(7):2657–61.
- Koli S, Shetty S. Ribosomal dormancy at the nexus of ribosome homeostasis and protein synthesis. Bioessays. 2024;46(7):e2300247.
- Lang M, Krin E, Korlowski C, Sismeiro O, Varet H, Coppee JY, et al. Sleeping ribosomes: Bacterial signaling triggers RaiA mediated persistence to aminoglycosides. iScience. 2021;24(10):103128.
- Tanaka K, Yoshizawa Y, Oda T, Sekine Y. Chloroplast Hibernation-Promoting Factor PSRP1 Prevents Ribosome Degradation Under Darkness Independently of 100S Dimer Formation. Plants (Basel). 2025;14(20).
- Krokowski D, Gaccioli F, Majumder M, Mullins MR, Yuan CL, Papadopoulou B, et al. Characterization of hibernating ribosomes in mammalian cells. Cell Cycle. 2011;10(16):2691–702.
- McLaren M, Conners R, Isupov MN, Gil-Diez P, Gambelli L, Gold VAM, et al. CryoEM reveals that ribosomes in microsporidian spores are locked in a dimeric hibernating state. Nat Microbiol. 2023;8(10):1834–45.
- Rahaman S, Schiffelholz N, Mittal N, Frohlich KE, Zavolan M, Becskei A. Heat shock induces silent ribosomes and reorganizes mRNA turnover. Cell Rep. 2025;44(10):116447.
- Maki Y, Yoshida H, Wada A. Two proteins, YfiA and YhbH, associated with resting ribosomes in stationary phase Escherichia coli. Genes Cells. 2000;5(12):965–74.
- Van Dyke N, Chanchorn E, Van Dyke MW. The Saccharomyces cerevisiae protein Stm1p facilitates ribosome preservation during quiescence. Biochem Biophys Res Commun. 2013;430(2):745–50.
- Li SH, Li Z, Park JO, King CG, Rabinowitz JD, Wingreen NS, et al. Escherichia coli translation strategies differ across carbon, nitrogen and phosphorus limitation conditions. Nat Microbiol. 2018;3(8):939–47.
- Hofmann S, Cherkasova V, Bankhead P, Bukau B, Stoecklin G. Translation suppression promotes stress granule formation and cell survival in response to cold shock. Mol Biol Cell. 2012;23(19):3786–800.
- Montero-Lomeli M, Morais BL, Figueiredo DL, Neto DC, Martins JR, Masuda CA. The initiation factor eIF4A is involved in the response to lithium stress in Saccharomyces cerevisiae. J Biol Chem. 2002;277(24):21542–8.
- Fresno M, Jimenez A, Vazquez D. Inhibition of translation in eukaryotic systems by harringtonine. Eur J Biochem. 1977;72(2):323–30.
- Shalgi R, Hurt JA, Krykbaeva I, Taipale M, Lindquist S, Burge CB. Widespread Regulation of Translation by Elongation Pausing in Heat Shock. Molecular Cell. 2013;49(3):439–52.
- Liu B, Han Y, Qian SB. Cotranslational response to proteotoxic stress by elongation pausing of ribosomes. Mol Cell. 2013;49(3):453–63.
- Barros GC, Guerrero S, Silva GM. The central role of translation elongation in response to stress. Biochem Soc Trans. 2023;51(3):959–69.
- Navarro IC, Tuorto F, Jordan D, Legrand C, Price J, Braukmann F, et al. Translational adaptation to heat stress is mediated by RNA 5-methylcytosine in Caenorhabditis elegans. EMBO J. 2021;40(6):e105496.
- Goyal A, Belardinelli R, Maracci C, Milon P, Rodnina MV. Directional transition from initiation to elongation in bacterial translation. Nucleic Acids Res. 2015;43(22):10700–12.
- Wang J, Johnson AG, Lapointe CP, Choi J, Prabhakar A, Chen DH, et al. eIF5B gates the transition from translation initiation to elongation. Nature. 2019;573(7775):605–8.
- Ben-Shem A, Garreau de Loubresse N, Melnikov S, Jenner L, Yusupova G, Yusupov M. The structure of the eukaryotic ribosome at 3.0 A resolution. Science. 2011;334(6062):1524–9.
- Van Dyke N, Baby J, Van Dyke MW. Stm1p, a ribosome-associated protein, is important for protein synthesis in Saccharomyces cerevisiae under nutritional stress conditions. J Mol Biol. 2006;358(4):1023–31.
- Balagopal V, Parker R. Polysomes, P bodies and stress granules: states and fates of eukaryotic mRNAs. Curr Opin Cell Biol. 2009;21(3):403–8.
- van den Elzen AM, Schuller A, Green R, Seraphin B. Dom34-Hbs1 mediated dissociation of inactive 80S ribosomes promotes restart of translation after stress. EMBO J. 2014;33(3):265–76.
- Gemin O, Gluc M, Rosa H, Purdy M, Niemann M, Peskova Y, et al. Ribosomes hibernate on mitochondria during cellular stress. Nat Commun. 2024;15(1):8666.
- Martinez MJ, Roy S, Archuletta AB, Wentzell PD, Anna-Arriola SS, Rodriguez AL, et al. Genomic analysis of stationary-phase and exit in Saccharomyces cerevisiae: gene expression and identification of novel essential genes. Mol Biol Cell. 2004;15(12):5295–305.
- Aguilar Rangel M, Stein K, Frydman J. A machine learning approach uncovers principles and determinants of eukaryotic ribosome pausing. Sci Adv. 2024;10(42):eado0738.
- Meydan S, Guydosh NR. A cellular handbook for collided ribosomes: surveillance pathways and collision types. Curr Genet. 2021;67(1):19–26.
- Love AM, Nair NU. Specific codons control cellular resources and fitness. Sci Adv. 2024;10(8):eadk3485.
- Zhao T, Chen YM, Li Y, Wang J, Chen S, Gao N, et al. Disome-seq reveals widespread ribosome collisions that promote cotranslational protein folding. Genome Biol. 2021;22(1):16.
- Anger AM, Armache JP, Berninghausen O, Habeck M, Subklewe M, Wilson DN, et al. Structures of the human and Drosophila 80S ribosome. Nature. 2013;497(7447):80–5.
- Shetty S, Hofstetter J, Battaglioni S, Ritz D, Hall MN. TORC1 phosphorylates and inhibits the ribosome preservation factor Stm1 to activate dormant ribosomes. EMBO J. 2023;42(5):e112344.
- Blandy A, Hopes T, Vasconcelos EJR, Turner A, Fatkhullin B, Agapiou M, et al. Translational activity of 80S monosomes varies dramatically across different tissues. Nucleic Acids Res. 2025;53(8).
- Zylber EA, Penman S. The effect of high ionic strength on monomers, polyribosomes, and puromycin-treated polyribosomes. Biochim Biophys Acta. 1970;204(1):221–9.
- Martin TE, Hartwell LH. Resistance of active yeast ribosomes to dissociation by KCl. J Biol Chem. 1970;245(6):1504–6.
- Biever A, Glock C, Tushev G, Ciirdaeva E, Dalmay T, Langer JD, et al. Monosomes actively translate synaptic mRNAs in neuronal processes. Science. 2020;367(6477).
- Popper B, Burkle M, Ciccopiedi G, Marchioretto M, Forne I, Imhof A, et al. Ribosome inactivation regulates translation elongation in neurons. J Biol Chem. 2024;300(2):105648.
- Martin S, Allan KC, Pinkard O, Sweet T, Tesar PJ, Coller J. Oligodendrocyte differentiation alters tRNA modifications and codon optimality-mediated mRNA decay. Nat Commun. 2022;13(1):5003.
- Liu B, Qian SB. Characterizing inactive ribosomes in translational profiling. Translation (Austin). 2016;4(1):e1138018.
- Tomuro K, Mito M, Toh H, Kawamoto N, Miyake T, Chow SYA, et al. Calibrated ribosome profiling assesses the dynamics of ribosomal flux on transcripts. Nat Commun. 2024;15(1):7061.
- Wolf DA, Bahler J, Wise JA. Schizosaccharomyces pombe Polysome Profile Analysis and RNA Purification. Cold Spring Harb Protoc. 2017;2017(4):pdb prot091637.
- Santos DA, Shi L, Tu BP, Weissman JS. Cycloheximide can distort measurements of mRNA levels and translation efficiency. Nucleic Acids Res. 2019;47(10):4974–85.
- Fraenkel, DG. Yeast intermediary metabolism. Cold Spring Harbor, N.Y.: Cold Spring Harbor Laboratory Press; 2011. ix, 434 p. p.
- Reier K, Liiv A, Remme J. Ribosome Protein Composition Mediates Translation during the Escherichia coli Stationary Phase. Int J Mol Sci. 2023;24(4).
- Cheng Z, Brar GA. Global translation inhibition yields condition-dependent de-repression of ribosome biogenesis mRNAs. Nucleic Acids Res. 2019;47(10):5061–73.
- Yin K, Tong M, Suttapitugsakul S, Xu S, Wu R. Global quantification of newly synthesized proteins reveals cell type- and inhibitor-specific effects on protein synthesis inhibition. PNAS Nexus. 2023;2(6):pgad168.
- Jaquet V, Wallerich S, Voegeli S, Turos D, Viloria EC, Becskei A. Determinants of the temperature adaptation of mRNA degradation. Nucleic Acids Res. 2022;50(2):1092–110.
- Becskei A, Rahaman S. The life and death of RNA across temperatures. Comput Struct Biotechnol J. 2022;20:4325–36.
- Rahaman S, Faravelli S, Voegeli S, Becskei A. Polysome propensity and tunable thresholds in coding sequence length enable differential mRNA stability. Sci Adv. 2023;9(39):eadh9545.
- Mondal S, Becskei A. Gene choice in cancer cells is exclusive in ion transport but concurrent in DNA replication. Comput Struct Biotechnol J. 2024;23:2534–47.
- Imai H, Utsumi D, Torihara H, Takahashi K, Kuroyanagi H, Yamashita A. Simultaneous measurement of nascent transcriptome and translatome using 4-thiouridine metabolic RNA labeling and translating ribosome affinity purification. Nucleic Acids Res. 2023;51(14):e76.
- Xie Y, Shu T, Liu T, Spindler MC, Mahamid J, Hocky GM, et al. Polysome collapse and RNA condensation fluidize the cytoplasm. Mol Cell. 2024;84(14):2698–716 e9.
- Wu Y, Wang X, Meng L, Liao Z, Ji W, Zhang P, et al. Translation landscape of stress granules. Sci Adv. 2025;11(40):eady6859.
- Fedorovskiy AG, Burakov AV, Terenin IM, Bykov DA, Lashkevich KA, Popenko VI, et al. A Solitary Stalled 80S Ribosome Prevents mRNA Recruitment to Stress Granules. Biochemistry (Mosc). 2023;88(11):1786–99.
- Brodiazhenko T, Johansson MJO, Takada H, Nissan T, Hauryliuk V, Murina V. Elimination of Ribosome Inactivating Factors Improves the Efficiency of Bacillus subtilis and Saccharomyces cerevisiae Cell-Free Translation Systems. Front Microbiol. 2018;9:3041.
- Ekemezie CL, Melnikov SV. Hibernating ribosomes as drug targets? Front Microbiol. 2024;15:1436579.
- Roshan P, Biswas A, Ahmed S, Anagnos S, Luebbers R, Harish K, et al. Sequestration of ribosomal subunits as inactive 80S by targeting eIF6 limits mitotic exit and cancer progression. Nucleic Acids Res. 2025;53(4).
| Year | Authors (First, Last) | Finding | Condition |
|---|---|---|---|
| 2006 | Van Dyke/Van Dyke (26) | Stm1 associates with ribosomes independently of mRNA; Viability of Δstm1 cells is reduced after long-term nitrogen-starvation | Long-term starvation in nitrogen-deficient medium (2 to 6 days). |
| 2011 | Balagopal/Parker (27) | Stm1 overexpression causes stronger growth inhibition in dom34Δ strains than in wild type, suggesting that Stm1 stalls ribosomes in vivo and that Dom34/Hbs1 releases Stm1-stalled ribosomes. | Growth on rich medium at mild cold shock (16 °C). |
| 2011 | Ben-Shem/Yusupov (25) | Crystal structure reveals only one non-ribosomal protein, Stm1. Stm1 is suggested to clamp the two subunits preventing their dissociation, and inhibits translation by excluding mRNA binding. | Short-term starvation in medium lacking glucose (30 °C, 10 min). |
| 2013 | Van Dyke/Van Dyke (14) | Ribosomal protein levels are similar between wild-type and Δstm1 after one day in quiescence but diverge after four days, and Stm1 overexpression prevents ribosome degradation. | Cells were kept in stationary state for 4 days. |
| 2014 | van den Elzen/Séraphin (28) | Stm1-bound 80S ribosomes are substrates for Dom34/Hbs1/Rli1-mediated subunit splitting in vitro; Deletion of STM1 suppresses the requirement for Dom34-Hbs1 to restart translation in vivo. | Short term glucose-starvation combined with mild cold shock (10 min, 16 °C). |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




