Submitted:
26 November 2025
Posted:
27 November 2025
Read the latest preprint version here
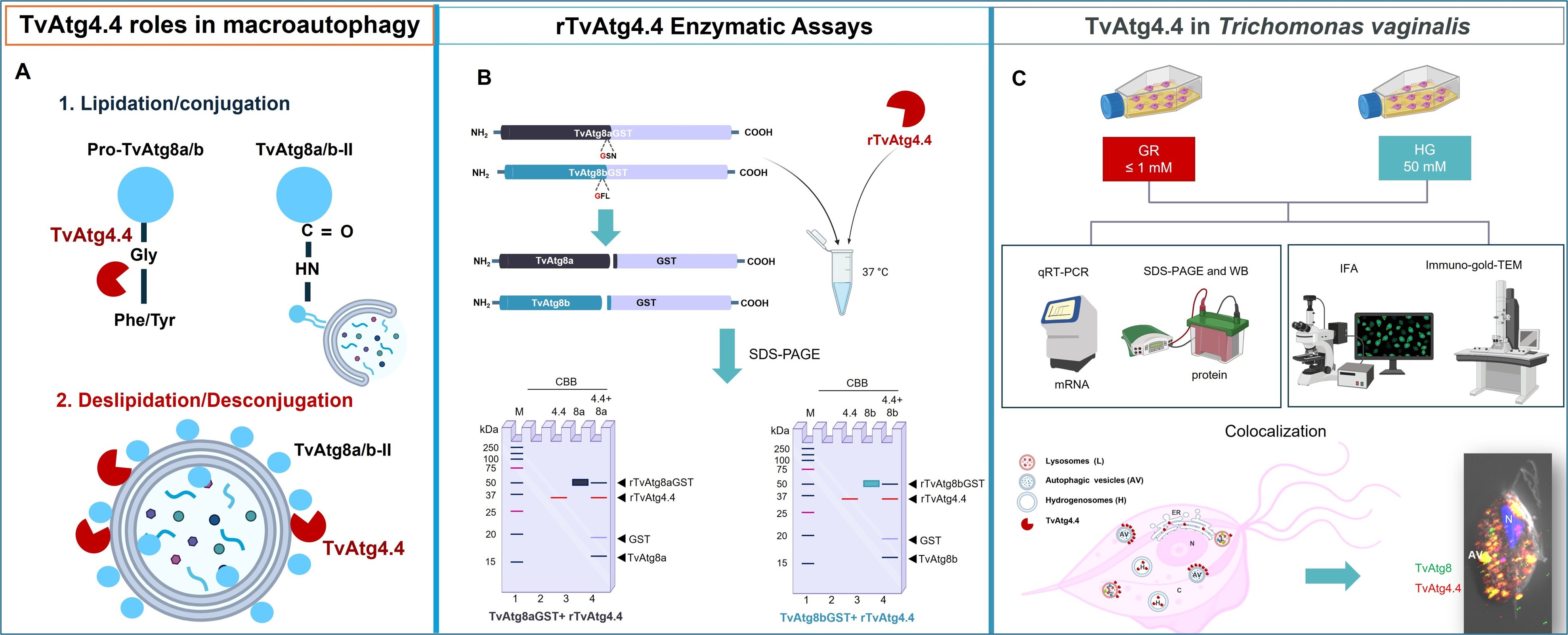
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. In Vitro Cultures of Trichomonas vaginalis
2.2. Cloning, Expression, and Purification of Recombinant TvAtg4.4, TvAtg8aGST, TvAtg8bGST, and GST Proteins
2.3. Production of Anti-rTvAtg4.4 Specific Antibodies in Mice and Rabbits
2.4. Enzymatic Assays of rTvAtg4.4 CP Against rTvAtg8aGST and rTvAtg8bGST Substrates
2.5. FarWestern Blotting (FWB) Assay
2.6. qRT-PCR Assay
2.7. Total Protein Extracts for SDS-PAGE Analysis
2.8. Western Blot (WB) Assay
2.9. Localization of TvAtg4.4 in Trichomonas by Indirect Immunofluorescence Assays
2.10. Gold Immunolabeling by Transmission Electron Microscopy
2.11. Statistical Analysis
3. Results
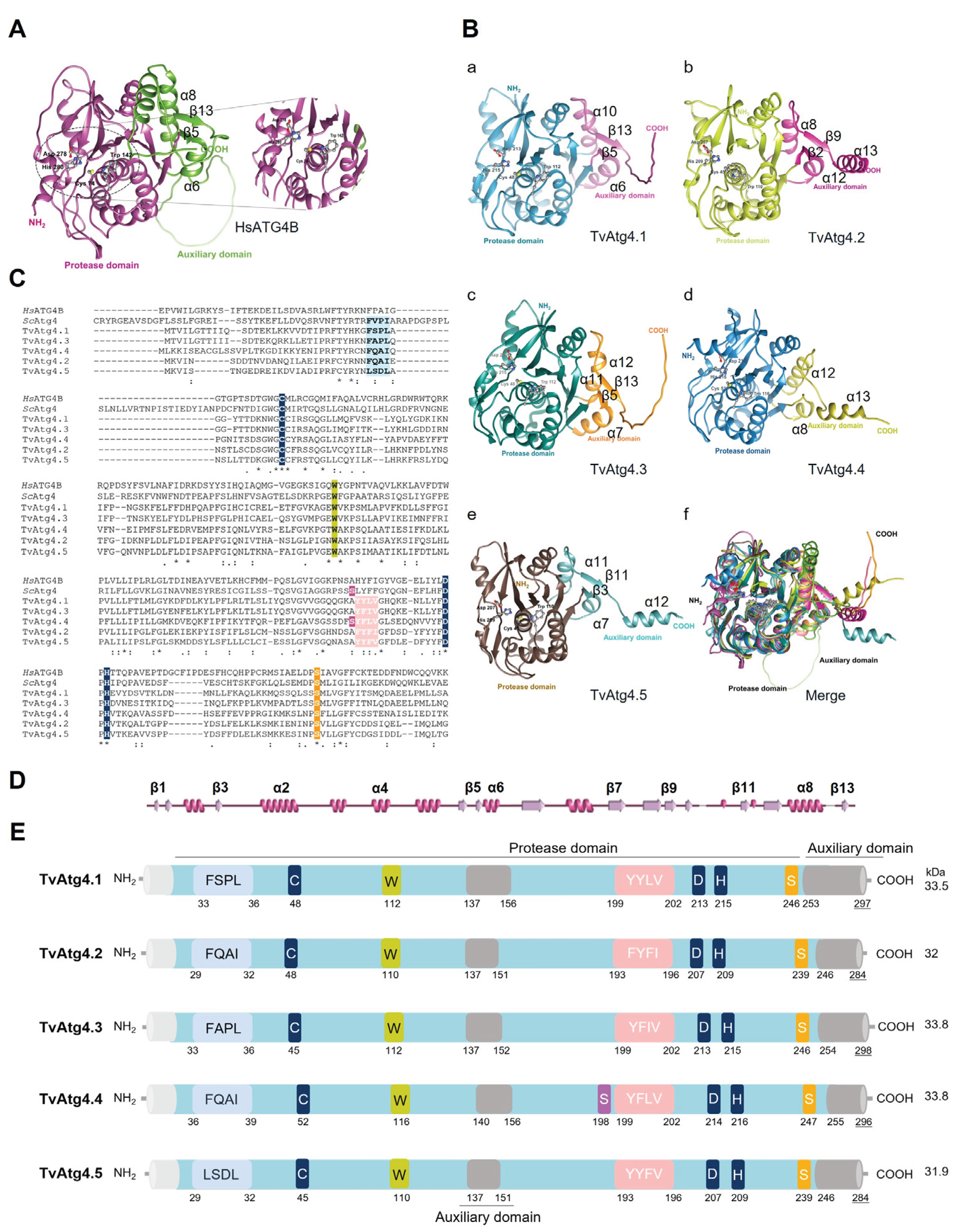
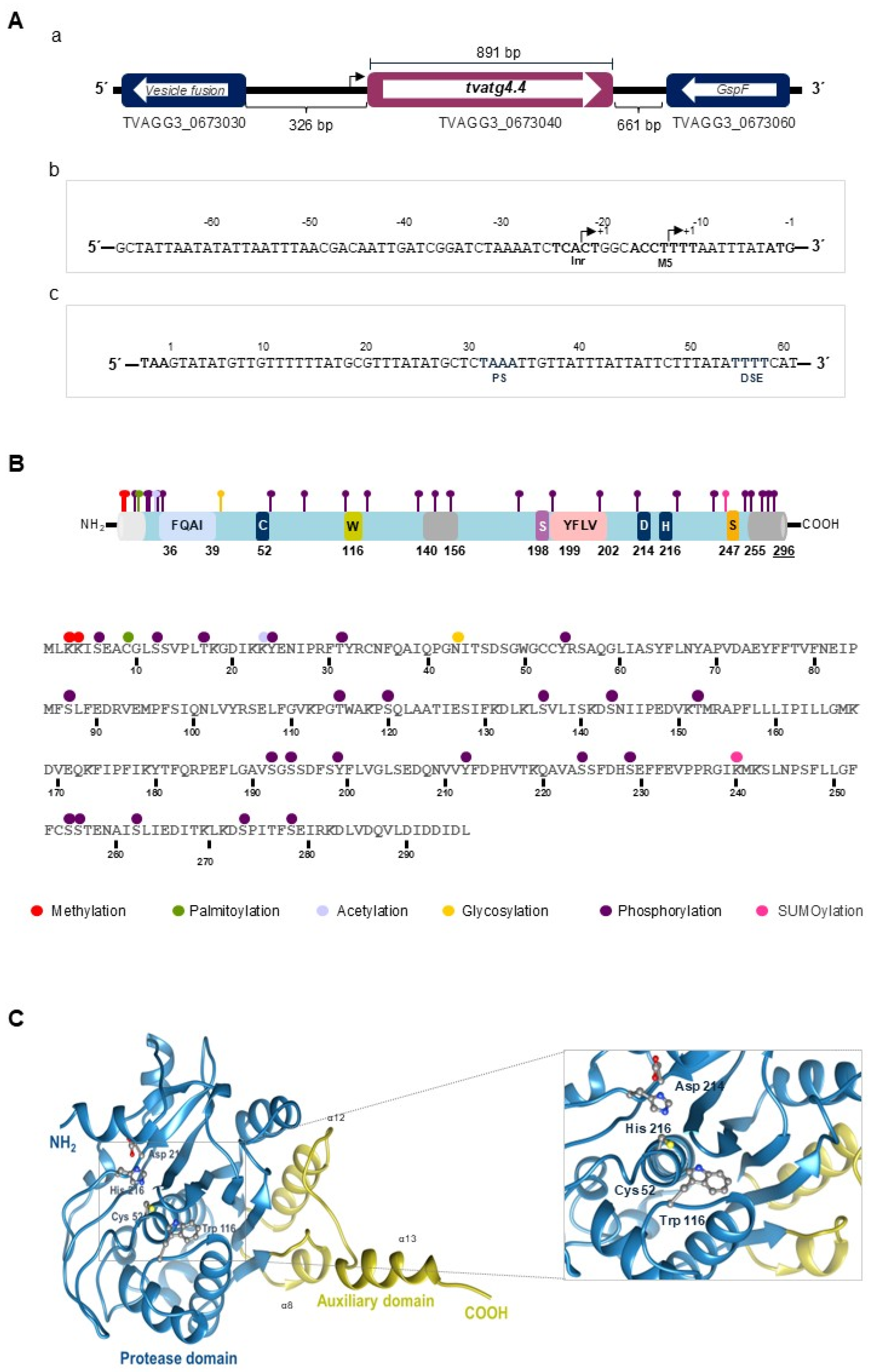
3.1. In Silico Analysis of the Genomic Sequence of T. vaginalis Autophagins
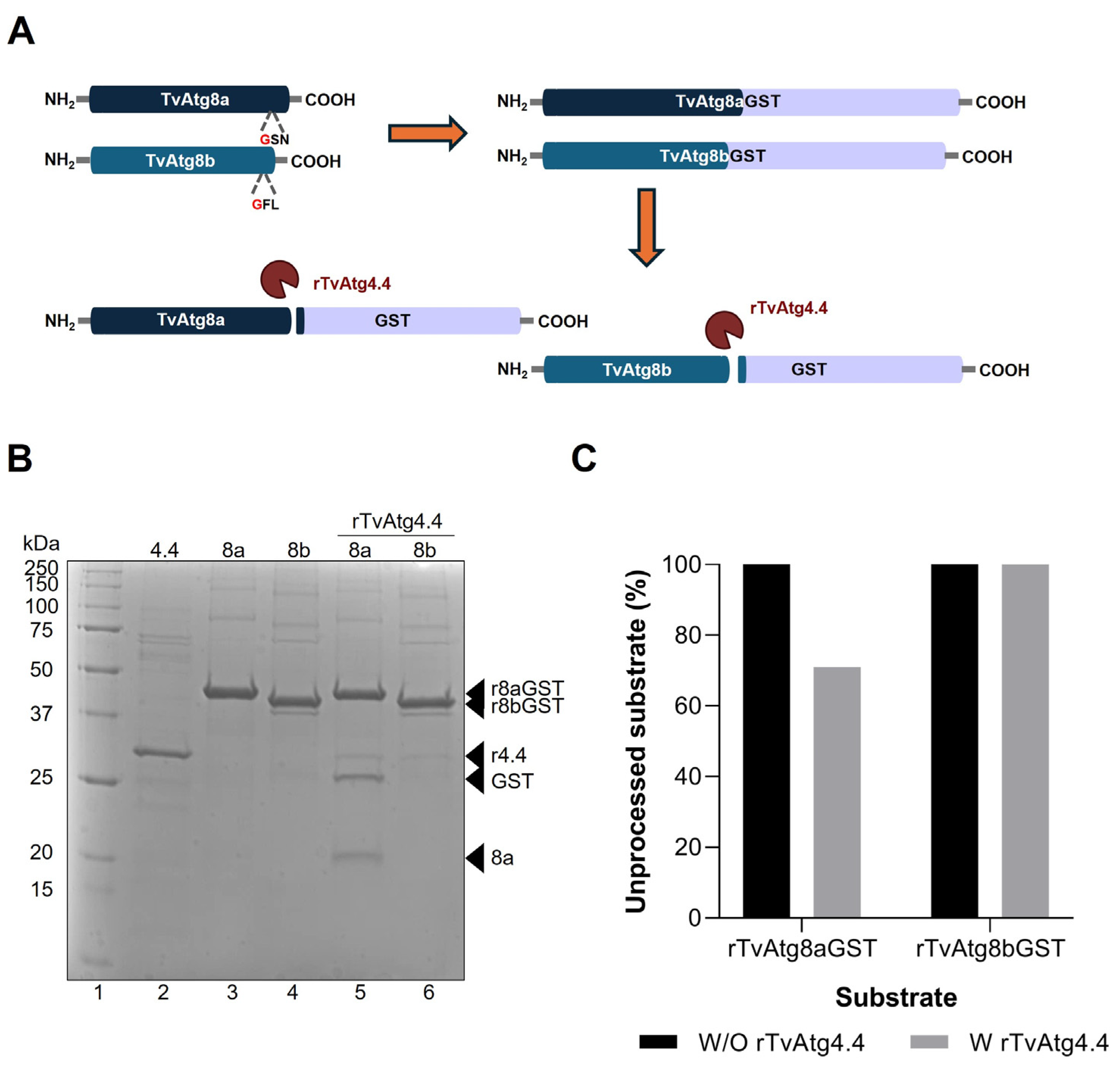
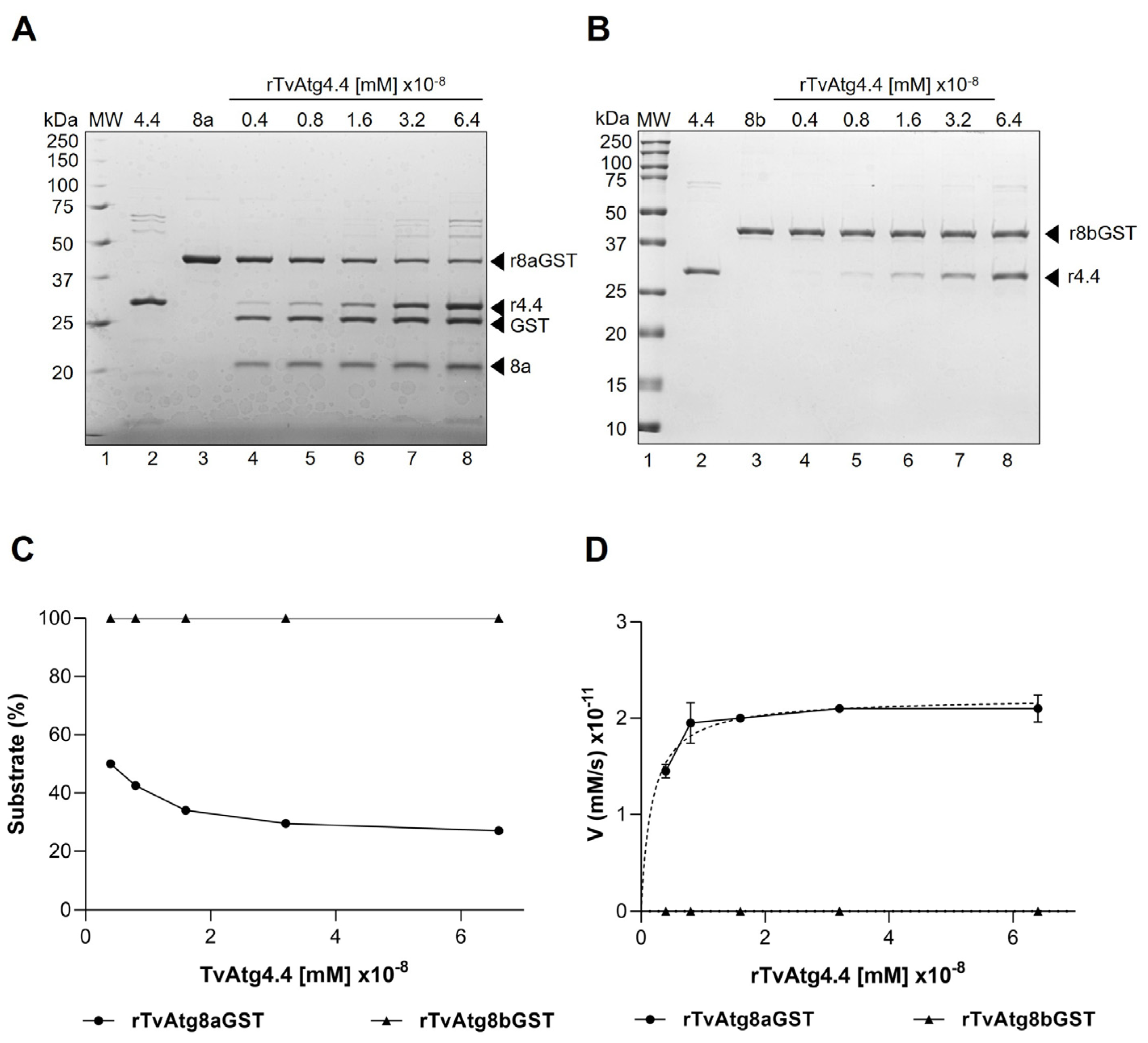
3.2. In Vitro Processing of rTvAtg8aGST and rTvAtg8bGST Substrates by rTvAtg4.4 Cysteine Protease
3.3. The Proteolytic Activity of rTvAtg4.4 is Concentration-Dependent
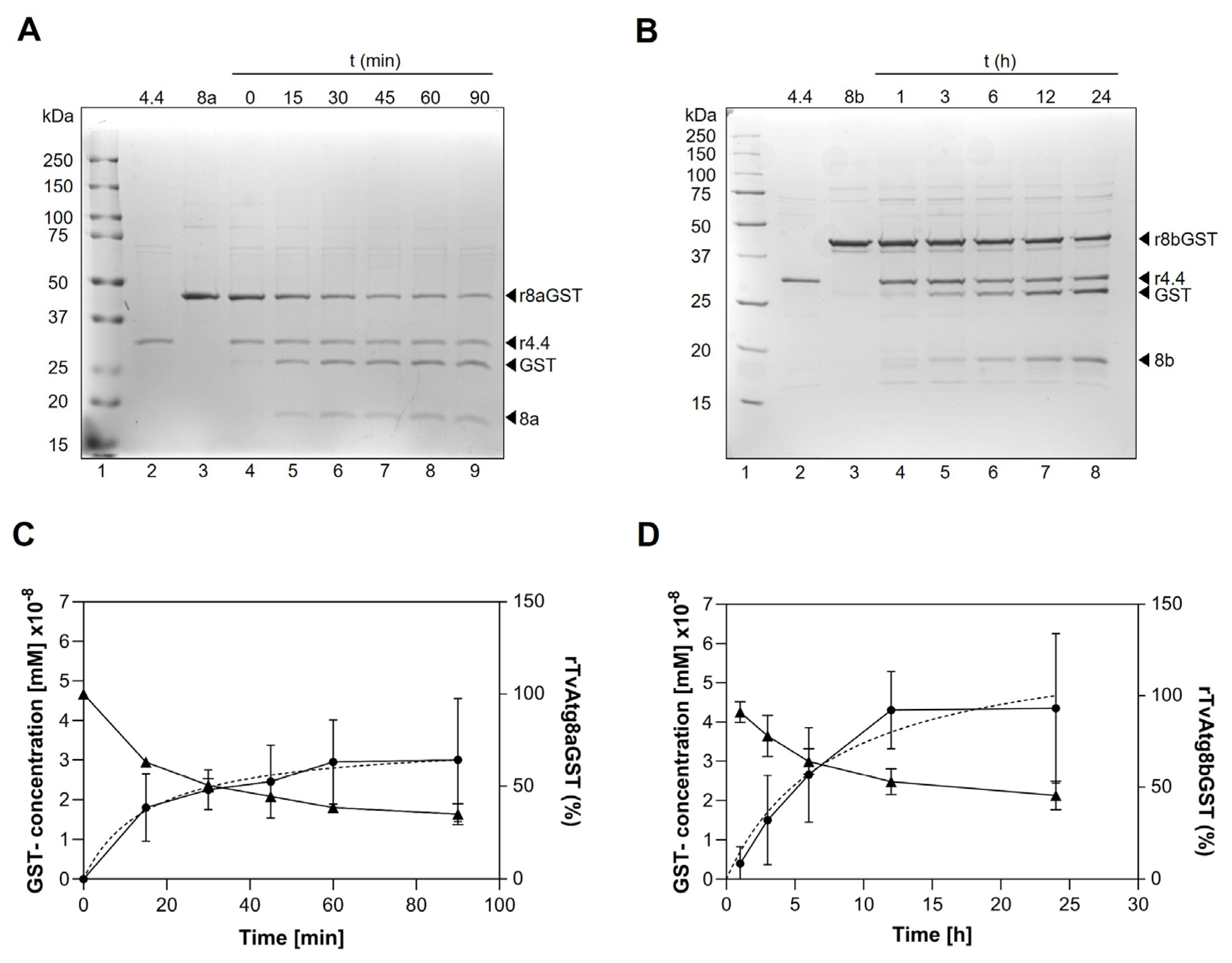
3.4. Effect of Time on rTvAtg4.4 Proteolytic Activity Towards rTvAtg8aGST and rTvAtg8bGST Substrates
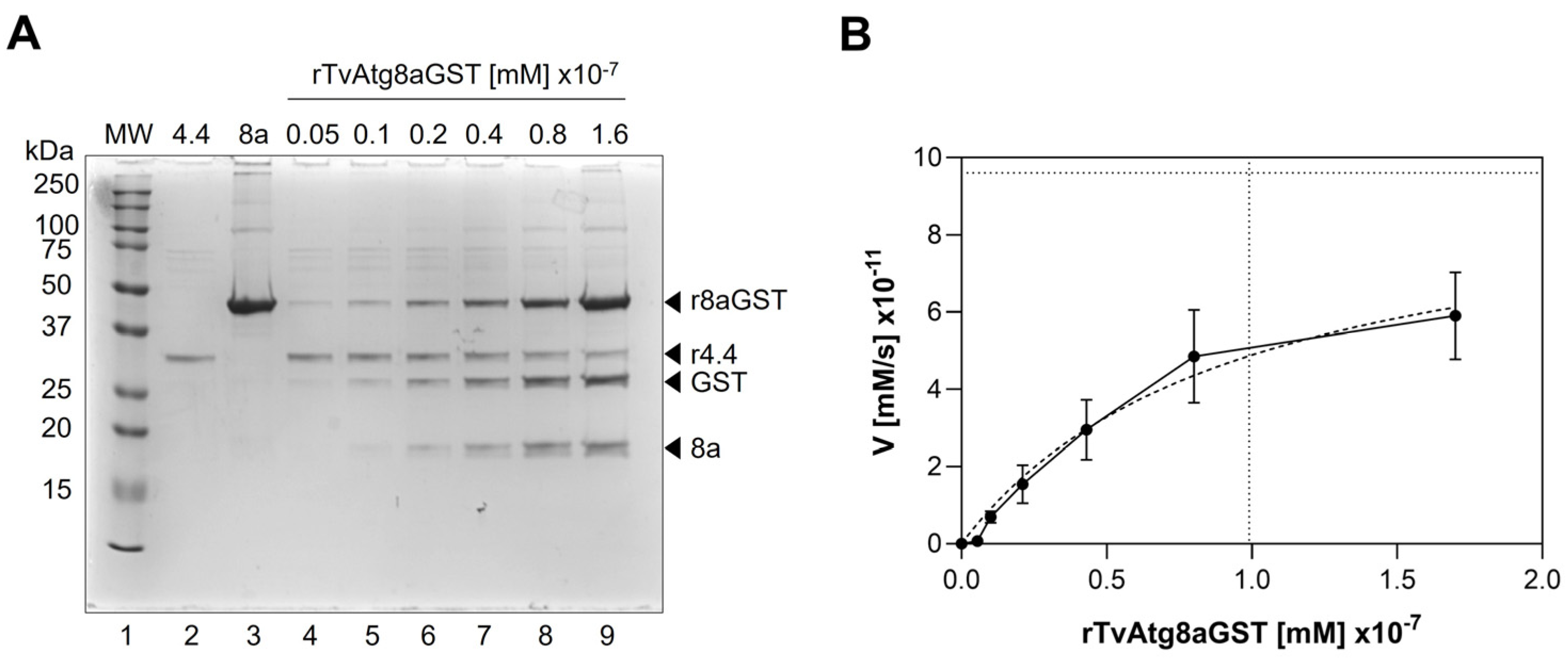
3.5. Enzymatic Kinetics for rTvAtg4.4 Autophagin
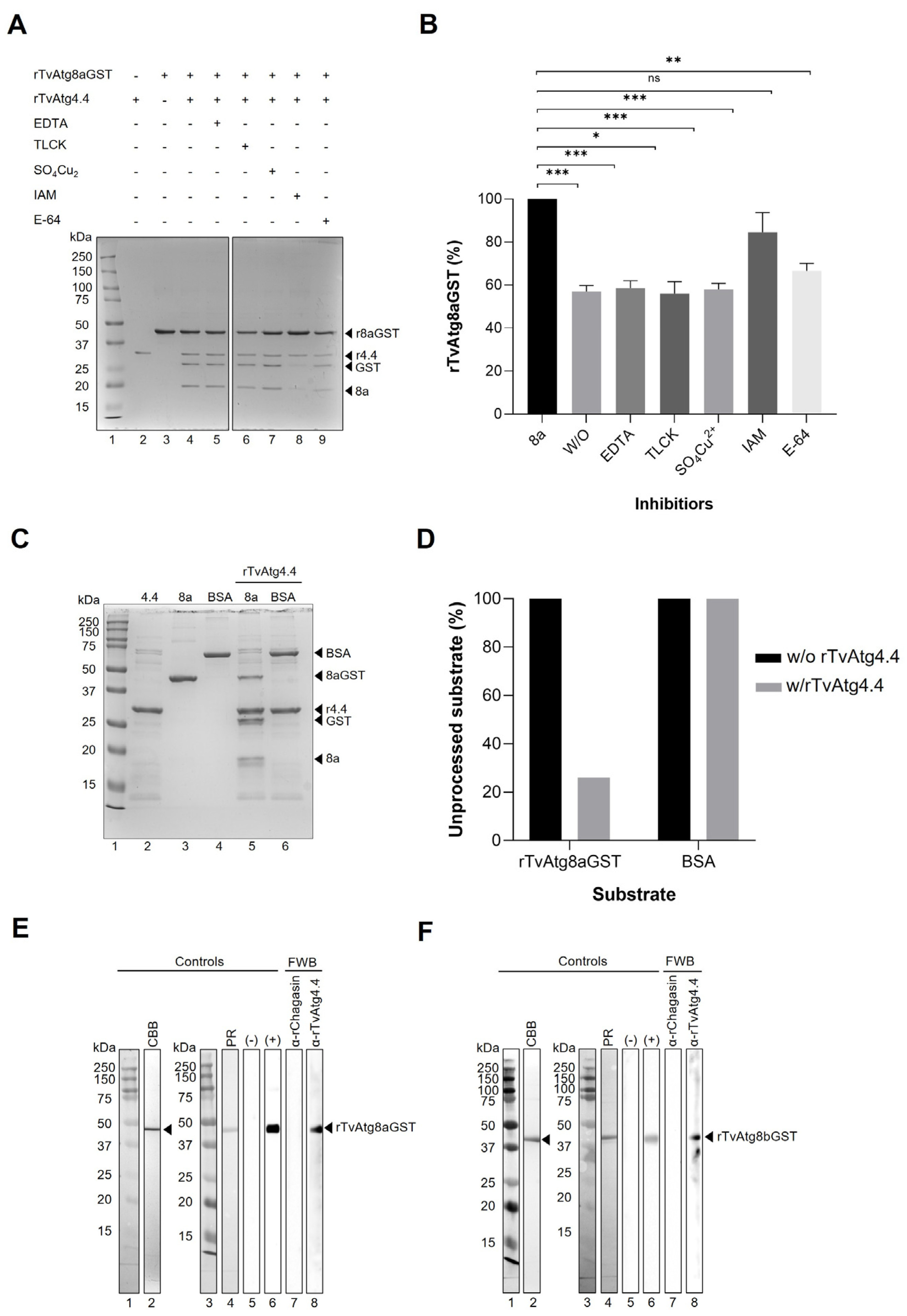
3.6. Interaction of rTvAtg4.4 Autophagin with rTvAtg8aGST and rTvAtg8bGST Substrates
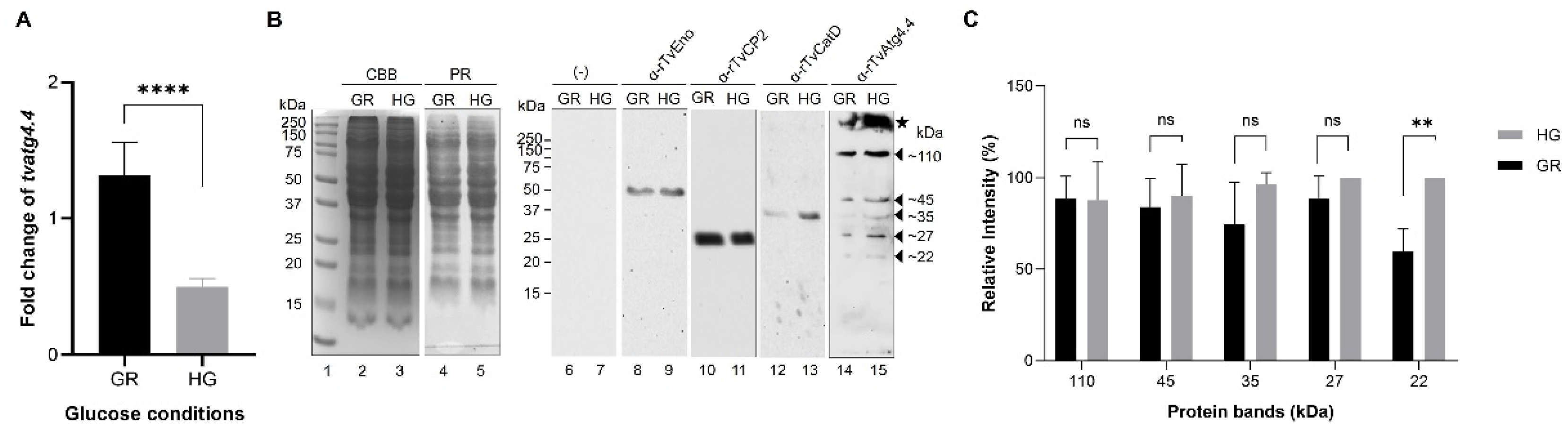
3.7. Effect of Glucose on Tvatg4.4 Transcription
3.8. Effect of Glucose on the Amount of TvAtg4.4 Protein
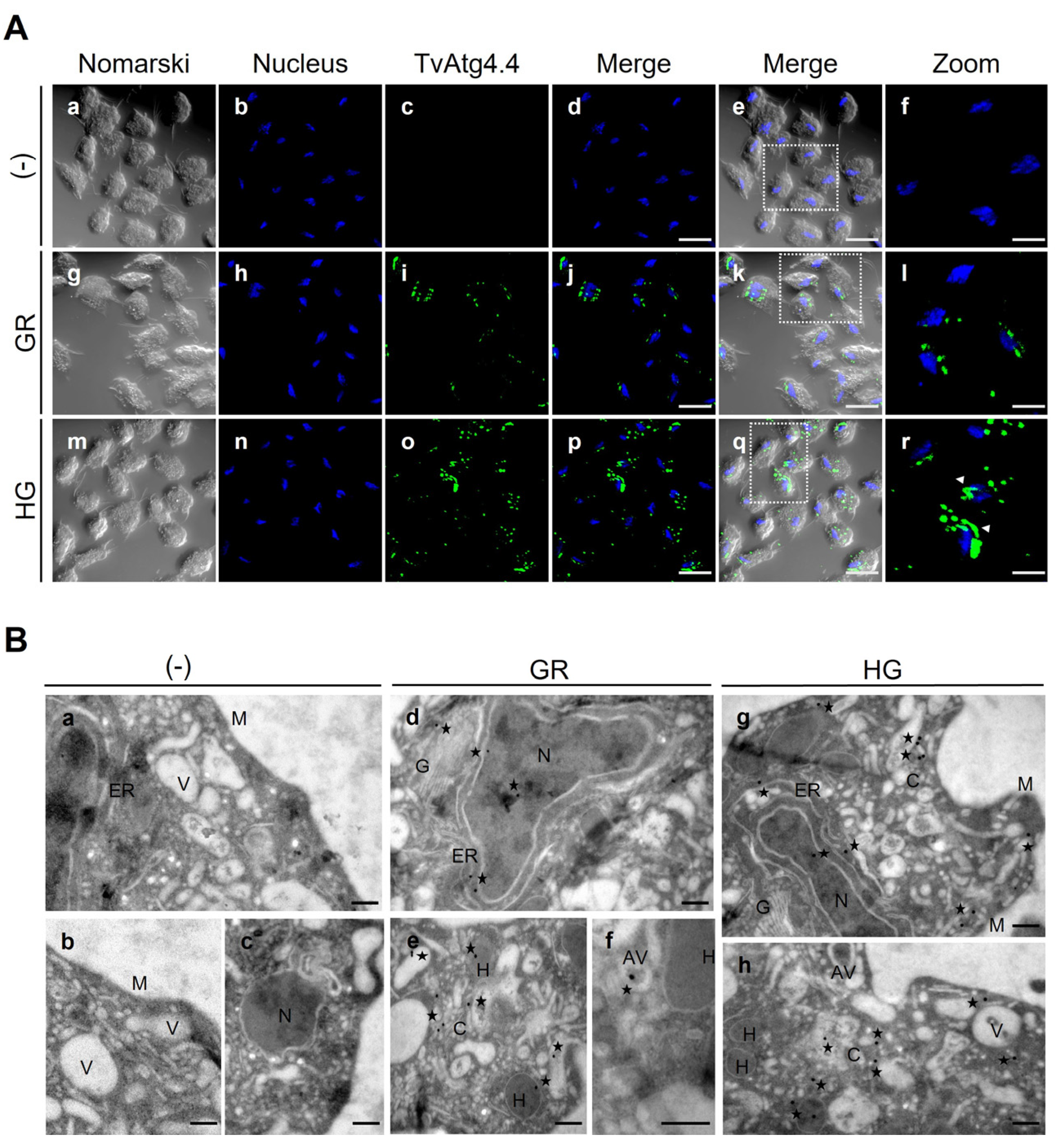
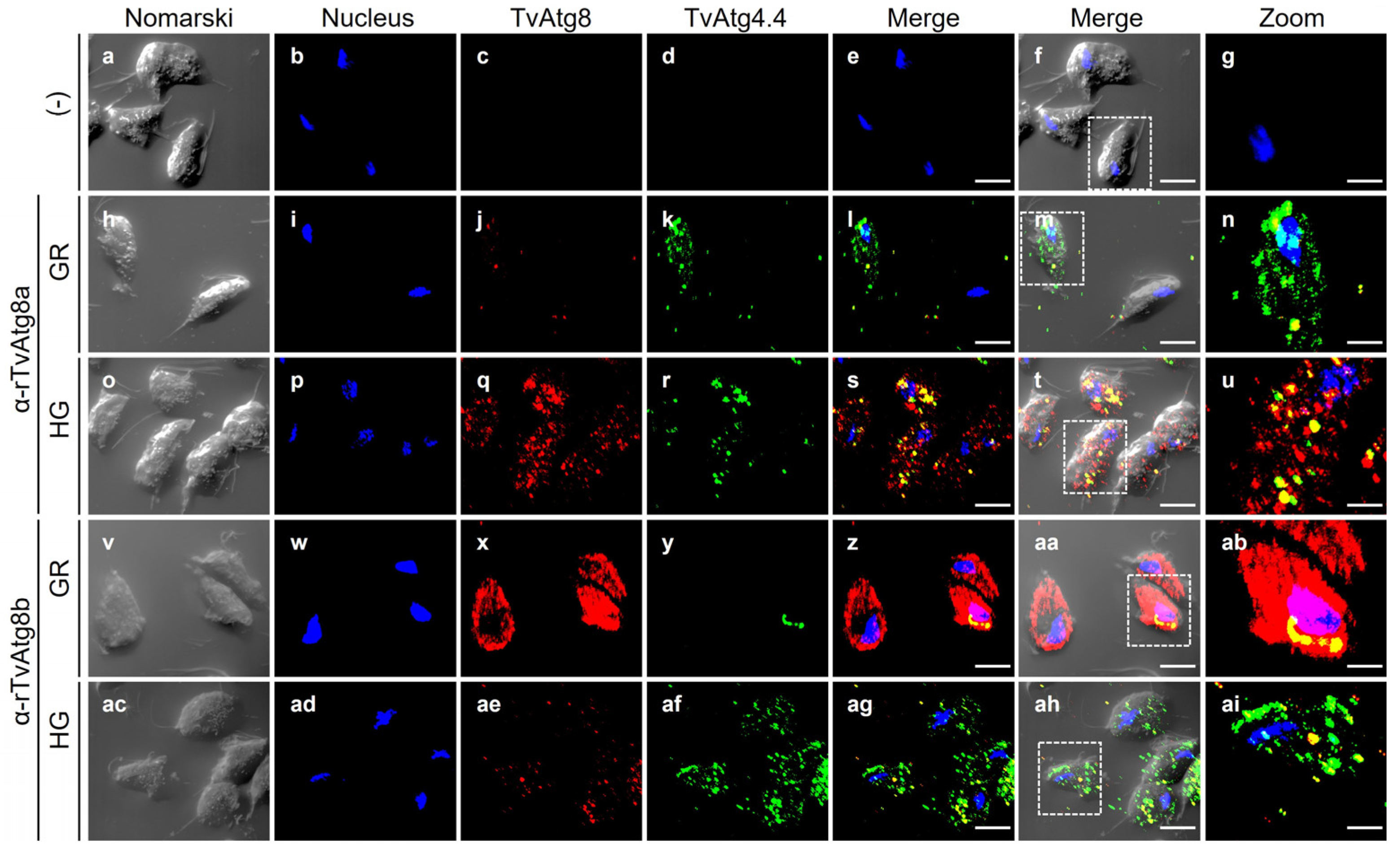
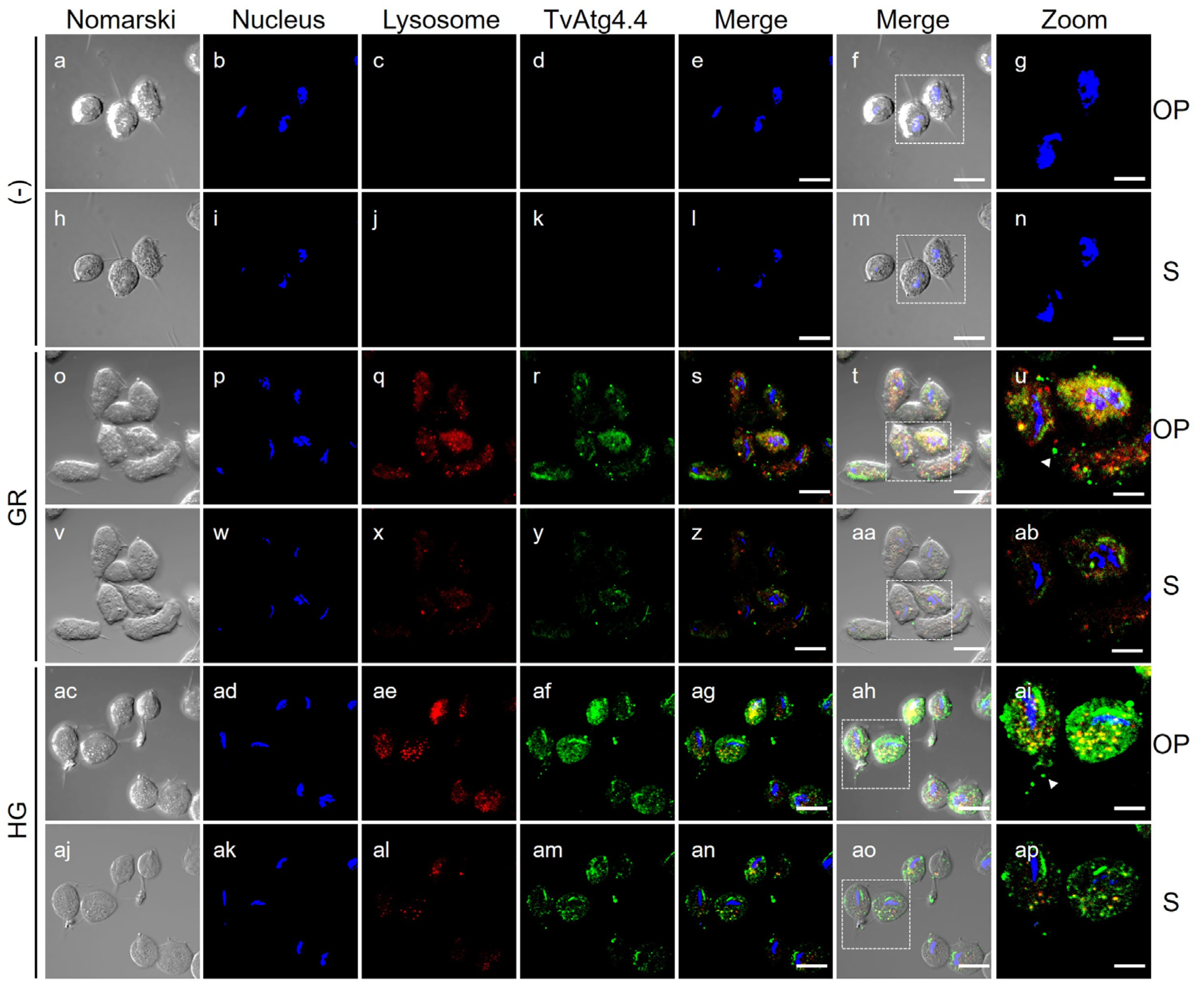
3.9. Localization of the TvAtg4.4 Autophagin of T. vaginalis Grown Under Different Glucose Conditions
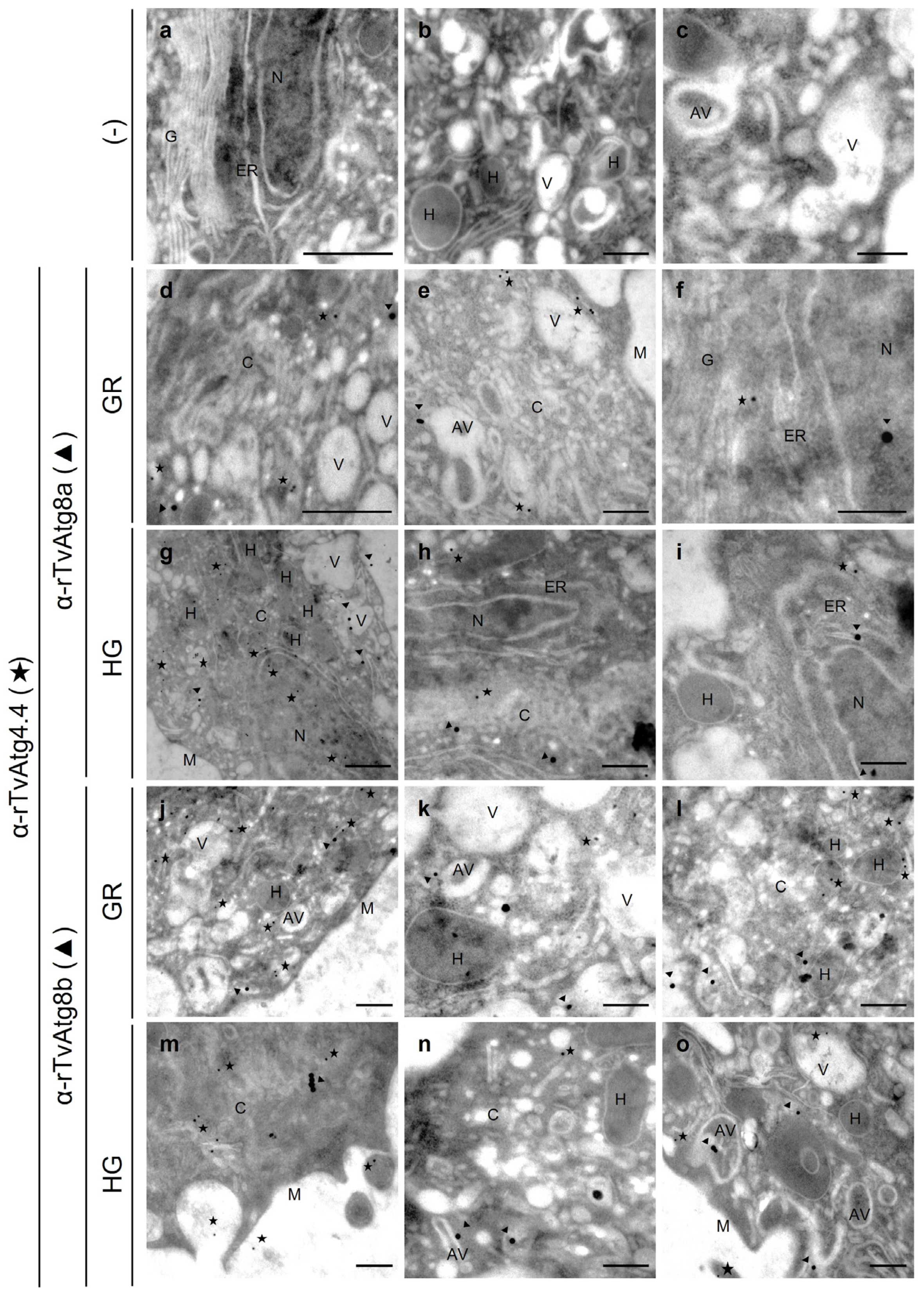
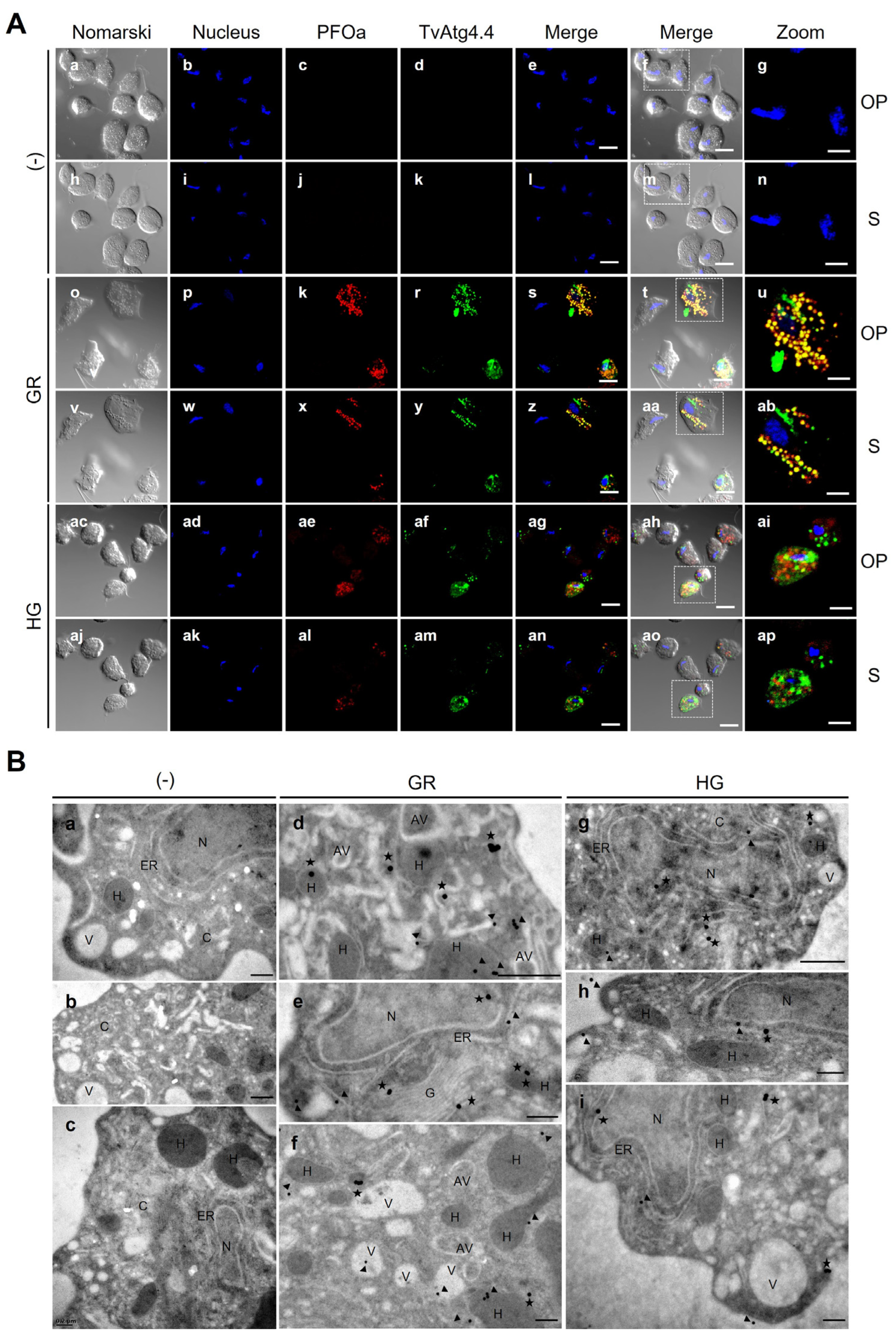
3.10. TvAtg4.4 Is Localized in Autophagic Vesicles Under Different Glucose Conditions
3.11. Localization of TvAtg4.4 Autophagin in Lysosomes
3.12. Localization of TvAtg4.4 Autophagin in Hydrogenosomes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Aman, Y., Schmauck-Medina, T., Hansen, M., Morimoto, R. I., Simon, A. K., Bjedov, I., Palikaras, K., Simonsen, A., Johansen, T., Tavernarakis, N., Rubinsztein, D. C., Partridge, L., Kroemer, G., Labbadia, J., & Fang, E. F. (2021). Autophagy in healthy aging and disease. Nature Aging, 1(8), 634–650. [CrossRef]
- Dikic, I., & Elazar, Z. (2018). Mechanism and medical implications of mammalian autophagy. Nature Reviews. Molecular Cell Biology, 19(6), 349–364. [CrossRef]
- Chang N. C. (2020). Autophagy and stem cells: Self-eating for self-renewal. Frontiers in Cell and Developmental Biology, 8, 138. [CrossRef]
- Yin, Z., Pascual, C., & Klionsky, D. J. (2016). Autophagy: machinery and regulation. Microbial Cell (Graz, Austria), 3(12), 588–596. [CrossRef]
- Yu, L., Chen, Y., & Tooze, S. A. (2018). Autophagy pathway: Cellular and molecular mechanisms. Autophagy, 14(2), 207–215. [CrossRef]
- Li, X., He, S., & Ma, B. (2020). Autophagy and autophagy-related proteins in cancer. Molecular Cancer, 19(1), 12. [CrossRef]
- Noda, N. N., Ohsumi, Y., & Inagaki, F. (2009). ATG systems from the protein structural point of view. Chemical Reviews, 109(4), 1587–1598. [CrossRef]
- Xie, Z., & Klionsky, D. J. (2007). Autophagosome formation: core machinery and adaptations. Nature Cell Biology, 9(10), 1102–1109. [CrossRef]
- Mizushima N. (2020). The ATG conjugation systems in autophagy. Current Opinion in Cell Biology, 63, 1–10. [CrossRef]
- Martens, S., & Fracchiolla, D. (2020). Activation and targeting of ATG8 protein lipidation. Cell Discovery, 6, 23. [CrossRef]
- Glick, D., Barth, S., & Macleod, K. F. (2010). Autophagy: cellular and molecular mechanisms. The Journal of Pathology, 221(1), 3–12. [CrossRef]
- Centers for Disease Control and Prevention. (CDC). About trichomoniasis. Retrieved October 22, 2025, from https://www.cdc.gov/trichomoniasis/es/about/acerca-de-la-tricomoniasis.html.
- Edwards, T., Burke, P., Smalley, H., & Hobbs, G. (2016). Trichomonas vaginalis: Clinical relevance, pathogenicity and diagnosis. Critical Reviews in Microbiology, 42(3), 406–417. [CrossRef]
- Poole, D. N., & McClelland, R. S. (2013). Global epidemiology of Trichomonas vaginalis. Sexually Transmitted Infections, 89(6), 418–422. [CrossRef]
- Kissinger P. (2015). Trichomonas vaginalis: a review of epidemiologic, clinical and treatment issues. BMC Infectious Diseases, 15, 307. [CrossRef]
- Huang, K. Y., Chen, Y. Y., Fang, Y. K., Cheng, W. H., Cheng, C. C., Chen, Y. C., Wu, T. E., Ku, F. M., Chen, S. C., Lin, R., & Tang, P. (2014). Adaptive responses to glucose restriction enhance cell survival, antioxidant capability, and autophagy of the protozoan parasite Trichomonas vaginalis. Biochimica et Biophysica Acta, 1840(1), 53–64. [CrossRef]
- Huang, K. Y., Chen, R. M., Lin, H. C., Cheng, W. H., Lin, H. A., Lin, W. N., Huang, P. J., Chiu, C. H., & Tang, P. (2019). Potential role of autophagy in proteolysis in Trichomonas vaginalis. Journal of Microbiology, Immunology, and Infection = Wei mian yu gan ran za zhi, 52(2), 336–344. [CrossRef]
- Hernández-García, M. S., Miranda-Ozuna, J. F. T., Salazar-Villatoro, L., Vázquez-Calzada, C., Ávila-González, L., González-Robles, A., Ortega-López, J., & Arroyo, R. (2019). Biogenesis of autophagosome in Trichomonas vaginalis during macroautophagy induced by rapamycin-treatment and iron or glucose starvation conditions. The Journal of Eukaryotic Microbiology, 66(4), 654–669. [CrossRef]
- Dias-Lopes, G., Wiśniewski, J. R., de Souza, N. P., Vidal, V. E., Padrón, G., Britto, C., Cuervo, P., & De Jesus, J. B. (2018). In-depth quantitative proteomic analysis of trophozoites and pseudocysts of Trichomonas vaginalis. Journal of Proteome Research, 17(11), 3704–3718. [CrossRef]
- Goulas, T., Cuppari, A., Garcia-Castellanos, R., Snipas, S., Glockshuber, R., Arolas, J. L., & Gomis-Rüth, F. X. (2014). The pCri system: a vector collection for recombinant protein expression and purification. PloS One, 9(11), e112643. [CrossRef]
- Li, M., Fu, Y., Yang, Z., & Yin, X. M. (2017). Measurement of the activity of the Atg4 cysteine proteases. Methods in Enzymology, 587, 207–225. [CrossRef]
- Cárdenas-Guerra, R. E., Montes-Flores, O., Nava-Pintor, E. E., Resendiz-Cardiel, G., Flores-Pucheta, C. I., Rodriguez-Gavaldon, Y. I., Arroyo, R., Bottazzi, M. E., Hotez, P. J., and Ortega-Lopez, J. (2024) Chagasin from Trypanosoma cruzi as a molecular scaffold to express epitopes of TSA-1 as soluble recombinant chimeras, Protein Expr Purif 218, 106458.
- Dos Santos, O., de Vargas Rigo, G., Frasson, A. P., Macedo, A. J., & Tasca, T. (2015). Optimal reference genes for gene expression normalization in Trichomonas vaginalis. PloS One, 10(9), e0138331. [CrossRef]
- Euceda-Padilla, E. A., Mateo-Cruz, M. G., Ávila-González, L., Flores-Pucheta, C. I., Ortega-López, J., Talamás-Lara, D., Velazquez-Valassi, B., Jasso-Villazul, L., & Arroyo, R. (2024). Trichomonas vaginalis legumain-2, TvLEGU-2, is an immunogenic cysteine peptidase expressed during trichomonal infection. Pathogens (Basel, Switzerland), 13(2), 119. [CrossRef]
- Meza-Cervantez, P., González-Robles, A., Cárdenas-Guerra, R. E., Ortega-López, J., Saavedra, E., Pineda, E., & Arroyo, R. (2011). Pyruvate:ferredoxin oxidoreductase (PFO) is a surface-associated cell-binding protein in Trichomonas vaginalis and is involved in trichomonal adherence to host cells. Microbiology (Reading, England), 157(Pt 12), 3469–3482. [CrossRef]
- Kumanomidou, T., Mizushima, T., Komatsu, M., Suzuki, A., Tanida, I., Sou, Y. S., Ueno, T., Kominami, E., Tanaka, K., & Yamane, T. (2006). The crystal structure of human Atg4b, a processing and de-conjugating enzyme for autophagosome-forming modifiers. Journal of Molecular Biology, 355(4), 612–618. [CrossRef]
- Rogov, V. V., Nezis, I. P., Tsapras, P., Zhang, H., Dagdas, Y., Noda, N. N., Nakatogawa, H., Wirth, M., Mouilleron, S., McEwan, D. G., Behrends, C., Deretic, V., Elazar, Z., Tooze, S. A., Dikic, I., Lamark, T., & Johansen, T. (2023). Atg8 family proteins, LIR/AIM motifs and other interaction modes. Autophagy Reports, 2(1), 2188523. [CrossRef]
- Satoo, K., Suzuki, N. N., Fujioka, Y., Mizushima, N., Ohsumi, Y., & Inagaki, F. (2007). Crystallization and preliminary crystallographic analysis of human Atg4B-LC3 complex. Acta crystallographica. Section F, Structural biology and crystallization communications, 63(Pt 2), 99–102. [CrossRef]
- Liston, D. R., & Johnson, P. J. (1999). Analysis of a ubiquitous promoter element in a primitive eukaryote: early evolution of the initiator element. Molecular and Cellular Biology, 19(3), 2380–2388. [CrossRef]
- Smith, A. J., Chudnovsky, L., Simoes-Barbosa, A., Delgadillo-Correa, M. G., Jonsson, Z. O., Wohlschlegel, J. A., & Johnson, P. J. (2011). Novel core promoter elements and a cognate transcription factor in the divergent unicellular eukaryote Trichomonas vaginalis. Molecular and Cellular Biology, 31(7), 1444–1458. [CrossRef]
- Espinosa, N., Hernández, R., López-Griego, L., & López-Villaseñor, I. (2002). Separable putative polyadenylation and cleavage motifs in Trichomonas vaginalis mRNAs. Gene, 289(1-2), 81–86. [CrossRef]
- Kirisako, T., Ichimura, Y., Okada, H., Kabeya, Y., Mizushima, N., Yoshimori, T., Ohsumi, M., Takao, T., Noda, T., & Ohsumi, Y. (2000). The reversible modification regulates the membrane-binding state of Apg8/Aut7 essential for autophagy and the cytoplasm to vacuole targeting pathway. The Journal of Cell Biology, 151(2), 263–276. [CrossRef]
- Hirata, E., Ohya, Y., & Suzuki, K. (2017). Atg4 plays an important role in efficient expansion of autophagic isolation membranes by cleaving lipidated Atg8 in Saccharomyces cerevisiae. Plos One, 12 (7), e0181047. [CrossRef]
- Zhou, Y., Wang, Z., Huang, Y., Bai, C., Zhang, X., Fang, M., Ju, Z., & Liu, B. (2022). Membrane dynamics of ATG4B and LC3 in autophagosome formation. Journal of Molecular Cell Biology, 13(12), 853–863. [CrossRef]
- Abreu, S., Kriegenburg, F., Gómez-Sánchez, R., Mari, M., Sánchez-Wandelmer, J., Skytte Rasmussen, M., Soares Guimarães, R., Zens, B., Schuschnig, M., Hardenberg, R., Peter, M., Johansen, T., Kraft, C., Martens, S., & Reggiori, F. (2017). Conserved Atg8 recognition sites mediate Atg4 association with autophagosomal membranes and Atg8 deconjugation. EMBO Reports, 18(5), 765–780. [CrossRef]
- Robinson P. K. (2015). Enzymes: principles and biotechnological applications. Essays in Biochemistry, 59, 1–41. [CrossRef]
- Rendón, J. L., & Pardo, J. P. (2025). Time-dependent kinetic complexities in enzyme assays: A review. Biomolecules, 15(5), 641. [CrossRef]
- Li, M., Hou, Y., Wang, J., Chen, X., Shao, Z. M., & Yin, X. M. (2011). Kinetics comparisons of mammalian Atg4 homologues indicate selective preferences toward diverse Atg8 substrates. The Journal of Biological Chemistry, 286(9), 7327–7338. [CrossRef]
- Shu, C. W., Drag, M., Bekes, M., Zhai, D., Salvesen, G. S., & Reed, J. C. (2010). Synthetic substrates for measuring activity of autophagy proteases: autophagins (Atg4). Autophagy, 6(7), 936–947. [CrossRef]
- Tanida, I., Sou, Y. S., Minematsu-Ikeguchi, N., Ueno, T., & Kominami, E. (2006). Atg8L/Apg8L is the fourth mammalian modifier of mammalian Atg8 conjugation mediated by human Atg4B, Atg7 and Atg3. The FEBS Journal, 273(11), 2553–2562. [CrossRef]
- Silverstein T. P. (2019). When both Km and Vmax are altered, Is the enzyme inhibited or activated? Biochemistry and Molecular Biology Education: a bimonthly publication of the International Union of Biochemistry and Molecular Biology, 47(4), 446–449. [CrossRef]
- Rajković, J., Poreba, M., Caglič, D., Vidmar, R., Wilk, A., Borowik, A., Salvesen, G., Turk, V., Drag, M., & Turk, B. (2015). Biochemical characterization and substrate specificity of autophagin-2 from the parasite Trypanosoma cruzi. The Journal of Biological Chemistry, 290(47), 28231–28244. [CrossRef]
- Rawlings, N. D., Barrett, A. J., Thomas, P. D., Huang, X., Bateman, A., & Finn, R. D. (2018). The MEROPS database of proteolytic enzymes, their substrates and inhibitors in 2017 and a comparison with peptidases in the PANTHER database. Nucleic Acids Research, 46(D1), D624–D632. [CrossRef]
- Yang, N., Matthew, M. A., & Yao, C. (2023). Roles of cysteine proteases in biology and pathogenesis of parasites. Microorganisms, 11(6), 1397. [CrossRef]
- Hernández-Gutiérrez, R., Avila-González, L., Ortega-López, J., Cruz-Talonia, F., Gómez-Gutierrez, G., & Arroyo, R. (2004). Trichomonas vaginalis: characterization of a 39-kDa cysteine proteinase found in patient vaginal secretions. Experimental Parasitology, 107(3-4), 125–135. [CrossRef]
- Mancilla-Olea, M. I., Ortega-López, J., Figueroa-Angulo, E. E., Avila-González, L., Cárdenas-Guerra, R. E., Miranda-Ozuna, J. F. T., González-Robles, A., Hernández-García, M. S., Sánchez-Ayala, L., & Arroyo, R. (2018). Trichomonas vaginalis cathepsin D-like aspartic proteinase (Tv-CatD) is positively regulated by glucose and degrades human hemoglobin. The International Journal of Biochemistry & Cell Biology, 97, 1–15. [CrossRef]
- Grant G. A. (2017). Modification of cysteine. Current Protocols in Protein Science, 87, 15.1.1–15.1.23. [CrossRef]
- Matsumoto, K., Mizoue, K., Kitamura, K., Tse, W. C., Huber, C. P., & Ishida, T. (1999). Structural basis of inhibition of cysteine proteases by E-64 and its derivatives. Biopolymers, 51(1), 99–107. [CrossRef]
- Liu, W. L., Wen, Z. H., Li, Q. Y., Liu, H. B., Li, Q. L., Deng, S. Z., Zeng, Z. Y., Luo, M. C., Tang, A. X., & Liu, Y. Y. (2025). New insights into exploring new functional enzymes through the enzyme promiscuity. International Journal of Biological Macromolecules, 304(Pt 1), 140576. [CrossRef]
- Zhang, L., Li, J., Ouyang, L., Liu, B., & Cheng, Y. (2016). Unraveling the roles of Atg4 proteases from autophagy modulation to targeted cancer therapy. Cancer Letters, 373(1), 19–26. [CrossRef]
- Wu, Y., Li, Q., & Chen, X. Z. (2007). Detecting protein-protein interactions by Far western blotting. Nature Protocols, 2(12), 3278–3284. [CrossRef]
- Miranda-Ozuna, J. F., Hernández-García, M. S., Brieba, L. G., Benítez-Cardoza, C. G., Ortega-López, J., González-Robles, A., & Arroyo, R. (2016). The glycolytic enzyme triosephosphate isomerase of Trichomonas vaginalis is a surface-associated protein induced by glucose that functions as a laminin- and fibronectin-binding protein. Infection and Immunity, 84(10), 2878–2894. [CrossRef]
- Vaulont, S., Vasseur-Cognet, M., & Kahn, A. (2000). Glucose regulation of gene transcription. The Journal of Biological Chemistry, 275(41), 31555–31558. [CrossRef]
- Zheng, X., Yang, Z., Gu, Q., Xia, F., Fu, Y., Liu, P., Yin, X. M., & Li, M. (2020). The protease activity of human ATG4B is regulated by reversible oxidative modification. Autophagy, 16(10), 1838–1850. [CrossRef]
- Pérez-Pérez, M. E., Lemaire, S. D., & Crespo, J. L. (2021). The ATG4 protease integrates redox and stress signals to regulate autophagy. Journal of Experimental Botany, 72(9), 3340–3351. [CrossRef]
- Pérez-Pérez, M. E., Zaffagnini, M., Marchand, C. H., Crespo, J. L., & Lemaire, S. D. (2014). The yeast autophagy protease Atg4 is regulated by thioredoxin. Autophagy, 10(11), 1953–1964. [CrossRef]
- Cleland W. W. (1964). Dithiothreitol, a new protective reagent for SH groups. Biochemistry, 3, 480–482. [CrossRef]
- Xie, Z., Nair, U., & Klionsky, D. J. (2008). Atg8 controls phagophore expansion during autophagosome formation. Molecular Biology of the Cell, 19(8), 3290–3298. [CrossRef]
- Fujita, N., Hayashi-Nishino, M., Fukumoto, H., Omori, H., Yamamoto, A., Noda, T., & Yoshimori, T. (2008). An Atg4B mutant hampers the lipidation of LC3 paralogues and causes defects in autophagosome closure. Molecular Biology of the Cell, 19(11), 4651–4659. [CrossRef]
- Luzio, J. P., Pryor, P. R., & Bright, N. A. (2007). Lysosomes: fusion and function. Nature Reviews. Molecular Cell Biology, 8(8), 622–632. [CrossRef]
- Schneider, R. E., Brown, M. T., Shiflett, A. M., Dyall, S. D., Hayes, R. D., Xie, Y., Loo, J. A., & Johnson, P. J. (2011). The Trichomonas vaginalis hydrogenosome proteome is highly reduced relative to mitochondria, yet complex compared with mitosomes. International Journal for Parasitology, 41(13-14), 1421–1434. [CrossRef]
- Embley, T. M., van der Giezen, M., Horner, D. S., Dyal, P. L., & Foster, P. (2003). Mitochondria and hydrogenosomes are two forms of the same fundamental organelle. Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences, 358(1429), 191–202. [CrossRef]
- Smutná, T., Dohnálková, A., Sutak, R., Narayanasamy, R. K., Tachezy, J., & Hrdý, I. (2022). A cytosolic ferredoxin-independent hydrogenase possibly mediates hydrogen uptake in Trichomonas vaginalis. Current Biology: CB, 32(1), 124–135.e5. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



