Submitted:
18 November 2025
Posted:
18 November 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Experimental Materials and Treatment
2.2. Identification and Physicochemical Property Analysis of the AMT Gene Family in Wheat
2.3. Phylogenetic Analysis of TaAMT Proteins
2.4. Analysis of Gene Structure, Protein Features, and Cis-Regulatory Elements
2.5. Chromosomal Distribution, Gene Duplication Events, and Synteny Analysis of TaAMTs
2.6. Analysis of Tissue Expression Patterns of TaAMTs
2.7. Analysis of Expression Patterns of TaAMTs Under Different Stress Conditions
3. Results
3.1. Identification and Basic Physicochemical Properties of TaAMT Members
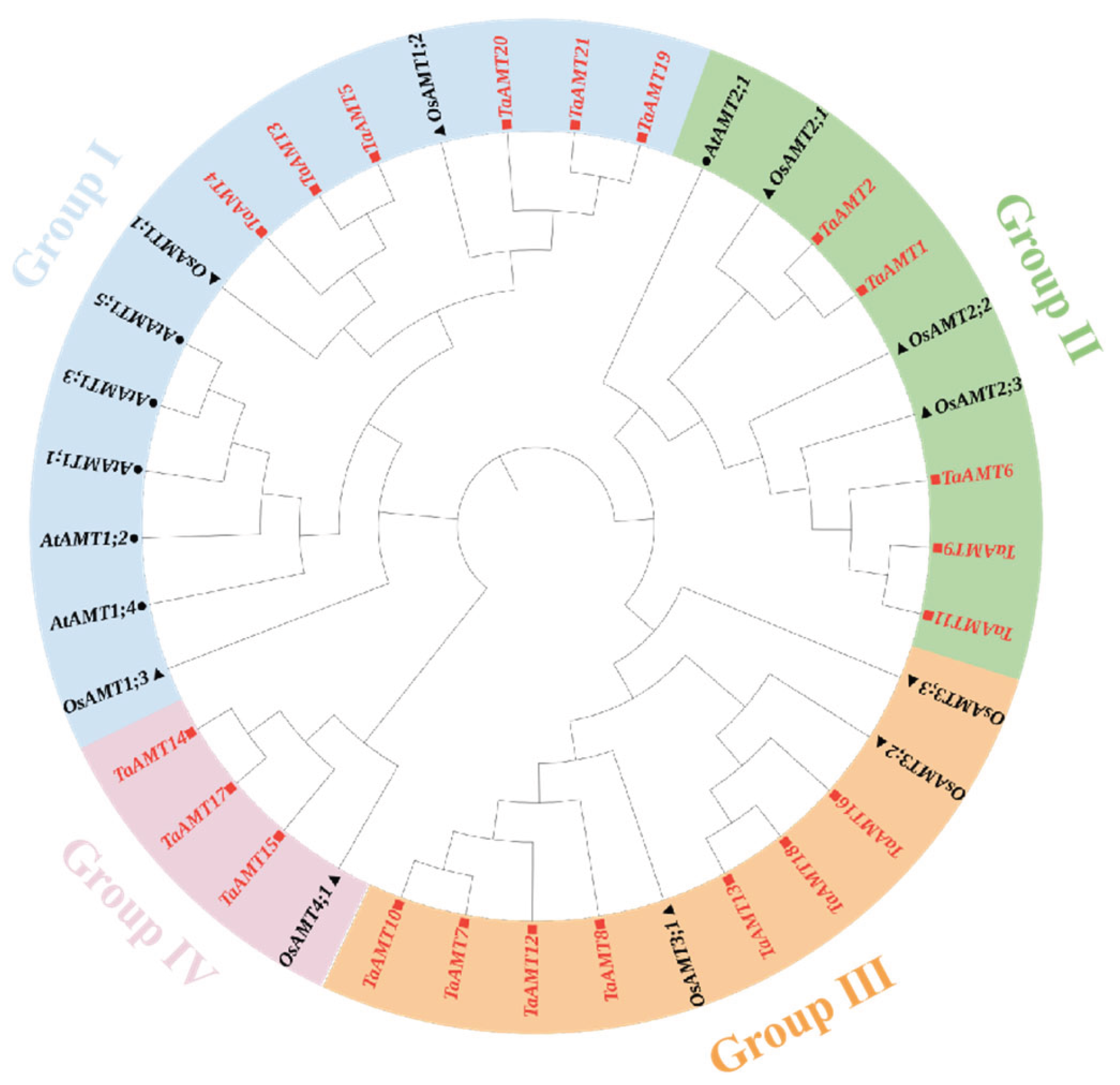
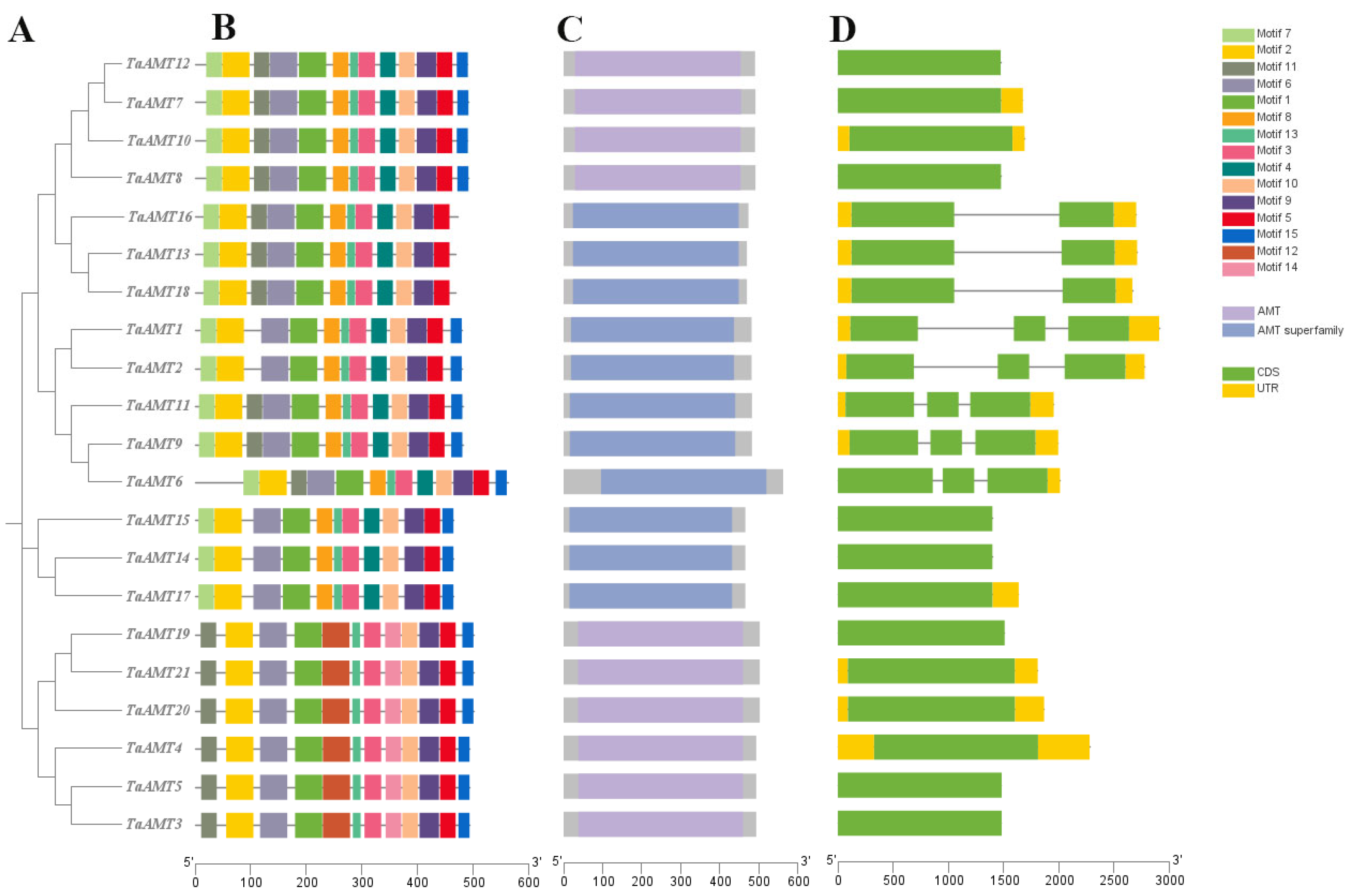
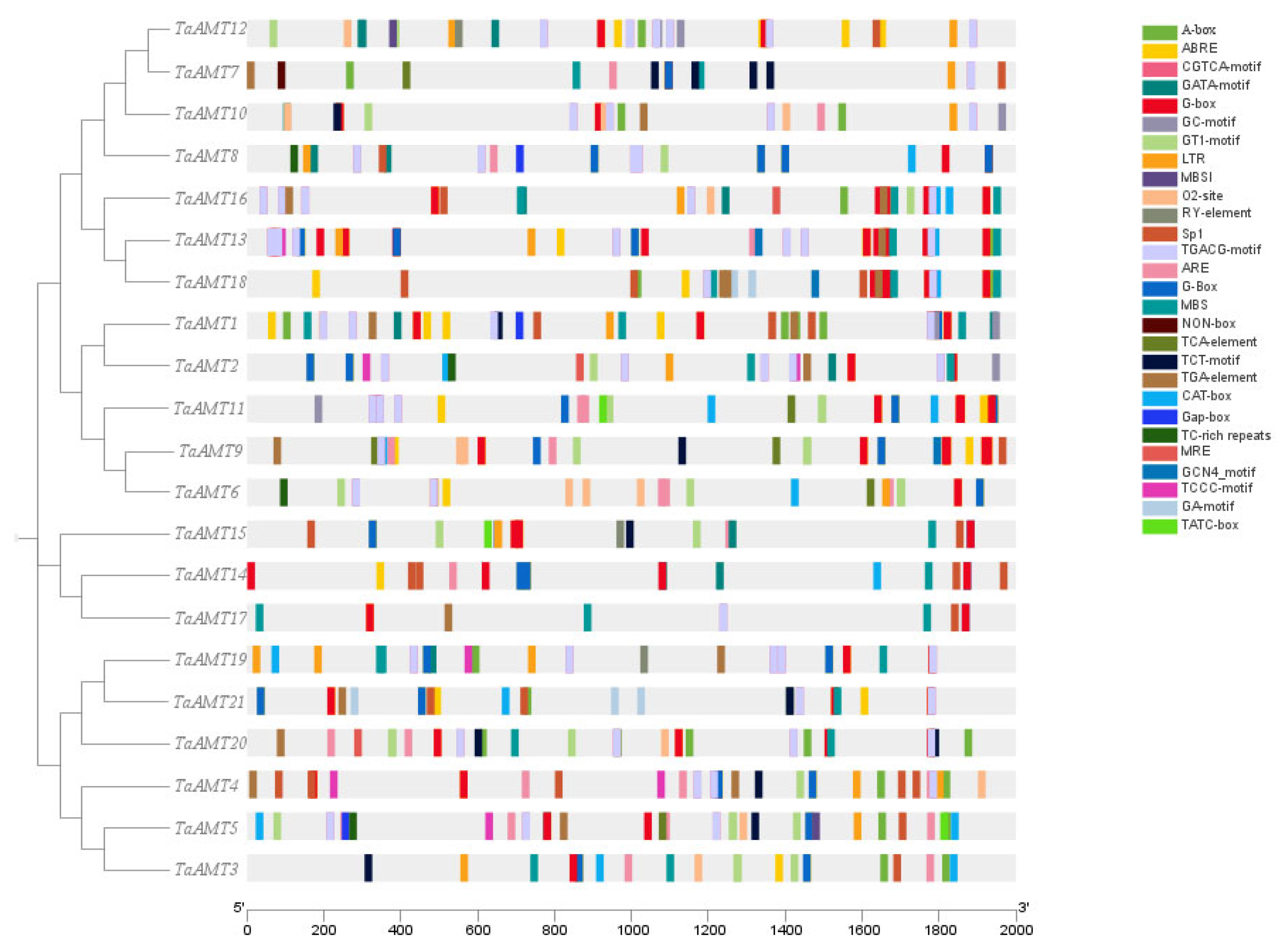
3.2. Phylogenetic Tree Analysis, Conserved Domains, and Gene Structural Features
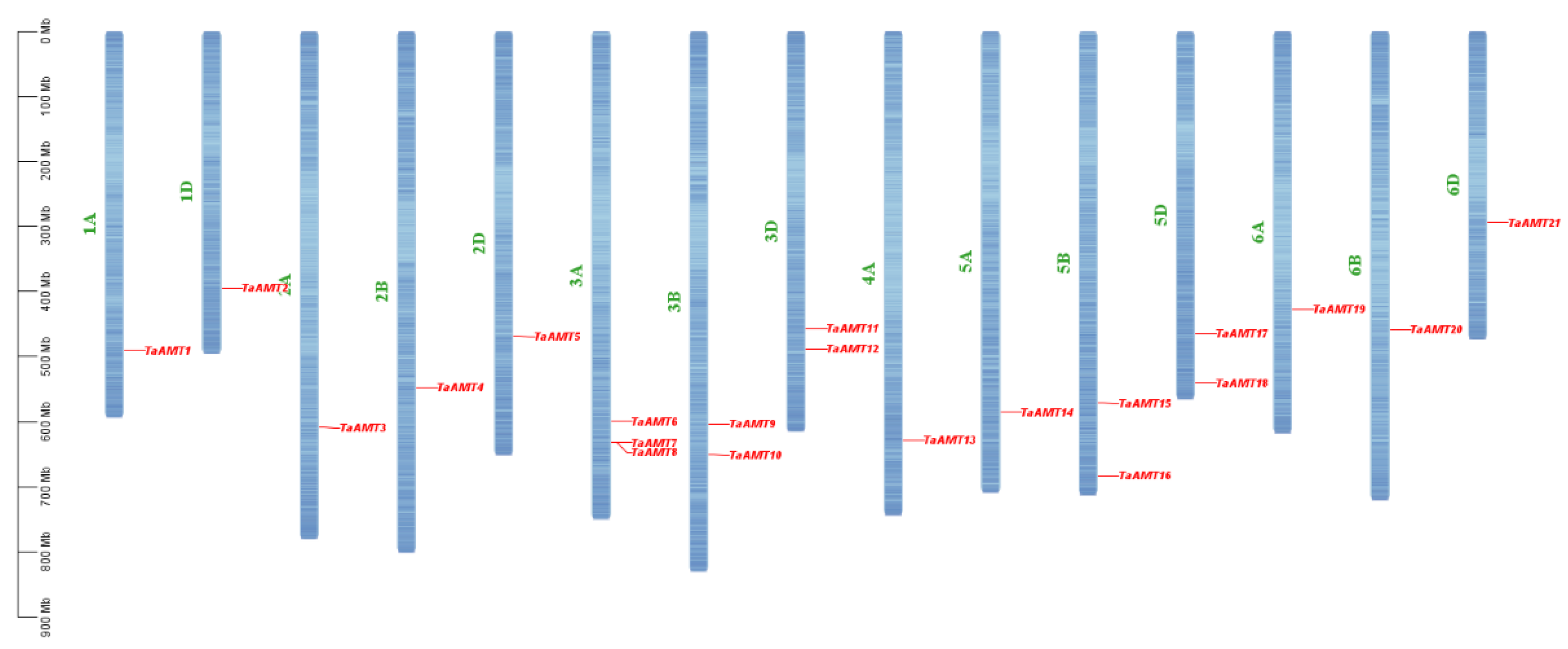
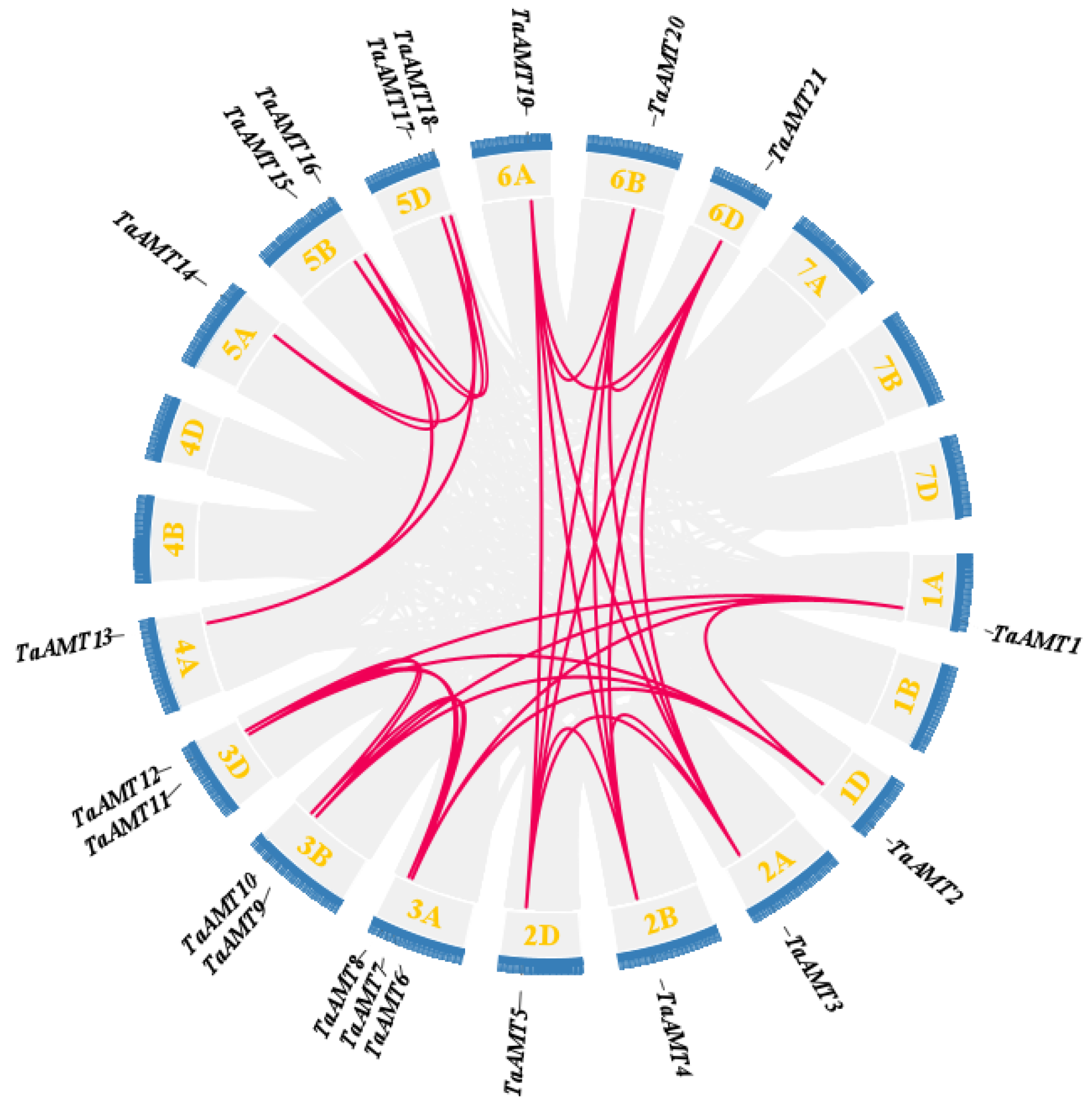
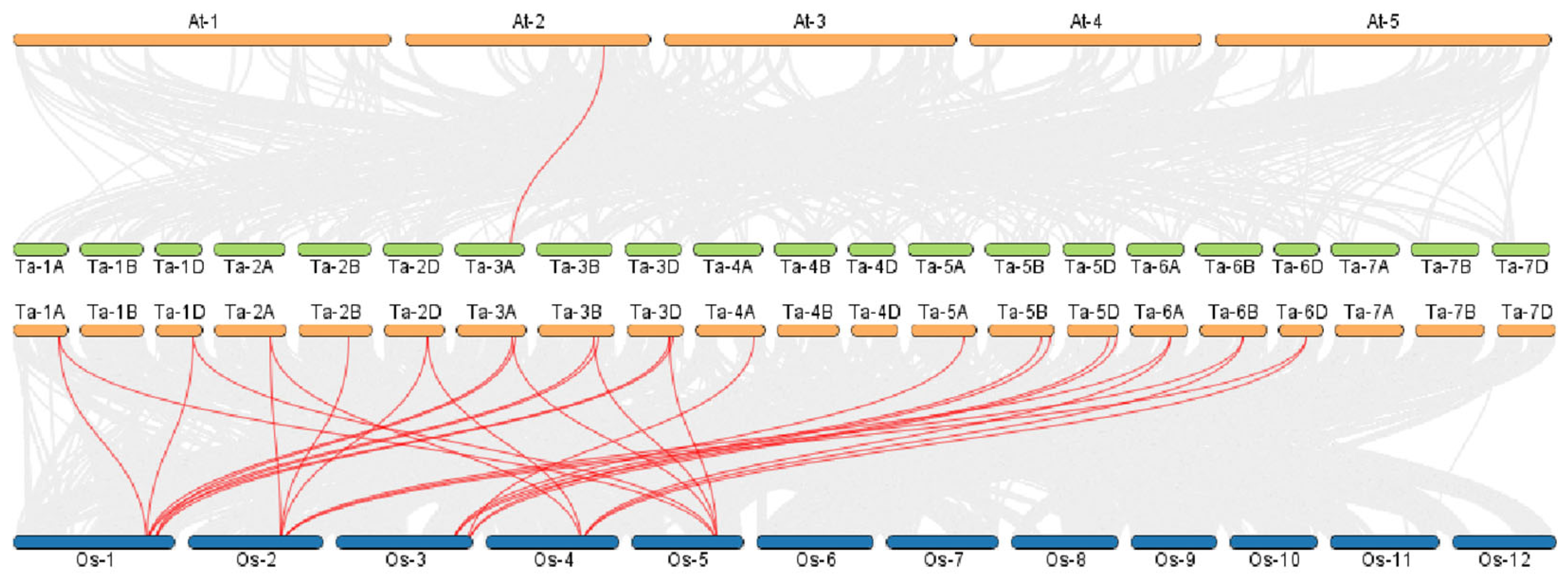
3.3. Chromosomal Localization, Synteny, and Interspecific Evolution
3.4. Cis-Acting Element Analysis
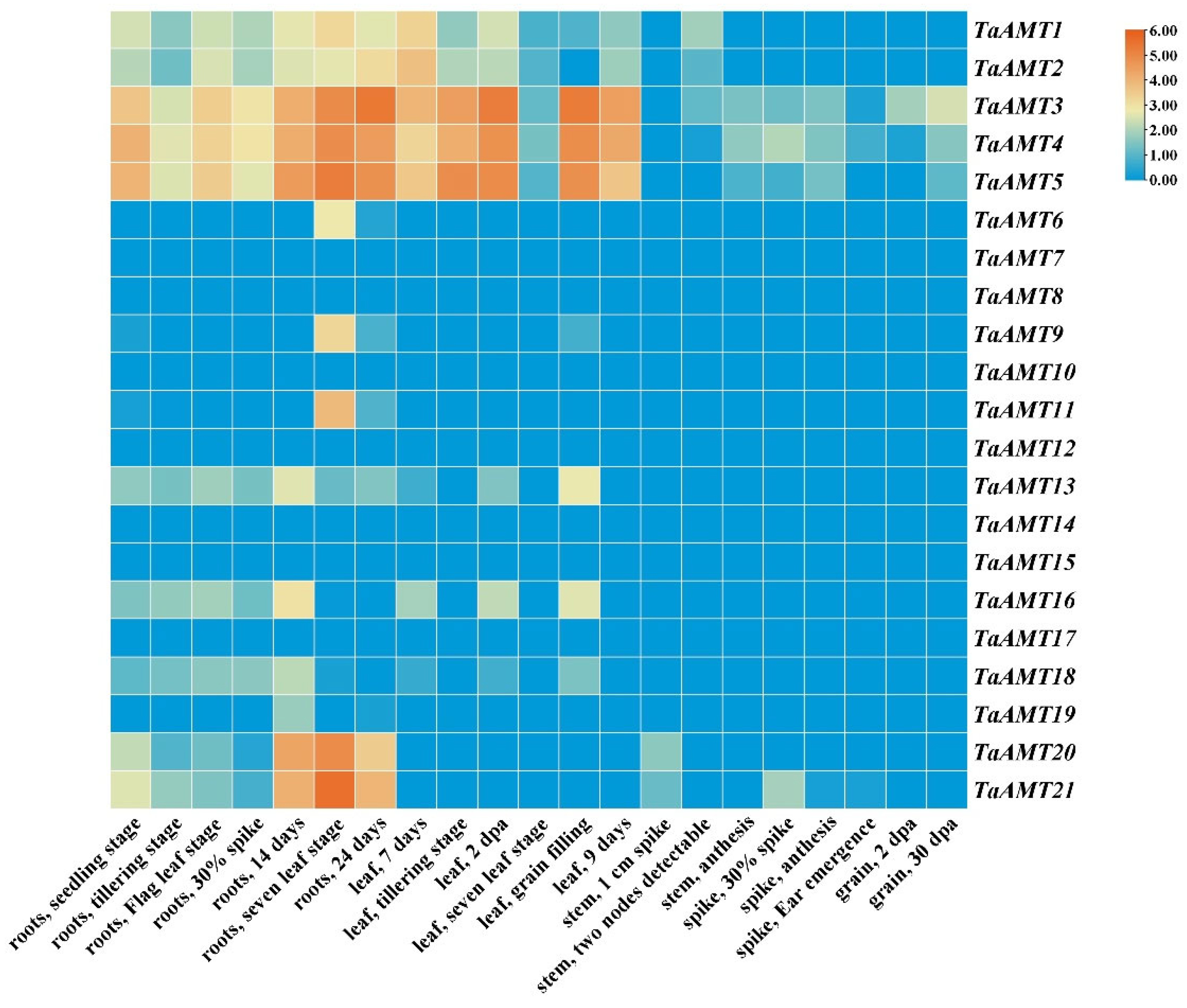
3.5. Tissue-Specific Expression Pattern Analysis
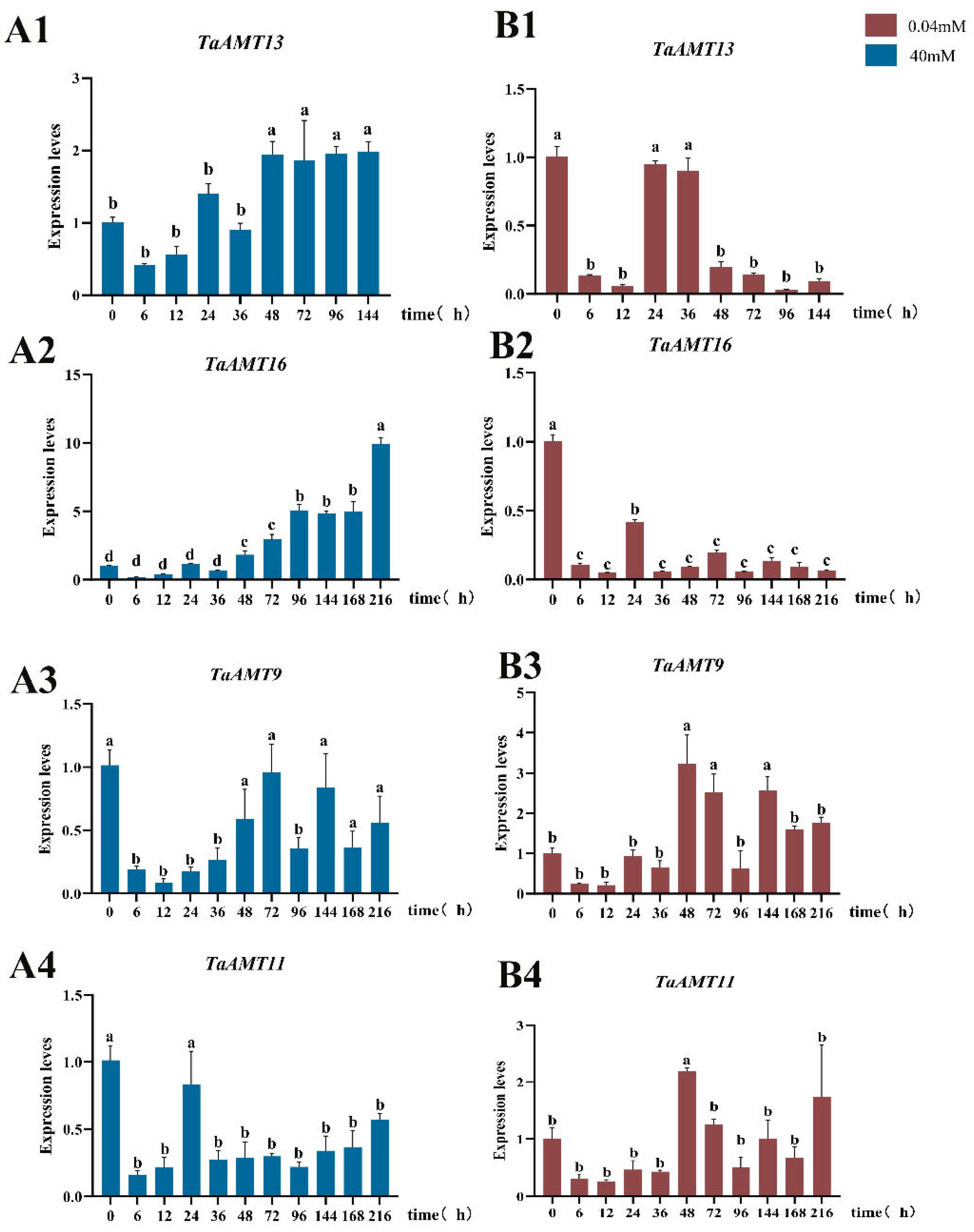
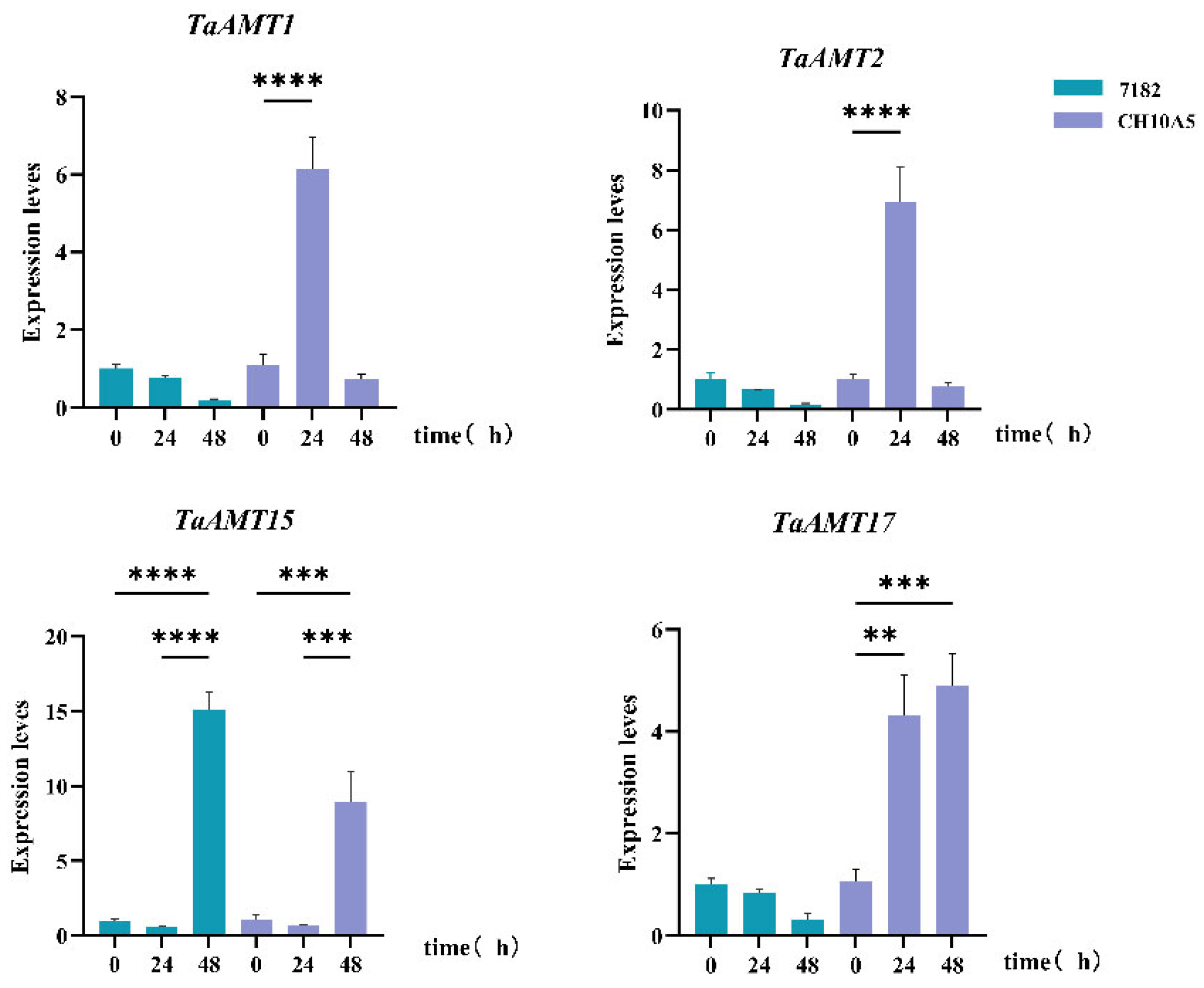
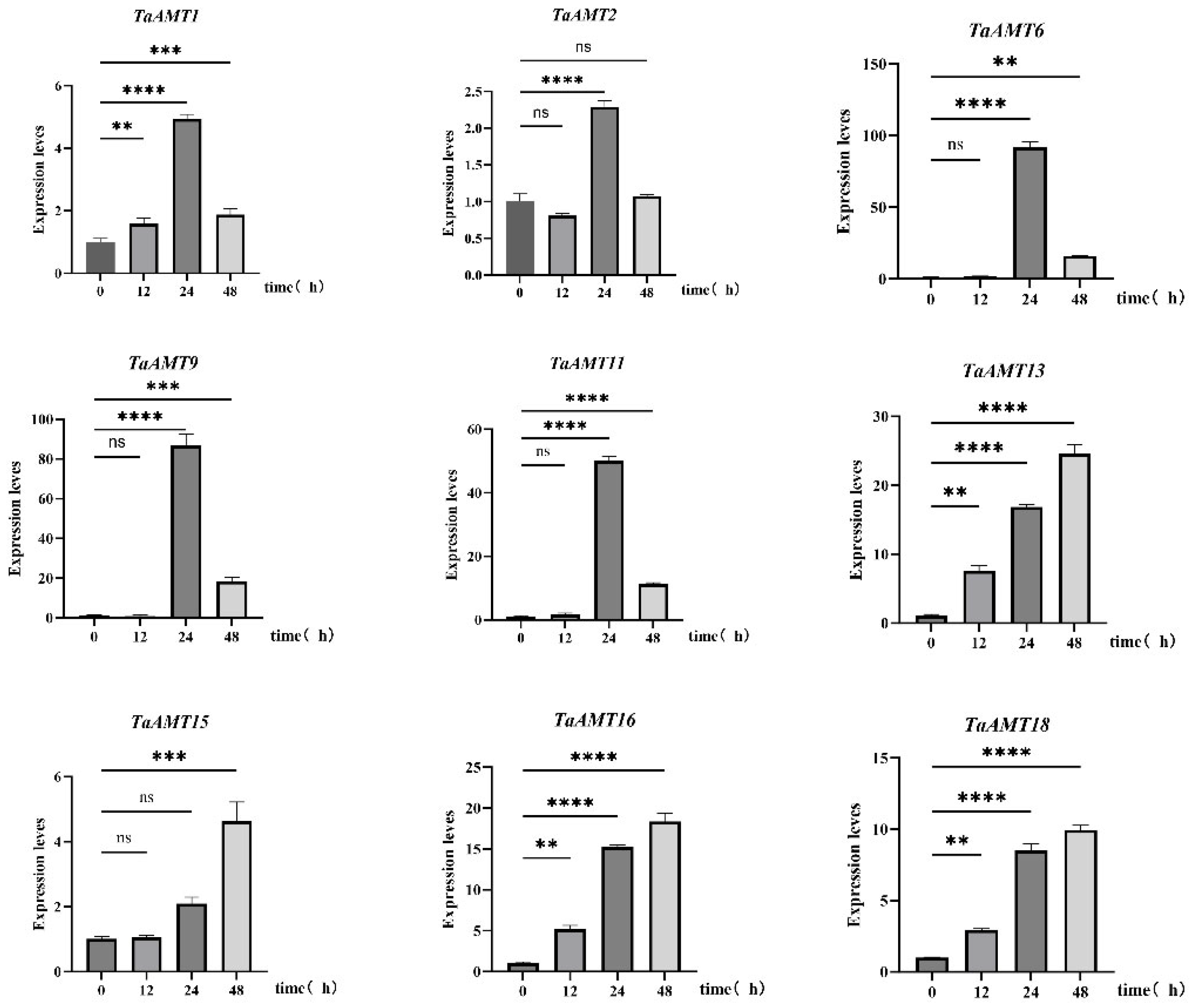
3.6. Analysis of Expression Patterns of TaAMT Members Under Three Stress Conditions
4. Discussion
4.1. Evolutionary Dynamics and Functional Divergence of TaAMT Gene Family Members
4.2. Cis-Regulatory Elements and Stress Response Mechanisms
4.3. Tissue-Specific Expression and Dynamic Regulation of Nitrogen Uptake and Utilization
4.4. Pathogen Infection Responses and the “Nitrogen-Disease” Nexus
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AMT | Ammonium transporter |
| RT-qPCR | Reverse Transcription Quantitative Polymerase Chain Reaction |
| GRAVY | Grand Average of Hydropathy |
| pI | isoelectric points |
| ABA | abscisic acid |
| MBS | Matrix Binding Site |
| LTR | Long Terminal Repeat |
| MRE | Metal Response Element |
| MBSI | Matrix Binding Site I |
Appendix A
| Gene | Forward primer sequence | Reverse primer sequence |
| TaAMT1 | GGGTGGGACCGAGGCTGAAG | CGTGATGTTGGCGGCGTAGC |
| TaAMT2 | CAGTTCGAGTTCGCCGCCATC | TGTAGGAGAGCATGAGCCAGAGC |
| TaAMT6 | GGTCCGCCCTCCTCCAGAAG | TGGGTGTGCAGGGTGGTGAG |
| TaAMT9 | CCCACGCAGCCCTACTACCC | TGAACGCCATCCACGCCTTG |
| TaAMT11 | CTGCTCCTCTCCTACACCGTCTG | ATGACGTAGCCGCCGGAGTAG |
| TaAMT13 | CGTCGTCATCCGCTTAGTTGTCC | GTGTCATCGTAGTGCTCGCCATC |
| TaAMT15 | CAGTTCGTCTTCGCCGCCATC | CCGACGGTGTAGGAGAAGGTGAG |
| TaAMT16 | TCTGTGTTGTCATCCGCCITGTC | GTGTCATCGTAGTGCTCGCCATC |
| TaAMT17 | CAGTTCGTCTTCGCCGCCATC | CCGACGGTGTAGGAGAAGGTGAG |
| TaAMT18 | GTCGTCATCCGCCTTGTCGTC | GTGTCATCGTAGTGCTCGCCATC |
| TaActin | ACCTTCAGTTGCCCAGCAAT | CAGAGTCGAGCACAATACCAGTTG |
References
- Wei, W.; Aifu, L.; Zhihua, Z.; Chengcai, C. Posttranslational Modifications: Regulation of Nitrogen Utilization and Signaling. Plant Cell Physiol 2021, 62. [CrossRef]
- Fujita, T.; Beier, M.P.; Tabuchi-Kobayashi, M.; Hayatsu, Y.; Nakamura, H.; Umetsu-Ohashi, T.; Sasaki, K.; Ishiyama, K.; Murozuka, E.; Kojima, M.; et al. Cytosolic Glutamine Synthetase GS1;3 Is Involved in Rice Grain Ripening and Germination. Front Plant Sci 2022, 13, 835835. [CrossRef]
- Oldroyd, G.E.D.; Leyser, O. A plant’s diet, surviving in a variable nutrient environment. Science 2020, 368. [CrossRef]
- Hao, D.L.; Zhou, J.Y.; Yang, S.Y.; Qi, W.; Yang, K.J.; Su, Y.H. Function and Regulation of Ammonium Transporters in Plants. Int J Mol Sci 2020, 21. [CrossRef]
- Ma, L.; Qin, D.B.; Sun, L.; Zhang, K.; Yu, X.; Dang, A.K.; Hou, S.; Zhao, X.; Yang, Y.; Wang, Y.; et al. Salt overly sensitive2 and ammonium transporter1;1 contribute to plant salt tolerance by maintaining ammonium uptake. Plant Cell 2025, 37. [CrossRef]
- Fang, L.; Wang, M.; Chen, X.; Zhao, J.; Wang, J.; Liu, J. Analysis of the AMT gene family in chili pepper and the effects of arb uscular mycorrhizal colonization on the expression patterns of CaAMT2 genes. BMC genomics 2023, 24, 158. [CrossRef]
- Chen, M.; Zhu, K.; Xie, J.; Liu, J.; Tan, P.; Peng, F. Genome-wide Identification and expression analysis of AMT and NRT gene family in Pecan (Carya illinoinensis) Seedlings Revealed a Preference for NH4+-N. Int J Mol Sci 2022, 23, 13314. [CrossRef]
- Wu, X.; Yang, H.; Qu, C.; Xu, Z.; Li, W.; Hao, B.; Yang, C.; Sun, G.; Liu, G. Sequence and expression analysis of the AMT gene family in poplar. Front Plant Sci 2015, 6, 337. [CrossRef]
- Wang, X.; Wu, H.; Manzoor, N.; Dongcheng, W.; Su, Y.; Liu, Z.; Lin, C.; Mao, Z. The identification of AMT family genes and their expression, function, and regulation in Chenopodium quinoa. Plants (Basel) 2024, 13, 3524. [CrossRef]
- Sonoda, Y.; Ikeda, A.; Saiki, S.; Yamaya, T.; Yamaguchi, J. Feedback regulation of the ammonium transporter gene family AMT1 by glutamine in rice. Plant Cell Physiol 2003, 44, 1396-1402. [CrossRef]
- Gu, R.; Duan, F.; An, X.; Zhang, F.; von Wirén, N.; Yuan, L. Characterization of AMT-mediated high-affinity ammonium uptake in roots of maize (Zea mays L.). Plant Cell Physiol 2013, 54, 1515-1524. [CrossRef]
- Hui, J.; An, X.; Li, Z.; Neuhäuser, B.; Ludewig, U.; Wu, X.; Schulze, W.X.; Chen, F.; Feng, G.; Lambers, H.; et al. The mycorrhiza-specific ammonium transporter ZmAMT3;1 mediates mycorrhiza-dependent nitrogen uptake in maize roots. Plant Cell 2022, 34, 4066-4087. [CrossRef]
- Loque, D.; Yuan, L.; Kojima, S.; Gojon, A.; Wirth, J.; Gazzarrini, S.; Ishiyama, K.; Takahashi, H.; von Wiren, N. Additive contribution of AMT1;1 and AMT1;3 to high-affinity ammonium uptake across the plasma membrane of nitrogen-deficient Arabidopsis roots. Plant J 2006, 48, 522-534. [CrossRef]
- Wang, Q.; Zhao, Y.; Luo, W.; Li, R.; He, Q.; Fang, X.; Michele, R.D.; Ast, C.; von Wirén, N.; Lin, J. Single-particle analysis reveals shutoff control of the Arabidopsis ammonium transporter AMT1;3 by clustering and internalization. Proc Natl Acad Sci U S A 2013, 110, 13204-13209. [CrossRef]
- Yuan, L.; Loque, D.; Kojima, S.; Rauch, S.; Ishiyama, K.; Inoue, E.; Takahashi, H.; von Wiren, N. The organization of high-affinity ammonium uptake in Arabidopsis roots depends on the spatial arrangement and biochemical properties of AMT1-type transporters. Plant Cell 2007, 19, 2636-2652. [CrossRef]
- von Wiren, N.; Gazzarrini, S.; Gojon, A.; Frommer, W.B. The molecular physiology of ammonium uptake and retrieval. Curr Opin Plant Biol 2000, 3, 254-261. PMID: 10837267.
- Loque, D.; Lalonde, S.; Looger, L.L.; von Wiren, N.; Frommer, W.B. A cytosolic trans-activation domain essential for ammonium uptake. Nature 2007, 446, 195-198. [CrossRef]
- Wu, X.; Liu, T.; Zhang, Y.; Duan, F.; Neuhauser, B.; Ludewig, U.; Schulze, W.X.; Yuan, L. Ammonium and nitrate regulate NH4+ uptake activity of Arabidopsis ammonium transporter AtAMT1;3 via phosphorylation at multiple C-terminal sites. J Exp Bot 2019, 70, 4919-4930. [CrossRef]
- Sasaki, A.; Ashikari, M.; Ueguchi-Tanaka, M.; Itoh, H.; Nishimura, A.; Swapan, D.; Ishiyama, K.; Saito, T.; Kobayashi, M.; Khush, G.S.; et al. Green revolution: a mutant gibberellin-synthesis gene in rice. Nature 2002, 416, 701-702. [CrossRef]
- Peng, J.; Richards, D.E.; Hartley, N.M.; Murphy, G.P.; Devos, K.M.; Flintham, J.E.; Beales, J.; Fish, L.J.; Worland, A.J.; Pelica, F.; et al. ‘Green revolution’ genes encode mutant gibberellin response modulators. Nature 1999, 400, 256-261. [CrossRef]
- Li, S.; Tian, Y.; Wu, K.; Ye, Y.; Yu, J.; Zhang, J.; Liu, Q.; Hu, M.; Li, H.; Tong, Y.; et al. Modulating plant growth-metabolism coordination for sustainable agriculture. Nature 2018, 560, 595-600. [CrossRef]
- Ray, D.K.; Sloat, L.L.; Garcia, A.S.; Davis, K.F.; Ali, T.; Xie, W. Crop harvests for direct food use insufficient to meet the UN’s food security goal. Nat Food 2022, 3, 367-374. [CrossRef]
- Wang, C.; Shen, Y.; Fang, X.T.; Xiao, S.Q.; Liu, G.Y.; Wang, L.G.; Gu, B.J.; Zhou, F.; Chen, D.L.; Tian, H.Q.; et al. Reducing soil nitrogen losses from fertilizer use in global maize and wheat production. Nature Geoscience 2024, 17, 1008-1015. [CrossRef]
- Wang, F.; Yoshida, H.; Matsuoka, M. Making the ‘Green Revolution’ truly green: improving crop nitrogen use efficiency. Plant Cell Physiol 2021, 62, 942-947. [CrossRef]
- Islam, S.; Zhang, J.; Zhao, Y.; She, M.; Ma, W. Genetic regulation of the traits contributing to wheat nitrogen use efficiency. Plant Sci 2021, 303, 110759. [CrossRef]
- Liu, Q.; Wu, K.; Song, W.; Zhong, N.; Wu, Y.; Fu, X. Improving crop nitrogen use efficiency toward sustainable green revolution. Annu Rev Plant Biol 2022, 73, 523-551. [CrossRef]
- Xu, G.; Fan, X.; Miller, A.J. Plant nitrogen assimilation and use efficiency. Annu Rev Plant Biol 2012, 63, 153-182. [CrossRef]
- Vidal, E.A.; Alvarez, J.M.; Araus, V.; Riveras, E.; Brooks, M.D.; Krouk, G.; Ruffel, S.; Lejay, L.; Crawford, N.M.; Coruzzi, G.M.; et al. Nitrate in 2020: Thirty tears from transport to signaling networks. Plant cell 2020, 32, 2094-2119. [CrossRef]
- Li, L.; Liu, K.H.; Sheen, J. Dynamic nutrient signaling networks in plants. Annu Rev Cell Dev Biol 2021, 37, 341-367. [CrossRef]
- Shanks, C.M.; Rothkegel, K.; Brooks, M.D.; Cheng, C.Y.; Alvarez, J.M.; Ruffel, S.; Krouk, G.; Gutiérrez, R.A.; Coruzzi, G.M. Nitrogen sensing and regulatory networks: it’s about time and space. Plant Cell 2024, 36, 1482-1503. [CrossRef]
- Shi, H.W.; Wang, W.C.; Gao, L.F.; Wu, J.R.; Hu, C.M.; Yan, H.S.; Shi, Y.G.; Li, N.; Ma, Y.Z.; Zhou, Y.B.; et al. Genome-wide association study of seedling nitrogen-use efficiency-associated traits in common wheat. Crop J 2024, 12, 222-231. [CrossRef]
- Wu, X.X.; Yuan, P.; Chen, H.; Kumar, V.; Kang, S.M.; Jia, B.; Xuan, Y.H. Ammonium transporter 1 increases rice resistance to sheath blight by promoting nitrogen assimilation and ethylene signalling. Plant Biotechnol J 2022, 20, 1085-1097. [CrossRef]
- Li, T.; Liao, K.; Xu, X.; Gao, Y.; Wang, Z.; Zhu, X.; Jia, B.; Xuan, Y. Wheat Ammonium Transporter (AMT) gene gamily: diversity and possible role in Host-Pathogen interaction with stem rust. Front Plant Sci 2017, 8, 1637. [CrossRef]
- Jiang, J.; Zhao, J.; Duan, W.; Tian, S.; Wang, X.; Zhuang, H.; Fu, J.; Kang, Z. TaAMT2;3a, a wheat AMT2-type ammonium transporter, facilitates the infection of stripe rust fungus on wheat. BMC Plant Biol 2019, 19, 239. [CrossRef]
- Muratore, C.; Espen, L.; Prinsi, B. Nitrogen Uptake in Plants: The plasma membrane root transport systems from a physiological and proteomic perspective. Plants (Basel) 2021, 10. [CrossRef]
- Wang Y, Cao Q, Zhang J, Wang S, Chen C, Wang C, Zhang H, Wang Y, Ji W. Cytogenetic Analysis and Molecular Marker Development for a new wheat-Thinopyrum ponticum 1Js (1D) disomic substitution line with resistance to stripe rust and powdery mildew. Front Plant Sci 2020, 11, 1282. [CrossRef]
- Bolser, D.; Staines, D.M.; Pritchard, E.; Kersey, P. Ensembl Plants: integrating tools for visualizing, mining, and analyzing plant genomics data. Methods Mol Bio 2016, 1374, 115-40. [CrossRef]
- Chenna, R.; Sugawara, H.; Koike, T.; Lopez, R.; Gibson, T.J.; Higgins, D.G.; Thompson, J.D. Multiple sequence alignment with the Clustal series of programs. Nucleic Acids Res 2003, 31, 3497-3500. [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol Biol Evol 2016, 33, 1870-1874. [CrossRef]
- Letunic, I.; Bork, P. Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res 2024, 52, W78-W82. [CrossRef]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Ren, J.Y.; Li, W.W.; Noble, W.S. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res 2009, 37, W202-W208. [CrossRef]
- Ma, S.; Wang, M.; Wu, J.; Guo, W.; Chen, Y.; Li, G.; Wang, Y.; Shi, W.; Xia, G.; Fu, D.; et al. WheatOmics: A platform combining multiple omics data to accelerate functional genomics studies in wheat. Mol Plant 2021, 14, 1965-1968. [CrossRef]
- Lescot, M.; Dehais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouze, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 2002, 30, 325-327. [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol Plant 2020, 13, 1194-1202. [CrossRef]
- Zorb, C.; Ludewig, U.; Hawkesford, M.J. Perspective on wheat yield and quality with reduced nitrogen supply. Trends Plant Sci 2018, 23, 1029-1037. [CrossRef]
- Hawkesford, M.J. Genetic variation in traits for nitrogen use efficiency in wheat. J Exp Bot 2017, 68, 2627-2632. [CrossRef]
- Xu, G.; Fan, X.; Miller, A.J. Plant nitrogen assimilation and use efficiency. Annu Rev Plant Biol 2012, 63, 153-182. [CrossRef]
- Li, H.; Zhu, X.; Wang, J.; Wei, Y.; Nai, F.; Yu, H.; Wang, X. Unraveling differential characteristics and mechanisms of nitrogen uptake in wheat cultivars with varied nitrogen use efficiency. Plant Physiol Biochem 2024, 206, 108278. [CrossRef]
- Keerthana, K.; Ramakrishnan, M.; Ahmad, Z.; Amali, P.; Vijayakanth, V.; Wei, Q. Root-derived small peptides: Key regulators of plant development, stress resilience, and nutrient acquisition. Plant Sci 2025, 354, 112433. [CrossRef]
- Tsay, Y.-F.; Ho, C.-H.; Chen, H.-Y.; Lin, S.-H. Integration of Nitrogen and Potassium Signaling. Annual Review of Plant Biology 2011, 62, 207-226. [CrossRef]
- Wang, H.; Ma, Q.; Shan, F.; Tian, L.; Gong, J.; Quan, W.; Yang, W.; Hou, Q.; Zhang, F.; Zhang, S. Transcriptional regulation mechanism of wheat varieties with different nitrogen use efficiencies in response to nitrogen deficiency stress. BMC Genomics 2022, 23, 727. [CrossRef]
- Zhang, Y.; Zhao, Y.; Sun, L.; Han, P.; Bai, X.; Lin, R.; Xiao, K. The N uptake-associated physiological processes at late growth stage in wheat (Triticum aestivum) under N deprivation combined with deficit irrigation condition. Plant Physiol Biochem 2021, 164, 160-172. [CrossRef]
- Liu, B.; Xu, W.; Niu, Y.; Li, Q.; Cao, B.; Qi, J.; Zhao, Y.; Zhou, Y.; Song, L.; Cui, D.; et al. TaTCP6 is required for efficient and balanced utilization of nitrate and phosphorus in wheat. Nat Commun 2025, 16, 1683. [CrossRef]
- Zhang, H.; Jin, Z.; Cui, F.; Zhao, L.; Zhang, X.; Chen, J.; Zhang, J.; Li, Y.; Li, Y.; Niu, Y.; et al. Epigenetic modifications regulate cultivar-specific root development and metabolic adaptation to nitrogen availability in wheat. Nat Commun 2023, 14, 8238. [CrossRef]
- International Wheat Genome Sequencing, C. Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 2018, 361, eaar7191. [CrossRef]
- Li, T.; Kong, C.; Deng, P.; Li, C.; Zhao, G.; Li, H.; Gao, L.; Cui, D.; Jia, J. Intra-Varietal diversity and its contribution to wheat evolution, domestication, and improvement in wheat. Int J Mol Sci 2023, 24. [CrossRef]
- Howitt, S.M.; Udvardi, M.K. Structure, function and regulation of ammonium transporters in plants. Biochim Biophys Acta 2000, 1465, 152-170. [CrossRef]
- Hao, D.L.; Yang, S.Y.; Liu, S.X.; Zhou, J.Y.; Huang, Y.N.; Very, A.A.; Sentenac, H.; Su, Y.H. Functional Characterization of the Arabidopsis Ammonium Transporter AtAMT1;3 With the Emphasis on Structural Determinants of Substrate Binding and Permeation Properties. Front Plant Sci 2020, 11, 571. [CrossRef]
- Zhu, J.K. Abiotic Stress Signaling and Responses in Plants. Cell 2016, 167, 313-324. [CrossRef]
- Chinnusamy, V.; Schumaker, K.; Zhu, J.K. Molecular genetic perspectives on cross-talk and specificity in abiotic stress signalling in plants. J Exp Bot 2004, 55, 225-236. [CrossRef]
- Ma, X.; Wang, W.; Zhang, J.; Jiang, Z.; Xu, C.; Zhu, W.; Shi, B.; Yang, W.; Su, H.; Wang, X.; et al. NRT1.1B acts as an abscisic acid receptor in integrating compound environmental cues for plants. Cell 2025. [CrossRef]
- Garnica, M.; Houdusse, F.; Zamarreño, A.M.; Garcia-Mina, J.M. The signal effect of nitrate supply enhances active forms of cytokinins and indole acetic content and reduces abscisic acid in wheat plants grown with ammonium. J Plant Physiol 2010, 167, 1264-1272. [CrossRef]
- Oka, M.; Shimoda, Y.; Sato, N.; Inoue, J.; Yamazaki, T.; Shimomura, N.; Fujiyama, H. Abscisic acid substantially inhibits senescence of cucumber plants (Cucumis sativus) grown under low nitrogen conditions. J Plant Physiol 2012, 169, 789-796, doi:/10.1016/j.jplph.2012.02.001.
- Liu, H.; Carvalhais, L.C.; Kazan, K.; Schenk, P.M. Development of marker genes for jasmonic acid signaling in shoots and roots of wheat. Plant Signal Behav 2016, 11, e1176654. [CrossRef]
- Fernandez-Milmanda, G.L.; Crocco, C.D.; Reichelt, M.; Mazza, C.A.; Kollner, T.G.; Zhang, T.; Cargnel, M.D.; Lichy, M.Z.; Fiorucci, A.S.; Fankhauser, C.; et al. A light-dependent molecular link between competition cues and defence responses in plants. Nat Plants 2020, 6, 223-230. [CrossRef]
- Kumar, A.; Silim, S.N.; Okamoto, M.; Siddiqi, M.Y.; Glass, A.D. Differential expression of three members of the AMT1 gene family encoding putative high-affinity NH4+ transporters in roots of Oryza sativa subspecies indica. Plant Cell Environ 2003, 26, 907-914. [CrossRef]
- Lu, Y.; Gao, L.; Hu, J.; Liu, X.; Jiang, D.; Cao, W.; Dai, T.; Tian, Z. Low nitrogen priming improves nitrogen uptake and assimilation adaptation to nitrogen deficit stress in wheat seedling. Planta 2024, 259, 107. [CrossRef]
- Liu, Y.; von Wiren, N. Ammonium as a signal for physiological and morphological responses in plants. J Exp Bot 2017, 68, 2581-2592. [CrossRef]
- Zhao, R.; Cao, Y.; Ge, Y.; Xu, J.; Li, R.; Yang, M.; Chen, Y.; Wu, D.; Xiao, J.; Li, R. Single-Molecule and Vesicle Trafficking Analysis of Ubiquitination Involved in the Activity of Ammonium Transporter AMT1;3 in Arbidopsis under high ammonium Stress. Cells-Basel 2022, 11. [CrossRef]
- Devadas, R.; Simpfendorfer, S.; Backhouse, D.; Lamb, D.W. Effect of stripe rust on the yield response of wheat to nitrogen. Crop J 2014, 2, 201-206. [CrossRef]
- Bolton, M.D.; Thomma, B.P.H.J. The complexity of nitrogen metabolism and nitrogen-regulated gene expression in plant pathogenic fungi. Physiol Mol Plant P 2008, 72, 104-110. [CrossRef]
- Sun, Y.; Wang, M.; Mur, L.A.J.; Shen, Q.; Guo, S. Unravelling the roles of nitrogen nutrition in plant disease defences. Int J Mol Sci 2020, 21. [CrossRef]
| Name | Number of Amino Acid | Molecular Weight | Theoretical pI | Instability Index | Aliphatic Index | Grand Average of Hydropathicity | Subcellular Localization |
| TaAMT1 | 482 | 51273.1 | 7.75 | 28.77 | 102.45 | 0.541 | Plasma Membrane |
| TaAMT2 | 482 | 51336.2 | 8.67 | 29.15 | 101.43 | 0.537 | Plasma Membrane |
| TaAMT3 | 494 | 52190.17 | 7.15 | 19.04 | 94.11 | 0.520 | Plasma Membrane |
| TaAMT4 | 494 | 52298.33 | 7.15 | 19.59 | 93.32 | 0.523 | Plasma Membrane |
| TaAMT5 | 494 | 52224.19 | 7.15 | 19.04 | 93.32 | 0.518 | Plasma Membrane |
| TaAMT6 | 563 | 59889.76 | 9.13 | 38.08 | 100.36 | 0.427 | Plasma Membrane |
| TaAMT7 | 492 | 53474.26 | 7.63 | 34.41 | 96.30 | 0.420 | Plasma Membrane |
| TaAMT8 | 492 | 53458.29 | 7.10 | 34.55 | 97.13 | 0.452 | Plasma Membrane |
| TaAMT9 | 483 | 51327.01 | 8.28 | 31.13 | 107.52 | 0.583 | Plasma Membrane |
| TaAMT10 | 491 | 53372.22 | 7.63 | 33.25 | 96.90 | 0.434 | Plasma Membrane |
| TaAMT11 | 483 | 51280.94 | 8.28 | 31.53 | 107.70 | 0.580 | Plasma Membrane |
| TaAMT12 | 491 | 53385.21 | 7.63 | 34.29 | 96.90 | 0.428 | Plasma Membrane |
| TaAMT13 | 470 | 50042.43 | 6.49 | 34.26 | 103.11 | 0.625 | Plasma Membrane |
| TaAMT14 | 466 | 50040.55 | 7.61 | 33.81 | 102.62 | 0.613 | Plasma Membrane |
| TaAMT15 | 466 | 49917.42 | 7.07 | 32.81 | 104.72 | 0.630 | Plasma Membrane |
| TaAMT16 | 474 | 50497.9 | 6.30 | 32.68 | 102.05 | 0.596 | Plasma Membrane |
| TaAMT17 | 466 | 49910.41 | 7.61 | 32.72 | 102.83 | 0.626 | Plasma Membrane |
| TaAMT18 | 470 | 50056.46 | 6.49 | 34.08 | 103.11 | 0.625 | Plasma Membrane |
| TaAMT19 | 503 | 52851.72 | 7.62 | 23.94 | 91.87 | 0.514 | Plasma Membrane |
| TaAMT20 | 503 | 52865.74 | 7.62 | 23.94 | 92.07 | 0.514 | Plasma Membrane |
| TaAMT21 | 503 | 52865.1 | 7.62 | 24.10 | 91.67 | 0.509 | Plasma Membrane |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




