Submitted:
14 November 2025
Posted:
14 November 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Data Sets of Protein Kinase Inhibitors and Small Molecules
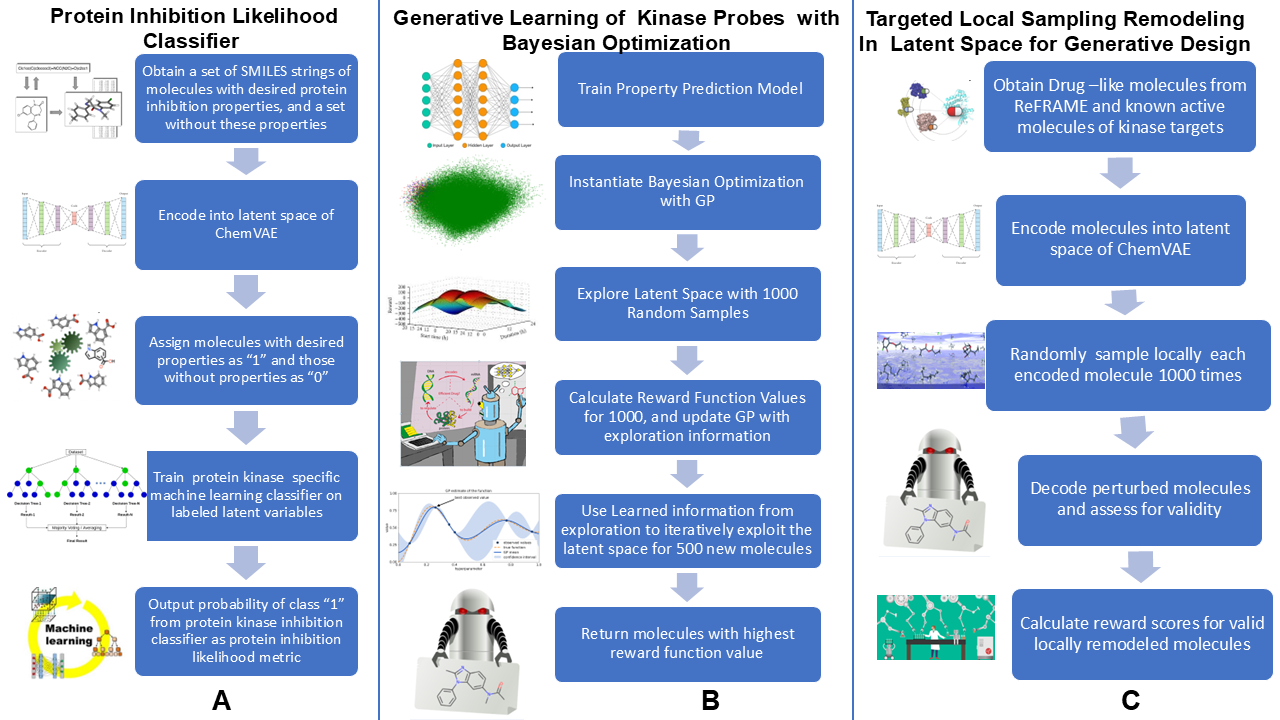
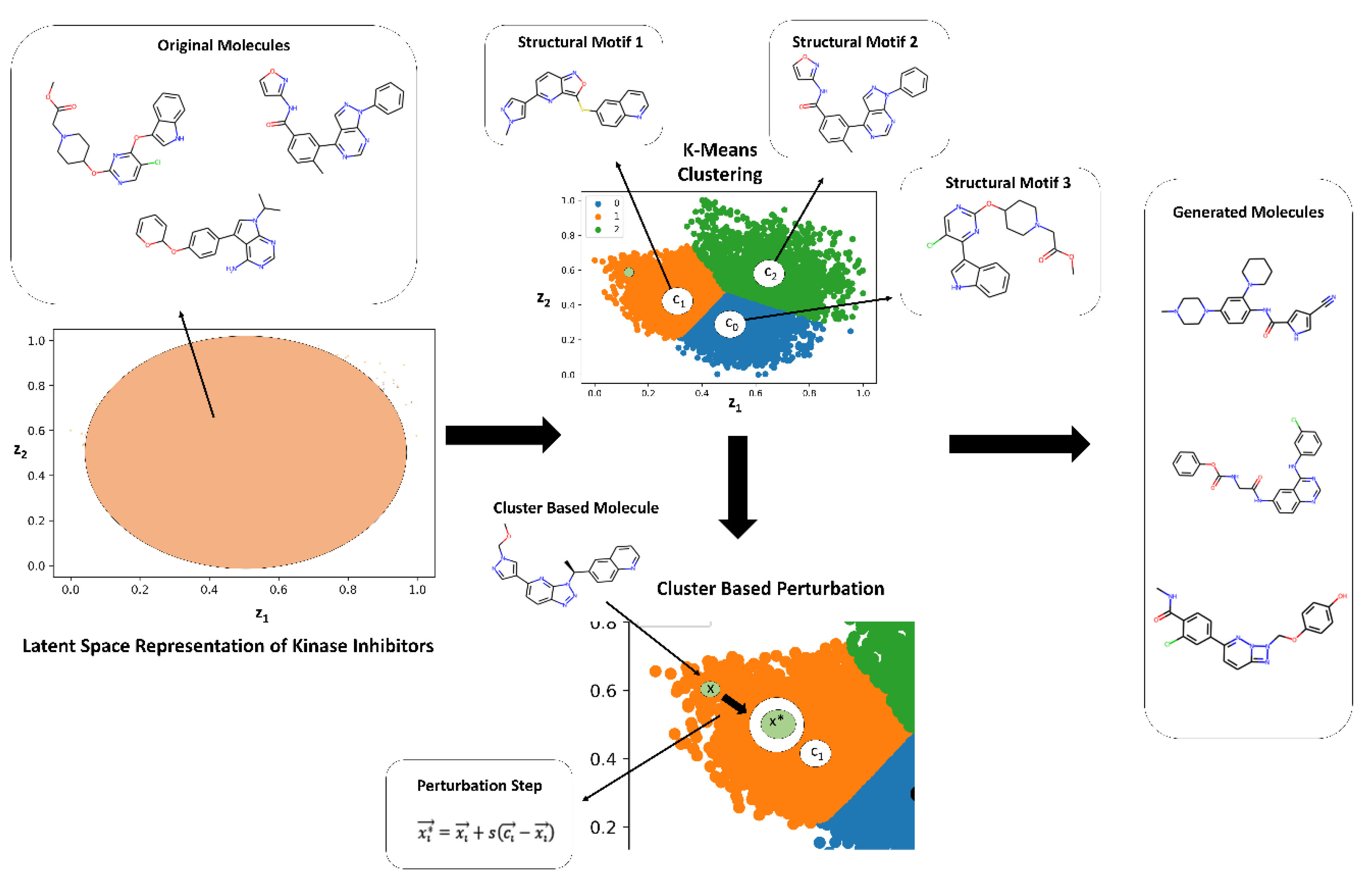
2.2. Guided Remodeling of Latent Neighborhoods via Cluster-Directed Sampling
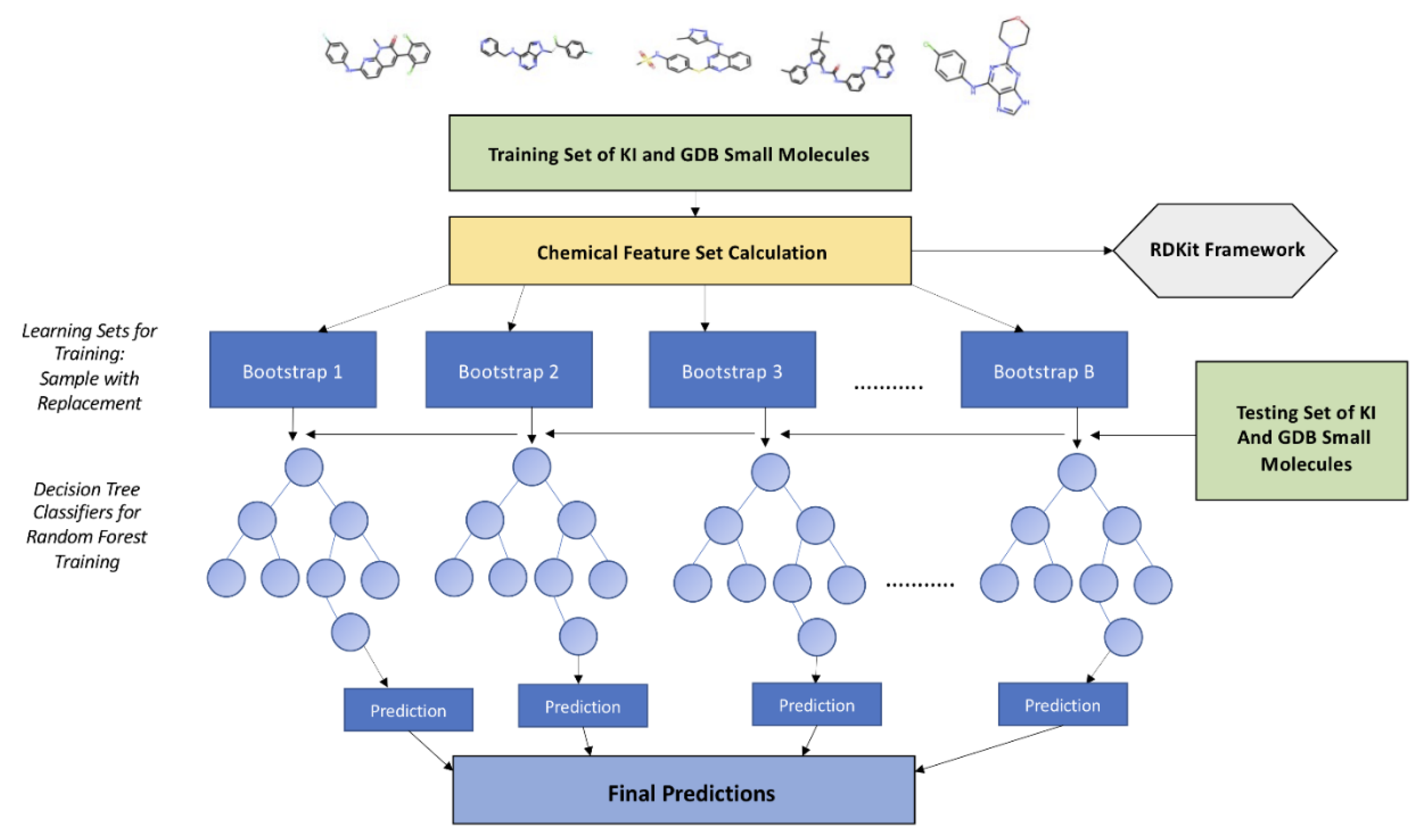
2.3. Kinase Inhibition Likelihood Classifiers
3. Results and Discussion
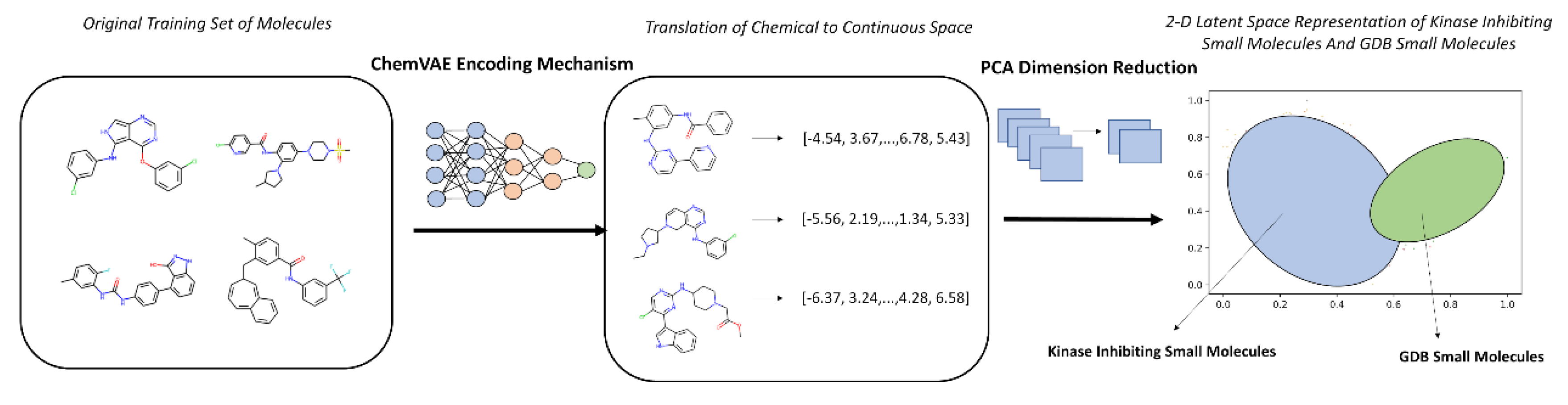
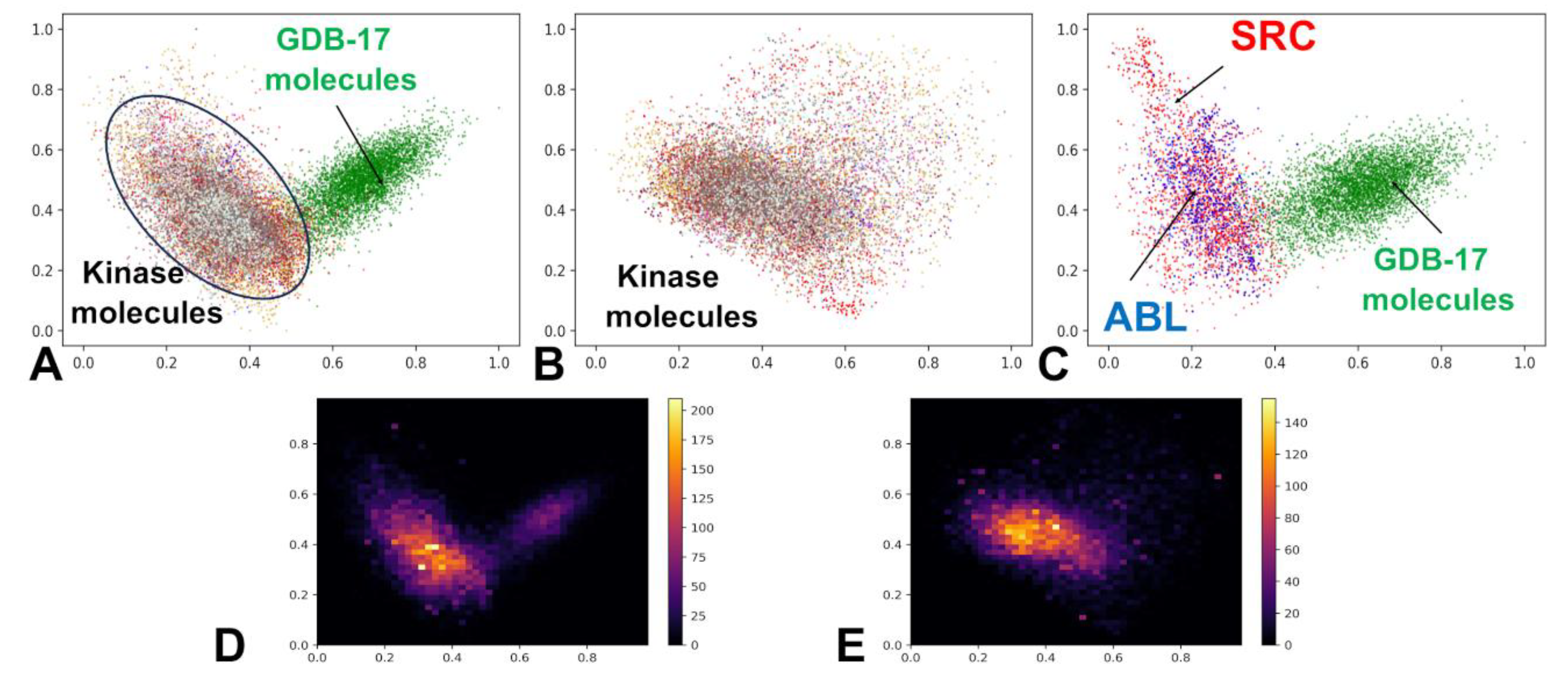
3.1. The Kinase Inhibitor Dataset and Its Embedding Reveals Organized Kinome Manifold in Latent Space
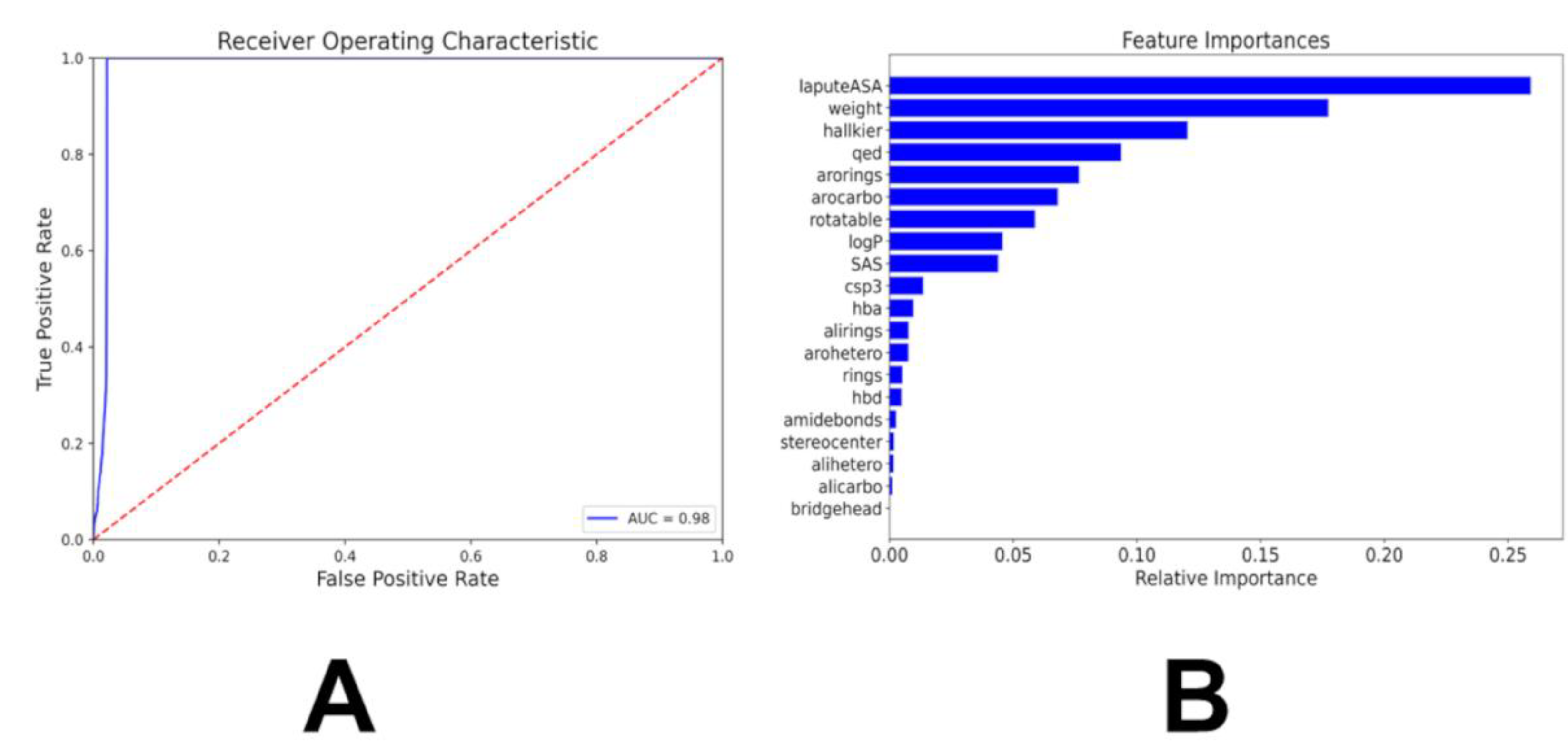
3.2. Multiclass and Binary Kinase Inhibition Likelihood Classifiers
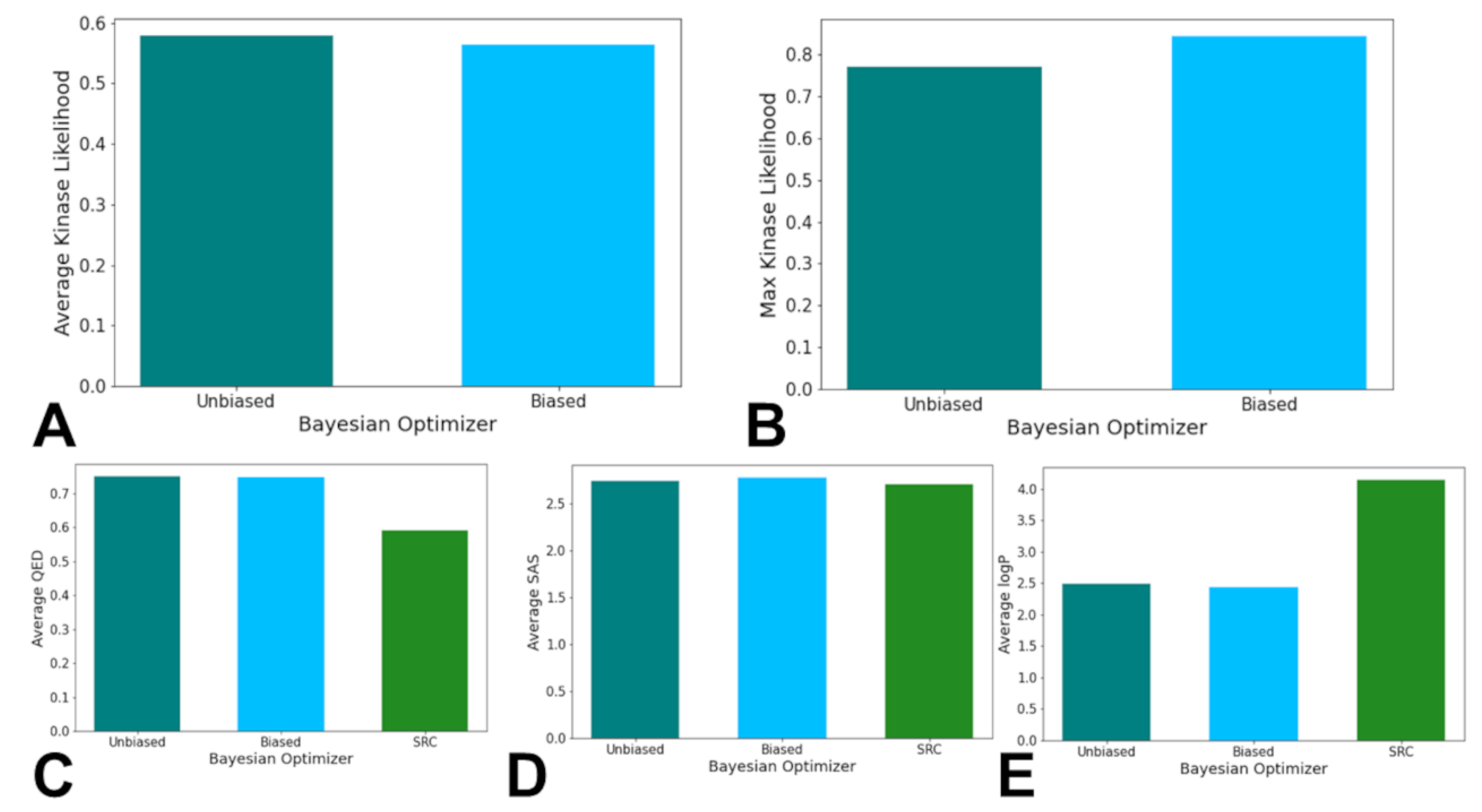
3.3. Bayesian Optimization Enables Efficient Exploration of SRC Kinase Inhibitor Chemical Space
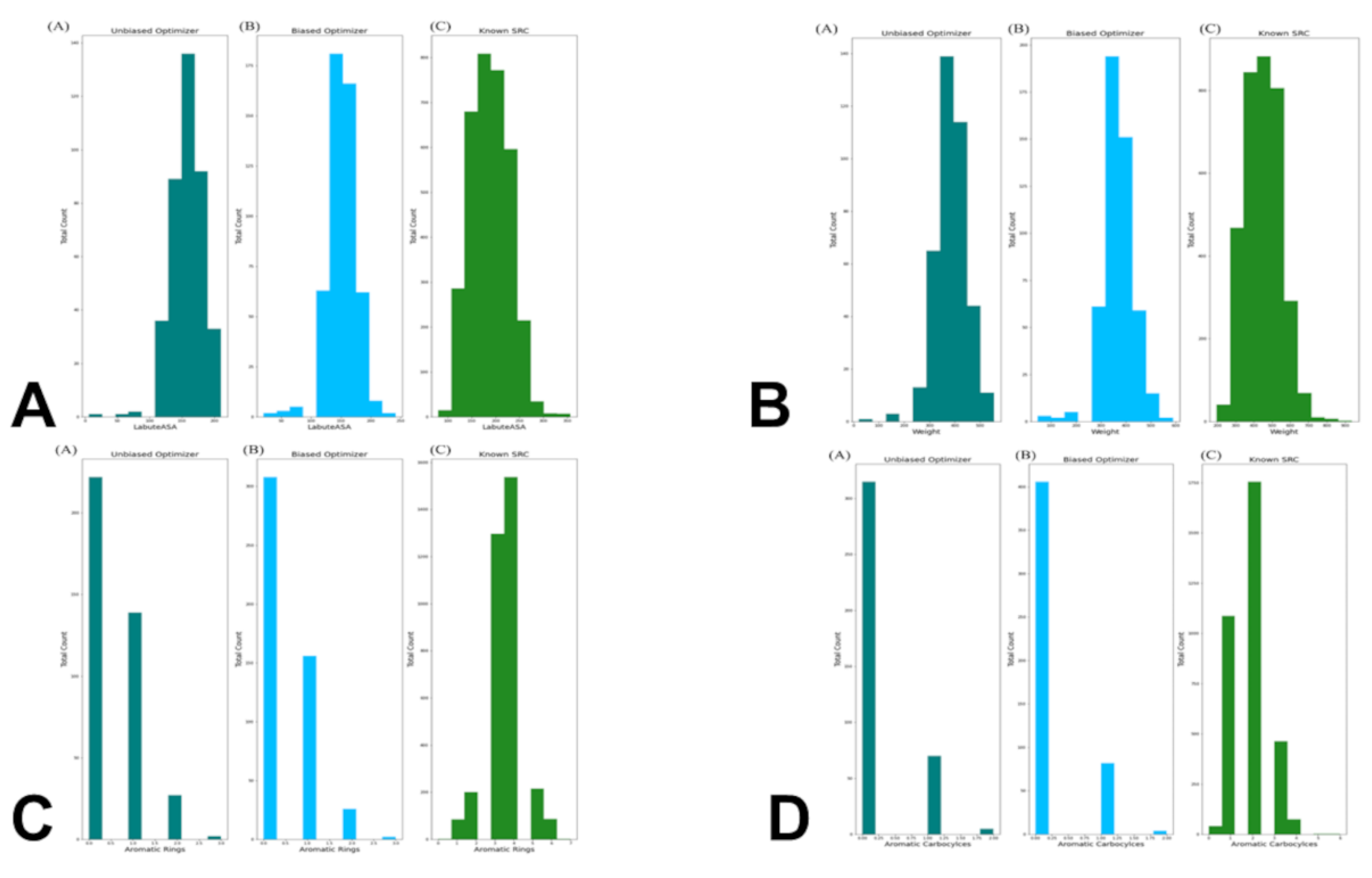
3.4. Targeted Local Latent Neighborhood Sampling Recovers Pharmacophoric Complexity
3.5. Computational Docking Validation of Generated Molecules Reveals High-Affinity Binding to the SRC Kinase Active Site
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- White, D.; Wilson, R. C. Generative models for chemical structures. J. Chem. Inf. Model. 2010, 50, 1257−1274. [CrossRef]
- Goh, G. B.; Hodas, N. O.; Vishnu, A. Deep learning for computational chemistry. J Comput Chem 2017, 38, 1291-1307. [CrossRef]
- Mater, A. C.; Coote, M. L., Deep Learning in Chemistry. J Chem Inf Model 2019, 59, 2545-2559. [CrossRef]
- Chen, H.; Engkvist, O.; Wang, Y.; Olivecrona, M.; Blaschke, T. The rise of deep learning in drug discovery. Drug Discov Today 2018, 23, 1241-1250. [CrossRef]
- Popova, M.; Isayev, O.; Tropsha, A. Deep reinforcement learning for de novo drug design. Sci Adv 2018, 4, eaap7885. [CrossRef]
- Dimitrov, T.; Kreisbeck, C.; Becker, J. S.; Aspuru-Guzik, A.; Saikin, S. K. Autonomous Molecular Design: Then and Now. ACS Appl Mater Interfaces 2019, 11, 24825-24836. [CrossRef]
- Korotcov, A.; Tkachenko, V.; Russo, D. P.; Ekins, S. Comparison of Deep Learning With Multiple Machine Learning Methods and Metrics Using Diverse Drug Discovery Data Sets. Mol Pharm 2017, 14, 4462-4475. [CrossRef]
- Sanchez-Lengeling, B.; Aspuru-Guzik, A. Inverse molecular design using machine learning: Generative models for matter engineering. Science 2018, 361, 360−365. [CrossRef]
- Gomez-Bombarelli, R.; Wei, J. N.; Duvenaud, D.; Hernández- Lobato, J. M.; S ́anchez- Lengeling, B.; Sheberla, D.; Aguilera- Iparraguirre, J.; Hirzel, T. D.; Adams, R. P.; Aspuru-Guzik, A. Automatic Chemical Design Using a Data-Driven Continuous Representation of Molecules. ACS Cent. Sci. 2018, 4, 268−276. [CrossRef]
- Yuan, W.; Jiang, D.; Nambiar, D. K.; Liew, L. P.; Hay, M. P.; Bloomstein, J.; Lu, P.; Turner, B.; Le, Q.-T.; Tibshirani, R.; Khatri, P.; Moloney, M. G.; Koong, A. C. Chemical Space Mimicry for Drug Discovery. J. Chem. Inf. Model. 2017, 57, 875−882. [CrossRef]
- Segler, M. H.; Kogej, T.; Tyrchan, C.; Waller, M. P. Generating focused molecule libraries for drug discovery with recurrent neural networks. ACS Cent. Sci. 2018, 4, 120−131. [CrossRef]
- Elton, D. C.; Boukouvalas, Z.; Fuge, M. D.; Chung, P. W. Deep learning for molecular design - a review of the state of the art. Mol. Syst. Des. Eng. 2019, 4, 828−849. [CrossRef]
- Bian, Y.; Xie, X.-Q. Generative chemistry: drug discovery with deep learning generative models. J. Mol. Model. 2021, 27, 71. [CrossRef]
- Chen, H.; Engkvist, O.; Wang, Y.; Olivecrona, M.; Blaschke, T. The rise of deep learning in drug discovery. Drug Discov. Today 2018, 23, 1241−1250. [CrossRef]
- Vamathevan, J.; Clark, D.; Czodrowski, P.; Dunham, I.; Ferran, E.; Lee, G.; Li, B.; Madabhushi, A.; Shah, P.; Spitzer, M.; Zhao, S. Applications of machine learning in drug discovery and development. Nat. Rev. Drug Discov. 2019, 18, 463−477. [CrossRef]
- Sousa, T.; Correia, J.; Pereira, V.; Rocha, M. Generative Deep Learning for Targeted Compound Design. J. Chem. Inf. Model. 2021, 61, 5343-5361. [CrossRef]
- Goodfellow, I.; Pouget-Abadie, J.; Mirza, M.; Xu, B.; Warde-Farley, D.; Ozair, S.; Courville, A.; Bengio, Y. Generative Adversarial Nets. Adv. Neural Inf. Proc. Syst. 2014, 2672−2680.
- Yu, L.; Zhang, W.; Wang, J.; Yu, Y. SeqGAN: Sequence Generative Adversarial Nets With Policy Gradient. Proceedings of the Thirty-First AAAI Conference on Artificial Intelligence (AAAI-17), 2017, pp 2852−2858.
- Guimaraes G. L.; Sanchez-Lengeling B.; Outeiral C.; Farias P. L. C.; Aspuru-Guzik A. Objective-reinforced generative adversarial networks (ORGAN) for sequence generation models. arXiv, 2017, 1705.10843.
- Olivecrona, M.; Blaschke, T.; Engkvist, O.; Chen, H. Molecular de-novo design through deep reinforcement learning. J Cheminform 2017, 9, 48. [CrossRef]
- Sanchez-Lengeling, B.; Outeiral, C.; L, G.; Aspuru-Guzik, A. Optimizing distributions over molecular space. An Objective-Reinforced Generative Adversarial Network for Inverse-design Chemistry (ORGANIC). ChemRxiv.org 2017, 5309668. [CrossRef]
- Prykhodko, O.; Johansson, S.V.; Kotsias, P.C.; Arús-Pous, J.; Bjerrum, E.J.; Engkvist, O.; Chen, H. A de novo molecular generation method using latent vector based generative adversarial network. J. Cheminform. 2019, 11, 74. [CrossRef]
- Kadurin, A.; Nikolenko, S.; Khrabrov, K.; Aliper, A.; Zhavoronkov, A. druGAN: An Advanced Generative Adversarial Autoencoder Model for de Novo Generation of New Molecules with Desired Molecular Properties in Silico. Mol. Pharm. 2017, 14, 3098-3104. [CrossRef]
- Putin, E.; Asadulaev, A.; Ivanenkov, Y.; Aladinskiy, V.; Sanchez-Lengeling, B.; Aspuru-Guzik, A.; Zhavoronkov, A. Reinforced Adversarial Neural Computer for de Novo Molecular Design. J. Chem. Inf. Model. 2018, 58, 1194-1204. [CrossRef]
- Putin, E.; Asadulaev, A.; Vanhaelen, Q.; Ivanenkov, Y.; Aladinskaya, A. V.; Aliper, A.; Zhavoronkov, A. Adversarial Threshold Neural Computer for Molecular de Novo Design. Mol. Pharm. 2018, 15, 4386-4397. [CrossRef]
- Gupta, A.; Muller, A. T.; Huisman, B. J. H.; Fuchs, J. A.; Schneider, P.; Schneider, G. Generative Recurrent Networks for De Novo Drug Design. Mol. Inform. 2018, 37 (1-2). [CrossRef]
- Kadurin, A.; Aliper, A.; Kazennov, A.; Mamoshina, P.; Vanhaelen, Q.; Khrabrov, K.; Zhavoronkov, A. The cornucopia of meaningful leads: Applying deep adversarial autoencoders for new molecule development in oncology. Oncotarget 2017, 8, 10883-10890. [CrossRef]
- Polykovskiy, D.; Zhebrak, A.; Vetrov, D.; Ivanenkov, Y.; Aladinskiy, V.; Mamoshina, P.; Bozdaganyan, M.; Aliper, A.; Zhavoronkov, A.; Kadurin, A. Entangled Conditional Adversarial Autoencoder for de Novo Drug Discovery. Mol. Pharm. 2018, 15 , 4398-4405. [CrossRef]
- Dr Cao, N.; Kipf, T. MolGAN: An implicit generative model for small molecular graphs. arXiv.org, 2018, 1805.11973. [CrossRef]
- Zhu, J-Y.; Park,T.; Isola, P.; Efros. A.A. Unpaired Image-to-Image Translation using Cycle-Consistent Adversarial Networks. arXiv.org, 2018, 1703.10593v6. [CrossRef]
- Maziarka, L.; Pocha, A.; Kaczmarczyk, J.; Rataj, K.; Warchol, M. Mol-CycleGAN - a generative mode, for molecular optimization. J. Cheminf. 2020, 12, 2. [CrossRef]
- Racz, A.; Bajusz, D.; Heberger, K. Multi-Level Comparison of Machine Learning Classifiers and Their Performance Metrics. Molecules 2019, 24, 2811. [CrossRef]
- Olson, R. S.; La Cava, W.; Orzechowski, P.; Urbanowicz, R. J.; Moore, J. H. PMLB: a large benchmark suite for machine learning evaluation and comparison. BioData Min 2017, 10, 36. [CrossRef]
- Polykovskiy, D.; Zhebrak, A.; Sanchez-Lengeling, B.; Golovanov, S.; Tatanov, O.; Belyaev, S.; Kurbanov, R.; Artamonov, A.; Aladinskiy, V.; Veselov, M.; Kadurin, A.; Nikolenko, S.; Aspuru-Guzik, A.; Zhavoronkov, A. Molecular Sets (MOSES): A Benchmarking Platform for Molecular Generation Models. Front. Pharmacol. 2020, 11, 565644. [CrossRef]
- Preuer, K.; Renz, P.; Unterthiner, T.; Hochreiter, S.; Klambauer, G. Fréchet ChemNet Distance: A Metric for Generative Models for Molecules in Drug Discovery. J. Chem. Inf. Model. 2018, 58, 1736-1741. [CrossRef]
- Brown, N.; Fiscato, M.; Segler, M.H.S.; Vaucher, A.C. GuacaMol: Benchmarking Models for de Novo Molecular Design. J. Chem. Inf. Model. 2019, 59, 1096-1108. [CrossRef]
- Bickerton, G.R.; Paolini, G.V.; Besnard, J.; Muresan, S.; Hopkins, A.L. Quantifying the chemical beauty of drugs. Nat. Chem. 2012, 4, 90-98. [CrossRef]
- Ertl, P.; Schuffenhauer, A. Estimation of synthetic accessibility score of drug-like molecules based on molecular complexity and fragment contributions. J. Cheminform. 2009, 1, 8. [CrossRef]
- Buchwald, P.; Bodor, N. Octanol-water partition: searching for predictive models. Curr. Med. Chem. 1998, 5, 353-380.
- Blaschke, T.; Arús-Pous, J.; Chen, H.; Margreitter, C.; Tyrchan, C.; Engkvist, O.; Papadopoulos, K.; Patronov, A. REINVENT 2.0: An AI Tool for De Novo Drug Design. J. Chem. Inf. Model. 2020, 60, 5918–5922. [CrossRef]
- Loeffler, H. H.; He, J.; Tibo, A.; Janet, J. P.; Voronov, A.; Mervin, L. H.; Engkvist, O. Reinvent 4: Modern AI–Driven Generative Molecule Design. J. Cheminform. 2024, 16, 20. [CrossRef]
- Zhavoronkov, A.; Ivanenkov, Y.A.; Aliper, A.; Veselov, M.S.; Aladinskiy, V.A.; Aladinskaya, A.V.; Terentiev, V.A.; Polykovskiy, D.A.; Kuznetsov, M.D.; Asadulaev, A.; Volkov, Y.; Zholus, A.; Shayakhmetov, R.R.; Zhebrak, A.; Minaeva, L.I.; Zagribelnyy, B.A.; Lee, L.H.; Soll, R.; Madge, D.; Xing, L.; Guo, T.; Aspuru-Guzik, A. Deep learning enables rapid identification of potent DDR1 kinase inhibitors. Nat. Biotechnol. 2019, 37, 1038-1040. [CrossRef]
- Dollar, O.; Joshi, N.; Beck, D.A.C.; Pfaendtner, J. Attention-based generative models for de novo molecular design. Chem. Sci. 2021, 12, 8362-8372. [CrossRef]
- Winter, R.; Montanari, F.; Noé, F.; Clevert, D.A. Learning continuous and data-driven molecular descriptors by translating equivalent chemical representations. Chem. Sci. 2018, 10, 1692-1701. [CrossRef]
- Winter, R.; Montanari, F.; Steffen, A.; Briem, H.; Noé, F.; Clevert, D.A. Efficient multi-objective molecular optimization in a continuous latent space. Chem. Sci. 2019, 10, 8016-8024. [CrossRef]
- Winter, R.; Retel, J.; Noé, F.; Clevert, D.A.; Steffen, A. grünifai: interactive multiparameter optimization of molecules in a continuous vector space. Bioinformatics 2020, 36, 4093-4094. [CrossRef]
- Hoffman, S.C.; Chenthamarakshan, V.; Wadhawan, K.; Cen, P-Y., Das, P. Optimizing molecules using efficient queries from property evaluations. Nat. Mach. Intell. 2022, 4, 21–31 . [CrossRef]
- Wang, M.; Sun, H.; Wang, J.; Pang, J.; Chai, X.; Xu, L.; Li, H.; Cao, D.; Hou, T. Comprehensive assessment of deep generative architectures for de novo drug design. Brief. Bioinform. 2022, 23, bbab544. [CrossRef]
- Wang, S.; Guo, Y.; Wang, Y.; Sun, H.; Huang, J. Smiles-Bert: large scale unsupervised pre-training for molecular property prediction. Proceedings of the 10th ACM International Conference on Bioinformatics, Computational Biology and Health Informatics, 2019, 429–436. [CrossRef]
- Irwin, R.; Dimitriadis, S.; He, J.; Bjerrum, E. J. Chemformer: A Pre-Trained Transformer for Computational Chemistry. Mach. Learn.: Sci. Technol. 2022, 3 (1), 015022. [CrossRef]
- Westerlund, A. M.; Manohar Koki, S.; Kancharla, S.; Tibo, A.; Saigiridharan, L.; Kabeshov, M.; Mercado, R.; Genheden, S. Do Chemformers Dream of Organic Matter? Evaluating a Transformer Model for Multistep Retrosynthesis. J. Chem. Inf. Model. 2024, 64, 3021–3033. [CrossRef]
- Zhou, Z.; Kearnes, S.; Li, L.; Zare, R. N.; Riley, P. Author Correction: Optimization of Molecules via Deep Reinforcement Learning. Sci Rep 2020, 10478. [CrossRef]
- Xu, Y.; Lin, K.; Wang, S.; Wang, L.; Cai, C.; Song, C.; Lai, L.; Pei, J. Deep learning for molecular generation. Future Med Chem. 2019, 11, 567-597. [CrossRef]
- Zhang, J.; Chen, H. De Novo Molecule Design Using Molecular Generative Models Constrained by Ligand-Protein Interactions. J. Chem. Inf. Model. 2022. [CrossRef]
- Li, Y.; Pei, J.; Lai, L. Structure-based de novo drug design using 3D deep generative models. Chem. Sci. 2021, 12, 13664-13675. [CrossRef]
- Xie, W.; Wang, F.; Li, Y.; Lai, L.; Pei, J. Advances and Challenges in De Novo Drug Design Using Three-Dimensional Deep Generative Models. J. Chem. Inf. Model. 2022, 62, 2269-2279. [CrossRef]
- Gilmer, J.; Schoenholz, S. S.; Riley, P. F. ; Vinyals, O.; Dahl, G. E. Neural Message Passing for Quantum Chemistry. arXiv 2017. [CrossRef]
- Kearnes, S.; McCloskey, K.; Berndl, M.; Pande, V.; Riley, P. Molecular Graph Convolutions: Moving beyond Fingerprints. J Comput Aided Mol. Des. 2016, 30, 595–608. [CrossRef]
- Veličković, P.; Cucurull, G.; Casanova, A.; Romero, A.; Liò, P.; Bengio, Y. Graph Attention Networks. arXiv 2017, 1710.10903 . [CrossRef]
- Yang, K.; Swanson, K.; Jin, W.; Coley, C.; Eiden, P.; Gao, H.; Guzman-Perez, A.; Hopper, T.; Kelley, B.; Mathea, M.; Palmer, A.; Settels, V.; Jaakkola, T.; Jensen, K.; Barzilay, R. Analyzing Learned Molecular Representations for Property Prediction. J. Chem. Inf. Model. 2019, 59, 3370–3388. [CrossRef]
- Jin, W.; Barzilay, R.; Jaakkola, T. Junction Tree Variational Autoencoder for Molecular Graph Generation. arXiv 2018. 1802.04364. [CrossRef]
- Shi, C.; Xu, M.; Zhu, Z.; Zhang, W.; Zhang, M.; Tang, J. GraphAF: A Flow-Based Autoregressive Model for Molecular Graph Generation. arXiv 2020. [CrossRef]
- Bengio, E.; Jain, M.; Korablyov, M.; Precup, D.; Bengio, Y. Flow network based generative models for non-iterative diverse candidate generation. In Advances in Neural Information Processing Systems 34: Proceedings of the Neural Information Processing Systems Conference (NeurIPS 2021), 2021 , 7924-7936.
- Jain, M.; Deleu, T.; Hartford, J.; Liu, C.-H.; Hernandez-Garcia, A.; Bengio, Y. GFlowNets for AI-Driven Scientific Discovery. Digital Discovery 2023, 2, 557–577. [CrossRef]
- Schütt, K. T.; Kindermans, P-J.; Sauceda, H.E.; Chmiela, S.; Tkatchenko, A.; Müller, K. R. SchNet: A continuous-filter convolutional neural network for modeling quantum interactions. NIPS'17: Proceedings of the 31st International Conference on Neural Information Processing Systems, 2017, 992 – 1002.
- Schütt, K. T.; Arbabzadah, F.; Chmiela, S.; Müller, K. R.; Tkatchenko, A. Quantum-Chemical Insights from Deep Tensor Neural Networks. Nat Commun 2017, 8, 13890. [CrossRef]
- Gasteiger, J.; Groß, J.; Günnemann, S. Directional Message Passing for Molecular Graphs. arXiv 2020. [CrossRef]
- Gasteiger, J.; Giri, S.; Margraf, J. T.; Günnemann, S. Fast and Uncertainty-Aware Directional Message Passing for Non-Equilibrium Molecules. arXiv 2020. [CrossRef]
- Stärk, H.; Ganea, O.-E.; Pattanaik, L.; Barzilay, R.; Jaakkola, T. EquiBind: Geometric Deep Learning for Drug Binding Structure Prediction. arXiv 2022. [CrossRef]
- Lu, W.; Wu, Q.; Zhang, J.; Rao, J.; Li, C.; Zheng, S. TANKBind: Trigonometry-Aware Neural NetworKs for Drug-Protein Binding Structure Prediction. bioRxiv 2022, 2022.06.06.495043;. [CrossRef]
- Liu, S.; Wang, H.; Liu, W.; Lasenby, J.; Guo, H.; Tang, J. Pre-Training Molecular Graph Representation with 3D Geometry. arXiv 2021. [CrossRef]
- Stärk, H.; Beaini, D.; Corso, G.; Tossou, P.; Dallago, C.; Günnemann, S.; Liò, P. 3D Infomax Improves GNNs for Molecular Property Prediction. arXiv 2021. [CrossRef]
- Wang, X.; Zhao, H.; Tu, W.; Yao, Q. Automated 3D Pre-Training for Molecular Property Prediction. arXiv 2023. [CrossRef]
- Xu, M.; Yu, L.; Song, Y.; Shi, C.; Ermon, S.; Tang, J. GeoDiff: A Geometric Diffusion Model for Molecular Conformation Generation. arXiv 2022. [CrossRef]
- Cai, H.; Zhang, Z.; Wang, M.; Zhong, B.; Li, Q.; Zhong, Y.; Wu, Y.; Ying, T.; Tang, J. Pretrainable Geometric Graph Neural Network for Antibody Affinity Maturation. Nat Commun 2024, 15, 7785. [CrossRef]
- Jing, B.; Corso, G.; Chang, J.; Barzilay, R.; Jaakkola, T. Torsional Diffusion for Molecular Conformer Generation. arXiv 2022. [CrossRef]
- Corso, G.; Stärk, H.; Jing, B.; Barzilay, R.; Jaakkola, T. DiffDock: Diffusion Steps, Twists, and Turns for Molecular Docking. arXiv 2022. [CrossRef]
- Corso, G.; Deng, A.; Fry, B.; Polizzi, N.; Barzilay, R.; Jaakkola, T. Deep Confident Steps to New Pockets: Strategies for Docking Generalization. arXiv 2024. [CrossRef]
- Zheng, J.; Yi, H.-C.; You, Z.-H. Equivariant 3D-Conditional Diffusion Model for De Novo Drug Design. IEEE J. Biomed. Health Inform. 2025, 29, 1805–1816. [CrossRef]
- Watson, J. L.; Juergens, D.; Bennett, N. R.; Trippe, B. L.; Yim, J.; Eisenach, H. E.; Ahern, W.; Borst, A. J.; Ragotte, R. J.; Milles, L. F.; Wicky, B. I. M.; Hanikel, N.; Pellock, S. J.; Courbet, A.; Sheffler, W.; Wang, J.; Venkatesh, P.; Sappington, I.; Torres, S. V.; Lauko, A.; De Bortoli, V.; Mathieu, E.; Ovchinnikov, S.; Barzilay, R.; Jaakkola, T. S.; DiMaio, F.; Baek, M.; Baker, D. De Novo Design of Protein Structure and Function with RFdiffusion. Nature 2023, 620, 1089–1100. [CrossRef]
- Dauparas, J.; Anishchenko, I.; Bennett, N.; Bai, H.; Ragotte, R. J.; Milles, L. F.; Wicky, B. I. M.; Courbet, A.; de Haas, R. J.; Bethel, N.; Leung, P. J. Y.; Huddy, T. F.; Pellock, S.; Tischer, D.; Chan, F.; Koepnick, B.; Nguyen, H.; Kang, A.; Sankaran, B.; Bera, A. K.; King, N. P.; Baker, D. Robust Deep Learning–Based Protein Sequence Design Using ProteinMPNN. Science 2022, 378, 49–56. [CrossRef]
- Hayes, T.; Rao, R.; Akin, H.; Sofroniew, N. J.; Oktay, D.; Lin, Z.; Verkuil, R.; Tran, V. Q.; Deaton, J.; Wiggert, M.; Badkundri, R.; Shafkat, I.; Gong, J.; Derry, A.; Molina, R. S.; Thomas, N.; Khan, Y. A.; Mishra, C.; Kim, C.; Bartie, L. J.; Nemeth, M.; Hsu, P. D.; Sercu, T.; Candido, S.; Rives, A. Simulating 500 Million Years of Evolution with a Language Model. Science 2025, 387, 850–858. [CrossRef]
- Ingraham, J. B.; Baranov, M.; Costello, Z.; Barber, K. W.; Wang, W.; Ismail, A.; Frappier, V.; Lord, D. M.; Ng-Thow-Hing, C.; Van Vlack, E. R.; Tie, S.; Xue, V.; Cowles, S. C.; Leung, A.; Rodrigues, J. V.; Morales-Perez, C. L.; Ayoub, A. M.; Green, R.; Puentes, K.; Oplinger, F.; Panwar, N. V.; Obermeyer, F.; Root, A. R.; Beam, A. L.; Poelwijk, F. J.; Grigoryan, G. Illuminating Protein Space with a Programmable Generative Model. Nature 2023, 623, 1070–1078. [CrossRef]
- Fey, M.; Lenssen, J. E. Fast Graph Representation Learning with PyTorch Geometric. arXiv 2019. [CrossRef]
- Wang, M.; Zheng, D.; Ye, Z.; Gan, Q.; Li, M.; Song, X.; Zhou, J.; Ma, C.; Yu, L.; Gai, Y.; Xiao, T.; He, T.; Karypis, G.; Li, J.; Zhang, Z. Deep Graph Library: A Graph-Centric, Highly-Performant Package for Graph Neural Networks. arXiv 2019. [CrossRef]
- Huang, K.; Fu, T.; Gao, W.; Zhao, Y.; Roohani, Y.; Leskovec, J.; Coley, C. W.; Xiao, C.; Sun, J.; Zitnik, M. Therapeutics Data Commons: Machine Learning Datasets and Tasks for Drug Discovery and Development. arXiv 2021. [CrossRef]
- Krishnan, K.; Kassab, R.; Agajanian, S.; Verkhivker, G. Interpretable Machine Learning Models for Molecular Design of Tyrosine Kinase Inhibitors Using Variational Autoencoders and Perturbation-Based Approach of Chemical Space Exploration. Int. J. Mol. Sci. 2022, 23, 11262. [CrossRef]
- Davies, M.; Nowotka, M.; Papadatos, G.; Dedman, N.; Gaulton, A.; Atkinson, F.; Bellis, L.; Overington, J. P. ChEMBL web services: streamlining access to drug discovery data and utilities. Nucleic Acids Res 2015, 43 (W1), W612-W620. [CrossRef]
- Wishart, D. S.; Feunang, Y. D.; Guo, A. C.; Lo, E. J.; Marcu, A.; Grant, J. R.; Sajed, T.; Johnson, D.; Li, C.; Sayeeda, Z.; Assempour, N.; Iynkkaran, I.; Liu, Y.; Maciejewski, A.; Gale, N.; Wilson, A.; Chin, L.; Cummings, R.; Le, D.; Pon, A.; Knox, C.; Wilson, M. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res 2018, 46 (D1), D1074-D1082. [CrossRef]
- Knox, C.; Wilson, M.; Klinger, C. M.; Franklin, M.; Oler, E.; Wilson, A.; Pon, A.; Cox, J.; Chin, N. E. (Lucy); Strawbridge, S. A.; Garcia-Patino, M.; Kruger, R.; Sivakumaran, A.; Sanford, S.; Doshi, R.; Khetarpal, N.; Fatokun, O.; Doucet, D.; Zubkowski, A.; Rayat, D. Y.; Jackson, H.; Harford, K.; Anjum, A.; Zakir, M.; Wang, F.; Tian, S.; Lee, B.; Liigand, J.; Peters, H.; Wang, R. Q. (Rachel); Nguyen, T.; So, D.; Sharp, M.; da Silva, R.; Gabriel, C.; Scantlebury, J.; Jasinski, M.; Ackerman, D.; Jewison, T.; Sajed, T.; Gautam, V.; Wishart, D. S. DrugBank 6.0: The DrugBank Knowledgebase for 2024. Nucleic Acids Res. 2024 , 52(D1), D1265-D1275. [CrossRef]
- Liu, T.; Hwang, L.; Burley, S. K.; Nitsche, C. I.; Southan, C.; Walters, W. P.; Gilson, M. K. BindingDB in 2024: A FAIR Knowledgebase of Protein-Small Molecule Binding Data. Nucleic Acids Res. 2025 , 53(D1), D1633-D1644. [CrossRef]
- Ahmed, A.; Smith, R. D.; Clark, J. J.; Dunbar, J. B., Jr.; Carlson, H. A. Recent improvements to Binding MOAD: a resource for protein-ligand binding affinities and structures. Nucleic Acids Res 2015, 43, D465-D469. [CrossRef]
- Hastings, J.; de Matos, P.; Dekker, A.; Ennis, M.; Harsha, B.; Kale, N.; Muthukrishnan, V.; Owen, G.; Turner, S.; Williams, M.; Steinbeck, C. The ChEBI reference database and ontology for biologically relevant chemistry: enhancements for 2013. Nucleic Acids Res 2013, 41, D456-D463. [CrossRef]
- Irwin, J. J.; Sterling, T.; Mysinger, M. M.; Bolstad, E. S.; Coleman, R. G. ZINC: a free tool to discover chemistry for biology. J. Chem. Inf. Model. 2012, 52, 1757-1768. [CrossRef]
- Sterling, T.; Irwin, J. J. ZINC 15--Ligand Discovery for Everyone. J. Chem. Inf. Model. 2015, 55, 2324-2337. [CrossRef]
- Irwin, J. J.; Tang, K. G.; Young, J.; Dandarchuluun, C.; Wong, B. R.; Khurelbaatar, M.; Moroz, Y. S.; Mayfield, J.; Sayle, R. A. ZINC20—A Free Ultralarge-Scale Chemical Database for Ligand Discovery. J. Chem. Inf. Model. 2020, 60, 6065–6073. [CrossRef]
- Ruddigkeit, L.; van Deursen, R.; Blum, L. C.; Reymond, J. L. Enumeration of 166 billion organic small molecules in the chemical universe database GDB-17. J. Chem. Inf. Model. 2012, 52, 2864-2875. [CrossRef]
- Ruddigkeit, L.; Blum, L. C.; Reymond, J. L. Visualization, and virtual screening of the chemical universe database GDB-17. J. Chem. Inf. Model. 2013, 53, 56-65. [CrossRef]
- Visini, R.; Awale, M.; Reymond, J. L. Fragment Database FDB-17. J. Chem. Inf. Model. 2017, 57, 700-709.
- Xerxa, E.; Bajorath, J. Data Sets of Human and Mouse Protein Kinase Inhibitors With Curated Activity Data Including Covalent Inhibitors. Future Sci OA. 2023 , 9, FSO892. [CrossRef]
- Koch, S.; Kullmann, E.; Bajorath, J. Opportunities for Protein Kinase Drug Discovery – 2025 Update on the Chemically Underexplored Human Kinome. Eur J Med Chem Rep 2025, 15, 100305. [CrossRef]
- Hu, H.; Laufkötter, O.; Miljković, F.; Bajorath, J. Data set of competitive and allosteric protein kinase inhibitors confirmed by X-ray crystallography. Data Brief. 2021, 35, 106816. [CrossRef]
- Laufkötter, O.; Hu, H.; Miljković, F.; Bajorath, J. Structure- and Similarity-Based Survey of Allosteric Kinase Inhibitors, Activators, and Closely Related Compounds. J. Med. Chem. 2022, 65, 922-934. [CrossRef]
- Hu, H.; Laufkötter, O.; Miljković, F.; Bajorath, J. Systematic comparison of competitive and allosteric kinase inhibitors reveals common structural characteristics. Eur. J. Med. Chem. 2021, 214, 113206. [CrossRef]
- Kanev, G. K.; de Graaf, C.; Westerman, B. A.; de Esch, I. J. P.; Kooistra, A. J. KLIFS: An Overhaul after the First 5 Years of Supporting Kinase Research. Nucleic Acids Res. 2021 , 49(D1), D562-D569. [CrossRef]
- Xerxa, E.; Laufkötter, O.; Bajorath, J. Systematic Analysis of Covalent and Allosteric Protein Kinase Inhibitors. Molecules 2023, 28, 5805. [CrossRef]
- Bento, A.P.; Hersey, A.; Félix, E.; Landrum, G.; Gaulton, A.; Atkinson, F.; Bellis, L.J.; De Veij, M.; Leach, A.R. An open-source chemical structure curation pipeline using RDKit. J. Cheminform. 2020, 12, 51. [CrossRef]
- Kruger, F.; Stiefl, N.; Landrum, G.A. rdScaffoldNetwork: The Scaffold Network Implementation in RDKit. J. Chem. Inf. Model. 2020, 60, 3331-3335. [CrossRef]
- Abadi, M.; Agarwal, A.; Barham, P.; Brevdo, E.; Chen, Z.; Citro, C.; Corrado, G. S.; Davis, A.; Dean, J.; Devin, M.; Ghemawat, S.; Goodfellow, I.; Harp, A.; Irving, G.; Isard, M.; Jia, Y.; Jozefowicz, R.; Kaiser, L.; Kudlur, M.; Levenberg, J.; Mane, D.; Monga, R.; Moore, S.; Murray, D.; Olah, C.; Schuster, M.; Shlens, J.; Steiner, B.; Sutskever, I.; Talwar, K.; Tucker, P.; Vanhoucke, V.; Vasudevan, V.; Viegas, F.; Vinyals, O.; Warden, P.; Wattenberg, M.; Wicke, M.; Yu, Y.; Zheng, X. TensorFlow: A System for Large-Scale Machine Learning. In 12th USENIX Symposium on Operating Systems Design and Implementation (OSDI 16) 2016, 16, 265-283. [CrossRef]
- Boulesteix, A.; Janitza, S.; Kruppa, J.; König, I. Overview of random forest methodology and practical guidance with emphasis on computational biology and bioinformatics. Wiley Interdisciplinary Reviews: Data Mining and Knowledge Discovery, 2012, 2, 493-507. [CrossRef]
- Godden, J.W.; Xue, L.; Bajorath, J. Combinatorial preferences affect molecular similarity/diversity calculations using binary fingerprints and Tanimoto coefficients. J. Chem. Inf. Comput. Sci. 2000, 40, 163-166. [CrossRef]
| Family | Min Range | Max Range | Min Average | Max Average |
Min Stand Dev |
Max Stand Dev |
|---|---|---|---|---|---|---|
| ABL1 | -5.89215 | 5.97272 | -1.34594 | 1.2609 | 0.78482 | 1.46389 |
| SRC | -5.89215 | 6.20087 | -1.38016 | 1.30248 | 0.86567 | 1.63218 |
| CSF1R | -5.19233 | 6.84467 | -1.19730 | 1.21217 | 0.65711 | 1.46416 |
| EGFR | -6.18875 | 6.55361 | -1.25954 | 1.22010 | 0.82409 | 1.39603 |
| FLT3 | -5.00162 | 6.45221 | -1.17921 | 1.15374 | 0.69147 | 1.42987 |
| KDR | -6.15671 | 7.05822 | -1.37088 | 1.32073 | 0.80067 | 1.35351 |
| LCK | -6.15671 | 6.62534 | -1.38279 | 1.39623 | 0.81684 | 1.55863 |
| MAPK10 | -5.08671 | 5.98541 | -1.16237 | 1.14753 | 0.68575 | 1.29511 |
| MAPK14 | -6.15671 | 6.89392 | -1.52617 | 1.44791 | 0.73652 | 1.29781 |
| MET | -6.13674 | 6.49813 | -1.45546 | 1.52347 | 0.79279 | 1.53428 |
| Precision | Recall | F1-Score | Support | |
| 0 | 0.99 | 0.98 | 0.98 | 23530 |
| 1 | 0.71 | 0.86 | 0.78 | 1502 |
| Macro Avg | 0.85 | 0.92 | 0.88 | 25032 |
| Weighted Avg | 0.97 | 0.97 | 0.97 | 25032 |
| Precision | Recall | F1-Score | Support | |
| ABL1 | 0.51 | 0.58 | 0.55 | 409 |
| SRC | 0.57 | 0.56 | 0.56 | 660 |
| CSF1R | 0.69 | 0.54 | 0.61 | 142 |
| EGFR | 0.69 | 0.74 | 0.71 | 795 |
| FLT3 | 0.55 | 0.46 | 0.50 | 194 |
| KDR | 0.58 | 0.59 | 0.58 | 916 |
| LCK | 0.47 | 0.41 | 0.44 | 313 |
| MAPK10 | 0.77 | 0.55 | 0.64 | 163 |
| MAPK14 | 0.75 | 0.80 | 0.78 | 722 |
| MET | 0.74 | 0.72 | 0.73 | 421 |
| Macro Avg | 0.63 | 0.59 | 0.61 | 4735 |
| Weighted Avg | 0.63 | 0.63 | 0.63 | 4735 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




