Submitted:
25 February 2026
Posted:
26 February 2026
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
1.1. L-DMD – A Rational Hypothesis
1.2. The Special Properties of Ionizable Lipids
1.3. Powerful Signaling Effects of Phosphoinositides (PIPs)
2. Lipid Nanoparticles for mRNA Delivery: Biological Properties and Effects on Cellular Systems
2.1. Factors Influencing Nanoparticle Bioactivity
2.2. The Biocorona and Biodistribution of the LNPs
2.3. Endosomal Escape and Membrane Destabilization Due to Ionizable Lipids
2.4. Spread to Distant Sites via Exosomes
2.5. LNP Metabolism and Oxidative Stress Mechanism
2.6. Activation of the Immune System
2.7. Delivery Architecture as a Determinant of Membrane Stress and Possible Systemic Risk
3. Omics Data indicating Membrane Dysfunction Secondary to LNP Transfection
3.1. Mouse Data
3.2. Human Data
3.3. Convergent Findings Across Studies and Platforms
- (1)
- Multiple lines of evidence indicate modulation of phosphoinositide-related pathways. These include downregulation of PIK3C2G (class II PI3K) across independent mouse datasets, phosphoinositide-dependent signaling alterations observed in human serum proteomics, and dysregulation of ESCRT-associated pathways involved in membrane repair and endocytic trafficking.
- (2)
- Inflammatory and stress-associated signaling pathways are reproducibly engaged, including NF-κB activation, upregulation of TNF-α, IL-6, and IL-1β, chemokine induction, and evidence of NLRP3 inflammasome involvement. TLR4/MyD88-dependent signaling is consistently implicated, with indications of pathway bias, depending on cellular and experimental context.
- (3)
- Metabolic and detoxification pathways are affected in a context-dependent manner. These include downregulation of cytochrome P450-associated xenobiotic metabolism, suppression of PPAR and AMPK signaling, and attenuation of TCA cycle activity, consistent with altered lipid and energy homeostasis.
- (4)
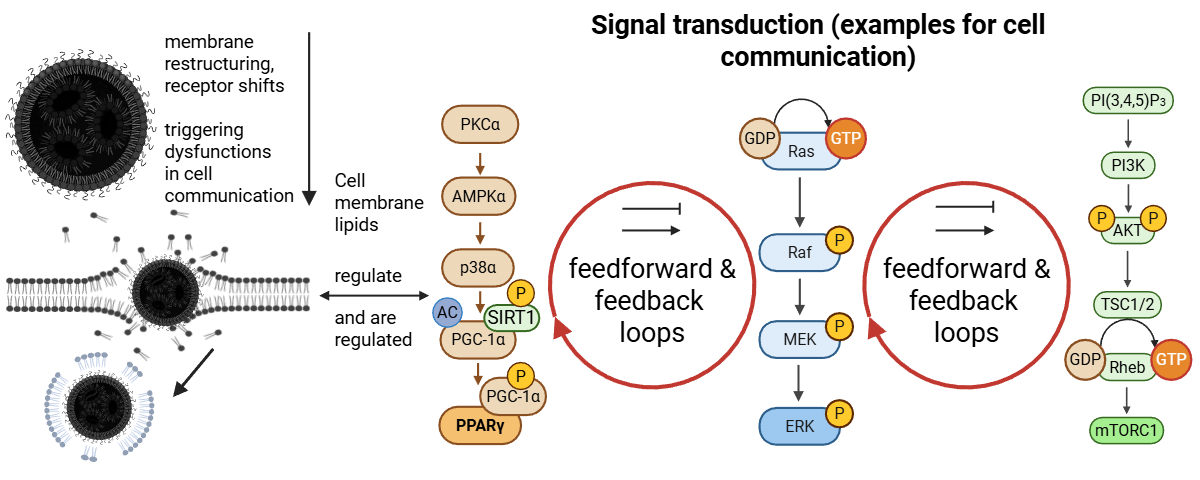
- Multiple signaling cascades downstream of membrane-proximal events are activated, including RAS/MAPK, PI3K/AKT/mTOR, and JAK-STAT pathways, as observed in both mouse and human datasets.
4. Proposed Mechanistic Hypothesis Derived from the Omics Data: L-DMD as a Central Node
4.1. Disruption of the ESCRT Circuit and Phosphatidylinositol Signaling (Hickey et al.)
4.2. Downregulated Xenobiotic Metabolism by Cytochrome P450 Enzymes (Ndeupen et al., Hickey et al.)
4.3. Are the TLR4 Reactions Biased? What Mouse Data Reveal (Ndeupen et al., Korzun et al.)
4.4. Upregulation of Multiple Inflammatory Markers (Ndeupen et al., Korzun et al., Knabl et al.)
4.5. Complement Activation (Korzun et al. & Luo et al.)
4.6. Downregulation of PPAR and AMPK Signaling (Ndeupen et al.)
4.7. RAS (Rat Sarkoma) Signaling and the MAPK (Hickey et al., Knabl. et al., Korzun et al.)
4.8. Transcription Factors (E2F1, E2F8) (Knabl et al.) and Mechanistic Target of Rapamycin Complex (mTORC) (Knabl et al., Hickey et al.)
4.9. Conceptual Consolidation of Section 3 and Section 4
5. Breaching the Plasma Membrane: Important Roles for Phosphoinositides
5.1. Brief Overview of the Phosphatidylinositol(PI)-Cycle
5.2. Signaling Through Phosphorylation States of Phosphatidylinositol
5.3. The LNP Components and Their Effects
5.3.1. Ionizable Lipids
5.3.2. DSPC (1,2-distearoyl-sn-glycero-3-phosphocholine)
5.3.3. Oxysterol-Binding Proteins
5.3.4. A Role for Lipid Impurities
5.4. Small Perturbations Can Lead to Major Shifts in PIP Signaling
6. Discussion
6.1. LNP Perturbation and Functional modRNA Persistence
6.2. Experimental Validation
6.3. Inter-Individual Variability
6.4. Pharmacovigilance and Monitoring Gaps
6.5. Implications for Nanomedicine Design
- Rational LNP design
- 2.
- Spatiotemporal control of signaling
- 3.
- Predictive biomarker development
6.6. Broader Relevance and Methodological Outlook
6.7. Limitations
7. Conclusions
Funding
Authors’ Contributions
Use of Generative AI
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AAV | adeno-associated viruses |
| AKT | Protein Kinase B |
| AMPK | AMP-Activated Protein Kinase |
| BUB1 | Budding Uninhibited by Benzimidazole 1 |
| CAR | Constitutive Androstane Receptor |
| CARPA | Complement Activation-Related Pseudoallergy |
| CCL(X) | C-C Motif Chemokine Ligand (x) |
| CCNA1 | Cyclin A1 |
| CD1d | Cluster of Differentiation 1d |
| CDC25A | Cell Division Cycle 25A |
| CDP-DAG | CDP-Diacylglycerol |
| CHAC1 | ChaC glutathione specific gamma-glutamylcyclotransferase 1 |
| COVID-19 | Coronavirus Disease 2019 |
| CRP | C-Reactive Protein |
| CSF2 | Colony Stimulating Factor 2 |
| CSF3 | Colony Stimulating Factor 3 |
| CXCL | C-X-C Motif Chemokine Ligand (X) |
| CYP | Cytochrome P450 |
| DEG | Differentially Expressed Gene |
| DEP | Differentially Expressed Protein |
| DSPC | Distearoylphosphatidylcholine |
| E2F | E2F Transcription Factor |
| E2F1 | E2F Transcription Factor 1 |
| E2F8 | E2F Transcription Factor 8 |
| Eif | Eukaryotic Initiation Factor |
| ELISA | Enzyme-Linked Immunosorbent Assay |
| eLNP | Empty Lipid Nanoparticle |
| EMA | European Medicines Agency |
| ENaC | Epithelial Sodium Channel |
| ER | Endoplasmic Reticulum |
| ERK | Extracellular signal-Regulated Kinase |
| ESCRT | Endosomal Sorting Complex Required for Transport |
| FDA | Food and Drug Administration |
| FDR | False Discovery Rate |
| GalNAc | N-Acetylgalactosamine |
| GAP43 | Growth-Associated Protein 43 |
| GO | Gene Ontology |
| GPCR | G-Protein Coupled Receptor |
| GSEA | Gene Set Enrichment Analysis |
| IL-1β | Interleukin-1 beta |
| IL-6 | Interleukin-6 |
| INS | Insulin |
| JAK | Janus Kinase |
| JAK-STAT | Janus Kinase-Signal Transducer and Activator of Transcription |
| JNK | Jun N-terminal Kinase |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| KRAS | Kirsten Rat Sarcoma |
| L-DMD | Lipid-Driven Membrane Dysfunction |
| LDL | Low-Density Lipoprotein |
| LDLR | Low-Density Lipoprotein Receptor |
| LCN2 | Lipocalin 2 |
| LNP | Lipid Nanoparticle |
| LNPs | Lipid Nanoparticles |
| MAPK | Mitogen-Activated Protein Kinase |
| MARCKS | Myristoylated Alanine-Rich C Kinase Substrate |
| MEK | MAPK/ERK Kinase |
| modRNA | Modified (messenger)RNA |
| mRNA | Messenger RNA |
| mTOR | Mechanistic Target of Rapamycin |
| mTORC1 | mTOR Complex 1 |
| mTORC2 | mTOR Complex 2 |
| MVB | Multivesicular Body |
| NF-κB | Nuclear Factor kappa B |
| NLRP3 | NOD-Like Receptor Protein 3 |
| OSBP | Oxysterol-Binding Protein |
| P38 | p38-mitogenaktivierte Proteinkinasen |
| PA | Phosphatidic Acid |
| PBS | Phosphate Buffered Saline |
| PEG | Polyethylene Glycol |
| PEI | Polyethylenimine |
| PI | Phosphatidylinositol / Phosphoinositide |
| PI(3,4)P₂ | Phosphatidylinositol-3,4-bisphosphate |
| PI(3,4,5)P₃ | Phosphatidylinositol-3,4,5-trisphosphate |
| PI(3,5)P₂ | Phosphatidylinositol-3,5-bisphosphate |
| PI(4,5)P₂ | Phosphatidylinositol-4,5-bisphosphate |
| PI3K | Phosphoinositide 3-Kinase |
| PI4P | Phosphatidylinositol 4-phosphate |
| PIK3C2G | Phosphoinositide 3-Kinase Class 2 Gamma |
| PIP | Phosphoinositide |
| PIP₂ | Phosphoinositide bisphosphate |
| PIP₃ | Phosphoinositide trisphosphate |
| pKa | Negative base-10 logarithm of the acid dissociation constant |
| PKB | Serine/threonine-specific protein kinases |
| PLC | Phospholipase C |
| PPAR | Peroxisome Proliferator-Activated Receptor |
| PPARγ | Peroxisome Proliferator-Activated Receptor gamma |
| PRR | Pattern Recognition Receptor |
| PXR | Pregnane X Receptor |
| qPCR | Quantitative Polymerase Chain Reaction |
| RAF | Rapidly Accelerated Fibrosarcoma (kinase) |
| RAS | Rat Sarcoma (GTPase family) |
| RNA-Seq | RNA Sequencing |
| Rps | Ribosomal Protein |
| RRM2 | Ribonucleotide Reductase M2 |
| saRNA | Self-Amplifying RNA |
| SARS-CoV-2 | Severe Acute Respiratory Syndrome Coronavirus 2 |
| siRNA | Small Interfering RNA |
| STAT | Signal Transducer and Activator of Transcription |
| TAK-242 | Resatorvid, a small molecule inhibitor of toll-like receptor 4 signaling |
| TFH | T follicular helper |
| TGA | Therapeutic Goods Administration |
| TLR | Toll-Like Receptor |
| TLR4 | Toll-Like Receptor 4 |
| TLR7 | Toll-Like Receptor 7 |
| TLR8 | Toll-Like Receptor 8 |
| TNF-α | Tumor Necrosis Factor alpha |
| WHO | World Health Organization |
References
- Khurana, A; Allawadhi, P; Khurana, I; Allwadhi, S; Weiskirchen, R; Banothu, AK; et al. Role of nanotechnology behind the success of mRNA vaccines for COVID-19. Nano Today 2021, 38, 101142. [Google Scholar] [CrossRef]
- Hald Albertsen, C; Kulkarni, JA; Witzigmann, D; Lind, M; Petersson, K; Simonsen, JB. The role of lipid components in lipid nanoparticles for vaccines and gene therapy. Adv Drug Deliv Rev 2022, 188, 114416. [Google Scholar] [CrossRef]
- Swetha, K; Kotla, NG; Tunki, L; Jayaraj, A; Bhargava, SK; Hu, H; et al. Recent advances in the lipid nanoparticle-mediated delivery of mRNA vaccines. Vaccines (Basel) 2023, 11(3), 658. [Google Scholar] [CrossRef] [PubMed]
- Suk, JS; Xu, Q; Kim, N; Hanes, J; Ensign, LM. PEGylation as a strategy for improving nanoparticle-based drug and gene delivery. Adv Drug Deliv Rev 2016, 99 Pt A, 28–51. [Google Scholar] [CrossRef] [PubMed]
- Kloczewiak, M; Banks, JM; Jin, L; Brader, ML. Perspective on higher-order structure and thermal stability of mRNA vaccines. Mol Pharmaceutics 2022, 19(7), 2022–2031. [Google Scholar] [CrossRef]
- Trollmann, MFW; Böckmann, RA. Decoding pH-driven phase transition of lipid nanoparticles. Small Epub ahead of print. 2026, e11381. [Google Scholar] [CrossRef] [PubMed]
- Haghighi, E; Abolmaali, SS; Dehshahri, A; Mousavi Shaegh, SA; Azarpira, N; Tamaddon, AM. Navigating the intricate in-vivo journey of lipid nanoparticles tailored for the targeted delivery of RNA therapeutics: a quality-by-design approach. J Nanobiotechnology 2024, 22(1), 710. [Google Scholar] [CrossRef]
- Brader, ML; Williams, SJ; Banks, JM; Hui, WH; Zhou, ZH; Jin, L. Encapsulation state of messenger RNA inside lipid nanoparticles. Biophys J 2021, 120(14), 2766–2770. [Google Scholar] [CrossRef]
- Eygeris, Y; Gupta, M; Kim, J; Sahay, G. Chemistry of lipid nanoparticles for RNA delivery. Acc Chem Res 2022, 55(1), 2–12. [Google Scholar] [CrossRef]
- Kulkarni, JA; Cullis, PR; van der Meel, R. Lipid nanoparticles enabling gene therapies: From concepts to clinical utility. Nucleic Acid Ther 2018, 28(3), 146–157. [Google Scholar] [CrossRef]
- Simonsen, JB. A perspective on bleb and empty LNP structures. J Control Release 2024, 373, 952–961. [Google Scholar] [CrossRef]
- Trollmann, MFW; Böckmann, RA. mRNA lipid nanoparticle phase transition. Biophys J 2022, 121(20), 3927–3939. [Google Scholar] [CrossRef]
- Hou, X; Zaks, T; Langer, R; Dong, y. Lipid nanoparticles for mRNA delivery. Nat Rev Mater 2021, 6, 1078–1094. [Google Scholar] [CrossRef] [PubMed]
- Cárdenas, M; Campbell, RA; Arteta, MY; Lawrence, MJ; Sebastiani, F. Review of structural design guiding the development of lipid nanoparticles for nucleic acid delivery. Current Opinion in Colloid & Interface Science 2023, 66, 101705. [Google Scholar] [CrossRef]
- Nele, V; Campani, V; Alia Moosavian, S; De Rosa, G. Lipid nanoparticles for RNA delivery: Self-assembling vs driven-assembling strategies. Adv Drug Deliv Rev 2024, 208, 115291. [Google Scholar] [CrossRef] [PubMed]
- Manning, AM; Tilstra, G; Khan, AB; Couture-Senécal, J; Lau, YMA; Pang, J; et al. Ionizable lipid with supramolecular chemistry features for RNA delivery in vivo. Small 2023, 19(41), e2302917. [Google Scholar] [CrossRef]
- Čechová, P; Paloncýová, M; Šrejber, M; Otyepka, M. Mechanistic insights into interactions between ionizable lipid nanodroplets and biomembranes. J Biomol Struct Dyn 2025, 43(17), 9983–9993. [Google Scholar] [CrossRef]
- Ermilova, I; Swenson, J. Ionizable lipids penetrate phospholipid bilayers with high phase transition temperatures: perspectives from free energy calculations. Chem Phys Lipids 2023, 253, 105294. [Google Scholar] [CrossRef]
- Atmuri, NDP; Saadati, F; Kulkarni, J; Witzigmann, D; Cullis, PR; Ciufolini, MA. Design of cationic ionizable lipids for the delivery of therapeutic nucleic acids. Mol Ther Methods Clin Dev 2025, 33(4), 101585. [Google Scholar] [CrossRef]
- Gurtovenko, AA; Lyulina, AS. Electroporation of asymmetric phospholipid membranes. J Phys Chem B 2014, 118(33), 9909–18. [Google Scholar] [CrossRef] [PubMed]
- McLaughlin, S; Murray, D. Plasma membrane phosphoinositide organization by protein electrostatics. Nature 2005, 438(7068), 605–11. [Google Scholar] [CrossRef] [PubMed]
- Fung, SYS; Xǔ, XJ; Wu, M. Nonlinear dynamics in phosphoinositide metabolism. Curr Opin Cell Biol 2024, 88, 102373. [Google Scholar] [CrossRef]
- Ellenbroek, WG; Wang, YH; Christian, DA; Discher, DE; Janmey, PA; Liu, AJ. Divalent cation-dependent formation of electrostatic PIP2 clusters in lipid monolayers. Biophys J 2011, 101(9), 2178–84. [Google Scholar] [CrossRef]
- Mandala, VS; MacKinnon, R. The membrane electric field regulates the PIP2-binding site to gate the KCNQ1 channel. Proc Natl Acad Sci U S A 2023, 120(21), e2301985120. [Google Scholar] [CrossRef]
- Yue, Q; Al-Khalili, O; Moseley, A; Yoshigi, M; Wynne, BM; Ma, H; Eaton, DC. PIP2 interacts electrostatically with MARCKS-like Protein-1 and ENaC in renal epithelial cells. Biology (Basel) 2022, 11(12), 1694. [Google Scholar] [CrossRef]
- Yamaga, M; Martin, TFJ. PI(4,5)P2 is a master regulator for Ca2+-triggered vesicle exocytosis. Biochim Biophys Acta Mol Cell Biol Lipids 2025, 1870(6), 159651. [Google Scholar] [CrossRef]
- Gada, KD; Logothetis, DE. PKC regulation of ion channels: The involvement of PIP2. J Biol Chem 2022, 298(6), 102035. [Google Scholar] [CrossRef]
- Tariq, K; Luikart, BW. Striking a balance: PIP2 and PIP3 signaling in neuronal health and disease. Explor Neuroprotective Ther 2021, 1, 86–100. [Google Scholar] [CrossRef]
- Clarke, RJ. Electrostatic switch mechanisms of membrane protein trafficking and regulation. Biophys Rev 2023, 15(6), 1967–1985. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, T; Legat, A; Adam, E; Steuve, J; Gatot, JS; Vandenbranden, M; et al. DiC14-amidine cationic liposomes stimulate myeloid dendritic cells through Toll-like receptor 4. Eur J Immunol. 2008, 38(5), 1351–7. [Google Scholar] [CrossRef] [PubMed]
- Glaviano, A; Foo, ASC; Lam, HY; Yap, KCH; Jacot, W; Jones, RH; et al. PI3K/AKT/mTOR signaling transduction pathway and targeted therapies in cancer. Mol Cancer 2023, 22(1), 138. [Google Scholar] [CrossRef] [PubMed]
- Ivanova, A; Atakpa-Adaji, P; Rao, S; Marti-Solano, M; Taylor, CW. Dual regulation of IP3 receptors by IP3 and PIP2 controls the transition from local to global Ca2+ signals. Mol Cell 2024, 84(20), 3997–4015.e7. [Google Scholar] [CrossRef]
- Putney, JW; Tomita, T. Phospholipase C signaling and calcium influx. Adv Biol Reg 2012, 52(1), 152–164. [Google Scholar] [CrossRef]
- Cao, S; Chung, S; Kim, S; Li, Z; Manor, D; Buck, M. K-Ras G-domain binding with signaling lipid phosphatidylinositol (4,5)-phosphate (PIP2): membrane association, protein orientation, and function. J Biol Chem 2019, 294(17), 7068–7084. [Google Scholar] [CrossRef]
- Janmey, PA; Bucki, R; Radhakrishnan, R. Regulation of actin assembly by PI(4,5)P2 and other inositol phospholipids: An update on possible mechanisms. Biochem Biophys Res Commun 2018, 506(2), 307–314. [Google Scholar] [CrossRef]
- Abbasi, R; Shineh, G; Mobaraki, M; Doughty, S; Tayebi, L. Structural parameters of nanoparticles affecting their toxicity for biomedical applications: a review. J Nanopart Res 2023, 25(3), 43. [Google Scholar] [CrossRef]
- Yuan, Z; Yan, R; Fu, Z; Wu, T; Ren, C. Impact of physicochemical properties on biological effects of lipid nanoparticles: Are they completely safe. Sci Total Environ 2024, 927, 172240. [Google Scholar] [CrossRef] [PubMed]
- Szebeni, J; Kiss, B; Bozó, T; Turjeman, K; Levi-Kalisman, Y; Barenholz, Y; et al. Insights into the structure of Comirnaty COVID-19 vaccine: A theory on soft, partially bilayer-covered nanoparticles with hydrogen bond-stabilized mRNA-lipid complexes. ACS Nano 2023, 17(14), 13147–13157. [Google Scholar] [CrossRef]
- Münter, R; Larsen, JB; Andresen, TL. The vast majority of nucleic acid-loaded lipid nanoparticles contain cargo. J Colloid Interface Sci 2024, 674, 139–144. [Google Scholar] [CrossRef]
- Li, S; Hu, Y; Li, A; Lin, J; Hsieh, K; Schneiderman, Z; et al. Payload distribution and capacity of mRNA lipid nanoparticles. Nat Commun 2022, 13(1), 5561. [Google Scholar] [CrossRef] [PubMed]
- https. [CrossRef]
- Chen, X; Ye, Y; Li, M; Zuo, T; Xie, Z; Ke, Y; et al. Structural characterization of mRNA lipid nanoparticles (LNPs) in the presence of mRNA-free LNPs. J Control Release 2025, 386, 114082. [Google Scholar] [CrossRef]
- Müller, JA; Schäffler, N; Kellerer, T; Schwake, G; Ligon, TS; Rädler, JO. Kinetics of RNA-LNP delivery and protein expression. Eur J Pharm Biopharm 2024, 197, 114222. [Google Scholar] [CrossRef] [PubMed]
- Liao, S; Wang, S; Wadhwa, A; Birkenshaw, A; Fox, K; Cheng, MHY; et al. Transfection potency of lipid nanoparticles containing mRNA depends on relative loading levels. ACS Appl Mater Interfaces 2025, 17(2), 3097–3105. [Google Scholar] [CrossRef]
- Grumelot, S; Mohammed, N; Colonrosado, J; Sadeghi, SA; Fang, F; Hilsen, K. Lipid nanoparticle protein coronas form via lipoprotein fusion rather than shell-like adsorption. bioRxiv Preprint 2025. [CrossRef]
- Rampado, R; Crotti, S; Caliceti, P; Pucciarelli, S; Agostini, M. Recent advances in understanding the protein corona of nanoparticles and in the formulation of "stealthy" nanomaterials. Front Bioeng Biotechnol 2020, 8, 166. [Google Scholar] [CrossRef]
- Sun, Y; Zhou, Y; Rehman, M; Wang, YF; Guo, S. Protein corona of nanoparticles: Isolation and analysis. Chem Bio Eng. 2024, 1(9), 757–772. [Google Scholar] [CrossRef] [PubMed]
- Voke, E; Arral, ML; Squire, HJ; Lin, TJ; Zheng, L; Coreas, R; et al. Protein corona formed on lipid nanoparticles compromises delivery efficiency of mRNA cargo. Nat Commun 2025, 16(1), 8699. [Google Scholar] [CrossRef] [PubMed]
- Sebastiani, F; Yanez Arteta, M; Lerche, M; Porcar, L; Lang, C; Bragg, RA; Elmore, CS; et al. Apolipoprotein E binding drives structural and compositional rearrangement of mRNA-containing lipid nanoparticles. ACS Nano 2021, 15(4), 6709–6722. [Google Scholar] [CrossRef]
- Kim, W; Ly, NK; He, Y; Li, Y; Yuan, Z; Yeo, Y. Protein corona: Friend or foe? Co-opting serum proteins for nanoparticle delivery. Adv Drug Deliv Rev 2023, 192, 114635. [Google Scholar] [CrossRef]
- Paunovska, K; Da Silva Sanchez, AJ; Lokugamage, MP; Loughrey, D; Echeverri, ES; Cristian, A; et al. The extent to which lipid nanoparticles require apolipoprotein E and low-density lipoprotein receptor for delivery changes with ionizable lipid structure. Nano Lett 2022, 22(24), 10025–10033. [Google Scholar] [CrossRef]
- Naasani, I. Establishing the pharmacokinetics of genetic vaccines is essential for maximising their safety and efficacy. Clin Pharmacokinet 2022, 61(7), 921–927. [Google Scholar] [CrossRef]
- Hosseini-Kharat, M; Bremmell, KE; Prestidge, CA. Why do lipid nanoparticles target the liver? Understanding of biodistribution and liver-specific tropism. Mol Ther Methods Clin Dev 2025, 33(1), 101436. [Google Scholar] [CrossRef]
- European Medicines Agency. Assessment report: Comirnaty. EMA/707383/2020 Corr.2*1,2. February 19, 2021. 2020. [Google Scholar]
- Available online: https://www.ema.europa.eu/en/documents/assessment-report/comirnaty-epar-public-assessment-report_en.pdf.
- Therapeutic Goods Administration. Nonclinical Evaluation Report BNT162b2 [mRNA] COVID-19 vaccine (COMIRNATY); Health, Ed.; Department of Health and Aged Care, 2021; Vol. FOI 2389, Available online: https://www.tga.gov.au/sites/default/files/foi-2389-06.pdf.
- Neves, AR; Queiroz, JF; Costa Lima, SA; Figueiredo, F; Fernandes, R; Reis, S. Cellular uptake and transcytosis of lipid-based nanoparticles across the intestinal barrier: Relevance for oral drug delivery. J Colloid Interface Sci 2016, 463, 258–65. [Google Scholar] [CrossRef] [PubMed]
- Khare, P; Edgecomb, SX; Hamadani, CM; Tanner, EEL; S Manickam, D. Lipid nanoparticle-mediated drug delivery to the brain. Adv Drug Deliv Rev 2023, 197, 114861. [Google Scholar] [CrossRef]
- Chen, J; Xu, Y; Zhou, M; Xu, S; Varley, AJ; Golubovic, A; et al. Combinatorial design of ionizable lipid nanoparticles for muscle-selective mRNA delivery with minimized off-target effects. Proc Natl Acad Sci U S A 2023, 120(50), e2309472120. [Google Scholar] [CrossRef]
- Younis, MA; Sato, Y; Elewa, YHA; Kon, Y; Harashima, H. Self-homing nanocarriers for mRNA delivery to the activated hepatic stellate cells in liver fibrosis. J Control Release 2023, 353, 685–698. [Google Scholar] [CrossRef]
- Kent, SJ; Li, S; Amarasena, TH; Reynaldi, A; Lee, WS; Leeming, MG; et al. Blood distribution of SARS-CoV-2 lipid nanoparticle mRNA vaccine in humans. ACS Nano 2024, 18(39), 27077–27089. [Google Scholar] [CrossRef] [PubMed]
- Buckley, M; Araínga, M; Maiorino, L; Pires, IS; Kim, BJ; Michaels, KK; et al. Visualizing lipid nanoparticle trafficking for mRNA vaccine delivery in non-human primates. Mol Ther 2025, 33(3), 1105–1117. [Google Scholar] [CrossRef] [PubMed]
- Balcorta, HV; Mata Corral, MY; Gallegos, A; Chavez, J; Perez, J; Balivada, S; et al. Development of chemical tags for universal lipid nanoparticle visualization and tracking in 2D and 3D imaging. Nano Lett 2025, 25(19), 7682–7689. [Google Scholar] [CrossRef]
- Akhter, MH; Khalilullah, H; Gupta, M; Alfaleh, MA; Alhakamy, NA; Riadi, Y; et al. Impact of protein corona on the biological identity of nanomedicine: Understanding the fate of nanomaterials in the biological milieu. Biomedicines 2021, 9(10), 1496. [Google Scholar] [CrossRef]
- Behzadi, S; Serpooshan, V; Tao, W; Hamaly, MA; Alkawareek, MY; Dreaden, EC; et al. Cellular uptake of nanoparticles: journey inside the cell. Chem Soc Rev 2017, 46(14), 4218–4244. [Google Scholar] [CrossRef]
- Lavington, S; Watts, A. Lipid nanoparticle technologies for the study of G protein-coupled receptors in lipid environments. Biophys Rev 2020, 12(6), 1287–302. [Google Scholar] [CrossRef]
- Chatterjee, S; Kon, E; Sharma, P; Peer, D. Endosomal escape: A bottleneck for LNP-mediated therapeutics. Proc Natl Acad Sci U S A 2024, 121(11), e2307800120. [Google Scholar] [CrossRef]
- Vermeulen, LMP; Brans, T; Samal, SK; Dubruel, P; Demeester, J; De Smedt, SC; et al. Endosomal size and membrane leakiness influence proton sponge-based rupture of endosomal vesicles. ACS Nano 2018, 12(3), 2332–2345. [Google Scholar] [CrossRef] [PubMed]
- Johansson, JM; Du Rietz, H; Hedlund, H; Eriksson, HC; Oude Blenke, E; Pote, A; et al. Cellular and biophysical barriers to lipid nanoparticle mediated delivery of RNA to the cytosol. Nat Commun 2025, 16(1), 5354. [Google Scholar] [CrossRef] [PubMed]
- Er-Rafik, M; Ferji, K; Combet, J; Sandre, O; Lecommandoux, S; Schmutz, M; et al. Tear of lipid membranes by nanoparticles. Soft Matter 2022, 18(17), 3318–3322. [Google Scholar] [CrossRef]
- Pilkington, EH; Suys, EJA; Trevaskis, NL; Wheatley, AK; Zukancic, D; Algarni, A; et al. From influenza to COVID-19: Lipid nanoparticle mRNA vaccines at the frontiers of infectious diseases. Acta Biomater 2021, 131, 16–40. [Google Scholar] [CrossRef]
- Dowdy, SF. Endosomal escape of RNA therapeutics: How do we solve this rate-limiting problem? RNA 2023, 29(4), 396–401. [Google Scholar] [CrossRef] [PubMed]
- Sabnis, S; Kumarasinghe, ES; Salerno, T; Mihai, C; Ketova, T; Senn, JJ; et al. A novel amino lipid series for mRNA delivery: Improved endosomal escape and sustained pharmacology and safety in non-human primates. Mol Ther 2018, 26(6), 1509–1519. [Google Scholar] [CrossRef]
- Maugeri, M; Nawaz, M; Papadimitriou, A; Angerfors, A; Camponeschi, A; Na, M; et al. Linkage between endosomal escape of LNP-mRNA and loading into EVs for transport to other cells. Nat Commun 2019, 10, 4333. [Google Scholar] [CrossRef]
- Gilleron, J; Querbes, W; Zeigerer, A; Borodovsky, A; Marsico, G; Schubert, U; et al. Image-based analysis of lipid nanoparticle-mediated siRNA delivery, intracellular trafficking and endosomal escape. Nat Biotechnol 2013, 31(7), 638–46. [Google Scholar] [CrossRef]
- Aliakbarinodehi, N; Niederkofler, S; Emilsson, G; Parkkila, P; Olsén, E; Jing, Y; et al. Time-resolved inspection of ionizable lipid-facilitated lipid nanoparticle disintegration and cargo release at an early endosomal membrane mimic. ACS Nano 2024, 18(34), 22989–23000. [Google Scholar] [CrossRef]
- Schlich, M; Palomba, R; Costabile, G; Mizrahy, S; Pannuzzo, M; Peer, D; et al. Cytosolic delivery of nucleic acids: The case of ionizable lipid nanoparticles. Bioeng Transl Med 2021, 6(2), e10213. [Google Scholar] [CrossRef]
- Paramasivam, P; Franke, C; Stöter, M; Höijer, A; Bartesaghi, S; Sabirsh, A; et al. Endosomal escape of delivered mRNA from endosomal recycling tubules visualized at the nanoscale. J Cell Biol 2022, 221(2), e202110137. [Google Scholar] [CrossRef]
- Sahay, G; Querbes, W; Alabi, C; Eltoukhy, A; Sarkar, S; Zurenko, C; et al. Efficiency of siRNA delivery by lipid nanoparticles is limited by endocytic recycling. Nat Biotechnol 2013, 31(7), 653–8. [Google Scholar] [CrossRef]
- Bitounis, D; Jacquinet, E; Rogers, MA; Amiji, MM. Strategies to reduce the risks of mRNA drug and vaccine toxicity. Nat Rev Drug Discov. 2024, 23(4), 281–300. [Google Scholar] [CrossRef] [PubMed]
- Moghimi, SM; Simberg, D. Pro-inflammatory concerns with lipid nanoparticles. Mol Ther 2022, 30(6), 2109–2110. [Google Scholar] [CrossRef]
- Bates, SM; Munson, MJ; Trovisco, V; Pereira, S; Miller, SR; Sabirsh, A; et al. The kinetics of endosomal disruption reveal differences in lipid nanoparticle induced cellular toxicity. J Control Release 2025, 386, 114047. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L; Bandara, SR; Tan, Z; Leal, C. Lipid nanoparticle topology regulates endosomal escape and delivery of RNA to the cytoplasm. Proc Natl Acad Sci U S A 2023, 120(27), e2301067120. [Google Scholar] [CrossRef] [PubMed]
- Lu, ZR; Sun, D. Mechanism of pH-sensitive amphiphilic endosomal escape of ionizable lipid nanoparticles for cytosolic nucleic acid delivery. Pharm Res 2025l, 42(7), 1065–1077. [Google Scholar] [CrossRef]
- Su, J; Song, Y; Zhu, Z; Huang, X; Fan, J; Qiao, J; et al. Cell-cell communication: new insights and clinical implications. Signal Transduct Target Ther 2024, 9(1), 196. [Google Scholar] [CrossRef]
- Pateev, I; Seregina, K; Ivanov, R; Reshetnikov, V. Biodistribution of RNA vaccines and of their products: Evidence from human and animal studies. Biomedicines 2023, 12(1), 59. [Google Scholar] [CrossRef]
- Fertig, TE; Chitoiu, L; Marta, DS; Ionescu, VS; Cismasiu, VB; Radu, E; et al. Vaccine mRNA can be detected in blood at 15 days post-vaccination. Biomedicines 2022, 10(7), 1538. [Google Scholar] [CrossRef] [PubMed]
- Sahin, U; Karikó, K; Türeci, Ö. mRNA-based therapeutics--developing a new class of drugs. Nat Rev Drug Discov 2014, 13(10), 759–80. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y; Zhao, E; Yang, X; Luo, C; Zi, G; Wang, R; et al. Entrapment of lipid nanoparticles in peripheral endosomes but not lysosomes impairs intracellular trafficking and endosomal escape. Int J Pharm 2025, 669, 125024. [Google Scholar] [CrossRef]
- Omo-Lamai, S; Wang, Y; Patel, MN; Milosavljevic, A; Zuschlag, D; Poddar, S; et al. Limiting endosomal damage sensing reduces inflammation triggered by lipid nanoparticle endosomal escape. Nat Nanotechnol. 2025, 20(9), 1285–1297. [Google Scholar] [CrossRef]
- Bansal, S; Perincheri, S; Fleming, T; Poulson, C; Tiffany, B; Bremner, RM; et al. Cutting edge: Circulating exosomes with COVID spike protein are induced by BNT162b2 (Pfizer-BioNTech) vaccination prior to development of antibodies: A novel mechanism for immune activation by mRNA vaccines. J Immunol 2021, 207(10), 2405–2410. [Google Scholar] [CrossRef] [PubMed]
- Jörgensen, AM; Wibel, R; Bernkop-Schnürch, A. Biodegradable cationic and ionizable cationic lipids: A roadmap for safer pharmaceutical excipients. Small 2023, 19(17), e2206968. [Google Scholar] [CrossRef]
- Knaggs, KLM; Sun, Y; Walz, BA; Pang, J; Khan, OF. The role of excipients in lipid nanoparticle metabolism: implications for enhanced therapeutic effect. Ther Deliv 2025, 16(7), 687–700. [Google Scholar] [CrossRef]
- Wang, W; Deng, S; Lin, J; Ouyang, D. Modeling on in vivo disposition and cellular transportation of RNA lipid nanoparticles via quantum mechanics/physiologically-based pharmacokinetic approaches. Acta Pharm Sin B 2024, 14(10), 4591–4607. [Google Scholar] [CrossRef]
- Yu, Z; Li, Q; Wang, J; Yu, Y; Wang, Y; Zhou, Q; et al. Reactive oxygen species-related nanoparticle toxicity in the biomedical field. Nanoscale Res Lett 2020, 15(1), 115. [Google Scholar] [CrossRef] [PubMed]
- Packer, M; Gyawali, D; Yerabolu, R; Schariter, J; White, P. A novel mechanism for the loss of mRNA activity in lipid nanoparticle delivery systems. Nat Commun 2021, 12(1), 6777. [Google Scholar] [CrossRef]
- Hashiba, K; Taguchi, M; Sakamoto, S; Otsu, A; Maeda, Y; Ebe, H; et al. Overcoming thermostability challenges in mRNA-lipid nanoparticle systems with piperidine-based ionizable lipids. Commun Biol. 2024, 7(1), 556. [Google Scholar] [CrossRef]
- Cordes, J; Zhao, S; Engel, CM; Stingele, J. Cellular responses to RNA damage. Cell 2025, 188(4), 885–900. [Google Scholar] [CrossRef]
- Maelfait, J; Liverpool, L; Rehwinkel, J. Nucleic acid sensors and programmed cell death. J Mol Biol 2020, 432(2), 552–568. [Google Scholar] [CrossRef]
- Li, Y; Zhao, T; Li, J; Xia, M; Li, Y; Wang, X; et al. Oxidative stress and 4-hydroxy-2-nonenal (4-HNE): Implications in the pathogenesis and treatment of aging-related diseases. J Immunol Res. 2022, 2022, 2233906. [Google Scholar] [CrossRef]
- Zhou, ZH; Stone, CA, Jr.; Jakubovic, B; Phillips, EJ; Sussman, G; Park, J; et al. Anti-PEG IgE in anaphylaxis associated with polyethylene glycol. J Allergy Clin Immunol Pract 2021, 9(4), 1731–1733.e3. [Google Scholar] [CrossRef] [PubMed]
- Kozma, GT; Shimizu, T; Ishida, T; Szebeni, J. Anti-PEG antibodies: Properties, formation, testing and role in adverse immune reactions to PEGylated nano-biopharmaceuticals. Adv Drug Deliv Rev 2020, 154-155, 163–175. [Google Scholar] [CrossRef] [PubMed]
- Fu, S; Zhu, X; Huang, F; Chen, X. Anti-PEG antibodies and their biological impact on PEGylated drugs: Challenges and strategies for optimization. Pharmaceutics 2025, 17(8), 1074. [Google Scholar] [CrossRef]
- Szebeni, J. Complement activation-related pseudoallergy: a new class of drug-induced acute immune toxicity. Toxicology 2005, 216(2-3), 106–21. [Google Scholar] [CrossRef]
- Szebeni, J; Simberg, D; González-Fernández, Á; Barenholz, Y; Dobrovolskaia, MA. Roadmap and strategy for overcoming infusion reactions to nanomedicines. Nat Nanotechnol 2018, 13(12), 1100–1108. [Google Scholar] [CrossRef]
- Dézsi, L; Mészáros, T; Kozma, G; H-Velkei, M; Oláh, CZ; Szabó, M; et al. A naturally hypersensitive porcine model may help understand the mechanism of COVID-19 mRNA vaccine-induced rare (pseudo) allergic reactions: complement activation as a possible contributing factor. Geroscience 2022, 44(2), 597–618. [Google Scholar] [CrossRef] [PubMed]
- https. [CrossRef]
- Bakos, T; Mészáros, T; Kozma, GT; Berényi, P; Facskó, R; Farkas, H; et al. mRNA-LNP COVID-19 vaccine lipids induce complement activation and production of proinflammatory cytokines: Mechanisms, effects of complement inhibitors, and relevance to adverse reactions. Int J Mol Sci 2024, 25(7), 3595. [Google Scholar] [CrossRef]
- Song, J; Su, D; Wu, H; Guo, J. Implications of anaphylaxis following mRNA-LNP vaccines: It is urgent to eliminate PEG and find alternatives. Pharmaceutics 2025, 17(6), 798. [Google Scholar] [CrossRef]
- European Medicines Agency. Onpattro: European Public Assessment Report (EPAR) EMA/554262/2018; CHMP, Ed.; Amsterdam, NL, 2018. [Google Scholar]
- Wang, J; Ding, Y; Chong, K; Cui, M; Cao, Z; Tang, C; et al. Recent advances in lipid nanoparticles and their safety concerns for mRNA delivery. Vaccines (Basel) 2024, 12(10), 1148. [Google Scholar] [CrossRef]
- Dai, W; Xing, M; Sun, L; Lv, L; Wang, X; Wang, Y; et al. Lipid nanoparticles as adjuvant of norovirus VLP vaccine augment cellular and humoral immune responses in a TLR9- and type I IFN-dependent pathway. J Virology 2024, 98(12), e0169924. [Google Scholar] [CrossRef] [PubMed]
- https. [CrossRef]
- Chaudhary, N; Kasiewicz, LN; Newby, AN; Arral, ML; Yerneni, SS; Melamed, JR; et al. Amine headgroups in ionizable lipids drive immune responses to lipid nanoparticles by binding to the receptors TLR4 and CD1d. Nat Biomed Eng 2024, 8(11), 1483–1498. [Google Scholar] [CrossRef] [PubMed]
- Di Fusco, SA; Maggioni, AP; Bernelli, C; Perone, F; De Marzo, V; Conte, E; et al. Inclisiran: A new pharmacological approach for hypercholesterolemia. Rev Cardiovasc Med 2022, 23(11), 375. [Google Scholar] [CrossRef]
- Wright, RS; Koenig, W; Landmesser, U; Leiter, LA; Raal, FJ; Schwartz, GG; et al. Safety and tolerability of Inclisiran for treatment of hypercholesterolemia in 7 clinical trials. J Am Coll Cardiol 2023, 82(24), 2251–2261. [Google Scholar] [CrossRef] [PubMed]
- Urits, I; Swanson, D; Swett, MC; Patel, A; Berardino, K; Amgalan, A; et al. A Review of patisiran (ONPATTRO®) for the treatment of polyneuropathy in people with hereditary transthyretin amyloidosis. Neurol Ther. 2020, 9(2), 301–315. [Google Scholar] [CrossRef] [PubMed]
- Thi, TTH; Suys, EJA; Lee, JS; Nguyen, DH; Park, KD; Truong, NP. Lipid-based nanoparticles in the clinic and clinical trials: From cancer nanomedicine to COVID-19 vaccines. Vaccines (Basel) 2021, 9(4), 359. [Google Scholar] [CrossRef]
- European Medicines Agency. Kostaive (zapomeran). Public Assessment report Sections 2.4.3 (Pharmacokinetics) and 2.4.6 (Discussion on non-clinical aspects). 12 December 2024. Available online: https://www.ema.europa.eu/en/documents/assessment-report/kostaive-epar-public-assessment-report_en.pdf.
- Committee for Medicinal Products for Human Use (CHMP). Covid-19 Vaccine Moderna European Public Assessment Report (EPAR). Procedure No. EMEA/H/C/005791/0000. European Medicines Agency. 11 March 2021. Available online: https://www.ema.europa.eu/en/documents/assessment-report/spikevax-previously-covid-19-vaccine-moderna-epar-public-assessment-report_en.pdf.
- Wong, HL; Hu, M; Zhou, CK; Lloyd, PC; Amend, KL; Beachler, DC; et al. Risk of myocarditis and pericarditis after the COVID-19 mRNA vaccination in the USA: a cohort study in claims databases. Lancet 2022, 399(10342), 2191–2199. [Google Scholar] [CrossRef]
- Avraham, R; Haseley, N; Brown, D; Penaranda, C; Jijon, HB; Trombetta, JJ; et al. Pathogen cell-to-cell variability drives heterogeneity in host immune responses. Cell Erratum in: Cell 2015;163(2):523. 2015, 162(6), 1309–21. [Google Scholar] [CrossRef] [PubMed]
- Ndeupen, S; Qin, Z; Jacobsen, S; Bouteau, A; Estanbouli, H; Igyártó, BZ. The mRNA-LNP platform's lipid nanoparticle component used in preclinical vaccine studies is highly inflammatory. iScience 2021, 24(12), 103479. [Google Scholar] [CrossRef]
- Finney, DJ. The median lethal dose and its estimation. Arch Toxicol. 1985, 56(4), 215–8. [Google Scholar] [CrossRef] [PubMed]
- Avila, AM; Bebenek, I; Bonzo, JA; Bourcier, T; Davis Bruno, KL; Carlson, DB; et al. An FDA/CDER perspective on nonclinical testing strategies: Classical toxicology approaches and new approach methodologies (NAMs). Regul Toxicol Pharmacol 2020, 114, 104662. [Google Scholar] [CrossRef]
- Luster, AD. The role of chemokines in linking innate and adaptive immunity. Curr Opin Immunol. 2002, 14(1), 129–135. [Google Scholar] [CrossRef]
- Esche, C; Stellato, C; Beck, LA. Chemokines: key players in innate and adaptive immunity. J Invest Dermatol 2005, 125(4), 615–628. [Google Scholar] [CrossRef]
- Tao, T; Jiang, G; Su, Y; He, D; Zhu, L; Jiang, Q; Su, W. The multiple roles of GM-CSF in autoimmune and autoinflammatory uveitis. Biochem Pharmacol. 2025, 240, 117090. [Google Scholar] [CrossRef]
- Rincón, M; Anguita, J; Nakamura, T; Fikrig, E; Flavell, RA. Interleukin (IL)-6 directs the differentiation of IL-4-producing CD4+ T cells. J Exper Med. 1997, 185(3), 461–469. [Google Scholar] [CrossRef]
- de Jong, AJ; Pollastro, S; Kwekkeboom, JC; Andersen, SN; Dorjée, AL; Bakker, AM; et al. Functional and phenotypical analysis of IL-6-secreting CD4+ T cells in human adipose tissue. Eur J Immunol. 2018, 48(3), 471–481. [Google Scholar] [CrossRef]
- Jeannin, P; Jaillon, S; Delneste, Y. Pattern recognition receptors in the immune response against dying cells. Curr Opin Immunol 2008, 20(5), 530–537. [Google Scholar] [CrossRef] [PubMed]
- Nagai, Y; Garrett, KP; Ohta, S; Bahrun, U; Kouro, T; Akira, S; et al. Toll-like receptors on hematopoietic progenitor cells stimulate innate immune system replenishment. Immunity 2006, 24(6), 801–812. [Google Scholar] [CrossRef] [PubMed]
- Fritz, JH. Nod-like receptors have a grip on stem cells. Cell Host Microbe 2014, 15(6), 659–661. [Google Scholar] [CrossRef]
- Ji, Y; Kumar, R; Gokhale, A; Chao, HP; Rycaj, K; Chen, X; Li, Q; Tang, DG. LRIG1, a regulator of stem cell quiescence and a pleiotropic feedback tumor suppressor. Semin Cancer Biol 2022, 82, 120–133. [Google Scholar] [CrossRef] [PubMed]
- Parhiz, H; Brenner, JS; Patel, PN; Papp, TE; Shahnawaz, H; Li, Q; et al. Added to pre-existing inflammation, mRNA-lipid nanoparticles induce inflammation exacerbation (IE). J Control Release 2022, 344, 50–61. [Google Scholar] [CrossRef]
- Zelkoski, AE; Lu, Z; Sukumar, G; Dalgard, C; Said, H; Alameh, MG; et al. Ionizable lipid nanoparticles of mRNA vaccines elicit NF-κB and IRF responses through toll-like receptor 4. NPJ Vaccines 2025, 10(1), 73. [Google Scholar] [CrossRef]
- Raftery, N; Stevenson, NJ. Advances in anti-viral immune defence: revealing the importance of the IFN JAK/STAT pathway. Cell Mol Life Sci 2017, 74(14), 2525–2535. [Google Scholar] [CrossRef] [PubMed]
- Korzun, T; Moses, AS; Jozic, A; Grigoriev, V; Newton, S; Kim, J; et al. Lipid nanoparticles elicit reactogenicity and sickness behavior in mice via toll-like receptor 4 and myeloid differentiation protein 88 axis. ACS Nano 2024, 18(36), 24842–24859. [Google Scholar] [CrossRef] [PubMed]
- Mendoza, MC; Er, EE; Blenis, J. The Ras-ERK and PI3K-mTOR pathways: cross-talk and compensation. Trends Biochem Sci 2011, 36(6), 320–8. [Google Scholar] [CrossRef]
- Soriano, O; Alcón-Pérez, M; Vicente-Manzanares, M; Castellano, E. The crossroads between RAS and RHO signaling pathways in cellular transformation, motility and contraction. Genes (Basel) 2021, 12(6), 819. [Google Scholar] [CrossRef] [PubMed]
- Luo, J; Molbay, M; Chen, Y; Horvath, I; Kadletz, K; Kick, B; et al. Nanocarrier imaging at single-cell resolution across entire mouse bodies with deep learning. Nat Biotechnol 2025, 43, 2009–2022. [Google Scholar] [CrossRef]
- Knabl, L; Lee, HK; Wieser, M; Mur, A; Zabernigg, A; Knabl, L, Sr.; et al. BNT162b2 vaccination enhances interferon-JAK-STAT-regulated antiviral programs in COVID-19 patients infected with the SARS-CoV-2 Beta variant. Commun Med (Lond) 2022, 2(1), 17. [Google Scholar] [CrossRef]
- Hickey, TE; Mudunuri, U; Hempel, HA; Kemp, TJ; Roche, NV; Talsania, K; et al. Proteomic and serologic assessments of responses to mRNA-1273 and BNT162b2 vaccines in human recipient sera. Front Immunol 2025, 15, 1502458. [Google Scholar] [CrossRef]
- Panwar, V; Singh, A; Bhatt, M; Tonk, RK; Azizov, S; Raza, AS; et al. Multifaceted role of mTOR (mammalian target of rapamycin) signaling pathway in human health and disease. Signal Transduct Target Ther 2023, 8(1), 375. [Google Scholar] [CrossRef]
- Yu, H; Lin, L; Zhang, Z; Zhang, H; Hu, H. Targeting NF-κB pathway for the therapy of diseases: mechanism and clinical study. Signal Transduct Target Ther 2020, 5(1), 209. [Google Scholar] [CrossRef]
- Guo, Q; Jin, Y; Chen, X; Ye, X; Shen, X; Lin, M; et al. NF-κB in biology and targeted therapy: new insights and translational implications. Signal Transduct Target Ther 2024, 9(1), 53. [Google Scholar] [CrossRef]
- Bassermann, F; Eichner, R; Pagano, M. The ubiquitin proteasome system - implications for cell cycle control and the targeted treatment of cancer. Biochim Biophys Acta 2014, 1843(1), 150–62. [Google Scholar] [CrossRef]
- Yasir, M; Goyal, A; Sonthalia, S. Corticosteroid Adverse Effects. In StatPearls [Internet]; StatPearls Publishing: Treasure Island (FL), 3 Jul 2023; Available online: https://www.ncbi.nlm.nih.gov/books/NBK531462/.
- Gulluni, F; Martini, M; Hirsch, E. Cytokinetic Abscission: Phosphoinositides and ESCRTs direct the final cut. J Cell Biochem Erratum in: J Cell Biochem 2024;125(8):e30626. https://doi.org/10.1002/jcb.30626. 2017, 118(11), 3561–3568. [Google Scholar] [CrossRef]
- Tan, X; Thapa, N; Choi, S; Anderson, RA. Emerging roles of PtdIns(4,5)P2 -- beyond the plasma membrane. J Cell Sci 2015, 128(22), 4047–56. [Google Scholar] [CrossRef]
- Ajazi, A; Bruhn, C; Shubassi, G; Lucca, C; Ferrari, E; Cattaneo, A; et al. Endosomal trafficking and DNA damage checkpoint kinases dictate survival to replication stress by regulating amino acid uptake and protein synthesis. Dev Cell 2021, 56(18), 2607–2622.e6. [Google Scholar] [CrossRef] [PubMed]
- Giannini, C; Ponzone, L; Barroero, N; Hirsch, E. The interplay between phosphoinositides and ESCRT proteins. Adv Biol Regul 2025, 9, 101126. [Google Scholar] [CrossRef]
- Forster, III J; Nandi, D; Kulkarni, A. mRNA-carrying lipid nanoparticles that induce lysosomal rupture activate NLRP3 inflammasome and reduce mRNA transfection efficiency. Biomater Sci 2022, 10(19), 5566–5582. [Google Scholar] [CrossRef] [PubMed]
- Hurley, JH; Coyne, AN; Miączyńska, M; Stenmark, H. The expanding repertoire of ESCRT functions in cell biology and disease. Nature 2025, 642, 877–888. [Google Scholar] [CrossRef] [PubMed]
- Baylon, JL; Lenov, IL; Sligar, SG; Tajkhorshid, E. Characterizing the membrane-bound state of cytochrome P450 3A4: structure, depth of insertion, and orientation. J Am Chem Soc. 2013, 135(23), 8542–51. [Google Scholar] [CrossRef]
- Hackett, JC. Membrane-embedded substrate recognition by cytochrome P450 3A4. J Biol Chem 2018, 293(11), 403704046. [Google Scholar] [CrossRef]
- Mukherjee, G; Nandekar, PP; Wade, RC. An electron transfer competent structural ensemble of membrane-bound cytochrome P450 1A1 and cytochrome P450 oxidoreductase. Commun Biol. 2021, 4(1), 55. [Google Scholar] [CrossRef]
- Liu, F; Aulin, LBS; Manson, ML; Krekels, EHJ; van Hasselt, JGC. Unraveling the effects of acute inflammation on pharmacokinetics: A model-based analysis focusing on renal glomerular filtration rate and cytochrome P450 3A4-mediated metabolism. Eur J Drug Metab Pharmacokinet 2023, 48(6), 623–631. [Google Scholar] [CrossRef] [PubMed]
- Stanke-Labesque, F; Gautier-Veyret, E; Chhun, S; Guilhaumou, R; French Society of Pharmacology and Therapeutics. Inflammation is a major regulator of drug metabolizing enzymes and transporters: Consequences for the personalization of drug treatment. Pharmacol Ther 2020, 215, 107627. [Google Scholar] [CrossRef] [PubMed]
- Koozi, H; Lengquist, M; Frigyesi, A. C-reactive protein as a prognostic factor in intensive care admissions for sepsis: A Swedish multicenter study. J Crit Care 2020, 56, 73–79. [Google Scholar] [CrossRef]
- Bayraktar, İ; Yalçın, N; Demirkan, K. The potential interaction between COVID-19 vaccines and clozapine: A novel approach for clinical trials. Int J Clin Pract 2021, 75(8), e14441. [Google Scholar] [CrossRef]
- Imai, T; Ochiai, S; Ishimaru, T; Daitoku, H; Miyagawa, Y; Fukuhara, R; et al. A case report: Clozapine-induced leukopenia and neutropenia after mRNA COVID-19 vaccination. Neuropsychopharmacol Rep 2022, 42(2), 238–240. [Google Scholar] [CrossRef]
- Thompson, D; Delorme, CM; White, RF; Honer, WG. Elevated clozapine levels and toxic effects after SARS-CoV-2 vaccination. J Psychiatry Neurosci 2021, 46(2), E210–E211. [Google Scholar] [CrossRef]
- Eiermann, B; Engel, G; Johansson, I; Zanger, UM; Bertilsson, L. The involvement of CYP1A2 and CYP3A4 in the metabolism of clozapine. Br J Clin Pharmacol 1997, 44(5), 439–46. [Google Scholar] [CrossRef]
- Villemure, S; Trenaman, SC; Goralski, KB. The impact of COVID-19 infection on cytochrome P450 3A4-mediated drug metabolism and drug interactions. Expert Opin Drug Metab Toxicol 2023, 19(6), 329–332. [Google Scholar] [CrossRef]
- Lim, SYM; Al Bishtawi, B; Lim, W. Role of cytochrome P450 2C9 in COVID-19 treatment: Current status and future directions. Eur J Drug Metab Pharmacokinet 2023, 48(3), 221–240. [Google Scholar] [CrossRef]
- Hossam Abdelmonem, B; Abdelaal, NM; Anwer, EKE; Rashwan, AA; Hussein, MA; Ahmed, YF; et al. Decoding the role of CYP450 enzymes in metabolism and disease: A comprehensive review. Biomedicines 2024, 12(7), 1467. [Google Scholar] [CrossRef] [PubMed]
- Akanchise, T; Luo, F; Angelov, B; Deng, Y; Manna, G; Angelova, A. Rapid structural transformation of ionizable lipid nanoparticles involving Omega-3 polyunsaturated fatty acids enhances antioxidant defense and mitochondrial proteins activity in pH-responsive drug delivery. J Colloid Interface Sci. 2026, 704 Pt 2, 139420. [Google Scholar] [CrossRef]
- Ruysschaert, JM; Lonez, C. Role of lipid microdomains in TLR-mediated signalling. Biochim Biophys Acta 2015, 1848(9), 1860–1867. [Google Scholar] [CrossRef] [PubMed]
- Chaluvadi, MR; Nyagode, BA; Kinloch, RD; Morgan, ET. TLR4-dependent and -independent regulation of hepatic cytochrome P450 in mice with chemically induced inflammatory bowel disease. Biochem Pharmacol. 2009, 77(3), 464–471. [Google Scholar] [CrossRef]
- Li, Z; Carter, J; Santos, L; Webster, C; van der Walle, CF; Li, P; Rogers, SE; Lu, JR. Acidification-induced structure evolution of lipid nanoparticles correlates with their in vitro gene transfections. ACS Nano 2023, 17(2), 979–90. [Google Scholar] [CrossRef]
- Kim, KH; Ahn, T; Yun, CH. Membrane properties induced by anionic phospholipids and phosphatidylethanolamine are critical for the membrane binding and catalytic activity of human cytochrome P450 3A4. Biochemistry 2003, 42(51), 15377–15387. [Google Scholar] [CrossRef]
- Mustafa, G; Nandekar, PP; Mukherjee, G; Bruce, NJ; Wade, RC. The Effect of Force-Field Parameters on Cytochrome P450-Membrane Interactions: Structure and Dynamics. Sci Rep. 2020, 10(1), 7284. [Google Scholar] [CrossRef]
- Asady, B; Sampels, V; Romano, JD; et al. Function and regulation of a steroidogenic CYP450 enzyme in the mitochondrion of Toxoplasma gondii. PLoS Pathog 2023, 19(8), e1011566. [Google Scholar] [CrossRef]
- Huang, Z; Xu, X; Zhu, G. Association between coronavirus disease 2019 vaccination and hypercholesterolemia: A cross-sectional study from the 2023 National Health Interview Survey. Medicine (Baltimore) 2025, 104(48), e46221. [Google Scholar] [CrossRef] [PubMed]
- Luo, L; Wall, AA; Yeo, JC; Condon, ND; Norwood, SJ; Schoenwaelder, S; et al. Rab8a interacts directly with PI3Kγ to modulate TLR4-driven PI3K and mTOR signalling. Nat Commun. 2014, 5, 4407. [Google Scholar] [CrossRef]
- Płóciennikowska, A; Hromada-Judycka, A; Dembińska, J; Roszczenko, P; Ciesielska, A; Kwiatkowska, K. Contribution of CD14 and TLR4 to changes of the PI(4,5)P2 level in LPS-stimulated cells. J Leukoc Biol. 2016, 100(6), 1363–1373. [Google Scholar] [CrossRef] [PubMed]
- López-Haber, C; Levin-Konigsberg, R; Zhu, Y; Bi-Karchin, J; Balla, T; Grinstein, S; et al. Phosphatidylinositol-4-kinase IIα licenses phagosomes for TLR4 signaling and MHC-II presentation in dendritic cells. Proc Natl Acad Sci U S A 2020, 117(45), 28251–28262. [Google Scholar] [CrossRef]
- Quan, Y; Yang, H; Li, W; Li, L. mRNA vaccines: immunogenicity and quality characteristics. J Nanobiotechnology 2025, 24(1), 6. [Google Scholar] [CrossRef]
- Shi, Y; Ruan, H; Xu, Y; Zou, C. Cholesterol, eukaryotic lipid domains, and an evolutionary perspective of transmembrane signaling. Cold Spring Harb Perspect Biol 2023, 15(11), a041418. [Google Scholar] [CrossRef]
- Barnett, KC; Kagan, JC. Lipids that directly regulate innate immune signal transduction. Innate Immunity 2019, 26(1), 4–14. [Google Scholar] [CrossRef]
- Zheng, Y; Sun, L; Jiang, T; Zhang, D; He, D; Nie, H. TNFα promotes Th17 cell differentiation through IL-6 and IL-1β produced by monocytes in rheumatoid arthritis. J Immunol Res 2014, 2014, 385352. [Google Scholar] [CrossRef]
- Marasco, MR; Conteh, AM; Reissaus, CA; Cupit, JE, 5th; Appleman, EM; Mirmira, RG; et al. Interleukin-6 reduces β-cell oxidative stress by linking autophagy with the antioxidant response. Diabetes 2018, 67(8), 1576–1588. [Google Scholar] [CrossRef]
- Deretic, V; Saitoh, T; Akira, S. Autophagy in infection, inflammation and immunity. Nat Rev Immunol 2013, 13(10), 722–37. [Google Scholar] [CrossRef]
- Chi, DS; Fitzgerald, SM; Pitts, S; Cantor, K; King, E; Lee, SA; et al. MAPK-dependent regulation of IL-1- and beta-adrenoreceptor-induced inflammatory cytokine production from mast cells: implications for the stress response. BMC Immunol 2004, 5, 22. [Google Scholar] [CrossRef] [PubMed]
- https. [CrossRef]
- Yang, HT; Cohen, P; Rousseau, S. IL-1beta-stimulated activation of ERK1/2 and p38alpha MAPK mediates the transcriptional up-regulation of IL-6, IL-8 and GRO-alpha in HeLa cells. Cell Signal 2008, 20(2), 375–80. [Google Scholar] [CrossRef] [PubMed]
- Tengesdal, IW; Dinarello, A; Powers, NE; Burchill, MA; Joosten, LAB; Marchetti, C; et al. Tumor NLRP3-derived IL-1β drives the IL-6/STAT3 axis resulting in sustained MDSC-mediated immunosuppression. Front Immunol 2021, 12, 661323. [Google Scholar] [CrossRef] [PubMed]
- Li, H; Wu, M; Zhao, X. Role of chemokine systems in cancer and inflammatory diseases. MedComm (2020) 2022, 3(2), e147. [Google Scholar] [CrossRef] [PubMed]
- Unver, N. Macrophage chemoattractants secreted by cancer cells: Sculptors of the tumor microenvironment and another crucial piece of the cancer secretome as a therapeutic target. Cytokine Growth Factor Rev 2019, 50, 13–18. [Google Scholar] [CrossRef] [PubMed]
- Nakatsumi, H; Matsumoto, M; Nakayama, KI. Noncanonical pathway for regulation of CCL2 expression by an mTORC1-FOXK1 axis promotes recruitment of tumor-associated macrophages. Cell Rep 2017, 21(9), 2471–2486. [Google Scholar] [CrossRef]
- Huda, N; Khambu, B; Liu, G; Nakatsumi, H; Yan, S; Chen, X; et al. Senescence connects autophagy deficiency to inflammation and tumor progression in the liver. Cell Mol Gastroenterol Hepatol 2022, 14(2), 333–355. [Google Scholar] [CrossRef]
- Guo, L; Li, N; Yang, Z; Li, H; Zheng, H; Yang, J; et al. Role of CXCL5 in regulating chemotaxis of innate and adaptive leukocytes in infected lungs upon pulmonary influenza infection. Front Immunol 2021, 12, 785457. [Google Scholar] [CrossRef]
- Jastrzebska, B; Debinski, A; Filipek, S; Palczewski, K. Role of membrane integrity on G protein-coupled receptors: Rhodopsin stability and function. Prog Lipid Res. 2011, 50(3), 267–77. [Google Scholar] [CrossRef]
- Gimpl, G. Interaction of G protein coupled receptors and cholesterol. Chem Phys Lipids 2016, 199, 61–73. [Google Scholar] [CrossRef]
- Prasanna, X; Sengupta, D; Chattopadhyay, A. Cholesterol-dependent conformational plasticity in GPCR dimers. Sci Rep 2016, 6, 31858. [Google Scholar] [CrossRef]
- Prasanna, X; Mohole, M; Chattopadhyay, A; Sengupta, D. Role of cholesterol-mediated effects in GPCR heterodimers. Chem Phys Lipids 2020, 227, 104852. [Google Scholar] [CrossRef]
- Kumar, GA; Chattopadhyay, A. Cholesterol-dependent endocytosis of GPCRs: implications in pathophysiology and therapeutics. Biophys Rev 2021, 13(6), 1007–1017. 12. [Google Scholar] [CrossRef]
- Ray, AP; Thakur, N; Pour, NG; Eddy, MT. Dual mechanisms of cholesterol-GPCR interactions that depend on membrane phospholipid composition. Structure 2023, 31(7), 836–847.e6. [Google Scholar] [CrossRef]
- Lin, Y; Bian, L; Zhu, G; Zhang, B. Vitronectin promotes proliferation and metastasis of cervical cancer cells via the epithelial-mesenchymal transition. Front Oncol 2024, 14, 1466264. [Google Scholar] [CrossRef]
- Gao, P; Lu, Y; Tang, K; Wang, W; Wang, T; Zhu, Y; et al. Ficolin-1 ameliorates pulmonary fibrosis via directly binding to TGF-β1. J Transl Med 2024, 22(1), 1051. [Google Scholar] [CrossRef] [PubMed]
- Sozio, MS; Lu, C; Zeng, Y; Liangpunsakul, S; Crabb, DW. Activated AMPK inhibits PPAR-α and PPAR-γ transcriptional activity in hepatoma cells. Am J Physiol Gastrointest Liver Physiol 2011, 301(4), G739–47. [Google Scholar] [CrossRef] [PubMed]
- Sun, H; Zhu, X; Lin, W; Zhou, Y; Cai, W; Qiu, L. Interactions of TLR4 and PPARγ, dependent on AMPK signalling pathway contribute to anti-inflammatory effects of vaccariae hypaphorine in endothelial cells. Cell Physiol Biochem 2017, 42(3), 1227–1239. [Google Scholar] [CrossRef]
- Harada, K; Isse, K; Kamihira, T; Shimoda, S; Nakanuma, Y. Th1 cytokine-induced downregulation of PPARγ in human biliary cells relates to cholangitis in primary biliary cirrhosis. Hepatology 2005, 41(6), 1329–38. [Google Scholar] [CrossRef]
- Yin, R; Dong, YG; Li, HL. PPARγ phosphorylation mediated by JNK MAPK: a potential role in macrophage-derived foam cell formation. Acta Pharmacol Sin 2006, 27(9), 1146–52. [Google Scholar] [CrossRef]
- Su, AC; Zhang, LY; Zhang, JG; Hu, YY; Liu, XY; Li, SC; et al. The regulation of autophagy by p38 MAPK-PPARγ signaling during the brain ischemic tolerance induced by cerebral ischemic preconditioning. DNA Cell Biol 2022, 41(9), 838–849. [Google Scholar] [CrossRef] [PubMed]
- Ballav, S; Biswas, B; Sahu, VK; Ranjan, A; Basu, S. PPARγ partial agonists in disease-fate decision with special reference to cancer. Cells 2022, 11(20), 3215. [Google Scholar] [CrossRef]
- McCubrey, JA; Steelman, LS; Chappell, WH; Abrams, SL; Montalto, G; Cervello, M; et al. Mutations and deregulation of Ras/Raf/MEK/ERK and PI3K/PTEN/Akt/mTOR cascades which alter therapy response. Oncotarget 2012, 3(9), 954–87. [Google Scholar] [CrossRef]
- Nigam, M; Punia, B; Dimri, DB; Mishra, AP; Radu, AF; Bungau, G. Reactive oxygen species: A double-edged sword in the modulation of cancer signaling pathway dynamics. Cells 2025, 14(15), 1207. [Google Scholar] [CrossRef]
- Worthylake, RA; Burridge, K. RhoA and ROCK promote migration by limiting membrane protrusions. J Biol Chem 2003, 278(15), 13578–13584. [Google Scholar] [CrossRef]
- Yoshizaki, H; Ohba, Y; Parrini, MC; et al. Cell type-specific regulation of RhoA activity during cytokinesis. J Biol Chem 2004, 279(43), 44756–44762. [Google Scholar] [CrossRef]
- Omerovic, J; Prior, IA. Compartmentalized signalling: Ras proteins and signalling nanoclusters. FEBS J 2009, 276(7), 1817–1825. [Google Scholar] [CrossRef]
- https. [CrossRef]
- Schmick, M; Vartak, N; Papke, B; Kovacevic, M; Truxius, DC; Rossmannek, L; Bastiaens, PIH. KRas localizes to the plasma membrane by spatial cycles of solubilization, trapping and vesicular transport. Cell 2014, 10;157(2), 459–471. [Google Scholar] [CrossRef]
- Zhou, Y; Prakash, PS; Liang, H; Gorfe, AA; Hancock, JF. The KRAS and other prenylated polybasic domain membrane anchors recognize phosphatidylserine acyl chain structure. Proc Natl Acad Sci U S A 2021, 118(6), e2014605118. [Google Scholar] [CrossRef] [PubMed]
- Munro, E. Anillin puts RhoA in touch with PIP2. Dev Cell 2019, 49(6), 819–820. [Google Scholar] [CrossRef]
- Hancock, JF; Parton, RG. Ras plasma membrane signalling platforms. Biochem J 2005, 389 Pt 1, 1–11. [Google Scholar] [CrossRef]
- Jang, H; Abraham, SJ; Chavan, TS; Hitchinson, B; Khavrutskii, L; Tarasova, NI; et al. Mechanisms of membrane binding of small GTPase K-Ras4B farnesylated hypervariable region. J Biol Chem 2015, 290(15), 9465–77. [Google Scholar] [CrossRef]
- Mosaddeghzadeh, N; Kazemein Jasemi, NS; Majolée, J; Zhang, SC; Hordijk, PL; Dvorsky, R; et al. Electrostatic forces mediate the specificity of RHO GTPase-GDI interactions. Int J Mol Sci 2021, 22(22), 12493. [Google Scholar] [CrossRef] [PubMed]
- Guillot-Ferriols, M; Lanceros-Méndez, S; Gómez Ribelles, JL; Gallego Ferrer, G. Electrical stimulation: Effective cue to direct osteogenic differentiation of mesenchymal stem cells? Biomater Adv 2022, 138, 212918. [Google Scholar] [CrossRef]
- Chen, Z; Chen, Y; Zhe, M; Jiang, J; Liu, H; Qin, L; et al. Engineered smart piezoelectric materials facilitate bone defect regeneration. Materials & Design 2026, 262, 115501. [Google Scholar] [CrossRef]
- Chen, Y; Gong, W; Dai, W; Jiang, H; Xu, X. E2F1/2/4 mRNA is associated with immune infiltration and are potential biomarkers for the prognosis of human gastric carcinoma. Transl Cancer Res 2021, 10(6), 2801–2811. [Google Scholar] [CrossRef]
- Daigh, LH; Saha, D; Rosenthal, DL; Ferrick, KR; Meyer, T. Uncoupling of mTORC1 from E2F activity maintains DNA damage and senescence. Nat Commun 2024, 15, 9181. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y; Hao, X; Han, G; Lu, Y; Chen, Z; Zhang, L; et al. E2F1-mediated GINS2 transcriptional activation promotes tumor progression through PI3K/AKT/mTOR pathway in hepatocellular carcinoma. Am J Cancer Res 2022, 12(4), 1707–1726. [Google Scholar] [PubMed]
- Calzone, L; Gelay, A; Zinovyev, A; Radvanyi, F; Barillot, E. A comprehensive modular map of molecular interactions in RB/E2F pathway. Mol Syst Biol 2008, 4, 173. [Google Scholar] [CrossRef]
- Bertonnier-Brouty, L; Andersson, J; Kaprio, T; Hagström, J; Bsharat, S; Asplund, O; et al. E2F transcription factors promote tumorigenicity in pancreatic ductal adenocarcinoma. Cancer Med 2024, 13(9), e7187. [Google Scholar] [CrossRef]
- Wasserman, D; Nachum, S; Cohen, M; Enrico, TP; Noach-Hirsh, M; Parasol, J; et al. Cell cycle oscillators underlying orderly proteolysis of E2F8. Mol Biol Cell 2020, 31(8), 725–740. [Google Scholar] [CrossRef]
- Timmers, C; Sharma, N; Opavsky, R; Maiti, B; Wu, L; Wu, J; Orringer, D; et al. E2f1, E2f2, and E2f3 control E2F target expression and cellular proliferation via a p53-dependent negative feedback loop. Mol Cell Biol. Erratum in: Mol Cell Biol 2012;32(9):1758. 2007, 27(1), 65–78. [Google Scholar] [CrossRef]
- Spitschak, A; Dhar, P; Singh, KP; Casalegno Garduño, R; Gupta, SK; Vera, J; et al. E2F1-induced autocrine IL-6 inflammatory loop mediates cancer-immune crosstalk that predicts T cell phenotype switching and therapeutic responsiveness. Front Immunol 2024, 15, 1470368. [Google Scholar] [CrossRef] [PubMed]
- Li, J; Ran, C; Li, E; Gordon, F; Comstock, G; Siddiqui, H; et al. Synergistic function of E2F7 and E2F8 is essential for cell survival and embryonic development. Dev Cell 2008, 14(1), 62–75. [Google Scholar] [CrossRef]
- Zheng, J; Huang, J; Xia, J; Zhou, W; Dai, L; Lin, S; et al. Transcription factor E2F8 is a therapeutic target in the basal-like subtype of breast cancer. Front Oncol 2023, 13, 1038787. [Google Scholar] [CrossRef] [PubMed]
- Vigo, E; Müller, H; Prosperini, E; Hateboer, G; Cartwright, P; Moroni, MC; et al. CDC25A phosphatase is a target of E2F and is required for efficient E2F-induced S phase. Mol Cell Biol 1999, 19(9), 6379–95. [Google Scholar] [CrossRef] [PubMed]
- Delgado, M; Washam, CL; Urbaniak, A; Heflin, B; Storey, AJ; Lan, RS; et al. Phosphoproteomics provides novel insights into the response of primary acute lymphoblastic leukemia cells to microtubule depolymerization in G1 phase of the cell cycle. ACS Omega 2021, 6(38), 24949–24959. [Google Scholar] [CrossRef]
- Wang, X; Hills, LB; Huang, YH. Lipid and protein co-regulation of PI3K effectors Akt and Itk in lymphocytes. Front Immunol. 2015, 6, 117. [Google Scholar] [CrossRef]
- Sun, F; Yang, Y; Tu, H; Cai, H. Gradients of PI(4,5)P2 and PI(3,5)P2 jointly participate in shaping the back state of dictyostelium cells. Front Cell Dev Biol 2022, 10, 835185. [Google Scholar] [CrossRef]
- Sun, J; Zalejski, J; Song, S; Sharma, A; Wang, W; Hu, Y; et al. PI(3,5)P 2 controls the signaling activity of class I PI3K. bioRxiv Preprint 2025, 2023.01.25.525550. [Google Scholar] [CrossRef]
- Yang, M; Lu, Y; Piao, W; Jin, H. The Translational Regulation in mTOR Pathway. Biomolecules 2022, 12(6), 802. [Google Scholar] [CrossRef]
- Shveygert, M; Kaiser, C; Bradrick, SS; Gromeier, M. Regulation of eukaryotic initiation factor 4E (eIF4E) phosphorylation by mitogen-activated protein kinase occurs through modulation of Mnk1-eIF4G interaction. Mol Cell Biol 2010, 30(21), 5160–7. [Google Scholar] [CrossRef] [PubMed]
- Qin, Z; Bouteau, A; Herbst, C; Igyártó, BZ. Pre-exposure to mRNA-LNP inhibits adaptive immune responses and alters innate immune fitness in an inheritable fashion. PLoS Pathog 2022, 18(9), e1010830. [Google Scholar] [CrossRef]
- Lonez, C; Vandenbranden, M; Ruysschaert, JM. Cationic liposomal lipids: From gene carriers to cell signaling. Prog Lipid Res 2008, 47(5), 340–7. [Google Scholar] [CrossRef]
- Lonez, C; Vandenbranden, M; Ruysschaert, JM. Cationic lipids activate intracellular signaling pathways. Adv Drug Deliv Rev 2012, 64(15), 1749–58. [Google Scholar] [CrossRef]
- Connors, J; Joyner, D; Mege, NJ; Cusimano, GM; Bell, MR; Marcy, J; et al. Lipid nanoparticles (LNP) induce activation and maturation of antigen presenting cells in young and aged individuals. Commun Biol Erratum in: Commun Biol 2025;8(1):285. https://doi.org/10.1038/s42003-025-07603-0. 2023, 6(1), 188. [Google Scholar] [CrossRef]
- Amor, NP; Guo, K; Zhang, S; Xia, J; Yang, Y; Lin, A. Lipid nanoparticle: Beyond delivery vehicle-unveiling its immunological adjuvant potential. FASEB J 2025, 39(10), e70641. [Google Scholar] [CrossRef]
- Yamaguchi, Y; Kato, Y; Edahiro, R; Søndergaard, JN; Murakami, T; Amiya, S; et al. Consecutive BNT162b2 mRNA vaccination induces short-term epigenetic memory in innate immune cells. JCI Insight 2022, 7(22), e163347. [Google Scholar] [CrossRef]
- Simonis, A; Theobald, SJ; Koch, AE; Mummadavarapu, R; Mudler, JM; Pouikli, A; et al. Persistent epigenetic memory of SARS-CoV-2 mRNA vaccination in monocyte-derived macrophages. Mol Syst Biol 2025, 21(4), 341–360. [Google Scholar] [CrossRef] [PubMed]
- Hifdi, N; Vaucourt, M; Hnia, K; Panasyuk, G; Vandromme, M. Phosphoinositide signaling in the nucleus: Impacts on chromatin and transcription regulation. Biol Cell 2025, 117(1), e2400096. [Google Scholar] [CrossRef]
- Chytła, A; Rattay, S; Akgül, B; Sztacho, M. Plasma membrane and nuclear phosphatidylinositol 4,5-bisphosphate signalling in cancer. Lipids Health Dis 2025, 24(1), 39. [Google Scholar] [CrossRef] [PubMed]
- McColl, ER; Croyle, MA; Zamboni, WC; Honer, WG; Heise, M; Piquette-Miller, M; et al. COVID-19 Vaccines and the Virus: Impact on Drug Metabolism and Pharmacokinetics. Drug Metab Dispos 2023, 51(1), 130–141. [Google Scholar] [CrossRef]
- Jung, SW; Jeon, JJ; Kim, YH; Choe, SJ; Lee, S. Long-term risk of autoimmune diseases after mRNA-based SARS-CoV2 vaccination in a Korean, nationwide, population-based cohort study. Nat Commun 2024, 15, 6181. [Google Scholar] [CrossRef]
- Sellegounder, D; Ferrucci, L; Anbazhagan, R; Basisty, N. Editorial: Molecular crosstalk between endocrine factors, paracrine signals, and the immune system during aging. Front Endocrinol (Lausanne) 2023, 14, 1203755. [Google Scholar] [CrossRef]
- Chen, M; Tan, JX; Sun, Y; Thapa, N; Cryns, VL; Anderson, RA. Agonist- and stress-driven compartmentalized phosphoinositide signaling in cells. Biochim Biophys Acta Mol Cell Biol Lipids 2025, 1870(6), 159662. [Google Scholar] [CrossRef]
- Ebner, M; Sinkovics, B; Szczygieł, M; Ribeiro, DW; Yudushkin, I. Localization of mTORC2 activity inside cells. J Cell Biol 2017, 216(2), 343–353. [Google Scholar] [CrossRef]
- Rädler, PD; Wehde, BL; Wagner, KU. Crosstalk between STAT5 activation and PI3K/AKT functions in normal and transformed mammary epithelial cells. Mol Cell Endocrinol 2017, 451, 31–39. [Google Scholar] [CrossRef]
- Flori, E; Cavallo, A; Mosca, S; et al. JAK/STAT inhibition normalizes lipid composition in 3D human epidermal equivalents challenged with Th2 cytokines. Cells 2024, 13(9), 760. [Google Scholar] [CrossRef] [PubMed]
- Koester, AM; Tao, K; Szczepaniak, M; Rames, MJ; Nan, X. Nanoscopic spatial association between Ras and phosphatidylserine on the cell membrane studied with multicolor super resolution microscopy. Biomolecules 2022, 12(8), 1033. [Google Scholar] [CrossRef] [PubMed]
- Panariti, A; Miserocchi, G; Rivolta, I. The effect of nanoparticle uptake on cellular behavior: disrupting or enabling functions? Nanotechnol Sci Appl 2012, 5, 87–100. [Google Scholar] [CrossRef]
- Voigt, J; Christensen, J; Shastri, VP. Differential uptake of nanoparticles by endothelial cells through polyelectrolytes with affinity for caveolae. Proc Natl Acad Sci U S A 2014, 111(8), 2942–7. [Google Scholar] [CrossRef] [PubMed]
- Wang, T; Bai, J; Jiang, X; Nienhaus, GU. Cellular uptake of nanoparticles by membrane penetration: a study combining confocal microscopy with FTIR spectroelectrochemistry. ACS Nano 2012, 6(2), 1251–9. [Google Scholar] [CrossRef]
- Gerelli, Y. Chapter Three: Exploring interactions between lipid membranes and nanoparticles through neutron and X-ray reflectometry techniques. In Advances in Biomembranes and Lipid Self-Assembly; Iglič, A., Rappolt, M., Losada-Pérez, P., Eds.; Academic Press, 2023; Volume 38, pp. 37–61. [Google Scholar] [CrossRef]
- Lavagna, E; Barnoud, J; Rossi, G; Monticelli, L. Size-dependent aggregation of hydrophobic nanoparticles in lipid membranes. Nanoscale 2020, 12, 9452–9461. [Google Scholar] [CrossRef]
- Cooper, GM. The Cell: A Molecular Approach, 2nd edition; Sinauer Associates: Sunderland (MA), 2000. [Google Scholar]
- Hammond, GRV; Burke, JE. Novel roles of phosphoinositides in signaling, lipid transport, and disease. Curr Opin Cell Biol 2020, 63, 57–67. [Google Scholar] [CrossRef]
- Skotland, T; Kavaliauskiene, S; Sandvig, K. The role of lipid species in membranes and cancer-related changes. Cancer Metastasis Rev 2020, 39(2), 343–360. [Google Scholar] [CrossRef]
- https. [CrossRef]
- Heimburg, T. The excitable fluid mosaic. Biochim Biophys Acta Biomembr 2023, 1865(3), 184104. [Google Scholar] [CrossRef]
- Lupyan, D; Mezei, M; Logothetis, DE; Osman, R. A molecular dynamics investigation of lipid bilayer perturbation by PIP2. Biophys J 2010, 98(2), 240–7. [Google Scholar] [CrossRef] [PubMed]
- Pike, LJ. Lipid rafts: bringing order to chaos. J Lipid Res 2003, 44(4), 655–67. [Google Scholar] [CrossRef]
- Levin, R; Grinstein, S; Schlam, D. Phosphoinositides in phagocytosis and macropinocytosis. Biochim Biophys Acta 2015, 1851(6), 805–23. [Google Scholar] [CrossRef] [PubMed]
- Weiner, OD; Neilsen, PO; Prestwich, GD; Kirschner, MW; Cantley, LC; Bourne, HR. A PtdInsP(3)- and Rho GTPase-mediated positive feedback loop regulates neutrophil polarity. Nat Cell Biol 2002, 4(7), 509–13. [Google Scholar] [CrossRef]
- Guan, K; Curtis, ER; Lew, DJ; Elston, TC. Particle-based simulations reveal two positive feedback loops allow relocation and stabilization of the polarity site during yeast mating. PLoS Comput Biol 2023, 19(10), e1011523. [Google Scholar] [CrossRef] [PubMed]
- Fallahi-Sichani, M; Linderman, JJ. Lipid raft-mediated regulation of G-protein coupled receptor signaling by ligands which influence receptor dimerization: a computational study. PLoS One 2009, 4(8), e6604. [Google Scholar] [CrossRef]
- Wang, X; Shi, X; Wang, R. Regulating mRNA endosomal escape through lipid rafts: A review. Int J Pharm 2025, 675, 125571. [Google Scholar] [CrossRef] [PubMed]
- Sarkar, S; Carroll, B; Buganim, Y; Maetzel, D; Ng, AH; Cassady, JP; et al. Impaired autophagy in the lipid-storage disorder Niemann-Pick type C1 disease. Cell Rep 2013, 5(5), 1302–15. [Google Scholar] [CrossRef]
- Dall'Armi, C; Devereaux, KA; Di Paolo, G. The role of lipids in the control of autophagy. Curr Biol 2013, 23(1), R33–45. [Google Scholar] [CrossRef]
- Zhang, S; Peng, X; Yang, S; Li, X; Huang, M; Wei, S; et al. The regulation, function, and role of lipophagy, a form of selective autophagy, in metabolic disorders. Cell Death Dis 2022, 13(2), 132. [Google Scholar] [CrossRef] [PubMed]
- Jarocki, M; Turek, K; Saczko, J; Tarek, M; Kulbacka, J. Lipids associated with autophagy: mechanisms and therapeutic targets. Cell Death Discov 2024, 10(1), 460. [Google Scholar] [CrossRef]
- Karim, M; Mishra, M; Lo, CW; Saul, S; Cagirici, HB; Gourdelier, M; et al. PIP4K2C inhibition reverses autophagic flux impairment induced by SARS-CoV-2. Nat Commun 2025, 16(1), 6397. [Google Scholar] [CrossRef]
- Haucke, V; Kozlov, MM. Membrane remodeling in clathrin-mediated endocytosis. J Cell Sci 2018, 131(17), jcs216812. [Google Scholar] [CrossRef]
- Vaithianathan, T; Bukiya, A; Liu, J; Liu, P; Asuncion-Chin, M; Fan, Z; Dopico, A. Direct regulation of BK channels by phosphatidylinositol 4,5-bisphosphate as a novel signaling pathway. J Gen Physiol 2008, 132(1), 13–28. [Google Scholar] [CrossRef]
- Blunsom, NJ; Cockcroft, S. Phosphatidylinositol synthesis at the endoplasmic reticulum. Biochim Biophys Acta Mol Cell Biol Lipids 2020, 1865(1), 158471. [Google Scholar] [CrossRef]
- Posor, Y; Jang, W; Haucke, V. Phosphoinositides as membrane organizers. Nat Rev Mol Cell Biol 2022, 23(12), 797–816. [Google Scholar] [CrossRef]
- Lolicato, F; Nickel, W; Haucke, V; Ebner, M. Phosphoinositide switches in cell physiology - From molecular mechanisms to disease. J Biol Chem 2024, 300(3), 105757. [Google Scholar] [CrossRef]
- Balla, T. Phosphoinositides: Tiny lipids with giant impact on cell regulation. Physiol Rev 2013, 93(3), 1019–137. [Google Scholar] [CrossRef]
- Eramo, MJ; Mitchell, CA. Regulation of PtdIns(3,4,5)P3/Akt signalling by inositol polyphosphate 5-phosphatases. Biochem Soc Trans 2016, 44(1), 240–52. [Google Scholar] [CrossRef] [PubMed]
- Xu, S; Cao, B; Xuan, G; Xu, S; An, Z; Zhu, C; et al. Function and regulation of Rab GTPases in cancers. Cell Biol Toxicol 2024, 40(1), 28. [Google Scholar] [CrossRef] [PubMed]
- Koike, S; Jahn, R. Rab GTPases and phosphoinositides fine-tune SNAREs dependent targeting specificity of intracellular vesicle traffic. Nat Commun 2024, 15(1), 2508. [Google Scholar] [CrossRef]
- Puranik, A; Lenehan, PJ; Silvert, E; Niesen, MJM; Corchado-Garcia, J; O'Horo, JC; et al. Comparative effectiveness of mRNA-1273 and BNT162b2 against symptomatic SARS-CoV-2 infection. Med 2022, 3(1), 28–41.e8. [Google Scholar] [CrossRef] [PubMed]
- Granados-Riveron, JT; Aquino-Jarquin, G. Engineering of the current nucleoside-modified mRNA-LNP vaccines against SARS-CoV-2. Biomed Pharmacother 2021, 142, 111953. [Google Scholar] [CrossRef]
- Harvey, RD; Ara, N; Heenan, RK; Barlow, DJ; Quinn, PJ; Lawrence, MJ. Stabilization of distearoylphosphatidylcholine lamellar phases in propylene glycol using cholesterol. Mol Pharm 2013, 10(12), 4408–17. [Google Scholar] [CrossRef]
- Li, J; Wang, X; Zhang, T; Wang, C; Huang, Z; Luo, X; Deng, Y. A review on phospholipids and their main applications in drug delivery systems. Asian J Pharm Sci 2015, 10(2), 81–98. [Google Scholar] [CrossRef]
- McMaster, CR. From yeast to humans - roles of the Kennedy pathway for phosphatidylcholine synthesis. FEBS Lett 2018, 592(8), 1256–1272. [Google Scholar] [CrossRef]
- Cummings, R; Parinandi, N; Wang, L; Usatyuk, P; Natarajan, V. Phospholipase D/phosphatidic acid signal transduction: role and physiological significance in lung. Mol Cell Biochem 2002, 234-235(1-2), 99–109. [Google Scholar] [CrossRef]
- Wagner, K; Brezesinski, G. Phospholipase D activity is regulated by product segregation and the structure formation of phosphatidic acid within model membranes. Biophys J 2007, 93(7), 2373–83. [Google Scholar] [CrossRef]
- Bruntz, RC; Lindsley, CW; Brown, HA. Phospholipase D signaling pathways and phosphatidic acid as therapeutic targets in cancer. Pharmacol Rev 2014, 66(4), 1033–79. [Google Scholar] [CrossRef]
- Semenkovich, CF; Goldberg, AC; Goldberg, IJ. Disorders of Lipid Metabolism. Chapter 37 in Melmed S, Polonsky KS, Larsen PR, Kronenberg HM, Eds. Williams Textbook of Endocrinology (Thirteenth Edition). Elsevier; Volume 2016, pp. 1660–1700.
- https. [CrossRef]
- Pearce, B; Jakobson, K; Morrow, C; Murphy, S. Phosphatidic acid promotes phosphoinositide metabolism and DNA synthesis in cultured cortical astrocytes. Neurochem Int 1994, 24(2), 165–71. [Google Scholar] [CrossRef] [PubMed]
- Quick, J; Santos, ND; Cheng, MHY; Chander, N; Brimacombe, CA; Kulkarni, J; et al. Lipid nanoparticles to silence androgen receptor variants for prostate cancer therapy. J Control Release 2022, 349, 174–183. [Google Scholar] [CrossRef] [PubMed]
- Jeschek, D; Lhota, G; Wallner, J; Vorauer-Uhl, K. A versatile, quantitative analytical method for pharmaceutical relevant lipids in drug delivery systems. J Pharm Biomed Anal 2016, 119, 37–44. [Google Scholar] [CrossRef]
- Rezaei, S; Blick, EE; Mineart, KP; Kelley, EG. Chapter Three - Linking chemical degradation and physical instability of lipid vesicles. Advances in Biomembranes and Lipid Self-Assembly 2025, 41, 47–64. [Google Scholar] [CrossRef]
- Nishimura, T; Gecht, M; Covino, R; Hummer, G; Surma, MA; Klose, C; et al. Osh proteins control nanoscale lipid organization necessary for PI(4,5)P2 synthesis. Mol Cell 2019, 75(5), 1043–1057.e8. [Google Scholar] [CrossRef]
- Heckle, LA; Kozminski, KG. Osh-dependent and -independent regulation of PI4P levels during polarized growth of Saccharomyces cerevisiae. Mol Biol Cell 2023, 34(11), ar104. [Google Scholar] [CrossRef]
- Raychaudhuri, S; Prinz, WA. The diverse functions of oxysterol-binding proteins. Annu Rev Cell Dev Biol 2010, 26, 157–77. [Google Scholar] [CrossRef]
- Leonarduzzi, G; Robbesyn, F; Poli, G. Signaling kinases modulated by 4-hydroxynonenal. Free Radic Biol Med. 2004, 37(11), 1694–702. [Google Scholar] [CrossRef]
- Forman, HJ; Fukuto, JM; Miller, T; Zhang, H; Rinna, A; Levy, S. The chemistry of cell signaling by reactive oxygen and nitrogen species and 4-hydroxynonenal. Arch Biochem Biophys. 2008, 477(2), 183–95. [Google Scholar] [CrossRef]
- Thiemicke, A; Neuert, G. Rate thresholds in cell signaling have functional and phenotypic consequences in non-linear time-dependent environments. Front Cell Dev Biol 2023, 11, 1124874. [Google Scholar] [CrossRef]
- Hou, X; Chen, Y; Carrillo, ND; Cryns, VL; Anderson, RA; Sun, J; et al. Phosphoinositide signaling at the cytoskeleton in the regulation of cell dynamics. Cell Death Dis 2025, 16(1), 296. [Google Scholar] [CrossRef]
- Adhikari, H; Cullen, PJ. Role of phosphatidylinositol phosphate signaling in the regulation of the filamentous-growth mitogen-activated protein kinase pathway. Eukaryot Cell 2015, 14(4), 427–40. [Google Scholar] [CrossRef]
- Houthaeve, G; De Smedt, SC; Braeckmans, K; De Vos, WH. The cellular response to plasma membrane disruption for nanomaterial delivery. Nano Converg 2022, 9(1), 6. [Google Scholar] [CrossRef]
- Papafilippou, L; Nicolaou, A; Kendall, AC; Camacho-Muñoz, D; Hadjidemetriou, M. The lipidomic profile of the nanoparticle-biomolecule corona reflects the diversity of plasma lipids. Nanoscale 2023, 15, 11038–11051. [Google Scholar] [CrossRef]
- Gual, P; Grémeaux, T; Gonzalez, T; Le Marchand-Brustel, Y; Tanti, JF. MAP kinases and mTOR mediate insulin-induced phosphorylation of insulin receptor substrate-1 on serine residues 307, 612 and 632. Diabetologia 2003, 46(11), 1532–1542. [Google Scholar] [CrossRef]
- Minard, AY; Tan, SX; Yang, P; et al. mTORC1 Is a Major Regulatory Node in the FGF21 Signaling Network in Adipocytes. Cell Rep. 2016, 17(1), 29–36. [Google Scholar] [CrossRef]
- Le, TKC; Dao, XD; Nguyen, DV; Luu, DH; Bui, TMH; Le, TH; et al. Insulin signaling and its application. Front Endocrinol (Lausanne) 2023, 14, 1226655. [Google Scholar] [CrossRef]
- Krauson, AJ; Casimero, FVC; Siddiquee, Z; Stone, JR. Duration of SARS-CoV-2 mRNA vaccine persistence and factors associated with cardiac involvement in recently vaccinated patients. NPJ Vaccines 2023, 8(1), 141. [Google Scholar] [CrossRef]
- Rosati, M; Terpos, E; Homan, P; Bergamaschi, C; Karaliota, S; Ntanasis-Stathopoulos, I; et al. Rapid transient and longer-lasting innate cytokine changes associated with adaptive immunity after repeated SARS-CoV-2 BNT162b2 mRNA vaccinations. Front Immunol. 2023, 14, 1292568. [Google Scholar] [CrossRef]
- Lee, Y; Jeong, M; Park, J; Jung, H; Lee, H. Immunogenicity of lipid nanoparticles and its impact on the efficacy of mRNA vaccines and therapeutics. Exp Mol Med. 2023, 55(10), 2085–2096. [Google Scholar] [CrossRef]
- Iordanov, MS; Paranjape, JM; Zhou, A; Wong, J; Williams, BR; Meurs, EF; et al. Activation of p38 mitogen-activated protein kinase and c-Jun NH(2)-terminal kinase by double-stranded RNA and encephalomyocarditis virus: involvement of RNase L, protein kinase R, and alternative pathways. Mol Cell Biol. 2000, 20(2), 617–27. [Google Scholar] [CrossRef]
- Jurkiewicz, A; Graczyk, D. MAP kinases are involved in RNA polymerase III regulation upon LPS treatment in macrophages. Gene 2022, 831, 146548. [Google Scholar] [CrossRef]
- Zhang, G; He, LS; Him Wong, Y; Xu, Y; Zhang, Y; Qian, PY. p38 MAPK regulates PKAα and CUB-serine protease in Amphibalanus amphitrite cyprids. Sci Rep. 2015, 5, 14767. [Google Scholar] [CrossRef]
- Tang, T; Zhu, Q; Li, X; Zhu, G; Deng, S; Wang, Y; et al. Protease Nexin I is a feedback regulator of EGF/PKC/MAPK/EGR1 signaling in breast cancer cells metastasis and stemness. Cell Death Dis. Erratum. 2019, 10(9), 649. [Google Scholar] [CrossRef]
- Sala-Gaston, J; Costa-Sastre, L; Pedrazza, L; Martinez-Martinez, A; Ventura, F; Rosa, JL. Regulation of MAPK signaling pathways by the large HERC ubiquitin ligases. Int J Mol Sci. 2023, 24(5), 4906. [Google Scholar] [CrossRef]
- Lutz, J; Lazzaro, S; Habbeddine, M; Schmidt, KE; Baumhof, P; Mui, BL; et al. Unmodified mRNA in LNPs constitutes a competitive technology for prophylactic vaccines. NPJ Vaccines 2017, 2, 29. [Google Scholar] [CrossRef]
- Pang, SJ; Jiang, TY; Wang, NG; Cui, XW; Wang, H; Pan, YF; et al. RNA polymerase II subunit 5-mediating protein limits TLR4-induced innate immune activation in macrophages by inhibiting IKKβ/NF-κB signaling during sepsis. Cell Commun Signal. 2025, 23(1), 274. [Google Scholar] [CrossRef]
- Ostendorf, T; Zillinger, T; Andryka, K; Schlee-Guimaraes, TM; Schmitz, S; Marx, S; et al. Immune sensing of synthetic, bacterial, and protozoan RNA by toll-like receptor 8 requires coordinated processing by RNase T2 and RNase 2. Immunity 2020, 52(4), 591–605.e6. [Google Scholar] [CrossRef]
- Lee, D; Le Pen, J; Yatim, A; Dong, B; Aquino, Y; Ogishi, M; et al. Inborn errors of OAS-RNase L in SARS-CoV-2-related multisystem inflammatory syndrome in children. Science 2023, 379(6632), eabo3627. [Google Scholar] [CrossRef]
- Bucci, M; Vellecco, V; Harrington, L; Brancaleone, V; Roviezzo, F; Mattace Raso, G; et al. Cross-talk between toll-like receptor 4 (TLR4) and proteinase-activated receptor 2 (PAR(2) ) is involved in vascular function. Br J Pharmacol. 2013, 168(2), 411–20. [Google Scholar] [CrossRef]
- Zhong, B; Liu, X; Wang, X; Liu, X; Li, H; Darnay, BG; et al. Ubiquitin-specific protease 25 regulates TLR4-dependent innate immune responses through deubiquitination of the adaptor protein TRAF3. Sci Signal. 2013, 6(275), ra35. [Google Scholar] [CrossRef]
- Papendorf, JJ; Krüger, E; Ebstein, F. Proteostasis perturbations and their roles in causing sterile inflammation and autoinflammatory diseases. Cells 2022, 11(9), 1422. [Google Scholar] [CrossRef] [PubMed]
- Guo, X; Wang, H; Li, Y; Leng, X; Huang, W; Ma, Y; et al. Transfection reagent Lipofectamine triggers type I interferon signaling activation in macrophages. Immunol Cell Biol. Erratum in: Immunol Cell Biol. 2020;98(1):88. https://doi.org/10.1111/imcb.12300. 2019, 97(1), 92–96. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Causality assessment of an adverse event following immunization (AEFI): user manual for the revised WHO classification, 2nd ed.; Pharmacovigilance 2019 Geneva, Switzerland: Department, 2019; Available online: https://www.who.int/publications/i/item/9789241516990.
- Bellavite, P. Causality assessment of adverse events following immunization: the problem of multifactorial pathology. F1000research 2020, 9, 170. [Google Scholar] [CrossRef]
- Islam, S; Bhattacharya, S. Dynamical systems theory as an organizing principle for single-cell biology. NPJ Syst Biol Appl. 2025, 11(1), 85. [Google Scholar] [CrossRef] [PubMed]
- Philipps, M; Schmid, N; Hasenauer, J. Current state and open problems in universal differential equations for systems biology. NPJ Syst Biol Appl. 2025, 11(1), 101. [Google Scholar] [CrossRef]
- Simons, BD; Karin, O. Cell cycle criticality as a mechanism for robust cell population control. Mol Syst Biol. 2025, 22, 241–258. [Google Scholar] [CrossRef] [PubMed]
- Fischer, DS; Villanueva, MA; Winter, PS; Shalek, AK. Adapting systems biology to address the complexity of human disease in the single-cell era. Nat Rev Genet. 2025, 26, 514–531. [Google Scholar] [CrossRef]
- U. S. Pharmacopeia (USP). Analytical procedures for mRNA Vaccine Quality. 3rd Edition. 2024. Available online: https://go.usp.org/mRNAVaccineQuality (accessed on 13 February 2026).
- Committee for Medicinal Products for Human Use (CHMP). Guideline on the quality aspects of mRNA vaccines: Draft. Euopean Medicines Agency. Amsterdam, 2025. Available online: https://www.ema.europa.eu/en/documents/scientific-guideline/draft-guideline-quality-aspects-mrna-vaccines_en.pdf.
- Hassett, KJ; Higgins, J; Woods, A; Levy, B; Xia, Y; Hsiao, CJ; et al. Impact of lipid nanoparticle size on mRNA vaccine immunogenicity. J Control Release 2021, 335, 237–246. [Google Scholar] [CrossRef]
- Arral, ML; Whitehead, KA. Design principles of lipid nanoparticles for RNA delivery. Nat Rev Bioeng Epub ahead of print. 2026. [Google Scholar] [CrossRef]
- Karikó, K; Ni, H; Capodici, J; Lamphier, M; Weissman, D. mRNA is an endogenous ligand for Toll-like receptor 3. J Biol Chem 2004, 279(13), 12542–50. [Google Scholar] [CrossRef]
- Karikó, K; Buckstein, M; Ni, H; Weissman, D. Suppression of RNA recognition by Toll-like receptors: the impact of nucleoside modification and the evolutionary origin of RNA. Immunity 2005, 23(2), 165–75. [Google Scholar] [CrossRef]
- Karikó, K; Muramatsu, H; Welsh, FA; Ludwig, J; Kato, H; Akira, S; et al. Incorporation of pseudouridine into mRNA yields superior nonimmunogenic vector with increased translational capacity and biological stability. Mol Ther 2008, 16(11), 1833–40. [Google Scholar] [CrossRef]
- Monroe, J; Eyler, DE; Mitchell, L; Deb, I; Bojanowski, A; Srinivas, P; et al. N1-Methylpseudouridine and pseudouridine modifications modulate mRNA decoding during translation. Nat Commun 2024, 15(1), 8119. [Google Scholar] [CrossRef]
- Ghosh, TK; Mickelson, DJ; Solberg, JC; Lipson, KE; Inglefield, JR; Alkan, SS. TLR-TLR cross talk in human PBMC resulting in synergistic and antagonistic regulation of type-1 and 2 interferons, IL-12 and TNF-alpha. Int Immunopharmacol 2007, 7(8), 1111–21. [Google Scholar] [CrossRef]
- Xu, XH; Shah, PK; Faure, E; Equils, O; Thomas, L; Fishbein, MC. Toll-like receptor-4 is expressed by macrophages in murine and human lipid-rich atherosclerotic plaques and upregulated by oxidized LDL. Circulation 2001, 104(25), 3103–8. [Google Scholar] [CrossRef]
- Hovland, A; Jonasson, L; Garred, P; Yndestad, A; Aukrust, P; Lappegård, KT; et al. The complement system and toll-like receptors as integrated players in the pathophysiology of atherosclerosis. Atherosclerosis 2015, 241(2), 480–94. [Google Scholar] [CrossRef] [PubMed]
- Luo, L; Wall, AA; Tong, SJ; Hung, Y; Xiao, Z; Tarique, AA; et al. TLR crosstalk activates LRP1 to recruit Rab8a and PI3Kγ for suppression of inflammatory responses. Cell Rep 2018, 24(11), 3033–3044. [Google Scholar] [CrossRef] [PubMed]
- Köberlin, MS; Heinz, LX; Superti-Furga, G. Functional crosstalk between membrane lipids and TLR biology. Curr Opin Cell Biol 2016, 39, 28–36. [Google Scholar] [CrossRef] [PubMed]
- Kornilov, FD; Shabalkina, AV; Lin, C; Volynsky, PE; Kot, EF; Kayushin, AL; et al. The architecture of transmembrane and cytoplasmic juxtamembrane regions of Toll-like receptors. Nat Commun. 2023, 14(1), 1503. [Google Scholar] [CrossRef]
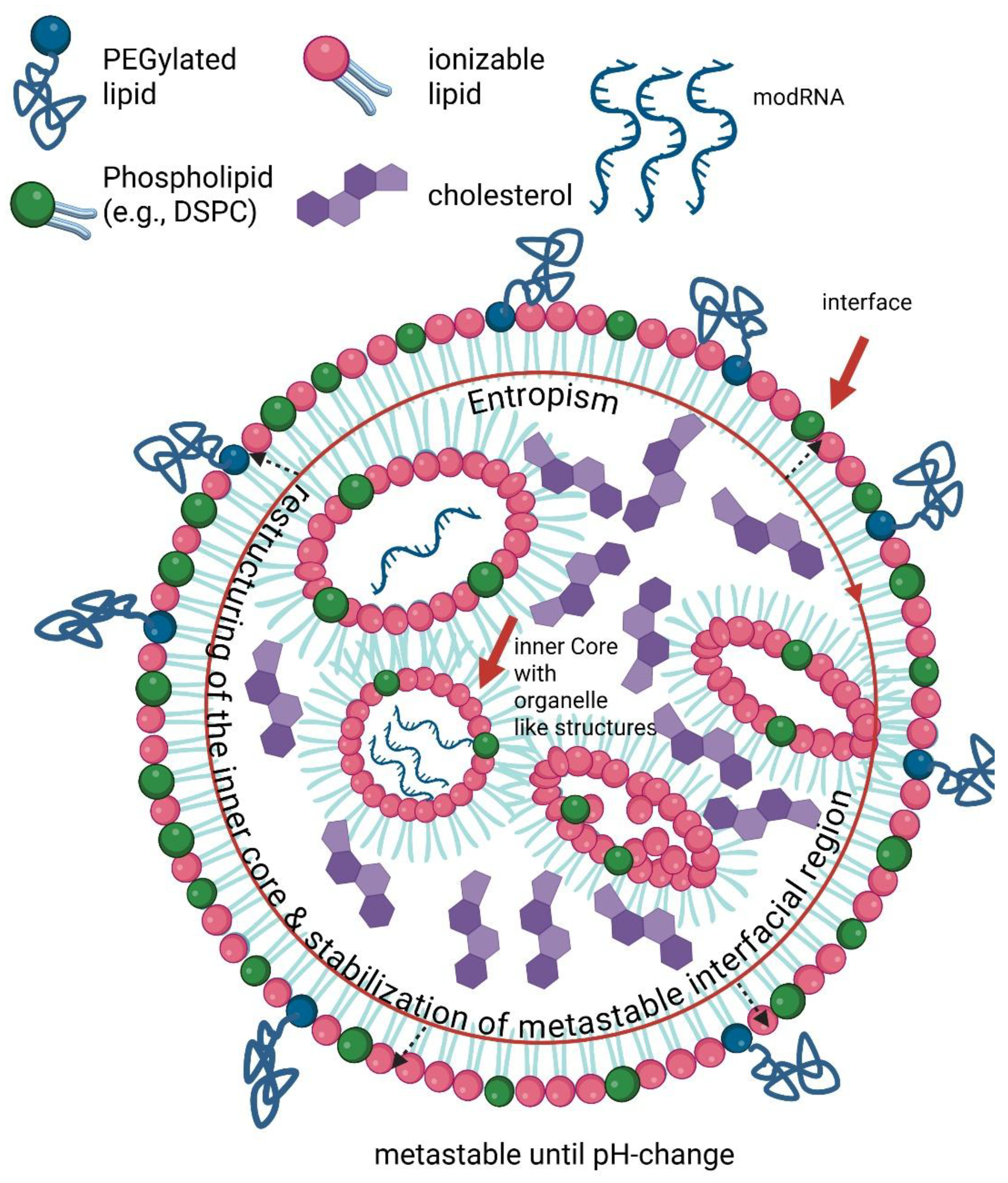
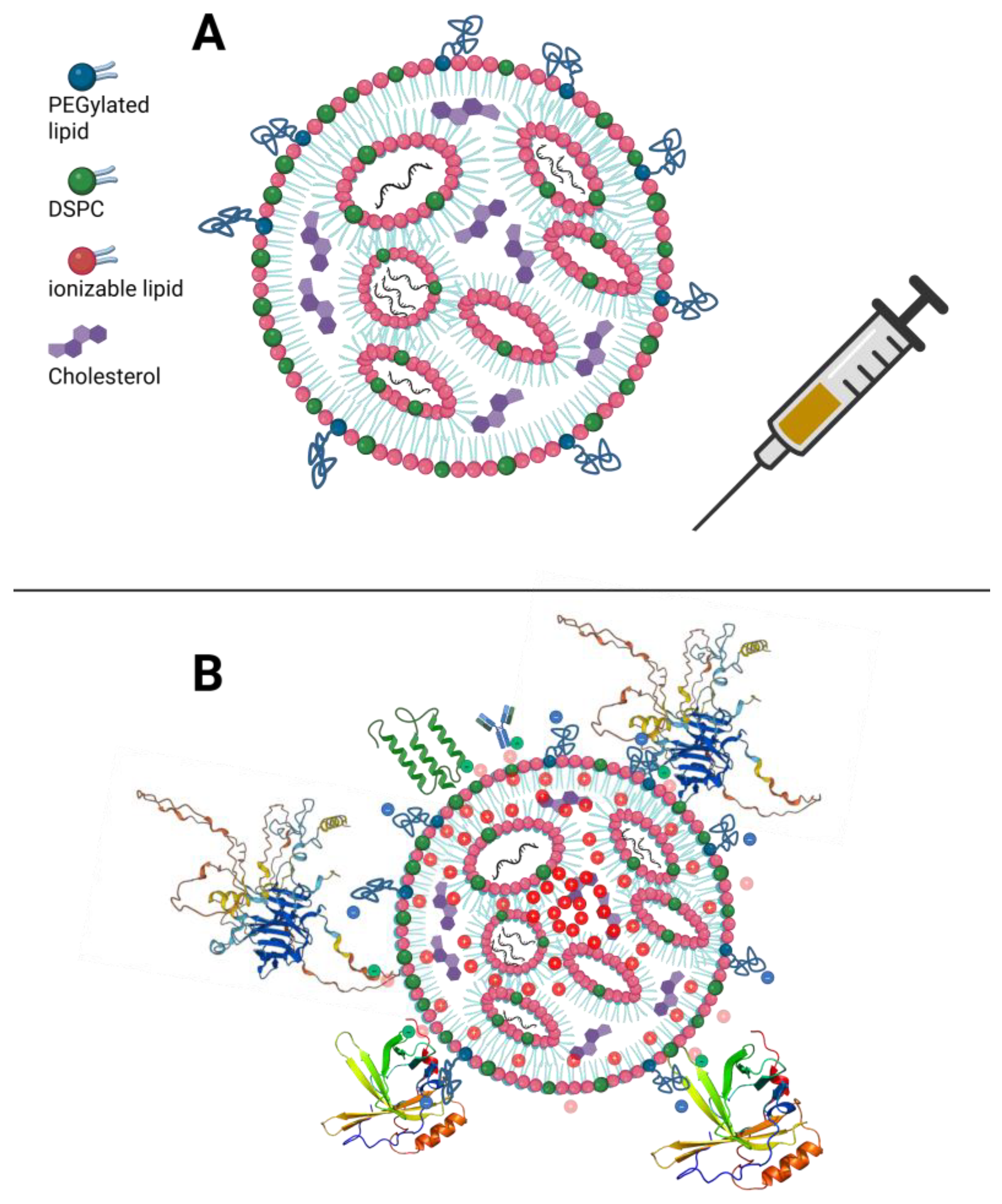
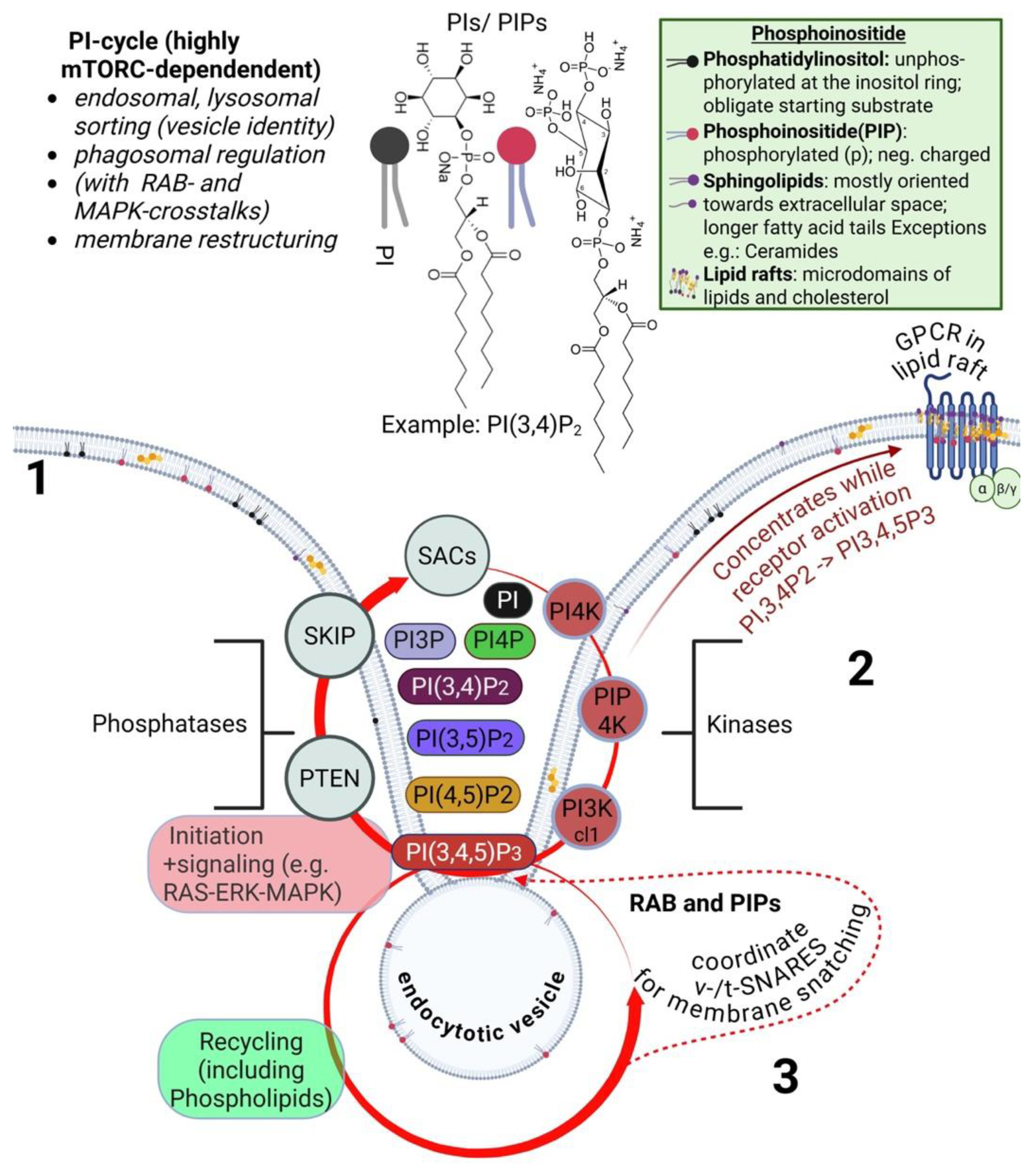
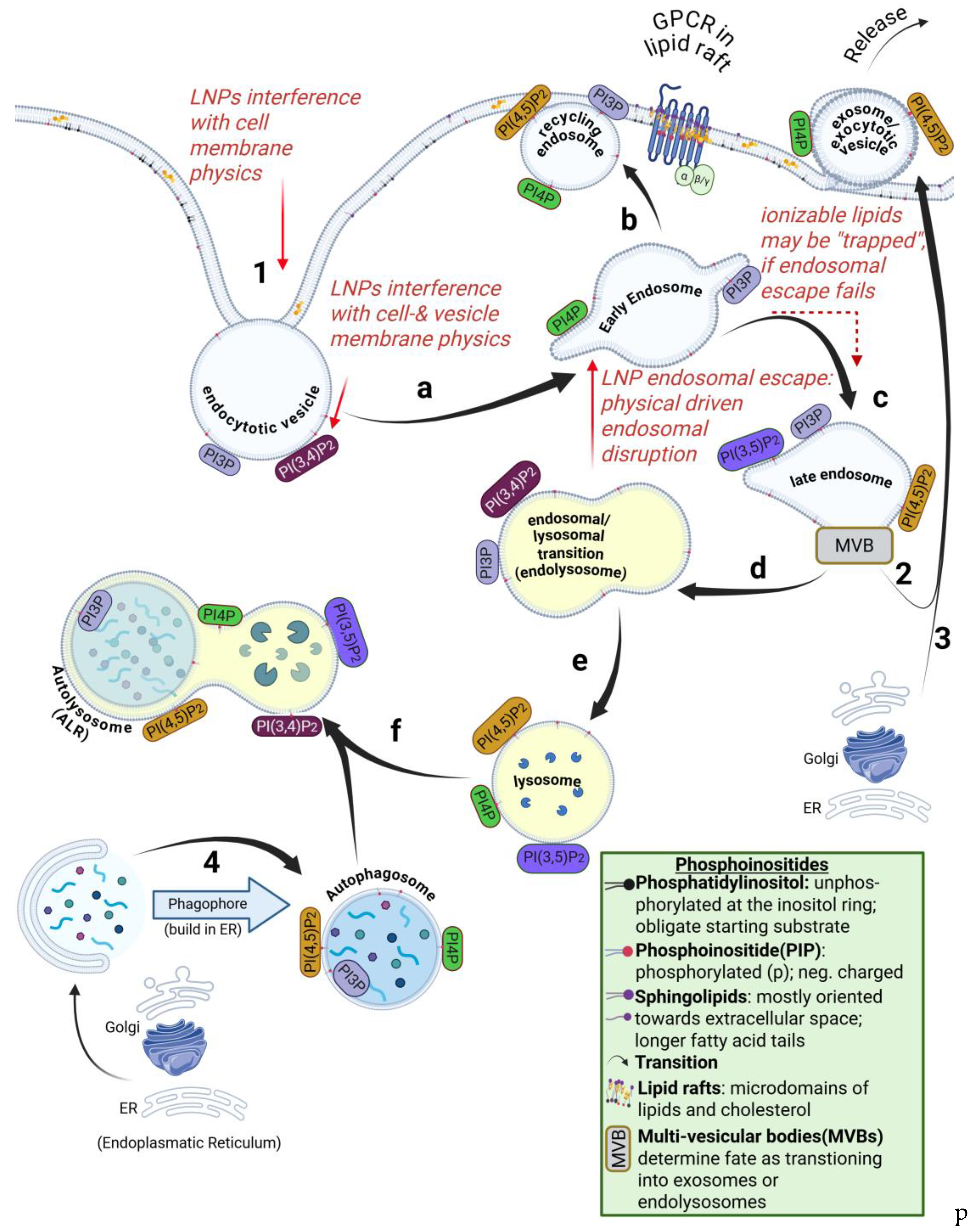
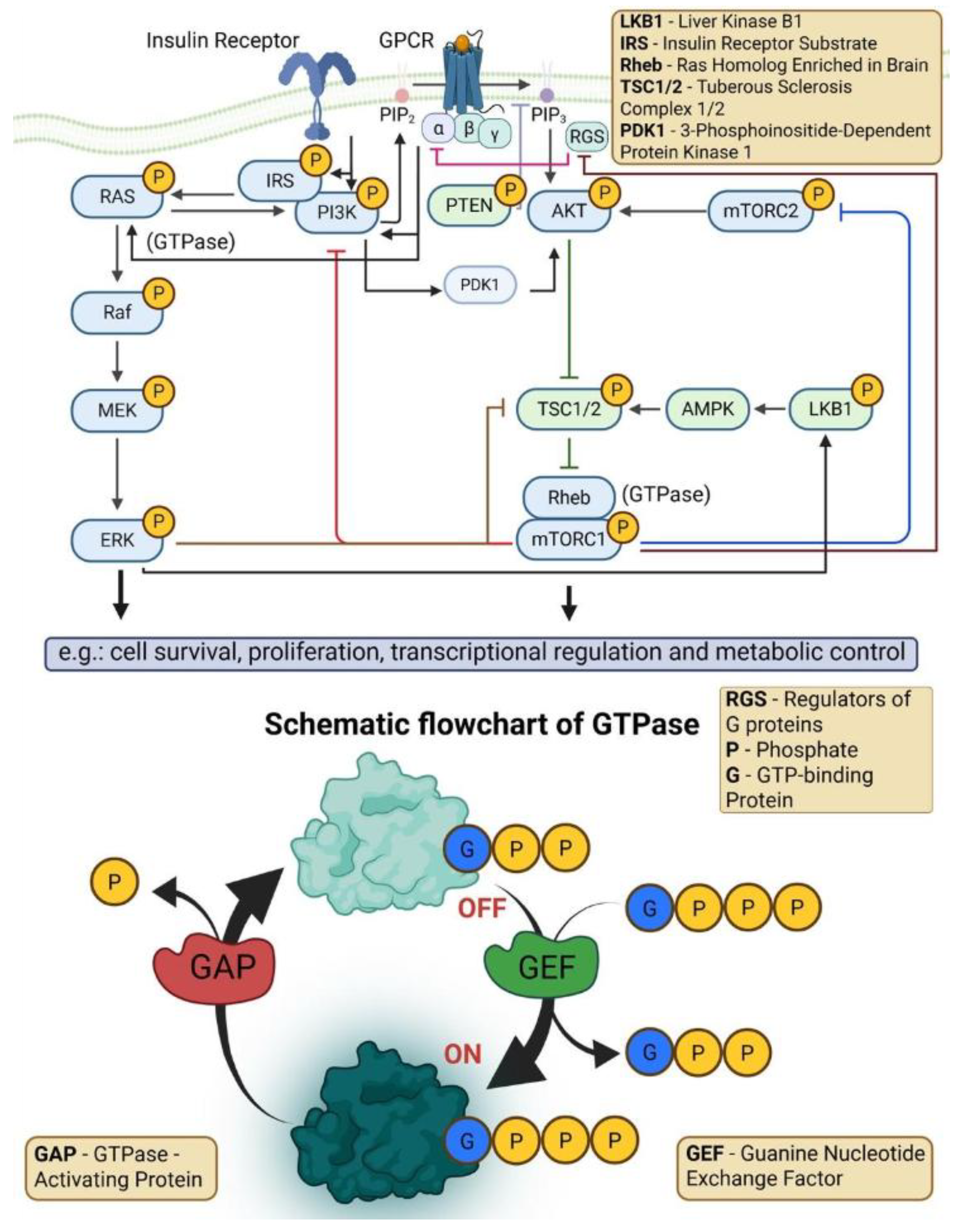
| Platform (example) | Route/ Vehicle |
Primary Delivery Mechanism | Approximate Endosomal Escapea | Membrane Activityb | Potential L-DMD Riskc |
Common safety signals e. |
Refs |
|---|---|---|---|---|---|---|---|
| > GalNAc–siRNA (inclisiran) | SC; Ligand–siRNA conjugate; No LNP |
Hepatocyte-targeted receptor uptake (ASGPR) | Sufficient at low doses (no LNP) |
Minimal: negligible disruptive capacity | √ (no ionizable lipids) |
Primarily injection site only | 111, 112 |
| > siRNA–LNP (patisiran) | IV; Ionizable lipid; (DLin-MC3-DMA) |
Hepatic fenestration + ApoE mediated LDL-R | ~1% (model-dependent) | Modest; Transient endosomal lesions repaired by ESCRT |
√√ (hepatotropic, low pKa 6.4) |
↑Transaminases; ↓ vitamin A levels; infusion reactions (CARPA) |
107, 113 |
| >
modRNA–LNP (SARS-CoV-2 vaccines) |
IM; Ionizable lipid; (ALC-0315/ SM-102) |
Immune activation + lymphatic drainage + systemic distribution | ≤~15% (context-dependent) | High; sustained endosomal/ lysosomal engagement |
√√√ (systemic distribution, pKa 6.1-6.7) |
Systemic reactogenicity (fever myalgias); Rare myocarditis; Rare anaphylaxis |
53, 116, 117 |
| >
saRNA–LNP (zapomeran/ SARS-CoV-2 vaccine) |
IM; Ionizable lipid; (ATX-126/LUNAR lipids) |
Prolonged RNA replication + front-loaded lipid-mediated delivery | Variable; amplification design | High; ionizable lipid with saRNA prolonging innate activation |
√√√ (very slow clearance, pKa undefined) |
Systemic reactogenicity ≈modRNA-LNPs; |
19, 115 |
| Section | Pathway | Membrane Connection | Representative Evidence | Reference |
|---|---|---|---|---|
| Membrane Structure and Maintenance | ||||
|
4.1 |
ESCRT/ endocytosis |
Membrane repair following endosomal stress. Recruitment depends on phosphoinositides. |
H-Px: ↓ESCRT proteins and ↓endocytic activity after modRNA-LNP injection. | Hickey et al.139 |
| 4.2 | CYP | ER/mitochondrial embedded membranas. Catalytic geometry depends on local phospholipids and charge. |
↕ M-Tx (eLNP), H-Px, H-Obs: ↓CYP metabolism post IM injection in mice; ↑Phase I enzymes (♂ only, BNT162b2 specific, hepatocellular leakage†); clozapine toxicity post-vaccination (functional corroboration‡). | Ndeupen et al. 119 Hickey et al. 139 Thompson et al. 159 |
| Innate Immune Signaling | ||||
| 4.3 | TLR4 axis (MyD88/ TRIF bias) |
Plasma lipid membrane organization. Intact phosphoinositides needed for balanced signaling. |
M-Tx (eLNP), M-Fx: MyD88-dominant NFκB activation, reversible with TLR-4 blockade (TAK-242) in human THP-1 cells in vitro NFκB.-κB | Ndeupen et al. 119 Korzun et al. 134 *Zelkoski et al.132 |
| 4.4 | Pro-inflammatory cytokines | Membrane-associated PRR signaling (NF-κB/MAPK). Functional organization of opes within inner-leaflet lipid raft. |
↕ (M-Tx, M-Tx (eLNP)), H-Obs: ↑NFκB targets and canonical cytokines in murine eLNP; Concordant upregulation in human datasets; cytokine-mediated drug interactions documented NFκB. -κB |
Ndeupen et al. 119 Korzun et al. 134 McColl et al. 242 |
| 4.4 | Β-Cytokines, CXC- chemokines, IL6, IFNs |
Membrane proximal NF-κB activation. | ↕ (MTx, MFx (eLNP)), H-Tx: Concordant CCL family upregulation in murine eLNP and human buffy-coat transcriptomes (CXCL 9,10).-Tx | Ndeupen et al. 119 Korzun et al. 134 Knabl et al. 138 |
| 4.5 | Complement (C3, lectin) |
Membrane-bound GPCR receptors (C3aR). Protein corona effects. |
M-Tx (eLNP): ↑C3; M-Px: LNP-vitronectin/ficolin-1 binding facilitates C3 activation events |
Korzun et al. 134 Luo et al. 137 |
| Growth and Metabolic Signaling | ||||
| 4.7 | RAS-RAF-MEK-ERK (MAPK) | RAS GTPases lipid anchors at plasma membrane nanodomains. Translate local charge shift to signal amplification. |
↕ M-Tx (eLNP), H-Tx, H-Px: ERK1/KRAS pathway upregulation across species omes. | Korzun et al.134 Knabl et al. 138 Hickey et al. 139 |
| 4.7 | PI3K-AKT-mTOR | Membrane-localized PI3K. Partial mTORC1 engagement indicates upstream dysregulated lipid signaling via RAS. |
↕ M-Tx (eLNP), H-Tx, H-Px: Partial ERK1/KRAS pathway + PI3K modulation post LNP exposure. |
Korzun et al.134 Knabl et al. 138 Hickey et al. 139 |
|
4.4 |
PI3K Class II (PIK3C2G) | Often activated downstream of RTKs + GPCRs. Act independently of regulatory subunits characteristic of PI3Ks. |
M-Tx (eLNP): Consistent ↓PIK3C2G across datasets. | Korzun et al. 134 |
| 4.6 | PPARγ | Integrates lipid-derived signals. Suppression favors NF-κB activation. |
↕ M-Tx (eLNP), H-Obs: ↓PPARγ pathway → loss of anti-inflammatory tone + lipid metabolism imbalance; post-vaccination hypercholesterolemia (OR = 1.54, 95% CI: 1.36–1.74; population level corroboration)§. | Ndeupen et al. 119 Huang et al. 171 |
| 4.6 | AMPK | Senses membrane-linked stress. Suppression favors NF-κB activation. |
M-Tx (eLNP): ↓AMPK transcripts → reduced oxidative metabolism and ↑NFκB. | Ndeupen et al. 119 |
| Cell Cycle and Stress Response | ||||
| 4.8 | p53 + E2F Network | Downstream of RAS-MAPK and PI3K-mTOR signaling. Cell-cycle checkpoint. |
H-Tx: p53 + E2F8; partial MTORC1 activation. | Knabl et al. 138 |
| Study and Reference | Model / Species | Material Tested | Route / Dose | Data Type / Readout | Timeline / Endpoints |
|---|---|---|---|---|---|
| Ndeupen et al. [119] | Mouse (C57BL/6, WT). |
eLNP (no mRNA). |
IM 10 µg in PBS |
M-Tx; (Whole tissue RNAseq + GSEA)-seq + GSEA |
~24 h post injection. |
| Korzun et al. [134] | Mouse (WT + KO lines) |
Luc mRNA LNP + eLNP. | Luc mRNA + eLNP groups received 5ug in 100μl PBS; IP single dose (acute); or IP once every 24h for 3 doses (chronic). = 4 groups total eLNP groups received TAK-242 inhibitor 2 hours prior to each dose |
M-Tx, M-FX; (behavior + ELISA). |
6h post last dose |
| Luo et al. [137] | Mouse / Cell models | eLNP (no payload) + comparators. |
Multiple routes (IN, IM, IV, PO, ID); 0.0005–0.5 mg/kg. |
M-Px; (quantitative proteomics). |
≤ 24h post dose-dose |
| Hickey et al. [139] | Human (adults post BNT162b2 / mRNA1273 vaccination)**- | ModRNA-LNP vaccination. | IM; clinical dose. |
H-Px ; (SomaScan v4.1 proteomics). |
1 month and 6 months post-dose dose 3. |
| Knabl et al. [138] | Human (patients and healthy volunteers)** Elderly patients (3 doses; postvaccination or infection, treatment with dexamethasone) Younger healthy group (naive, 1st dose) |
ModRNA-LNP vaccination/ infection cohort. |
IM; clinical dose. |
H-Tx; (whole blood buffy coat transcriptomics) |
Elderly Days 7–60 Younger Days 7–10 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




