Submitted:
21 October 2025
Posted:
21 October 2025
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. Mechanistic Synthesis
2.1. Nutritional Signaling and the AMPK–mTOR Axis
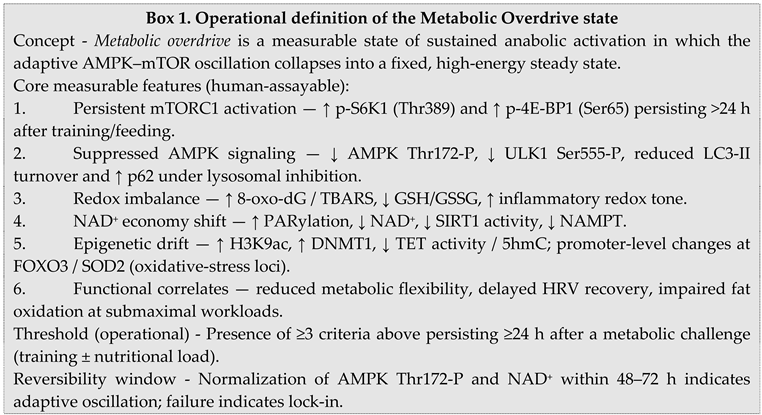

2.2. Molecular Parallels between Performance Enhancement and Tumorigenesis
2.3. Epigenetic Modulation under Metabolic Stress
2.4. Case Contexts: Elite Sport and Doping Paradigms
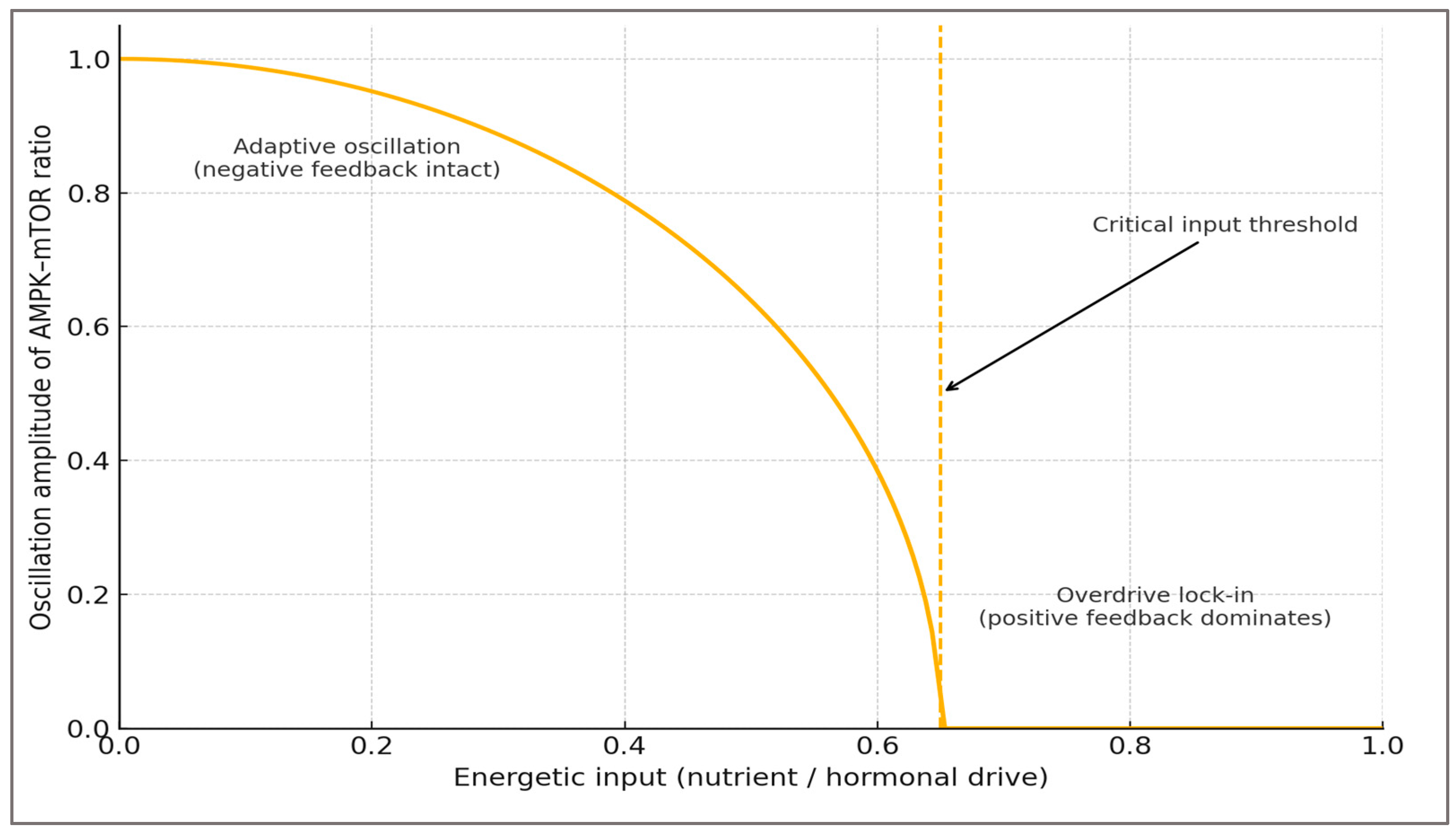
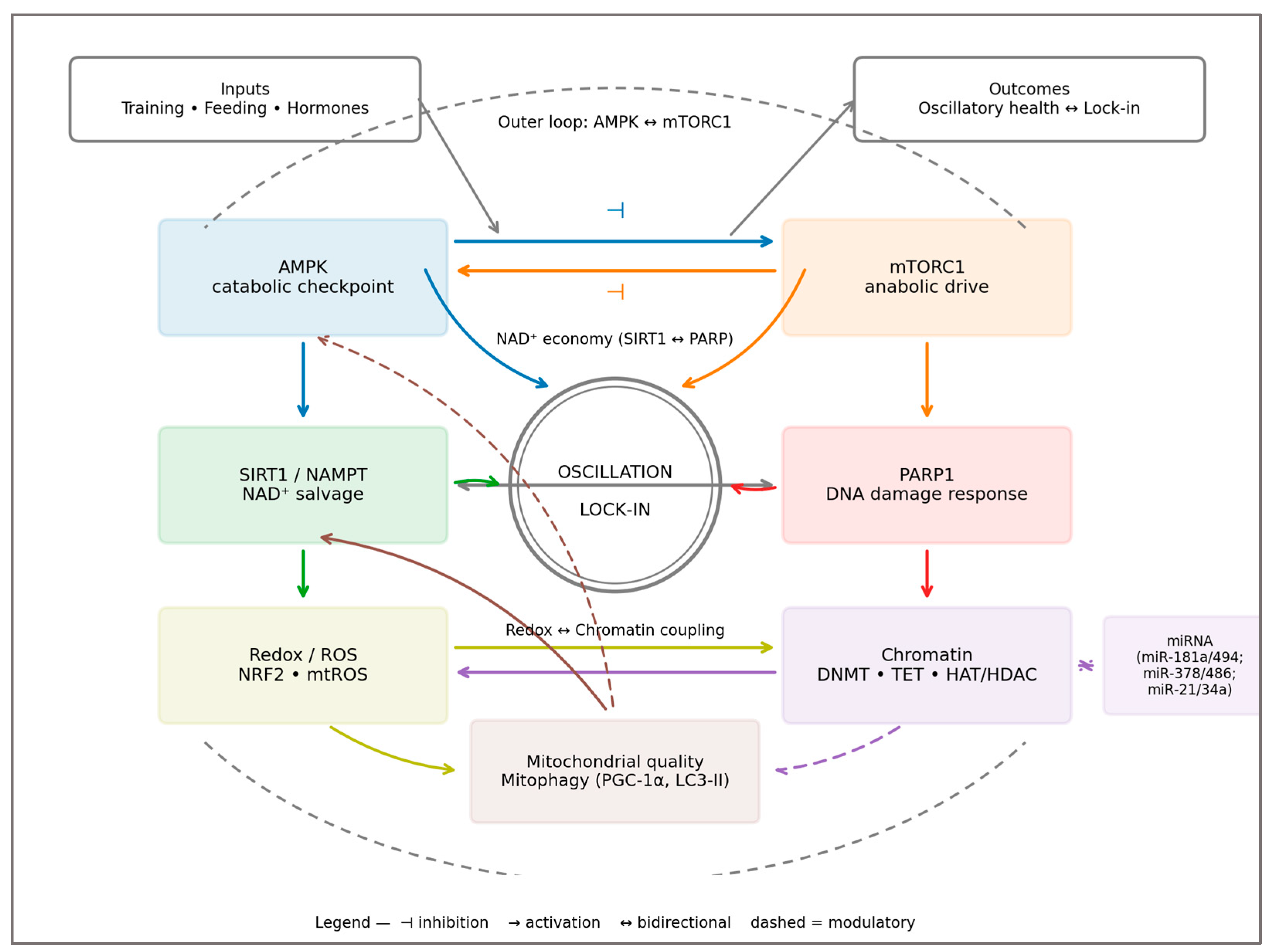
2.5. The Metabolic Overdrive Model
2.6. Testable Predictions and Biomarkers
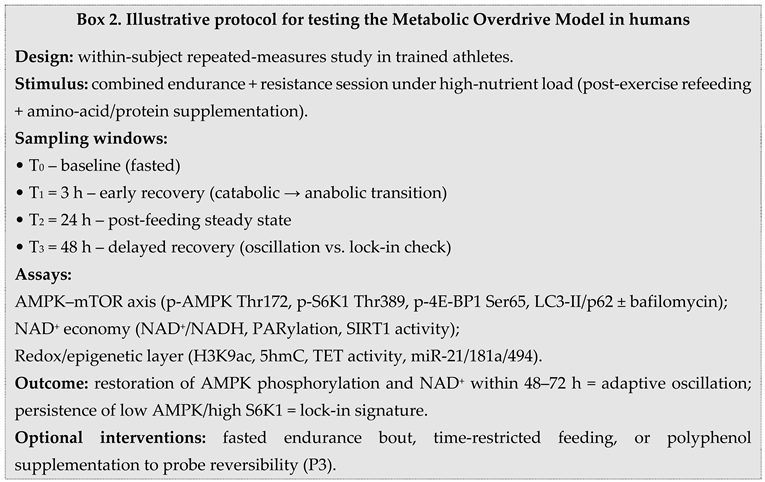

3. Discussion
4. Materials and Methods - Mechanistic Evidence Mapping and Causal-Loop Synthesis (Non-Systematic)
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of interest
Abbreviations
References
- González, A.; Hall, M.N.; Lin, S.C.; Hardie, D.G. AMPK and TOR: The Yin and Yang of Cellular Nutrient Sensing and Growth Control. Cell Metab. 2020, 31, 472–492. [Google Scholar] [CrossRef]
- Perry, C.G.R.; Hawley, J.A. Molecular Basis of Exercise-Induced Skeletal Muscle Mitochondrial Biogenesis: Historical Advances, Current Knowledge, and Future Challenges. Cold Spring Harb. Perspect. Med. 2018, 8, a029686. [Google Scholar] [CrossRef] [PubMed]
- Saxton, R.A.; Sabatini, D.M. mTOR signaling in growth, metabolism, and disease. Cell 2017, 169, 361–371. [Google Scholar] [CrossRef] [PubMed]
- Esteves, J.V.; Stanford, K.I. Exercise as a tool to mitigate metabolic disease. Am. J. Physiol. Cell Physiol. 2024, 327, C587–C598. [Google Scholar] [CrossRef] [PubMed]
- Camera, D.M.; Smiles, W.J.; Hawley, J.A. Exercise-induced skeletal muscle signaling pathways and the regulation of metabolism. Biochem. J. 2016, 473, 2319–2354. [Google Scholar] [CrossRef]
- Mentch, S.J.; Locasale, J.W. One-carbon metabolism and epigenetics: Understanding the specificity of the methyl donor pathway. Nat. Rev. Mol. Cell Biol. 2016, 17, 561–571. [Google Scholar] [CrossRef]
- Seaborne, R.A.; Strauss, J.; Cocks, M.; Shepherd, S.O.; O’Brien, T.D.; van Someren, K.A.; Bell, P.G.; Murgatroyd, C.; Morton, J.P.; Stewart, C.E. Human Skeletal Muscle Possesses an Epigenetic Memory of Hypertrophy. Sci. Rep. 2018, 8, 1898. [Google Scholar] [CrossRef]
- Barres, R.; Zierath, J.R. The role of diet and exercise in the transgenerational epigenetic landscape of T2DM. Nat. Rev. Endocrinol. 2016, 12, 441–451. [Google Scholar] [CrossRef]
- Mihaylova, M.M.; Shaw, R.J. The AMPK signaling pathway coordinates cell growth, autophagy, and metabolism. Nat. Cell Biol. 2011, 13, 1016–1023. [Google Scholar] [CrossRef]
- Sharples, A.P. Skeletal Muscle Possesses an Epigenetic Memory of Exercise: Role of Nucleus Type-Specific DNA Methylation. Function (Oxf.) 2021, 2, zqab047. [Google Scholar] [CrossRef]
- Clemente-Suárez, V.J.; Martín-Rodríguez, A.; Redondo-Flórez, L.; Ruisoto, P.; Navarro-Jiménez, E.; Ramos-Campo, D.J.; Tornero-Aguilera, J.F. Metabolic Health, Mitochondrial Fitness, Physical Activity, and Cancer. Cancers 2023, 15, 814. [Google Scholar] [CrossRef]
- Morton, R.W.; McGlory, C.; Phillips, S.M. Nutritional interventions to augment resistance training-induced skeletal muscle hypertrophy. Front. Physiol. 2015, 6, 245. [Google Scholar] [CrossRef] [PubMed]
- Powers, S.K.; Deminice, R.; Ozdemir, M.; Yoshihara, T.; Bomkamp, M.P.; Hyatt, H. Exercise-induced oxidative stress: Friend or foe? J. Sport Health Sci. 2020, 9, 415–425. [Google Scholar] [CrossRef] [PubMed]
- Herzig, S.; Shaw, R.J. AMPK: Guardian of Metabolism and Mitochondrial Homeostasis. Nat. Rev. Mol. Cell Biol. 2018, 19, 121–135. [Google Scholar] [CrossRef] [PubMed]
- Karasawa, T.; Choi, R.H.; Meza, C.A.; Maschek, J.A.; Cox, J.E.; Funai, K. Skeletal muscle PGC-1α remodels mitochondrial phospholipidome but does not alter energy efficiency for ATP synthesis. bioRxiv 2024, 2024.05.22.595374. [CrossRef]
- Goul, C.; Peruzzo, R.; Zoncu, R. The molecular basis of nutrient sensing and signalling by mTORC1 in metabolism regulation and disease. Nat. Rev. Mol. Cell Biol. 2023, 24, 857–875. [Google Scholar] [CrossRef]
- Martinez-Lopez, N.; Mattar, P.; Toledo, M.; Aoun, M.L.; Sharma, M.; McIntire, L.B.; Gunther-Cummins, L.; Macaluso, F.P.; Aguilan, J.T.; Sidoli, S.; et al. mTORC2–NDRG1–CDC42 axis couples fasting to mitochondrial fission. Nat. Cell Biol. 2023, 25, 989–1003. [Google Scholar] [CrossRef]
- Knudsen, J.R.; Blaschke, S.; et al. Exercise increases phosphorylation of the putative mTORC2 activity marker NDRG1 Thr346 in human skeletal muscle. Am. J. Physiol. Endocrinol. Metab. 2022, 322, E465–E475. [Google Scholar] [CrossRef]
- Li, J.; Wang, Z.; Li, C.; Song, Y.; Wang, Y.; Bo, H.; Zhang, Y. Impact of Exercise and Aging on Mitochondrial Homeostasis in Skeletal Muscle: Roles of ROS and Epigenetics. Cells 2022, 11, 2086. [Google Scholar] [CrossRef]
- Jaguri, A.; Nahar, P.; Srinivasan, S.; et al. Exercise Metabolome: Insights for Health and Performance. Metabolites 2023, 13, 694. [Google Scholar] [CrossRef]
- Schwalm, C.; Jamart, C.; Benoit, N.; Naslain, D.; Premont, C.; Prevet, J.; Van Thienen, R.; Deldicque, L.; Francaux, M. Activation of Autophagy in Human Skeletal Muscle Is Dependent on Exercise Intensity and AMPK Activation. FASEB J. 2015, 29, 3515–3526. [Google Scholar] [CrossRef]
- Mouchiroud, L.; et al. The NAD+/Sirtuin pathway modulates longevity through mitochondrial UPR. Cell 2013, 154, 430–441. [Google Scholar] [CrossRef]
- Powers, S.K.; Morton, A.B.; Ahn, B.; Smuder, A.J. Redox control of skeletal muscle atrophy. Free Radic. Biol. Med. 2016, 98, 208–217. [Google Scholar] [CrossRef]
- Fanti, M.; Longo, V.D. Nutrition, GH/IGF-1 Signaling, and Cancer. Endocr. Relat. Cancer 2024, 31, e230048. [Google Scholar] [CrossRef] [PubMed]
- Jewell, J.L.; Kim, Y.C.; Russell, R.C.; Yu, F.X.; Park, H.W.; Plouffe, S.W.; Tagliabracci, V.S.; Guan, K.L. Metabolism. Differential regulation of mTORC1 by leucine and glutamine. Science 2015, 347, 194–198. [Google Scholar] [CrossRef] [PubMed]
- Zubrzycki, I.Z.; Ossowski, Z.; Przybylski, S.; Wiacek, M.; Clarke, A.; Trąbka, B. Supplementation with Silk Amino Acids Improves Physiological Parameters Defining Stamina in Elite Fin-Swimmers. J. Int. Soc. Sports Nutr. 2014, 11, 57. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Courneya, K.S.; Friedenreich, C.M. The Physical Activity and Cancer Control (PACC) framework: Update on the evidence, guidelines, and future research priorities. Br. J. Cancer 2024, 131, 957–969. [Google Scholar] [CrossRef]
- McDermott, K.T.; Noake, C.; Wolff, R.; Bauld, L.; Espina, C.; Foucaud, J.; Steindorf, K.; Thorat, M.A.; Weijenberg, M.P.; Schüz, J.; Kleijnen, J. Digital interventions to moderate physical inactivity and/or nutrition in young people: A Cancer Prevention Europe overview of systematic reviews. Front. Digit. Health 2023, 5, 1185586. [Google Scholar] [CrossRef]
- Cantó, C.; Menzies, K.J.; Auwerx, J. NAD(+) Metabolism and the Control of Energy Homeostasis: A Balancing Act between Mitochondria and the Nucleus. Cell Metab. 2015, 22, 31–53. [Google Scholar] [CrossRef]
- Longo, V.D.; Panda, S. Fasting, circadian rhythms, and time-restricted feeding in healthy lifespan. Cell Metab. 2016, 23, 1048–1059. [Google Scholar] [CrossRef]
- Hargreaves, M.; Spriet, L.L. Skeletal Muscle Energy Metabolism during Exercise. Nat. Metab. 2020, 2, 817–828. [Google Scholar] [CrossRef]
- Sirago, G.; Maggi, L.; et al. Mammalian Target of Rapamycin (mTOR) Signaling at the Crossroad of Multicellular Muscle Adaptation. Int. J. Mol. Sci. 2022, 23, 13823. [Google Scholar] [CrossRef]
- Zhou, M.; Wei, Y.; Feng, Y.; Zhang, S.; Ma, N.; Wang, K.; Tan, P.; Zhao, Y.; Zhao, J.; Ma, X. Arginine Regulates Skeletal Muscle Fiber Type Formation via mTOR Signaling Pathway. Int. J. Mol. Sci. 2024, 25, 6184. [Google Scholar] [CrossRef]
- Fu, W.; Wu, G. Targeting mTOR for Anti-Aging and Anti-Cancer Therapy. Molecules 2023, 28, 3157. [Google Scholar] [CrossRef] [PubMed]
- Fingar, D.C.; Blenis, J. Target of rapamycin (TOR): An integrator of nutrient and growth factor signals and coordinator of cell growth and cell cycle progression. Oncogene 2024, 23, 3151–3171. [Google Scholar] [CrossRef] [PubMed]
- Mănescu, D.C. Computational Analysis of Neuromuscular Adaptations to Strength and Plyometric Training: An Integrated Modeling Study. Sports 2025, 13, 298. [Google Scholar] [CrossRef] [PubMed]
- Clemmons, D.R. Metabolic actions of insulin-like growth factor-I in normal physiology and diabetes. Endocrinol. Metab. Clin. North Am. 2012, 41, 425–443. [Google Scholar] [CrossRef]
- Galifi, C.A.; Colalillo, S.; Bertuzzi, A.; Bianchini, M.; Silletti, S. Insulin-Like Growth Factor-1 Receptor Crosstalk with Integrins in Cancer Progression. Endocr. Relat. Cancer 2023, 30, ERC–23. [Google Scholar] [CrossRef]
- Tran, D.; Bergholz, J.; Zhang, H.; He, H.; Wang, Y.; Zhang, Y.; Li, Q.; Kirkland, J.L.; Xiao, Z.; Wang, C.; et al. Insulin-like growth factor-1 regulates the SIRT1–p53 pathway in cellular senescence. Aging Cell 2014, 13, 669–678. [Google Scholar] [CrossRef]
- Guan, Y.; Yan, Z. Molecular Mechanisms of Exercise and Healthspan. Cells 2022, 11, 872. [Google Scholar] [CrossRef]
- Jacko, D.; Schaaf, K.; Masur, L.; Windoffer, H.; Aussieker, T.; Schiffer, T.; Zacher, J.; Bloch, W.; Gehlert, S. Repeated and Interrupted Resistance Exercise Induces the Desensitization and Re-Sensitization of mTOR-Related Signaling in Human Skeletal Muscle Fibers. Int. J. Mol. Sci. 2022, 23, 5431. [Google Scholar] [CrossRef]
- Maciejczyk, M.; Palka, T.; Wiecek, M.; Masel, S.; Szygula, Z. The Effects of Intermittent Hypoxic Training on Anaerobic Performance in Young Men. Applied Sciences 2024, 14, 676. [Google Scholar] [CrossRef]
- Kang, J.S.; Lee, C.; Kim, S.; et al. Baf155 Regulates Skeletal Muscle Metabolism via HIF-1α Signalling. PLoS Biol. 2023, 21, e3002192. [Google Scholar] [CrossRef] [PubMed]
- Yin, W.; Noguchi, C.T. The Role of Erythropoietin in Metabolic Regulation. Cells 2025, 14, 280. [Google Scholar] [CrossRef] [PubMed]
- Man, M.C.; Ganera, C.; Bărbuleţ, G.D.; Krzysztofik, M.; Panaet, A.E.; Cucui, A.I.; Tohănean, D.I.; Alexe, D.I. The Modifications of Haemoglobin, Erythropoietin Values and Running Performance While Training at Mountain vs. Hilltop vs. Seaside. Int. J. Environ. Res. Public Health 2021, 18, 9486. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Chen, C.; Teo, E.-C.; Zhang, Y.; Huang, J.; Xu, Y.; Gu, Y. Intracellular Oxidative Stress Induced by Physical Exercise in Adults: Systematic Review and Meta-Analysis. Antioxidants 2022, 11, 1751. [Google Scholar] [CrossRef]
- Meng, Q.; Su, C-H. The Impact of Physical Exercise on Oxidative and Nitrosative Stress: Balancing the Benefits and Risks. Antioxidants 2024, 13, 573. [Google Scholar] [CrossRef]
- Sabharwal, S.S.; Schumacker, P.T. Mitochondrial ROS in cancer: Initiators, amplifiers or an Achilles' heel? Nat. Rev. Cancer 2014, 14, 709–721. [Google Scholar] [CrossRef]
- Sullivan, L.B.; Chandel, N.S. Mitochondrial reactive oxygen species and cancer. Cancer Metab. 2014, 2, 17. [Google Scholar] [CrossRef]
- Barba, I.; Carrillo-Bosch, L.; Seoane, J. Targeting the Warburg Effect in Cancer: Where Do We Stand? Int. J. Mol. Sci. 2024, 25, 3142. [Google Scholar] [CrossRef]
- Maddocks, O.D.K.; Labuschagne, C.F.; Adams, P.D.; Vousden, K.H. Serine metabolism supports the methionine cycle and DNA/RNA methylation through de novo ATP synthesis in cancer cells. Mol. Cell 2016, 61, 210–221. [Google Scholar] [CrossRef]
- Chen, C.; Chauhan, A.; Dai, Y. Connections between metabolism and epigenetics: From basic mechanisms to clinical implications. Frontiers in Pharmacology 2022, 13, 935536. [Google Scholar] [CrossRef] [PubMed]
- Pouikli, A.; Tessarz, P. Metabolism and chromatin: A dynamic duo that regulates development and ageing. BioEssays 2021, 43, e2000273. [Google Scholar] [CrossRef] [PubMed]
- Xie, N.; Zhang, L.; Wang, J.; Zhao, Y.; Dong, Y. NAD+ metabolism: Pathophysiologic mechanisms and therapeutic potential. Signal Transduction and Targeted Therapy 2020, 5, 227. [Google Scholar] [CrossRef] [PubMed]
- Mentch, S.J.; Locasale, J.W. One-carbon metabolism and epigenetics: Understanding the specificity of the methyl donor pathway. Nat. Rev. Mol. Cell Biol. 2016, 17, 561–571. [Google Scholar] [CrossRef]
- Janke, R.; Dodson, A.E.; Rine, J. Metabolism and epigenetics. Annu. Rev. Cell Dev. Biol. 2015, 31, 473–496. [Google Scholar] [CrossRef]
- Barrès, R.; Yan, J.; Egan, B.; Treebak, J.T.; Rasmussen, M.; Fritz, T.; Caidahl, K.; Krook, A.; O’Gorman, D.J.; Zierath, J.R. Acute exercise remodels promoter methylation in human skeletal muscle. Cell Metab. 2012, 15, 405–411. [Google Scholar] [CrossRef]
- Steinberg, G.R.; Kemp, B.E. AMPK in health and disease. Physiol. Rev. 2019, 89, 1025–1078. [Google Scholar] [CrossRef]
- Seaborne, R.A.; Strauss, J.; Cocks, M.; Shepherd, S.O.; O’Brien, T.D.; van Someren, K.A.; Bell, P.G.; Murgatroyd, C.; Morton, J.P.; Stewart, C.E. Human skeletal muscle possesses an epigenetic memory of hypertrophy. Sci. Rep. 2018, 8, 1898. [Google Scholar] [CrossRef]
- Barrès, R.; Zierath, J.R. The role of diet and exercise in the transgenerational epigenetic landscape of T2DM. Nat. Rev. Endocrinol. 2016, 12, 441–451. [Google Scholar] [CrossRef]
- Kim, M.; Costello, J. DNA methylation: An epigenetic mark of cellular memory. Exp. Mol. Med. 2017, 49, e322. [Google Scholar] [CrossRef]
- Baylin, S.B.; Jones, P.A. A decade of exploring the cancer epigenome—Biological and translational implications. Nat. Rev. Cancer 2011, 11, 726–734. [Google Scholar] [CrossRef]
- Shvedunova, M.; Akhtar, A. Modulation of cellular processes by histone and non-histone lysine acetylation: Mechanisms, targets, and physiological significance. Nat. Rev. Mol. Cell Biol. 2022, 23, 304–326. [Google Scholar] [CrossRef] [PubMed]
- Cantó, C.; Sauve, A.A.; Bai, P. Crosstalk between poly(ADP-ribose) polymerase and sirtuin enzymes. Mol. Aspects Med. 2013, 34, 1168–1201. [Google Scholar] [CrossRef] [PubMed]
- Afzal, M.; Greco, F.; Quinzi, F.; Scionti, F.; Maurotti, S.; Montalcini, T.; Mancini, A.; Buono, P.; Emerenziani, G.P. The Effect of Physical Activity/Exercise on miRNA Expression and Function in Non-Communicable Diseases—A Systematic Review. Int. J. Mol. Sci. 2024, 25, 6813. [Google Scholar] [CrossRef] [PubMed]
- Grieb, A.; Schmitt, A.; Fragasso, A.; Widmann, M.; Mattioni Maturana, F.; Burgstahler, C.; Erz, G.; Schellhorn, P.; Nieß, A.M.; Munz, B. Skeletal Muscle MicroRNA Patterns in Response to a Single Bout of Exercise in Females: Biomarkers for Subsequent Training Adaptation? Biomolecules 2023, 13, 884. [Google Scholar] [CrossRef]
- Quintanilha, B.J.; Reis, B.Z.; Duarte, G.B.S.; Cozzolino, S.M.F.; Rogero, M.M. Nutrimiromics: Role of microRNAs and nutrition in modulating inflammation and chronic diseases. Nutrients 2017, 9, 1168. [Google Scholar] [CrossRef]
- Radom-Aizik, S.; et al. Effects of exercise training on microRNA expression in young and older men. Cell Physiol. Biochem. 2022, 29, 831–842. [Google Scholar] [CrossRef]
- Fu, J.; Zhang, X.; Li, H.; Liu, Y.; Wang, X. MicroRNA-34 Family in Cancers: Role, Mechanism, and Therapeutic Potential. Cancers 2023, 15, 4723. [Google Scholar] [CrossRef]
- Xiao, M.; Yang, H.; Xu, W.; Ma, S.; Lin, H.; Zhu, H.; Liu, L.; Liu, Y.; Yang, C.; Xu, Y.; et al. Inhibition of α-KG–dependent histone and DNA demethylases by fumarate and succinate that are accumulated in mutations of FH and SDH tumor suppressors. Genes Dev. 2012, 26, 1326–1338. [Google Scholar] [CrossRef]
- Walzik, D.; Jonas, W.; Joisten, N.; Belen, S.; Wüst, R.C.I.; Guillemin, G.; Zimmer, P. Tissue-Specific Effects of Exercise as NAD+-Boosting Strategy: Current Knowledge and Future Perspectives. Acta Physiol. 2023, 237, e13921. [Google Scholar] [CrossRef]
- Horvath, S.; Raj, K. DNA methylation-based biomarkers and the epigenetic clock theory of ageing. Nat. Rev. Genet. 2018, 19, 371–384. [Google Scholar] [CrossRef]
- Mănescu, D.C.; Mănescu, A.M. Artificial Intelligence in the Selection of Top-Performing Athletes for Team Sports: A Proof-of-Concept Predictive Modeling Study. Appl. Sci. 2025, 15, 9918. [Google Scholar] [CrossRef]
- Mănescu, D.C. Big Data Analytics Framework for Decision-Making in Sports Performance Optimization. Data 2025, 10, 116. [Google Scholar] [CrossRef]
- Meeusen, R.; Duclos, M.; Foster, C.; Fry, A.; Gleeson, M.; Nieman, D.; Raglin, J.; Rietjens, G.; Steinacker, J.; Urhausen, A. Prevention, diagnosis, and treatment of the overtraining syndrome. Eur. J. Sport Sci. 2013, 13, 1–24. [Google Scholar] [CrossRef]
- Fernández-Lázaro, D.; Mielgo-Ayuso, J.; Caballero-García, A.; Calleja-González, J.; Herrador-Pérez, M.; Armegod-Benítez, M.; Crespo-Otín, S.; Carratalá-Tejada, R. Adequacy of an Altitude Fitness Program (Living and Training at Moderate Altitude) Supplemented with Intermittent Hypoxic Training: Effects on Sports Performance, Blood Biomarkers and Safety Profiles in Elite Athletes. Int. J. Environ. Res. Public Health 2022, 19, 9095. [Google Scholar] [CrossRef] [PubMed]
- Maciejczyk, M.; Palka, T.; Wiecek, M.; Szymura, J.; Kusmierczyk, J.; Bawelski, M.; Masel, S.; Szygula, Z. Effects of Intermittent Hypoxic Training on Aerobic Capacity and Second Ventilatory Threshold in Untrained Men. Appl. Sci. 2023, 13, 9954. [Google Scholar] [CrossRef]
- Goliniewski, J.; Czuba, M.; Płoszczyca, K.; Chalimoniuk, M.; Gajda, R.; Niemaszyk, A.; Kaczmarczyk, K.; Langfort, J. The Impact of Normobaric Hypoxia and Intermittent Hypoxic Training on Cardiac Biomarkers in Endurance Athletes: A Pilot Study. Int. J. Mol. Sci. 2024, 25, 4584. [Google Scholar] [CrossRef]
- Andersen, A.B.; Bejder, J.; Bonne, T.C.; Nordsborg, N.B. Contemporary Blood Doping—Performance, Mechanism, and Detection. Scand. J. Med. Sci. Sports 2024, 34, e14243. [Google Scholar] [CrossRef]
- Gordeuk, V.R.; Key, N.S.; Prchal, J.T. Re-Evaluation of Hematocrit as a Determinant of Thrombotic Risk in Erythrocytosis. Haematologica 2019, 104, 653–658. [Google Scholar] [CrossRef]
- Chen, Z.; Han, F.; Du, Y.; Shi, H.; Zhou, W. Hypoxic Microenvironment in Cancer: Molecular Mechanisms and Therapeutic Interventions. Signal Transduct. Target. Ther. 2023, 8, 70. [Google Scholar] [CrossRef]
- Yasuoka, Y.; Izumi, Y.; Sands, J.M.; Kawahara, K.; Nonoguchi, H. Progress in the Detection of Erythropoietin in Blood, Urine, and Tissue. Molecules 2023, 28, 4446. [Google Scholar] [CrossRef]
- Nagel, M.J.; Jarrard, C.P.; Lalande, S. Effect of a Single Session of Intermittent Hypoxia on Erythropoietin and Oxygen-Carrying Capacity. Int. J. Environ. Res. Public Health 2020, 17, 7257. [Google Scholar] [CrossRef] [PubMed]
- Manescu, D.C. Alimentaţia în fitness şi bodybuilding. 2010, Editura AS.
- Hurst, P.; Kavussanu, M.; Davies, R.; Dallaway, N.; Ring, C. Use of Sport Supplements and Doping Substances by Athletes: Prevalence and Relationships. J. Clin. Med. 2024, 13, 7132. [Google Scholar] [CrossRef] [PubMed]
- Park, J. ; The mechanisms of anabolic steroids, selective androgen receptor modulators and resistance exercise in skeletal muscle hypertrophy. Korean J. Sports Med. 2022, 40, 67–77. [Google Scholar] [CrossRef]
- Waskiw-Ford, M.; Hodson, N.; Fung, H.J.W.; West, D.W.D.; Apong, P.; Bashir, R.; Moore, D.R. Essential Amino Acid Ingestion Facilitates Leucine Retention and Attenuates Myofibrillar Protein Breakdown following Bodyweight Resistance Exercise in Young Adults in a Home-Based Setting. Nutrients 2022, 14, 3532. [Google Scholar] [CrossRef]
- Sivalokanathan, S.; Małek, Ł.A.; Malhotra, A. The Cardiac Effects of Performance-Enhancing Medications: Caffeine vs. Anabolic Androgenic Steroids. Diagnostics 2021, 11, 324. [Google Scholar] [CrossRef]
- Nielsen, J.L.; Rasmussen, J.J.; Frandsen, M.N.; Fredberg, J.; Brandt-Jacobsen, N.H.; Aagaard, P.; Kistorp, C. Higher Myonuclei Density in Muscle Fibers Persists Among Former Users of Anabolic Androgenic Steroids. The Journal of Clinical Endocrinology & Metabolism 2024, 109. [CrossRef]
- Herlitz, L.C.; Markowitz, G.S.; Farris, A.B. ; Development of Focal Segmental Glomerulosclerosis after Anabolic Steroid Abuse. J. Am. Soc. Nephrol. 2010, 21, 163–172. [Google Scholar] [CrossRef]
- Brennan, B.P.; Kanayama, G.; Hudson, J.I.; Pope, H.G. Jr. Human Growth Hormone Abuse in Male Weightlifters. Am. J. Addict. 2011, 20, 9–13. [Google Scholar] [CrossRef]
- Hoshino, D.; Yoshida, Y.; Holloway, G.P.; Lally, J.; Hatta, H.; Bonen, A. Clenbuterol, a β2-Adrenergic Agonist, Reciprocally Alters PGC-1α and RIP140 and Reduces Fatty Acid and Pyruvate Oxidation in Rat Skeletal Muscle. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2012, 302, R373–R384. [Google Scholar] [CrossRef]
- Wiacek, M.; Trąbka, B.; Tomasiuk, R.; Zubrzycki, I.Z. Changes in Health-Related Parameters Associated with Sports Performance Enhancement Drugs. Int. J. Sports Med. 2023, 44, 206–214. [Google Scholar] [CrossRef]
- La Gerche, A.; Wasfy, M.M.; Brosnan, M.J.; Claessen, G.; Fatkin, D.; Heidbüchel, H.; Thompson, P.D. The Athlete’s Heart—Challenges and Controversies. J. Am. Coll. Cardiol. 2022, 80, 1346–1362. [Google Scholar] [CrossRef]
- Mănescu, A.M.; Grigoroiu, C.; Smîdu, N.; Dinciu, C.C.; Mărgărit, I.R.; Iacobini, A.; Mănescu, D.C. Biomechanical Effects of Lower Limb Asymmetry During Running: An OpenSim Computational Study. Symmetry 2025, 17, 1348. [Google Scholar] [CrossRef]
- Furrer, R.; Handschin, C. Molecular Aspects of the Exercise Response and Training Adaptation in Skeletal Muscle. Free Radic. Biol. Med. 2024, 223, 53–68. [Google Scholar] [CrossRef] [PubMed]
- Zubrzycki, I.Z.; Wiacek, M.; Trąbka, B.; Ossowski, Z. Anabolic Steroids in a Contest Preparation of the Top World-Class Bodybuilder. Int. J. Med. Pharm. Case Rep. 2015, 5, 1–8. [Google Scholar] [CrossRef]
- Plaza-Díaz, J.; Izquierdo, D.; Torres-Martos, Á.; Baig, A.T.; Aguilera, C.M.; Ruiz-Ojeda, F.J. Impact of Physical Activity and Exercise on the Epigenome in Skeletal Muscle and Effects on Systemic Metabolism. Biomedicines 2022, 10, 126. [Google Scholar] [CrossRef] [PubMed]
- Clemente-Suárez, V.J.; Bustamante-Sánchez, Á.; Mielgo-Ayuso, J.; Martínez-Guardado, I.; Martín-Rodríguez, A.; Tornero-Aguilera, J.F. Antioxidants and Sports Performance. Nutrients 2023, 15, 2371. [Google Scholar] [CrossRef]
- Li, J.; Zhang, H.; Zhou, H.; Liu, Q.; Xu, W.; Chen, X. Impact of Exercise and Aging on Mitochondrial Function via Epigenetic Modifications. Cells 2022, 11, 2086. [Google Scholar] [CrossRef]
- Williams, A.; Wadsworth, D.D.; Geetha, T. Exercise, Epigenetics, and Body Composition: Molecular Connections. Cells 2025, 14, 1553. [Google Scholar] [CrossRef]
- Zeng, Z.; Centner, C.; Gollhofer, A.; König, D. Effects of Dietary Strategies on Exercise-Induced Oxidative Stress: A Narrative Review of Human Studies. Antioxidants 2021, 10, 542. [Google Scholar] [CrossRef]
- Kolodziej, F.; O’Halloran, K.D. Re-Evaluating the Oxidative Phenotype: Can Endurance Exercise Save the Western World? Antioxidants 2021, 10, 609. [Google Scholar] [CrossRef]
- Moreno-Villanueva M, Kramer A, Hammes T, Venegas-Carro M, Thumm P, Bürkle A, Gruber, M. Influence of Acute Exercise on DNA Repair and PARP Activity before and after Irradiation in Lymphocytes from Trained and Untrained Individuals. Int J Mol Sci. 2019, 20, 2999. [Google Scholar] [CrossRef]
- Yasemi, M.; Jolicoeur, M. Modelling Cell Metabolism: A Review on Constraint-Based Steady-State and Kinetic Approaches. Processes 2021, 9, 322. [Google Scholar] [CrossRef]
- Manescu, D.C. Nutritional tips for muscular mass hypertrophy. Marathon 2016, 8, 79–83. [Google Scholar]
- Volkova, S.; Matos, M.R.A.; Mattanovich, M.; Marín de Mas, I. Metabolic Modelling as a Framework for Metabolomics Data Integration and Analysis. Metabolites 2020, 10, 303. [Google Scholar] [CrossRef]
| Pathway/Component | Nutritional or Energetic Trigger | Upstream Regulator(s) | Primary Molecular Effect | Downstream Target(s) | Adaptive Role (Physiological Context) | Potential Dysregulation (Overdrive State) |
| AMPK | Energy deficit, exercise, fasting | AMP/ATP ratio, LKB1 | Activates catabolic fluxes; inhibits mTORC1 | ACC, ULK1, PGC-1α | Enhances mitochondrial biogenesis, endurance, autophagy | Chronic suppression → anabolic dominance, insulin resistance |
| mTORC1 | Amino acids (leucine), insulin, IGF-1 | PI3K–AKT, Rheb, Rag GTPases | Stimulates protein synthesis via p70S6K and 4E-BP1 | Ribosomal proteins, eIF4E | Promotes muscle growth and recovery | Persistent activation → oxidative stress, epigenetic drift |
| SIRT1 | NAD+ levels, caloric restriction | AMPK, NAMPT | Deacetylates transcriptional regulators (PGC-1α, FOXO) | PGC-1α, p53, NF-κB | Increases oxidative metabolism and stress resistance | NAD+ depletion → loss of redox control and mitochondrial decline |
| IGF-1/AKT axis | Protein intake, GH, insulin | GH/IGF-1 signaling | Activates mTORC1, inhibits AMPK | FOXO, TSC2, GSK3β | Supports hypertrophy and tissue regeneration | Chronic activation → reduced autophagy, oncogenic signaling |
| Leucine–AMPK–mTOR crosstalk | Branched-chain amino acids | Ragulator complex | Coordinates anabolism–catabolism balance | mTORC1, AMPK | Fine-tunes training–fuel coupling, metabolic flexibility | Excess BCAA intake → insulin desensitization, ROS accumulation |
| NAD+/PARP balance | Oxidative load, DNA repair demand | PARP1, SIRT1 | Competes for NAD+ substrate | SIRT1, PGC-1α | Links redox balance to mitochondrial maintenance | PARP hyperactivation → NAD+ depletion, energy collapse |
| FOXO transcription factors | Energy deficit, oxidative stress | AMPK, SIRT1, AKT | Regulate autophagy, antioxidant enzymes | Catalase, MnSOD, LC3 | Antioxidant defense and longevity | AKT-mediated inhibition → reduced oxidative protection |
| Mechanistic domain | Representative trigger or intervention |
Physiological driver (Elite sport) |
Oncogenic analogue (Cancer biology) |
Key molecular mediator(s) | Adaptive outcome | Pathological counterpart (Overdrive state) |
| Anabolic signaling (mTOR/AKT) | Resistance training, growth hormone, anabolic steroids | Transient anabolic activation for muscle hypertrophy and recovery | PI3K–AKT–mTOR hyperactivation promoting uncontrolled cell proliferation | mTORC1, AKT, S6K1, 4E-BP1, TSC2 | Enhanced protein synthesis and tissue remodeling | Persistent anabolic signaling, autophagy suppression, tumor-like growth |
| Hypoxia and angiogenesis | Altitude exposure, EPO doping, hypoxic training | HIF-1α activation and VEGF upregulation for capillary and mitochondrial biogenesis | Tumor hypoxia, neovascularization | HIF-1α, VEGF, EPO, PGC-1α | Improved oxygen transport and energy efficiency | Pathological angiogenesis, oxidative stress, DNA instability |
| Oxidative stress and redox signaling | Intense endurance training, overreaching, nutrient excess | ROS-mediated activation of antioxidant defense and mitochondrial adaptation | Chronic ROS accumulation and oxidative DNA damage | NRF2, NF-κB, SIRT1, p53 | Adaptive hormesis, redox-dependent signaling plasticity | Genomic instability, lipid peroxidation, mitochondrial dysfunction |
| Autophagy–UPR balance | Caloric restriction, intermittent fasting, recovery phases | Controlled proteostasis and organelle turnover | Autophagy dysregulation and ER stress adaptation | AMPK, LC3, ATF4, GRP78 | Protein quality control and metabolic recycling | Proteotoxic stress, ER overload, resistance to apoptosis |
| Epigenetic reprogramming | Nutrient timing, methyl donor availability, supplementation | Transient changes in DNA methylation and histone acetylation for gene regulation | Stable oncogenic methylation, miRNA dysregulation | DNMT1, HDACs, TETs, miR-21 | Flexible gene expression, adaptive remodeling | Epigenetic drift, transcriptional instability, dedifferentiation |
| Hormonal and growth factor signaling | GH/IGF-1 therapy, insulin use, anabolic stacking | Enhanced recovery, anabolic sensitivity | Endocrine-driven tumor growth, hyperinsulinemia | IGF-1R, STAT5, ERK1/2 | Accelerated regeneration, anabolic drive | Endocrine disruption, persistent proliferative signaling |
| Metabolic reprogramming (Warburg-like effect) | High-intensity training, glucose or amino acid loading | Glycolytic shift for ATP and biosynthetic precursor supply | Aerobic glycolysis in tumors (Warburg effect) | HK2, PKM2, LDHA, MYC | Efficient energy flux for adaptation | Persistent glycolysis, one-carbon imbalance, redox exhaustion |
| Epigenetic layer | Primary metabolic cofactor or pathway | Key enzymes/regulators | Physiological role (adaptive) | Pathological outcome (overdrive state) | Representative molecular targets | Evidence from elite sport | Potential interventions/modulators |
| DNA methylation | SAM / one-carbon metabolism | DNMT1, TET1–3, MTHFR, MTR | Dynamic methylation of metabolic and mitochondrial genes; exercise-induced “epigenetic memory” | Hypermethylation of antioxidant genes; hypomethylation of anabolic/proliferative loci; methylation drift | PGC-1α, PDK4, TFAM, FOXO3, MYC | Endurance athletes show promoter hypomethylation of PGC-1α and TFAM after repeated training cycles | Folate and methionine balance, caloric restriction, AMPK activation |
| Histone acetylation | Acetyl-CoA, NAD+ / AMPK–SIRT1 axis | SIRT1, p300/CBP, GCN5, HDACs | Chromatin compaction under energy deficit; regulation of repair and mitochondrial genes | Hyperacetylation and persistent transcription of anabolic programs | H3K9ac, H4K16ac, PGC-1α, p53 | Training fasted or under caloric restriction enhances SIRT1 activity and deacetylation | Resveratrol, curcumin, exercise–fasting cycles |
| Histone and DNA demethylation | α-KG / TCA flux | TETs, Jumonji demethylases | Removal of repressive marks; mitochondrial-nuclear communication | α-KG depletion, succinate/fumarate inhibition of demethylases | H3K27me3, H3K9me3, IDH2, SDH | Overreaching phases show transient TET up-regulation linked to oxidative flux | Endurance exercise, antioxidant restoration |
| NAD+-dependent regulation | NAD+ salvage, PARP vs. sirtuins | PARP1, SIRT1, NAMPT | Balanced DNA repair and mitochondrial biogenesis | PARP overactivation → NAD+ depletion, SIRT1 silencing, redox collapse | PGC-1α, FOXO, PARP1, SIRT1 | Overtraining decreases muscle NAD+ and SIRT1, paralleling redox fatigue | Caloric restriction, AMPK activators, NAD+ precursors |
| MicroRNA regulation | Energy/redox-dependent transcription | miR-1, miR-21, miR-34a, miR-486, miR-494 | Fine-tuning of hypertrophy, oxidative capacity, stress defense | Aberrant miRNA expression driving fibrosis and oncogenic signaling | miR-21, miR-34a, miR-133a, miR-206 | Endurance ↑ miR-181a, miR-494; resistance ↑ miR-378, miR-486; overtraining ↑ miR-21 | Training modulation, antioxidant support |
| One-carbon and methyl donor flux | Methionine–folate cycle | MAT2A, BHMT, SHMT1/2 | Maintenance of methyl balance for DNA/histone regulation | SAM/SAH imbalance, global hypermethylation, redox-linked drift | DNMT3A, HMTs, MTHFD1L | High-protein diets alter plasma methionine and SAM/SAH ratios in athletes | Controlled protein intake, B-vitamin support |
| System component | Primary molecular driver | Key sensors or enzymes | Physiological feedback (adaptive) | Pathological feedback (overdrive state) | Representative molecular signature | Potential modulators / countermeasures |
| Mitochondrial ROS generation | Electron transport flux, NADH/NAD+ ratio | Complex I–III, NOX, SOD2 | ROS act as signaling molecules activating AMPK and antioxidant genes | Chronic ROS accumulation leads to mtDNA damage, lipid peroxidation, and nuclear stress signaling | ↑ ROS, ↑ SOD2, ↑ NRF2 | Endurance training, antioxidant periodization, redox-adaptive nutrition |
| NAD+ metabolism | NAMPT salvage pathway, PARP activity | NAMPT, PARP1, SIRT1 | Balanced NAD+ use supports DNA repair and mitochondrial biogenesis | PARP overactivation depletes NAD+, silencing SIRT1 and impairing repair | ↓ NAD+, ↓ SIRT1, ↑ PARylation | Caloric restriction, resveratrol, niacinamide, AMPK activation |
| AMPK–SIRT1–PGC-1α axis | Energy sensing, NAD+/AMP ratio | AMPK, SIRT1, PGC-1α | Enhances oxidative metabolism, mitophagy, and chromatin integrity | Collapse of AMPK–SIRT1 feedback causes autophagy failure and metabolic rigidity | ↑ PGC-1α, ↑ LC3-II, ↓ FOXO | Exercise-induced AMPK activation, fasting cycles |
| DNA and histone modifications | SAM/SAH ratio, α-KG availability | DNMTs, TETs, HDACs, HATs | Dynamic methylation/acetylation maintains gene expression flexibility | Methylation drift and histone hyperacetylation stabilize maladaptive transcription | ↑ H3K9ac, ↓ TET activity, ↑ DNMT1 | Balanced methyl donor intake, one-carbon flux restoration |
| Inflammatory redox signaling | NF-κB, NLRP3, cytokine ROS loops | NF-κB, IL-6, TNF-α | Transient activation supports repair and immune remodeling | Chronic activation sustains oxidative stress and metabolic block | ↑ NF-κB, ↑ IL-6, ↑ TNF-α | Polyphenols, omega-3s, anti-inflammatory recovery |
| Epigenetic memory and drift | ROS/NAD+-dependent enzyme regulation | SIRT1, PARP1, DNMT1, miRNAs | Transient chromatin remodeling encodes adaptive responses | Persistent oxidative stress leads to irreversible epigenetic drift | Hypomethylated oncogenes, hyperacetylated histones | Controlled recovery, antioxidant therapy, NAD+ support |
| System-level outcome | Redox–epigenetic coupling | Integrated signaling through AMPK–mTOR–SIRT1 | Self-limiting oscillation ensures resilience | Feedback saturation locks system in pathological anabolism | ↓ AMPK, ↑ mTOR, ↑ ROS | Nutritional periodization, training load modulation |
| Regulatory axis | Adaptive (oscillatory) state |
Overdrive (lock-in) state |
Representative molecular indicators |
Functional outcome (physiological/ clinical) |
Evidence or relevance in elite sport | Reversibility potential |
| Nutritional signaling (AMPK–mTOR) | Alternating activation maintains energy balance and anabolic–catabolic cycling; mTOR activity transient and self-limiting. | Chronic nutrient or hormonal stimulation suppresses AMPK; mTORC1 locked in constitutive activation. | Adaptive: ↑ AMPK (Thr172-P), oscillatory ↑ mTOR (Ser2448-P); Overdrive: ↑ S6K1, ↓ AMPK. | Efficient recovery, balanced hypertrophy, metabolic flexibility. | Observed in endurance vs. bulking athletes; chronic protein or insulin use suppresses AMPK signaling. | High — restored through fasting, caloric periodization, AMPK activators. |
| Redox homeostasis (ROS–NAD+ balance) | Controlled ROS pulses activate NRF2–SIRT1 defense; NAD+ recycling sustains redox tone. | ROS accumulation exceeds detox capacity; PARP activation depletes NAD+, silencing SIRT1. | Adaptive: ↑ NRF2, ↑ SIRT1, stable NAD+/NADH; Overdrive: ↑ PARP1, ↓ NAD+, ↑ oxidative adducts. | Enhanced mitochondrial efficiency and repair capacity. | Seen in overtraining and altitude exposure; redox imbalance linked to delayed recovery and fatigue. | Moderate — restored with NAD+ boosters, antioxidant periodization. |
| Epigenetic regulation (chromatin remodeling) | Reversible histone deacetylation and DNA demethylation maintain transcriptional flexibility; stress memory remains functional. | Hyperacetylation and aberrant methylation fix anabolic programs; transcriptional noise accumulates. | Adaptive: ↑ TET1–3, ↑ HDACs; Overdrive: ↑ DNMT1, ↑ HATs, ↑ H3K9ac. | Improved adaptation memory, genomic stability. | Exercise-induced hypomethylation of PGC-1α and TFAM vs. hypermethylation under chronic overload. | Variable — reversible early, lost under prolonged overload. |
| Mitochondrial dynamics | Balanced fusion–fission and autophagic recycling sustain bioenergetic quality. | Hyperpolarization, impaired mitophagy, ROS leakage disrupt energy homeostasis. | Adaptive: ↑ PGC-1α, ↑ MFN2, ↑ LC3-II; Overdrive: ↑ DRP1, ↑ ROS. | Optimal endurance and energy turnover. | Endurance training enhances mitophagy; chronic supplementation or doping impairs mitochondrial turnover. | Moderate — restored through recovery and redox normalization. |
| System-level behavior | Oscillatory negative feedback ensures renewal after stress; system remains dynamically stable. | Positive feedback loops reinforce anabolic and oxidative stress; system enters chaotic saturation. | Adaptive: oscillatory AMPK/mTOR ratio; Overdrive: flattened rhythm, ↑ entropy. | Resilient adaptation, sustained performance. | Elite athletes under chronic load exhibit reduced HRV and hormonal adaptability—biomarkers of systemic lock-in. | Low — requires full metabolic reset through deloading and restoration cycles. |
| Mechanistic axis |
Primary human readouts (assay & specimen) |
Predicted direction in overdrive | Evidence base (cell/animal/human) |
Reversibility signal (periodic AMPK) |
Preferred sampling window |
| AMPK–mTOR balance | AMPK Thr172-P (WB/ELISA; muscle or PBMCs), p-S6K1 (Thr389) / p-4E-BP1 (Ser65) (WB), LC3-II/p62 (autophagic flux with lysosomal inhibition) | ↓ AMPK Thr172-P; ↑ p-S6K1/4E-BP1; ↓ autophagic flux | Strong (cells/rodents) + human acute & training blocks |
↑ AMPK Thr172-P; normalization of p-S6K1; ↑ LC3-II | 24–48 h post-stimulus, spanning feeding |
| NAD+ economy (SIRT1–PARP) | NAD+/NADH (enzymatic cycling; muscle/PBMCs), PARylation (WB), SIRT1 activity (fluor./ELISA) | ↓ NAD+; ↑ PARylation; ↓ SIRT1 activity | Moderate–strong (cells/rodents) + human acute (exercise/irradiation) |
↑ NAD+; ↓ PARylation; ↑ SIRT1 | 0–6 h and 24 h post-stimulus |
| Redox status | 8-oxo-dG (DNA damage), GSH/GSSG, TBARS/MDA, SOD2/Catalase (WB/activity) | ↑ oxidative adducts; ↑ inflammatory redox tone | Strong across models; robust human acute | ↓ adducts; normalization of GSH/GSSG | Immediately post + 24–48 h |
| Epigenetic layer | H3K9ac (ChIP-WB), 5mC/5hmC (LC-MS/MS; targeted bisulfite), TET activity, DNMT1; target loci: PGC-1α, TFAM, PDK4 | ↑ H3K9ac; ↓ TET/5hmC at oxidative loci; ↑ DNMT1; methylation drift | Moderate (cells/rodents) + human acute/longitudinal |
↓ H3K9ac; ↑ 5hmC/TET upon AMPK restoration | 24–72 h (miRNAs earlier) |
| miRNA remodeling | Plasma/serum miR-181a/494 (oxidative), miR-378/486 (hypertrophic), miR-21/34a (fibro-oncogenic) | ↑ miR-21/34a; rigidified profile | Human: acute + training periods |
Shift back toward oxidative profile (miR-181a/494) | 0–24 h |
| Mitochondrial quality | PGC-1α, MFN2, DRP1, mtDNA damage, LC3-II | ↓ mitophagy; ↑ DRP1; ↑ mtROS | Robust experimental; emerging human data | ↑ mitophagy; ↑ PGC-1α; reduced mtROS | 24–72 h |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




