Submitted:
16 October 2025
Posted:
17 October 2025
You are already at the latest version
Abstract
1) Background: The emergence of multidrug-resistant (MDR)-Salmonella enterica poses a major threat to global public health underscoring the urgent need for alternative therapeutic strategies where bacteriophages are a promising candidate due to their high specificity and potent ability to lyse MDR strains.2) Methods: In this study, we isolated a novel MDR-Salmonella Enteritidis-targeting bacteriophage from Lebanese sewage and characterized its host range, thermal and PH stability, and infection dynamics. Whole-genome sequencing was performed by Illumina to determine genetic features and taxonomic position.3) Results: the bacteriophage belongs to genus Jerseyvirus and Class Caudoviricetes with a 43kb dsDNA genome and 66 ORFs. It demonstrated extraordinary thermal stability, retaining infectivity after prolonged incubation at 65 °C, showed a broad host range, produced large, clear plaques, displayed rapid adsorption (>97% within 3 min), a short latent period (20 min), and a burst size of ~320 PFU per cell. Genome analysis revealed no lysogeny, virulence, or resistance genes, confirming its strictly lytic nature and supporting its potential use as a biocontrol agent. 4) Conclusions: These findings identify SA01 as a novel, strictly lytic, and thermally stable bacteriophage with strong potential as a biocontrol agent against multidrug-resistant Salmonella Enteritidis, supporting its potential activity also against other Salmonella enterica serovars for future applications in food safety and biotechnology.
Keywords:
1. Introduction
2. Materials and Methods
Bacterial Strain and Growth Conditions
Screening and Isolation of S01 Bacteriophage
Bacteriophage Bacteriolytic Activity and Host Range
Bacteriophage One-Step Growth Curve and Adsorption Rate Assay
Bacteriophage Thermal and pH Stability
Bacteriophage Genome Sequencing and Analysis
3. Results
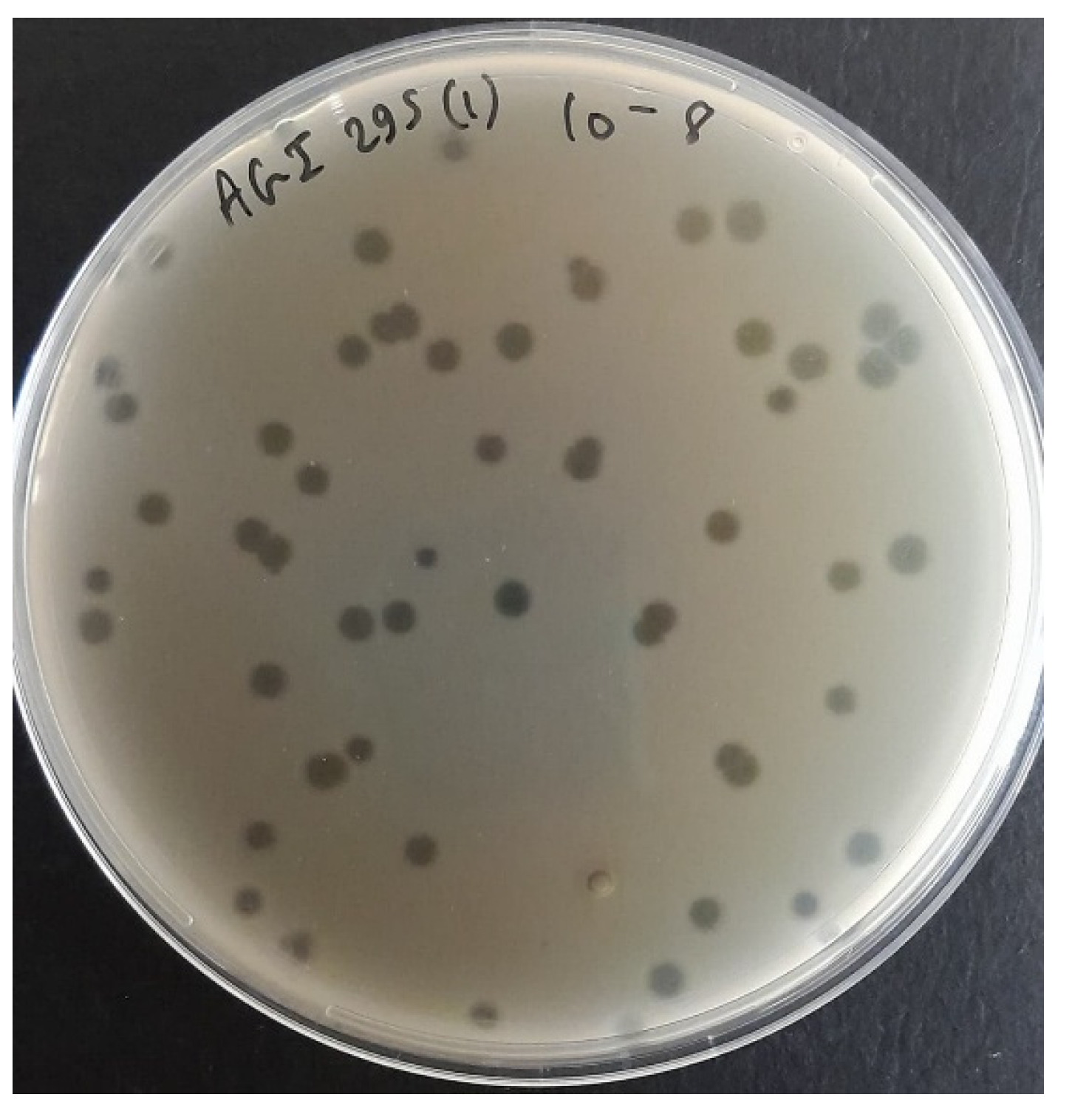
3.1. Bacteriophage Isolation and Purification
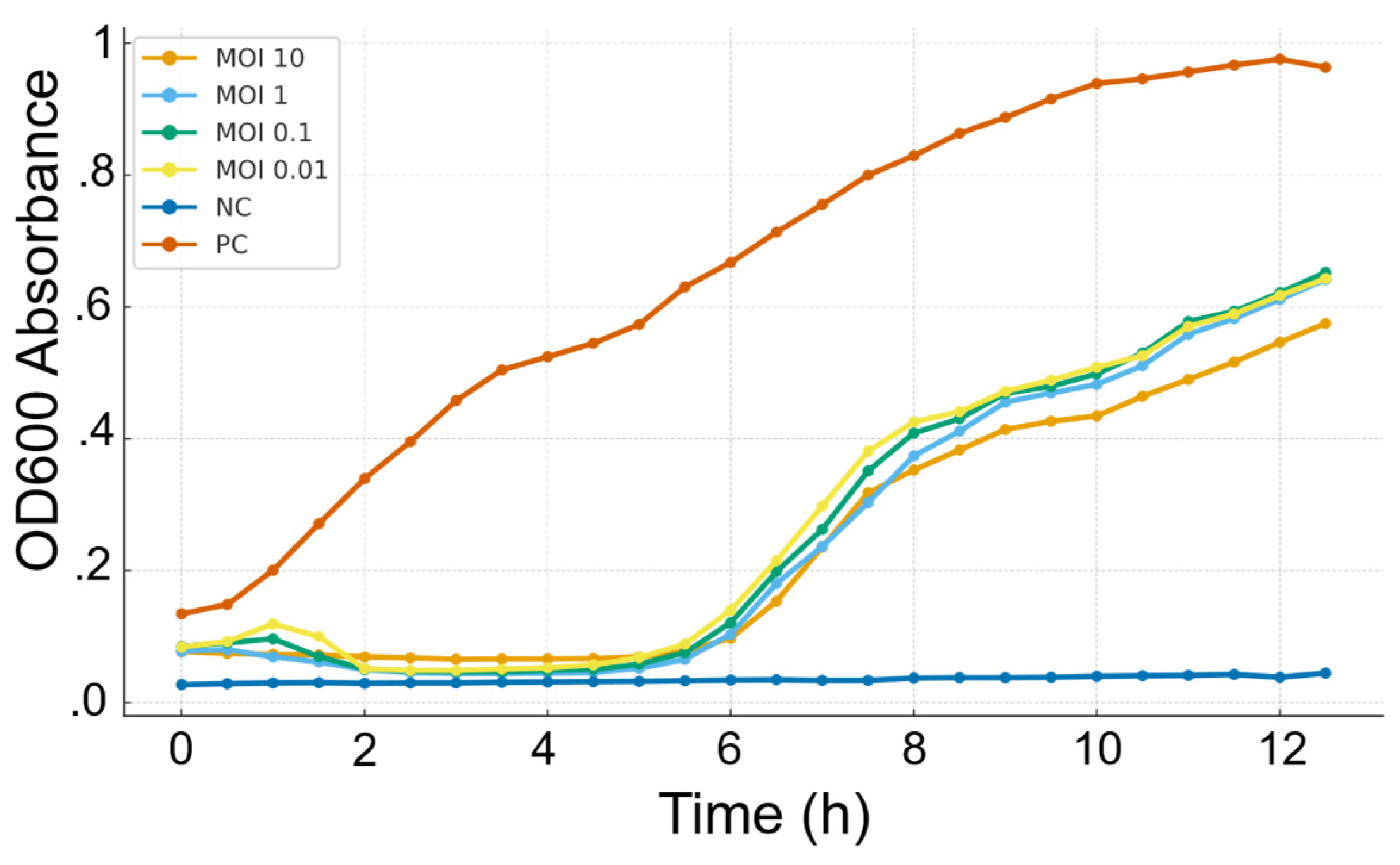
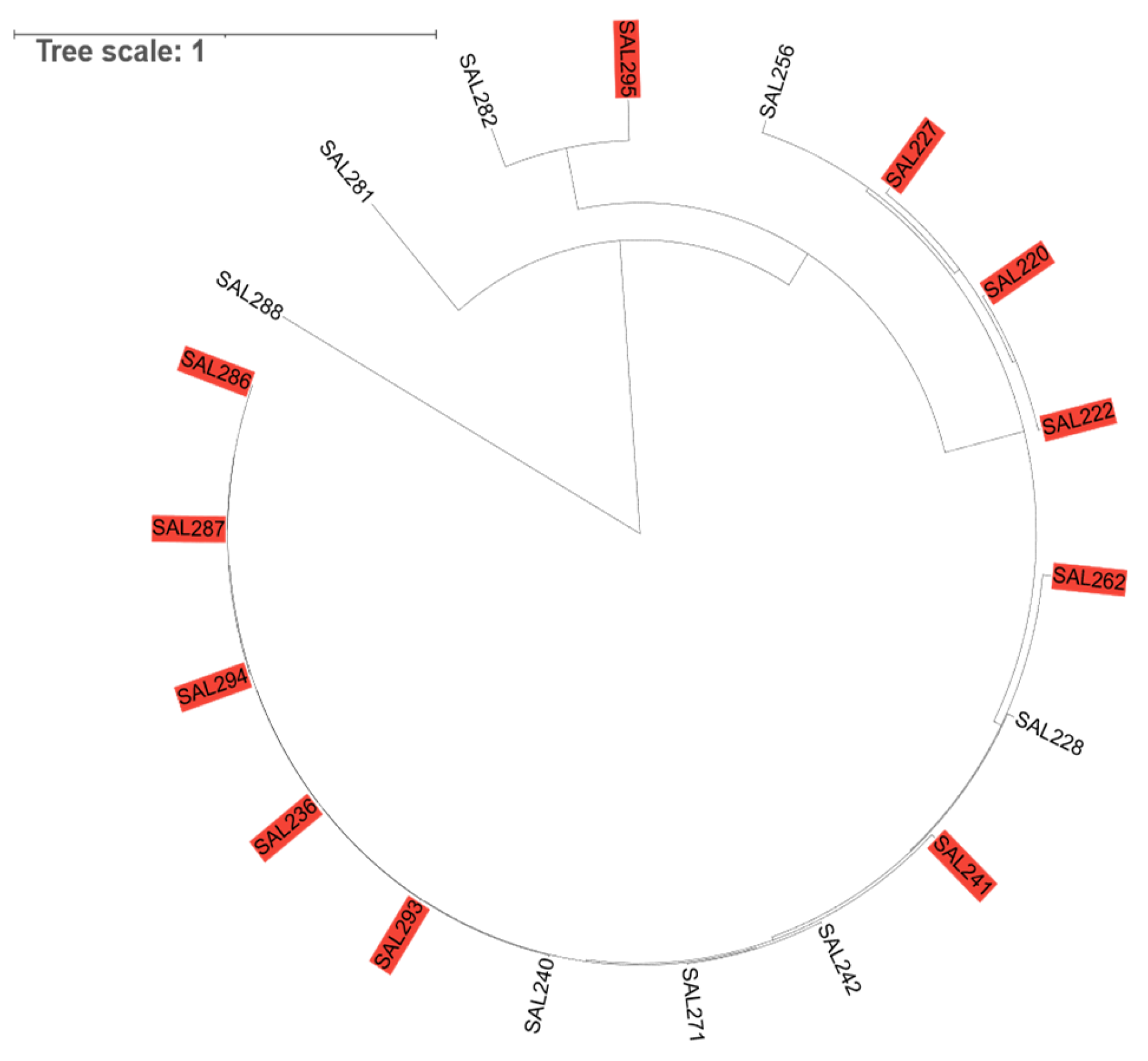
3.2. Bacteriolytic Activity and Host Range
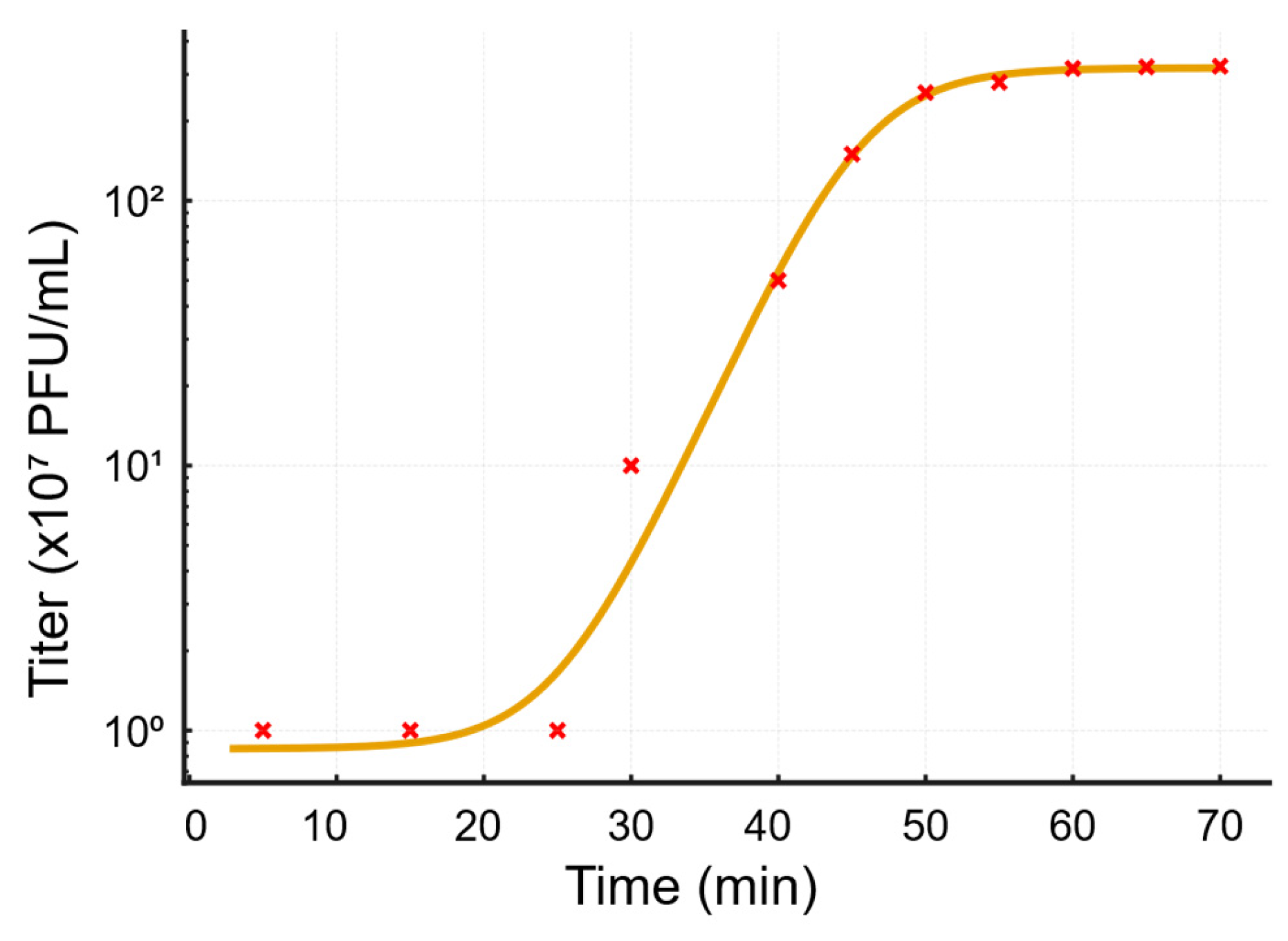
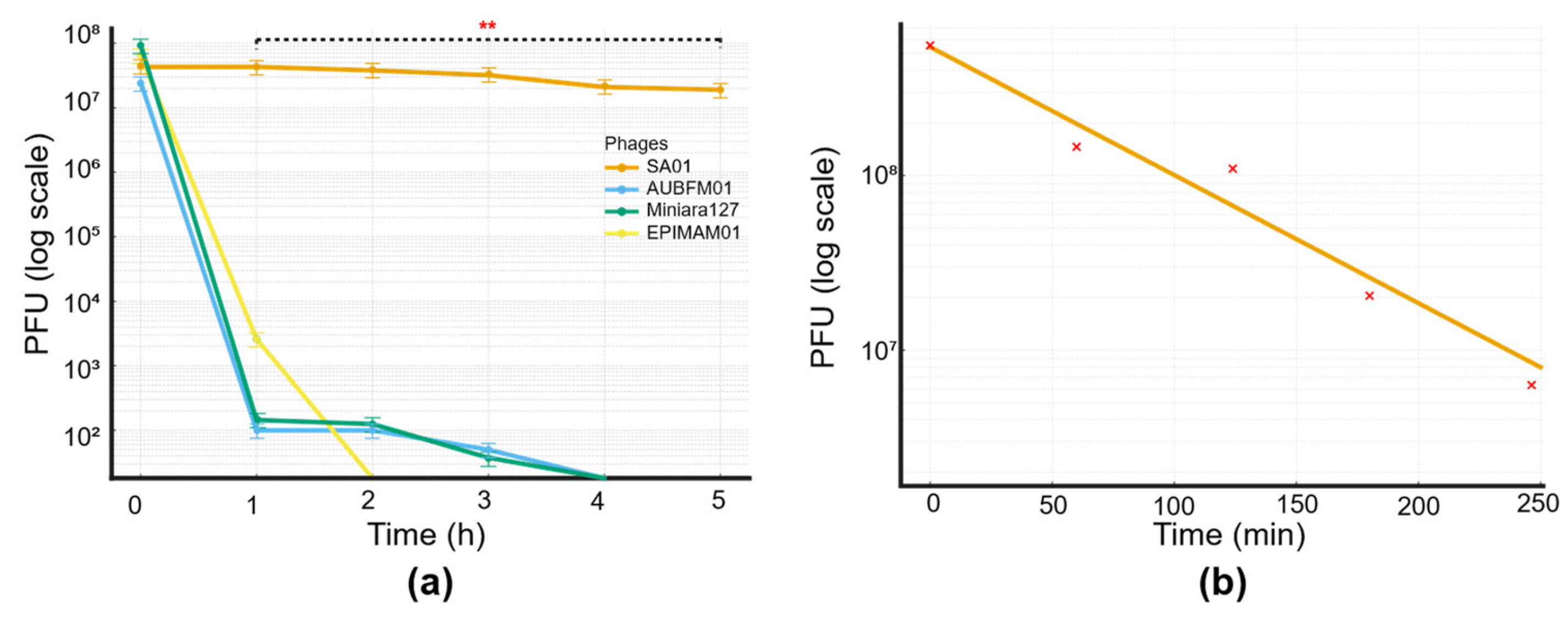
3.3. BacterioPhage–Host Interaction Dynamics
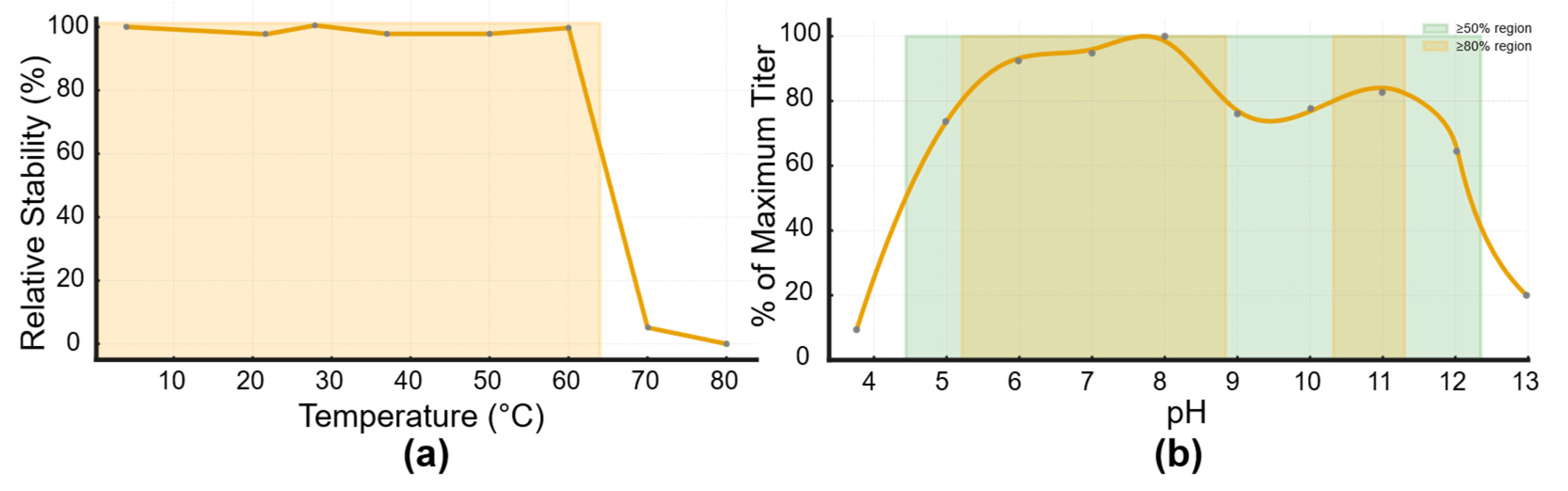
3.4. Thermal and PH Stability
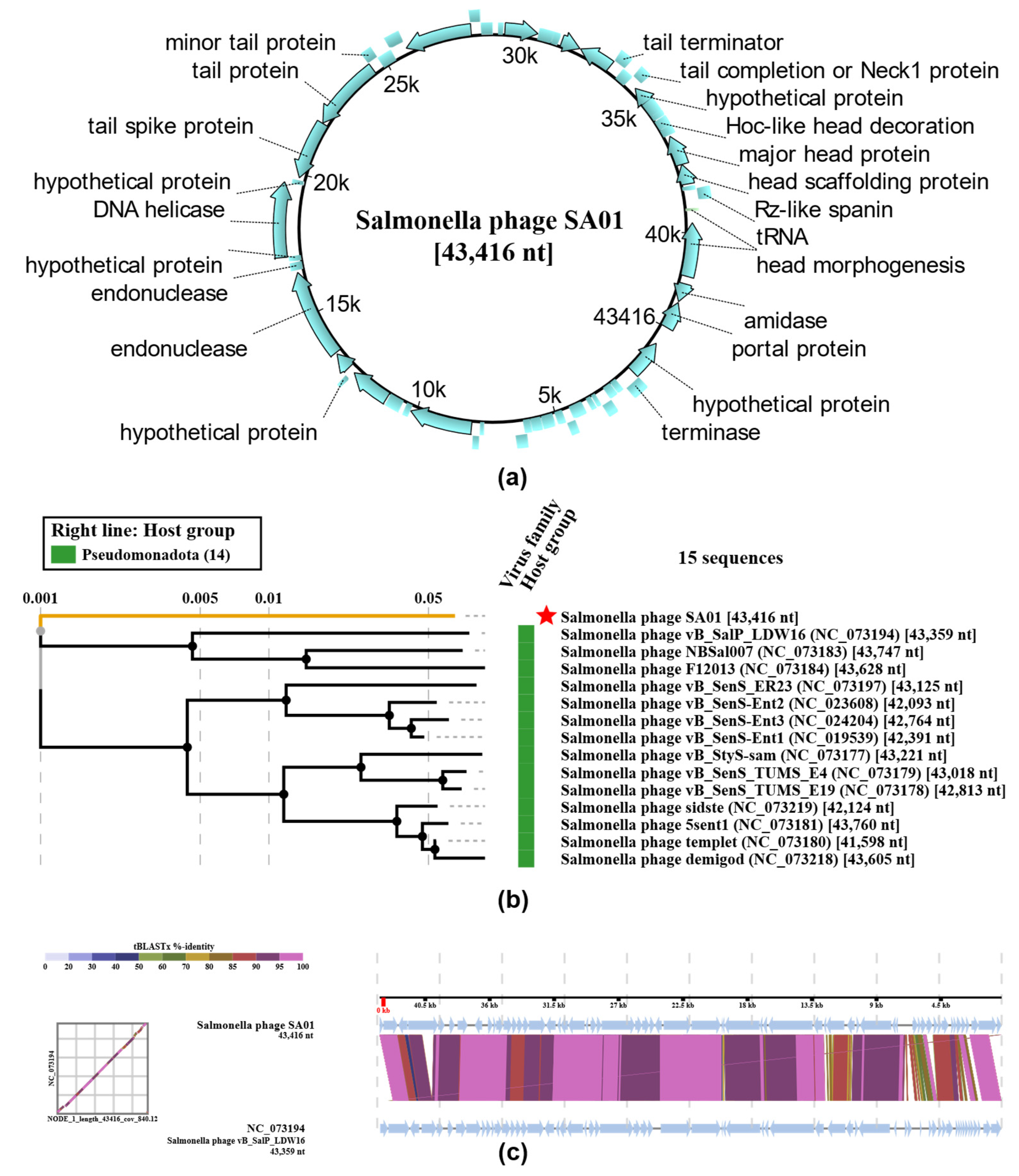
3.5. Whole-Genome Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| MDR | Multi-drug resistant |
| RTE | Ready-to-eat food |
| AMR | antimicrobial-resistant |
| ATCC | American Type Culture Collection |
| WHO | World Health Organization |
| FDA | Food and Drug Administration |
| GRAS | Generally recognized as safe |
| LB | Luria–Bertani |
| OD | optical density |
| PFU | Plaque forming unit |
| MOI | multiplicity of infection |
| ORF | open reading frame |
| tBLASTx | Translated Basic Local Alignment Search Tool |
Appendix A
Appendix A.1
| ID | Genus | Serovar | ST | BioProject | BioSample | Source (site) | Plasmids |
| SAL220 | Salmonella | Enteritidis | 11 | PRJNA613441 | SAMN43315390 | Human stool | IncFIB(S) |
| SAL222 | Salmonella | Enteritidis | 11 | PRJNA613441 | SAMN43315391 | Human stool | IncFIB(S) |
| SAL226 | Salmonella | Enteritidis | 11 | PRJNA613441 | SAMN43315395 | Human stool | IncFIB(S), IncFII(S) |
| SAL227 | Salmonella | Enteritidis | 11 | PRJNA613441 | SAMN43315396 | Human stool | IncFIB(S) |
| SAL228 | Salmonella | Enteritidis | 11 | PRJNA613441 | SAMN43315397 | Human stool | IncFIB(S), IncFII(S) |
| SAL236 | Salmonella | Enteritidis | 11 | PRJNA613441 | SAMN43315404 | Human stool | IncFIB(S), IncFII(S) |
| SAL240 | Salmonella | Enteritidis | 11 | PRJNA613441 | SAMN43315408 | Human stool | IncFIB(S), IncFII(S) |
| SAL241 | Salmonella | Enteritidis | 11 | PRJNA613441 | SAMN43315409 | Human stool | IncFIB(S), IncFII(S) |
| SAL242 | Salmonella | Enteritidis | 11 | PRJNA613441 | SAMN43315410 | Human stool | IncFIB(S), IncFII(S) |
| SAL256 | Salmonella | Enteritidis | 11 | PRJNA613441 | SAMN43309870 | Human stool | IncFIB(S) |
| SAL262 | Salmonella | Enteritidis | 11 | PRJNA613441 | SAMN43309873 | Human stool | IncFIB(S),IncFII(S) |
| SAL281 | Salmonella | Enteritidis | 11 | PRJNA613442 | SAMN52031898 | Human stool | IncFIB(S),IncFII(S) |
| SAL282 | Salmonella | Enteritidis | 11 | PRJNA613443 | SAMN52031899 | Human stool | IncFIB(S), IncFII(S), IncI1-I(Alpha) |
| SAL286 | Salmonella | Enteritidis | 11 | PRJNA613444 | SAMN52031900 | Human stool | IncFIB(S), IncFII(S) |
| SAL287 | Salmonella | Enteritidis | 11 | PRJNA613445 | SAMN52031901 | Human stool | IncFIB(S), IncFII(S) |
| SAL288 | Salmonella | Saintpaul | 11 | PRJNA613446 | SAMN52031902 | Human stool | IncI1-I(Alpha), IncP1 |
| SAL293 | Salmonella | Enteritidis | 11 | PRJNA613445 | SAMN52031905 | Human stool | IncFIB(S), IncFII(S) |
| SAL294 | Salmonella | Enteritidis | 11 | PRJNA613445 | SAMN52033040 | Human stool | IncFIB(S), IncFII(S) |
| SAL295 | Salmonella | Enteritidis | 11 | PRJNA613445 | SAMN52033041 | Human stool | IncFIB(S), IncFII(S), IncI1-I(Alpha) |
| SAL271 | Salmonella | Enteritidis | 11 | PRJNA613446 | SAMN43309880 | Human Stool | IncFIB(S),IncFII(S) |
Appendix A.2
| ID | Ceftriaxone CRO (R≥ 4) VITEK | Meropenem (R≥ 4) | Ampicillin (R≥ 32) | Amoxicillin-clavulanic acid -Amoclan (≥8) EUCAST | Gentamicin (R>2) EUCAST | Ciprofloxacin (R≥ 1) |
| SAL220 | >64 R | >16 R | <0.25 S | <2 S | >16 R | 4 R |
| SAL222 | <1 S | <0.25 S | <0.25 S | <2 S | >16 R | 4 R |
| SAL226 | <1 S | <0.25 S | <0.25 S | <2 S | >16 R | 4 R |
| SAL227 | <1 S | <0.25 S | <0.25 S | <2 S | <1 S | 4 R |
| SAL228 | <1 S | <0.25 S | 6 R | 32 R | >16 R | 4 R |
| SAL236 | <1 S | <0.25 S | <2 S | <2 S | >16 R | <0.25 S |
| SAL240 | <1 S | <0.25 S | <2 S | <8 S | <1 S | <0.25 S |
| SAL241 | <1 S | <0.25 S | <2 S | <8 S | >16 R | <0.25 S |
| SAL242 | <1 S | <0.25 S | <2 S | <8 S | >16 R | <0.25 S |
| SAL256 | 1 S | 0.25 S | 2 S | 8 R | >16 R | 0.25 S |
| SAL262 | 1 S | 0.25 S | 2 S | 8 R | >16 R | >16 R |
| SAL281 | 1 S | 0.25 S | 2 S | 8 R | >16 R | >16 R |
| SAL282 | 1 S | 0.25 S | 2 S | 8 R | >16 R | >16 R |
| SAL286 | 1 S | 0.25 S | 2 S | 8 R | >16 R | >16 R |
| SAL287 | 1 S | 0.25 S | 2 S | 8 R | >16 R | >16 R |
| SAL288 | 1 S | 0.25 S | 2 S | 8 R | >16 R | >16 R |
| SAL293 | S | S | R | R | R | R |
| SAL294 | S | S | R | S | R | R |
| SAL295 | R | S | S | R | R | R |
| SAL271 | 1 S | 0.25 S | 2 S | 8 R | >16 R | >16 R |
| ID | Nalidixic acid (≥32) VITEK | Tetracycline (R≥ 16) VITEK | Chloramphenicol (R≥ 32) BMD | Colistin (>2) BMD |
Trimethoprim-Sulfamethoxazole ( ≥4/76) VITEK | Azithromycin (≥32) BMD |
| SAL220 | 64 R | >16 R | 64 R | 1 S | >512 R | >512 R |
| SAL222 | >512 R | 20 S | 8 S | 8 R | 512 R | 64 R |
| SAL226 | >512 R | >16 R | 64 R | 2 S | >512 R | 64 R |
| SAL227 | >512 R | >16 R | 16 S | 1 S | >512 R | 16 S |
| SAL228 | >512 R | >16 R | 64 R | 1 S | >512 R | 64 R |
| SAL236 | >512 R | <1 S | 64 R | 0.5 S | >512 R | 32 R |
| SAL240 | >32 R | <1 S | 64 R | <0.25 S | >512 R | 2 S |
| SAL241 | >32 R | <1 S | 32 R | <0.25 S | >512 R | 4 S |
| SAL242 | >32 R | <1 S | 16 S | 16 R | <20 S | 2 S |
| SAL256 | 32 R | 16 R | 128 R | 1 S | 16 R | 4 S |
| SAL262 | 32 R | 16 R | 32 R | 0.5 S | 16 R | 4 S |
| SAL281 | 32 R | 16 R | >512 R | <0.25 S | 20 S | 64 R |
| SAL282 | 32 R | 16 R | >512 R | <0.25 S | 20 S | 128 R |
| SAL286 | 32 R | 16 R | >512 R | <0.25 S | 20 S | 64 R |
| SAL287 | 32 R | 16 R | >512 R | <0.25 S | 20 S | 128 R |
| SAL288 | 32 R | 16 R | >512 R | 16 R | 16 R | 64 R |
| SAL293 | R | R | S | S | R | S |
| SAL294 | R | 16 R | S | S | R | S |
| SAL295 | R | 16 R | S | S | S | S |
| SAL271 | 32 R | 1 S | 256 R | 2 S | 16 R | 4 S |
Appendix A.3
| ID | Resistance Profile |
| SAL220 | XDR |
| SAL222 | MDR |
| SAL226 | MDR |
| SAL227 | MDR |
| SAL228 | MDR |
| SAL236 | MDR |
| SAL240 | MDR |
| SAL241 | MDR |
| SAL242 | MDR |
| SAL256 | MDR |
| SAL262 | MDR |
| SAL281 | MDR |
| SAL282 | MDR |
| SAL286 | MDR |
| SAL287 | MDR |
| SAL293 | MDR |
| SAL294 | MDR |
| SAL295 | MDR |
| SAL271 | MDR |
References
- B. Coburn, G. A. Grassl, and B. B. Finlay, “Salmonella, the host and disease: a brief review,” Immunology & Cell Biology, vol. 85, no. 2, pp. 112–118, 2007, . [CrossRef]
- J. D. Stanaway et al., “The global burden of non-typhoidal salmonella invasive disease: a systematic analysis for the Global Burden of Disease Study 2017,” Lancet Infect. Dis., vol. 19, no. 12, pp. 1312–1324, 2019, . [CrossRef]
- ”Animal sources of salmonellosis in humans in: Journal of the American Veterinary Medical Association Volume 221 Issue 4 ().” Accessed: Sept. 10, 2025. [Online]. Available: https://avmajournals.avma.org/view/journals/javma/221/4/javma.2002.221.492.xml.
- ”Salmonella enterica Serovar Gallinarum Biovars Pullorum and Gallinarum in Poultry: Review of Pathogenesis, Antibiotic Resistance, Diagnosis and Control in the Genomic Era.” Accessed: Sept. 10, 2025. [Online]. Available: https://www.mdpi.com/2079-6382/13/1/23.
- N. Shivaning Karabasanavar, C. Benakabhat Madhavaprasad, S. Agalagandi Gopalakrishna, J. Hiremath, G. Shivanagowda Patil, and S. B Barbuddhe, “Prevalence of Salmonella serotypes S. Enteritidis and S. Typhimurium in poultry and poultry products,” J. Food Saf., vol. 40, no. 6, 2020, . [CrossRef]
- ”Tackling antibiotic resistance: the environmental framework—PubMed.” Accessed: Sept. 07, 2025. [Online]. Available: https://pubmed.ncbi.nlm.nih.gov/25817583/.
- Elbehiry et al., “Microbial Food Safety and Antimicrobial Resistance in Foods: A Dual Threat to Public Health,” Microorganisms, vol. 13, no. 7, p. 1592, July 2025, . [CrossRef]
- ”Salmonella Infection in Poultry: A Review on the Pathogen and Control Strategies.” Accessed: Sept. 10, 2025. [Online]. Available: https://www.mdpi.com/2076-2607/11/11/2814.
- A. Tolooe, M. Alizadeh, K. Blake, J. S. Doost, and S. Sharif, “Control of Salmonella in poultry: The role of host immunity and vaccines,” Poultry Science, vol. 104, no. 11, p. 105692, Nov. 2025, . [CrossRef]
- E. K. Silbergeld, J. Graham, and L. B. Price, “Industrial food animal production, antimicrobial resistance, and human health,” Annu Rev Public Health, vol. 29, pp. 151–169, 2008, . [CrossRef]
- T. F. Landers, B. Cohen, T. E. Wittum, and E. L. Larson, “A review of antibiotic use in food animals: perspective, policy, and potential,” Public Health Rep, vol. 127, no. 1, pp. 4–22, 2012, . [CrossRef]
- T. P. Robinson et al., “Antibiotic resistance is the quintessential One Health issue,” Trans R Soc Trop Med Hyg, vol. 110, no. 7, pp. 377–380, July 2016, . [CrossRef]
- European Food Safety Authority and European Centre for Disease Prevention and Control, “The European Union summary report on antimicrobial resistance in zoonotic and indicator bacteria from humans, animals and food in 2016,” EFSA J, vol. 16, no. 2, p. e05182, Feb. 2018, . [CrossRef]
- T. A. Ahmad, S. E. Houjeiry, S. S. Kanj, G. M. Matar, and E. S. Saba, “From forgotten cure to modern medicine: The resurgence of bacteriophage therapy,” Journal of Global Antimicrobial Resistance, vol. 39, pp. 231–239, Dec. 2024, . [CrossRef]
- ”Phage Products for Fighting Antimicrobial Resistance.” Accessed: Sept. 07, 2025. [Online]. Available: https://www.mdpi.com/2076-2607/10/7/1324.
- ”Comprehensive Evaluation of the Safety and Efficacy of BAFASAL® Bacteriophage Preparation for the Reduction of Salmonella in the Food Chain.” Accessed: Sept. 07, 2025. [Online]. Available: https://www.mdpi.com/1999-4915/12/7/742.
- K. Shahin et al., “Bio-control of O157:H7, and colistin-resistant MCR-1-positive Escherichia coli using a new designed broad host range phage cocktail,” LWT, vol. 154, p. 112836, Jan. 2022, . [CrossRef]
- Y. Nishimura, T. Yoshida, M. Kuronishi, H. Uehara, H. Ogata, and S. Goto, “ViPTree: the viral proteomic tree server,” Bioinformatics, vol. 33, no. 15, pp. 2379–2380, Aug. 2017, . [CrossRef]
- K. Yukgehnaish et al., “PhageLeads: Rapid Assessment of Phage Therapeutic Suitability Using an Ensemble Machine Learning Approach,” Viruses, vol. 14, no. 2, p. 342, Feb. 2022, . [CrossRef]
- P. Hyman, “Phages for Phage Therapy: Isolation, Characterization, and Host Range Breadth,” Pharmaceuticals, vol. 12, no. 1, p. 35, Mar. 2019, . [CrossRef]
- S. A. Strathdee, G. F. Hatfull, V. K. Mutalik, and R. T. Schooley, “Phage therapy: From biological mechanisms to future directions,” Cell, vol. 186, no. 1, pp. 17–31, 2023, . [CrossRef]
- H. Ge, Y. Wang, and X. Zhao, “Research on the drug resistance mechanism of foodborne pathogens,” Microb. Pathog., vol. 162, 2022, . [CrossRef]
- E. F. S. Authority and European Centre for Disease Prevention and Control, “The European Union One Health 2021 Zoonoses Report,” EFSA Journal, vol. 20, no. 12, p. e07666, 2022, . [CrossRef]
- ”Global meat consumption, World, 1961 to 2050.” Accessed: Sept. 11, 2025. [Online]. Available: https://ourworldindata.org/grapher/global-meat-projections-to-2050.
- S. T. Abedon, “Detection of Bacteriophages: Phage Plaques,” in Bacteriophages, Springer, Cham, 2021, pp. 507–538. [CrossRef]
- Z. Sun, Mandlaa, H. Wen, L. Ma, and Z. Chen, “Isolation, characterization and application of bacteriophage PSDA-2 against Salmonella Typhimurium in chilled mutton,” PLoS One, vol. 17, no. 1, p. e0262946, Jan. 2022, . [CrossRef]
- Huang et al., “Isolation, Characterization, and Application of Bacteriophage LPSE1 Against Salmonella enterica in Ready to Eat (RTE) Foods,” Front Microbiol, vol. 9, p. 1046, May 2018, . [CrossRef]
- X. Chen, C. Zhang, J. Guo, X. Huang, R. Lv, and X. Quan, “Thermal and Chemical Inactivation of Bacillus Phage BM-P1,” Journal of Food Protection, vol. 87, no. 3, p. 100223, Mar. 2024, . [CrossRef]
- M. Müller-Merbach, T. Rauscher, and J. Hinrichs, “Inactivation of bacteriophages by thermal and high-pressure treatment,” International Dairy Journal, vol. 15, no. 6, pp. 777–784, June 2005, . [CrossRef]
- ”Quiberoni: Inactivation of Lactobacillus delbrueckii...—Google Scholar.” Accessed: Sept. 13, 2025. [Online]. Available: https://scholar.google.com/scholar_lookup?title=Inactivation%20of%20Lactobacillus%20delbrueckii%20bacteriophages%20by%20heat%20and%20biocides&publication_year=2002&author=A.%20Quiberoni&author=D.M.%20Guglielmotti&author=J.A.%20Reinheimer.
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




