Submitted:
02 October 2025
Posted:
04 October 2025
You are already at the latest version
Abstract
Keywords:
Introduction
Results
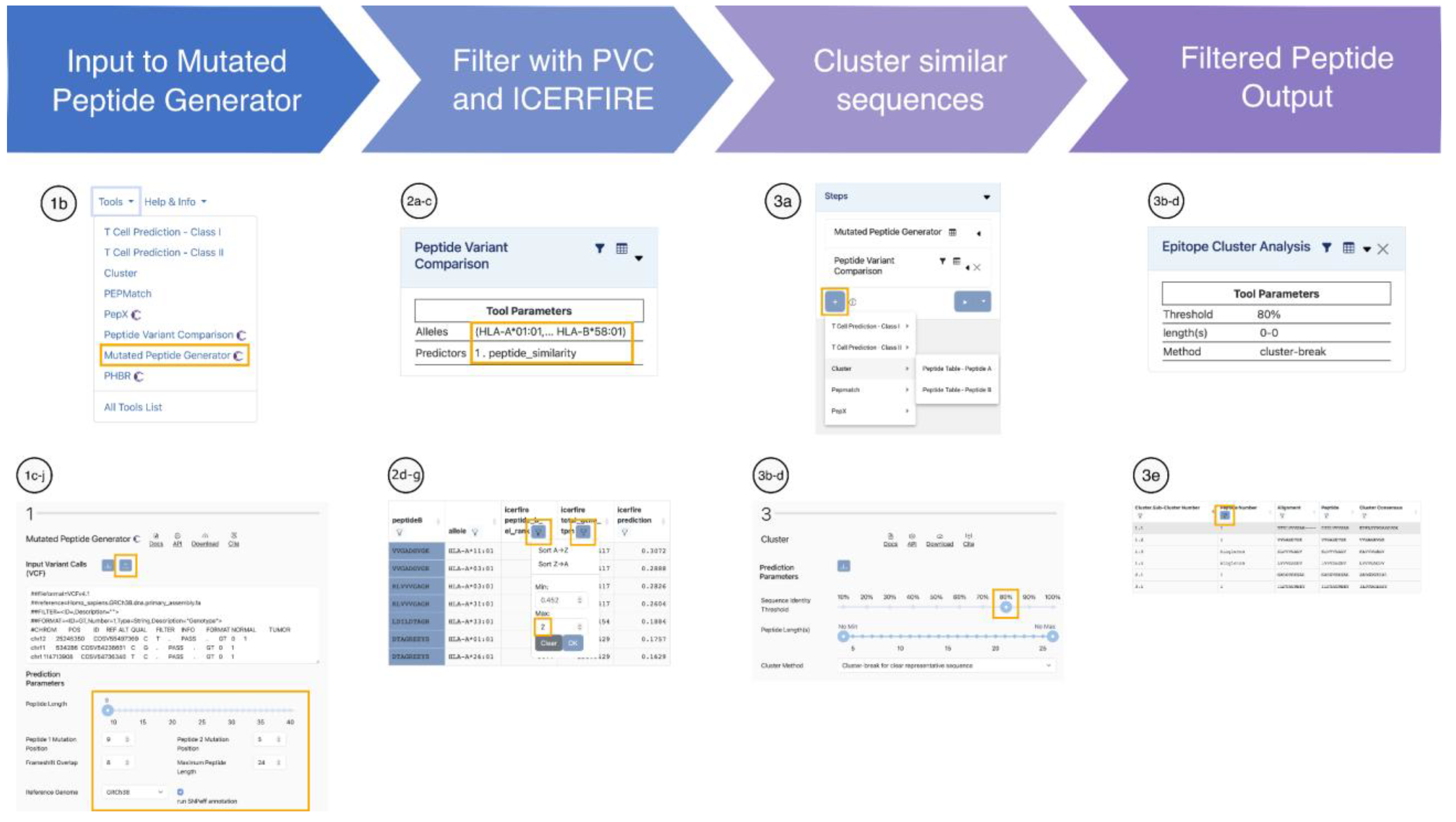
Mutated Peptide Generator (MPG)
Peptide Expression Annotation (PepX)
T Cell Prediction – Class I
T Cell Prediction – Class II
Peptide Variant Comparison (PVC)
PEPMatch
Patient Harmonic-Mean Best Rank (PHBR)
Clustering
Antigen eXpression based Epitope Likelihood-Function (AXEL-F)
Peptide Synthesis Score (PepSySco)
TCRMatch
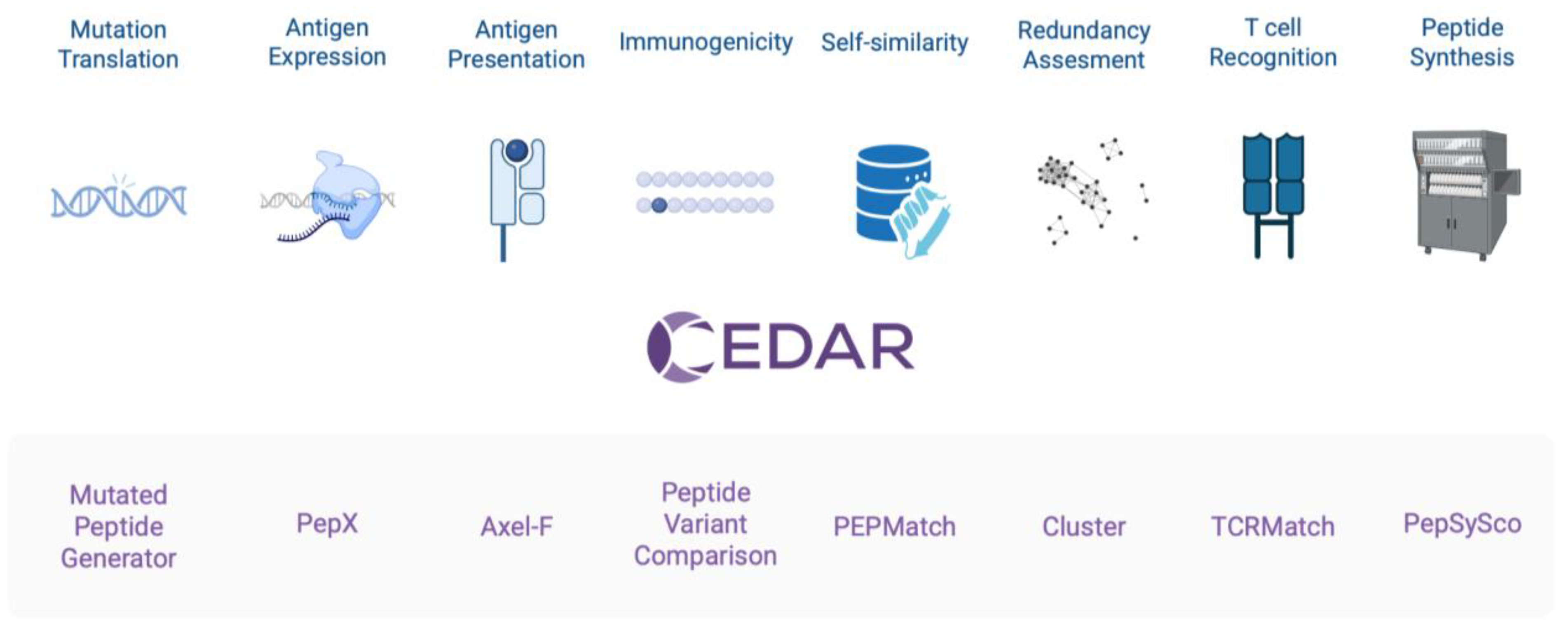
Pipeline Integration
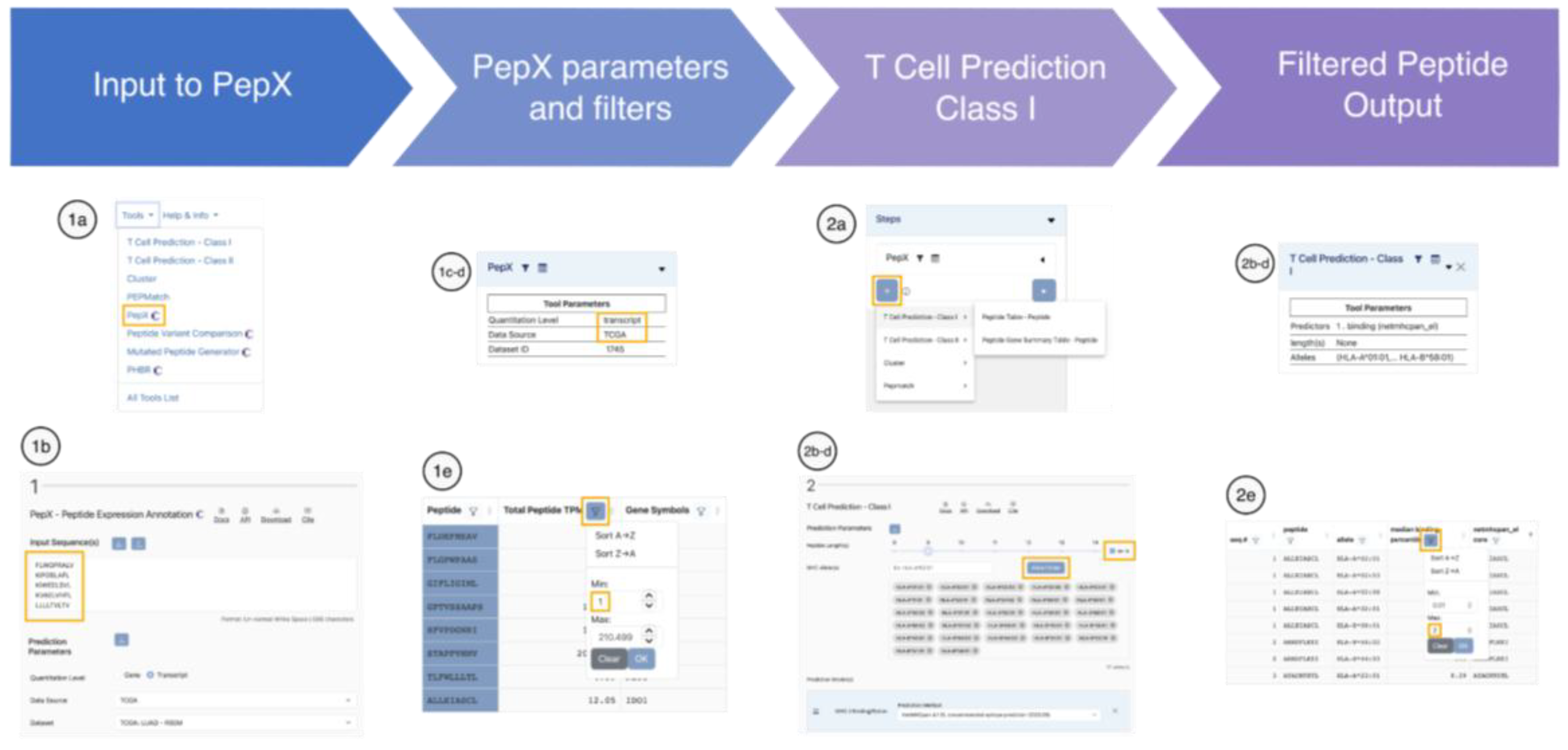
Case Scenario I: Expression-Based Filtering of Candidate Tumor-Associated Antigens in NSCLC
-
Use PepX to retrieve the expression levels of each peptide
- From the tools menu drop-down, select PepX.
- Paste the list of 9–11mer peptides derived from the tumor-associated proteins of interest into the input box.
- Under ‘Prediction Parameters’ and ‘Quantitation Level’ select ‘Transcript’.
- Under ‘Data Source’ select TCGA and under ‘Dataset’ select LUAD to retrieve malignancy-specific values and click on ‘Run’.
- In the results table, inspect the column ‘Total Transcript TPM’ and use the sciphon icon to filter out peptides that are not sufficiently expressed (e.g., select a minimum value of 1 TPM). Click on ‘Save Table State’.
-
Add MHC class I binding predictions
- On the left side, under ‘Pipeline Map’, click on ‘+’ and select ‘T Cell Prediction - Class I’ and select the Peptide Table. The T Cell Prediction - Class I interface appears below the PepX results.
- bSelect the Peptide Length ‘as-is’ by clicking the checkbox.
- By default, the predictions are run for HLA-A*02:01, however this selection can be changed. Click on the button ‘Allele Finder’ and in the pop-up window select ‘27 Allele Panel’. This MHC panel was developed to cover >97% of the population [48]. Click on ‘Submit’, which will close the pop-up and populate the ‘MHC Alleles’ field with the selected 27 alleles.
- Under ‘Prediction Model’ select ‘NetMHCpan 4.1 EL’, which is currently the recommended tool, and click on ‘Run’.
- In the results table, examine the column labeled ‘median binding percentile’. This number represents the percentile rank of the predicted peptide relative to a background of random natural peptides, avoiding biases arising from MHC alleles with inherently higher or lower predicted affinities. Peptides with lower ranks are more likely to be presented, with strong binders having scores ≤ 0.5 and weak binders with scores < 2 [33]. Use the sciphon icon to filter out peptides that are not predicted to bind.
- Download the final list of filtered peptides by clicking ‘Download’, ‘All rows’, in comma-separated format (CSV).
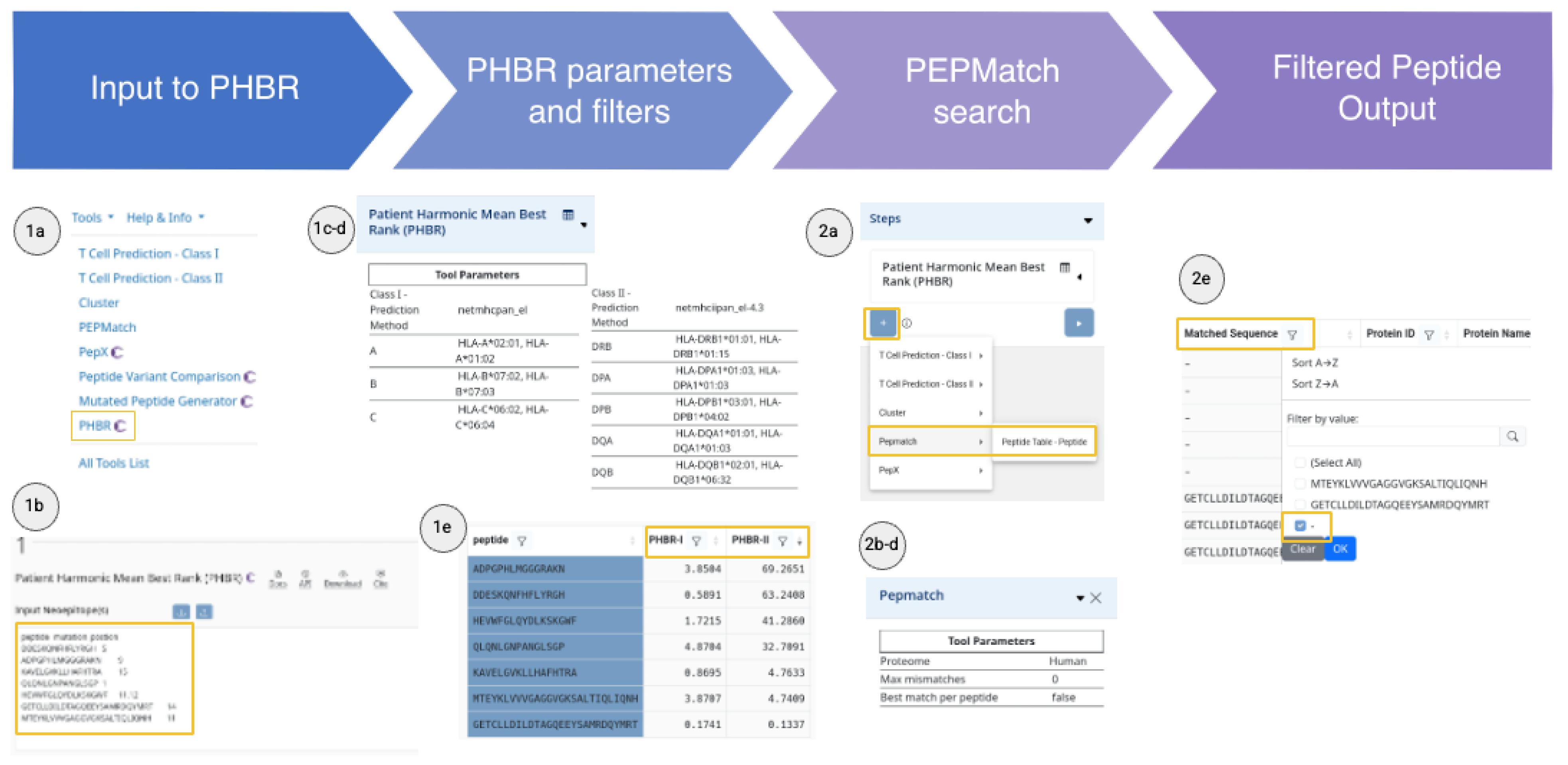
Case Scenario II: Neoepitope Discovery for Personalized Immunotherapy in Glioblastoma
-
Prioritize mutations likely to be presented in the patient’s MHC molecules with PHBR
- From the tools menu drop-down, select the Patient Harmonic Mean Best Rank (PHBR) tool.
- Paste the list of long mutated peptides (e.g., 21mers) with their corresponding mutation position(s) using a tab-delimiter from the sequence and use comma-separation for multiple positions.
- Select NetMHCpan 4.1 EL and NetMHCIIpan 4.3 EL models for MHC class I and II antigen presentation prediction, respectively.
- Introduce the MHC alleles of the patient. In the ‘Allele 1/2’ text box, type the patient’s MHC alleles, and a drop-down menu suggests alleles as you type. Repeat this for all the alleles and click on ‘Run’.
- Sort in ascending order by PHBR I and II, and select the top mutations with the lowest scores for either of the PHBR predictions.
-
Exclude self-peptides with PEPMatch
- On the left side, under ‘Pipeline Map’, click on ‘+’, select ‘PEPMatch’, and choose the ‘Peptide Table – Peptide’. The ‘PEPMatch’ interface appears below the PHBR results.
- Set mismatches to 0 using the slider.
- Select ‘All matches’.
- Check ‘Include unmatched peptides’ and click on ‘Run’.
- Select only the peptides not found in the human reference proteome. Use the sciphon icon in the ‘Matched Sequence’ column and select ‘-‘.
- Download the final list of filtered peptides by clicking ‘Download’ and ‘All rows’, in comma-separated format (CSV).
Conclusion and Discussion
Funding
Conflicts of Interest
References
- Karl Erik Hellström, IH. Tumor Antigens. In: Bertino JR, editor. Encyclopedia of Cancer (Second Edition). Academic Press; 2002. pp. 459–466.
- Jhunjhunwala, S.; Hammer, C.; Delamarre, L. Antigen presentation in cancer: insights into tumour immunogenicity and immune evasion. Nat. Rev. Cancer 2021, 21, 298–312. [Google Scholar] [CrossRef]
- Xie, N.; Shen, G.; Gao, W.; Huang, Z.; Huang, C.; Fu, L. Neoantigens: promising targets for cancer therapy. Signal Transduct. Target. Ther. 2023, 8, 1–38. [Google Scholar] [CrossRef] [PubMed]
- Peng, K.; Zhao, X.; Fu, Y.-X.; Liang, Y. Eliciting antitumor immunity via therapeutic cancer vaccines. Cell. Mol. Immunol. 2025, 22, 840–868. [Google Scholar] [CrossRef]
- Gopanenko, A.V.; Kosobokova, E.N.; Kosorukov, V.S. Main Strategies for the Identification of Neoantigens. Cancers 2020, 12, 2879. [Google Scholar] [CrossRef]
- Koşaloğlu-Yalçın, Z.; Lanka, M.; Frentzen, A.; Premlal, A.L.R.; Sidney, J.; Vaughan, K.; Greenbaum, J.; Robbins, P.; Gartner, J.; Sette, A.; et al. Predicting T cell recognition of MHC class I restricted neoepitopes. OncoImmunology 2018, 7, e1492508. [Google Scholar] [CrossRef]
- Sahin, U.; Derhovanessian, E.; Miller, M.; Kloke, B.-P.; Simon, P.; Löwer, M.; Bukur, V.; Tadmor, A.D.; Luxemburger, U.; Schrörs, B.; et al. Personalized RNA mutanome vaccines mobilize poly-specific therapeutic immunity against cancer. Nature 2017, 547, 222–226. [Google Scholar] [CrossRef]
- Peters, B.; Nielsen, M.; Sette, A. T Cell Epitope Predictions. Annu. Rev. Immunol. 2020, 38, 123–145. [Google Scholar] [CrossRef]
- Wan, Y.-T.R.; Koşaloğlu-Yalçın, Z.; Peters, B.; Nielsen, M. A large-scale study of peptide features defining immunogenicity of cancer neo-epitopes. NAR Cancer 2024, 6, zcae002. [Google Scholar] [CrossRef]
- Borch, A.; Carri, I.; Reynisson, B.; Alvarez, H.M.G.; Munk, K.K.; Montemurro, A.; Kristensen, N.P.; Tvingsholm, S.A.; Holm, J.S.; Heeke, C.; et al. IMPROVE: a feature model to predict neoepitope immunogenicity through broad-scale validation of T-cell recognition. Front. Immunol. 2024, 15, 1360281. [Google Scholar] [CrossRef]
- Koşaloğlu-Yalçın, Z.; Lee, J.; Greenbaum, J.; Schoenberger, S.P.; Miller, A.; Kim, Y.J.; Sette, A.; Nielsen, M.; Peters, B. Combined assessment of MHC binding and antigen abundance improves T cell epitope predictions. iScience 2022, 25, 103850. [Google Scholar] [CrossRef]
- Alvarez, H.M.G.; Koşaloğlu-Yalçın, Z.; Peters, B.; Nielsen, M. The role of antigen expression in shaping the repertoire of HLA presented ligands. iScience 2022, 25, 104975. [Google Scholar] [CrossRef]
- Sarkizova, S.; Klaeger, S.; Le, P.M.; Li, L.W.; Oliveira, G.; Keshishian, H.; Hartigan, C.R.; Zhang, W.; Braun, D.A.; Ligon, K.L.; et al. A large peptidome dataset improves HLA class I epitope prediction across most of the human population. Nat. Biotechnol. 2019, 38, 199–209. [Google Scholar] [CrossRef]
- 14. Vita R, Blazeska N, Marrama D, IEDB Curation Team Members, Duesing S, Bennett J, et al. The Immune Epitope Database (IEDB): 2024 update. Nucleic Acids Res. 2025;53: D436–D443.
- Vita, R.; Blazeska, N.; Marrama, D.; Members, I.C.T.; Shackelford, D.; Zalman, L.; Foos, G.; Zarebski, L.; Chan, K.; Reardon, B.; et al. The Immune Epitope Database (IEDB): 2024 update. Nucleic Acids Res. 2024, 53, D436–D443. [Google Scholar] [CrossRef] [PubMed]
- Koşaloğlu-Yalçın, Z.; Blazeska, N.; Vita, R.; Carter, H.; Nielsen, M.; Schoenberger, S.; Sette, A.; Peters, B. The Cancer Epitope Database and Analysis Resource (CEDAR). Nucleic Acids Res. 2022, 51, D845–D852. [Google Scholar] [CrossRef] [PubMed]
- Yan Z, Kim K, Kim H, Ha B, Gambiez A, Bennett J, et al. Next-generation IEDB tools: a platform for epitope prediction and analysis. Nucleic Acids Res. 2024;52: W526–W532.
- Yan, Z.; Kim, K.; Kim, H.; Ha, B.; Gambiez, A.; Bennett, J.; Mendes, M.F.d.A.; Trevizani, R.; Mahita, J.; Richardson, E.; et al. Next-generation IEDB tools: a platform for epitope prediction and analysis. Nucleic Acids Res. 2024, 52, W526–W532. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Chen, T.-T.; Li, X.; Lan, A.-L.; Ji, P.-F.; Zhu, Y.-J.; Ma, X.-Y. Advances and challenges in neoantigen prediction for cancer immunotherapy. Front. Immunol. 2025, 16, 1617654. [Google Scholar] [CrossRef]
- Reynisson, B.; Alvarez, B.; Paul, S.; Peters, B.; Nielsen, M. NetMHCpan-4.1 and NetMHCIIpan-4.0: improved predictions of MHC antigen presentation by concurrent motif deconvolution and integration of MS MHC eluted ligand data. Nucleic Acids Res. 2020, 48, W449–W454. [Google Scholar] [CrossRef]
- O’dOnnell, T.J.; Rubinsteyn, A.; Laserson, U. MHCflurry 2.0: Improved Pan-Allele Prediction of MHC Class I-Presented Peptides by Incorporating Antigen Processing. Cell Syst. 2020, 11, 418–419. [Google Scholar] [CrossRef]
- Koşaloğlu-Yalçın, Z.; Blazeska, N.; Carter, H.; Nielsen, M.; Cohen, E.; Kufe, D.; Conejo-Garcia, J.; Robbins, P.; Schoenberger, S.P.; Peters, B.; et al. The Cancer Epitope Database and Analysis Resource: A Blueprint for the Establishment of a New Bioinformatics Resource for Use by the Cancer Immunology Community. Front. Immunol. 2021, 12. [Google Scholar] [CrossRef]
- Dhanda, S.K.; Mahajan, S.; Paul, S.; Yan, Z.; Kim, H.; Jespersen, M.C.; Jurtz, V.; Andreatta, M.; A Greenbaum, J.; Marcatili, P.; et al. IEDB-AR: immune epitope database—analysis resource in 2019. Nucleic Acids Res. 2019, 47, W502–W506. [Google Scholar] [CrossRef]
- Chronister, W.D.; Crinklaw, A.; Mahajan, S.; Vita, R.; Koşaloğlu-Yalçın, Z.; Yan, Z.; Greenbaum, J.A.; Jessen, L.E.; Nielsen, M.; Christley, S.; et al. TCRMatch: Predicting T-Cell Receptor Specificity Based on Sequence Similarity to Previously Characterized Receptors. Front. Immunol. 2021, 12. [Google Scholar] [CrossRef]
- Gutman, I.; Gutman, R.; Sidney, J.; Chihab, L.; Mishto, M.; Liepe, J.; Chiem, A.; Greenbaum, J.; Yan, Z.; Sette, A.; et al. Predicting the Success of Fmoc-Based Peptide Synthesis. ACS Omega 2022, 7, 23771–23781. [Google Scholar] [CrossRef]
- Cingolani, P.; Platts, A.; Wang, L.L.; Coon, M.; Nguyen, T.; Wang, L.; Land, S.J.; Lu, X.; Ruden, D.M. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff. Fly 2012, 6, 80–92. [Google Scholar] [CrossRef] [PubMed]
- Frentzen, A.; Greenbaum, J.A.; Kim, H.; Peters, B.; Koşaloğlu-Yalçın, Z. Estimating tissue-specific peptide abundance from public RNA-Seq data. Front. Genet. 2023, 14, 1082168. [Google Scholar] [CrossRef] [PubMed]
- Cancer Genome Atlas Research Network, Weinstein JN, Collisson EA, Mills GB, Shaw KRM, Ozenberger BA, et al. The Cancer Genome Atlas Pan-Cancer analysis project. Nat Genet. 2013;45: 1113–1120.
- Carithers, L.J.; Moore, H.M. The Genotype-Tissue Expression (GTEx) Project. Biopreservation Biobanking 2015, 13, 307–308. [Google Scholar] [CrossRef]
- Uhlen, M.; Oksvold, P.; Fagerberg, L.; Lundberg, E.; Jonasson, K.; Forsberg, M.; Zwahlen, M.; Kampf, C.; Wester, K.; Hober, S.; et al. Towards a knowledge-based Human Protein Atlas. Nat. Biotechnol. 2010, 28, 1248–1250. [Google Scholar] [CrossRef]
- Ghandi M, Huang FW, Jané-Valbuena J, Kryukov GV, Lo CC, McDonald ER 3rd, et al. Next-generation characterization of the Cancer Cell Line Encyclopedia. Nature. 2019;569: 503–508.
- Moutaftsi, M.; Peters, B.; Pasquetto, V.; Tscharke, D.C.; Sidney, J.; Bui, H.-H.; Grey, H.; Sette, A. A consensus epitope prediction approach identifies the breadth of murine TCD8+-cell responses to vaccinia virus. Nat. Biotechnol. 2006, 24, 817–819. [Google Scholar] [CrossRef]
- Jurtz, V.; Paul, S.; Andreatta, M.; Marcatili, P.; Peters, B.; Nielsen, M. NetMHCpan-4.0: Improved Peptide–MHC Class I Interaction Predictions Integrating Eluted Ligand and Peptide Binding Affinity Data. J. Immunol. 2017, 199, 3360–3368. [Google Scholar] [CrossRef]
- Calis, J.J.A.; Maybeno, M.; Greenbaum, J.A.; Weiskopf, D.; De Silva, A.D.; Sette, A.; Keşmir, C.; Peters, B. Properties of MHC Class I Presented Peptides That Enhance Immunogenicity. PLOS Comput. Biol. 2013, 9, e1003266. [Google Scholar] [CrossRef]
- Larsen, M.V.; Lundegaard, C.; Lamberth, K.; Buus, S.; Brunak, S.; Lund, O.; Nielsen, M. An integrative approach to CTL epitope prediction: A combined algorithm integrating MHC class I binding, TAP transport efficiency, and proteasomal cleavage predictions. Eur. J. Immunol. 2005, 35, 2295–2303. [Google Scholar] [CrossRef]
- Stranzl, T.; Larsen, M.V.; Lundegaard, C.; Nielsen, M. NetCTLpan: pan-specific MHC class I pathway epitope predictions. Immunogenetics 2010, 62, 357–368. [Google Scholar] [CrossRef]
- Keşmir, C.; Nussbaum, A.K.; Schild, H.; Detours, V.; Brunak, S. Prediction of proteasome cleavage motifs by neural networks. Protein Eng. Des. Sel. 2002, 15, 287–296. [Google Scholar] [CrossRef] [PubMed]
- Paul, S.; Arlehamn, C.S.L.; Scriba, T.J.; Dillon, M.B.; Oseroff, C.; Hinz, D.; McKinney, D.M.; Pro, S.C.; Sidney, J.; Peters, B.; et al. Development and validation of a broad scheme for prediction of HLA class II restricted T cell epitopes. J. Immunol. Methods 2015, 422, 28–34. [Google Scholar] [CrossRef] [PubMed]
- Nilsson, J.B.; Kaabinejadian, S.; Yari, H.; Kester, M.G.D.; van Balen, P.; Hildebrand, W.H.; Nielsen, M. Accurate prediction of HLA class II antigen presentation across all loci using tailored data acquisition and refined machine learning. Sci. Adv. 2023, 9, eadj6367. [Google Scholar] [CrossRef]
- Dhanda, S.K.; Karosiene, E.; Edwards, L.; Grifoni, A.; Paul, S.; Andreatta, M.; Weiskopf, D.; Sidney, J.; Nielsen, M.; Peters, B.; et al. Predicting HLA CD4 Immunogenicity in Human Populations. Front. Immunol. 2018, 9, 1369. [Google Scholar] [CrossRef]
- Paul, S.; Karosiene, E.; Dhanda, S.K.; Jurtz, V.; Edwards, L.; Nielsen, M.; Sette, A.; Peters, B. Determination of a Predictive Cleavage Motif for Eluted Major Histocompatibility Complex Class II Ligands. Front. Immunol. 2018, 9, 1795. [Google Scholar] [CrossRef]
- Marrama, D.; Chronister, W.D.; Westernberg, L.; Vita, R.; Koşaloğlu-Yalçın, Z.; Sette, A.; Nielsen, M.; Greenbaum, J.A.; Peters, B. PEPMatch: a tool to identify short peptide sequence matches in large sets of proteins. BMC Bioinform. 2023, 24, 1–14. [Google Scholar] [CrossRef]
- Marty Pyke R, Thompson WK, Salem RM, Font-Burgada J, Zanetti M, Carter H. Evolutionary Pressure against MHC Class II Binding Cancer Mutations. Cell. 2018;175: 416–428.e13.
- Marty, R.; Kaabinejadian, S.; Rossell, D.; Slifker, M.J.; van de Haar, J.; Engin, H.B.; de Prisco, N.; Ideker, T.; Hildebrand, W.H.; Font-Burgada, J.; et al. MHC-I Genotype Restricts the Oncogenic Mutational Landscape. Cell 2017, 171, 1272–1283.e15. [Google Scholar] [CrossRef]
- Dhanda, S.K.; Vaughan, K.; Schulten, V.; Grifoni, A.; Weiskopf, D.; Sidney, J.; Peters, B.; Sette, A. Development of a novel clustering tool for linear peptide sequences. Immunology 2018, 155, 331–345. [Google Scholar] [CrossRef]
- Xiao, R.; Lin, M.; Liu, M.; Ma, Q. Single cells and TRUST4 reveal immunological features of the HFRS transcriptome. Front. Med. 2024, 11, 1403335. [Google Scholar] [CrossRef]
- Liu, Y.; Han, L. TRUST4 Interrogates the Immune Receptor Repertoire in Oncology and Immunotherapy. Cancer Immunol. Res. 2022, 10, 786–786. [Google Scholar] [CrossRef]
- Greenbaum, J.; Sidney, J.; Chung, J.; Brander, C.; Peters, B.; Sette, A. Functional classification of class II human leukocyte antigen (HLA) molecules reveals seven different supertypes and a surprising degree of repertoire sharing across supertypes. Immunogenetics 2011, 63, 325–335. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



