Submitted:
24 September 2025
Posted:
25 September 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Data Recording Details
2.1. Ethics Statement
2.2. Participants
- Early childhood (1–3 years): 2 patients (ages 2 and 3),
- Preschool age (4–6 years): 3 patients (ages 4, 5, and 6),
- Primary school age (7–11 years): 6 patients (ages 7, 8, 8, 8, 8, and 10),
- Adolescence (12–18 years): 1 patient (age 12).
2.3. Data Acquisition and Automatic Preprocessing by Neuroscope Software
- Sweep speed: 30 mm/s
- Sensitivity: 7–15 V/mm
- Low-frequency cutoff: 0.5 Hz
- High-frequency cutoff: 70 Hz
- Notch filter: 50 Hz (to suppress power-line interference)
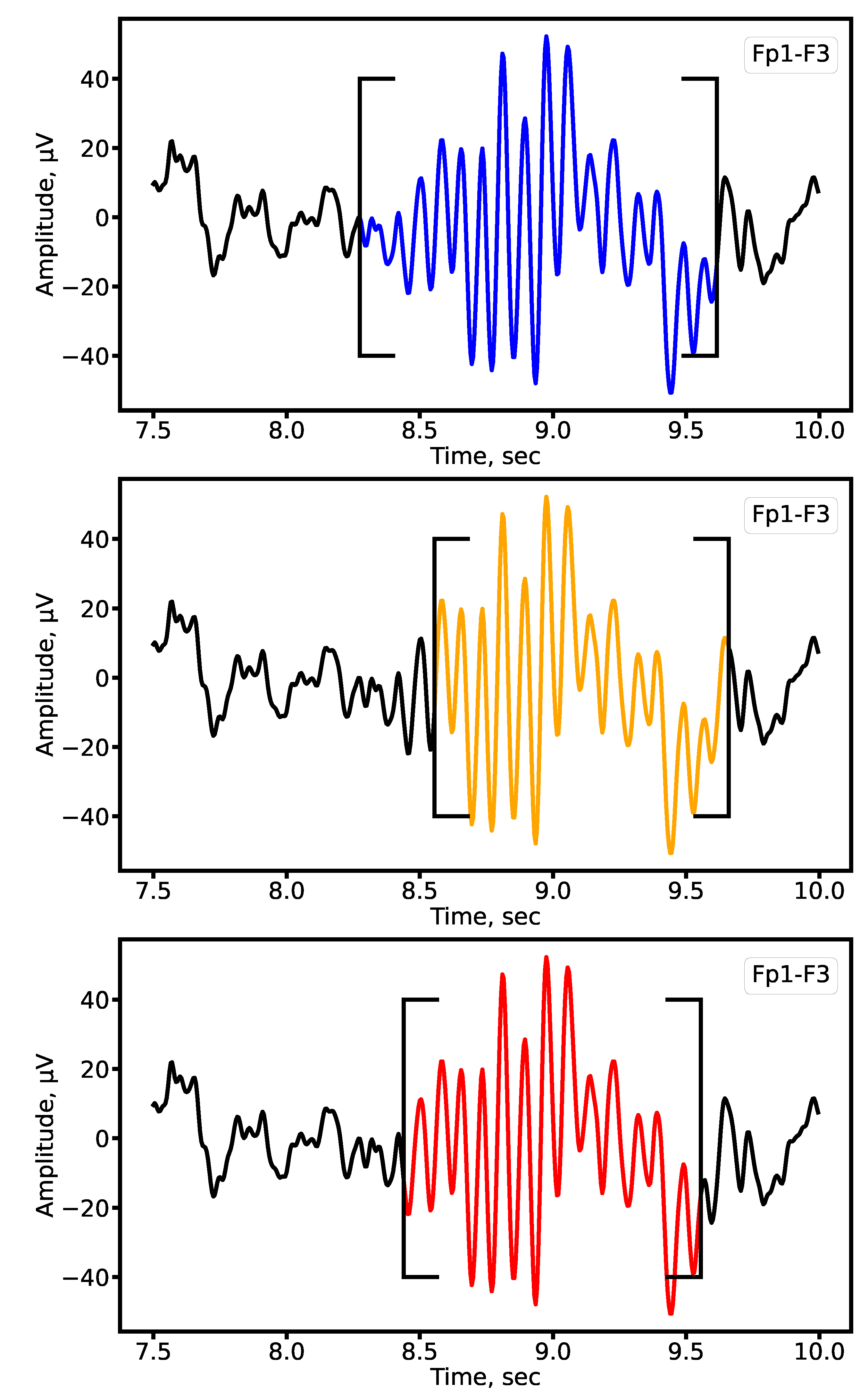
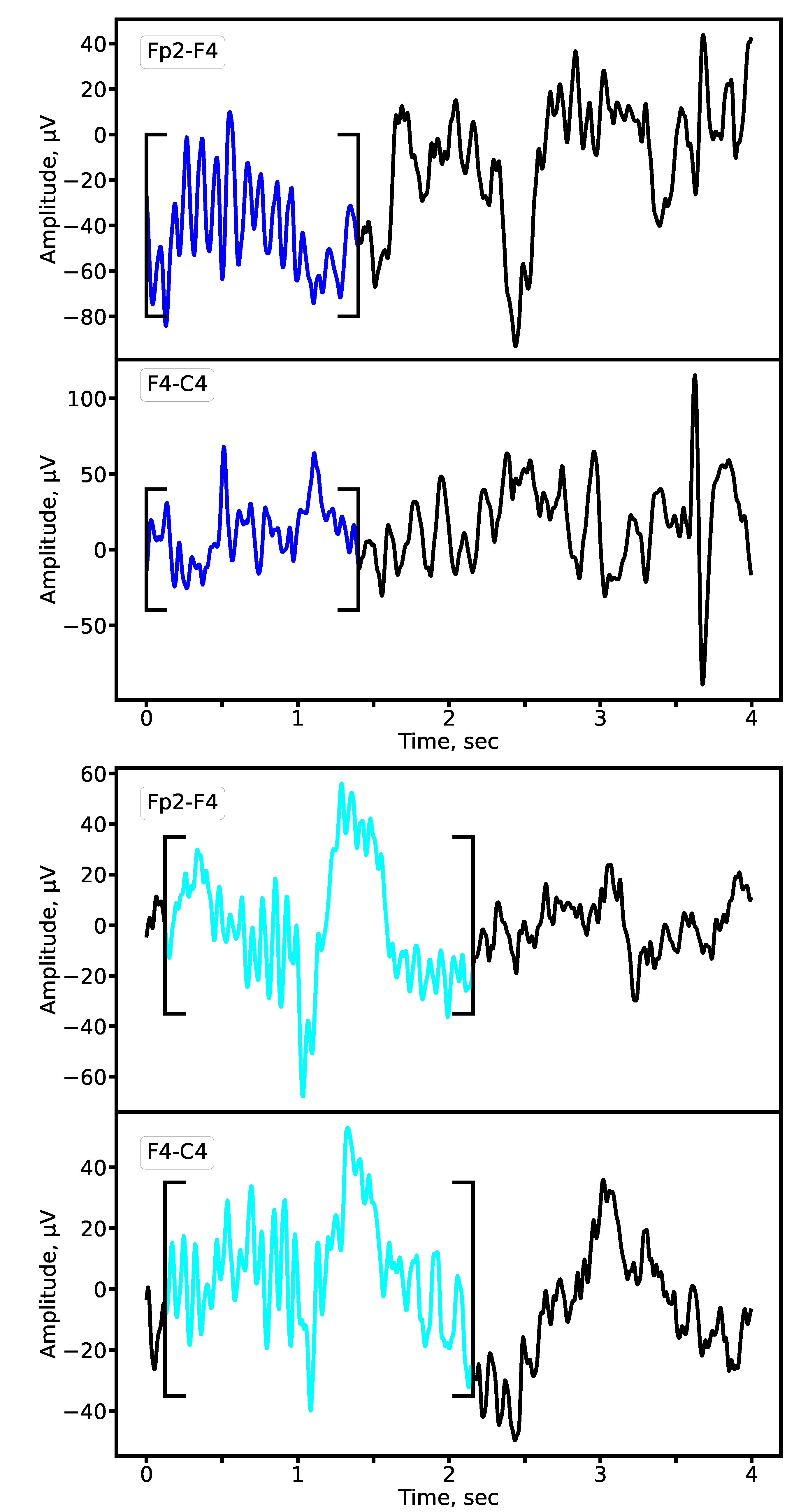
2.4. Sleep Spindle Annotation
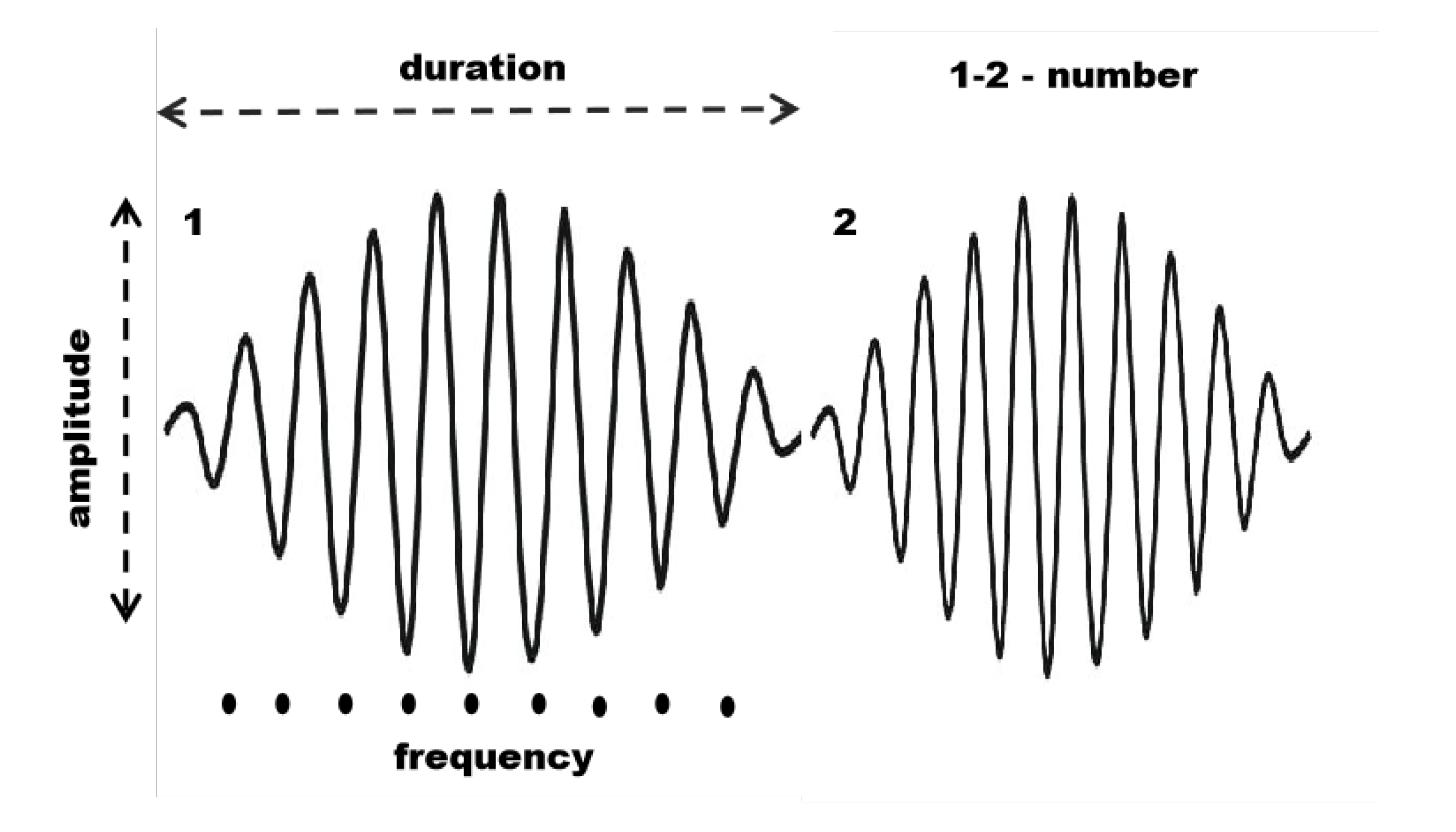
- Definite spindles (SP): amplitude V, duration s, located in fronto-central and vertex regions. SP serve as high-confidence “anchor” examples, offering clean prototypes of frequency, temporal profile, and waveform shape. They reduce noise in the training set and ensure stable learning during early stages.
- Probable spindles (PS): amplitude V, duration s, located in vertex and partially fronto-temporal regions. PS capture borderline cases between genuine spindles and other oscillations (e.g., partial arousals, artifacts). Including these examples improves the model’s ability to generalize and reduces overfitting to only idealized patterns.
- Dubious spindles (DS): amplitude 10–15 V or duration s, isolated in frontal or vertex regions, with unclear morphology. DS act as challenging negative or weakly positive examples. They train the network to reject artifacts and ambiguous oscillations, reflecting the complexity of real-world EEG recordings, particularly in epilepsy. Balanced sampling of these categories across training batches prevents overrepresentation of any one class and improves gradient stability.
2.5. EEG Data Sets Containing Sleep Spindles
2.6. The Montreal Archive of Sleep Studies (MASS)
3. Methodology
3.1. Data Preprocessing for Sleep Spindle Analysis
3.2. Sleep Spindle Characteristics
3.3. Performance of DNN Architectures on Pathological Spindles
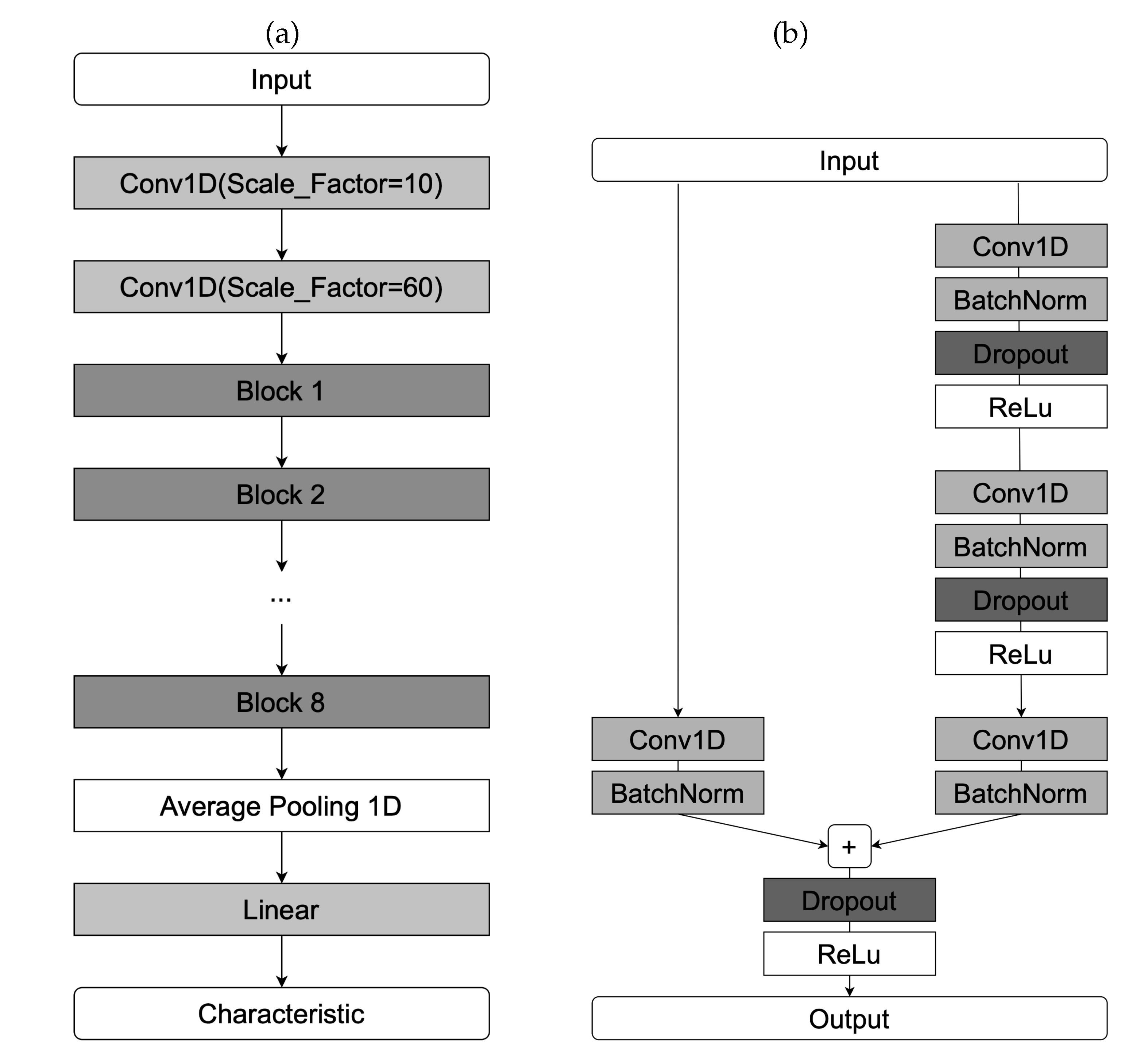
3.3.1. SlumberNet Architecture
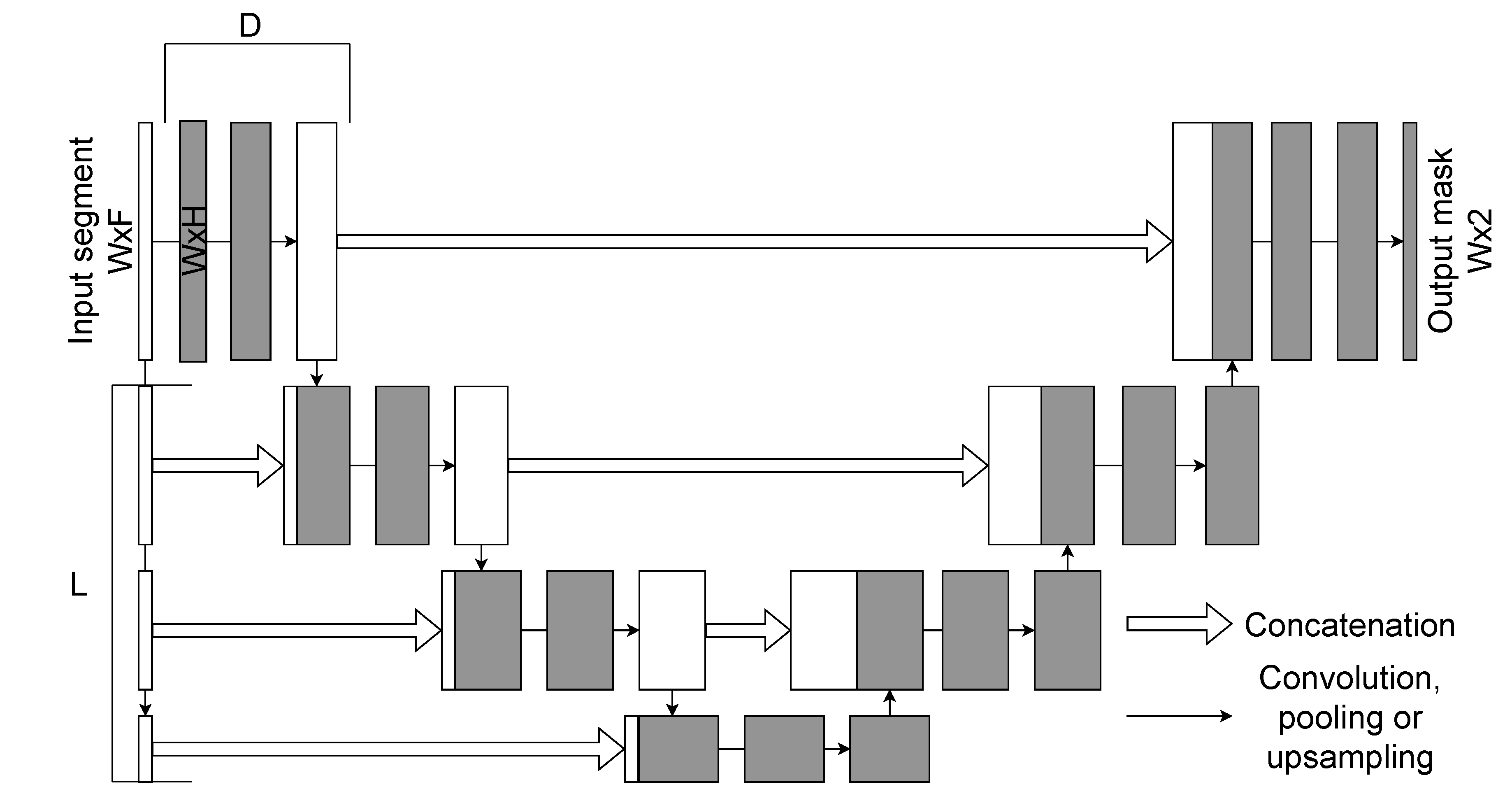
3.3.2. U-Net Architecture
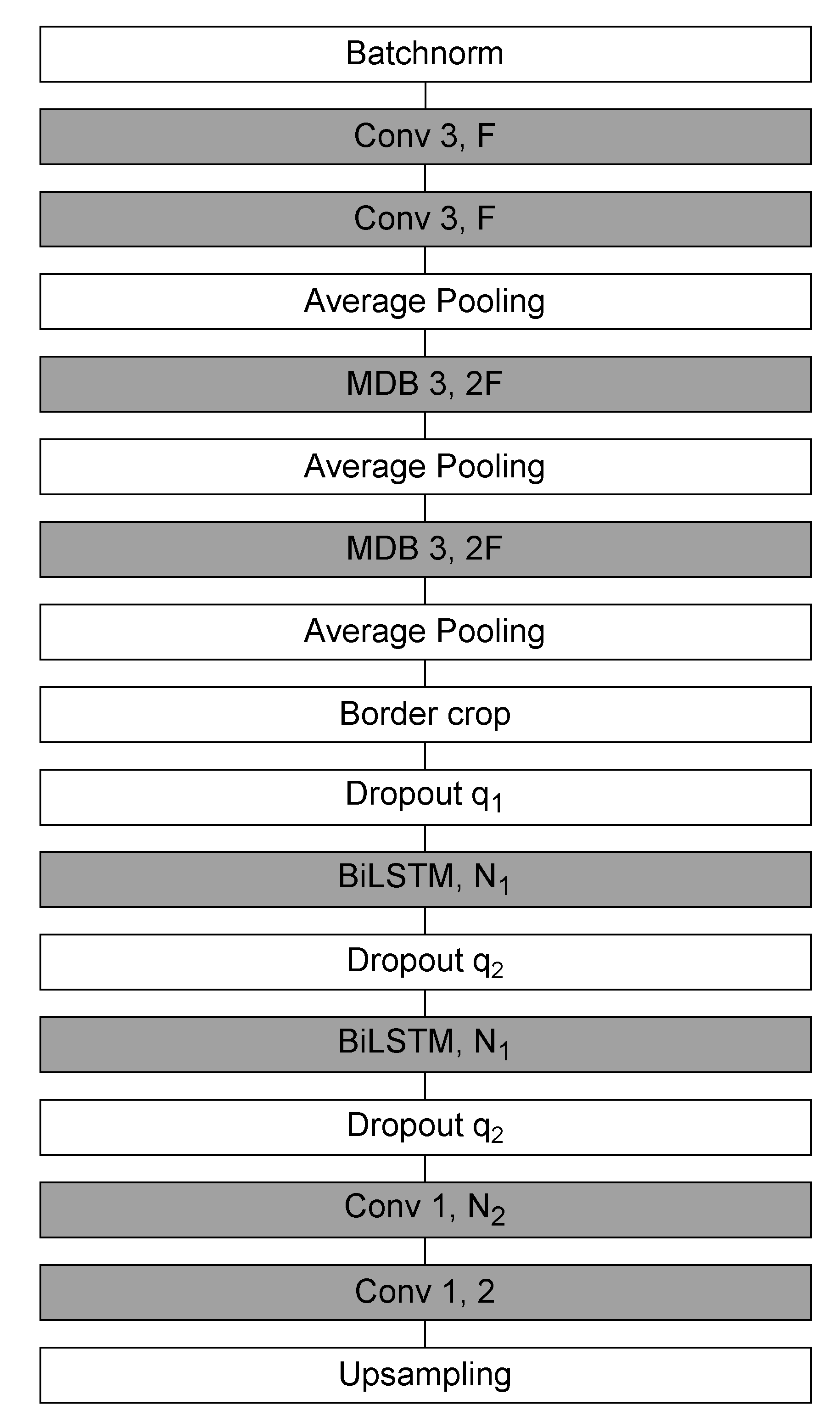
3.4. Sleep EEG Event Detector
3.5. Training Process and Evaluation Metrics
3.5.1. Segmentation-Based vs. Segmentation-Free Approaches
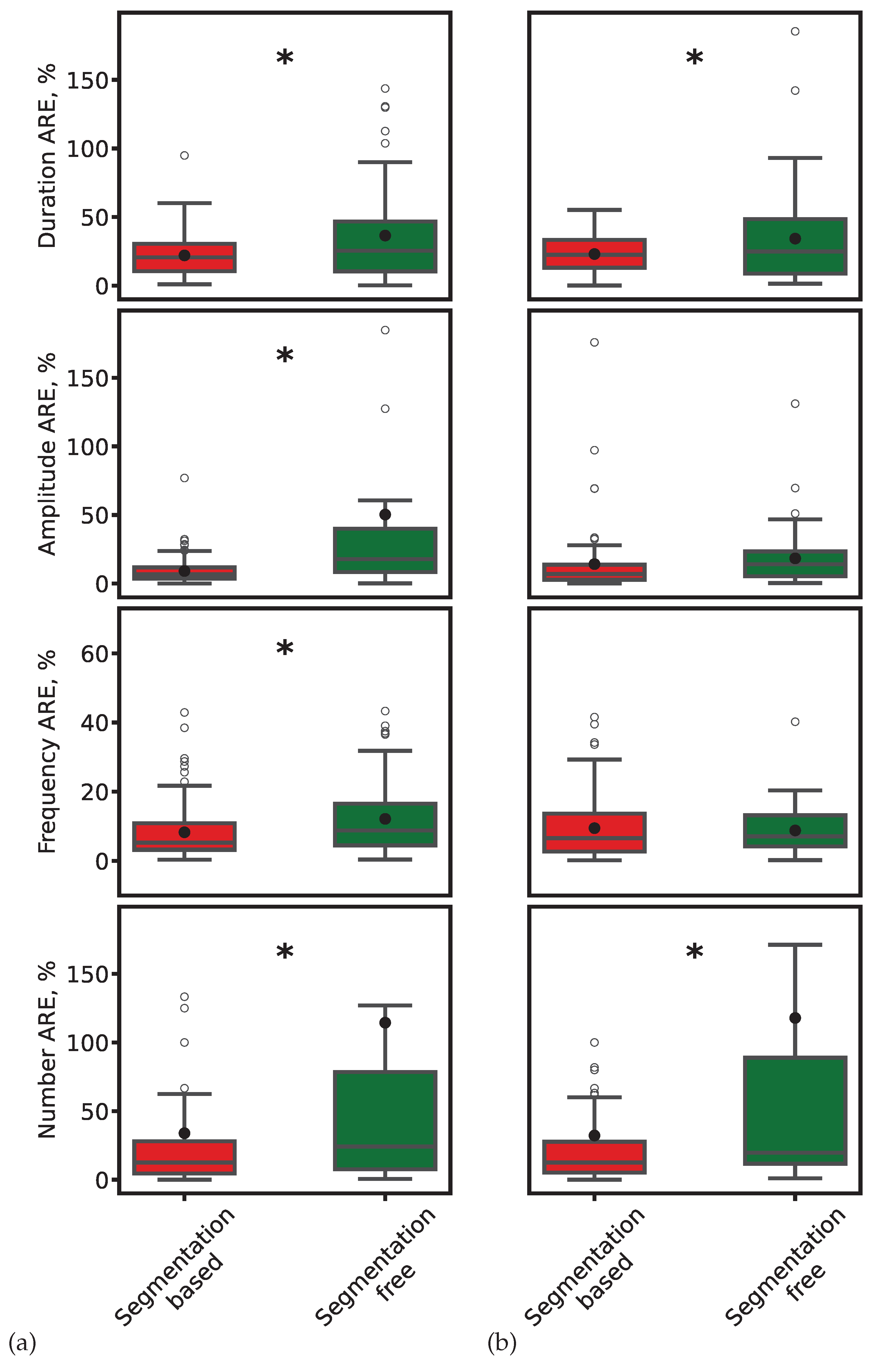
- Segmentation-based approach: Sleep spindles were first detected using DNN models in a sequence labeling framework, where each EEG time point was classified as spindle or non-spindle. Spindle characteristics were then computed from the resulting segmentation.
- Segmentation-free approach: Spindle characteristics were predicted directly from raw EEG segments using regression models, without an intermediate segmentation step.
3.5.2. Loss Functions
3.5.3. Evaluation Metrics
3.5.4. Hyperparameter Optimization
4. Results
4.1. Comparison of DL Architectures for Sleep Spindle Segmentation
4.2. Comparison of Segmentation-based and Segmentation-Free Approaches
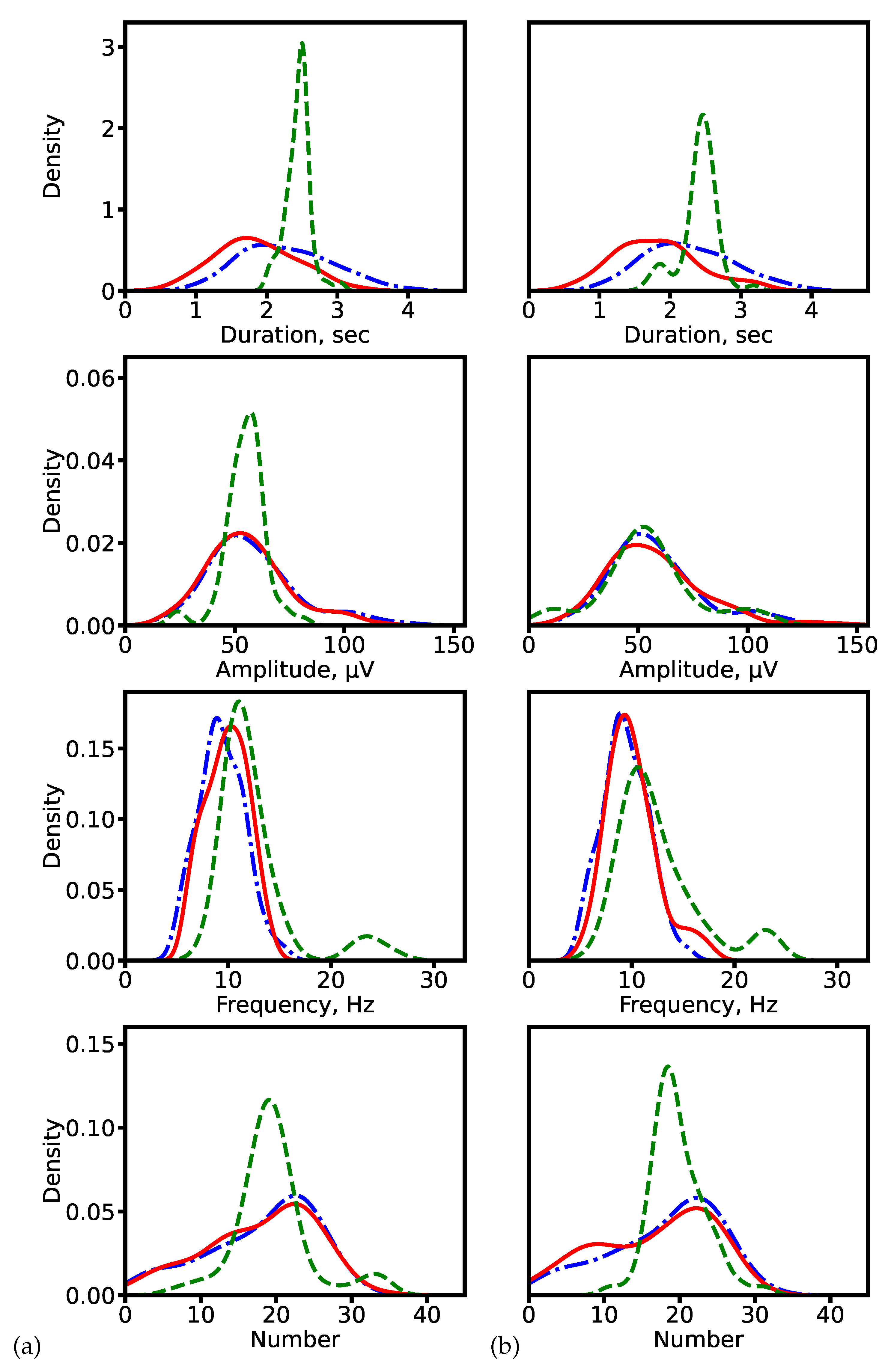
4.2.1. Distribution Analysis
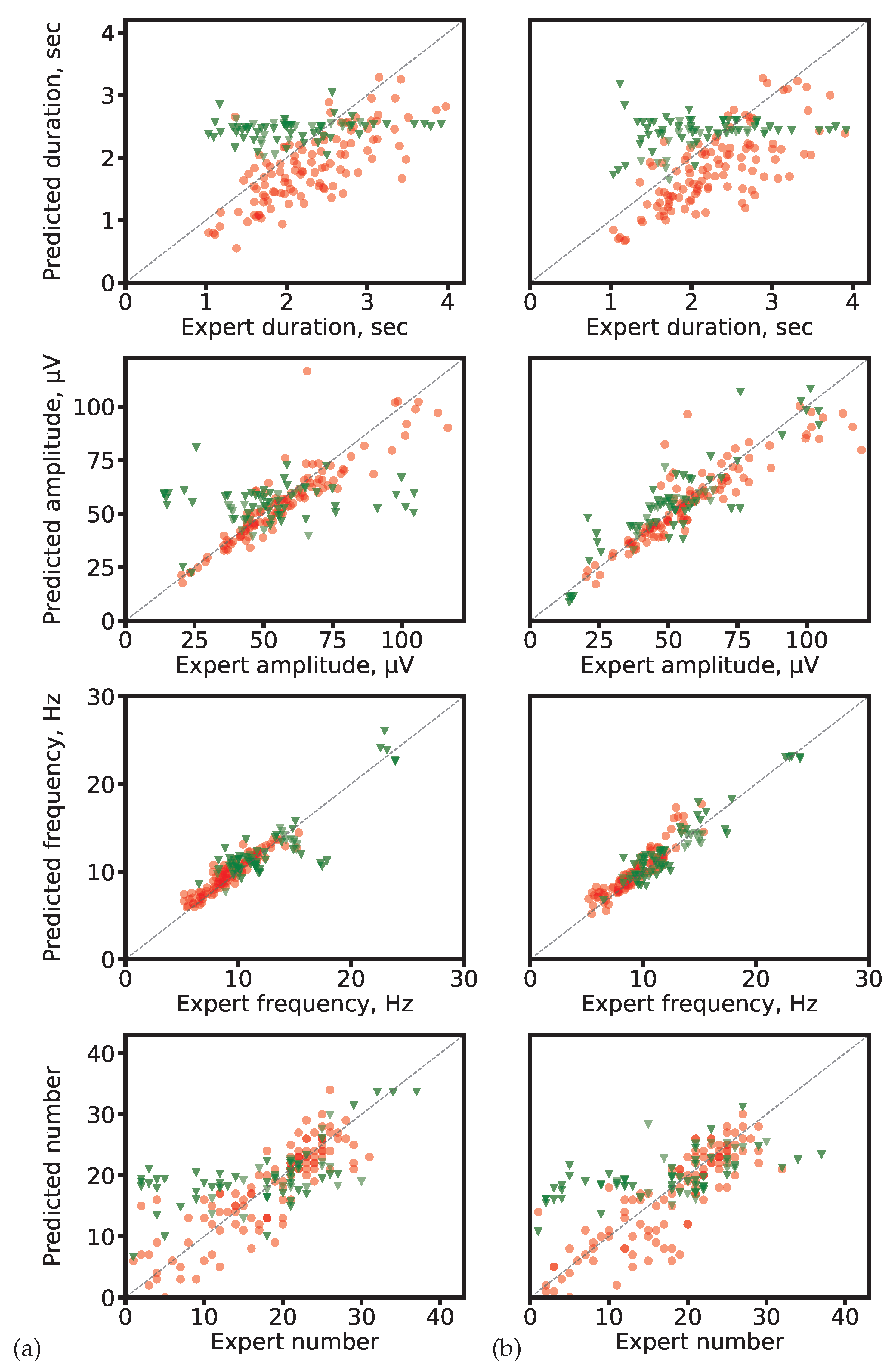
4.2.2. Prediction Accuracy
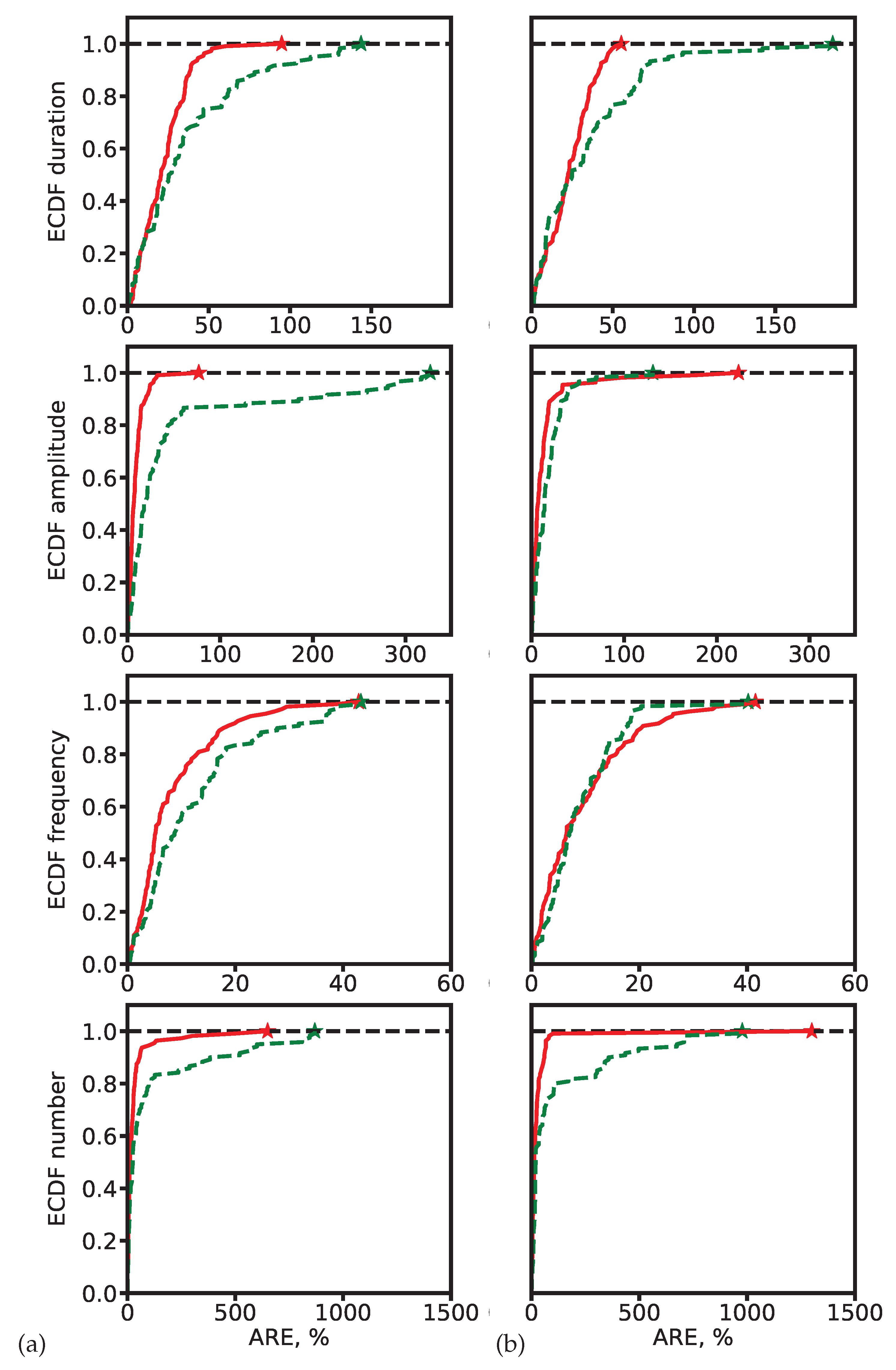
4.2.3. Error Analysis
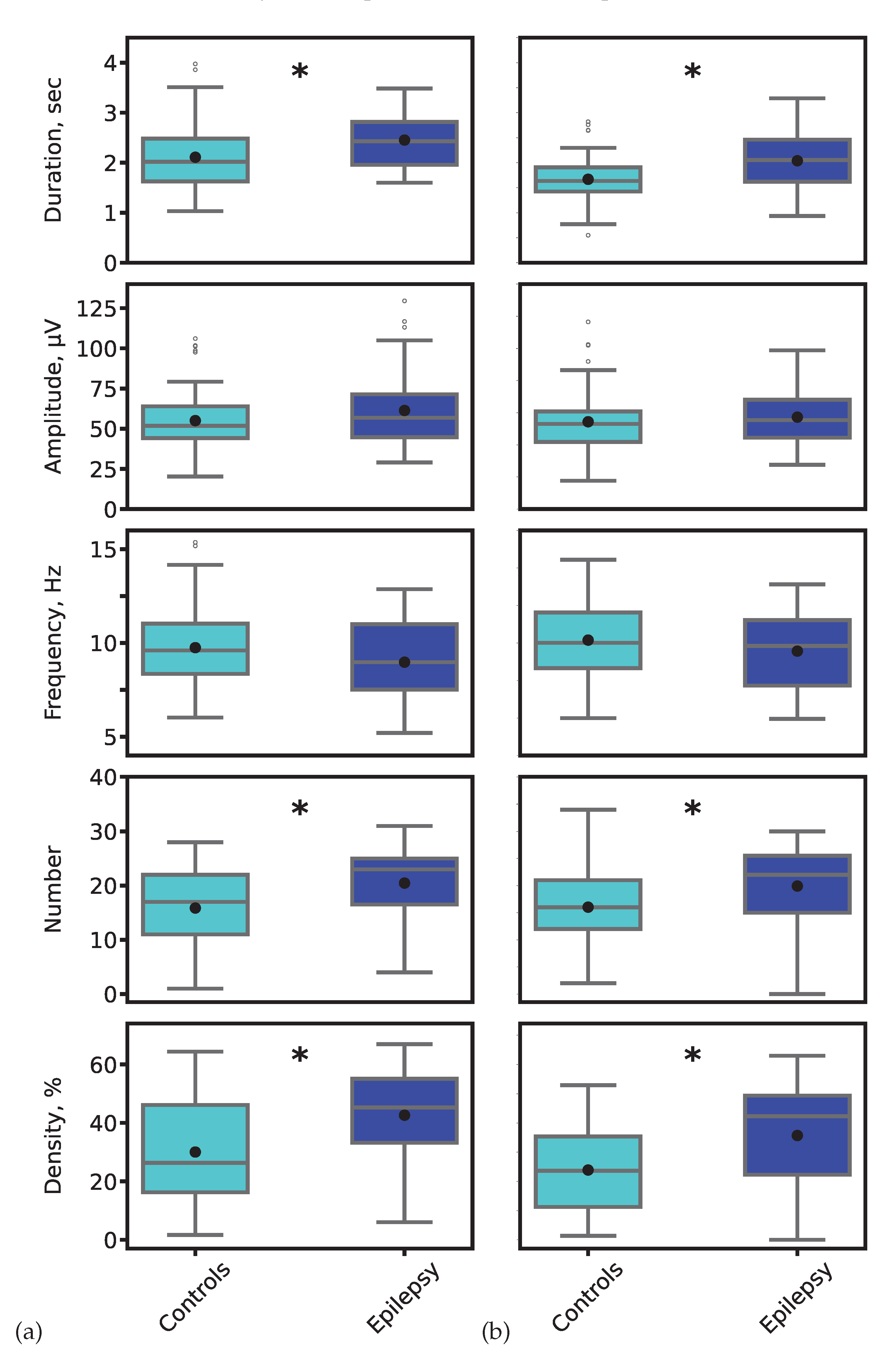
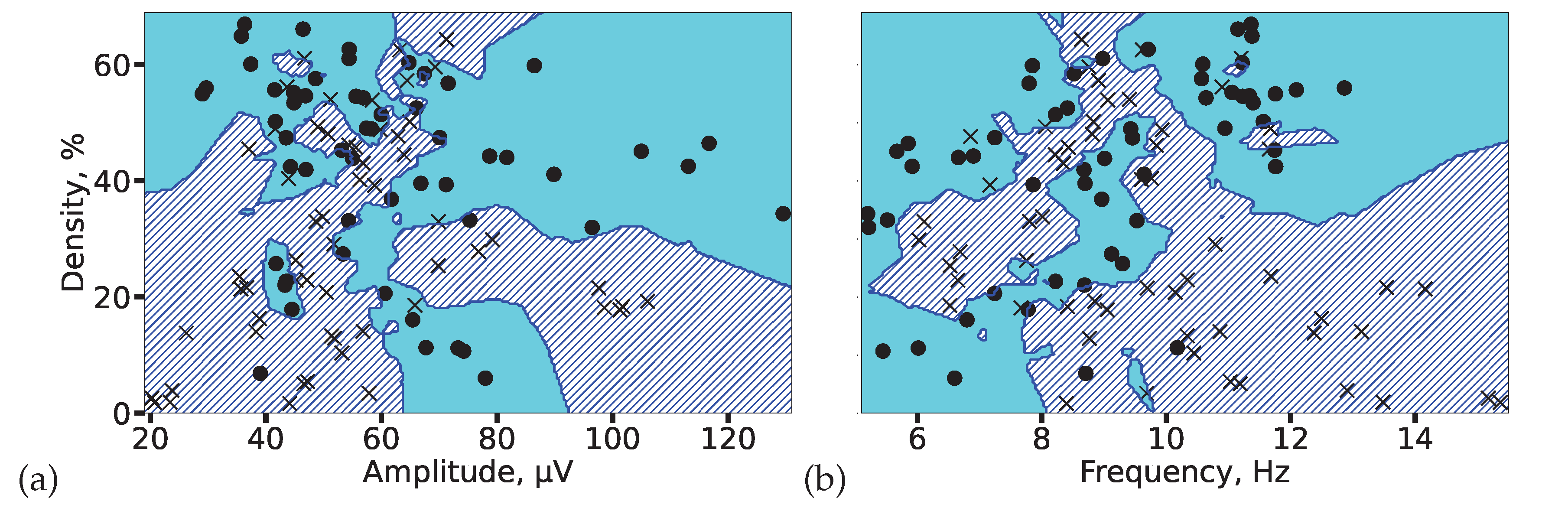
4.3. Alterations in Sleep Spindle Properties in Established Epilepsy
5. Discussion
5.1. Clinical Model Selection Framework
- High-Precision Option (e.g., SEED): Prioritizes minimizing false positives. This is advantageous in diagnostic contexts, where misclassifying epileptiform discharges or noise as spindles can corrupt biomarker quantification and lead to misleading conclusions. In this setting, SEED provides high confidence in detected spindles.
- High-Recall Option (e.g., 1D U-Net): Prioritizes capturing all true spindles, tolerating more false positives. This is valuable for screening or longitudinal monitoring, where the cost of missing altered spindle activity outweighs the burden of reviewing additional candidate events.
5.2. Synthesis of Challenges and Contributions
- Limited annotated clinical data. We mitigate this barrier by introducing a curated dataset of sleep spindles in pediatric epilepsy, showing that DNNs remain effective despite pathological spindle morphology. This resource provides a foundation for future model development and benchmarking in clinical populations.
- Building reliable clinical pipelines. We demonstrate and validate robust spindle segmentation as a key pipeline component. Automating this step reduces expert workload, improves reproducibility, and enables integration of spindle analysis into diagnostic and monitoring workflows.
5.3. The Critical Data Gap in Epilepsy Sleep Research
6. Conclusions and Future Directions
- (i)
- Dataset Expansion: Curating a larger, multi-center dataset encompassing diverse epilepsy syndromes and age groups to enhance model generalizability and robustness.
- (ii)
- Clinical Translation: Developing real-time detection algorithms and integrating spindle analysis with other EEG biomarkers (e.g., slow waves, epileptiform discharges) into a unified clinical dashboard to aid diagnosis and monitor therapy response.
- (iii)
- Validation: The essential next step towards clinical implementation is a rigorous external validation of our models and proposed pipeline on a completely independent patient cohort to confirm their efficacy and reliability.
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Loomis, A.L.; Harvey, E.N.; Hobart, G. Potential rhythms of the cerebral cortex during sleep. Science 1935, 81, 597–598. [Google Scholar] [CrossRef] [PubMed]
- Fernandez, L.M.; Lüthi, A. Sleep spindles: mechanisms and functions. Physiological reviews 2020, 100, 805–868. [Google Scholar] [CrossRef] [PubMed]
- Schönauer, M.; Pöhlchen, D. Sleep spindles. Current biology 2018, 28, R1129–R1130. [Google Scholar] [CrossRef] [PubMed]
- Schabus, M.; Hoedlmoser, K.; Pecherstorfer, T.; Anderer, P.; Gruber, G.; Parapatics, S.; Sauter, C.; Kloesch, G.; Klimesch, W.; Saletu, B.; et al. Interindividual sleep spindle differences and their relation to learning-related enhancements. Brain research 2008, 1191, 127–135. [Google Scholar] [CrossRef]
- Kumral, D.; Matzerath, A.; Leonhart, R.; Schönauer, M. Spindle-dependent memory consolidation in healthy adults: A meta-analysis. Neuropsychologia 2023, 189, 108661. [Google Scholar] [CrossRef]
- Staresina, B.P. Coupled sleep rhythms for memory consolidation. Trends in Cognitive Sciences 2024, 28, 339–351. [Google Scholar] [CrossRef]
- Dehnavi, F.; Koo-Poeggel, P.C.; Ghorbani, M.; Marshall, L. Memory ability and retention performance relate differentially to sleep depth and spindle type. Iscience 2023, 26. [Google Scholar] [CrossRef]
- Hahn, M.; Joechner, A.K.; Roell, J.; Schabus, M.; Heib, D.P.; Gruber, G.; Peigneux, P.; Hoedlmoser, K. Developmental changes of sleep spindles and their impact on sleep-dependent memory consolidation and general cognitive abilities: A longitudinal approach. Developmental science 2019, 22, e12706. [Google Scholar] [CrossRef]
- Champetier, P.; André, C.; Weber, F.D.; Rehel, S.; Ourry, V.; Laniepce, A.; Lutz, A.; Bertran, F.; Cabé, N.; Pitel, A.L.; et al. Age-related changes in fast spindle clustering during non-rapid eye movement sleep and their relevance for memory consolidation. Sleep 2023, 46, zsac282. [Google Scholar] [CrossRef]
- Reynolds, C.; Short, M.; Gradisar, M. Sleep spindles and cognitive performance across adolescence: A meta-analytic review. Journal of adolescence 2018, 66, 55–70. [Google Scholar] [CrossRef]
- Bernard, C.; Frauscher, B.; Gelinas, J.; Timofeev, I. Sleep, oscillations, and epilepsy. Epilepsia 2023, 64, S3–S12. [Google Scholar] [CrossRef]
- Sheybani, L.; Frauscher, B.; Bernard, C.; Walker, M.C. Mechanistic insights into the interaction between epilepsy and sleep. Nature Reviews Neurology 2025, pp. 1–16.
- Abdelaal, M.S.; Kato, T.; Natsubori, A.; Tanaka, K.F. Temporal and Potential Predictive Relationships between Sleep Spindle Density and Spike-and-Wave Discharges. Eneuro 2024, 11. [Google Scholar] [CrossRef]
- Hirsch, E.; French, J.; Scheffer, I.E.; Bogacz, A.; Alsaadi, T.; Sperling, M.R.; Abdulla, F.; Zuberi, S.M.; Trinka, E.; Specchio, N.; et al. ILAE definition of the idiopathic generalized epilepsy syndromes: position statement by the ILAE task force on nosology and definitions. Epilepsia 2022, 63, 1475–1499. [Google Scholar] [CrossRef]
- Kramer, M.A.; Stoyell, S.M.; Chinappen, D.; Ostrowski, L.M.; Spencer, E.R.; Morgan, A.K.; Emerton, B.C.; Jing, J.; Westover, M.B.; Eden, U.T.; et al. Focal sleep spindle deficits reveal focal thalamocortical dysfunction and predict cognitive deficits in sleep activated developmental epilepsy. Journal of Neuroscience 2021, 41, 1816–1829. [Google Scholar] [CrossRef]
- Roebber, J.K.; Lewis, P.A.; Crunelli, V.; Navarrete, M.; Hamandi, K. Effects of anti-seizure medication on sleep spindles and slow waves in drug-resistant epilepsy. Brain Sciences 2022, 12, 1288. [Google Scholar] [CrossRef]
- Gonzalez, C.; Jiang, X.; Gonzalez-Martinez, J.; Halgren, E. Human spindle variability. Journal of Neuroscience 2022, 42, 4517–4537. [Google Scholar] [CrossRef] [PubMed]
- Ujma, P.P. Sleep spindles and general cognitive ability–A meta-analysis. Sleep Spindles & Cortical Up States 2021, 2, 1–17. [Google Scholar]
- Bhattacharyya, S.; Ghoshal, S.; Biswas, A.; Mukhopadhyay, J.; Majumdar, A.K.; Majumdar, B.; Mukherjee, S.; Singh, A.K. Automatic sleep spindle detection in raw EEG signal of newborn babies. In Proceedings of the 2011 3rd International Conference on Electronics Computer Technology. IEEE, Vol. 1; 2011; pp. 73–77. [Google Scholar]
- Tsanas, A.; Clifford, G.D. Stage-independent, single lead EEG sleep spindle detection using the continuous wavelet transform and local weighted smoothing. Frontiers in human neuroscience 2015, 9, 181. [Google Scholar] [CrossRef] [PubMed]
- Patti, C.R.; Shahrbabaki, S.S.; Dissanayaka, C.; Cvetkovic, D. Application of random forest classifier for automatic sleep spindle detection. In Proceedings of the 2015 IEEE Biomedical Circuits and Systems Conference (BioCAS). IEEE; 2015; pp. 1–4. [Google Scholar]
- Wei, L.; Ventura, S.; Lowery, M.; Ryan, M.A.; Mathieson, S.; Boylan, G.B.; Mooney, C. Random forest-based algorithm for sleep spindle detection in infant EEG. In Proceedings of the 2020 42nd Annual International Conference of the IEEE Engineering in Medicine & Biology Society (EMBC). IEEE, 2020, pp. 58–61.
- Wei, L.; Ventura, S.; Mathieson, S.; Boylan, G.B.; Lowery, M.; Mooney, C. Spindle-AI: sleep spindle number and duration estimation in infant EEG. IEEE Transactions on Biomedical Engineering 2021, 69, 465–474. [Google Scholar] [CrossRef]
- Yasuhara, N.; Natori, T.; Hayashi, M.; Aikawa, N. A study on automatic detection of sleep spindles using a long short-term memory network. In Proceedings of the 2019 IEEE 62nd international midwest symposium on circuits and systems (MWSCAS). IEEE; 2019; pp. 45–48. [Google Scholar]
- Kulkarni, P.M.; Xiao, Z.; Robinson, E.J.; Jami, A.S.; Zhang, J.; Zhou, H.; Henin, S.E.; Liu, A.A.; Osorio, R.S.; Wang, J.; et al. A deep learning approach for real-time detection of sleep spindles. Journal of neural engineering 2019, 16, 036004. [Google Scholar] [CrossRef]
- Tan, D.; Zhao, R.; Sun, J.; Qin, W. Sleep spindle detection using deep learning: A validation study based on crowdsourcing. In Proceedings of the 2015 37th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC). IEEE; 2015; pp. 2828–2831. [Google Scholar]
- Devuyst, S.; Dutoit, T.; Stenuit, P.; Kerkhofs, M. Automatic sleep spindles detection—overview and development of a standard proposal assessment method. In Proceedings of the 2011 Annual international conference of the IEEE engineering in medicine and biology society. IEEE; 2011; pp. 1713–1716. [Google Scholar]
- Quan, S.F.; Howard, B.V.; Iber, C.; Kiley, J.P.; Nieto, F.J.; O’Connor, G.T.; Rapoport, D.M.; Redline, S.; Robbins, J.; Samet, J.M.; et al. The sleep heart health study: design, rationale, and methods. Sleep 1997, 20, 1077–1085. [Google Scholar] [CrossRef]
- Kemp, B.; Zwinderman, A.H.; Tuk, B.; Kamphuisen, H.A.; Oberye, J.J. Analysis of a sleep-dependent neuronal feedback loop: the slow-wave microcontinuity of the EEG. IEEE Transactions on Biomedical Engineering 2000, 47, 1185–1194. [Google Scholar] [CrossRef]
- O’reilly, C.; Gosselin, N.; Carrier, J.; Nielsen, T. Montreal Archive of Sleep Studies: an open-access resource for instrument benchmarking and exploratory research. Journal of sleep research 2014, 23, 628–635. [Google Scholar] [CrossRef] [PubMed]
- Sun, C.; Jing, J.; Turley, N.; Alcott, C.; Kang, W.Y.; Cole, A.J.; Goldenholz, D.M.; Lam, A.; Amorim, E.; Chu, C.; et al. Harvard Electroencephalography Database: A comprehensive clinical electroencephalographic resource from four Boston hospitals. Epilepsia 2025. [Google Scholar] [CrossRef] [PubMed]
- Al-Salman, W.; Li, Y.; Wen, P. Detecting sleep spindles in EEGs using wavelet fourier analysis and statistical features. Biomedical Signal Processing and Control 2019, 48, 80–92. [Google Scholar] [CrossRef]
- Adamczyk, M.; Genzel, L.; Dresler, M.; Steiger, A.; Friess, E. Automatic sleep spindle detection and genetic influence estimation using continuous wavelet transform. Frontiers in human neuroscience 2015, 9, 624. [Google Scholar] [CrossRef] [PubMed]
- Sitnikova, E.; Hramov, A.E.; Koronovsky, A.A.; van Luijtelaar, G. Sleep spindles and spike–wave discharges in EEG: their generic features, similarities and distinctions disclosed with Fourier transform and continuous wavelet analysis. Journal of neuroscience methods 2009, 180, 304–316. [Google Scholar] [CrossRef]
- Zhuang, X.; Li, Y.; Peng, N. Enhanced automatic sleep spindle detection: a sliding window-based wavelet analysis and comparison using a proposal assessment method. In Proceedings of the Applied Informatics. Springer, Vol. 3; 2016; pp. 1–9. [Google Scholar]
- Zhou, S.; Zhang, X.; Yu, Z. A sleep spindle detection algorithm based on SVM and WT. In Proceedings of the 2017 29th Chinese Control And Decision Conference (CCDC). IEEE; 2017; pp. 2213–2217. [Google Scholar]
- Hekmatmanesh, A.; Mikaeili, M.; Sadeghniiat-Haghighi, K.; Wu, H.; Handroos, H.; Martinek, R.; Nazeran, H. Sleep spindle detection and prediction using a mixture of time series and chaotic features. Advances in Electrical and Electronic Engineering 2017, 15, 435–447. [Google Scholar] [CrossRef]
- Duman, F.; Erdamar, A.; Erogul, O.; Telatar, Z.; Yetkin, S. Efficient sleep spindle detection algorithm with decision tree. Expert Systems with Applications 2009, 36, 9980–9985. [Google Scholar] [CrossRef]
- Jiang, D.; Ma, Y.; Wang, Y. A robust two-stage sleep spindle detection approach using single-channel EEG. Journal of Neural Engineering 2021, 18, 026026. [Google Scholar] [CrossRef]
- You, J.; Jiang, D.; Ma, Y.; Wang, Y. SpindleU-Net: An adaptive u-net framework for sleep spindle detection in single-channel EEG. IEEE transactions on neural systems and rehabilitation engineering 2021, 29, 1614–1623. [Google Scholar] [CrossRef]
- Fraiwan, M.; Khasawneh, N. Visual identification of sleep spindles in EEG waveform images using deep learning object detection (YOLOv4 vs YOLOX). Cluster Computing 2024, 27, 13607–13620. [Google Scholar] [CrossRef]
- Chambon, S.; Thorey, V.; Arnal, P.J.; Mignot, E.; Gramfort, A. DOSED: A deep learning approach to detect multiple sleep micro-events in EEG signal. Journal of neuroscience methods 2019, 321, 64–78. [Google Scholar] [CrossRef] [PubMed]
- Tapia, N.I.; Estévez, P.A. RED: Deep recurrent neural networks for sleep EEG event detection. In Proceedings of the 2020 international joint conference on neural networks (IJCNN). IEEE; 2020; pp. 1–8. [Google Scholar]
- Chen, P.; Chen, D.; Zhang, L.; Tang, Y.; Li, X. Automated sleep spindle detection with mixed EEG features. Biomedical Signal Processing and Control 2021, 70, 103026. [Google Scholar] [CrossRef]
- Kaulen, L.; Schwabedal, J.T.; Schneider, J.; Ritter, P.; Bialonski, S. SUMO: Advanced sleep spindle identification with neural networks. arXiv e-prints 2022, pp. arXiv–2202.
- Lacourse, K.; Yetton, B.; Mednick, S.; Warby, S.C. Massive online data annotation, crowdsourcing to generate high quality sleep spindle annotations from EEG data. Scientific data 2020, 7, 190. [Google Scholar] [CrossRef]
- Moradi, F.; Mohammadi, H.; Rezaei, M.; Sariaslani, P.; Razazian, N.; Khazaie, H.; Adeli, H. A novel method for sleep-stage classification based on sonification of sleep electroencephalogram signals using wavelet transform and recurrent neural network. European Neurology 2020, 83, 468–486. [Google Scholar] [CrossRef]
- Michielli, N.; Acharya, U.R.; Molinari, F. Cascaded LSTM recurrent neural network for automated sleep stage classification using single-channel EEG signals. Computers in biology and medicine 2019, 106, 71–81. [Google Scholar] [CrossRef]
- Perslev, M.; Darkner, S.; Kempfner, L.; Nikolic, M.; Jennum, P.J.; Igel, C. U-Sleep: resilient high-frequency sleep staging. NPJ digital medicine 2021, 4, 72. [Google Scholar] [CrossRef]
- Jha, P.K.; Valekunja, U.K.; Reddy, A.B. SlumberNet: deep learning classification of sleep stages using residual neural networks. Scientific Reports 2024, 14, 4797. [Google Scholar] [CrossRef]
- Tapia-Rivas, N.I.; Estévez, P.A.; Cortes-Briones, J.A. A robust deep learning detector for sleep spindles and K-complexes: towards population norms. Scientific Reports 2024, 14, 263. [Google Scholar] [CrossRef]
- Grieger, N.; Mehrkanoon, S.; Ritter, P.; Bialonski, S. From Sleep Staging to Spindle Detection: Evaluating End-to-End Automated Sleep Analysis. arXiv preprint arXiv:2505.05371, arXiv:2505.05371 2025.
- Coppieters’t Wallant, D.; Maquet, P.; Phillips, C. Sleep spindles as an electrographic element: description and automatic detection methods. Neural plasticity 2016, 2016, 6783812. [Google Scholar] [CrossRef]
- Schiller, K.; Avigdor, T.; Abdallah, C.; Sziklas, V.; Crane, J.; Stefani, A.; Peter-Derex, L.; Frauscher, B. Focal epilepsy disrupts spindle structure and function. Scientific reports 2022, 12, 11137. [Google Scholar] [CrossRef]
- Bandarabadi, M.; Herrera, C.; Gent, T.; Bassetti, C.; Schindler, K.; Adamantidis, A. A role for spindles in the onset of rapid eye movement sleep. Nat Commun 11, 5247, 2020.
| SEED | U-Net | |
|---|---|---|
| Precision | ||
| Recall | ||
| F1-score |
| SEED | U-Net | |
|---|---|---|
| Precision | ||
| Recall | ||
| F1-score |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




