Submitted:
23 September 2025
Posted:
25 September 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Site-Saturation Mutagenesis Library Construction for DFPH-a.01
2.2. Yeast Display Library Screening
2.3. NGS Analysis of Sorted Yeast Libraries
2.4. Antibody Expression and Purification
2.5. Virus Neutralization Assays
2.5.1. 20-Virus Panel
2.5.2. 208-Virus Panel
2.6. Affinity Measurements by Surface Plasmon Resonance
2.7. Stability Measurements
2.8. Cryo-EM Structure Determination
2.9. Quantification and Statistical Analysis
3. Results
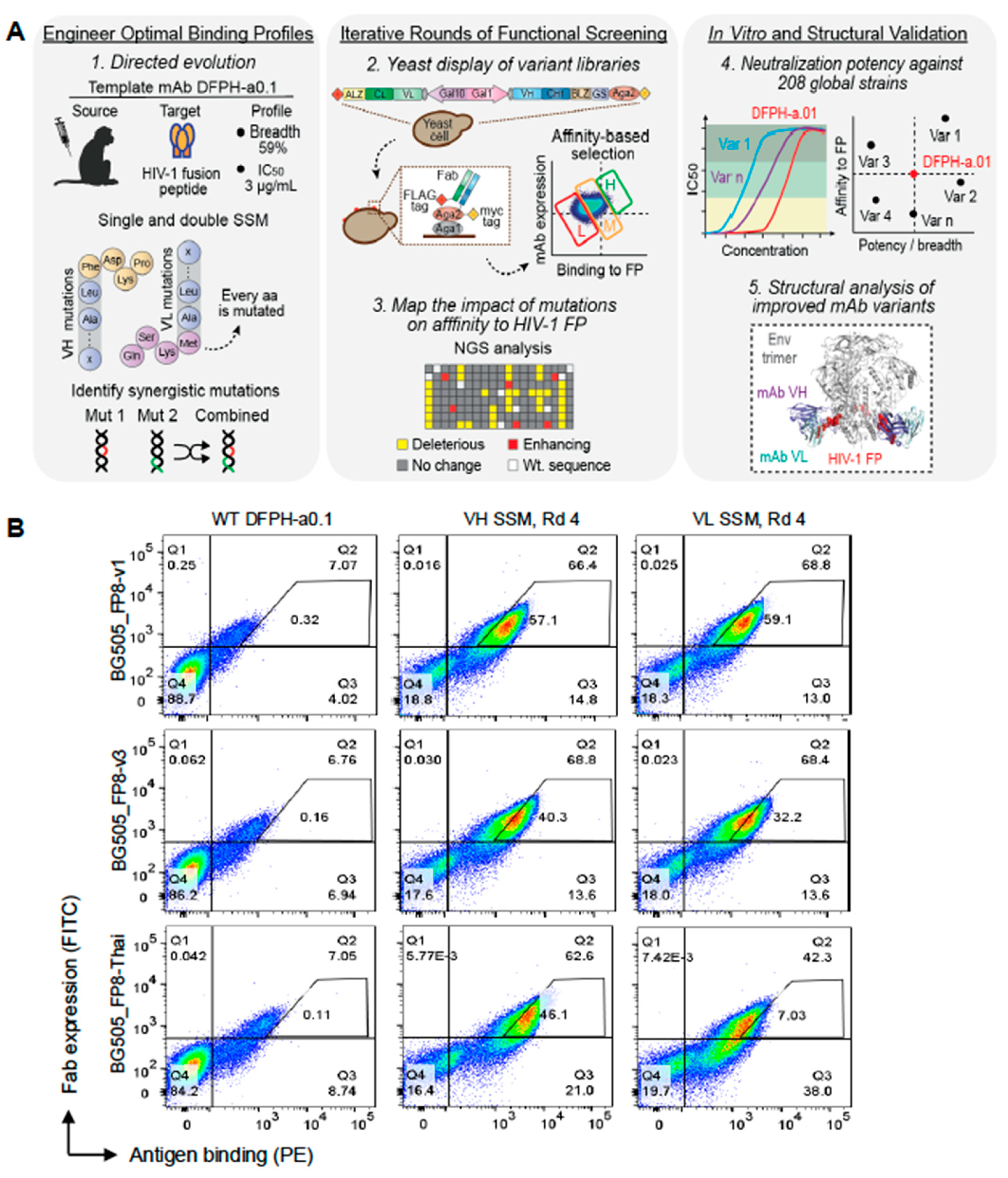
3.1. Precision Yeast Display Antibody Engineering of DFPH-a.01
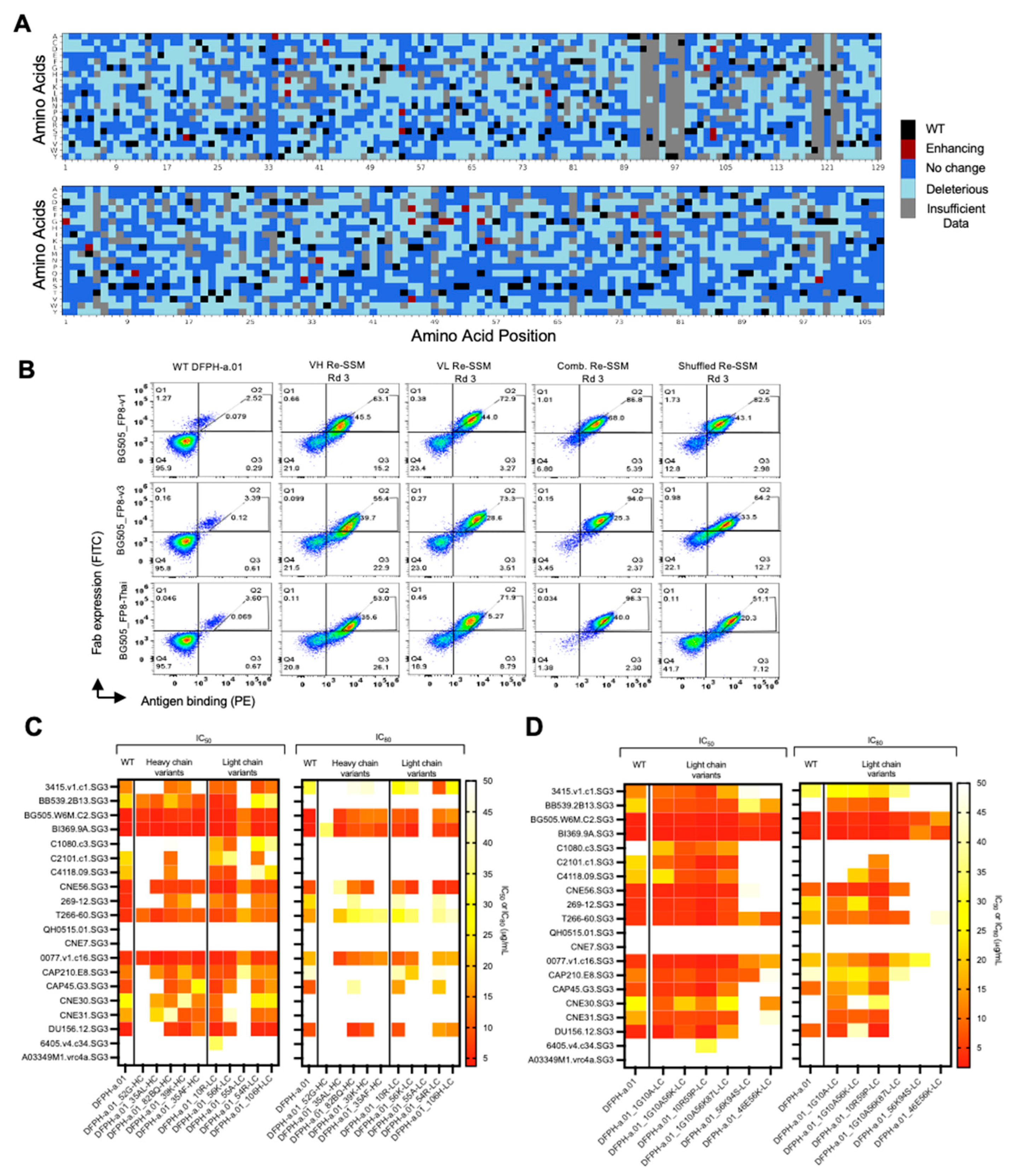
3.2. Multi-Mutation Library Generation and Screening
3.3. HIV-1 Neutralization Analysis of Mutational Variants Identified in Yeast Display
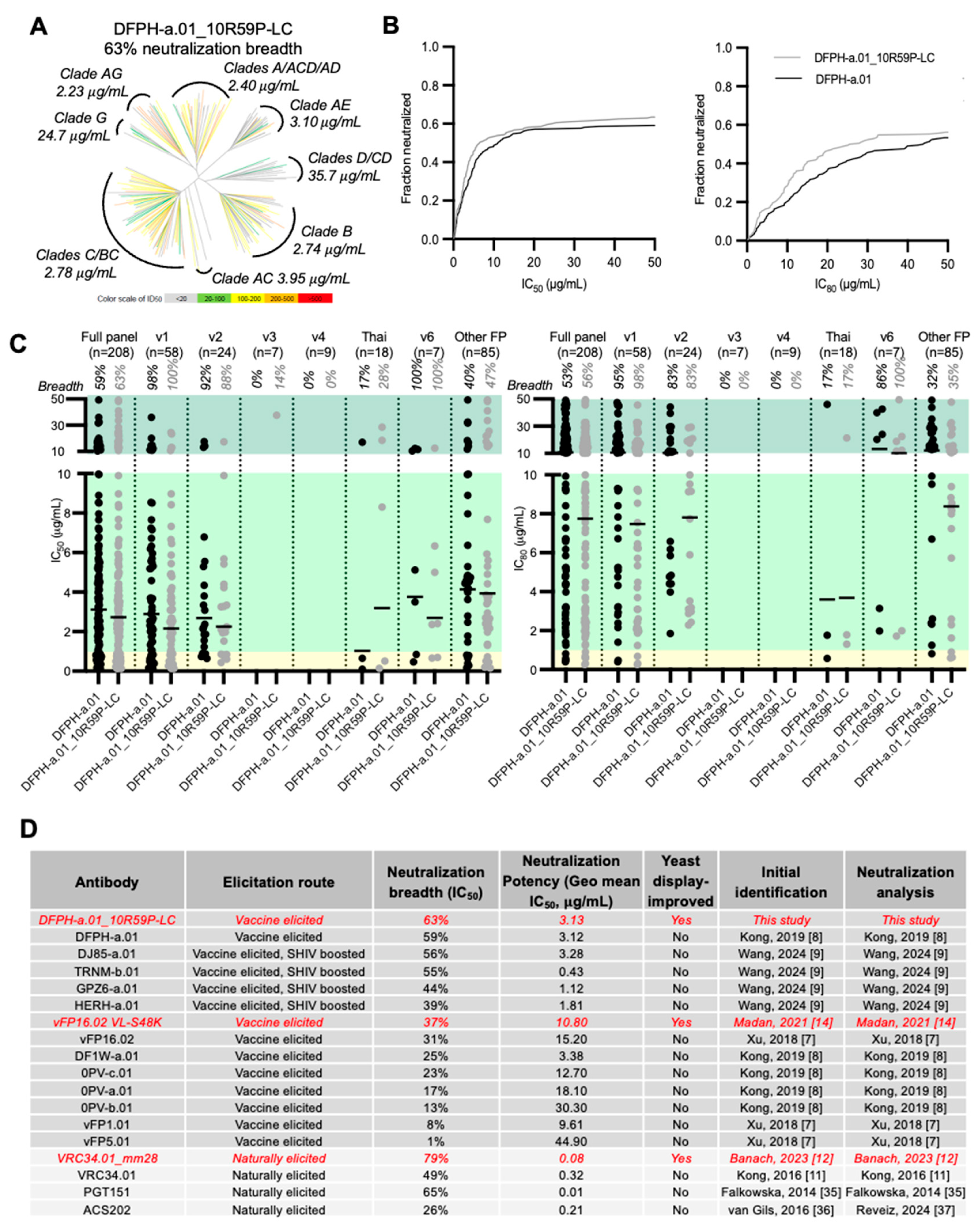
3.4. Breadth and Potency Analysis of DFPH-a.01_10R59P-LC
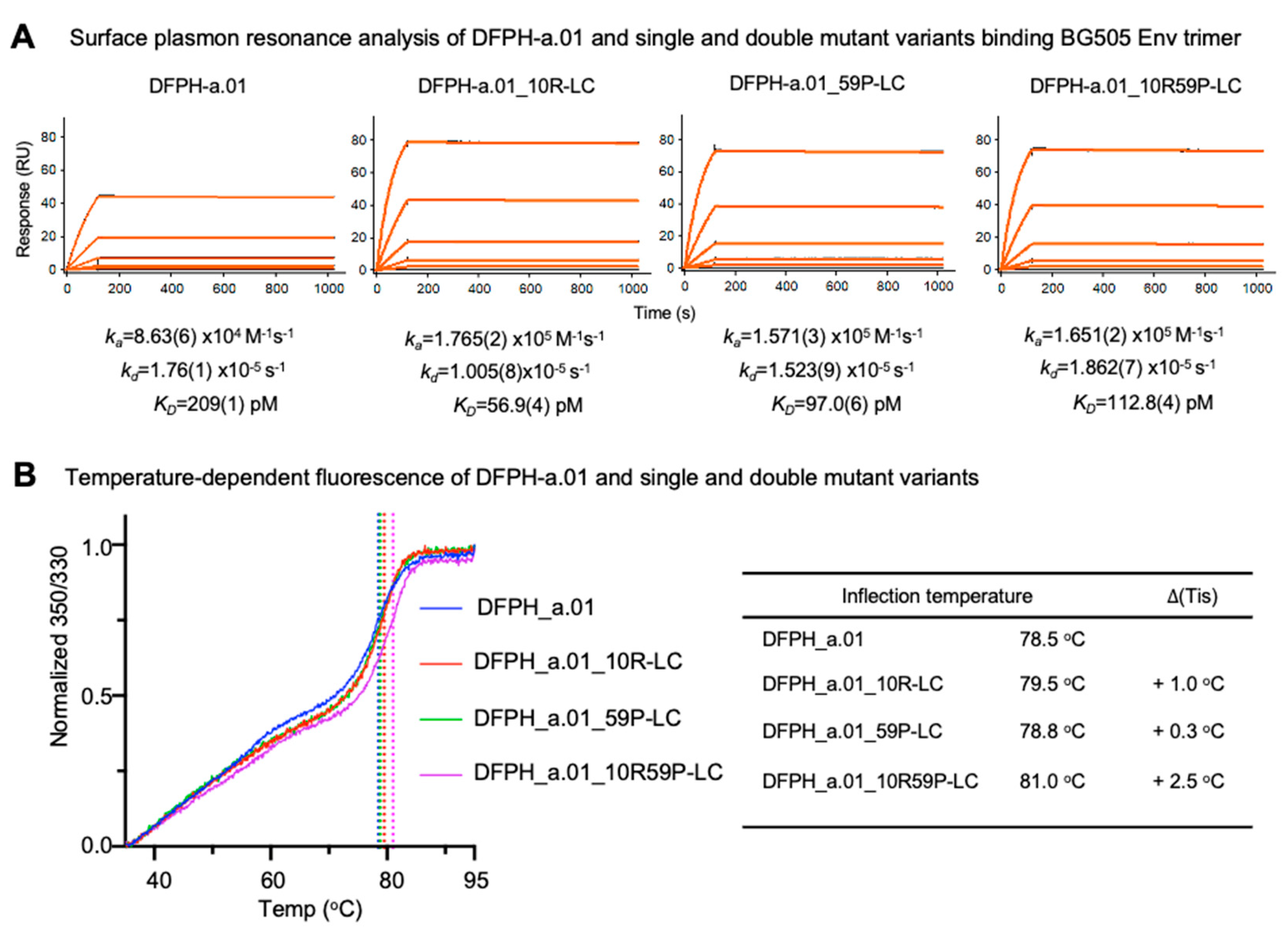
3.5. Surface Plasmon Resonance and Thermal Stability Analyses
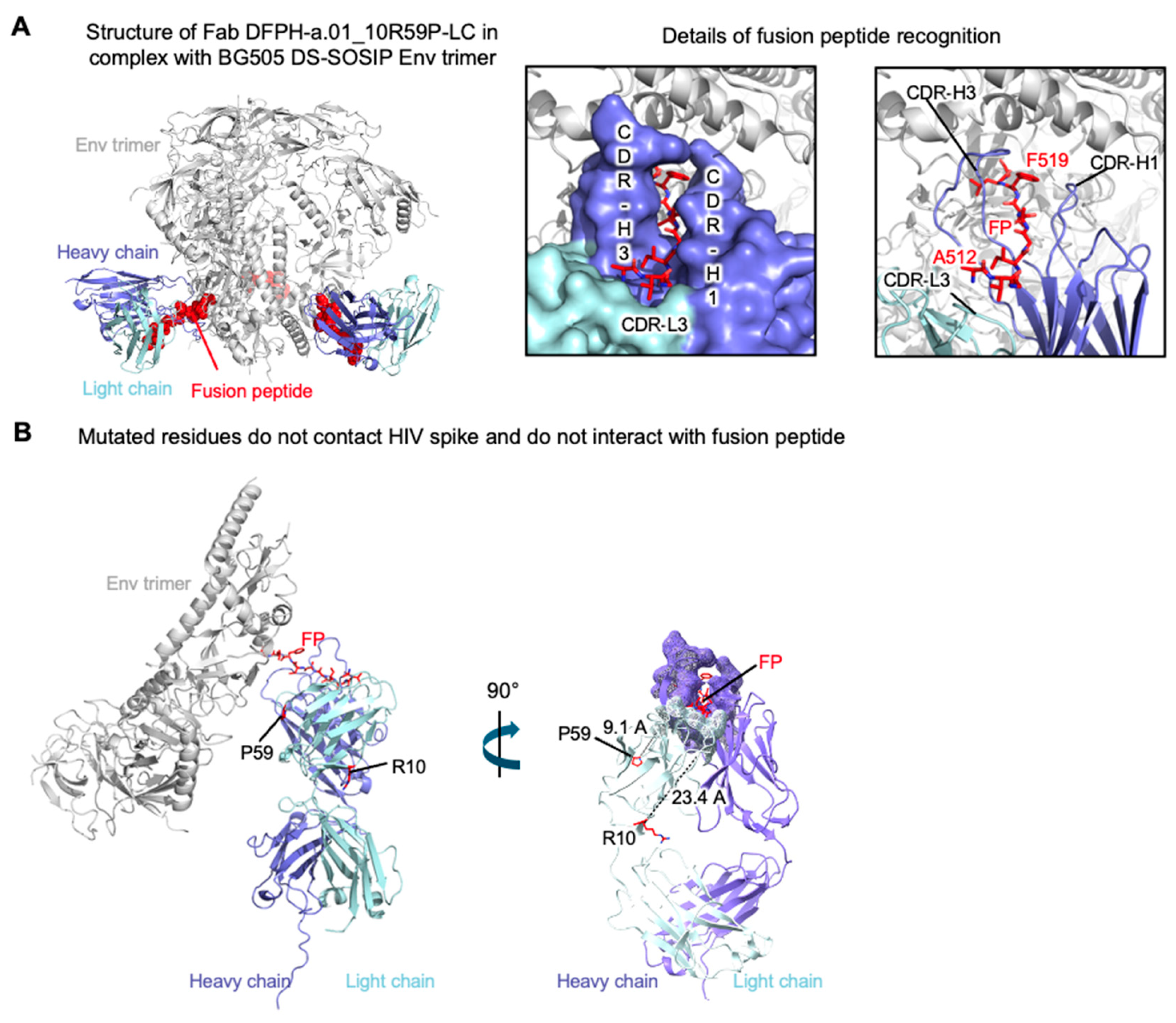
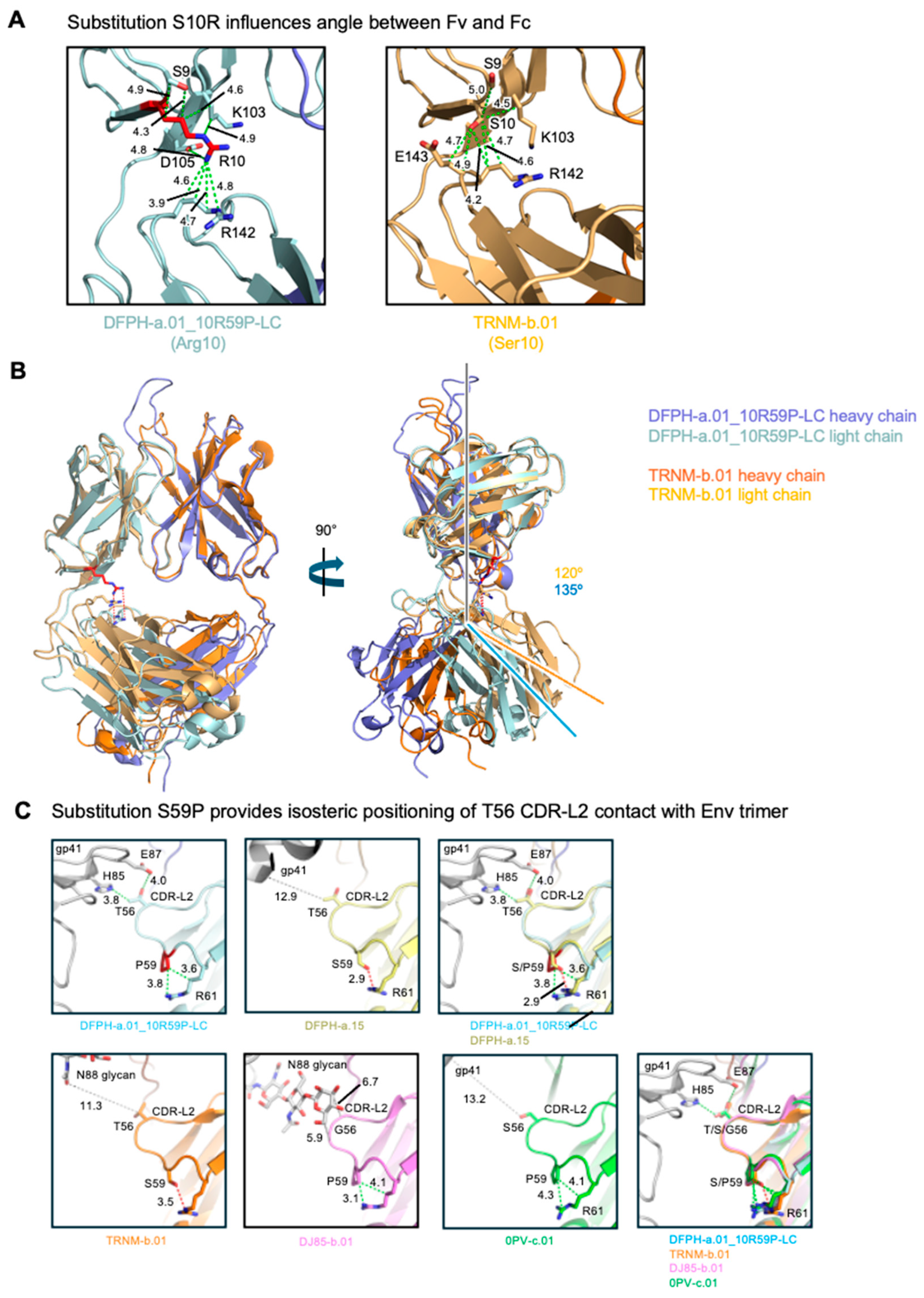
3.6. Cryo-EM Structure of Fab DFPH-a.01_10R59P-LC with BG505 DS-SOSIP Env Trimer
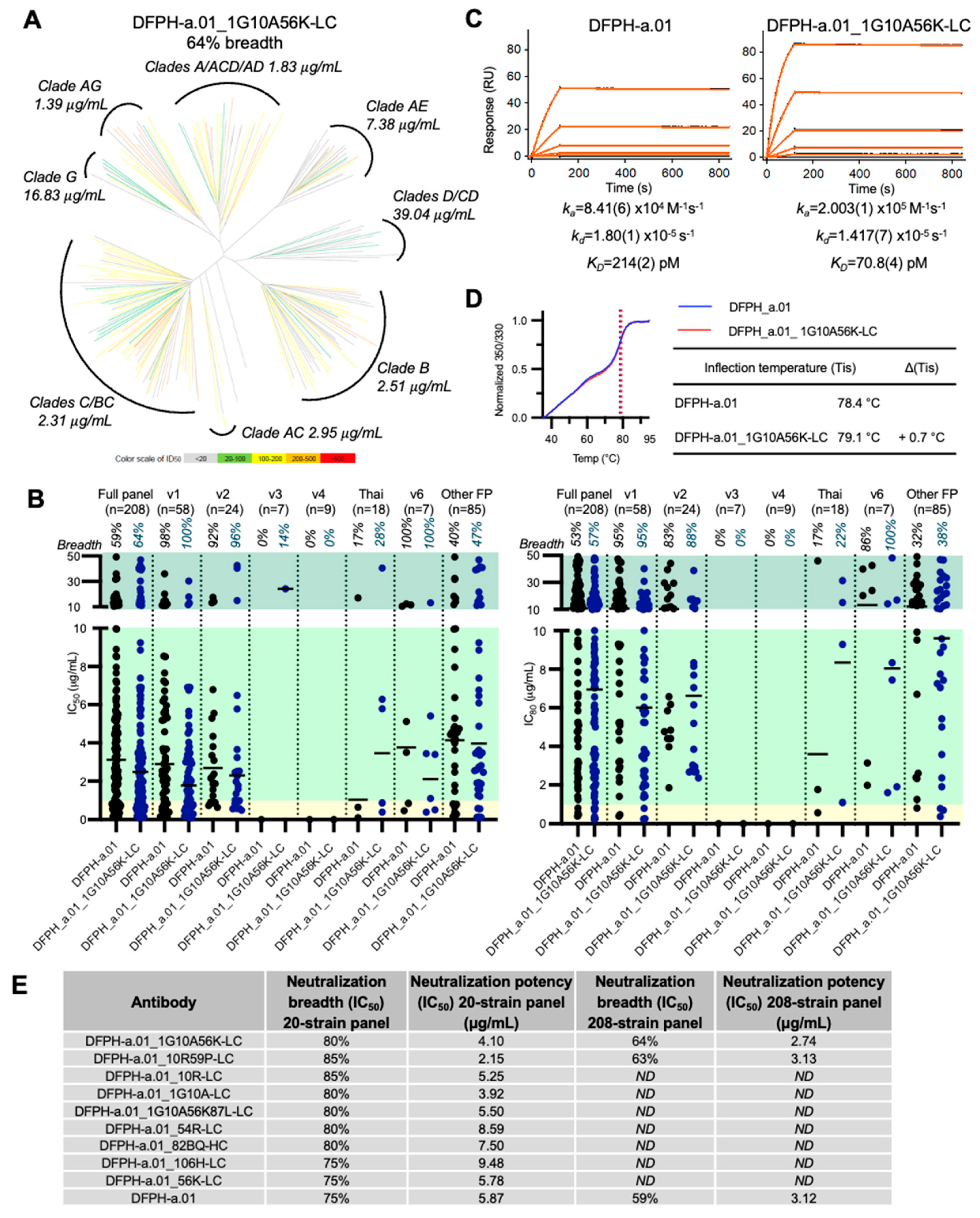
3.7. A Second Yeast Display-Identified Antibody, DFPH-a.01_1G10A56K-LC, with 64% Neutralization Breadth
3.8. Comparative Analysis of Improving FP Broadly Neutralizing Antibody Mutations Across Macaque, Mouse, and Human Studies
4. Discussion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| Å | Ångström (10⁻¹⁰ m, used for structural resolution/length) |
| BSA | Buried Surface Area |
| °C | Degrees Celsius |
| CDR | Complementarity-Determining Region |
| COOT | Model-Building Tools for Molecular Graphics |
| Cryo-EM | Cryogenic Electron Microscopy |
| CTF | Contrast Transfer Function |
| DDM | n-Dodecyl β-D-maltoside |
| DOAJ | Directory of Open Access Journals |
| Env | Envelope Glycoprotein |
| Fab | Fragment Antigen-Binding |
| FACS | Fluorescence-Activated Cell Sorting |
| FP | Fusion Peptide |
| FR | Framework Region |
| HIV | Human Immunodeficiency Virus |
| IC50/ IC80 |
Concentration of Antibody Required to Inhibit 50% / 80% of Infection |
| ID50 | Inhibitory Dilution 50 |
| KD | Equilibrium Dissociation Constant |
| Ka | Association Rate Constant |
| Kd | Dissociation Rate Constant |
| LD | Linear Dichroism |
| mg/ml | Milligrams per milliliter (concentration unit) |
| μg/ml | Micrograms per milliliter (concentration unit) |
| μL | Microliter (10⁻⁶ liter) |
| μM | Micromolar (10⁻⁶ molar concentration) |
| nM | Nanomolar (10⁻⁹ molar concentration) |
| NGS | Next-Generation Sequencing |
| PDB | Protein Data Bank |
| RU | Response Units (Surface Plasmon Resonance Readout) |
| SHIV | Simian-Human Immunodeficiency Virus |
| SHM | Somatic Hypermutation |
| SPR | Surface Plasmon Resonance |
| SSM | Site-Saturation Mutagenesis |
| VH | Variable Heavy Chain |
| VL | Variable Light Chain |
References
- Willis, J.R.; Berndsen, Z.T.; Ma, K.M.; Steichen, J.M.; Schiffner, T.; Landais, E.; Liguori, A.; Kalyuzhniy, O.; Allen, J.D.; Baboo, S.; et al. Human immunoglobulin repertoire analysis guides design of vaccine priming immunogens targeting HIV V2-apex broadly neutralizing antibody precursors. Immunity 2022, 55, 2149-2167 e2149. [CrossRef]
- Haynes, B.F.; Wiehe, K.; Borrow, P.; Saunders, K.O.; Korber, B.; Wagh, K.; McMichael, A.J.; Kelsoe, G.; Hahn, B.H.; Alt, F.; et al. Strategies for HIV-1 vaccines that induce broadly neutralizing antibodies. Nat Rev Immunol 2023, 23, 142-158. [CrossRef]
- Burton, D.R.; Hangartner, L. Broadly Neutralizing Antibodies to HIV and Their Role in Vaccine Design. Annu Rev Immunol 2016, 34, 635-659. [CrossRef]
- Medina-Ramirez, M.; Garces, F.; Escolano, A.; Skog, P.; de Taeye, S.W.; Del Moral-Sanchez, I.; McGuire, A.T.; Yasmeen, A.; Behrens, A.J.; Ozorowski, G.; et al. Design and crystal structure of a native-like HIV-1 envelope trimer that engages multiple broadly neutralizing antibody precursors in vivo. J Exp Med 2017, 214, 2573-2590. [CrossRef]
- Schleich, F.A.; Bale, S.; Guenaga, J.; Ozorowski, G.; Adori, M.; Lin, X.; Castro Dopico, X.; Wilson, R.; Chernyshev, M.; Cotgreave, A.T.; et al. Vaccination of nonhuman primates elicits a broadly neutralizing antibody lineage targeting a quaternary epitope on the HIV-1 Env trimer. Immunity 2025, 58, 1598-1613 e1598. [CrossRef]
- Dubrovskaya, V.; Tran, K.; Ozorowski, G.; Guenaga, J.; Wilson, R.; Bale, S.; Cottrell, C.A.; Turner, H.L.; Seabright, G.; O'Dell, S.; et al. Vaccination with Glycan-Modified HIV NFL Envelope Trimer-Liposomes Elicits Broadly Neutralizing Antibodies to Multiple Sites of Vulnerability. Immunity 2019, 51, 915-929 e917. [CrossRef]
- Xu, K.; Acharya, P.; Kong, R.; Cheng, C.; Chuang, G.Y.; Liu, K.; Louder, M.K.; O'Dell, S.; Rawi, R.; Sastry, M.; et al. Epitope-based vaccine design yields fusion peptide-directed antibodies that neutralize diverse strains of HIV-1. Nat Med 2018, 24, 857-867. [CrossRef]
- Kong, R.; Duan, H.; Sheng, Z.; Xu, K.; Acharya, P.; Chen, X.; Cheng, C.; Dingens, A.S.; Gorman, J.; Sastry, M.; et al. Antibody Lineages with Vaccine-Induced Antigen-Binding Hotspots Develop Broad HIV Neutralization. Cell 2019, 178, 567-584 e519. [CrossRef]
- Wang, H.; Cheng, C.; Dal Santo, J.L.; Shen, C.H.; Bylund, T.; Henry, A.R.; Howe, C.A.; Hwang, J.; Morano, N.C.; Morris, D.J.; et al. Potent and broad HIV-1 neutralization in fusion peptide-primed SHIV-infected macaques. Cell 2024, 187, 7214-7231 e7223. [CrossRef]
- Pegu, A.; Lovelace, S.E.; DeMouth, M.E.; Cully, M.D.; Morris, D.J.; Li, Y.; Wang, K.; Schmidt, S.D.; Choe, M.; Liu, C.; et al. Antibodies targeting the fusion peptide on the HIV envelope provide protection to rhesus macaques against mucosal SHIV challenge. Sci Transl Med 2024, 16, eadh9039. [CrossRef]
- Kong, R.; Xu, K.; Zhou, T.; Acharya, P.; Lemmin, T.; Liu, K.; Ozorowski, G.; Soto, C.; Taft, J.D.; Bailer, R.T.; et al. Fusion peptide of HIV-1 as a site of vulnerability to neutralizing antibody. Science 2016, 352, 828-833. [CrossRef]
- Banach, B.B.; Pletnev, S.; Olia, A.S.; Xu, K.; Zhang, B.; Rawi, R.; Bylund, T.; Doria-Rose, N.A.; Nguyen, T.D.; Fahad, A.S.; et al. Antibody-directed evolution reveals a mechanism for enhanced neutralization at the HIV-1 fusion peptide site. Nat Commun 2023, 14, 7593. [CrossRef]
- Banach, B.B.; Tripathi, P.; Da Silva Pereira, L.; Gorman, J.; Nguyen, T.D.; Dillon, M.; Fahad, A.S.; Kiyuka, P.K.; Madan, B.; Wolfe, J.R.; et al. Highly protective antimalarial antibodies via precision library generation and yeast display screening. J Exp Med 2022, 219. [CrossRef]
- Madan, B.; Zhang, B.; Xu, K.; Chao, C.W.; O'Dell, S.; Wolfe, J.R.; Chuang, G.Y.; Fahad, A.S.; Geng, H.; Kong, R.; et al. Mutational fitness landscapes reveal genetic and structural improvement pathways for a vaccine-elicited HIV-1 broadly neutralizing antibody. Proc Natl Acad Sci U S A 2021, 118. [CrossRef]
- Wrenbeck, E.E.; Klesmith, J.R.; Stapleton, J.A.; Adeniran, A.; Tyo, K.E.; Whitehead, T.A. Plasmid-based one-pot saturation mutagenesis. Nat Methods 2016, 13, 928-930. [CrossRef]
- Meyer, A.J.; Ellefson, J.W.; Ellington, A.D. Library generation by gene shuffling. Curr Protoc Mol Biol 2014, 105, Unit 15 12. [CrossRef]
- Whitehead, T.A.; Chevalier, A.; Song, Y.; Dreyfus, C.; Fleishman, S.J.; De Mattos, C.; Myers, C.A.; Kamisetty, H.; Blair, P.; Wilson, I.A.; et al. Optimization of affinity, specificity and function of designed influenza inhibitors using deep sequencing. Nat Biotechnol 2012, 30, 543-548. [CrossRef]
- Wang, B.; DeKosky, B.J.; Timm, M.R.; Lee, J.; Normandin, E.; Misasi, J.; Kong, R.; McDaniel, J.R.; Delidakis, G.; Leigh, K.E.; et al. Functional interrogation and mining of natively paired human V(H):V(L) antibody repertoires. Nat Biotechnol 2018, 36, 152-155. [CrossRef]
- McDaniel, J.R.; DeKosky, B.J.; Tanno, H.; Ellington, A.D.; Georgiou, G. Ultra-high-throughput sequencing of the immune receptor repertoire from millions of lymphocytes. Nat Protoc 2016, 11, 429-442. [CrossRef]
- Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010, 26, 2460-2461. [CrossRef]
- Fahad, A.S.; Gutierrez-Gonzalez, M.F.; Madan, B.; DeKosky, B.J. Clonal Variant Analysis of Antibody Engineering Libraries. Cold Spring Harb Protoc 2025, 2025, pdb prot108626. [CrossRef]
- Sarzotti-Kelsoe, M.; Bailer, R.T.; Turk, E.; Lin, C.L.; Bilska, M.; Greene, K.M.; Gao, H.; Todd, C.A.; Ozaki, D.A.; Seaman, M.S.; et al. Optimization and validation of the TZM-bl assay for standardized assessments of neutralizing antibodies against HIV-1. J Immunol Methods 2014, 409, 131-146. [CrossRef]
- Mastronarde, D.N. Automated electron microscope tomography using robust prediction of specimen movements. J Struct Biol 2005, 152, 36-51. [CrossRef]
- Punjani, A.; Rubinstein, J.L.; Fleet, D.J.; Brubaker, M.A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat Methods 2017, 14, 290-296. [CrossRef]
- Mirdita, M.; Schutze, K.; Moriwaki, Y.; Heo, L.; Ovchinnikov, S.; Steinegger, M. ColabFold: making protein folding accessible to all. Nat Methods 2022, 19, 679-682. [CrossRef]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Zidek, A.; Potapenko, A.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583-589. [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera--a visualization system for exploratory research and analysis. J Comput Chem 2004, 25, 1605-1612. [CrossRef]
- Emsley, P.; Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr D Biol Crystallogr 2004, 60, 2126-2132. [CrossRef]
- Emsley, P.; Lohkamp, B.; Scott, W.G.; Cowtan, K. Features and development of Coot. Acta Crystallogr D Biol Crystallogr 2010, 66, 486-501. [CrossRef]
- Liebschner, D.; Afonine, P.V.; Baker, M.L.; Bunkóczi, G.; Chen, V.B.; Croll, T.I.; Hintze, B.; Hung, L.W.; Jain, S.; McCoy, A.J.; et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr D Struct Biol 2019, 75, 861-877. [CrossRef]
- Davis, I.W.; Murray, L.W.; Richardson, J.S.; Richardson, D.C. MOLPROBITY: structure validation and all-atom contact analysis for nucleic acids and their complexes. Nucleic Acids Res 2004, 32, W615-619. [CrossRef]
- Chen, V.B.; Arendall, W.B., 3rd; Headd, J.J.; Keedy, D.A.; Immormino, R.M.; Kapral, G.J.; Murray, L.W.; Richardson, J.S.; Richardson, D.C. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr D Biol Crystallogr 2010, 66, 12-21. [CrossRef]
- Krissinel, E.; Henrick, K. Inference of macromolecular assemblies from crystalline state. J Mol Biol 2007, 372, 774-797. [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Meng, E.C.; Couch, G.S.; Croll, T.I.; Morris, J.H.; Ferrin, T.E. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci 2021, 30, 70-82. [CrossRef]
- Falkowska, E.; Le, K.M.; Ramos, A.; Doores, K.J.; Lee, J.H.; Blattner, C.; Ramirez, A.; Derking, R.; van Gils, M.J.; Liang, C.H.; et al. Broadly neutralizing HIV antibodies define a glycan-dependent epitope on the prefusion conformation of gp41 on cleaved envelope trimers. Immunity 2014, 40, 657-668. [CrossRef]
- van Gils, M.J.; van den Kerkhof, T.L.; Ozorowski, G.; Cottrell, C.A.; Sok, D.; Pauthner, M.; Pallesen, J.; de Val, N.; Yasmeen, A.; de Taeye, S.W.; et al. An HIV-1 antibody from an elite neutralizer implicates the fusion peptide as a site of vulnerability. Nat Microbiol 2016, 2, 16199. [CrossRef]
- Reveiz, M.; Xu, K.; Lee, M.; Wang, S.; Olia, A.S.; Harris, D.R.; Liu, K.; Liu, T.; Schaub, A.J.; Stephens, T.; et al. Vaccine-elicited and naturally elicited antibodies differ in their recognition of the HIV-1 fusion peptide. Front Immunol 2024, 15, 1484029. [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



