Submitted:
21 September 2025
Posted:
23 September 2025
You are already at the latest version
Abstract
A novel flavi–like virus such as Alongshan virus (ALSV) with segmented genomes have been early isolated from Ixodes ticks in Russia. In this study, the ALSV genetic markers were tested in 4458 ixodid ticks collected in 22 regions of European and Asian part of Russia by RT PCR. The highest infection rate of ALSV in ticks was detected for Khakassia Republic (3.3%) and Kemerovo region (2.4%) and low infection rate was more typical for European part of Russia (0.4–0.7%). The sequencing complete genomes (four segments) of the ALSV from 22 PCR positive Ixodes persulcatus ticks was carry out by high–throughput sequencing. The identity level nucleotide sequences for Asian ALSV isolates are 94.5–96.5% with ALSV isolates previously found in China and in 89–93% range for the European isolates. This data together with phylogenetic analysis are propose the existence of Asian and European subtypes of the ALSV, and they may be associated with I. persulcatus and I. ricinus ticks. The obtained results actualize spreading ALSV in Russia and also may be useful for diagnostics, prophylactic and treatment this infection.
Keywords:
1. Introduction
2. Materials and Methods
2.1. Collection and Processing of Ticks
2.2. Reverse–Transcriptase PCR (RT–PCR) and Sequencing of Amplified Products
2.3. NGS Sequencing and NGS Data Analysis
| Description | Primers | Sequences (5′–3′) | Size (bp) |
|---|---|---|---|
| Segment 1 | AL1_1F | GCCATGATTGTCCTGATAGTG | |
| AL1_1R | GCCCTGTCCATCTTCATTTCC | 982 | |
| AL1_2F | AGGAAAGACAGATCACTCAC | ||
| AL1_2R | GGACATCATGGACTTCTCCT | 1038 | |
| AL1_3F | AGAAGTCCATGATGTCCTCC | ||
| AL1_3R | GTTCATCCAGTCCTTGTAGTTTCC | 840 | |
| Segment 2 | AL2_1F | GTAACCTCCGTAGACTGTCCA | |
| AL2_1R | GTCCCTTCCGTTTGGTTGTG | 477 | |
| AL2_2F | CTTGCTACATCGGAATCATGCC | ||
| AL2_2R | GATAAGCCCTCTCGATACCTC | 1091 | |
| AL2_3F | TGGTACGACTGGCTTTCGAG | ||
| AL2_3R | ACTTGTTGTAGTCTGCAACCC | 1098 | |
| Segment 3 | AL3_1F | TCGTCCAAGACTACTTAACAG | |
| AL3_1R | GTATCGCCTGTCCTCTATCC | 721 | |
| AL3_2F | TGCTGTCCATAGCAATCATACC | ||
| AL3_2R | GTAGGACACGTCCTTTGCGA | 865 | |
| AL3_3F | GCAAAGGACGTGTCCTACGT | ||
| AL3_3R | TTACCACTTGCTGGTCACAG | 1314 | |
| Segment 4 | AL4_1F | ACTTTGATCTACATCCTCGCC | |
| AL4_1R | GTATCCAGCTCTTCCCTTCTC | 824 | |
| AL4_2F | GGAAGAGCTGGATACCGAACTG | ||
| AL4_2R | TGCCAGATGTGTAGCTTCCC | 1274 | |
| AL4_3F | CAGCACTGGCGAAGATAACC | ||
| AL4_3R | TGCCCTGATACCTCCTAGCA | 503 |
2.4. Phylogenetic Analysis
2.5. Nucleotide Sequence Accession Numbers
2.6. Biosafety
3. Results
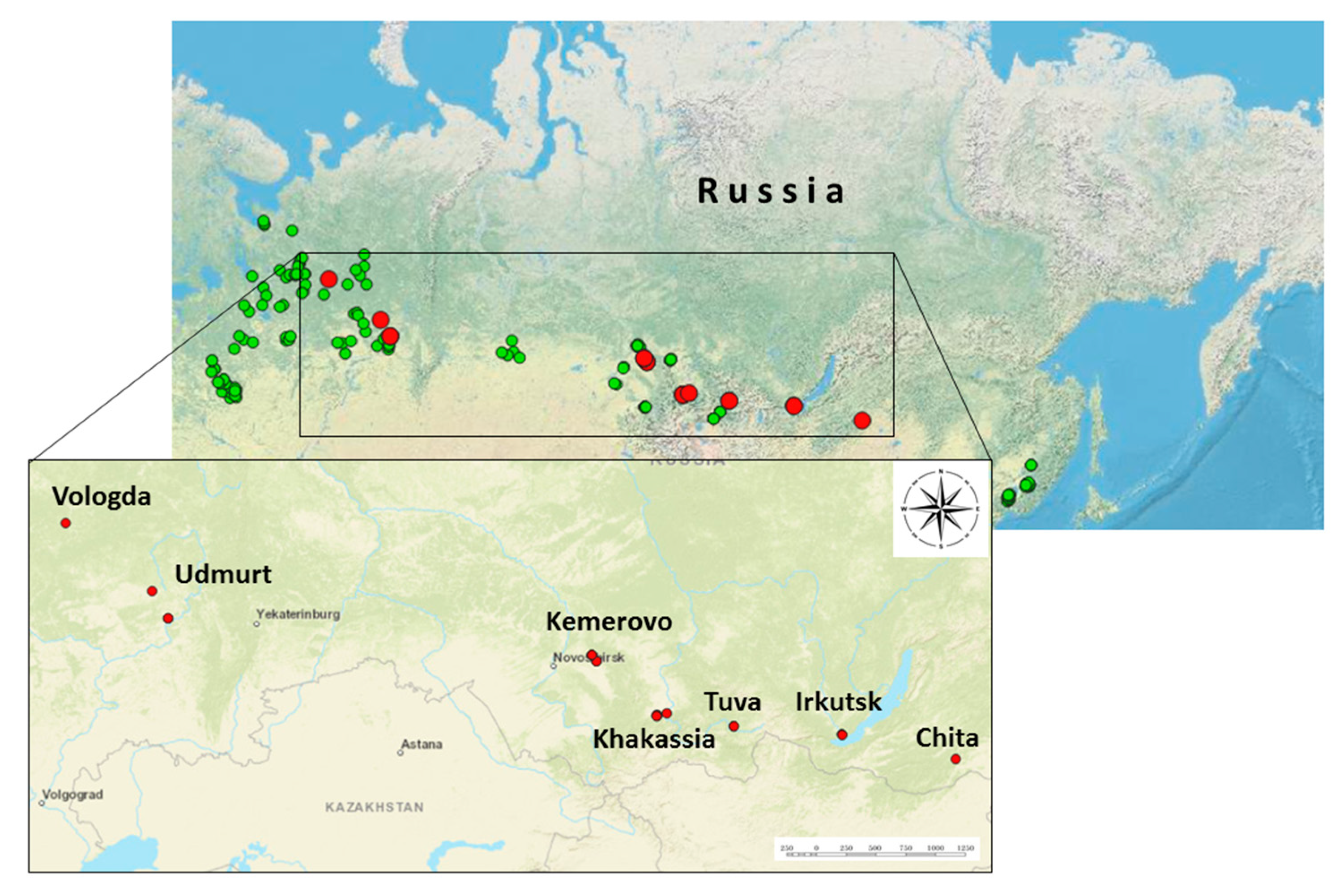
3.1. Tick Collection and ALSV Detection
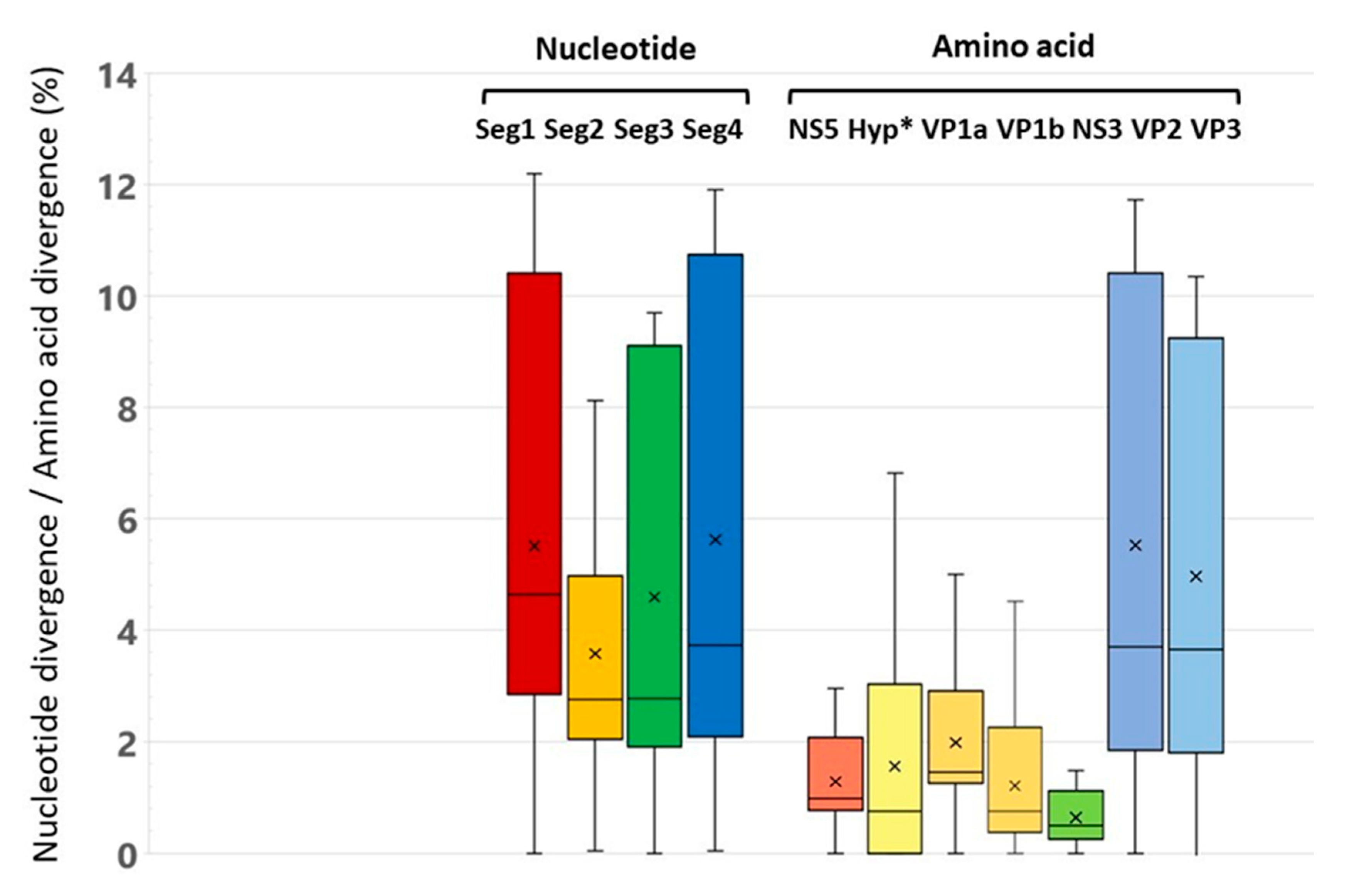
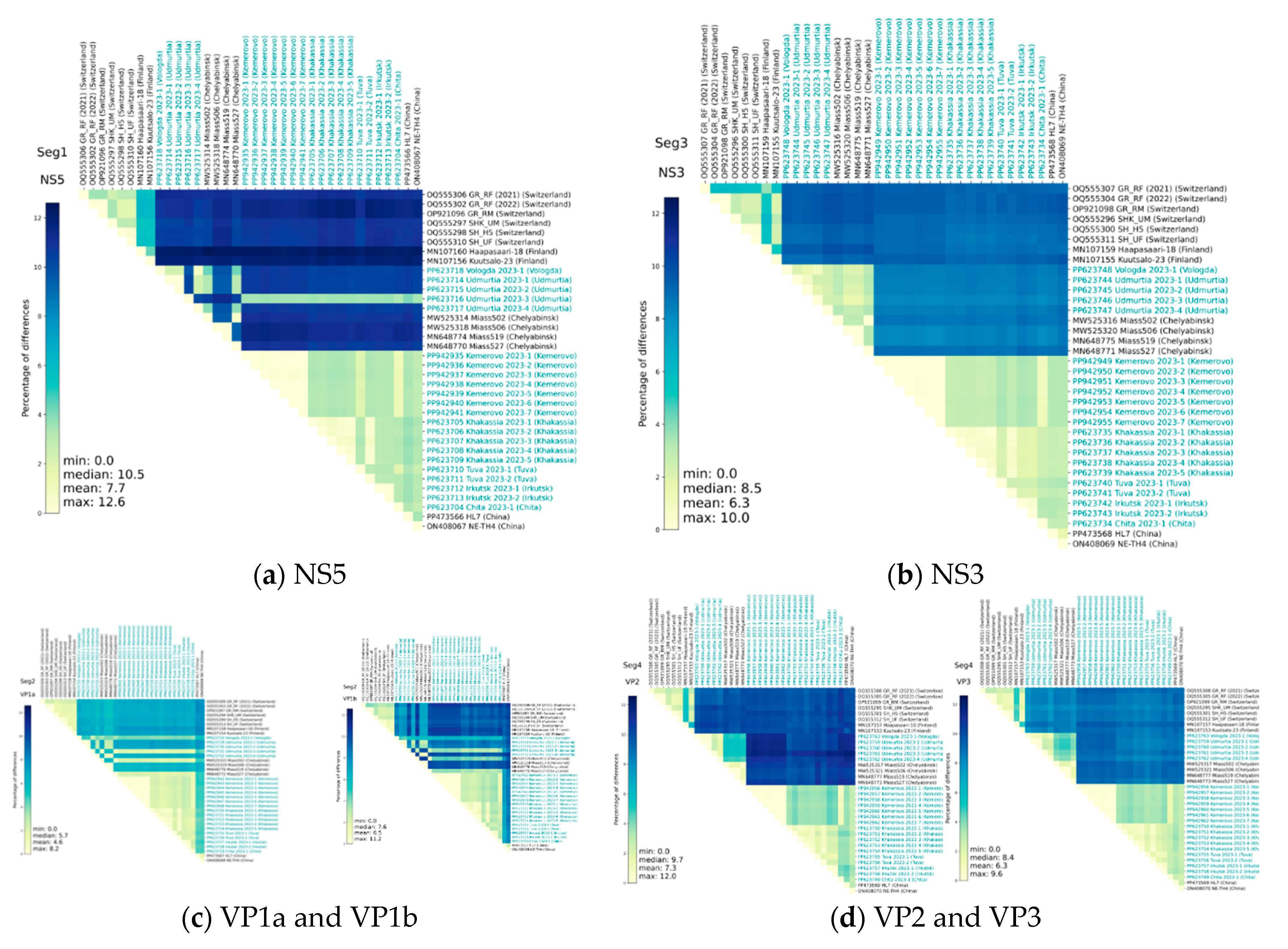
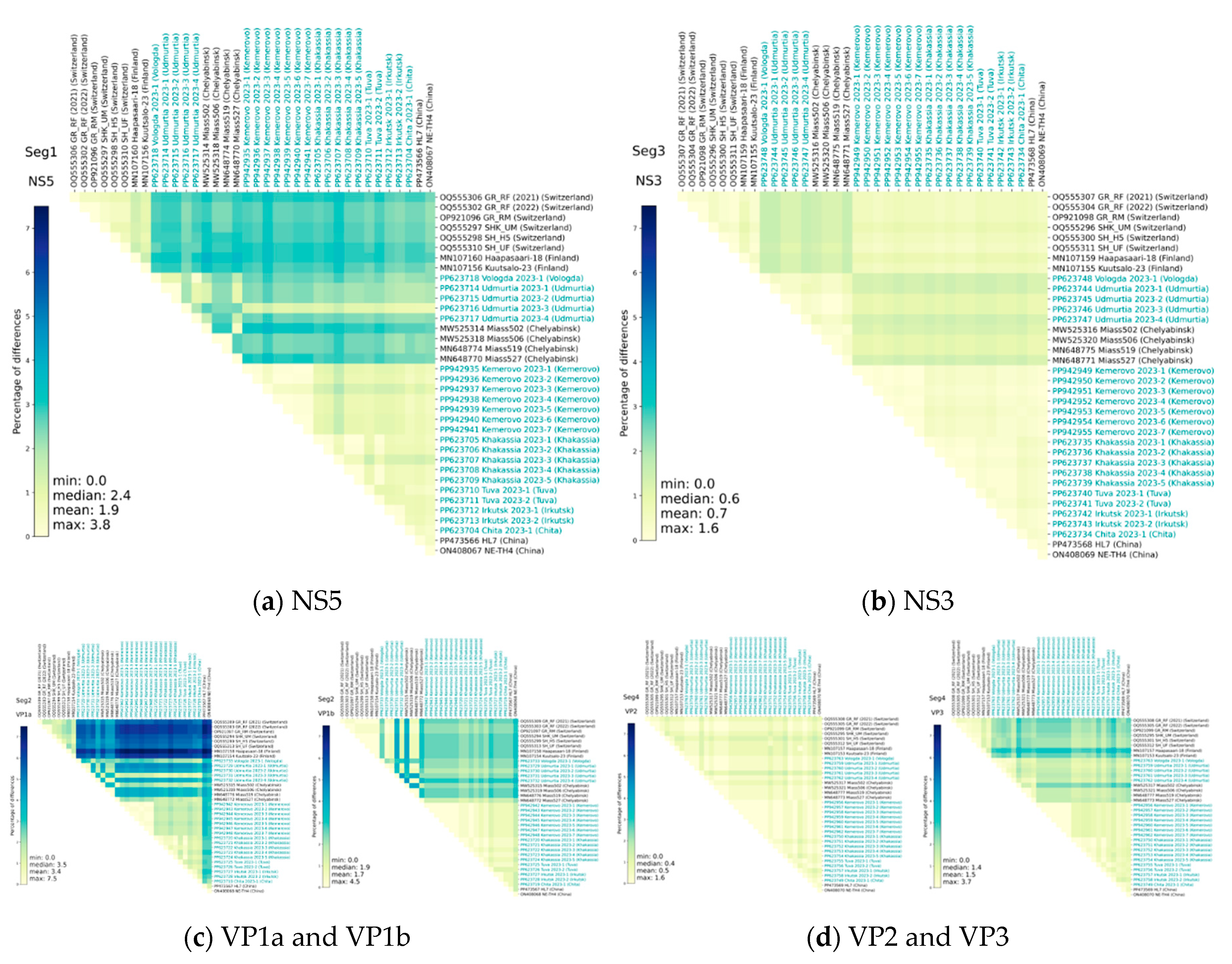
3.2. Analysis of Genome Identity
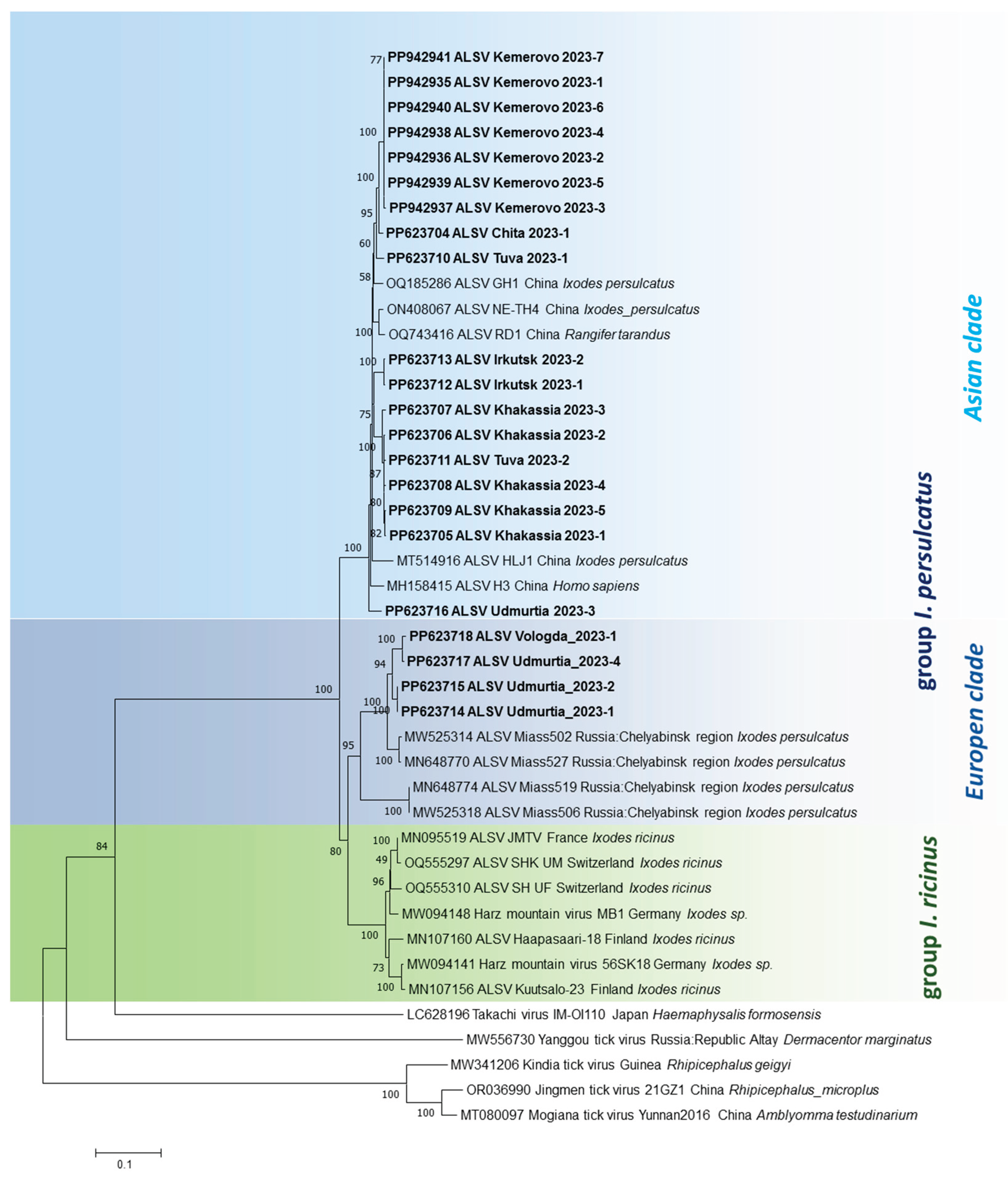
3.3. Phylogenetic Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Pierson, T.C. , Diamond, M. S. The continued threat of emerging flaviviruses. Nat Microbiol. 2020, 5, 796–812. [Google Scholar] [CrossRef]
- Leonova, G.N., Kondratov, I.G., Ternovoi, V.A., Romanova, E.V., Protopopova, E.V., Chausov, E.V., Pavlenko, E.V., Ryabchikova, E.I., Belikov, S.I., Loktev, V.B. Characterization of Powassan viruses from Far Eastern Russia. Arch Virol. 2009, 154(5), 811-820. [CrossRef]
- Ruzek, D. , Avšič Županc, T., Borde, J., Chrdle, A., Eyer, L., Karganova, G., Kholodilov, I., Knap, N., Kozlovskaya, L., Matveev, A., et al. Tick–borne encephalitis in Europe and Russia: Review of pathogenesis, clinical features, therapy, and vaccines. Antiviral Res. 2019, 164, 23–51. [Google Scholar] [CrossRef] [PubMed]
- Tyulko, Z.S., Fadeev, A.V., Vasilenko, A.G., Gradoboeva, E.A., Yakimenko, V.V., Komissarov, A.B. Analysis of changes in the genome of the Omsk hemorrhagic fever virus (Flaviviridae: Orthoflavivirus) during laboratory practices for virus preservation, Problems of Virology (Voprosy Virusologii). 2024, 69(6), 509–523. [CrossRef]
- Ternovoi, V.A. Ternovoi, V.A., Gladysheva, A.V., Ponomareva, E.P., Mikryukova, T.P., Protopopova, E.V., Shvalov, A.N., Konovalova, S.N., Chausov, E.V., Loktev, V.B. Variability in the 3’ untranslated regions of the genomes of the different tick–borne encephalitis virus subtypes. Virus Genes. 2019, 55, 448–457. [CrossRef]
- Postler, T.S., Beer, M., Blitvich, B.J., Bukh, J., de Lamballerie, X., Drexler, J.F., Imrie, A., Kapoor, A., Karganova, G.G., Lemey, P., et al. Renaming of the genus Flavivirus to Orthoflavivirus and extension of binomial species names within the family Flaviviridae. Arch Virol. 2023, 168, 224. [CrossRef]
- Qin, X.C.; Shi, M.; Tian, J.H.; Lin, X.D.; Gao, D.Y.; He, J.R.; Wang, J.B.; Li, C.X.; Kang, Y.J.; Yu, B.; et al. A tick–borne segmented RNA virus contains genome segments derived from unsegmented viral ancestors. Proc Natl Acad Sci U S A. 2014, 111, 6744–6749. [Google Scholar] [CrossRef] [PubMed]
- Villa, E.C., Maruyama, S.R., de Miranda–Santos, I.K.F., Palacios, G., Ladner, J.T. Complete Coding Genome Sequence for Mogiana Tick Virus, a Jingmenvirus Isolated from Ticks in Brazil. Genome Announc. 2017, 5, e00232–17. [CrossRef]
- Colmant, A.M.G. , Charrel, R.N., Coutard, B. Jingmenviruses: Ubiq–uitous, understudied, segmented flavi–like viruses. Front. Microbiol. 9970. [Google Scholar] [CrossRef]
- Ogola, E.O. , Roy, A., Wollenberg, K., Ochwoto, M., Bloom, M.E. Strange relatives: the enigmatic arbo–jingmenviruses and orthoflaviviruses. npj Viruses 2025, 3, 24. [Google Scholar] [CrossRef] [PubMed]
- Temmam, S., Bigot, T., Chrétien, D., Gondard, M., Pérot, P., Pommelet, V., Dufour, E., Petres, S., Devillers, E., Hoem, T., et al. Insights into the host range, genetic diversity, and geographical distribution of Jingmenviruses. mSphere. 2019; 4, e00645–19. [CrossRef]
- Valle, C., Parry, R.H., Coutard, B., Colmant, A.M.G. Discovery of additional genomic segments reveals the fluidity of jingmenvirus genomic organization. Virus Evolution, 2025, 11, veaf023. [CrossRef]
- Wang, Z.D.; Wang, B.; Wei, F.; Han, S.Z.; Zhang, L.; Yang, Z.T.; Yan, Y.; Lv, X.L.; Li, L.; Wang, S.C.; et al. A New Segmented Virus Associated with Human Febrile Illness in China. N Engl J Med. 2019, 380, 2116–2125. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.D., Wang, W., Wang, N.N., Qiu, K., Zhang, X., Tana. G., Liu, Q., Zhu, X.Q. Prevalence of the emerging novel Alongshan virus infection in sheep and cattle in Inner Mongolia, northeastern China. Parasit Vectors. 2019, 12, 450. [CrossRef]
- Kuivanen, S., Levanov, L., Kareinen, L., Sironen, T., Jääskeläinen, A.J., Plyusnin, I., Zakham, F., Emmerich, P., Schmidt–Chanasit, J., Hepojoki, J., Smura, T., Vapalahti, O. Detection of novel tick–borne pathogen, Alongshan virus, in Ixodes ricinus ticks, south–eastern Finland, 2019. Euro Surveill. 2019, 24, 1900394. [CrossRef]
- Stanojević, M., Li, K., Stamenković, G., Ilić, B., Paunović, M., Pešić, B., Maslovara, I.Đ., Šiljić, M., Ćirković, V., Zhang,.Y. Depicting the RNA virome of hematophagous arthropods from Belgrade, Serbia. Viruses. 2020, 12, 975. [CrossRef]
- Ebert, C.L., Söder, L., Kubinski, M., Glanz, J., Gregersen, E., Dümmer, K., Grund, D., Wöhler, A.S., Könenkamp, L., Liebig, K., et al. Detection and characterization of Alongshan virus in ticks and tick saliva from Lower Saxony, Germany with serological evidence for viral transmission to game and domestic animals. Microorganisms. 2023, 11, 543. [CrossRef]
- Stegmüller, S. , Fraefel, C., Kubacki, J. Genome Sequence of Alongshan virus from Ixodes ricinus ticks collected in Switzerland. Microbiol. Resour. Announc. 2023, 12, e0128722. [Google Scholar] [CrossRef] [PubMed]
- Kholodilov, I.S. , Litov, A.G., Klimentov, A.S., Belova, O.A., Polienko, A.E., Nikitin, N.A., Shchetinin, A.M., Ivannikova, A.Y., Bell–Sakyi, L., Yakovlev, A.S., et al. Isolation and characterisation of Alongshan virus in Russia. Viruses. 2020, 12, 362. [Google Scholar] [CrossRef]
- Kholodilov, I.S., Belova, O.A., Morozkin, E.S., Litov, A.G., Ivannikova, A.Y., Makenov, M.T., Shchetinin, A.M., Aibulatov, S.V., Bazarova, G.K., Bell–Sakyi, L., et al. Geographical and tick–dependent distribution of flavi–like Alongshan and Yanggou tick viruses in Russia. Viruses. 2021, 13, 458. [CrossRef]
- Kholodilov, I.S., Belova, O.A., Ivannikova, A.Y., Gadzhikurbanov, M.N., Makenov, M.T., Yakovlev, A.S., Polienko, A.E., Dereventsova, A.V., Litov, A.G., Gmyl, L.V., et al. Distribution and characterisation of tick–borne flavi–, flavi–like, and phenuiviruses in the Chelyabinsk Region of Russia. Viruses. 2022, 14, 2699. [CrossRef]
- Kartashov, M.Yu., Krivosheina, E.I., Kurushina, V.Yu., Moshkin, A.B., Khankhareev, S.S., Biche–ool, C.R., Pelevina, O.N., Popov, N.V., Bogomazova, O.L., Ternovoi, V.A. Prevalence and genetic diversity of the Alongshan virus (Flaviviridae) circulating in ticks in the south of Eastern Siberia. Problems of Virology (Voprosy Virusologii). 2024, 69(2), 151–161. (in Russian). [CrossRef]
- Tamura, K.; Nei, M. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol. Biol. Evol. 1993, 10, 512–526. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef] [PubMed]
- Ladner, J.T. , Wiley, M.R., Beitzel, B., Auguste, A.J., Dupuis, A.P., Lindquist, M.E., Sibley, S.D., Kota, K.P., Fetterer, D., Eastwood, G., et al. A Multicomponent Animal Virus Isolated from Mosquitoes. Cell Host Microbe. 2016, 20(3), 357–367. [Google Scholar] [CrossRef] [PubMed]
- Souza, W.M., Fumagalli, M.J., Torres Carrasco, A.O., Romeiro, M.F., Modha, S., Seki, M.C., Gheller, J.M., Daffre, S., Nunes, M.R.T., Murcia, P.R., et al. Viral diversity of Rhipicephalus microplus parasitizing cattle in southern Brazil. Sci Rep. 2018, 8, 16315. [CrossRef]
- Emmerich, P. , Jakupi, X., von Possel, R., Berisha, L., Halili, B., Günther, S., Cadar, D., Ahmeti, S., Schmidt–Chanasit, J. Viral metagenomics, genetic and evolutionary characteristics of Crimean–Congo hemorrhagic fever orthonairovirus in humans, Kosovo. Infect. Genet. Evol, 65. [CrossRef]
- Su, S., Cui, M.Y., Xing, L.L, Gao, R.J., Mu, L., Hong, M., Guo, Q.Q., Ren, H., Yu, J.F., Si, X.Y., Eerde, M. Metatranscriptomic analysis reveals the diversity of RNA viruses in ticks in Inner Mongolia, China. PLoS Negl Trop Dis. 2024, 18, e0012706. [CrossRef]
- Xu, W., Wang, W., Li, L., Li, N., Liu, Z., Che, L., Wang, G., Zhang, K., Feng, X., Wang, W.J., et al. Alongshan Virus Infection in Rangifer tarandus Reindeer, Northeastern China. Emerg Infect Dis. 2024, 30, 1434–1437. [CrossRef]
- Kartashov, M.Y., Krivosheina, E.I., Naidenova, E.V., Zakharov, K.S., Shvalov, A.N., Boumbaly, S., Ternovoi, V.A., Loktev, V.B. Simultaneous Detection and Genome Analysis of the Kindia Tick Virus in Cattle and Rhipicephalus Ticks in the Republic of Guinea. Vector Borne Zoonotic Dis. 2025, 25(7), 470–475. [CrossRef]
- Ternovoi, V.A., Gladysheva, A.V., Sementsova, A.O., Zaykovskaya, A.V., Volynkina, A.S., Kotenev, E.S., Agafonov, A.P., Loktev, V.B. Detection of the RNA for new multicomponent virus in patients with Crimean-Congo hemorrhagic fever in southern Russia. Annals of the Russian academy of medical sciences. 2020, 75(2):129-134. (in Russian). [CrossRef]
- Korobitsyn, I.G., Moskvitina, N.S., Tyutenkov, O.Y., Gashkov, S.I., Kononova, Y.V., Moskvitin, S.S., Romanenko, V.N., Mikryukova, T.P., Protopopova, E.V., Kartashov, M.Y., et al. Detection of tick–borne pathogens in wild birds and their ticks in Western Siberia and high level of their mismatch. Folia Parasitol (Praha). 2021, 68: 024. [CrossRef]
- Kartashov, M.Y., Gladysheva, A.V., Shvalov, A.N., Tupota, N.L., Chernikova, A.A., Ternovoi, V.A., Loktev, V.B. Novel Flavi–like virus in ixodid ticks and patients in Russia. Ticks Tick Borne Dis. 2023, 14(2). 102101. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




