Submitted:
20 September 2025
Posted:
23 September 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
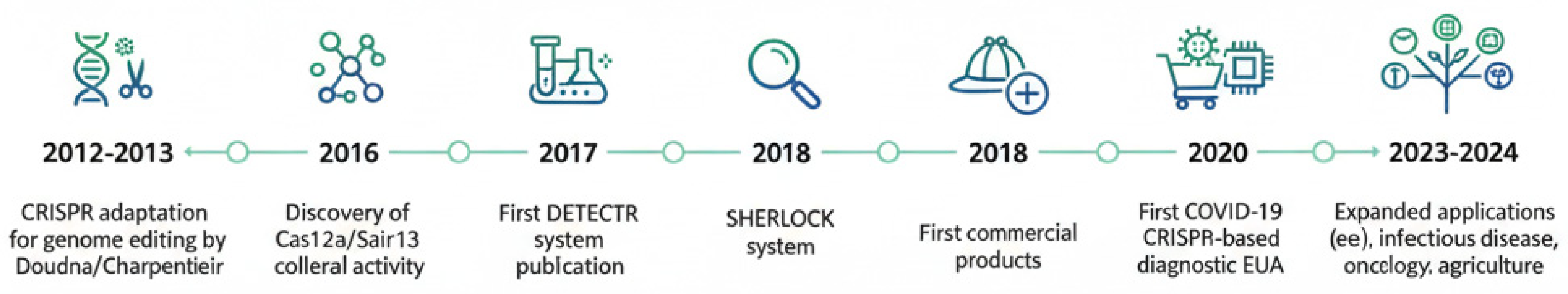
2. Core Mechanisms of CRISPR-Based Detection
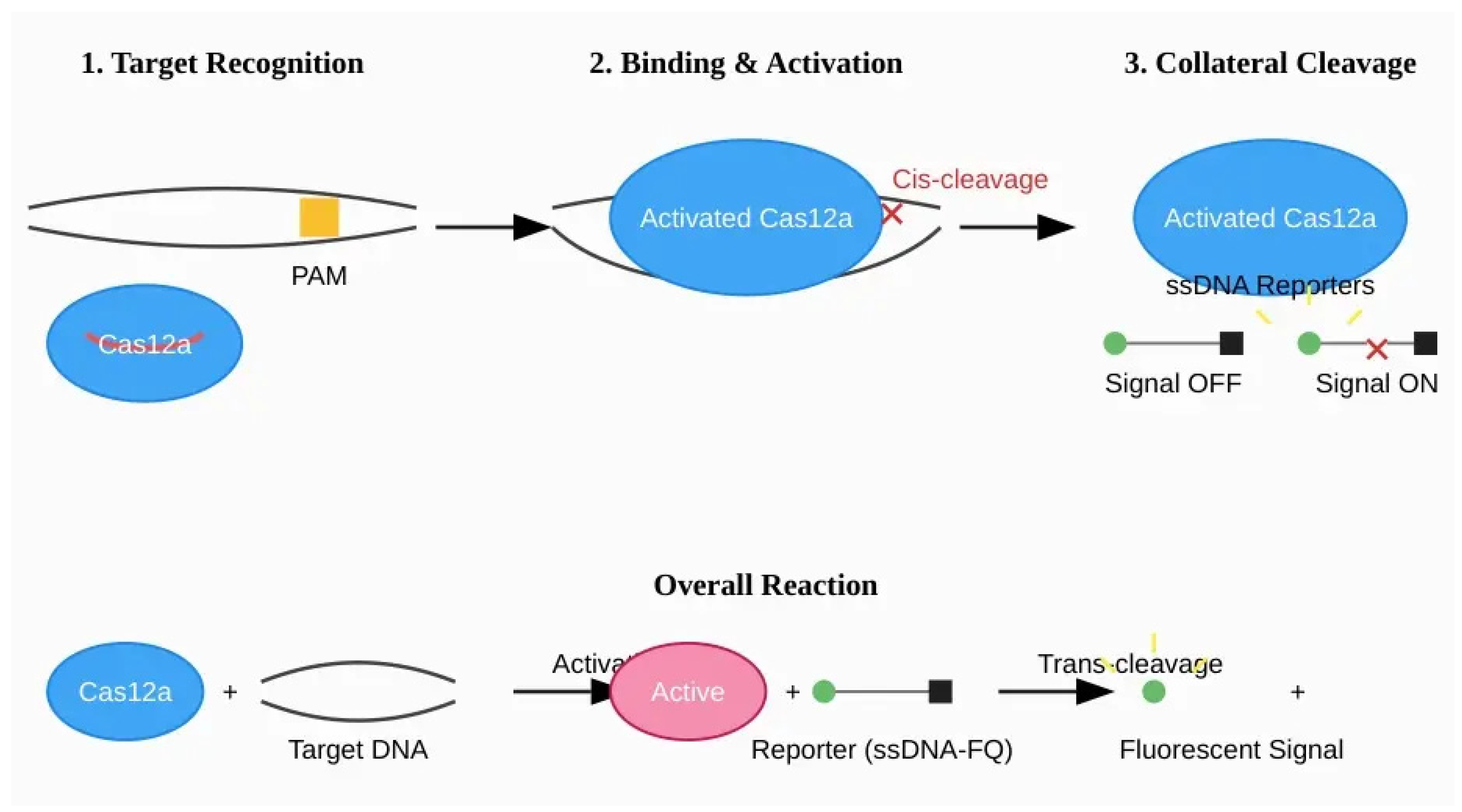
2.1. The Cas12 Family (e.g., DETECTR): DNA-Targeted Collateral Cleavage
- Cis-cleavage: The specific, staggered cleavage of both strands of the target dsDNA molecule.
- Trans-cleavage (collateral activity): Once activated, the Cas12a RNP becomes a non-specific DNase, indiscriminately cleaving any single-stranded DNA (ssDNA) molecules in its vicinity. This collateral activity is a high-turnover process, with a single target recognition event triggering the cleavage of thousands of bystander ssDNA molecules.
2.2. The Cas13 Family (e.g., SHERLOCK): RNA-Targeted Collateral Cleavage
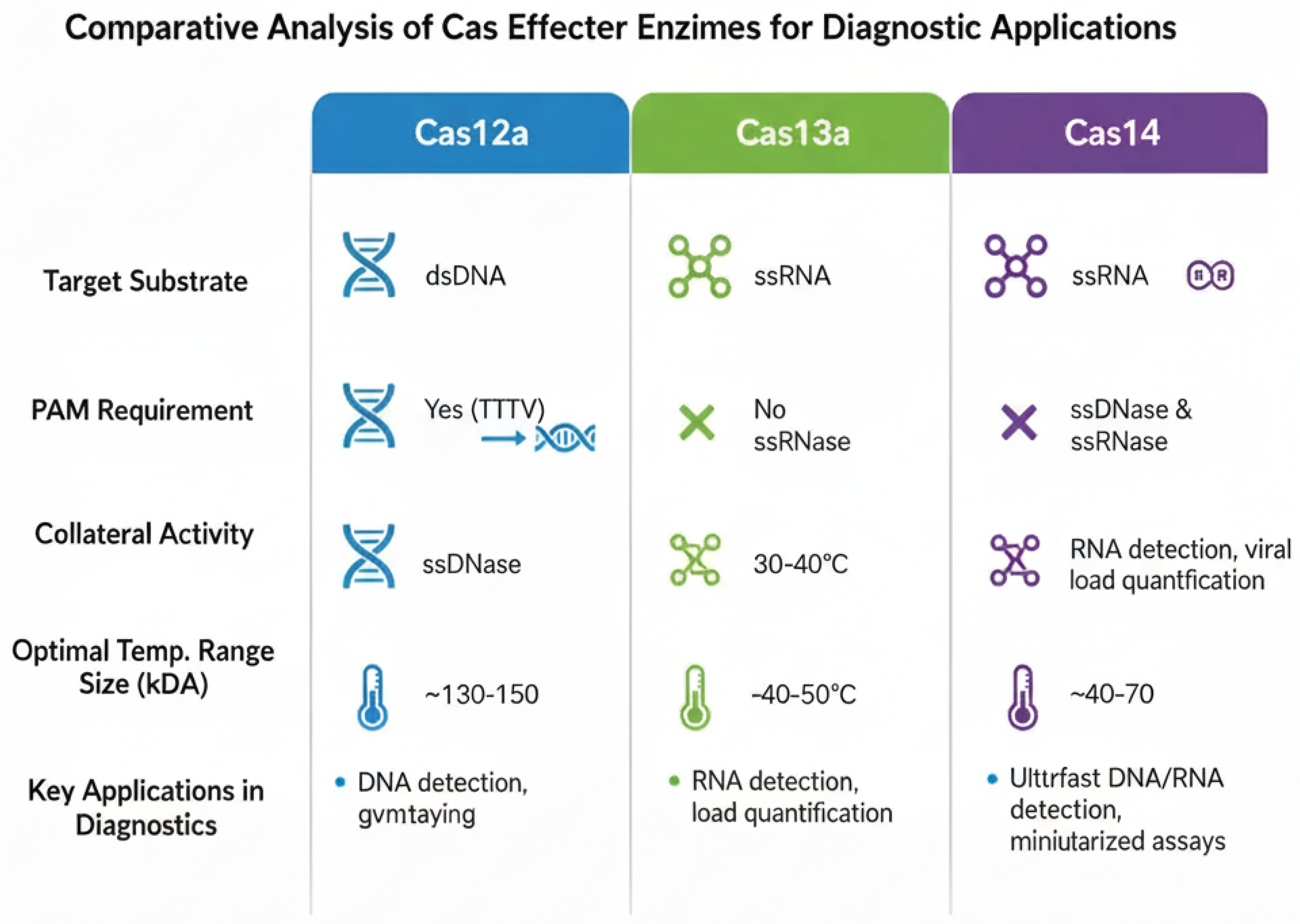
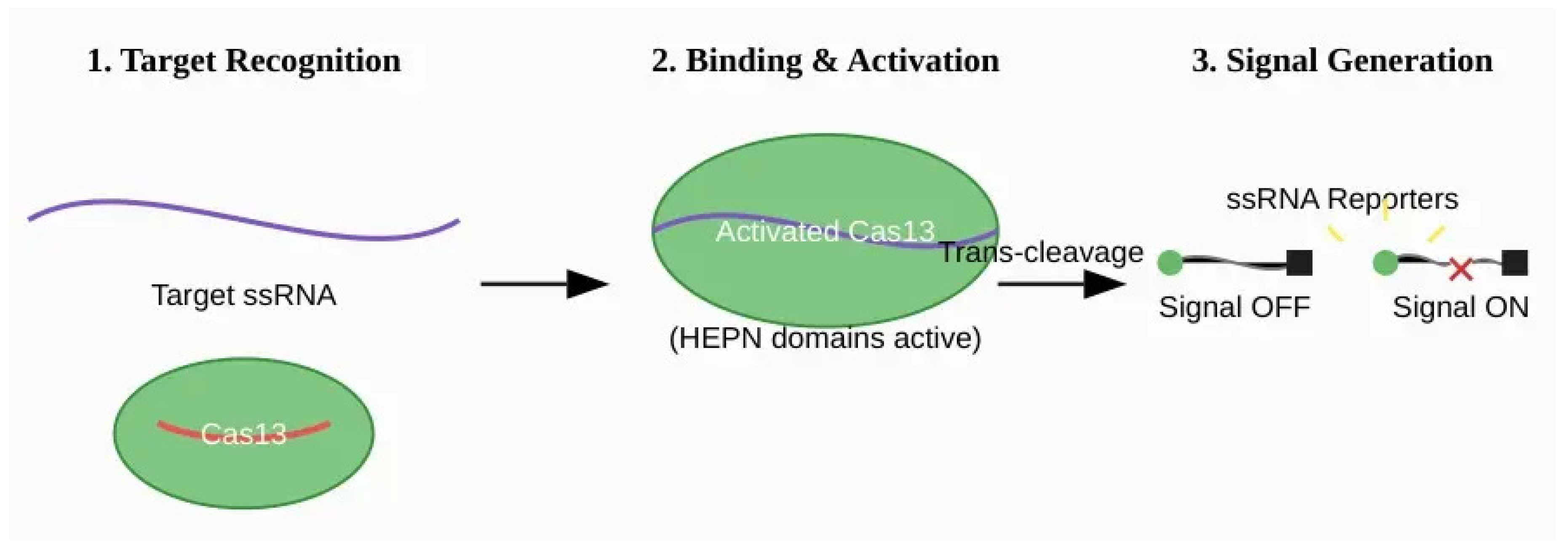
2.3. Signal Amplification and Readout
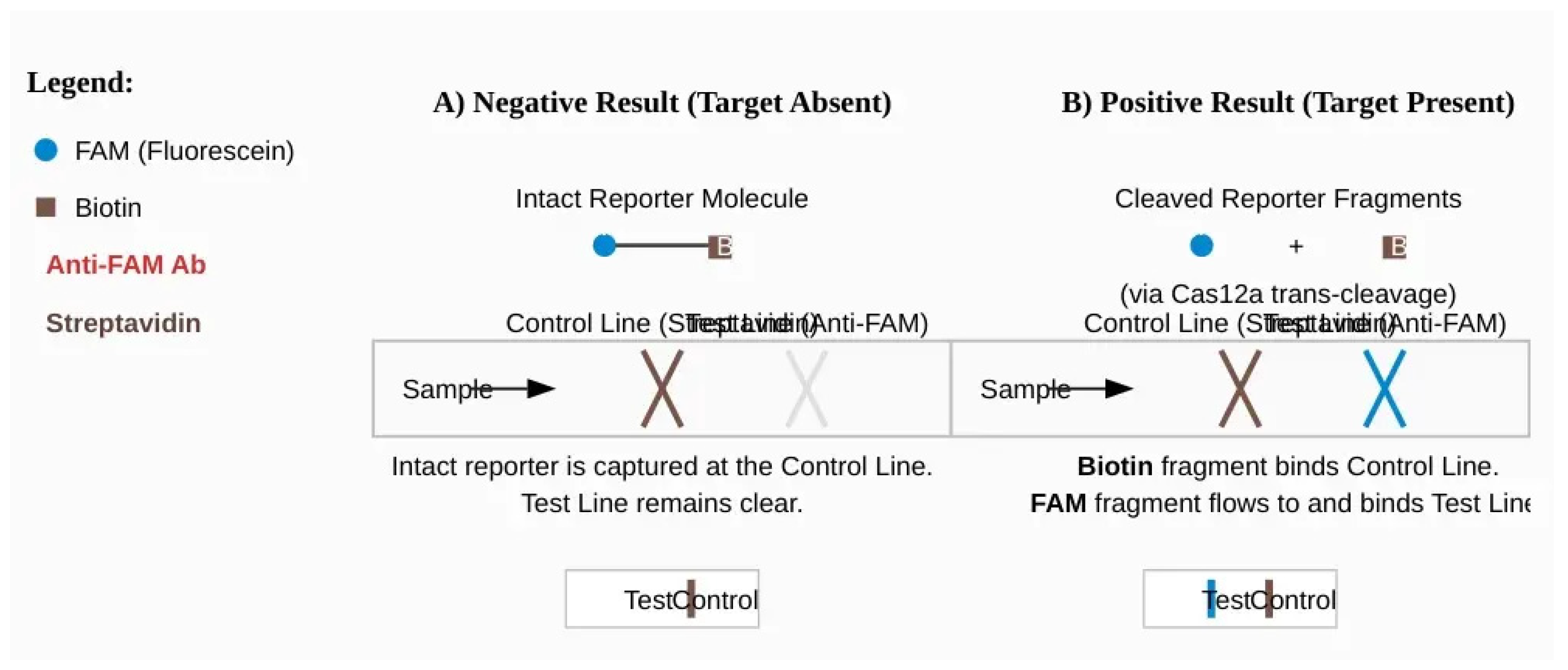
2.4. Isothermal Pre-Amplification: Achieving Attomolar Sensitivity
| Feature | Recombinase Polymerase Amplification (RPA) | Loop-Mediated Isothermal Amplification (LAMP) |
| Operating Temperature | 37–42°C (low, body temperature range) | 60–65°C (higher, requires simple heating block) |
| Reaction Time | Fast (typically <20 minutes) | Rapid (15-60 minutes) |
| Primer Design | Moderate complexity (one pair, 30-35 nt) | High complexity (4-6 specialized primers) |
| Specificity | High (based on 2 primers) | Very High (based on 6-8 target regions) |
| Key Advantage | Speed and low temperature operation | High efficiency and robustness against inhibitors |
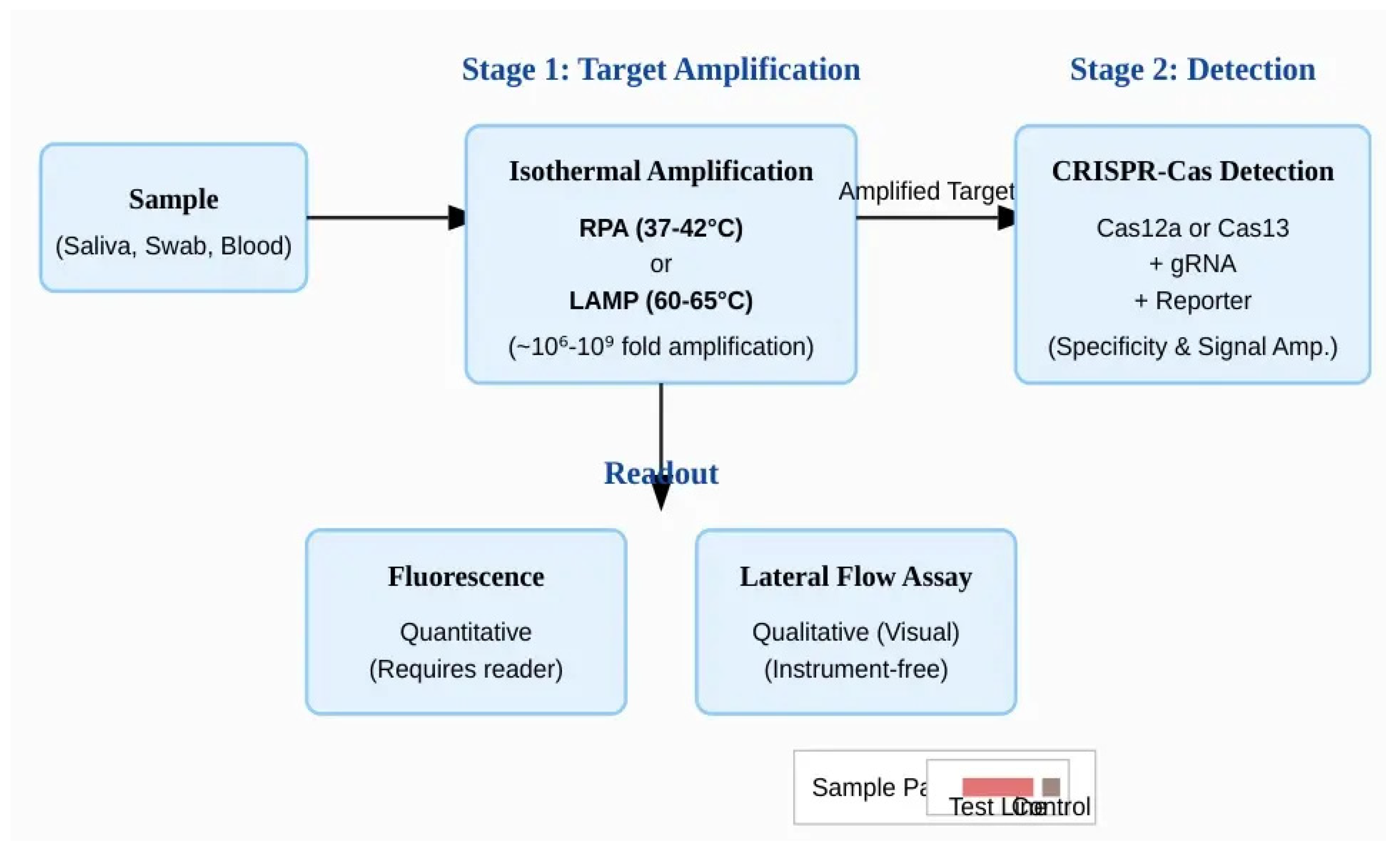
3. Key Applications in Global Health and Infectious Disease
3.1. Viral Pandemics: A Proving Ground for Rapid Response
3.2. Endemic Diseases
3.3. Antimicrobial Resistance (AMR)
3.4. Other Applications
4. Bridging the Gap: Overcoming Hurdles to Widespread Adoption
4.1. Technical and Logistical Challenges
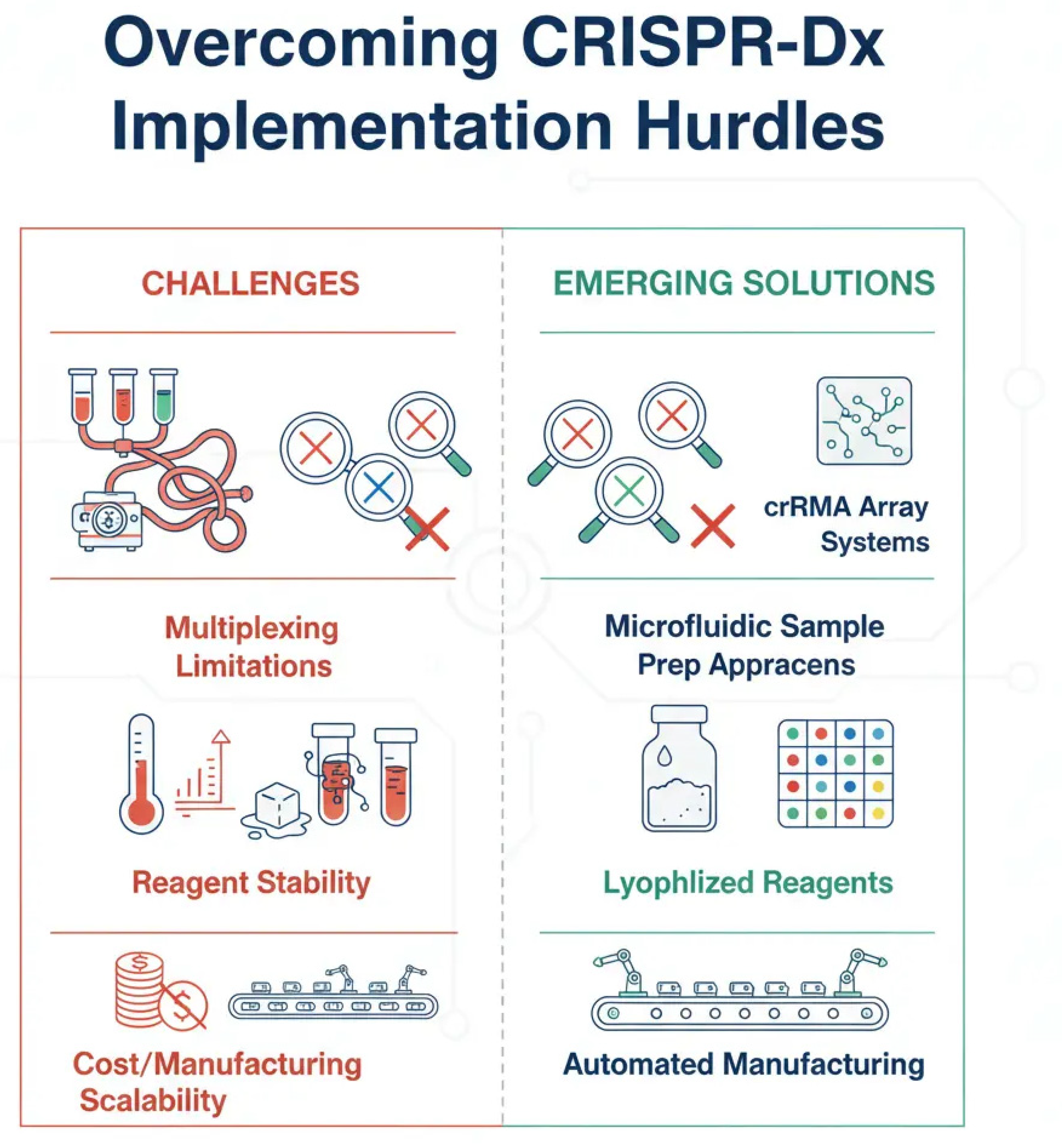
4.2. Regulatory and Commercial Landscape.
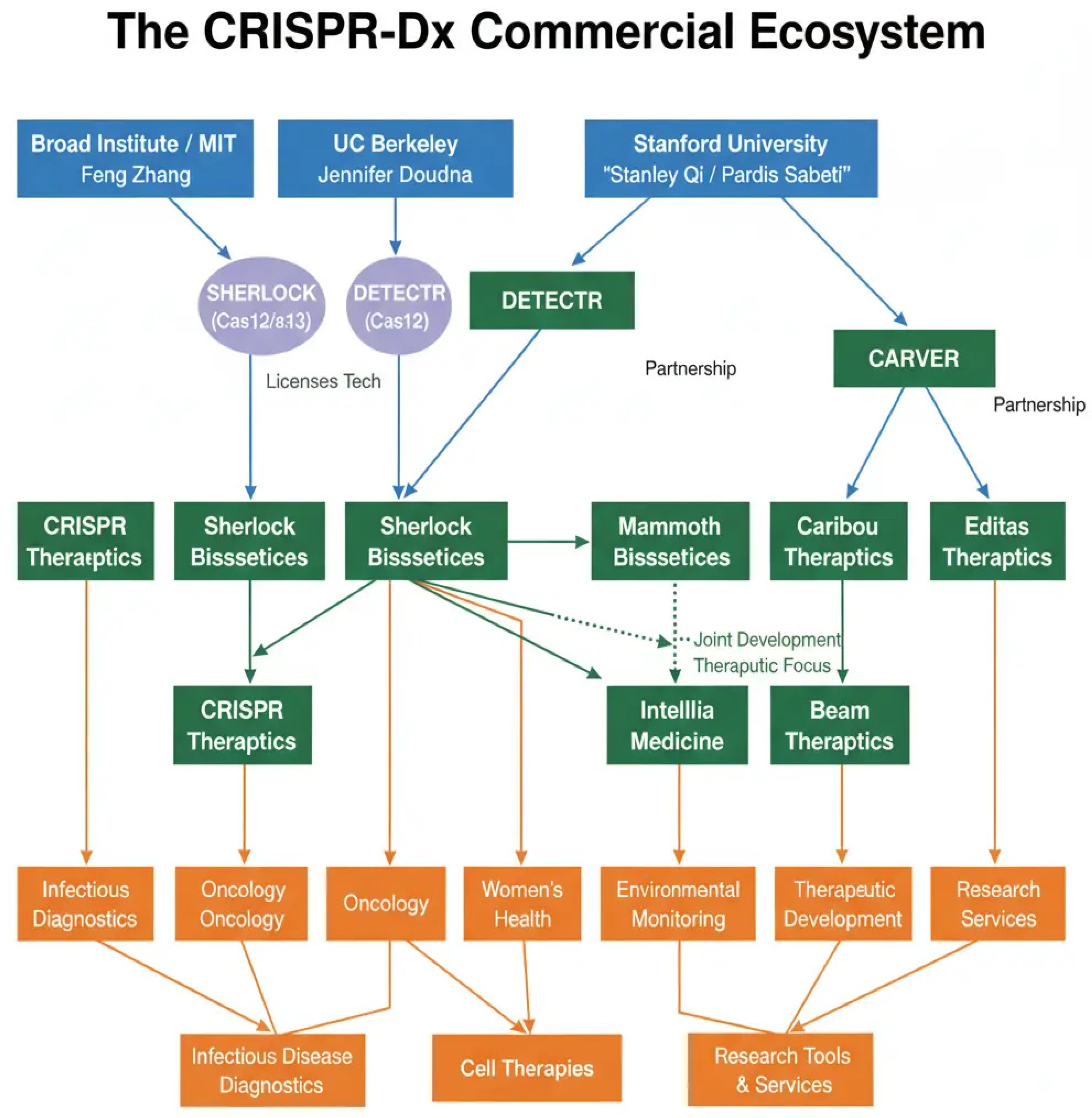
5. Concluding Remarks and Future Perspectives
References
- Unemo M, van der Veen S, Rösth O, et al. The challenge of introducing and implementing new and rapid point-of-care tests for sexually transmitted infections. PLoS Med. 2021;18(6):e1003655.
- World Health Organization. (2021). Diagnostic testing for SARS-CoV-2. WHO.
- World Health Organization. (2006). Sexually Transmitted Diseases Diagnostics Initiative (SDI): ASSURED criteria. WHO.
- Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 2012;337(6096):816-821.
- Myhrvold C, Freije CA, Gootenberg JS, et al. Field-deployable viral diagnostics using CRISPR-Cas13. Science. 2018;360(6387):444-448.
- Chen JS, Ma E, Harrington LB, et al. CRISPR-Cas12a target binding unleashes indiscriminate single-stranded DNase activity. Science. 2018;360(6387):436-439.
- Zetsche B, Gootenberg JS, Abudayyeh OO, et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell. 2015;163(3):759-771.
- Li SY, Cheng QX, Wang JM, et al. CRISPR-Cas12a-assisted nucleic acid detection. Cell Discov. 2018;4:20.
- Ramachandran A, Huyke DA, Sharma E, Sotomayor M, Santiago JG. Enzyme kinetics of CRISPR molecular diagnostics. Anal Chem. 2021;93(10):4341-4349.
- Gootenberg JS, Abudayyeh OO, Lee JW, et al. Multiplexed and portable nucleic acid detection platform with Cas13, Cas12a, and Csm6. Science. 2018;360(6387):439-444.
- Abudayyeh OO, Gootenberg JS, Konermann S, et al. RNA targeting with CRISPR-Cas13. Nature. 2017;550(7675):280-284.
- Feng W, et al. Signal Amplification by the trans-Cleavage Activity of CRISPR-Cas Systems: Kinetics and Performance. Anal Chem. 2023;95(1):401-409.
- Kaminski MM, Abudayyeh OO, Gootenberg JS, et al. CRISPR-based diagnostics. Nat Biomed Eng. 2021;5(7):643-656.
- Broughton JP, et al. CRISPR–Cas12-based detection of SARS-CoV-2. Nat Biotechnol. 2020;38(7):871-875.
- Cheng M, et al. Chain hybridization-based CRISPR-lateral flow assay for sensitive and specific detection of nucleic acids. Anal Chim Acta. 2023;1277:341778.
- Kaminski MM, Abudayyeh OO, Gootenberg JS, et al. CRISPR-based diagnostics. Nat Biomed Eng. 2021;5(7):643-656.
- Lobato IM, O’Sullivan DM. Recombinase polymerase amplification: Basics and applications. Adv Clin Chem. 2017;80:1-35. https://www.sciencedirect.com/science/article/pii/S006524231730030X.
- Notomi T, et al. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 2000;28(12):e63.
- Chen L, Hu M, Zhou X. Trends in developing one-pot CRISPR diagnostics strategies. Trends Biotechnol. 2024;42(5):495-507. [CrossRef]
- Broughton JP, et al. CRISPR–Cas12-based detection of SARS-CoV-2. Nat Biotechnol. 2020;38(7):871-875.
- Joung J, et al. Detection of SARS-CoV-2 with SHERLOCK One-Pot Testing. N Engl J Med. 2020;383(15):1492-1494.
- Broughton JP, et al. CRISPR–Cas12-based detection of SARS-CoV-2. Nat Biotechnol. 2020;38(7):871-875. https://www.nature.com/articles/s41587-020-0502-2.
- Joung J, et al. Point-of-care testing for COVID-19 using SHERLOCK diagnostics. Med. 2020;1(1):101-106.
- Qiu X, et al. An extraction-free one-step CRISPR-assisted detection platform and a potential Streptococcus pneumoniae at-home self-testing kit. Biosens Bioelectron. 2023;223:115001.
- Pardee K, et al. Rapid, Low-Cost Detection of Zika Virus Using Programmable Biomolecular Components. Cell. 2016;165(5):1255-1266.
- Zhang Y, et al. A one-pot method for universal Dengue virus detection by combining RT-RPA amplification and CRISPR/Cas12a assay. BMC Microbiol. 2024;24(1):1-13. [CrossRef]
- Li X, et al. Highly sensitive and rapid point-of-care testing for HIV-1 infection based on CRISPR-Cas13a system. BMC Infect Dis. 2023;23(1):577.
- Lee RA, et al. Ultrasensitive CRISPR-based diagnostic for field-applicable detection of Plasmodium species in symptomatic and asymptomatic malaria. Proc Natl Acad Sci U S A. 2020;117(40):24922-24931.
- Ai JW, et al. CRISPR-based rapid and ultra-sensitive diagnostic test for Mycobacterium tuberculosis. Emerg Microbes Infect. 2019;8(1):131-141.
- Di Carlo E, et al. State of the art CRISPR-based strategies for cancer diagnostics and treatment. Biomarker Research. 2024;12(1):1-19.
- Kaminski MM, Abudayyeh OO, Gootenberg JS, et al. CRISPR-based diagnostics. Nat Biomed Eng. 2021;5(7):643-656.
- Tao S, et al. The Application of the CRISPR-Cas System in Antibiotic Resistance. Infect Drug Resist. 2022;15:4591-4602.
- Wani AK, et al. CRISPR/Cas12a-based biosensors for environmental monitoring and diagnostics. Environ Technol Innov. 2024;34:103625.
- Li Y, et al. CRISPR-Cas-based detection for food safety problems: Current status, challenges, and opportunities. Compr Rev Food Sci Food Saf. 2022;21(4):3521-3543.
- Tanny T, et al. CRISPR/Cas-Based Diagnostics in Agricultural Applications. J Agric Food Chem. 2023;71(30):11087-11099.
- Ghouneimy A, et al. CRISPR-Based Diagnostics: Challenges and Potential Solutions Toward Point-of-Care Applications. ACS Synth Biol. 2022;11(1):11-29.
- Ghouneimy A, et al. CRISPR-Based Diagnostics: Challenges and Potential Solutions Toward Point-of-Care Applications. ACS Synth Biol. 2022;11(1):11-29.
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




