Submitted:
16 September 2025
Posted:
17 September 2025
You are already at the latest version
Abstract
Keywords:
Cancer as an Evolving Ecosystem of Hallmark Programs
The Spatial Organization of Hallmarks
The Temporal Evolution of Hallmarks
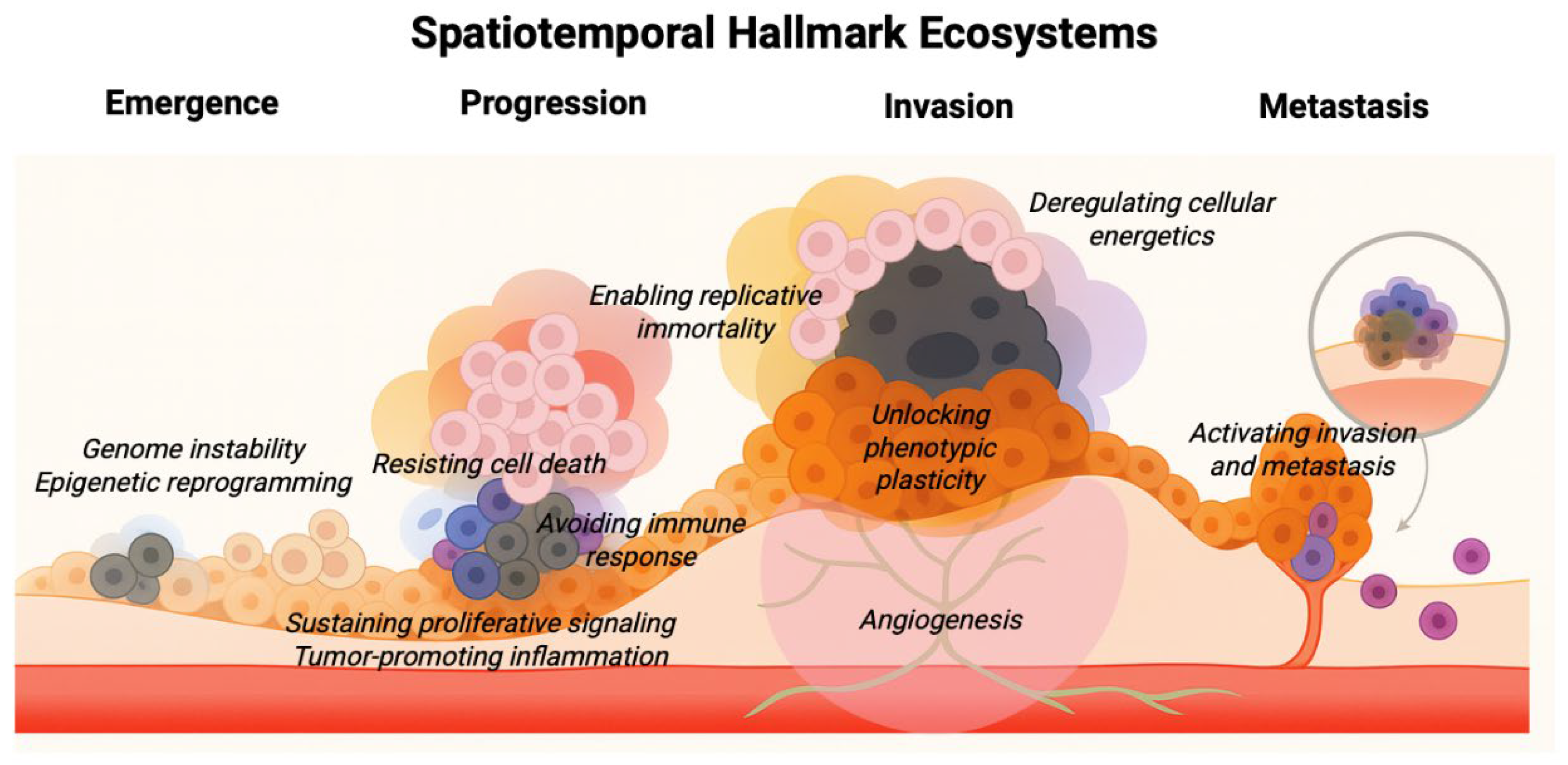
Spatiotemporal Hallmark Ecosystems: A Conceptual Framework
- Oncogenic mutations in normal tissue
- Intratumoral heterogeneity
- Therapeutic resistance
- Transformation events
- Metastatic colonization
Current Technical and Conceptual Limitations
Charting Hallmarks in Motion: Implications for Cancer Research
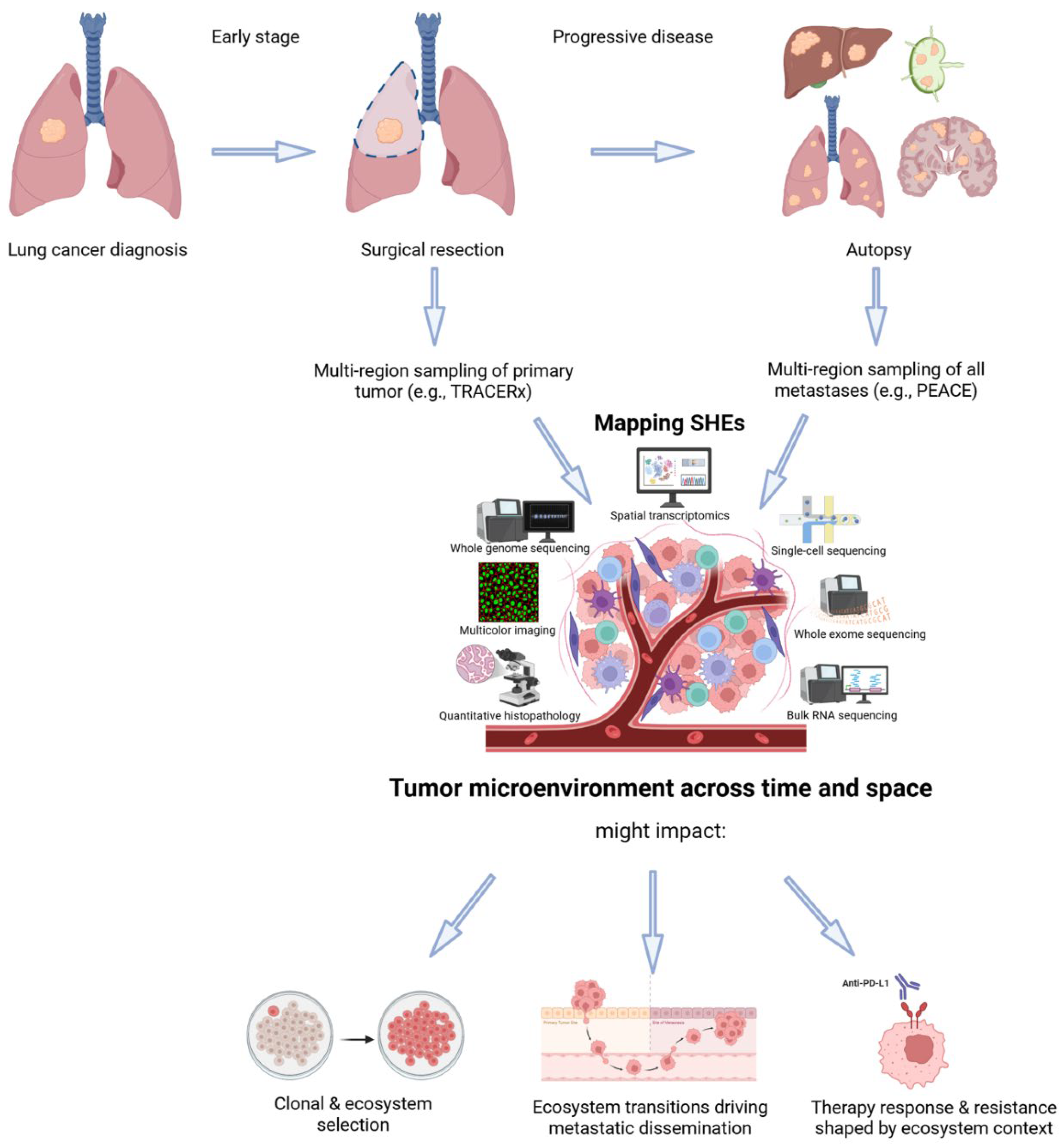
From Maps to Models: Clinical Applications of SHEs
Conclusions: A New Lens on Cancer Complexity
- Prediction 1: Therapeutic resistance is an emergent property of the tumour ecosystem, not just a feature of individual clones: If resistance arises from the interaction of tumour cells with their microenvironment, then ablating a resistant clone alone should not lead to durable response unless its supporting ecosystem is also disrupted.
- Prediction 2: Subclonal hallmark specialization reflects ecological trade-offs rather than purely genetic divergence: In tumours with high subclonal diversity, hallmark programs (e.g., proliferation, immune evasion) should segregate across clones in a mutually exclusive fashion, even when those clones share a similar genetic background.
- Prediction 3: Transformation to malignancy requires the spatial coordination of multiple hallmark programs: Premalignant lesions exhibiting spatially co-localized hallmark traits should show higher progression rates than lesions in which these traits remain uncoordinated.
- Prediction 4: The degree of spatial complementarity between tumour and TME hallmark programs predicts treatment response: Tumours with high cross-compartment coordination (e.g., evading growth suppressors in tumour cells coupled with sustained proliferative signaling from the TME) should respond more favorably to therapies that target both compartments.
- Prediction 5: Phenotypic divergence among genetically identical clones arises from local niche conditions, and can be reversed: Isogenic clones located in different spatial niches should exhibit divergent hallmark activity, which should homogenize if their niche environment is disrupted (e.g., through fibroblast depletion or cytokine blockade).
Acknowledgements
Conflicts of Interest
References
- Hanahan D, Weinberg RA. The hallmarks of cancer. Cell. 2000;100: 57–70.
- Weinberg, R.A. Coming Full Circle—From Endless Complexity to Simplicity and Back Again. Cell 2014, 157, 267–271. [Google Scholar] [CrossRef]
- Swanton, C.; Bernard, E.; Abbosh, C.; André, F.; Auwerx, J.; Balmain, A.; Bar-Sagi, D.; Bernards, R.; Bullman, S.; DeGregori, J.; et al. Embracing cancer complexity: Hallmarks of systemic disease. Cell 2024, 187, 1589–1616. [Google Scholar] [CrossRef]
- Nowell, P.C. The clonal evolution of tumor cell populations. Science 1976, 194, 23–28. [Google Scholar] [CrossRef]
- Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144: 646–674.
- Elhanani, O.; Ben-Uri, R.; Keren, L. Spatial profiling technologies illuminate the tumor microenvironment. Cancer Cell 2023, 41, 404–420. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D. Hallmarks of Cancer: New Dimensions. Cancer Discov. 2022, 12, 31–46. [Google Scholar] [CrossRef] [PubMed]
- Burkhardt, D.B.; Juan, B.P.S.; Lock, J.G.; Krishnaswamy, S.; Chaffer, C.L. Mapping Phenotypic Plasticity upon the Cancer Cell State Landscape Using Manifold Learning. Cancer Discov. 2022, 12, 1847–1859. [Google Scholar] [CrossRef] [PubMed]
- Flavahan, W.A.; Gaskell, E.; Bernstein, B.E. Epigenetic plasticity and the hallmarks of cancer. Science 2017, 357, eaal2380. [Google Scholar] [CrossRef]
- Gavish, A.; Tyler, M.; Greenwald, A.C.; Hoefflin, R.; Simkin, D.; Tschernichovsky, R.; Darnell, N.G.; Somech, E.; Barbolin, C.; Antman, T.; et al. Hallmarks of transcriptional intratumour heterogeneity across a thousand tumours. Nature 2023, 618, 598–606. [Google Scholar] [CrossRef]
- Oliveira MF de, Romero JP, Chung M, Williams SR, Gottscho AD, Gupta A, et al. High-definition spatial transcriptomic profiling of immune cell populations in colorectal cancer. Nat Genet. 2025;57: 1512–1523.
- Janesick, A.; Shelansky, R.; Gottscho, A.D.; Wagner, F.; Williams, S.R.; Rouault, M.; Beliakoff, G.; Morrison, C.A.; Oliveira, M.F.; Sicherman, J.T.; et al. High resolution mapping of the tumor microenvironment using integrated single-cell, spatial and in situ analysis. Nat. Commun. 2023, 14, 1–15. [Google Scholar] [CrossRef]
- Lucas, O.; Ward, S.; Zaidi, R.; Bunkum, A.; Frankell, A.M.; Moore, D.A.; Hill, M.S.; Liu, W.K.; Marinelli, D.; Lim, E.L.; et al. Characterizing the evolutionary dynamics of cancer proliferation in single-cell clones with SPRINTER. Nat. Genet. 2024, 57, 103–114. [Google Scholar] [CrossRef]
- Floor, S.L.; Dumont, J.E.; Maenhaut, C.; Raspe, E. Hallmarks of cancer: of all cancer cells, all the time? Trends Mol. Med. 2012, 18, 509–515. [Google Scholar] [CrossRef]
- Sibai, M.; Cervilla, S.; Grases, D.; Musulen, E.; Lazcano, R.; Mo, C.-K.; Davalos, V.; Fortian, A.; Bernat, A.; Romeo, M.; et al. The spatial landscape of cancer hallmarks reveals patterns of tumor ecological dynamics and drug sensitivity. Cell Rep. 2025, 44, 115229. [Google Scholar] [CrossRef] [PubMed]
- Arora, R.; Cao, C.; Kumar, M.; Sinha, S.; Chanda, A.; McNeil, R.; Samuel, D.; Arora, R.K.; Matthews, T.W.; Chandarana, S.; et al. Spatial transcriptomics reveals distinct and conserved tumor core and edge architectures that predict survival and targeted therapy response. Nat. Commun. 2023, 14, 1–19. [Google Scholar] [CrossRef]
- Somarelli, J.A. The Hallmarks of Cancer as Ecologically Driven Phenotypes. Front. Ecol. Evol. 2021, 9. [Google Scholar] [CrossRef]
- Skrupskelyte G, Rojo Arias JE, Dang Y, Han S, Bejar MT, Colom B, et al. Pre-cancerous Niche Remodelling Dictates Nascent Tumour Survival. bioRxiv. 2024. p. 2024.07.04.602022. [CrossRef]
- Kang, J.; Lee, J.H.; Cha, H.; An, J.; Kwon, J.; Lee, S.; Kim, S.; Baykan, M.Y.; Kim, S.Y.; An, D.; et al. Systematic dissection of tumor-normal single-cell ecosystems across a thousand tumors of 30 cancer types. Nat. Commun. 2024, 15, 1–17. [Google Scholar] [CrossRef]
- Zhou, D.C.; Jayasinghe, R.G.; Chen, S.; Herndon, J.M.; Iglesia, M.D.; Navale, P.; Wendl, M.C.; Caravan, W.; Sato, K.; Storrs, E.; et al. Spatially restricted drivers and transitional cell populations cooperate with the microenvironment in untreated and chemo-resistant pancreatic cancer. Nat. Genet. 2022, 54, 1390–1405. [Google Scholar] [CrossRef]
- Bhattacharya, R.; Avdieiev, S.S.; Bukkuri, A.; Whelan, C.J.; Gatenby, R.A.; Tsai, K.Y.; Brown, J.S. The Hallmarks of Cancer as Eco-Evolutionary Processes. Cancer Discov. 2025, 15, 685–701. [Google Scholar] [CrossRef]
- Grande E, Sibai M, Andrada E, Grases D, Reig O, Escobosa M, et al. Spatial biomarkers of response to neoadjuvant therapy in muscle-invasive bladder cancer: the DUTRENEO trial. medRxiv. 2025. [CrossRef]
- Colom, B.; Alcolea, M.P.; Piedrafita, G.; Hall, M.W.J.; Wabik, A.; Dentro, S.C.; Fowler, J.C.; Herms, A.; King, C.; Ong, S.H.; et al. Spatial competition shapes the dynamic mutational landscape of normal esophageal epithelium. Nat. Genet. 2020, 52, 604–614. [Google Scholar] [CrossRef]
- Ciwinska, M.; Messal, H.A.; Hristova, H.R.; Lutz, C.; Bornes, L.; Chalkiadakis, T.; Harkes, R.; Langedijk, N.S.M.; Hutten, S.J.; Menezes, R.X.; et al. Mechanisms that clear mutations drive field cancerization in mammary tissue. Nature 2024, 633, 198–206. [Google Scholar] [CrossRef] [PubMed]
- Chu, X.; Tian, Y.; Lv, C. Decoding the spatiotemporal heterogeneity of tumor-associated macrophages. Mol. Cancer 2024, 23, 1–24. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Sinjab, A.; Min, J.; Han, G.; Paradiso, F.; Zhang, Y.; Wang, R.; Pei, G.; Dai, Y.; Liu, Y.; et al. Conserved spatial subtypes and cellular neighborhoods of cancer-associated fibroblasts revealed by single-cell spatial multi-omics. Cancer Cell 2025, 43, 905–924.e6. [Google Scholar] [CrossRef]
- Tirosh, I.; Suva, M.L. Cancer cell states: Lessons from ten years of single-cell RNA-sequencing of human tumors. Cancer Cell 2024, 42, 1497–1506. [Google Scholar] [CrossRef]
- Lomakin, A.; Svedlund, J.; Strell, C.; Gataric, M.; Shmatko, A.; Rukhovich, G.; Park, J.S.; Ju, Y.S.; Dentro, S.; Kleshchevnikov, V.; et al. Spatial genomics maps the structure, nature and evolution of cancer clones. Nature 2022, 611, 594–602. [Google Scholar] [CrossRef]
- Tabassum, D.P.; Polyak, K. Tumorigenesis: it takes a village. Nat. Rev. Cancer 2015, 15, 473–483. [Google Scholar] [CrossRef]
- Shi, H.; Williams, M.J.; Satas, G.; Weiner, A.C.; McPherson, A.; Shah, S.P. Allele-specific transcriptional effects of subclonal copy number alterations enable genotype-phenotype mapping in cancer cells. Nat. Commun. 2024, 15, 1–13. [Google Scholar] [CrossRef]
- Campbell, N.R.; Rao, A.; Hunter, M.V.; Sznurkowska, M.K.; Briker, L.; Zhang, M.; Baron, M.; Heilmann, S.; Deforet, M.; Kenny, C.; et al. Cooperation between melanoma cell states promotes metastasis through heterotypic cluster formation. Dev. Cell 2021, 56, 2808–2825.e10. [Google Scholar] [CrossRef] [PubMed]
- Adler, M.; Chavan, A.R.; Medzhitov, R. Tissue Biology: In Search of a New Paradigm. Annu. Rev. Cell Dev. Biol. 2023, 39, 67–89. [Google Scholar] [CrossRef]
- Roehrig, A.; Hirsch, T.Z.; Pire, A.; Morcrette, G.; Gupta, B.; Marcaillou, C.; Imbeaud, S.; Chardot, C.; Gonzales, E.; Jacquemin, E.; et al. Single-cell multiomics reveals the interplay of clonal evolution and cellular plasticity in hepatoblastoma. Nat. Commun. 2024, 15, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Dujon, A.M.; Aktipis, A.; Alix-Panabières, C.; Amend, S.R.; Boddy, A.M.; Brown, J.S.; Capp, J.; DeGregori, J.; Ewald, P.; Gatenby, R.; et al. Identifying key questions in the ecology and evolution of cancer. Evol. Appl. 2020, 14, 877–892. [Google Scholar] [CrossRef] [PubMed]
- Gourmet L, Ramazzoti D, Mallick P, Walker-Samuel S, Zapata L. The temporal evolution of cancer hallmarks. bioRxiv. 2024. [CrossRef]
- Gerlinger, M.; Rowan, A.J.; Horswell, S.; Math, M.; Larkin, J.; Endesfelder, D.; Gronroos, E.; Martinez, P.; Matthews, N.; Stewart, A.; et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 2012, 366, 883–892. [Google Scholar] [CrossRef] [PubMed]
- Jamal-Hanjani, M.; Wilson, G.A.; McGranahan, N.; Birkbak, N.J.; Watkins, T.B.K.; Veeriah, S.; Shafi, S.; Johnson, D.H.; Mitter, R.; Rosenthal, R.; et al. Tracking the Evolution of Non–Small-Cell Lung Cancer. N. Engl. J. Med. 2017, 376, 2109–2121. [Google Scholar] [CrossRef]
- Gerstung M, Jolly C, Leshchiner I, Dentro SC, Gonzalez S, Rosebrock D, et al. The evolutionary history of 2,658 cancers. Nature. 2020;578: 122–128.
- Dijkstra, K.K.; Vendramin, R.; Karagianni, D.; Witsen, M.; Gálvez-Cancino, F.; Hill, M.S.; Foster, K.A.; Barbè, V.; Angelova, M.; Hynds, R.E.; et al. Subclonal immune evasion in non-small cell lung cancer. Cancer Cell 2025. [Google Scholar] [CrossRef]
- Rosenthal, R.; Cadieux, E.L.; Salgado, R.; Bakir, M.A.; Moore, D.A.; Hiley, C.T.; Lund, T.; Tanić, M.; Reading, J.L.; Joshi, K.; et al. Neoantigen-directed immune escape in lung cancer evolution. Nature 2019, 567, 479–485. [Google Scholar] [CrossRef]
- Martínez-Ruiz, C.; Black, J.R.M.; Puttick, C.; Hill, M.S.; Demeulemeester, J.; Cadieux, E.L.; Thol, K.; Jones, T.P.; Veeriah, S.; Naceur-Lombardelli, C.; et al. Genomic–transcriptomic evolution in lung cancer and metastasis. Nature 2023, 616, 543–552. [Google Scholar] [CrossRef]
- Frankell, A.M.; Dietzen, M.; Al Bakir, M.; Lim, E.L.; Karasaki, T.; Ward, S.; Veeriah, S.; Colliver, E.; Huebner, A.; Bunkum, A.; et al. The evolution of lung cancer and impact of subclonal selection in TRACERx. Nature 2023, 616, 525–533. [Google Scholar] [CrossRef]
- Al Bakir, M.; Huebner, A.; Martínez-Ruiz, C.; Grigoriadis, K.; Watkins, T.B.K.; Pich, O.; Moore, D.A.; Veeriah, S.; Ward, S.; Laycock, J.; et al. The evolution of non-small cell lung cancer metastases in TRACERx. Nature 2023, 616, 534–542. [Google Scholar] [CrossRef] [PubMed]
- Lu, W.-T.; Zalmas, L.-P.; Bailey, C.; Black, J.R.M.; Martinez-Ruiz, C.; Pich, O.; Gimeno-Valiente, F.; Usaite, I.; Magness, A.; Thol, K.; et al. TRACERx analysis identifies a role for FAT1 in regulating chromosomal instability and whole-genome doubling via Hippo signalling. Nat. Cell Biol. 2024, 27, 154–168. [Google Scholar] [CrossRef]
- Arner, E.N.; Rathmell, J.C. Metabolic programming and immune suppression in the tumor microenvironment. Cancer Cell 2023, 41, 421–433. [Google Scholar] [CrossRef] [PubMed]
- Dongre, A.; Weinberg, R.A. New insights into the mechanisms of epithelial-mesenchymal transition and implications for cancer. Nat. Rev. Mol. Cell Biol. 2018, 20, 69–84. [Google Scholar] [CrossRef]
- Gourmet, L.; Sottoriva, A.; Walker-Samuel, S.; Secrier, M.; Zapata, L. Immune evasion impacts the landscape of driver genes during cancer evolution. Genome Biol. 2024, 25, 1–15. [Google Scholar] [CrossRef]
- Semenza, G.L. HIF-1 mediates metabolic responses to intratumoral hypoxia and oncogenic mutations. J. Clin. Investig. 2013, 123, 3664–3671. [Google Scholar] [CrossRef]
- Winkler, J.; Abisoye-Ogunniyan, A.; Metcalf, K.J.; Werb, Z. Concepts of extracellular matrix remodelling in tumour progression and metastasis. Nat. Commun. 2020, 11, 5120. [Google Scholar] [CrossRef]
- Greten, F.R.; Grivennikov, S.I. Inflammation and Cancer: Triggers, Mechanisms, and Consequences. Immunity 2019, 51, 27–41. [Google Scholar] [CrossRef]
- Leonetti, A.; Sharma, S.; Minari, R.; Perego, P.; Giovannetti, E.; Tiseo, M. Resistance mechanisms to osimertinib in EGFR-mutated non-small cell lung cancer. Br. J. Cancer 2019, 121, 725–737. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.-L.; Chen, H.-H.; Zheng, L.-L.; Sun, L.-P.; Shi, L. Angiogenic signaling pathways and anti-angiogenic therapy for cancer. Signal Transduct. Target. Ther. 2023, 8, 198. [Google Scholar] [CrossRef]
- Puttick, C.; Jones, T.P.; Leung, M.M.; Galvez-Cancino, F.; Liu, J.; Varas-Godoy, M.; Rowan, A.; Pich, O.; Martinez-Ruiz, C.; Bentham, R.; et al. MHC Hammer reveals genetic and non-genetic HLA disruption in cancer evolution. Nat. Genet. 2024, 56, 2121–2131. [Google Scholar] [CrossRef]
- Shin, D.S.; Zaretsky, J.M.; Escuin-Ordinas, H.; Garcia-Diaz, A.; Hu-Lieskovan, S.; Kalbasi, A.; Grasso, C.S.; Hugo, W.; Sandoval, S.; Torrejon, D.Y.; et al. Primary Resistance to PD-1 Blockade Mediated by JAK1/2 Mutations. Cancer Discov. 2017, 7, 188–201. [Google Scholar] [CrossRef] [PubMed]
- Joller, N.; Anderson, A.C.; Kuchroo, V.K. LAG-3, TIM-3, and TIGIT: Distinct functions in immune regulation. Immunity 2024, 57, 206–222. [Google Scholar] [CrossRef] [PubMed]
- Hessey, S.; Fessas, P.; Zaccaria, S.; Jamal-Hanjani, M.; Swanton, C. Insights into the metastatic cascade through research autopsies. Trends Cancer 2023, 9, 490–502. [Google Scholar] [CrossRef]
- Fares, J.; Fares, M.Y.; Khachfe, H.H.; Salhab, H.A.; Fares, Y. Molecular principles of metastasis: a hallmark of cancer revisited. Signal Transduct. Target. Ther. 2020, 5, 28. [Google Scholar] [CrossRef]
- Geukens, T.; Maetens, M.; E Hooper, J.; Oesterreich, S.; Lee, A.V.; Miller, L.; Atkinson, J.M.; Rosenzweig, M.; Puhalla, S.; Thorne, H.; et al. Research autopsy programmes in oncology: shared experience from 14 centres across the world. J. Pathol. 2024, 263, 150–165. [Google Scholar] [CrossRef] [PubMed]
- Fischer, G.M.; Jalali, A.; Kircher, D.A.; Lee, W.-C.; McQuade, J.L.; Haydu, L.E.; Joon, A.Y.; Reuben, A.; de Macedo, M.P.; Carapeto, F.C.L.; et al. Molecular Profiling Reveals Unique Immune and Metabolic Features of Melanoma Brain Metastases. Cancer Discov. 2019, 9, 628–645. [Google Scholar] [CrossRef]
- Karimi, E.; Yu, M.W.; Maritan, S.M.; Perus, L.J.M.; Rezanejad, M.; Sorin, M.; Dankner, M.; Fallah, P.; Doré, S.; Zuo, D.; et al. Single-cell spatial immune landscapes of primary and metastatic brain tumours. Nature 2023, 614, 555–563. [Google Scholar] [CrossRef] [PubMed]
- Dong, W.; Sheng, J.; Cui, J.Z.; Zhao, H.; Wong, S.T. Systems immunology insights into brain metastasis. Trends Immunol. 2024, 45, 903–916. [Google Scholar] [CrossRef]
- Croucher, P.I.; McDonald, M.M.; Martin, T.J. Bone metastasis: the importance of the neighbourhood. Nat. Rev. Cancer 2016, 16, 373–386. [Google Scholar] [CrossRef] [PubMed]
- Best, S.A.; Gubser, P.M.; Sethumadhavan, S.; Kersbergen, A.; Abril, Y.L.N.; Goldford, J.; Sellers, K.; Abeysekera, W.; Garnham, A.L.; McDonald, J.A.; et al. Glutaminase inhibition impairs CD8 T cell activation in STK11-/Lkb1-deficient lung cancer. Cell Metab. 2022, 34, 874–887.e6. [Google Scholar] [CrossRef]
- Zavitsanou, A.-M.; Pillai, R.; Hao, Y.; Wu, W.L.; Bartnicki, E.; Karakousi, T.; Rajalingam, S.; Herrera, A.; Karatza, A.; Rashidfarrokhi, A.; et al. KEAP1 mutation in lung adenocarcinoma promotes immune evasion and immunotherapy resistance. Cell Rep. 2023, 42, 113295–113295. [Google Scholar] [CrossRef]
- Gui, P.; Bivona, T.G. Evolution of metastasis: new tools and insights. Trends Cancer 2022, 8, 98–109. [Google Scholar] [CrossRef]
- Shi, Q.; Chen, Y.; Li, Y.; Qin, S.; Yang, Y.; Gao, Y.; Zhu, L.; Wang, D.; Zhang, Z. Cross-tissue multicellular coordination and its rewiring in cancer. Nature 2025, 643, 529–538. [Google Scholar] [CrossRef]
- Feunteun, J.; Ostyn, P.; Delaloge, S. Tumor cell malignancy: A complex trait built through reciprocal interactions between tumors and tissue-body system. iScience 2022, 25, 104217. [Google Scholar] [CrossRef]
- Zahir, N.; Sun, R.; Gallahan, D.; Gatenby, R.A.; Curtis, C. Characterizing the ecological and evolutionary dynamics of cancer. Nat. Genet. 2020, 52, 759–767. [Google Scholar] [CrossRef] [PubMed]
- Maley, C.C.; Aktipis, A.; Graham, T.A.; Sottoriva, A.; Boddy, A.M.; Janiszewska, M.; Silva, A.S.; Gerlinger, M.; Yuan, Y.; Pienta, K.J.; et al. Classifying the evolutionary and ecological features of neoplasms. Nat. Rev. Cancer 2017, 17, 605–619. [Google Scholar] [CrossRef]
- Noble, R.; Burri, D.; Le Sueur, C.; Lemant, J.; Viossat, Y.; Kather, J.N.; Beerenwinkel, N. Spatial structure governs the mode of tumour evolution. Nat. Ecol. Evol. 2021, 6, 207–217. [Google Scholar] [CrossRef]
- Khaliq, A.M.; Rajamohan, M.; Saeed, O.; Mansouri, K.; Adil, A.; Zhang, C.; Turk, A.; Carstens, J.L.; House, M.; Hayat, S.; et al. Spatial transcriptomic analysis of primary and metastatic pancreatic cancers highlights tumor microenvironmental heterogeneity. Nat. Genet. 2024, 56, 2455–2465. [Google Scholar] [CrossRef]
- Aguadé-Gorgorió, G.; Anderson, A.R.; Solé, R. Modeling tumors as complex ecosystems. iScience 2024, 27, 110699. [Google Scholar] [CrossRef]
- Ma, H.; Srivastava, S.; Ho, S.W.T.; Xu, C.; Lian, B.S.X.; Ong, X.; Tay, S.T.; Sheng, T.; Lum, H.Y.J.; Ghani, S.A.B.A.; et al. Spatially Resolved Tumor Ecosystems and Cell States in Gastric Adenocarcinoma Progression and Evolution. Cancer Discov. 2025, 15, 767–792. [Google Scholar] [CrossRef] [PubMed]
- Béguelin, W.; Teater, M.; Meydan, C.; Hoehn, K.B.; Phillip, J.M.; Soshnev, A.A.; Venturutti, L.; Rivas, M.A.; Calvo-Fernández, M.T.; Gutierrez, J.; et al. Mutant EZH2 Induces a Pre-malignant Lymphoma Niche by Reprogramming the Immune Response. Cancer Cell 2020, 37, 655–673.e11. [Google Scholar] [CrossRef]
- Mourcin F, Verdière L, Roulois D, Amin R, Lamaison C, Sibut V, et al. Follicular lymphoma triggers phenotypic and functional remodeling of the human lymphoid stromal cell landscape. Immunity. 2021;54: 1788–1806.e7.
- Laurent, C.; Dietrich, S.; Tarte, K. Cell cross talk within the lymphoma tumor microenvironment: follicular lymphoma as a paradigm. Blood 2024, 143, 1080–1090. [Google Scholar] [CrossRef]
- Laplane, L.; Maley, C.C. The evolutionary theory of cancer: challenges and potential solutions. Nat. Rev. Cancer 2024, 24, 718–733. [Google Scholar] [CrossRef]
- Huang, S.; Soto, A.M.; Sonnenschein, C. The end of the genetic paradigm of cancer. PLOS Biol. 2025, 23, e3003052. [Google Scholar] [CrossRef]
- Lawson, A.R.J.; Abascal, F.; Coorens, T.H.H.; Hooks, Y.; O’nEill, L.; Latimer, C.; Raine, K.; Sanders, M.A.; Warren, A.Y.; Mahbubani, K.T.A.; et al. Extensive heterogeneity in somatic mutation and selection in the human bladder. Science 2020, 370, 75–82. [Google Scholar] [CrossRef]
- Wijewardhane, N.; Dressler, L.; Ciccarelli, F.D. Normal Somatic Mutations in Cancer Transformation. Cancer Cell 2021, 39, 125–129. [Google Scholar] [CrossRef]
- Bailey, M.H.; Tokheim, C.; Porta-Pardo, E.; Sengupta, S.; Bertrand, D.; Weerasinghe, A.; Colaprico, A.; Wendl, M.C.; Kim, J.; Reardon, B.; et al. Comprehensive Characterization of Cancer Driver Genes and Mutations. Cell 2018, 173, 371–385.E18. [Google Scholar] [CrossRef]
- Li, Y.; Porta-Pardo, E.; Tokheim, C.; Bailey, M.H.; Yaron, T.M.; Stathias, V.; Geffen, Y.; Imbach, K.J.; Cao, S.; Anand, S.; et al. Pan-cancer proteogenomics connects oncogenic drivers to functional states. Cell 2023, 186, 3921–3944.e25. [Google Scholar] [CrossRef] [PubMed]
- Porta-Pardo, E.; Valencia, A.; Godzik, A. Understanding oncogenicity of cancer driver genes and mutations in the cancer genomics era. FEBS Lett. 2020, 594, 4233–4246. [Google Scholar] [CrossRef] [PubMed]
- Barkley, D.; Rao, A.; Pour, M.; França, G.S.; Yanai, I. Cancer cell states and emergent properties of the dynamic tumor system. Genome Res. 2021, 31, 1719–1727. [Google Scholar] [CrossRef]
- Massagué, J.; Ganesh, K. Metastasis-Initiating Cells and Ecosystems. Cancer Discov. 2021, 11, 971–994. [Google Scholar] [CrossRef] [PubMed]
- Guilliams, M.; Bonnardel, J.; Haest, B.; Vanderborght, B.; Wagner, C.; Remmerie, A.; Bujko, A.; Martens, L.; Thoné, T.; Browaeys, R.; et al. Spatial proteogenomics reveals distinct and evolutionarily conserved hepatic macrophage niches. Cell 2022, 185, 379–396.e38. [Google Scholar] [CrossRef]
- Bhang, H.-E.C.; Ruddy, D.A.; Krishnamurthy Radhakrishna, V.; Caushi, J.X.; Zhao, R.; Hims, M.M.; Singh, A.P.; Kao, I.; Rakiec, D.; Shaw, P.; et al. Studying clonal dynamics in response to cancer therapy using high-complexity barcoding. Nat. Med. 2015, 21, 440–448. [Google Scholar] [CrossRef]
- Gerlinger, M.; Horswell, S.; Larkin, J.; Rowan, A.J.; Salm, M.P.; Varela, I.; Fisher, R.; McGranahan, N.; Matthews, N.; Santos, C.R.; et al. Genomic architecture and evolution of clear cell renal cell carcinomas defined by multiregion sequencing. Nat. Genet. 2014, 46, 225–233. [Google Scholar] [CrossRef]
- de Bruin, E.C.; McGranahan, N.; Mitter, R.; Salm, M.; Wedge, D.C.; Yates, L.; Jamal-Hanjani, M.; Shafi, S.; Murugaesu, N.; Rowan, A.J.; et al. Spatial and temporal diversity in genomic instability processes defines lung cancer evolution. Science 2014, 346, 251–256. [Google Scholar] [CrossRef]
- Yates, L.R.; Gerstung, M.; Knappskog, S.; Desmedt, C.; Gundem, G.; Van Loo, P.; Aas, T.; Alexandrov, L.B.; Larsimont, D.; Davies, H.; et al. Subclonal diversification of primary breast cancer revealed by multiregion sequencing. Nat. Med. 2015, 21, 751–759. [Google Scholar] [CrossRef]
- Carbone A, Roulland S, Gloghini A, Younes A, von Keudell G, López-Guillermo A, et al. Follicular lymphoma. Nat Rev Dis Primers. 2019;5: 83.
- Dyrskjøt L, Hansel DE, Efstathiou JA, Knowles MA, Galsky MD, Teoh J, et al. Bladder cancer. Nat Rev Dis Primers. 2023;9: 58.
- Halbrook CJ, Lyssiotis CA, Pasca di Magliano M, Maitra A. Pancreatic cancer: Advances and challenges. Cell. 2023;186: 1729–1754.
- Risom, T.; Glass, D.R.; Averbukh, I.; Liu, C.C.; Baranski, A.; Kagel, A.; McCaffrey, E.F.; Greenwald, N.F.; Rivero-Gutiérrez, B.; Strand, S.H.; et al. Transition to invasive breast cancer is associated with progressive changes in the structure and composition of tumor stroma. Cell 2022, 185, 299–310.e18. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Li, B.; Luo, M.; Huang, J.; Zhang, K.; Zheng, S.; Zhang, S.; Zhou, J. Progression from ductal carcinoma in situ to invasive breast cancer: molecular features and clinical significance. Signal Transduct. Target. Ther. 2024, 9, 1–28. [Google Scholar] [CrossRef]
- Chen, J.; Larsson, L.; Swarbrick, A.; Lundeberg, J. Spatial landscapes of cancers: insights and opportunities. Nat. Rev. Clin. Oncol. 2024, 21, 660–674. [Google Scholar] [CrossRef] [PubMed]
- Gong, D.; Arbesfeld-Qiu, J.M.; Perrault, E.; Bae, J.W.; Hwang, W.L. Spatial oncology: Translating contextual biology to the clinic. Cancer Cell 2024, 42, 1653–1675. [Google Scholar] [CrossRef]
- Johnson, J.A.; Bergman, D.R.; Rocha, H.L.; Zhou, D.L.; Cramer, E.; Mclean, I.C.; Dance, Y.W.; Booth, M.; Nicholas, Z.; Lopez-Vidal, T.; et al. Human interpretable grammar encodes multicellular systems biology models to democratize virtual cell laboratories. Cell 2025, 188, 4711–4733.e37. [Google Scholar] [CrossRef]
- Metzcar, J.; Wang, Y.; Heiland, R.; Macklin, P. A Review of Cell-Based Computational Modeling in Cancer Biology. JCO Clin. Cancer Informatics 2019, 3, 1–13. [Google Scholar] [CrossRef]
- Rocha, H.L.; Aguilar, B.; Getz, M.; Shmulevich, I.; Macklin, P. A multiscale model of immune surveillance in micrometastases gives insights on cancer patient digital twins. npj Syst. Biol. Appl. 2024, 10, 1–12. [Google Scholar] [CrossRef]
- Rocha, H.L.; Godet, I.; Kurtoglu, F.; Metzcar, J.; Konstantinopoulos, K.; Bhoyar, S.; Gilkes, D.M.; Macklin, P. A persistent invasive phenotype in post-hypoxic tumor cells is revealed by fate mapping and computational modeling. iScience 2021, 24, 102935. [Google Scholar] [CrossRef] [PubMed]
- Hernandez-Boussard, T.; Macklin, P.; Greenspan, E.J.; Gryshuk, A.L.; Stahlberg, E.; Syeda-Mahmood, T.; Shmulevich, I. Digital twins for predictive oncology will be a paradigm shift for precision cancer care. Nat. Med. 2021, 27, 2065–2066. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



