Submitted:
05 September 2025
Posted:
08 September 2025
You are already at the latest version
Abstract
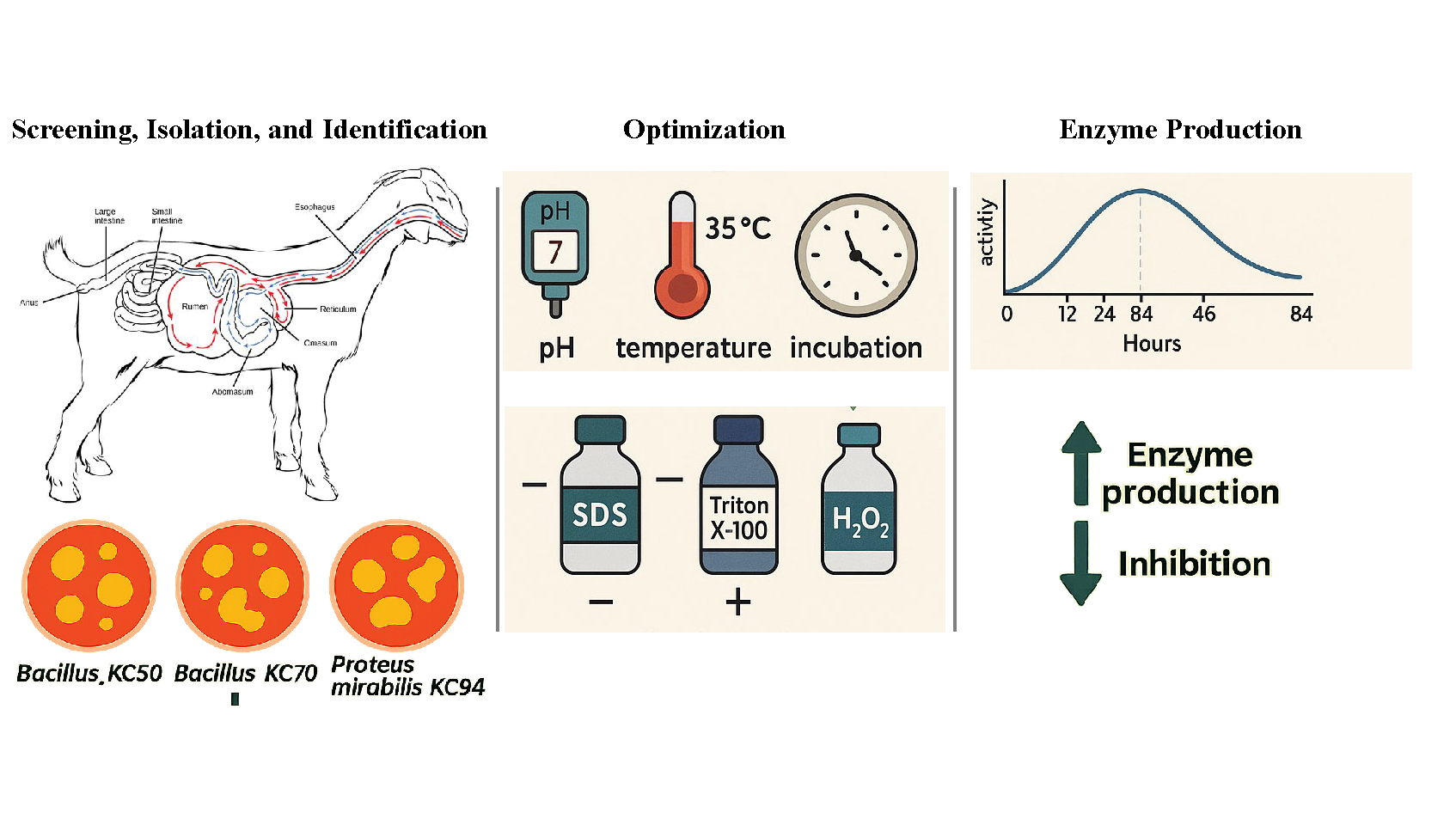
This study reports the isolation and optimization of cellulase-producing bacteria from the gastrointestinal tract of South African goats for the pretreatment of lignocellulosic biomass in bioenergy applications. Among the isolates, three strains, Bacillus KC50, Bacillus KC70, and Proteus mirabilis KC94, were identified by 16S rDNA sequencing. To our knowledge, this is the first report of cellulolytic optimization in P. mirabilis derived from a goat rumen. Enzyme production was optimized under varying pH, temperature, and incubation conditions. P. mirabilis KC94 exhibited robust enzyme activity at pH 7 and 35 °C, with stability across a broader range than the Bacillus strains. Peak activity occurred at 84 hours of incubation, reflecting strain-specific metabolic adaptation. The presence of organic solvents and surfactants inhibited enzyme activity, whereas mild oxidative stress induced by H₂O₂ stimulated cellulase production. Amplification of GH39, GH45, and GH48 genes revealed KC94’s strong genetic potential for efficient lignocellulose degradation. These findings highlight the biotechnological potential of rumen-derived cellulolytic bacteria, particularly P. mirabilis KC94, for advancing sustainable bioenergy systems.
Keywords:
1. Introduction
2. Materials and Methods
Ethical Statement
2.1. Bacterial Screening, Isolation, and Qualitative Assay
2.2. Identification of the Cellulase-Producing Bacteria by 16S rDNA Sequencing
2.3. Effects of pH, Temperature, and Incubation Period
2.4. Effects of Organic Solvents and Surfactants on Cellulase Production
2.5. PCR amplification of Possible Cellulase and Hemi-Cellulase Gene Fragments on KC94
2.6. Statistical Analysis
3. Results
3.1. Qualitative Assay of Cellulase-Producing Bacteria
3.2. Bacterial Identification
3.3. Optimization Studies
3.3.1. The Effect of pH
3.3.2. The Effects of Temperature
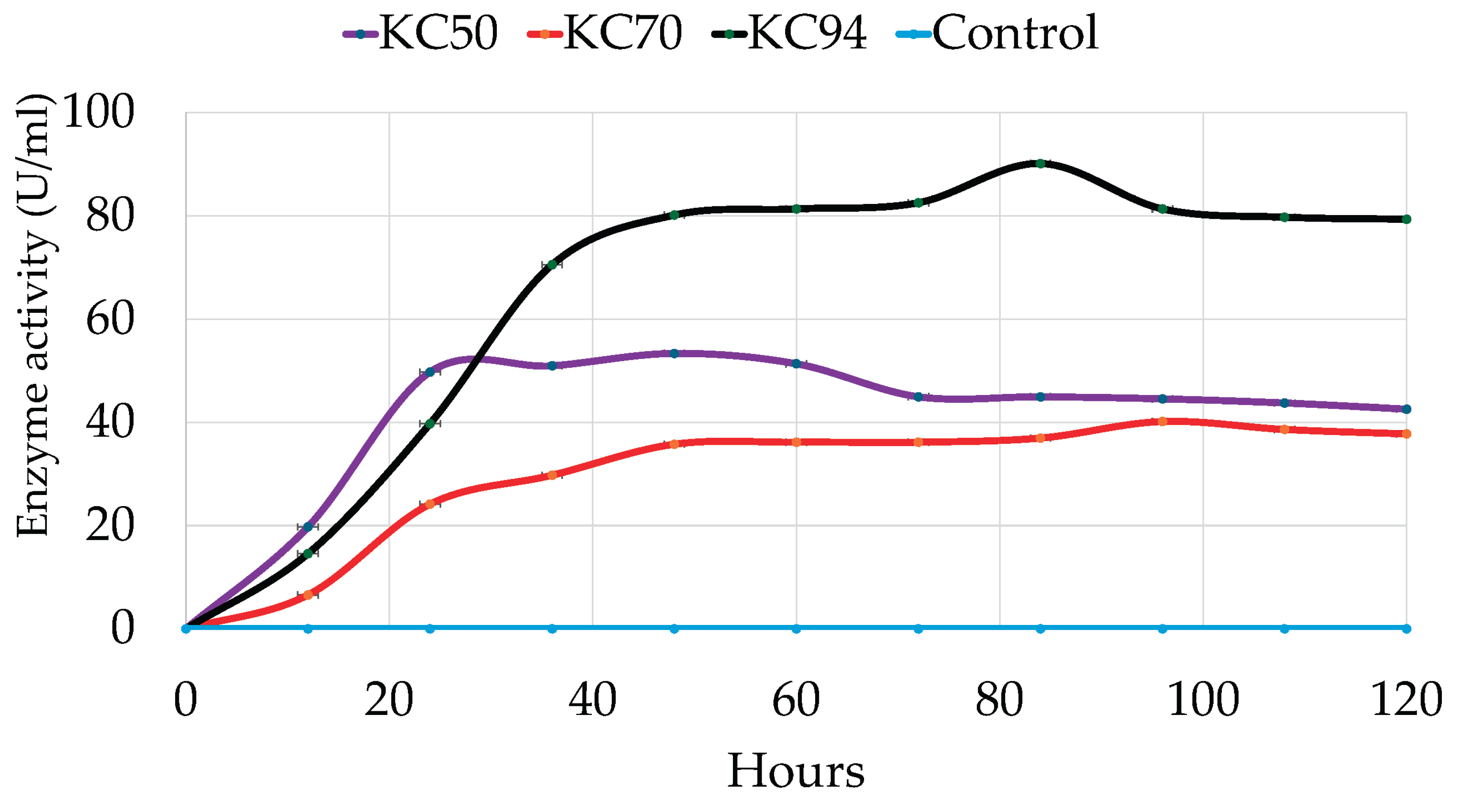
3.3.3. The Effects of the Incubation Period
3.4. The Effects of Organic Solvents, Surfactants, and Oxidizing agent on Cellulase Production
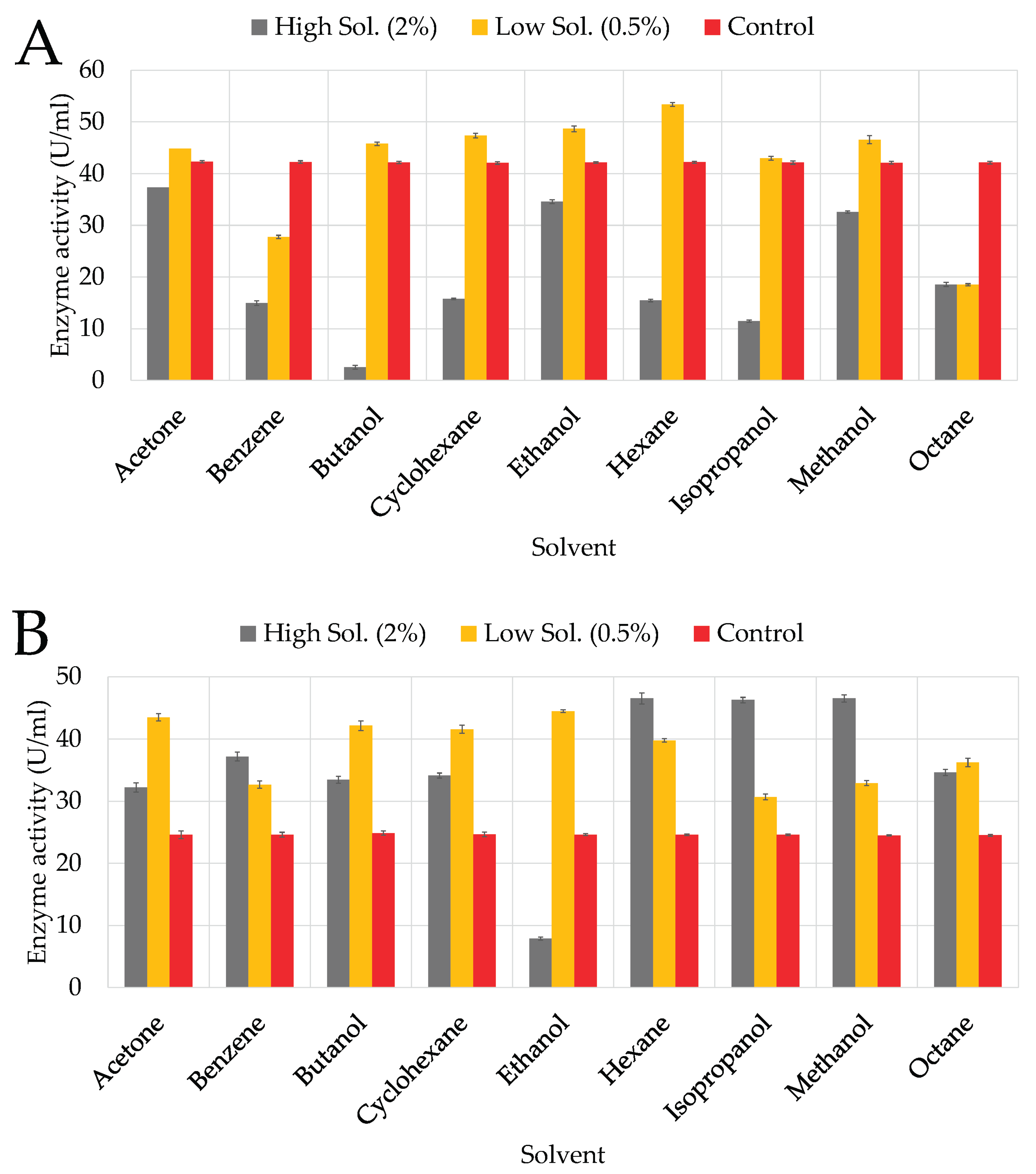
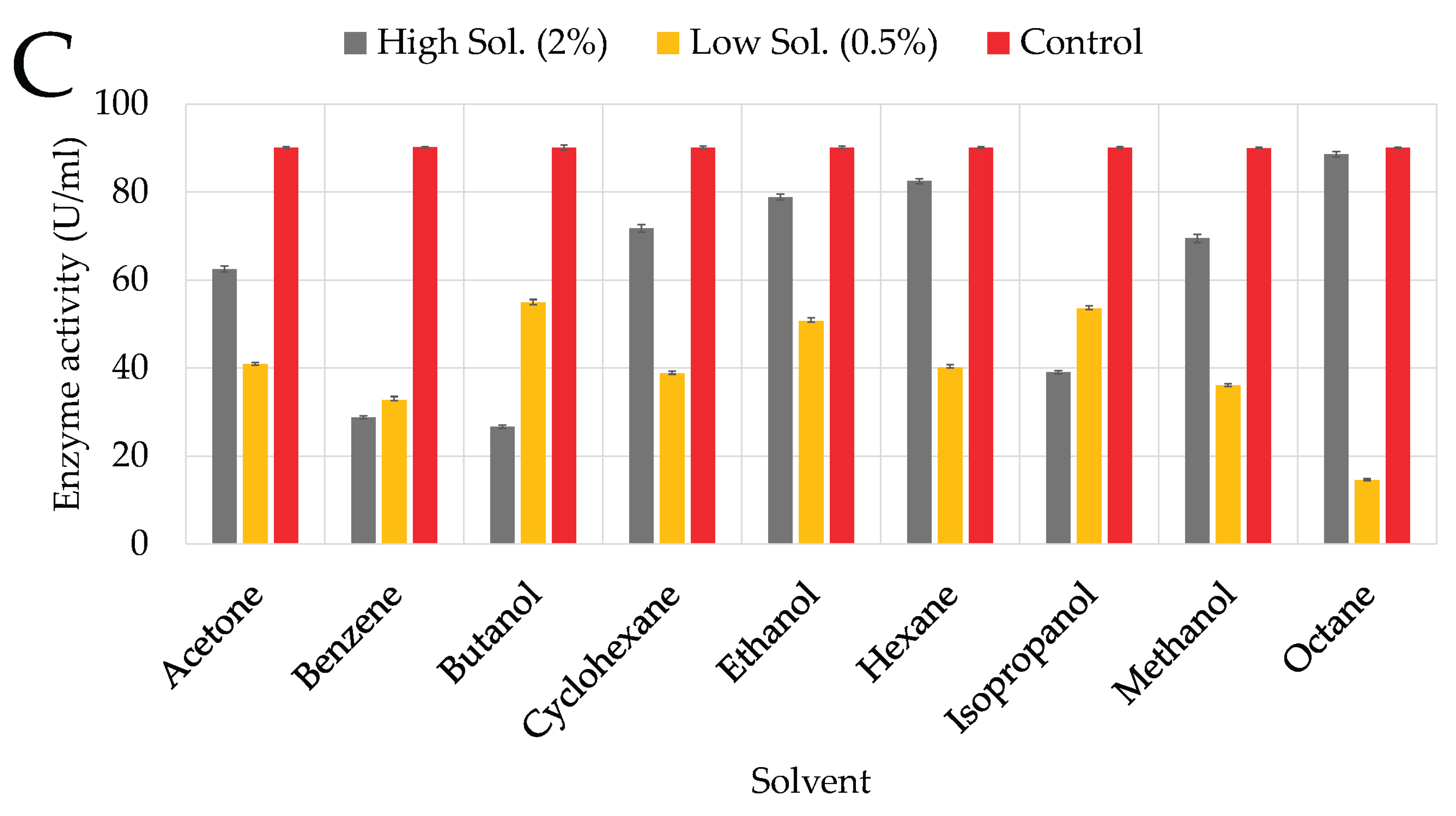
3.4.1. Solvents
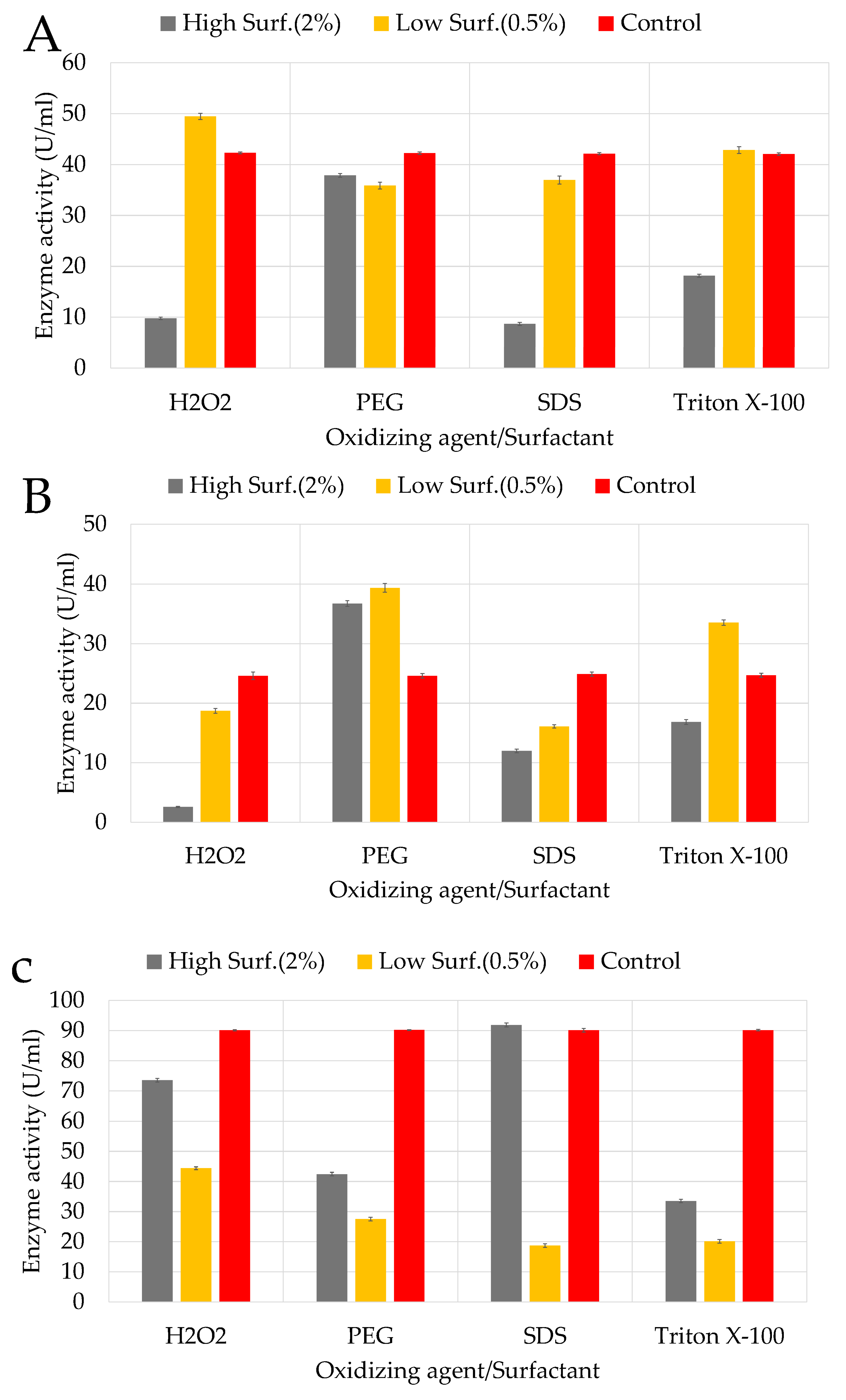
3.4.2. Surfactants and Oxidising Agent
3.5. Diversity of Possible Cellulase and Hemi-Cellulase Gene Fragments in KC94
4. Discussion
4.1. Qualitative Assay of Cellulase-Producing Bacteria
4.2. The effects of pH, temperature, and incubation period
4.2.1. pH
4.2.2. Temperature
4.2.3. Incubation Period
4.3. The Effects of Organic Solvents, Surfactants, and Oxidizing Agent on Cellulase Production
4.3.1. Organic Solvents
4.3.2. Surfactants and Oxidizing Agent
4.4. Diversity of Possible Cellulase and Hemicellulase Gene Fragments in KC94
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ajala, E., et al., Sugarcane bagasse: a biomass sufficiently applied for improving global energy, environment and economic sustainability. Bioresources and Bioprocessing, 2021. 8: p. 1-25. [CrossRef]
- Pan, S., et al., Technoeconomic and environmental perspectives of biofuel production from sugarcane bagasse: Current status, challenges and future outlook. Industrial Crops and Products, 2022. 188: p. 115684. [CrossRef]
- Kumar, A., V. Kumar, and B. Singh, Cellulosic and hemicellulosic fractions of sugarcane bagasse: Potential, challenges and future perspective. International Journal of Biological Macromolecules, 2021. 169: p. 564-582. [CrossRef]
- Negrão, D.R., et al., Inorganics in sugarcane bagasse and straw and their impacts for bioenergy and biorefining: A review. Renewable and Sustainable Energy Reviews, 2021. 148: p. 111268.
- Konde, K.S., et al., Sugarcane bagasse based biorefineries in India: potential and challenges. Sustainable Energy & Fuels, 2021. 5(1): p. 52-78. [CrossRef]
- Sarker, T.C., S.M.G.G. Azam, and G. Bonanomi, Recent advances in sugarcane industry solid by-products valorization. Waste and Biomass Valorization, 2017. 8: p. 241-266. [CrossRef]
- Liang, B., et al., Stepwise processing strategy for efficient and comprehensive utilization of sugarcane bagasse via enzymatic hydrolysis and catalytic conversion to produce 5-hydroxymethylfurfural. Industrial Crops and Products, 2024. 214: p. 118592. [CrossRef]
- Kim, Y.-K., et al., Isolation of cellulolytic Bacillus subtilis strains from agricultural environments. International Scholarly Research Notices, 2012. 2012(1): p. 650563. [CrossRef]
- Yin, L.-J., H.-H. Lin, and Z.-R. Xiao, Purification and characterization of a cellulase from Bacillus subtilis YJ1. Journal of Marine Science and Technology, 2010. 18(3): p. 19. [CrossRef]
- Mushtaq, Q., et al., Amylase and cellulase production from newly isolated Bacillus subtilis using acid treated potato peel waste. Microorganisms, 2024. 12(6): p. 1106. [CrossRef]
- Bhatt, B., et al., Production of a novel cellulase by Bacillus amyloliquefaciens OKB3 isolated from soil: Purification and characterization. International Journal of Biological Macromolecules, 2024. 282: p. 137454. [CrossRef]
- Liao, Y., et al., Cellulolytic Bacillus cereus produces a variety of short-chain fatty acids and has potential as a probiotic. Microbiology Spectrum, 2024. 12(4): p. e03267-23. [CrossRef]
- Pană, A.-M., et al., Study on the Biodegradation Process of D-Mannose Glycopolymers in Liquid Media and Soil. Polymers, 2023. 15(15): p. 3194. [CrossRef]
- Araki, Y., et al., Proteus mirabilis sp. intestinal microflora grow in a dextran sulfate sodium-rich environment. Int J Mol Med, 2010. 25(2): p. 203-8. [CrossRef]
- Mahapatra, S., et al., Screening, production, optimization and characterization of β-glucosidase using microbes from shellfish waste. 3 Biotech, 2016. 6: p. 1-10. [CrossRef]
- Patel, D.D., et al., Microbial and Carbohydrate Active Enzyme profile of buffalo rumen metagenome and their alteration in response to variation in the diet. Gene 545, 2014: p. 88-94. [CrossRef]
- Patel, V., et al., Characterization of the rumen microbiome of Indian Kankrej cattle (Bos indicus) adapted to different forage diet. Applied microbiology and biotechnology, 2014. 98(23): p. 9749-9761. [CrossRef]
- Rabapane, K.J. and T.S. Matambo, Profiling the dynamic adaptations of CAZyme-Producing microorganisms in the gastrointestinal tract of South African goats. Heliyon, 2024. 10(17). [CrossRef]
- Sarsan, S. and R. Merugu, Role of bioprocess parameters to improve cellulase production: Part II, in New and future developments in microbial biotechnology and bioengineering. 2019, Elsevier. p. 77-97. [CrossRef]
- Kumari, S., et al., Screening and molecular characterization of cellulase producing actinobacteria from Litchi Orchard. Current Chemical Biology, 2019. 13(1): p. 90-101. [CrossRef]
- Gaur, R. and S. Tiwari, Isolation, production, purification and characterization of an organic-solvent-thermostable alkalophilic cellulase from Bacillus vallismortis RG-07. BMC biotechnology, 2015. 15(1): p. 1-12. [CrossRef]
- Hussain, A.A., et al., Optimization and molecular identification of novel cellulose degrading bacteria isolated from Egyptian environment. Journal of Genetic Engineering and Biotechnology, 2017. 15(1): p. 77-85. [CrossRef]
- Irfan, M., et al., Carboxymethyl cellulase production optimization from newly isolated thermophilic Bacillus subtilis K-18 for saccharification using response surface methodology. AMB Express, 2017. 7(1): p. 29. [CrossRef]
- Hsiao, N.-W., et al., Purification and characterization of an aspartic protease from the Rhizopus oryzae protease extract, Peptidase R. Electronic Journal of Biotechnology, 2014. 17(2): p. 89-94. [CrossRef]
- Usman, A., S. Mohammed, and J. Mamo, Production, Optimization, and Characterization of an Acid Protease from a Filamentous Fungus by Solid-State Fermentation. International journal of microbiology, 2021. 2021(1): p. 6685963. [CrossRef]
- Weisburg, W.G., et al., 16S ribosomal DNA amplification for phylogenetic study. Journal of bacteriology, 1991. 173(2): p. 697-703. [CrossRef]
- Wilson, K.H., R.B. Blitchington, and R.C. Greene, Amplification of bacterial 16S ribosomal DNA with polymerase chain reaction. Journal of clinical microbiology, 1990. 28(9): p. 1942-1946. [CrossRef]
- Irfan, M., et al., Production and characterization of organic solvent-tolerant cellulase from Bacillus amyloliquefaciens AK9 isolated from hot spring. Applied Biochemistry and Biotechnology, 2017. 182: p. 1390-1402. [CrossRef]
- Hall, T.A. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. in Nucleic acids symposium series. 1999. [London]: Information Retrieval Ltd., c1979-c2000.
- Waheeb, M.S., et al., Optimized production and characterization of a thermostable cellulase from Streptomyces thermodiastaticus strain. AMB Express, 2024. 14(1): p. 129. [CrossRef]
- Potprommanee, L., et al., Characterization of a thermophilic cellulase from Geobacillus sp. HTA426, an efficient cellulase-producer on alkali pretreated of lignocellulosic biomass. PLoS one, 2017. 12(4): p. e0175004. [CrossRef]
- Nkuna, R. and T.S. Matambo, Determining the metabolic processes of metal-tolerant fungi isolated from mine tailings for bioleaching. Minerals, 2024. 14(3): p. 235. [CrossRef]
- Bernfeld, P., [17] Amylases, α and β, in Methods in Enzymology. 1955, Academic Press. p. 149-158. [CrossRef]
- Alamri, S.A., Y.S. Mostafa, and S.A. Alrumman, Isolation, Screening and Optimization of Geobacillus stearothermophilus Cellulase Production using Date Palm Cellulosic Wastes. Journal of Pure & Applied Microbiology, 2016. 10(1).
- Demissie, M.S., N.H. Legesse, and A.A. Tesema, Isolation and characterization of cellulase producing bacteria from forest, cow dung, Dashen brewery and agro-industrial waste. PLOS ONE, 2024. 19(4): p. e0301607. [CrossRef]
- Gebbie, L., et al., A snapshot of microbial diversity and function in an undisturbed sugarcane bagasse pile. BMC biotechnology, 2020. 20: p. 1-16. [CrossRef]
- Shareef, I., et al., Isolation and identification of cellulose degrading microbes. 2015. http://13.232.72.61:8080/jspui/handle/123456789/2245.
- Singh, S., et al., Developing efficient thermophilic cellulose degrading consortium for glucose production from different agro-residues. Frontiers in Energy Research, 2019. 7: p. 61. [CrossRef]
- Barman, D. and M.S. Dkhar, Characterization and purification of esterase from Cellulomonas fimi DB19 isolated from Zanthoxylum armatum with its possible role in diesel biodegradation. Archives of Microbiology, 2022. 204(9): p. 580. [CrossRef]
- Sheng, P., et al., High genetic diversity of microbial cellulase and hemicellulase genes in the hindgut of Holotrichia parallela larvae. International Journal of Molecular Sciences, 2015. 16(7): p. 16545-16559. [CrossRef]
- Islam, M., et al., Optimization of fermentation condition for cellulase enzyme production from Bacillus sp. Malaysian Journal of Halal Research, 2019. 2(2): p. 19-24. [CrossRef]
- El-Khamisi, E.F., et al., Optimization, gene cloning, expression, and molecular docking insights for enhanced cellulase enzyme production by Bacillus amyloliquefaciens strain elh1. Microbial Cell Factories, 2024. 23(1): p. 191. [CrossRef]
- Malik, W.A., H.M. Khan, and S. Javed, Bioprocess optimization for enhanced production of bacterial cellulase and hydrolysis of sugarcane bagasse. BioEnergy Research, 2021: p. 1-14. [CrossRef]
- Mobley, H.L.T., Proteus mirabilis Overview, in Proteus mirabilis: Methods and Protocols, M.M. Pearson, Editor. 2019, Springer New York: New York, NY. p. 1-4. [CrossRef]
- Patakova, P., et al., Comparative analysis of high butanol tolerance and production in clostridia. Biotechnology advances, 2018. 36(3): p. 721-738. [CrossRef]
- Russmayer, H., H. Marx, and M. Sauer, Microbial 2-butanol production with Lactobacillus diolivorans. Biotechnology for biofuels, 2019. 12: p. 1-11. [CrossRef]
- Zaks, A. and A.M. Klibanov, Enzymatic catalysis in nonaqueous solvents. Journal of Biological Chemistry, 1988. 263(7): p. 3194-3201. [CrossRef]
- Skovgaard, P.A. and H. Jørgensen, Influence of high temperature and ethanol on thermostable lignocellulolytic enzymes. Journal of Industrial Microbiology and Biotechnology, 2013. 40(5): p. 447-456. [CrossRef]
- Saini, A., N.K. Aggarwal, and A. Yadav, Isolation and screening of cellulose hydrolyzing bacteria from different ecological niches. Bioengineering and Bioscience, 2017. 5(1): p. 7-13. [CrossRef]
- Asha, B.M. and N. Sakthivel, Production, purification and characterization of a new cellulase from Bacillus subtilis that exhibit halophilic, alkalophilic and solvent-tolerant properties. Annals of Microbiology, 2014. 64(4): p. 1839-1848. [CrossRef]
- Rabapane, K.J., G.N. Ijoma, and T.S. Matambo, Insufficiency in functional genomics studies, data, and applications: A case study of bio-prospecting research in ruminant microbiome. Frontiers in Genetics, 2022. 13. [CrossRef]
- Zhang, Q., et al., The structural basis of the pH-homeostasis mediated by the Cl−/HCO3− exchanger, AE2. Nature Communications, 2023. 14(1): p. 1812. [CrossRef]
- Wasfi, R., et al., Proteus mirabilis biofilm: development and therapeutic strategies. Frontiers in cellular and infection microbiology, 2020. 10: p. 414. [CrossRef]
- Chakkour, M., et al., Overview of Proteus mirabilis pathogenicity and virulence. Insights into the role of metals. Frontiers in Microbiology, 2024. 15: p. 1383618. [CrossRef]
- Yang, W. and K. Beauchemin, Physically effective fiber: method of determination and effects on chewing, ruminal acidosis, and digestion by dairy cows. Journal of dairy science, 2006. 89(7): p. 2618-2633. [CrossRef]
- Hossain, M., Sub-acute ruminal acidosis in dairy cows: Its causes, consequences and preventive measures. Online J. Anim. Feed Res, 2020. 10(1): p. 302-312. [CrossRef]
- Lian, H., et al., In Vitro Gene Expression Responses of Bovine Rumen Epithelial Cells to Different pH Stresses. Animals, 2022. 12(19): p. 2621. [CrossRef]
- Russell, J.B. and D.B. Wilson, Why Are Ruminal Cellulolytic Bacteria Unable to Digest Cellulose at Low pH? Journal of Dairy Science, 1996. 79(8): p. 1503-1509. [CrossRef]
- Hyder, I., et al., Alteration in rumen functions and diet digestibility during heat stress in sheep, in Sheep production adapting to climate change. 2017, Springer. p. 235-265. [CrossRef]
- Castro-Costa, A., et al., Using wireless rumen sensors for evaluating the effects of diet and ambient temperature in nonlactating dairy goats. J Dairy Sci, 2015. 98(7): p. 4646-58. [CrossRef]
- Xu, Z., et al., Recent advances in the improvement of enzyme thermostability by structure modification. Critical reviews in biotechnology, 2020. 40(1): p. 83-98. [CrossRef]
- Rahban, M., et al., Thermal stability enhancement: Fundamental concepts of protein engineering strategies to manipulate the flexible structure. International journal of biological macromolecules, 2022. 214: p. 642-654. [CrossRef]
- Wu, H., et al., Overview of strategies for developing high thermostability industrial enzymes: Discovery, mechanism, modification and challenges. Critical Reviews in Food Science and Nutrition, 2023. 63(14): p. 2057-2073. [CrossRef]
- Günel, G., et al., Enhancing biomethane production from cattle manure by integrating rumen bacteria: a microbial analyses with next-generation sequencing and quantitative PCR. Biomass Conversion and Biorefinery, 2025. [CrossRef]
- Oztat, K., A.A. Yavuz, and C.F. Işçen, Optimization studies on laccase activity of Proteus mirabilis isolated from treatment sludge of textile industry factories. Brazilian Journal of Microbiology, 2024. 55(2): p. 1231-1241. [CrossRef]
- Koike, S. and Y. Kobayashi, Fibrolytic Rumen Bacteria: Their Ecology and Functions. Asian-Australasian Journal of Animal Sciences, 2009. 22. [CrossRef]
- Cheng, B., et al., Protein denaturation at a single-molecule level: the effect of nonpolar environments and its implications on the unfolding mechanism by proteases. Nanoscale, 2015. 7(7): p. 2970-2977. [CrossRef]
- Thomas, L., et al., Production, Optimization, and Characterization of Organic Solvent Tolerant Cellulases from a Lignocellulosic Waste-Degrading Actinobacterium, Promicromonospora sp. VP111. Applied Biochemistry and Biotechnology, 2016. 179(5): p. 863-879. [CrossRef]
- Liu, K., et al., Ruminal microbiota-host interaction and its effect on nutrient metabolism. Anim Nutr, 2021. 7(1): p. 49-55. [CrossRef]
- Silva, É.B.R.d., et al., A Review of the Rumen Microbiota and the Different Molecular Techniques Used to Identify Microorganisms Found in the Rumen Fluid of Ruminants. Animals, 2024. 14(10): p. 1448. [CrossRef]
- Celi, P., Oxidative Stress in Ruminants, in Studies on Veterinary Medicine, L. Mandelker and P. Vajdovich, Editors. 2011, Humana Press: Totowa, NJ. p. 191-231. [CrossRef]
- Celi, P., et al., Oxidative stress in ruminants: enhancing productivity through antioxidant supplementation. Feedipedia Broadening Horizons, 2014. 13.
- Hua, M., et al., Genetic diversity detection and gene discovery of novel glycoside hydrolase family 48 from soil environmental genomic DNA. Annals of Microbiology, 2018. 68(4): p. 163-174. [CrossRef]
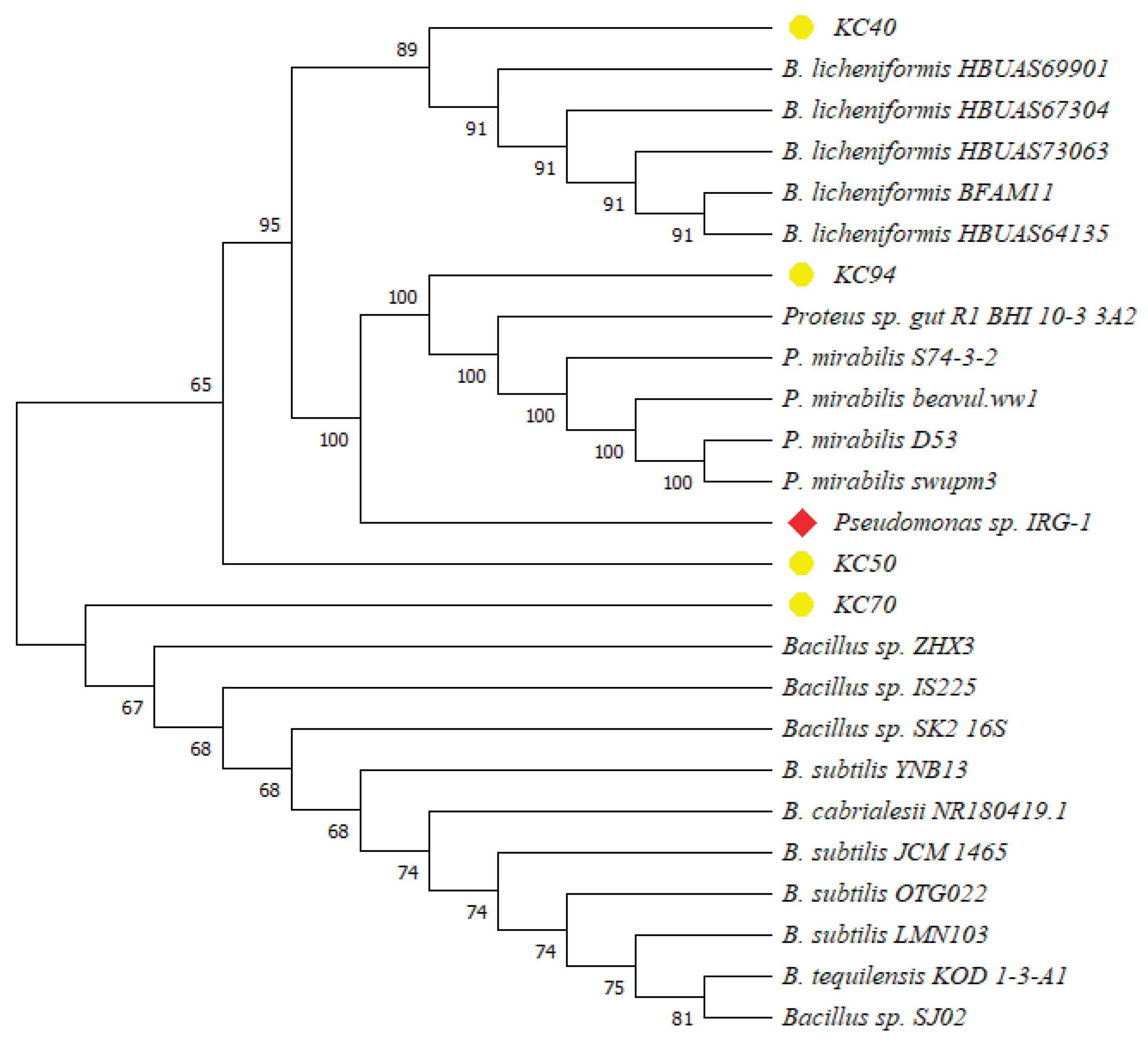
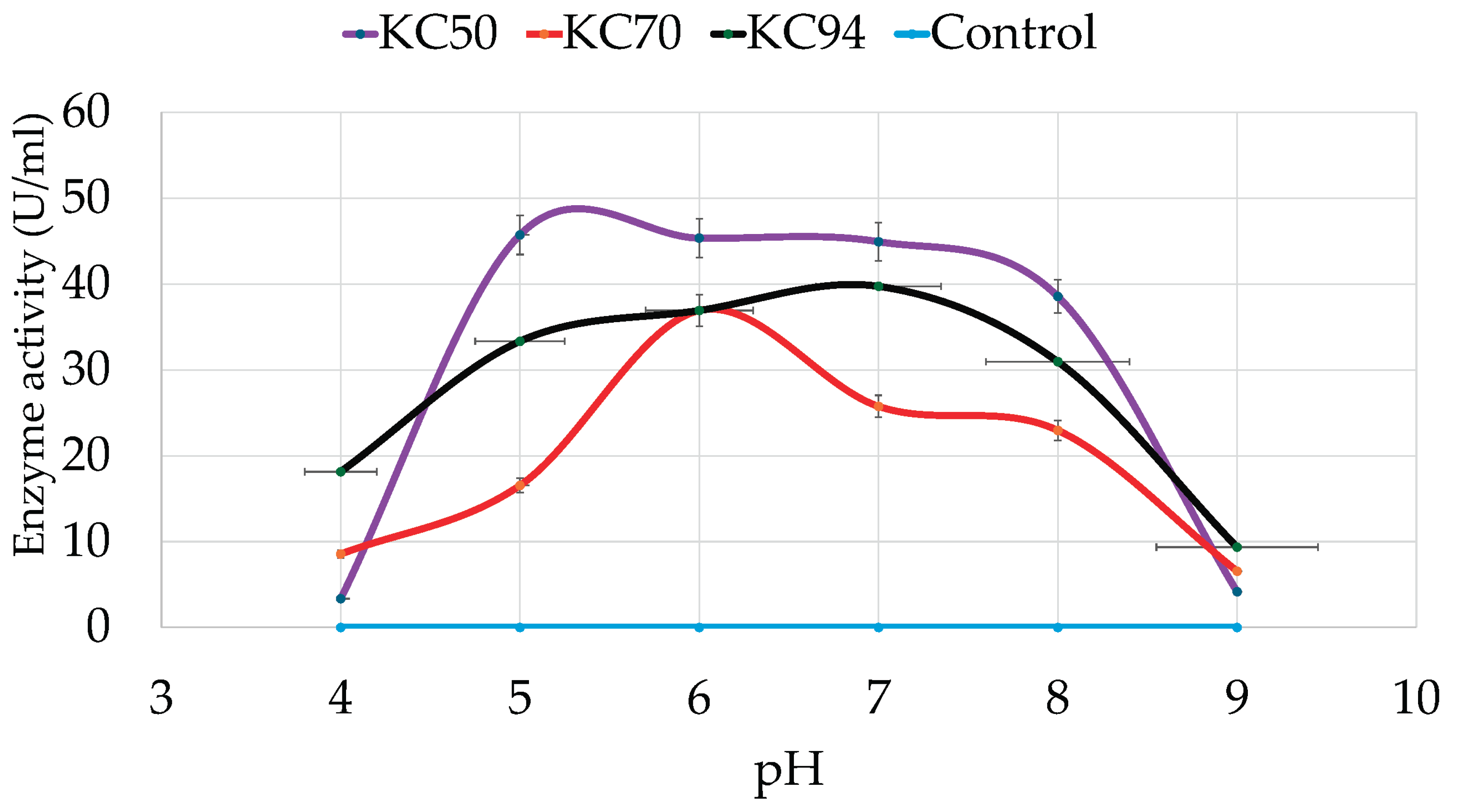
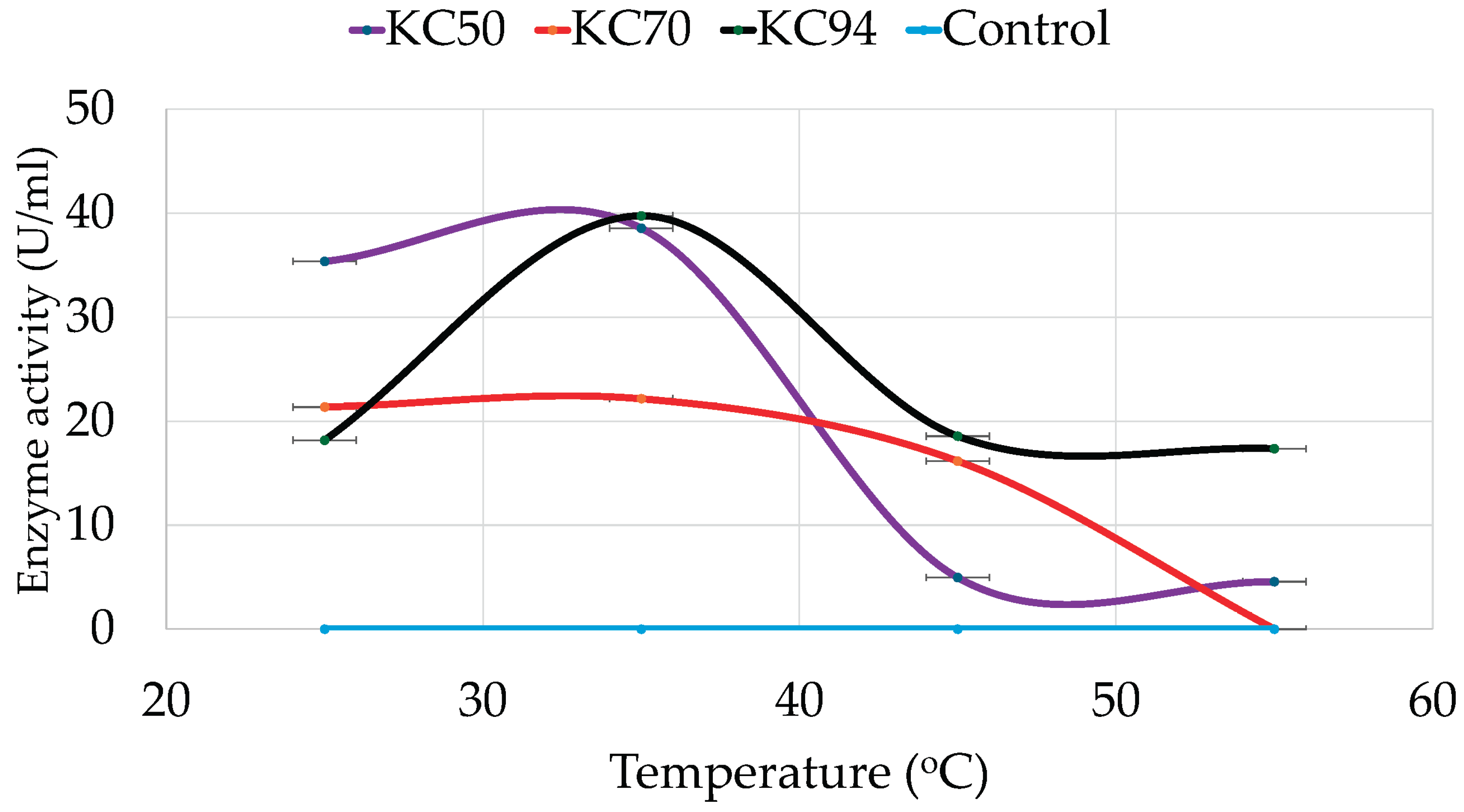

| Isolate | Colony Diameter (mm) | Clear Zone Diameter(mm) | Relative Enzyme Activity |
| KC40 | 7.33±0.12 | 16.21±0.12 | 2.21±0.03 |
| KC50 | 62.16±0.12 | 78.62±0.11 | 1.26±0.01 |
| KC70 | 8.81±0.10 | 23.41±0.10 | 2.66±0.02 |
| KC94 | 5.99±0.10 | 24.86±0.15 | 4.15±0.05 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




