Submitted:
25 August 2025
Posted:
26 August 2025
You are already at the latest version
Abstract
Keywords:
1. Maternal Effects: Beyond Genetic Inheritance
2. Epitranscriptomics: Adding Regulatory Dimension to RNA Biology
3. Early Developmental Windows and Maternal Regulation of the Fetal Epitranscriptome
4. Potential Mediators: Maternal RNAs and RNA-Modifying Enzymes
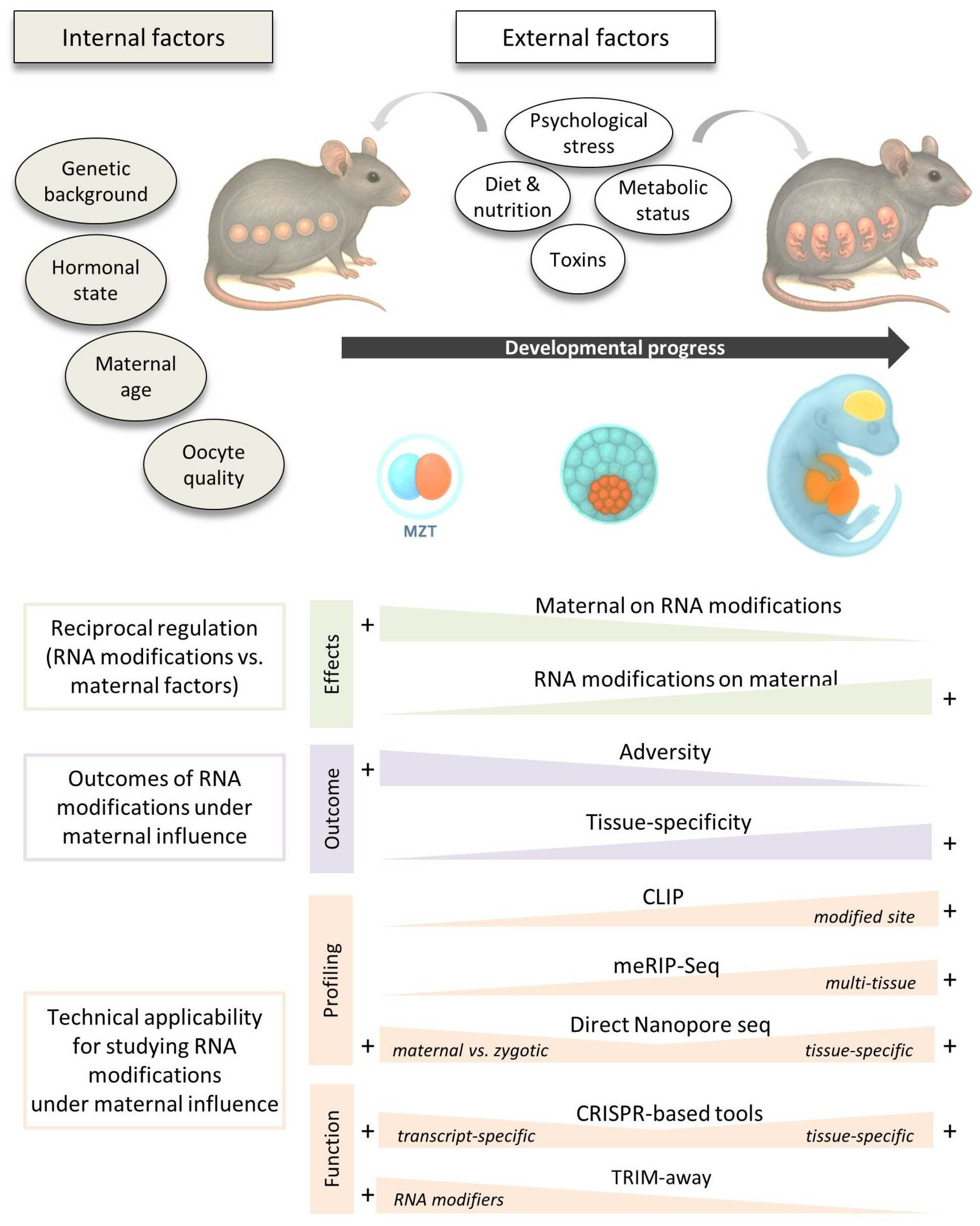
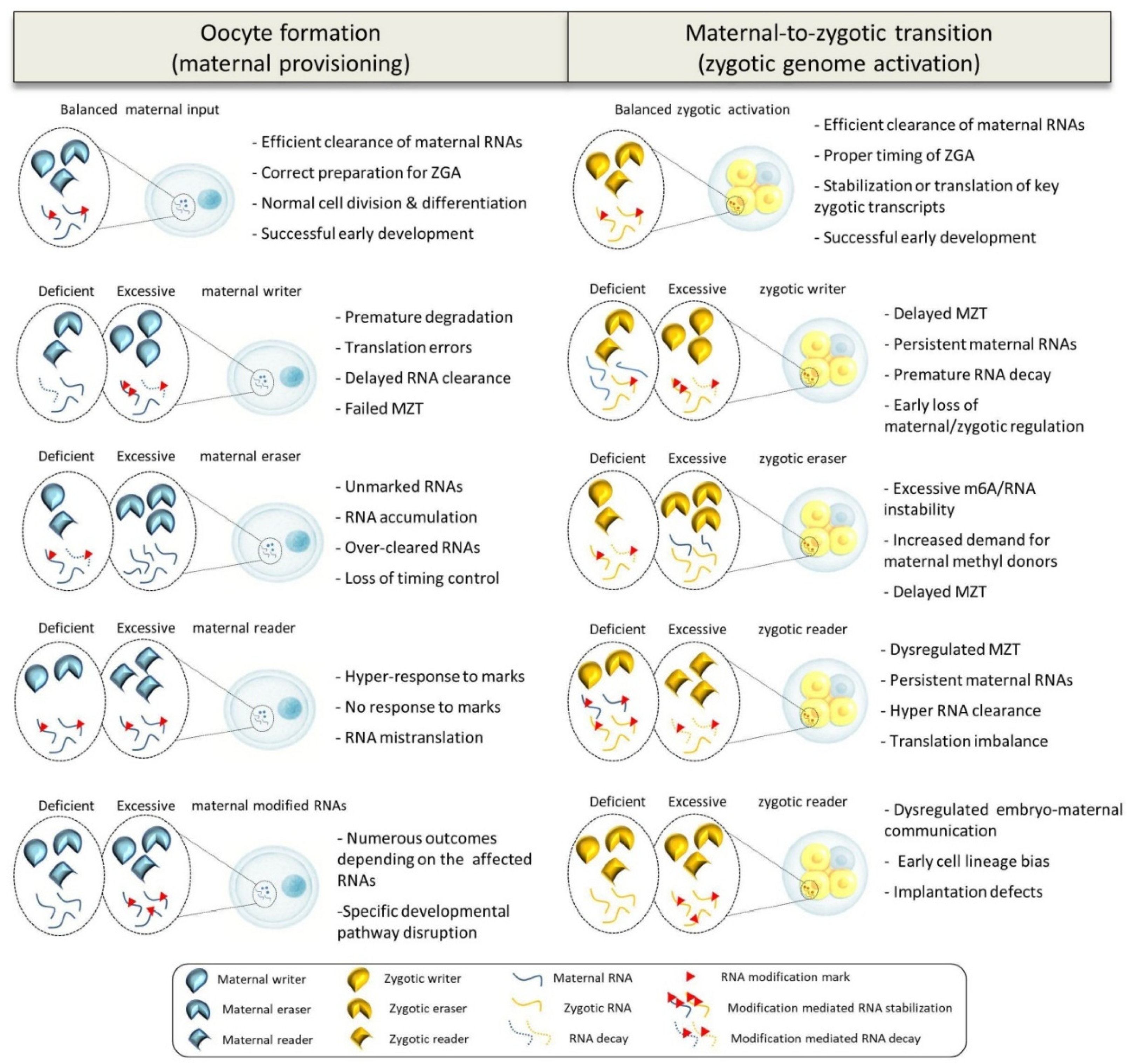
5. Evidence for Maternal-Epitranscriptomic Crosstalk: What Do We Know So Far?
6. Evolutionary and Ecological Implications of Maternal Epitranscriptomic Influence
7. Experimental Challenges and Emerging Tools to Uncover Mechanistic Links
8. Concluding Thoughts: Conceptual Bridges and Open Questions
| Box 1 | Key concepts referenced in this article |
|
Maternal effects: Non-genetic influences from the mother that shape offspring phenotype. Epitranscriptomics: Study of chemical modifications on RNA that regulate its function post-transcription. m6A (N6-methyladenosine): The most abundant mRNA modification; affects RNA stability, splicing, and translation. RNA-modifying enzymes: Proteins that install (writers), remove (erasers), or recognize (readers) RNA modifications. Maternal RNAs: Transcripts deposited into the oocyte, guiding early embryonic development. Maternal-to-zygotic transition (MZT): Developmental phase where embryonic control shifts from maternal RNAs to the zygotic genome. Placental interface: The maternal–fetal exchange site; transmits nutrients, signals, and possibly RNA regulators. Developmental plasticity: The capacity of an organism to alter its developmental trajectory in response to environmental cues. Non-genetic inheritance: Transmission of traits across generations independent of DNA sequence changes. Single-cell epitranscriptomics: Techniques that profile RNA modifications at the single-cell level for spatial and temporal precision. TRIM-away: A proteolysis method used to rapidly degrade endogenous proteins, including RNA-modifying enzymes. Phenotypic programming: The process by which environmental factors during development shape long-term traits. |
Ethics approval statement
Consent for publication
Data availability statement
Conflict of interest disclosure
Funding statement
Authors’ contributions
References
- Adrian-Kalchhauser, I., Walser, J.C., Schwaiger, M., Burkhardt-Holm, P., 2018. RNA sequencing of early round goby embryos reveals that maternal experiences can shape the maternal RNA contribution in a wild vertebrate. BMC Evol. Biol. 18, 1–14. [CrossRef]
- Agrelius, T.C., Dudycha, J.L., 2025. Maternal effects in the model system Daphnia: the ecological past meets the epigenetic future. Hered. 2025 1342 134, 142–154. [CrossRef]
- Aguilera, C., Wong, Y.S., Gutierrez-Reinoso, M.A., Velásquez, A.E., Melo-Báez, B., Cabezas, J., Caamaño, D., Navarrete, F., Castro, F.O., Rodriguez-Alvarez, L.L., 2024. Embryo-maternal communication mediated by extracellular vesicles in the early stages of embryonic development is modified by in vitro conditions. Theriogenology 214, 43–56. [CrossRef]
- Ahi, E.P., 2025. Regulation of Skeletogenic Pathways by m6A RNA Modification: A Comprehensive Review. Calcif. Tissue Int. 2025 1161 116, 1–23. [CrossRef]
- Ahi, E.P., Khorshid, M., 2025. Potentials of RNA biosensors in developmental biology. Dev. Biol. 526, 173–188. [CrossRef]
- Ahi, E.P., Singh, P., 2024. An emerging orchestrator of ecological adaptation: m6A regulation of post-transcriptional mechanisms. Mol. Ecol. 17545. [CrossRef]
- Almeida, M.V., de Jesus Domingues, A.M., Ketting, R.F., 2019. Maternal and zygotic gene regulatory effects of endogenous RNAi pathways. PLOS Genet. 15, e1007784. [CrossRef]
- Arzumanian, V.A., Dolgalev, G. V., Kurbatov, I.Y., Kiseleva, O.I., Poverennaya, E. V., 2022. Epitranscriptome: Review of Top 25 Most-Studied RNA Modifications. Int. J. Mol. Sci. 23, 13851. [CrossRef]
- Bai, S., Fu, K., Yin, H., Cui, Y., Yue, Q., Li, W., Cheng, L., Tan, H., Liu, X., Guo, Y., Zhang, Y., Xie, J., He, W., Wang, Y., Feng, H., Xin, C., Zhang, J., Lin, M., Shen, B., Sun, Z., Guo, X., Zheng, K., Ye, L., 2019. The maternal-to-zygotic transition revisited. Development 146. [CrossRef]
- Bedzhov, I., Graham, S.J.L., Leung, C.Y., Zernicka-Goetz, M., 2014. Developmental plasticity, cell fate specification and morphogenesis in the early mouse embryo. Philos. Trans. R. Soc. B Biol. Sci. 369. [CrossRef]
- Besson, A.A., Lagisz, M., Senior, A.M., Hector, K.L., Nakagawa, S., 2016. Effect of maternal diet on offspring coping styles in rodents: a systematic review and meta-analysis. Biol. Rev. 91, 1065–1080. [CrossRef]
- Bettegowda, A., Smith, G.W., 2007. Mechanisms of maternal mRNA regulation: implications for mammalian early embryonic development. Front. Biosci. 12, 3713–3726. [CrossRef]
- Bian, Y., Li, J., Shen, H., Li, Y., Hou, Y., Huang, L., Song, G., Qiao, C., 2022. WTAP dysregulation-mediated HMGN3-m6A modification inhibited trophoblast invasion in early-onset preeclampsia. FASEB J. 36, e22617. [CrossRef]
- Binan, L., Jiang, A., Danquah, S.A., Valakh, V., Simonton, B., Bezney, J., Manguso, R.T., Yates, K.B., Nehme, R., Cleary, B., Farhi, S.L., 2025. Simultaneous CRISPR screening and spatial transcriptomics reveal intracellular, intercellular, and functional transcriptional circuits. Cell 188, 2141-2158.e18. [CrossRef]
- Bjorklund, D.F., 2006. Mother knows best: Epigenetic inheritance, maternal effects, and the evolution of human intelligence. Dev. Rev. 26, 213–242. [CrossRef]
- Branco, M.R., King, M., Perez-Garcia, V., Bogutz, A.B., Caley, M., Fineberg, E., Lefebvre, L., Cook, S.J., Dean, W., Hemberger, M., Reik, W., 2016. Maternal DNA Methylation Regulates Early Trophoblast Development. Dev. Cell 36, 152–163. [CrossRef]
- Bresnahan, S.T., Lee, E., Clark, L., Ma, R., Rangel, J., Grozinger, C.M., Li-Byarlay, H., 2023. Examining parent-of-origin effects on transcription and RNA methylation in mediating aggressive behavior in honey bees (Apis mellifera). BMC Genomics 24, 1–13. [CrossRef]
- Breunig, C.T., Köferle, A., Neuner, A.M., Wiesbeck, M.F., Baumann, V., Stricker, S.H., 2021. Crispr tools for physiology and cell state changes: Potential of transcriptional engineering and epigenome editing. Physiol. Rev. 101, 177–211. [CrossRef]
- Burgueño, A.L., Juárez, Y.R., Genaro, A.M., Tellechea, M.L., 2020. Prenatal stress and later metabolic consequences: Systematic review and meta-analysis in rodents. Psychoneuroendocrinology 113, 104560. [CrossRef]
- Cayir, A., Byun, H.M., Barrow, T.M., 2020. Environmental epitranscriptomics. Environ. Res. 189, 109885. [CrossRef]
- Champagne, F.A., 2011. Maternal imprints and the origins of variation. Horm. Behav. 60, 4–11. [CrossRef]
- Champagne, F.A., Curley, J.P., 2009. Epigenetic mechanisms mediating the long-term effects of maternal care on development. Neurosci. Biobehav. Rev. 33, 593–600. [CrossRef]
- Che, Y.H., Lee, H., Kim, Y.J., 2022. New insights into the epitranscriptomic control of pluripotent stem cell fate. Exp. Mol. Med. 2022 5410 54, 1643–1651. [CrossRef]
- Chen, R., Wang, T., Tong, H., Zhang, X., Ruan, J., Qi, H., Liu, X., He, G., 2024. METTL3 and IGF2BP2 coordinately regulate FOSL1 mRNA via m6A modification, suppressing trophoblast invasion and contributing to fetal growth restriction. FASEB J. 38, e70154. [CrossRef]
- Chen, X., Liu, M., Lou, H., Lu, Y., Zhou, M.T., Ou, R., Xu, Y., Tang, K.F., 2019. Degradation of endogenous proteins and generation of a null-like phenotype in zebrafish using Trim-Away technology. Genome Biol. 20, 1–6. [CrossRef]
- Chille, E., Strand, E., Neder, M., Schmidt, V., Sherman, M., Mass, T., Putnam, H., 2021. Developmental series of gene expression clarifies maternal mRNA provisioning and maternal-to-zygotic transition in a reef-building coral. BMC Genomics 22, 1–17. [CrossRef]
- Clift, D., So, C., McEwan, W.A., James, L.C., Schuh, M., 2018. Acute and rapid degradation of endogenous proteins by Trim-Away. Nat. Protoc. 2018 1310 13, 2149–2175. [CrossRef]
- Crespo-García, E., Bueno-Costa, A., Esteller, M., 2024. Single-cell analysis of the epitranscriptome: RNA modifications under the microscope. RNA Biol. 21, 1–8. [CrossRef]
- Cummings, J.A., Clemens, L.G., Nunez, A.A., 2010. Mother counts: How effects of environmental contaminants on maternal care could affect the offspring and future generations. Front. Neuroendocrinol. 31, 440–451. [CrossRef]
- Das, A., Iwata-Otsubo, A., Destouni, A., Dawicki-McKenna, J.M., Boese, K.G., Black, B.E., Lampson, M.A., 2022. Epigenetic, genetic and maternal effects enable stable centromere inheritance. Nat. Cell Biol. 2022 245 24, 748–756. [CrossRef]
- Despic, V., Neugebauer, K.M., 2018. RNA tales - how embryos read and discard messages from mom. J. Cell Sci. 131. [CrossRef]
- Ding, Y., Zheng, Y., Wang, J., Li, H., Zhao, C., Tao, H., Li, Y., Xu, K., Huang, X., Gao, G., Chen, H., Bo, X., 2022. Recurrent RNA edits in human preimplantation potentially enhance maternal mRNA clearance. Commun. Biol. 5, 1400. [CrossRef]
- Du, R., Li, L., Wang, Y., 2022. N6-Methyladenosine-Related Gene Signature Associated With Monocyte Infiltration Is Clinically Significant in Gestational Diabetes Mellitus. Front. Endocrinol. (Lausanne). 13, 853857. [CrossRef]
- Du, Z.Y., Zhu, H.L., Chang, W., Zhang, Y.F., Ling, Q., Wang, K.W., Zhang, J., Zhang, Q.B., Kan, X.L., Wang, Q.N., Wang, H., Zhou, Y., 2024. Maternal prednisone exposure during pregnancy elevates susceptibility to osteoporosis in female offspring: The role of mitophagy/FNDC5 alteration in skeletal muscle. J. Hazard. Mater. 469, 133997. [CrossRef]
- Dvoran, M., Nemcova, L., Kalous, J., 2022. An Interplay between Epigenetics and Translation in Oocyte Maturation and Embryo Development: Assisted Reproduction Perspective. Biomed. 2022, Vol. 10, Page 1689 10, 1689. [CrossRef]
- Dworkin, M.B., Dworkin-Rastl, E., 1990. Functions of maternal mRNA in early development. Mol. Reprod. Dev. 26, 261–297. [CrossRef]
- Edwards, P.D., Lavergne, S.G., McCaw, L.K., Wijenayake, S., Boonstra, R., McGowan, P.O., Holmes, M.M., 2021. Maternal effects in mammals: Broadening our understanding of offspring programming. Front. Neuroendocrinol. 62, 100924. [CrossRef]
- Fabbiano, F., Corsi, J., Gurrieri, E., Trevisan, C., Notarangelo, M., D’Agostino, V.G., Giuseppe, V., Agostino, D.’, Risparmio, D., Trento, D., Rovereto, E., 2020. RNA packaging into extracellular vesicles: An orchestra of RNA-binding proteins? J. Extracell. Vesicles 10, e12043. [CrossRef]
- Fang, J., Wu, X., He, J., Zhang, Hanwen, Chen, X., Zhang, Hua, Novakovic, B., Qi, H., Yu, X., 2023. RBM15 suppresses hepatic insulin sensitivity of offspring of gestational diabetes mellitus mice via m6A-mediated regulation of CLDN4. Mol. Med. 29, 1–16. [CrossRef]
- Fang, L., Wang, W., Li, G., Zhang, L., Li, J., Gan, D., Yang, J., Tang, Y., Ding, Z., Zhang, M., Zhang, W., Deng, D., Song, Z., Zhu, Q., Cui, H., Hu, Y., Chen, W., 2020. CIGAR-seq, a CRISPR/Cas-based method for unbiased screening of novel mRNA modification regulators. Mol. Syst. Biol. 16, 10025. [CrossRef]
- Farley, B.M., Ryder, S.P., 2008. Regulation of Maternal mRNAs in Early Development. Crit. Rev. Biochem. Mol. Biol. 43, 135–162. [CrossRef]
- Fawcett, T.W., Frankenhuis, W.E., 2015. Adaptive explanations for sensitive windows in development. Front. Zool. 12, 1–14. [CrossRef]
- Frapin, M., Guignard, S., Meistermann, D., Grit, I., Moullé, V.S., Paillé, V., Parnet, P., Amarger, V., 2020. Maternal Protein Restriction in Rats Alters the Expression of Genes Involved in Mitochondrial Metabolism and Epitranscriptomics in Fetal Hypothalamus. Nutr. 2020, Vol. 12, Page 1464 12, 1464. [CrossRef]
- Frasch, M.G., Schulkin, J., Metz, G.A.S., Antonelli, M., 2017. Animal Models of Fetal Programming: Focus on Chronic Maternal Stress During Pregnancy and Neurodevelopment. Anim. Model. Study Hum. Dis. Second Ed. 839–849. [CrossRef]
- Galloway, L.F., 2005. Maternal effects provide phenotypic adaptation to local environmental conditions. New Phytol. 166, 93–100. [CrossRef]
- Groothuis, T.G.G., Hsu, B.Y., Kumar, N., Tschirren, B., 2019. Revisiting mechanisms and functions of prenatal hormone-mediated maternal effects using avian species as a model. Philos. Trans. R. Soc. B 374. [CrossRef]
- Grueber, C.E., Gray, L.J., Morris, K.M., Simpson, S.J., Senior, A.M., 2018. Intergenerational effects of nutrition on immunity: a systematic review and meta-analysis. Biol. Rev. 93, 1108–1124. [CrossRef]
- Haran, V., Lenka, N., 2019. Deciphering the Epitranscriptomic Signatures in Cell Fate Determination and Development. Stem Cell Rev. Reports 15, 474–496. [CrossRef]
- Harry, N.D., Zakas, C., 2023. Maternal patterns of inheritance alter transcript expression in eggs. BMC Genomics 24, 1–13. [CrossRef]
- Heng, J., Tian, M., Zhang, W., Chen, F., Guan, W., Zhang, S., 2019. Maternal heat stress regulates the early fat deposition partly through modification of m6A RNA methylation in neonatal piglets. Cell Stress Chaperones 24, 635–645. [CrossRef]
- Ho, D.H., 2014. Transgenerational Epigenetics: The Role of Maternal Effects in Cardiovascular Development. Integr. Comp. Biol. 54, 43–51. [CrossRef]
- Hu, Y., Feng, B., Wang, F., 2024. Analysis of maternal effect genes from maternal mRNA in eggs of Sogatella furcifera. Heliyon 10, e34014. [CrossRef]
- Izquierdo, V., Palomera-ávalos, V., Pallàs, M., Griñán-Ferré, C., 2021. Resveratrol Supplementation Attenuates Cognitive and Molecular Alterations under Maternal High-Fat Diet Intake: Epigenetic Inheritance over Generations. Int. J. Mol. Sci. 22, 1–18. [CrossRef]
- Jiang, Z.Y., Fan, H.Y., 2022. Five questions toward mRNA degradation in oocytes and preimplantation embryos: when, who, to whom, how, and why? Biol. Reprod. 107, 62–75. [CrossRef]
- Kaiser, S., Sachser, N., 2005. The effects of prenatal social stress on behaviour: mechanisms and function. Neurosci. Biobehav. Rev. 29, 283–294. [CrossRef]
- Kaspi, A., Khurana, I., Ziemann, M., Connor, T., Spolding, B., Zimmet, P., Walder, K., El-Osta, A., 2018. Diet during Pregnancy is Implicated in the Regulation of Hypothalamic RNA Methylation and Risk of Obesity in Offspring. Mol. Nutr. Food Res. 62, 1800134. [CrossRef]
- Kisliouk, T., Rosenberg, T., Ben-Nun, O., Ruzal, M., Meiri, N., 2020. Early-Life m6A RNA Demethylation by Fat Mass and Obesity-Associated Protein (FTO) Influences Resilience or Vulnerability to Heat Stress Later in Life. eNeuro 7, 1–14. [CrossRef]
- Kojima, M.L., Hoppe, C., Giraldez, A.J., 2024. The maternal-to-zygotic transition: reprogramming of the cytoplasm and nucleus. Nat. Rev. Genet. 2024 264 26, 245–267. [CrossRef]
- Kumar, R.P., Kumar, R., Ganguly, A., Ghosh, A., Ray, S., Islam, M.R., Saha, A., Roy, N., Dasgupta, P., Knowles, T., Niloy, A.J., Marsh, C., Paul, S., 2024. METTL3 shapes m6A epitranscriptomic landscape for successful human placentation. bioRxiv 2024.07.12.603294. [CrossRef]
- Kunovac, A., Hathaway, Q.A., Pinti, M. V., Durr, A.J., Taylor, A.D., Goldsmith, W.T., Garner, K.L., Nurkiewicz, T.R., Hollander, J.M., 2021. Enhanced antioxidant capacity prevents epitranscriptomic and cardiac alterations in adult offspring gestationally-exposed to ENM. Nanotoxicology 15, 812–831. [CrossRef]
- Kunovac, A., Hathaway, Q.A., Thapa, D., Durr, A.J., Taylor, A.D., Rizwan, S., Sharif, D., Valentine, S.J., Hollander, J.M., 2023. N6-methyladenosine (M6A) in fetal offspring modifies mitochondrial gene expression following gestational nano-TiO2 inhalation exposure. Nanotoxicology 17, 651–668. [CrossRef]
- Lecorguillé, M., Teo, S., Phillips, C.M., 2021. Maternal Dietary Quality and Dietary Inflammation Associations with Offspring Growth, Placental Development, and DNA Methylation. Nutr. 2021, Vol. 13, Page 3130 13, 3130. [CrossRef]
- Lee, M.T., Bonneau, A.R., Giraldez, A.J., 2014. Zygotic genome activation during the maternal-to-zygotic transition. Annu. Rev. Cell Dev. Biol. 30, 581–613. [CrossRef]
- Legoff, L., D’Cruz, S.C., Tevosian, S., Primig, M., Smagulova, F., 2019. Transgenerational Inheritance of Environmentally Induced Epigenetic Alterations during Mammalian Development. Cells 2019, Vol. 8, Page 1559 8, 1559. [CrossRef]
- Li, J., Gao, X., Wang, S., Yao, D., Shao, S., Wu, H., Xu, M., Yi, Q., Xie, L., Zhu, Z., Song, D., Li, H., 2023. The role of m6A methylation in prenatal maternal psychological distress and birth outcome. J. Affect. Disord. 338, 52–59. [CrossRef]
- Li, P., Lin, Y., Ma, H., Zhang, J., Zhang, Q., Yan, R., Fan, Y., 2025. Epigenetic regulation in female reproduction: the impact of m6A on maternal-fetal health. Cell Death Discov. 2025 111 11, 1–30. [CrossRef]
- Li, X., Yang, J., Zhu, Y., Liu, Y., Shi, X., Yang, G., 2016. Mouse Maternal High-Fat Intake Dynamically Programmed mRNA m6A Modifications in Adipose and Skeletal Muscle Tissues in Offspring. Int. J. Mol. Sci. 2016, Vol. 17, Page 1336 17, 1336. [CrossRef]
- Li, X., Zhao, S., Zhai, M., Ma, Y., Jiang, B., Jiang, Y., Chen, T., 2025. Extractable organic matter from PM2.5 inhibits cardiomyocyte differentiation via AHR-mediated m6A RNA methylation. J. Hazard. Mater. 486, 137110. [CrossRef]
- Li, Y., Wang, Y., Vera-Rodriguez, M., Lindeman, L.C., Skuggen, L.E., Rasmussen, E.M.K., Jermstad, I., Khan, S., Fosslie, M., Skuland, T., Indahl, M., Khodeer, S., Klemsdal, E.K., Jin, K.X., Dalen, K.T., Fedorcsak, P., Greggains, G.D., Lerdrup, M., Klungland, A., Au, K.F., Dahl, J.A., 2023. Single-cell m6A mapping in vivo using picoMeRIP–seq. Nat. Biotechnol. 2023 424 42, 591–596. [CrossRef]
- Liu, H., Zheng, J., Liao, A., 2022. The regulation and potential roles of m6A modifications in early embryonic development and immune tolerance at the maternal-fetal interface. Front. Immunol. 13, 988130. [CrossRef]
- Liu, J., Huang, T., Chen, W., Ding, C., Zhao, T., Zhao, X., Cai, B., Zhang, Y., Li, S., Zhang, L., Xue, M., He, X., Ge, W., Zhou, C., Xu, Y., Zhang, R., 2022. Developmental mRNA m5C landscape and regulatory innovations of massive m5C modification of maternal mRNAs in animals. Nat. Commun. 2022 131 13, 1–13. [CrossRef]
- Livneh, I., Moshitch-Moshkovitz, S., Amariglio, N., Rechavi, G., Dominissini, D., 2020. The m6A epitranscriptome: Transcriptome plasticity in brain development and function. Nat. Rev. Neurosci. 21, 36-51. [CrossRef]
- Lorenzo-Orts, L., Pauli, A., 2024. The molecular mechanisms underpinning maternal mRNA dormancy. Biochem. Soc. Trans. 52, 861. [CrossRef]
- Ma, J., Cain, K.D., 2025. Maternal effects on offspring immunity in fish. Fish Shellfish Immunol. 161, 110261. [CrossRef]
- Ma, X., Chen, X., Mu, X., Cao, M., Zhang, Y., 2025. Epigenetics of maternal-fetal interface immune microenvironment and placental related pregnancy complications. Front. Immunol. 16, 1549839. [CrossRef]
- Macartney, E.L., Crean, A.J., Bonduriansky, R., 2022. Parental dietary protein effects on offspring viability in insects and other oviparous invertebrates: a meta-analysis. Curr. Res. Insect Sci. 2, 100045. [CrossRef]
- Meaney, M.J., Szyf, M., 2005. Maternal care as a model for experience-dependent chromatin plasticity? Trends Neurosci. 28, 456–463. [CrossRef]
- Melnik, B.C., Weiskirchen, R., Stremmel, W., John, S.M., Schmitz, G., 2024. Risk of Fat Mass- and Obesity-Associated Gene-Dependent Obesogenic Programming by Formula Feeding Compared to Breastfeeding. Nutr. 2024, Vol. 16, Page 2451 16, 2451. [CrossRef]
- Meyer, K.D., 2019. m6A-mediated translation regulation. Biochim. Biophys. Acta - Gene Regul. Mech. 1862, 301–309. [CrossRef]
- Meyer, K.D., Jaffrey, S.R., 2014. The dynamic epitranscriptome: N6-methyladenosine and gene expression control. Nat. Rev. Mol. Cell Biol. 2014 155 15, 313–326. [CrossRef]
- Meylan, S., Miles, D.B., Clobert, J., 2012. Hormonally mediated maternal effects, individual strategy and global change. Philos. Trans. R. Soc. B Biol. Sci. 367, 1647–1664. [CrossRef]
- Mitchell, L.E., 2022. Maternal effect genes: Update and review of evidence for a link with birth defects. Hum. Genet. Genomics Adv. 3, 100067. [CrossRef]
- Moore, M.P., Whiteman, H.H., Martin, R.A., 2019. A mother’s legacy: the strength of maternal effects in animal populations. Ecol. Lett. 22, 1620–1628. [CrossRef]
- Moshitch-Moshkovitz, S., Dominissini, D., Rechavi, G., 2022. The epitranscriptome toolbox. Cell 185, 764–776. [CrossRef]
- Motorin, Y., Helm, M., 2022. RNA nucleotide methylation: 2021 update. Wiley Interdiscip. Rev. RNA 13, e1691. [CrossRef]
- Mousseau, T.A., Uller, T., Wapstra, E., Badyaev, A. V., 2009. Evolution of maternal effects: past and present. Philos. Trans. R. Soc. B Biol. Sci. 364, 1035–1038. [CrossRef]
- Navarro-Martín, L., Martyniuk, C.J., Mennigen, J.A., 2020. Comparative epigenetics in animal physiology: An emerging frontier. Comp. Biochem. Physiol. Part D Genomics Proteomics 36, 100745. [CrossRef]
- O’Brien, K., Wang, Y., 2023. The Placenta: A Maternofetal Interface. Annu. Rev. Nutr. 43, 301–325. [CrossRef]
- Park, C.W., Lee, S.M., Yoon, K.J., 2020. Epitranscriptomic regulation of transcriptome plasticity in development and diseases of the brain. BMB Rep. 53, 551. [CrossRef]
- Pollak, D.D., Weber-Stadlbauer, U., 2020. Transgenerational consequences of maternal immune activation. Semin. Cell Dev. Biol. 97, 181–188. [CrossRef]
- Potticary, A.L., Duckworth, R.A., 2020. Multiple environmental stressors induce an adaptive maternal effect. Am. Nat. 196, 487–500. [CrossRef]
- Qiu, W., Zhou, Y., Wu, H., Lv, X., Yang, L., Ren, Z., Tian, H., Yu, Q., Li, J., Lin, W., Zhao, L., Luo, S., Gao, J., 2021. RNA Demethylase FTO Mediated RNA m6A Modification Is Involved in Maintaining Maternal-Fetal Interface in Spontaneous Abortion. Front. Cell Dev. Biol. 9, 617172. [CrossRef]
- Quarto, G., Li Greci, A., Bizet, M., Penning, A., Primac, I., Murisier, F., Garcia-Martinez, L., Borges, R.L., Gao, Q., Cingaram, P.K.R., Calonne, E., Hassabi, B., Hubert, C., Herpoel, A., Putmans, P., Mies, F., Martin, J., Van der Linden, L., Dube, G., Kumar, P., Soin, R., Kumar, A., Misra, A., Lan, J., Paque, M., Gupta, Y.K., Blomme, A., Close, P., Estève, P.O., Caine, E.A., Riching, K.M., Gueydan, C., Daniels, D.L., Pradhan, S., Shiekhattar, R., David, Y., Morey, L., Jeschke, J., Deplus, R., Collignon, E., Fuks, F., 2025. Fine-tuning of gene expression through the Mettl3-Mettl14-Dnmt1 axis controls ESC differentiation. Cell 188, 998-1018.e26. [CrossRef]
- Räsänen, K., Kruuk, L.E.B., 2007. Maternal effects and evolution at ecological time-scales. Funct. Ecol. 21, 408–421. [CrossRef]
- Ren, J., Zhou, H., Zeng, H., Wang, C.K., Huang, J., Qiu, X., Sui, X., Li, Q., Wu, X., Lin, Z., Lo, J.A., Maher, K., He, Y., Tang, X., Lam, J., Chen, H., Li, B., Fisher, D.E., Liu, J., Wang, X., 2023. Spatiotemporally resolved transcriptomics reveals the subcellular RNA kinetic landscape. Nat. Methods 20, 695–705. [CrossRef]
- Ren, Z., He, J., Huang, X., Gao, Y., Wei, C., Wu, Z., Guo, W., Wang, F., Zhao, Q., Sun, X., Zhang, J., Cao, N., Lin, L., Wang, J., Cun, Y., 2025. Isoform characterization of m6A in single cells identifies its role in RNA surveillance. Nat. Commun. 2025 161 16, 1–19. [CrossRef]
- Ruebel, M.L., Latham, K.E., 2020. Listening to mother: Long-term maternal effects in mammalian development. Mol. Reprod. Dev. 87, 399–408. [CrossRef]
- Sağlam, B., Akgül, B., 2024. An Overview of Current Detection Methods for RNA Methylation. Int. J. Mol. Sci. 2024, Vol. 25, Page 3098 25, 3098. [CrossRef]
- Schier, A.F., 2007. The maternal-zygotic transition: death and birth of RNAs. Science 316, 406–407. [CrossRef]
- Schroeder, M., Fuenzalida, B., Yi, N., Shahnawaz, S., Gertsch, J., Pellegata, D., Ontsouka, E., Leiva, A., Gutiérrez, J., Müller, M., Brocco, M.A., Albrecht, C., 2024. LAT1-dependent placental methionine uptake is a key player in fetal programming of metabolic disease. Metabolism 153, 155793. [CrossRef]
- Segura-Benítez, M., Carbajo-García, M.C., Quiñonero, A., De Los Santos, M.J., Pellicer, A., Cervelló, I., Ferrero, H., 2025. Endometrial extracellular vesicles regulate processes related to embryo development and implantation in human blastocysts. Hum. Reprod. 40, 56–68. [CrossRef]
- Sha, Q.Q., Zhang, J., Fan, H.Y., 2019. A story of birth and death: mRNA translation and clearance at the onset of maternal-to-zygotic transition in mammals. Biol. Reprod. 101, 579–590. [CrossRef]
- Shama, L.N.S., Mark, F.C., Strobel, A., Lokmer, A., John, U., Mathias Wegner, K., 2016. Transgenerational effects persist down the maternal line in marine sticklebacks: gene expression matches physiology in a warming ocean. Evol. Appl. 9, 1096–1111. [CrossRef]
- Sharp, G.C., Lawlor, D.A., Richardson, S.S., 2018. It’s the mother!: How assumptions about the causal primacy of maternal effects influence research on the developmental origins of health and disease. Soc. Sci. Med. 213, 20–27. [CrossRef]
- Sharp, G.C., Salas, L.A., Monnereau, C., Allard, C., Yousefi, P., Everson, T.M., Bohlin, J., Xu, Z., Huang, R.C., Reese, S.E., Xu, C.J., Baïz, N., Hoyo, C., Agha, G., Roy, R., Holloway, J.W., Ghantous, A., Merid, S.K., Bakulski, K.M., Küpers, L.K., Zhang, H., Richmond, R.C., Page, C.M., Duijts, L., Lie, R.T., Melton, P.E., Vonk, J.M., Nohr, E.A., Williams-DeVane, C.L., Huen, K., Rifas-Shiman, S.L., Ruiz-Arenas, C., Gonseth, S., Rezwan, F.I., Herceg, Z., Ekström, S., Croen, L., Falahi, F., Perron, P., Karagas, M.R., Quraishi, B.M., Suderman, M., Magnus, M.C., Jaddoe, V.W.V., Taylor, J.A., Anderson, D., Zhao, S., Smit, H.A., Josey, M.J., Bradman, A., Baccarelli, A.A., Bustamante, M., Håberg, S.E., Pershagen, G., Hertz-Picciotto, I., Newschaffer, C., Corpeleijn, E., Bouchard, L., Lawlor, D.A., Maguire, R.L., Barcellos, L.F., Smith, G.D., Eskenazi, B., Karmaus, W., Marsit, C.J., Hivert, M.F., Snieder, H., Fallin, M.D., Melén, E., Munthe-Kaas, M.C., Arshad, H., Wiemels, J.L., Annesi-Maesano, I., Vrijheid, M., Oken, E., Holland, N., Murphy, S.K., Sørensen, T.I.A., Koppelman, G.H., Newnham, J.P., Wilcox, A.J., Nystad, W., London, S.J., Felix, J.F., Relton, C.L., 2017. Maternal BMI at the start of pregnancy and offspring epigenome-wide DNA methylation: findings from the pregnancy and childhood epigenetics (PACE) consortium. Hum. Mol. Genet. 26, 4067–4085. [CrossRef]
- Shen, W. Bin, Ni, J., Yao, R., Goetzinger, K.R., Harman, C., Reece, E.A., Wang, B., Yang, P., 2022. Maternal obesity increases DNA methylation and decreases RNA methylation in the human placenta. Reprod. Toxicol. 107, 90–96. [CrossRef]
- Song, Y.P., Lv, J.W., Zhang, Z.C., Qian, Q.H., Fan, Y.J., Chen, D.Z., Zhang, H., Xu, F.X., Zhang, C., Huang, Y., Wang, H., Wei, W., Xu, D.X., 2023. Effects of Gestational Arsenic Exposures on Placental and Fetal Development in Mice: The Role of Cyr61 m6A. Environ. Health Perspect. 131, 097004. [CrossRef]
- Sui, X., Hu, Y., Ren, C., Cao, Q., Zhou, S., Cao, Y., Li, M., Shu, W., Huo, R., 2020. METTL3-mediated m6A is required for murine oocyte maturation and maternal-to-zygotic transition. Cell Cycle 19, 391–404. [CrossRef]
- Sun, M.H., Jiang, W.J., Li, X.H., Lee, S.H., Heo, G., Zhou, D., Guo, J., Cui, X.S., 2023. High Temperature–Induced m6A Epigenetic Changes Affect Early Porcine Embryonic Developmental Competence in Pigs. Microsc. Microanal. 29, 2174–2183. [CrossRef]
- Taniguchi, K., Kawai, T., Kitawaki, J., Tomikawa, J., Nakabayashi, K., Okamura, K., Sago, H., Hata, K., 2020. Epitranscriptomic profiling in human placenta: N6-methyladenosine modification at the 5′-untranslated region is related to fetal growth and preeclampsia. FASEB J. 34, 494–512. [CrossRef]
- Vejnar, C.E., Messih, M.A., Takacs, C.M., Yartseva, V., Oikonomou, P., Christiano, R., Stoeckius, M., Lau, S., Lee, M.T., Beaudoin, J.D., Musaev, D., Darwich-Codore, H., Walther, T.C., Tavazoie, S., Cifuentes, D., Giraldez, A.J., 2019. Genome wide analysis of 3′ UTR sequence elements and proteins regulating mRNA stability during maternal-to-zygotic transition in zebrafish. Genome Res. 29, 1100–1114. [CrossRef]
- Videvall, E., Sletvold, N., Hagenblad, J., Agren, J., Hansson, B., 2016. Strong Maternal Effects on Gene Expression in Arabidopsis lyrata Hybrids. Mol. Biol. Evol. 33, 984–994. [CrossRef]
- Wagner, A., Schosserer, M., 2022. The epitranscriptome in ageing and stress resistance: A systematic review. Ageing Res. Rev. 81, 101700. [CrossRef]
- Wang, J., Gao, F., Zhao, X., Cai, Y., Jin, H., 2020. Integrated analysis of the transcriptome- wide m6A methylome in preeclampsia and healthy control placentas. PeerJ 8, e9880. [CrossRef]
- Wang, Q., Pan, M., Zhang, T., Jiang, Y., Zhao, P., Liu, X., Gao, A., Yang, L., Hou, J., 2022. Fear Stress During Pregnancy Affects Placental m6A-Modifying Enzyme Expression and Epigenetic Modification Levels. Front. Genet. 13, 927615. [CrossRef]
- Wang, S., Chen, S., Sun, J., Han, P., Xu, B., Li, X., Zhong, Y., Xu, Z., Zhang, P., Mi, P., Zhang, C., Li, L., Zhang, H., Xia, Y., Li, S., Heikenwalder, M., Yuan, D., 2023. m6A modification-tuned sphingolipid metabolism regulates postnatal liver development in male mice. Nat. Metab. 2023 55 5, 842–860. [CrossRef]
- Wei, S., Tao, H.Y., Duan, Z., Wang, Y., 2025. Environmental Exposure, Epitranscriptomic Perturbations, and Human Diseases. Environ. Sci. Technol. [CrossRef]
- Weir, E., McLinden, G., Alfandari, D., Cousin, H., 2021. Trim-Away mediated knock down uncovers a new function for Lbh during gastrulation of Xenopus laevis. Dev. Biol. 470, 74–83. [CrossRef]
- Wells, J.C.K., 2019. Developmental plasticity as adaptation: adjusting to the external environment under the imprint of maternal capital. Philos. Trans. R. Soc. B 374. [CrossRef]
- Wells, J.C.K., 2007. The thrifty phenotype as an adaptive maternal effect. Biol. Rev. 82, 143–172. [CrossRef]
- Wiener, D., Schwartz, S., 2020. The epitranscriptome beyond m6A. Nat. Rev. Genet. 2020 222 22, 119–131. [CrossRef]
- Winata, C.L., Korzh, V., 2018. The translational regulation of maternal mRNAs in time and space. FEBS Lett. 592, 3007–3023. [CrossRef]
- Winata, C.L., Łapinśki, M., Pryszcz, L., Vaz, C., Ismail, M.H. Bin, Nama, S., Hajan, H.S., Lee, S.G.P., Korzh, V., Sampath, P., Tanavde, V., Mathavan, S., 2018. Cytoplasmic polyadenylation-mediated translational control of maternal mRNAs directs maternal-to-zygotic transition. Dev. 145. [CrossRef]
- Wolf, J.B., 2000. Gene interactions from maternal effects. Evolution 54, 1882–1898. [CrossRef]
- Wolf, J.B., Wade, M.J., 2009. What are maternal effects (and what are they not)? Philos. Trans. R. Soc. B Biol. Sci. 364, 1107–1115. [CrossRef]
- Wu, Shengyu, Xie, H., Su, Y., Jia, X., Mi, Y., Jia, Y., Ying, H., 2023. The landscape of implantation and placentation: deciphering the function of dynamic RNA methylation at the maternal-fetal interface. Front. Endocrinol. (Lausanne). 14, 1205408. [CrossRef]
- Wu, Suqi, Liu, K., Zhou, B., Wu, Suwen, 2023. N6-methyladenosine modifications in maternal-fetal crosstalk and gestational diseases. Front. Cell Dev. Biol. 11, 1164706. [CrossRef]
- Wu, Y., Xu, X., Qi, M., Chen, C., Li, M., Yan, R., Kou, X., Zhao, Y., Liu, W., Li, Y., Liu, X., Zhang, M., Yi, C., Liu, H., Xiang, J., Wang, H., Shen, B., Gao, Y., Gao, S., 2022. N6-methyladenosine regulates maternal RNA maintenance in oocytes and timely RNA decay during mouse maternal-to-zygotic transition. Nat. Cell Biol. 2022 246 24, 917–927. [CrossRef]
- Xia, H., Zhong, C., Wu, X., Chen, J., Tao, B., Xia, X., Shi, M., Zhu, Z., Trudeau, V.L., Hu, W., 2018. Mettl3 mutation disrupts gamete maturation and reduces fertility in zebrafish. Genetics 208, 729–743. [CrossRef]
- Xiang, Y., Chang, H.-M., Leung, P.C.K., Bai, L., Zhu, Y., 2025. RNA modifications in female reproductive physiology and disease: emerging roles and clinical implications. Hum. Reprod. Update. [CrossRef]
- Xiao, S., Cao, S., Huang, Q., Xia, Linjian, Deng, M., Yang, M., Jia, G., Liu, X., Shi, J., Wang, W., Li, Y., Liu, S., Zhu, H., Tan, K., Luo, Q., Zhong, M., He, C., Xia, Laixin, 2019. The RNA N6-methyladenosine modification landscape of human fetal tissues. Nat. Cell Biol. 2019 215 21, 651–661. [CrossRef]
- Xiao, Y., Chen, J., Yang, S., Sun, H., Xie, L., Li, J., Jing, N., Zhu, X., 2023. Maternal mRNA deadenylation and allocation via Rbm14 condensates facilitate vertebrate blastula development. EMBO J. 42. [CrossRef]
- Xiao, Z., Liu, S., Li, Z., Cui, J., Wang, H., Wang, Zihan, Ren, Q., Xia, L., Wang, Zhijian, Li, Y., 2022. The Maternal Microbiome Programs the m6A Epitranscriptome of the Mouse Fetal Brain and Intestine. Front. Cell Dev. Biol. 10, 882994. [CrossRef]
- Xiong, Y.W., Zhu, H.L., Zhang, J., Geng, H., Tan, L.L., Zheng, X.M., Li, H., Fan, L.L., Wang, X.R., Zhang, X.D., Wang, K.W., Chang, W., Zhang, Y.F., Yuan, Z., Duan, Z.L., Cao, Y.X., He, X.J., Xu, D.X., Wang, H., 2024. Multigenerational paternal obesity enhances the susceptibility to male subfertility in offspring via Wt1 N6-methyladenosine modification. Nat. Commun. 2024 151 15, 1–20. [CrossRef]
- Yan, C., He, B., Wang, C., Li, W., Tao, S., Chen, J., Wang, Y., Yang, L., Wu, Y., Wu, Z., Liu, N., Qin, Y., 2025. Methionine in embryonic development: metabolism, redox homeostasis, epigenetic modification and signaling pathway. Crit. Rev. Food Sci. Nutr. [CrossRef]
- Yang, G., Xin, Q., Dean, J., 2024. Degradation and translation of maternal mRNA for embryogenesis. Trends Genet. 40, 238–249. [CrossRef]
- Yang, L., Ma, M., Gao, Y., Liu, J., 2025. Decoding N6-methyladenosine’s dynamic role in stem cell fate and early embryo development: insights into RNA–chromatin interactions. Curr. Opin. Genet. Dev. 91, 102311. [CrossRef]
- Yang, Y., Wang, L., Han, X., Yang, W.L., Zhang, M., Ma, H.L., Sun, B.F., Li, A., Xia, J., Chen, J., Heng, J., Wu, B., Chen, Y.S., Xu, J.W., Yang, X., Yao, H., Sun, J., Lyu, C., Wang, H.L., Huang, Y., Sun, Y.P., Zhao, Y.L., Meng, A., Ma, J., Liu, F., Yang, Y.G., 2019. RNA 5-Methylcytosine Facilitates the Maternal-to-Zygotic Transition by Preventing Maternal mRNA Decay. Mol. Cell 75, 1188-1202.e11. [CrossRef]
- Yao, Y., Liu, P., Li, Y., Wang, W., Jia, H., Bai, Y., Yuan, Z., Yang, Z., 2024. Regulatory role of m6A epitranscriptomic modifications in normal development and congenital malformations during embryogenesis. Biomed. Pharmacother. 173. [CrossRef]
- Yoon, K.J., Vissers, C., Ming, G. li, Song, H., 2018. Epigenetics and epitranscriptomics in temporal patterning of cortical neural progenitor competence. J. Cell Biol. 217, 1901–1914. [CrossRef]
- Yu, X., Xu, J., Song, B., Zhu, R., Liu, J., Liu, Y.F., Ma, Y.J., 2024. The role of epigenetics in women’s reproductive health: the impact of environmental factors. Front. Endocrinol. (Lausanne). 15, 1399757. [CrossRef]
- Zhang, G., Xu, Y., Wang, X., Zhu, Y., Wang, L., Zhang, W., Wang, Y., Gao, Y., Wu, X., Cheng, Y., Sun, Q., Chen, D., 2022. Dynamic FMR1 granule phase switch instructed by m6A modification contributes to maternal RNA decay. Nat. Commun. 2022 131 13, 1–16. [CrossRef]
- Zhang, M., Zhai, Y., Zhang, S., Dai, X., Li, Z., 2020. Roles of N6-Methyladenosine (m6A) in Stem Cell Fate Decisions and Early Embryonic Development in Mammals. Front. Cell Dev. Biol. 8, 566543. [CrossRef]
- Zhang, S., Meng, P., Cheng, S., Jiang, X., Zhang, J., Qin, X., Tang, Q., Bai, L., Zou, Z., Chen, C., 2020. Pregnancy exposure to carbon black nanoparticles induced neurobehavioral deficits that are associated with altered m6A modification in offspring. Neurotoxicology 81, 40–50. [CrossRef]
- Zhang, T.Y., Bagot, R., Parent, C., Nesbitt, C., Bredy, T.W., Caldji, C., Fish, E., Anisman, H., Szyf, M., Meaney, M.J., 2006. Maternal programming of defensive responses through sustained effects on gene expression. Biol. Psychol. 73, 72–89. [CrossRef]
- Zhao, B.S., Roundtree, I.A., He, C., 2016. Post-transcriptional gene regulation by mRNA modifications. Nat. Rev. Mol. Cell Biol. 2016 181 18, 31–42. [CrossRef]
- Zhao, B.S., Wang, X., Beadell, A. V., Lu, Z., Shi, H., Kuuspalu, A., Ho, R.K., He, C., 2017. M6 A-dependent maternal mRNA clearance facilitates zebrafish maternal-to-zygotic transition. Nature 542, 475–478. [CrossRef]
- Zheng, D., Jiang, J., Shen, A., Zhong, Y., Zhang, Y., Xiu, J., 2024. Maternal Hypertension Aggravates Vascular Dysfunction After Injury in Male Adult Offspring Through Transgenerational Transmission of N6-Methyladenosine. Hypertension. [CrossRef]
- Zheng, Z.H., Zhang, G. Le, Jiang, R.F., Hong, Y.Q., Zhang, Q.Y., He, J.P., Liu, X.R., Yang, Z.S., Yang, L., Jiang, X., Qu, L.J., Ding, C.H., Xu, Y.W., Yang, S.H., Liu, J.L., 2023. METTL3 is essential for normal progesterone signaling during embryo implantation via m6A-mediated translation control of progesterone receptor. Proc. Natl. Acad. Sci. U. S. A. 120, e2214684120. [CrossRef]
- Zhong, Z.D., Xie, Y.Y., Chen, H.X., Lan, Y.L., Liu, X.H., Ji, J.Y., Wu, F., Jin, L., Chen, J., Mak, D.W., Zhang, Z., Luo, G.Z., 2023. Systematic comparison of tools used for m6A mapping from nanopore direct RNA sequencing. Nat. Commun. 2023 141 14, 1–14. [CrossRef]
- Zhou, W., Xue, P., Yang, Y., Xia, L., Yu, B., 2022. Research progress on N6-methyladenosine in the human placenta. J. Perinat. Med. 50, 1115–1123. [CrossRef]
- Zhu, W., Ding, Y., Meng, J., Gu, L., Liu, W., Li, L., Chen, H., Wang, Y., Li, Z., Li, C., Sun, Y., Liu, Z., 2023. Reading and writing of mRNA m6A modification orchestrate maternal-to-zygotic transition in mice. Genome Biol. 24, 1–16. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




