Submitted:
22 August 2025
Posted:
25 August 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
Identification of CpG Islands and Primer Design
Cell Culture
DNA Extraction, Processing of Controls and Bisulfite Treatment
Bisulfite Sequencing PCR and Pyrosequencing
RNA Extraction and cDNA Synthesis
Plasmid Preparation and Linearization
Reverse Transcriptase Quantitative PCR
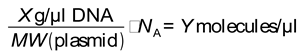
Statistical Analysis
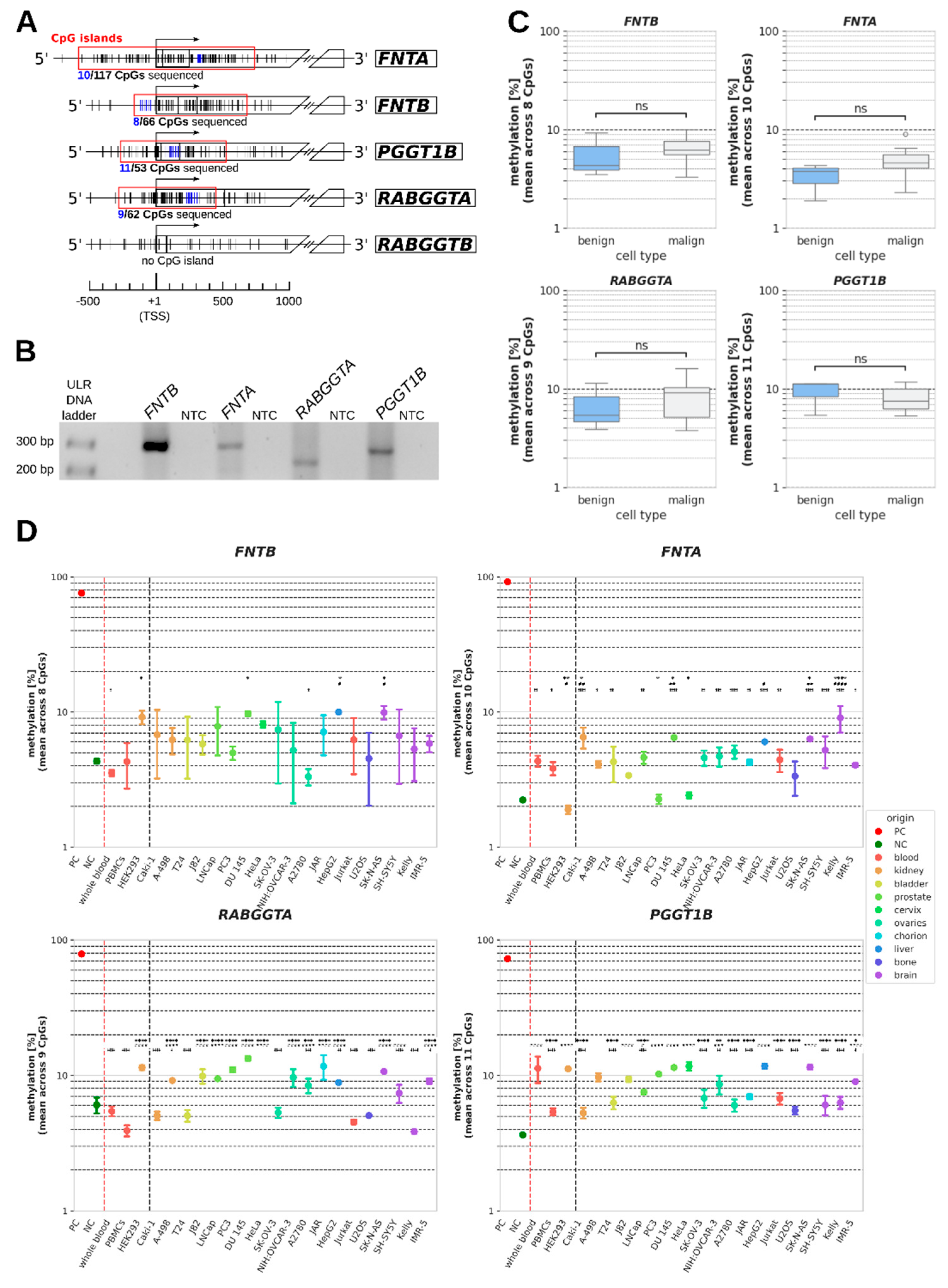
3. Results
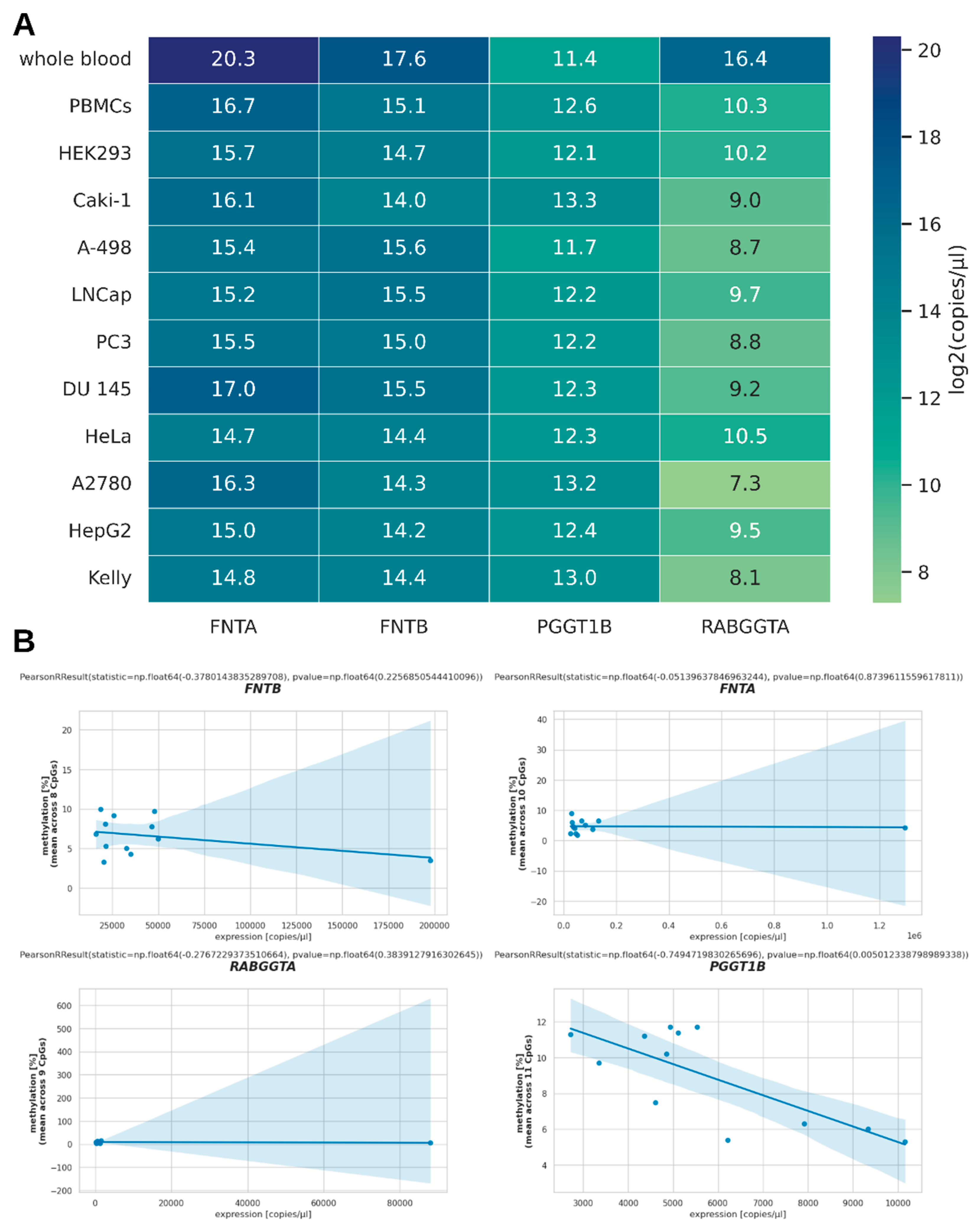
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Kuchay S, Wang H, Marzio A, Jain K, Homer H, Fehrenbacher N, Philips MR, Zheng N, Pagano M: GGTase3 is a newly identified geranylgeranyltransferase targeting a ubiquitin ligase. Nat Struct Mol Biol 2019, 26(7):628-636. [CrossRef]
- Shirakawa R, Goto-Ito S, Goto K, Wakayama S, Kubo H, Sakata N, Trinh DA, Yamagata A, Sato Y, Masumoto H et al: A SNARE geranylgeranyltransferase essential for the organization of the Golgi apparatus. Embo j 2020, 39(8):e104120. [CrossRef]
- Tanimoto H, Mehta KD, Parmley TH, Shigemasa K, Parham GP, Clarke J, O'Brien TJ: Expression of the farnesyltransferase beta-subunit gene in human ovarian carcinoma: correlation to K-ras mutation. Gynecol Oncol 1997, 66(2):308-312. [CrossRef]
- Lackner MR, Kindt RM, Carroll PM, Brown K, Cancilla MR, Chen C, de Silva H, Franke Y, Guan B, Heuer T et al: Chemical genetics identifies Rab geranylgeranyl transferase as an apoptotic target of farnesyl transferase inhibitors. Cancer Cell 2005, 7(4):325-336. [CrossRef]
- Khan SG, Dummer R, Siddiqui J, Bickers DR, Agarwal R, Mukhtar H: Farnesyltransferase activity and mRNA expression in human skin basal cell carcinomas. Biochem Biophys Res Commun 1996, 220(3):795-801.
- Zhou X, Qian J, Hua L, Shi Q, Liu Z, Xu Y, Sang B, Mo J, Yu R: Geranylgeranyltransferase I promotes human glioma cell growth through Rac1 membrane association and activation. J Mol Neurosci 2013, 49(1):130-139. [CrossRef]
- Sui GD, Zhang GY, Niu ZJ, Hu SY: Expression of farnesyltransferase in primary liver cancer. Chin Med J (Engl) 2012, 125(14):2427-2431.
- Kulis M, Esteller M: 2 - DNA Methylation and Cancer. In: Advances in Genetics. Volume 70, edn. Edited by Herceg Z, Ushijima T: Academic Press; 2010: 27-56.
- Wang XX, Ying P, Diao F, Wang Q, Ye D, Jiang C, Shen N, Xu N, Chen WB, Lai SS et al: Altered protein prenylation in Sertoli cells is associated with adult infertility resulting from childhood mumps infection. J Exp Med 2013, 210(8):1559-1574.
- He X, Chen Y, Xia Y, Hong X, You H, Zhang R, Liang Z, Cui Q, Zhang S, Zhou M et al: DNA methylation regulates biosynthesis of tanshinones and phenolic acids during growth of Salvia miltiorrhiza. Plant Physiol 2023. [CrossRef]
- Takai D, Jones PA: Comprehensive analysis of CpG islands in human chromosomes 21 and 22. Proc Natl Acad Sci U S A 2002, 99(6):3740-3745. [CrossRef]
- Kim Y, Kim T-H, Ergün T: The instability of the Pearson correlation coefficient in the presence of coincidental outliers. Finance Research Letters 2015, 13:243-257.
- Antequera F, Boyes J, Bird A: High levels of de novo methylation and altered chromatin structure at CpG islands in cell lines. Cell 1990, 62(3):503-514. [CrossRef]
- Heredia-Mendez AJ, Sánchez-Sánchez G, López-Camarillo C: Reprogramming of the Genome-Wide DNA Methylation Landscape in Three-Dimensional Cancer Cell Cultures. Cancers (Basel) 2023, 15(7):1991. [CrossRef]
- Ueki T, Walter KM, Skinner H, Jaffee E, Hruban RH, Goggins M: Aberrant CpG island methylation in cancer cell lines arises in the primary cancers from which they were derived. Oncogene 2002, 21(13):2114-2117. [CrossRef]
- Şükrüoğlu Erdoğan Ö, Kılıç Erciyas S, Bilir A, Buğra Tunçer Ş, Akdeniz Ödemiş D, Kurul S, Karanlık H, Cabıoğlu N, Yazıcı H: Methylation Changes of Primary Tumors, Monolayer, and Spheroid Tissue Culture Environments in Malignant Melanoma and Breast Carcinoma. Biomed Res Int 2019, 2019:1407167. [CrossRef]
- Barrera V, Peinado MA: Evaluation of single CpG sites as proxies of CpG island methylation states at the genome scale. Nucleic Acids Res 2012, 40(22):11490-11498. [CrossRef]
- van Vlodrop IJ, Niessen HE, Derks S, Baldewijns MM, van Criekinge W, Herman JG, van Engeland M: Analysis of promoter CpG island hypermethylation in cancer: location, location, location! Clin Cancer Res 2011, 17(13):4225-4231. [CrossRef]
- Iwata K, Nakabayashi K, Ishiwata K, Nakamura K, Kameno Y, Hata K, Matsuzaki H: Genome-wide DNA methylation profiles in the raphe nuclei of patients with autism spectrum disorder. Psychiatry Clin Neurosci 2025, 79(7):415-424. [CrossRef]
- Liu YX, Li QZ, Cao YN: The effect of key DNA methylation in different regions on gene expression in hepatocellular carcinoma. Mol Omics 2022, 18(1):57-70. [CrossRef]
| difference to reference cells |
FNTA (mean of 10 CpGs) |
difference to reference cells |
FNTB (mean of 8 CpGs) |
Gene | Origin | ||||
| Chr8: 43,056,611 -43,056,636 |
Chr14: 64,986,670 -64,986,749 |
CpG locationb | |||||||
| 2.2 ± 1.1 | 4.3 ± 1.6 | neg. control | |||||||
| 91.9 ± 7.7 | 75.8 ± 15.0 | pos. control | |||||||
| HEK293 | PBMCs | whole blood | HEK293 | PBMCs | whole blood | ||||
| ‡‡ 2.4 (0.6 to 4.3) |
- | 4.3 ± 1.2 | ‡ -5.7 (-11.2 to -0.1) |
- | 3.5 ± 0.9 | whole blood | blood | ||
| ‡ 1.9 (0.1 to 3.8) |
- | 3.8 ± 0.8 | - | 4.3 ± 0.8 | PBMCs | blood | |||
| - | # -1.9 (-3.8 to -0.1) |
** -2.4 (-4.3 to -0.6) |
1.9 ± 0.9 | - | * 5.7 (0.1 to 11.2) |
9.2 ± 3.7 | HEK293 | kidney | |
|
‡‡‡‡ 4.6 (2.7 to 6.5) |
## 2.7 (0.8 to 4.6) |
* 2.2 (0.3 to 4.1) |
6.5 ± 2.7 | 6.8 ± 2.1 | Caki-1 | kidney | |||
| ‡ 2.2 (0.3 to 4.1) |
4.1 ± 2.4 | 6.2 ± 2.3 | A-498 | kidney | |||||
| ‡‡ 2.4 (0.5 to 4.3) |
4.3 ± 1.7 | 6.2 ± 1.9 | T24 | bladder | |||||
| 3.4 ± 1.8 | 5.8 ± 2.2 | J82 | bladder | ||||||
| ‡‡ 2.7 (0.8 to 4.6) |
4.6 ± 1.7 | 7.8 ± 3.1 | LNCaP | prostate | |||||
| * -2.0 (-3.9 to -0.2) |
2.3 ± 1.0 | 5.0 ± 2.0 | PC3 | prostate | |||||
|
‡‡‡‡ 4.6 (2.7 to 6.5) |
## 2.7 (0.8 to 4.5) |
* 2.2 (0.3 to 4.0) |
6.5 ± 3.9 | * 6.2 (0.6 to 11.7) |
9.7 ± 5.1 | DU 145 | prostate | ||
| * -1.9 (-3.8 to -0.0) |
2.4 ± 1.0 | 8.1 ± 3.8 | HeLa | cervix | |||||
| ‡‡ 2.7 (0.8 to 4.6) |
4.6 ± 2.2 | 7.4 ± 2.0 | SK-OV-3 | ovaries | |||||
| ‡‡‡ 2.8 (0.9 to 4.7) |
4.7 ± 1.7 | 5.2 ± 1.3 | NIH:OVCAR-3 | ovaries | |||||
| ‡‡‡‡ 3.2 (1.3 to 5.1) |
5.1 ± 2.1 |
‡ -5.9 (-11.4 to -0.3) |
3.3 ± 0.9 | A2780 | ovaries | ||||
| ‡‡ 2.3 (0.5 to 4.2) |
4.2 ± 1.7 | 7.1 ± 2.9 | JAR | chorion | |||||
| ‡‡‡‡ 4.1 (2.2 to 6.0) |
# 2.2 (0.3 to 4.1) |
6.0 ± 3.6 | # 5.7 (0.2 to 11.2) |
* 6.5 (0.9 to 12.0) |
10.0 ± 4.9 | HepG2 | liver | ||
| ‡‡ 2.5 (0.7 to 4.4) |
4.4 ± 1.9 | 6.2 ± 1.0 | Jurkat | blood | |||||
| 3.4 ± 1.4 | 4.5 ± 1.2 | U2OS | bone | ||||||
|
‡‡‡‡ 4.4 (2.5 to 6.3) |
## 2.5 (0.6 to 4.4) |
* 2.0 (0.1 to 3.9) |
6.3 ± 4.0 | # 5.6 (0.1 to 11.2) |
* 6.4 (0.8 to 11.9) |
9.9 ± 5.2 | SK-N-AS | brain | |
| ‡‡‡‡ 3.3 (1.4 to 5.2) |
5.2 ± 2.4 | 6.7 ± 2.1 | SH-SY5Y | brain | |||||
|
‡‡‡‡ 7.1 (5.3 to 9.0) |
#### 5.2 (3.3 to 7.1) |
**** 4.7 (2.8 to 6.6) |
9.0 ± 4.2 | 5.3 ± 1.6 | Kelly | brain | |||
| ‡ 2.1 (0.3 to 4.0) |
4.0 ± 2.5 | 5.8 ± 2.3 | IMR-5 | brain | |||||
| difference to reference cells |
PGGT1B (mean of 11 CpGs) |
difference to reference cells |
RABGGTA (mean of 9 CpGs) |
Gene | Origin | ||||
| Chr5: 115,262,704 -115,262,816 |
Chr14: 24,271,320 -24,271,399 |
CpG locationb | |||||||
| 3.6 ± 1.1 | 6.0 ± 2.4 | neg. control | |||||||
| 72.7 ± 14.1 | 78.8 ± 11.2 | pos. control | |||||||
| HEK293 | PBMCs | whole blood | HEK293 | PBMCs | whole blood | ||||
| #### 5.9 (3.9 to 7.9) |
- | 11.3 ± 6.9 | ‡‡‡‡ -6.0 (-8.0 to -4.0) |
- | 5.4 ± 2.1 | whole blood | blood | ||
| ‡‡‡‡ -5.8 (-7.8 to -3.8) |
- | **** -5.9 (-7.9 to -3.9) |
5.4 ± 1.1 | ‡‡‡‡ -7.5 (-9.5 to -5.5) |
- | 3.9 ± 1.6 | PBMCs | blood | |
| - | #### 5.8 (3.8 to 7.8) |
11.2 ± 2.2 | - | #### 7.5 (5.5 to 9.5) |
**** 6.0 (4.0 to 8.0) |
11.4 ± 4.1 | HEK293 | kidney | |
| ‡‡‡‡ -5.9 (-7.9 to -3.9) |
**** -6.0 (-8.0 to -4.0) |
5.3 ± 1.2 | ‡‡‡‡ -6.4 (-8.4 to -4.4) |
5.0 ± 1.9 | Caki-1 | kidney | |||
| #### 4.3 (2.3 to 6.3) |
9.7 ± 2.2 |
‡ -2.2 (-4.3 to -0.3) |
#### 5.3 (3.3 to 7.3) |
**** 3.8 (1.7 to 5.7) |
9.2 ± 3.4 | A-498 | kidney | ||
| ‡‡‡‡ -4.9 (-6.9 to -2.9) |
**** -5.0 (-7.0 to -3.0) |
6.3 ± 1.4 | ‡‡‡‡ -6.4 (-8.4 to -4.4) |
5.0 ± 2.1 | T24 | bladder | |||
| #### 3.9 (1.9 to 6.0) |
9.3 ± 2.0 | #### 6.0 (4.0 to 8.0) |
**** 4.5 (2.4 to 6.4) |
9.9 ± 3.3 | J82 | bladder | |||
|
‡‡‡‡ -3.7 (-5.7 to -1.7) |
# 2.1 (0.1 to 4.1) |
**** -3.8 (-5.8 to -1.7) |
7.5 ± 1.9 | #### 5.6 (3.5 to 7.5) |
**** 4.1 (2.0 to 6.0) |
9.5 ± 3.6 | LNCaP | prostate | |
| #### 4.8 (2.8 to 6.8) |
10.2 ± 7.1 | #### 7.1 (5.1 to 9.1) |
**** 5.6 (3.6 to 7.6) |
11.0 ± 7.1 | PC3 | prostate | |||
| #### 6.0 (4.0 to 8.1) |
11.4 ± 2.5 | #### 9.4 (7.4 to 11.4) |
**** 7.9 (5.9 to 9.9) |
13.3 ± 4.8 | DU 145 | prostate | |||
| #### 6.3 (4.3 to 8.3) |
11.7 ± 2.6 |
‡‡‡‡ 4.6 (2.6 to 6.6) |
#### 12.1 (10.1 to 14.1) |
**** 10.6 (8.6 to 12.6) |
16.0 ± 5.4 | HeLa | cervix | ||
| ‡‡‡‡ -4.4 (-6.4 to -2.4) |
**** -4.5 (-6.5 to -2.5) |
6.8 ± 2.9 | ‡‡‡‡ -6.1 (-8.1 to -4.1) |
5.3 ± 2.0 | SK-OV-3 | ovaries | |||
|
‡‡ -2.6 (-4.6 to -0.6) |
### 3.2 (1.2 to 5.2) |
** -2.7 (-4.7 to -0.7) |
8.6 ± 2.1 |
#### 5.8 (3.8 to 7.7) |
**** 4.3 (2.2 to 6.2) |
9.7 ± 3.5 | NIH:OVCAR-3 | ovaries | |
| ‡‡‡‡ -5.2 (-7.2 to -3.2) |
**** -5.3 (-7.3 to -3.3) |
6.0 ± 1.6 |
‡‡‡ -3.0 (-5.0 to -1.0) |
#### 4.5 (2.5 to 6.5) |
*** 3.0 (1.0 to 5.0) |
8.4 ± 3.4 | A2780 | ovaries | |
| ‡‡‡‡ -4.2 (-6.2 to -2.2) |
**** -4.3 (-6.3 to -2.3) |
7.0 ± 1.7 | #### 7.8 (5.8 to 9.8) |
**** 6.3 (4.2 to 8.2) |
11.7 ± 4.5 | JAR | chorion | ||
| #### 6.3 (4.3 to 8.3) |
11.7 ± 2.5 |
‡‡ -2.5 (-4.6 to -0.6) |
#### 5.0 (2.9 to 6.9) |
**** 3.5 (1.4 to 5.4) |
8.9 ± 3.7 | HepG2 | liver | ||
| ‡‡‡‡ -4.5 (-6.5 to -2.4) |
**** -4.6 (-6.5 to -2.5) |
6.7 ± 1.5 | ‡‡‡‡ -6.9 (-8.9 to -4.9) |
4.5 ± 1.9 | Jurkat | blood | |||
| ‡‡‡‡ -5.7 (-7.7 to -3.7) |
**** -5.8 (-7.8 to -3.8) |
5.5 ± 1.3 | ‡‡‡‡ -6.3 (-8.4 to -4.4) |
5.1 ± 1.8 | U2OS | bone | |||
| #### 6.1 (4.1 to 8.1) |
11.5 ± 2.5 | #### 6.8 (4.8 to 8.8) |
**** 5.3 (3.2 to 7.2) |
10.7 ± 4.4 | SK-N-AS | brain | |||
| ‡‡‡‡ -5.1 (-7.1 to -3.1) |
**** -5.2 (-7.2 to -3.2) |
6.1 ± 1.3 | ‡‡‡‡ -4.0 (-6.0 to -2.0) |
#### 3.5 (1.5 to 5.5) |
7.4 ± 2.9 | SH-SY5Y | brain | ||
| ‡‡‡‡ -4.9 (-6.9 to -2.9) |
**** -5.0 (-7.0 to -3.0) |
6.3 ± 1.2 | ‡‡‡‡ -7.6 (-9.6 to -5.6) |
3.8 ± 1.5 | Kelly | brain | |||
|
‡ -2.2 (-4.2 to -0.2) |
#### 3.6 (1.6 to 5.6) |
* -2.3 (-4.3 to -0.3) |
9.0 ± 1.8 |
‡ -2.3 (-4.4 to -0.4) |
#### 5.2 (3.2 to 7.2) |
**** 3.7 (1.6 to 5.6) |
9.1 ± 3.3 | IMR-5 | brain |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




