Submitted:
01 August 2025
Posted:
05 August 2025
You are already at the latest version
Abstract
Keywords:
Findings
Introduction
Solution
- Web pages and their content should be open source and globally accessible.
- The service should be sufficiently intuitive and well-documented to allow a SIG to create a page without programming and web development knowledge, and without having to deploy additional services.
- The Engine should render a standardised template to provide a familiar and consistent user experience between Labs and between Galaxy servers.
- The Engine should provide mechanisms for customization by the requesting Galaxy server.
Live examples of Galaxy Labs
Conclusion
Methods
Architecture and Design
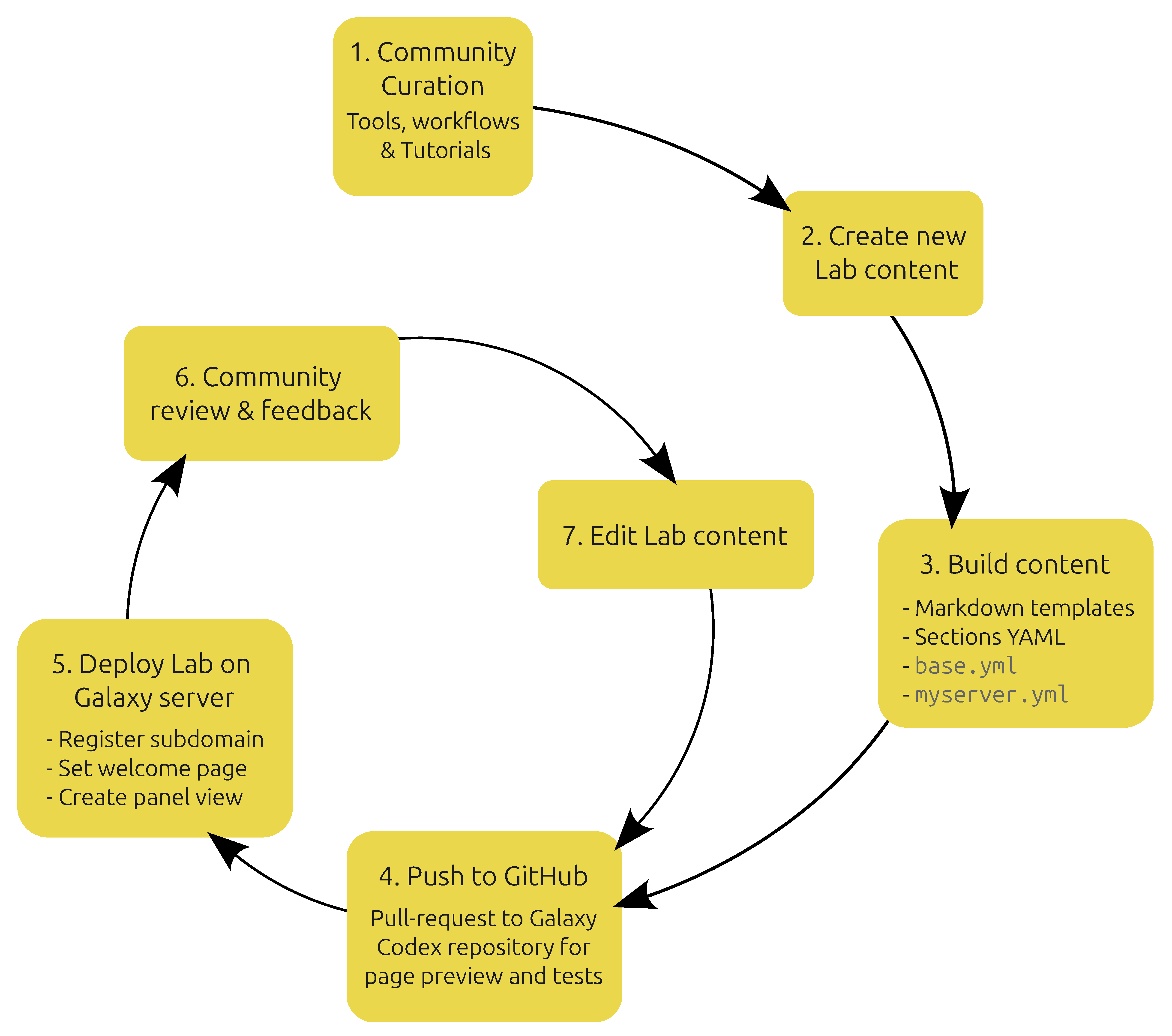
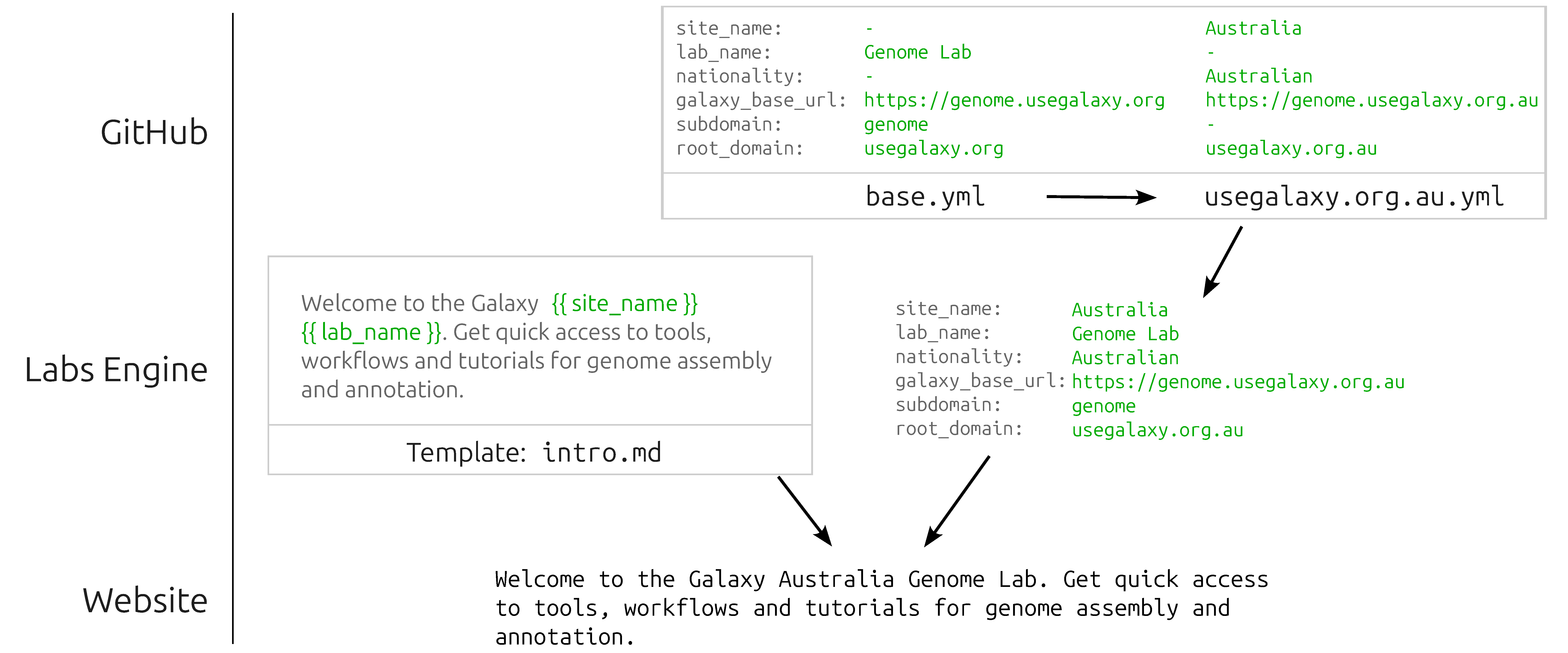
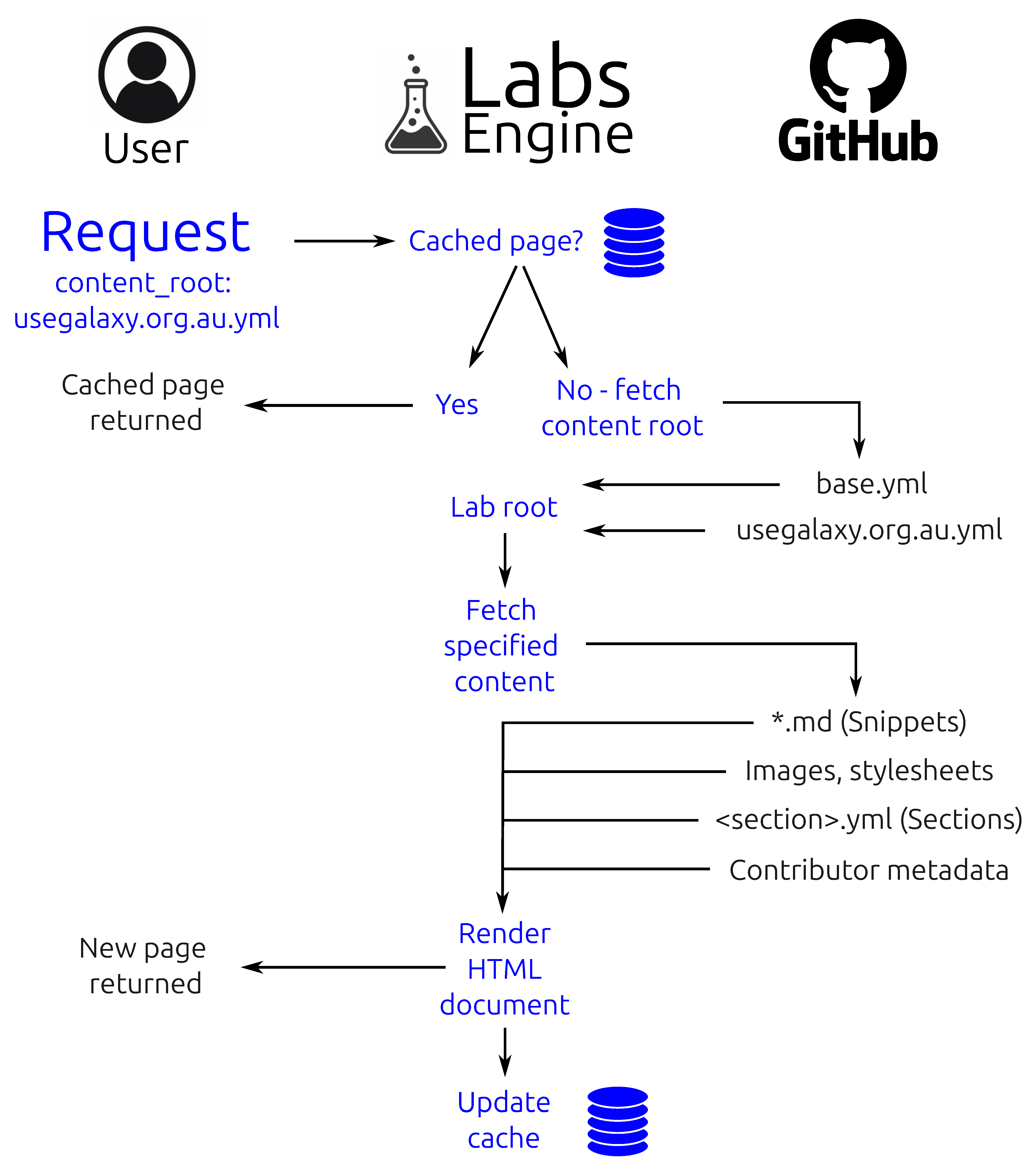
- Base template: the HTML base template from which all Lab pages are rendered.
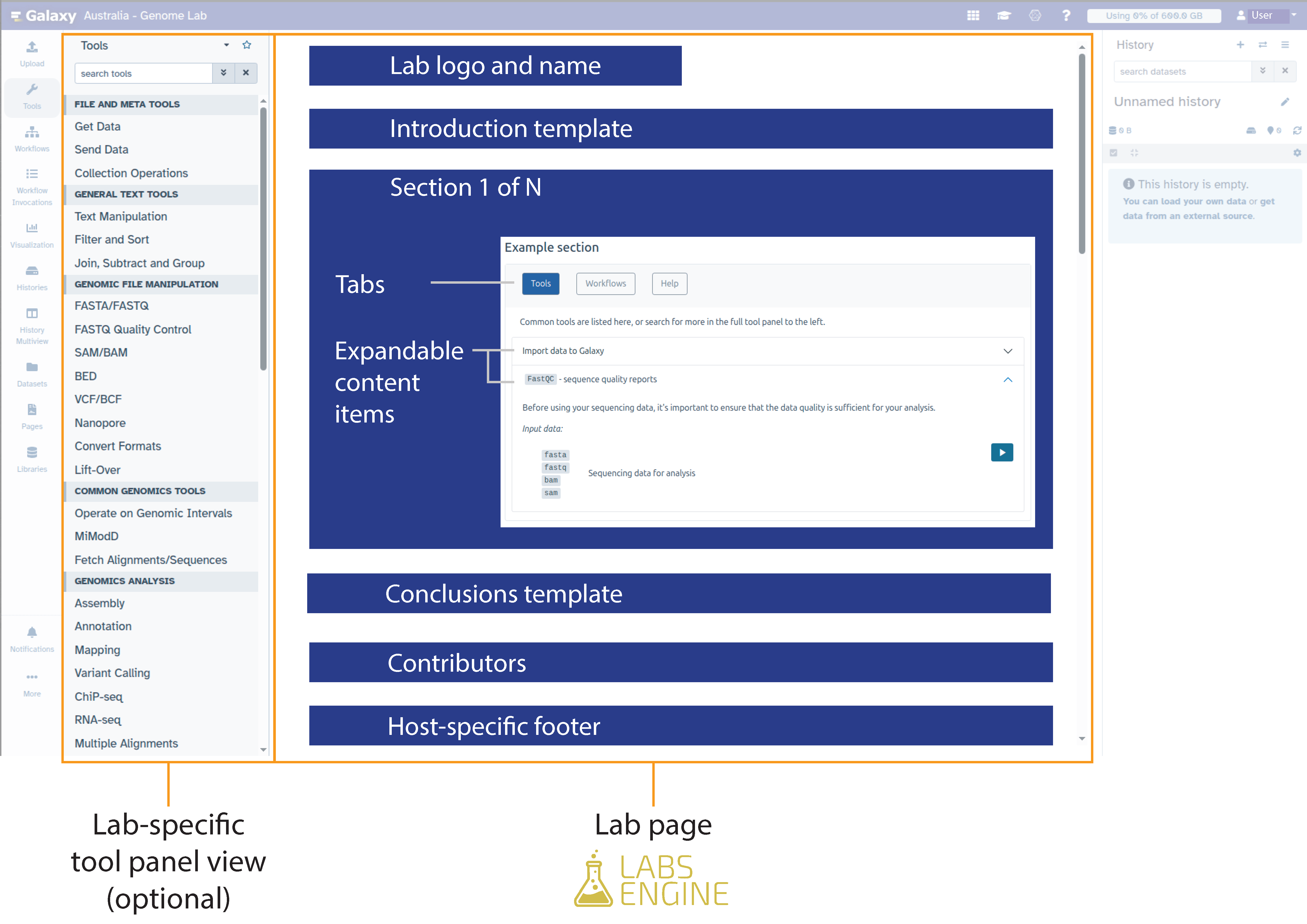
- Section schema: a schema which defines the accepted data structures for YAML content which populates the Sections component (Figure 1). This is the main body of the Lab which displays resources curated by the community with a structured user interface.
- Rendering engine: the web server endpoint which downloads YAML context, snippets and other arbitrary content (from a location on GitHub), validates it against the Section schema, and renders it all into the base template to produce the Lab web page.
- YAML context: A set of YAML files which provide the context for rendering templates and sections, while also declaring which icon and Snippets should be downloaded for the given Lab. Data defined in base.yml is overridden with a <server>.yml file in a cascade that provides fine-grained customization for each server that deploys a given Lab.
- Snippets: three Lab-specific Markdown templates (Introduction, Conclusion, Footer) which are injected into specific locations in the base template to provide server-specific customization.
Lab Content Structure
User Interface Design
Creating Lab Content
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- The Galaxy platform for accessible, reproducible, and collaborative data analyses: 2024 update. Nucleic Acids Res 2024, 52, W83–W94.
- Hiltemann S, Rasche H, Gladman S, Hotz H-R, Larivière D, Blankenberg D, et al. Galaxy Training: A powerful framework for teaching! PLoS Comput Biol 2023, 19, e1010752. [Google Scholar]
- Rhie A, McCarthy SA, Fedrigo O, Damas J, Formenti G, Koren S, et al. Towards complete and error-free genome assemblies of all vertebrate species. Nature 2021, 592, 737–746. [Google Scholar]
- Mehta S, Bernt M, Chambers M, Fahrner M, Föll MC, Gruening B, et al. A Galaxy of informatics resources for MS-based proteomics. Expert Rev Proteomics 2023, 20, 251–266. [Google Scholar]
- Afgan E, Sloggett C, Goonasekera N, Makunin I, Benson D, Crowe M, et al. Genomics virtual laboratory: a practical bioinformatics workbench for the cloud. PLoS One 2015, 10, e0140829. https://journals.plos.org/plosone/article/file?id=10.1371/journal.pone.0140829&type=printable.
- Libouban R, Mercier E, Chaussepied T, Batut B, Bretaudeau A, Le Corguillé G. Misconceptions about Galaxy debunked by the (French) Galaxy Community. In: jobim 2024; 2024. https://hal.science/hal-04694551/.
- Tekman M, Batut B, Ostrovsky A, Antoniewski C, Clements D, Ramirez F, et al. A single-cell RNA-sequencing training and analysis suite using the Galaxy framework. GigaScience 2020, 9, giaa102. https://academic.oup.com/gigascience/article-pdf/doi/10.1093/gigascience/giaa102/33956599/giaa102.pdf.
- Nasr E, Amato P, Bhardwaj A, Blankenberg D, Brites D, Cumbo F, et al. microGalaxy: A gateway to tools, workflows, and training for reproducible and FAIR analysis of microbial data. bioRxiv. 2024. https://www.biorxiv.org/content/10.1101/2024.12.23.629682.full.pdf.
- de Koning W, Miladi M, Hiltemann S, Heikema A, Hays JP, Flemming S, et al. NanoGalaxy: Nanopore long-read sequencing data analysis in Galaxy. GigaScience 2020, 9, giaa105. https://academic.oup.com/gigascience/article-pdf/9/10/giaa105/60688406/giaa105.pdf.
- Tekman M, Batut B, Ostrovsky A, Antoniewski C, Clements D, Ramirez F, et al. A single-cell RNA-sequencing training and analysis suite using the Galaxy framework. GigaScience 2020, 9, giaa102. [Google Scholar]
- Moreno P, Huang N, Manning JR, Mohammed S, Solovyev A, Polanski K, et al. User-friendly, scalable tools and workflows for single-cell RNA-seq analysis. Nat Methods 2021, 18, 327–328. [Google Scholar]
- Django (2024) Django (version 5.1.4). https://github.com/django/django/releases/tag/5.1.4.
- Python (2024) Python (version 3.12). https://github.com/python/cpython/releases/tag/v3.12.11.
- Batut B, Bacon W, Zierep P, Bernt M, Soranzo N, Gustafsson OJR. Galaxy CoDex for finding tools, workflows, and training. In: GCC 2024-Galaxy Community Conference; 2024. [CrossRef]
- https://doi.org/10.25953/3eje-q240.
- https://doi.org/10.5281/zenodo.3967970.
- https://github.com/galaxyproject/galaxy_codex/blob/main/communities/genome/lab/.
- https://singlecell.usegalaxy.org.
- https://singlecell.usegalaxy.eu.
- https://singlecell.usegalaxy.org.au.
- https://microbiology.usegalaxy.org.
- https://microbiology.usegalaxy.eu.
- https://microbiology.usegalaxy.fr.
- https://microbiology.usegalaxy.org.au.
| Galaxy Lab | A subdomain of a Galaxy server which shows the Galaxy service with a custom landing page and tool box (previously known as a “Flavour”) |
| GLE | Galaxy Labs Engine |
| Lab page | The custom landing page shown by a Galaxy Lab, rendered by the Galaxy Labs Engine |
| Lab content | A remote GitHub folder of documents that the Labs Engine uses to render a Lab page |
| Content root | A YAML file in the Lab content which defines a specific Lab page. Each Lab can have multiple content roots to render the Lab page differently for each Galaxy server. |
| Lab creator | The person who uses the Labs Engine to build Lab pages, typically a research community member or web developer |
| Lab user | The target end-user for a Lab, typically a researcher |
| SIG | Special Interest Group - research community members with a common interest e.g., Genomics |
| Subdomain | An alternative domain to serve a Galaxy Lab, e.g., genome.usegalaxy.org.au |
| Problem | Solution |
| Each Galaxy server must be able to customize each Lab page with local context | Galaxy servers can specify a <myserver>.yml to override base configuration, enabling server-specific variables and content overrides |
| Each Galaxy server must be able to customize the introductory text, footer and stylesheets to meet local requirements | All Markdown snippets and static files can be customized by placing them in a <HOSTNAME> folder and specifying the custom path in <myserver>.yml. |
| Redundant content across different servers | Introduces variable interpolation across YAML and Markdown/HTML to reuse content dynamically |
| Lack of flexibility in content formatting | Supports Markdown and HTML, with popular UI libraries available for use. Markdown “snippets” are injected in three locations in the page where the Lab developer has complete creative freedom. |
| Cluttered interfaces with lack of consistency between Labs | All Labs are rendered from the same base template, which includes a standardized UI layout to ensure that Lab pages are intuitive and consistent. Dense content is structured into sections, tabs and collapsible elements. |
| Managing sections that are specific to certain Galaxy server(s) | Allows exclusion of specific items from certain hosts using the exclude_from directive the YAML content |
| Requirement to host content on GitHub creates a slow development cycle | Local rendering capability via a Python-based CLI to instantly preview changes before pushing to GitHub |
| Delay in Lab page updates due to server-side caching | Supports a cache=false flag to skip the caching. Local rendering bypasses cache entirely. |
| Capability for hosting static content (e.g., images) for use in Lab pages | Images can be placed in the “static” folder of the content repository and then referenced in Markdown and YAML content. |
| Lab developers need constructive error messages when the expected Lab structure/schema is violated | YAML content is validated and sanitized using the “pydantic” library. Error feedback is rendered into the requested web page for interpretation by the user. |
| A standardized, accessible mechanism for adding contributors | The content repository can include a CONTRIBUTORS file which lists GitHub usernames of Lab contributors. These are rendered at the bottom of the Lab page. |
| Ability to embed content from other websites into Lab pages | Since extended Markdown is supported, iframe elements can be used to embed content from any publicly visible website. |
| Links in the Lab page need to be screened to guarantee that the “Galaxy in Galaxy” anomaly does not occur. | All URLs in the rendered Lab page are programatically evaluated and configured to open in a new tab if they originate from the current web host. |
| Robust documentation for building new Lab pages | The GLE landing page provides interactive documentation for building a Galaxy Lab page |
| Landing on a new resource for the first time can be daunting for inexperienced users | GLE accepts a video_url attribute in the <server>.yml metadata, which enables a YouTube video to be easily embedded in the Introduction of the Lab. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




