Submitted:
30 July 2025
Posted:
31 July 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Related Work
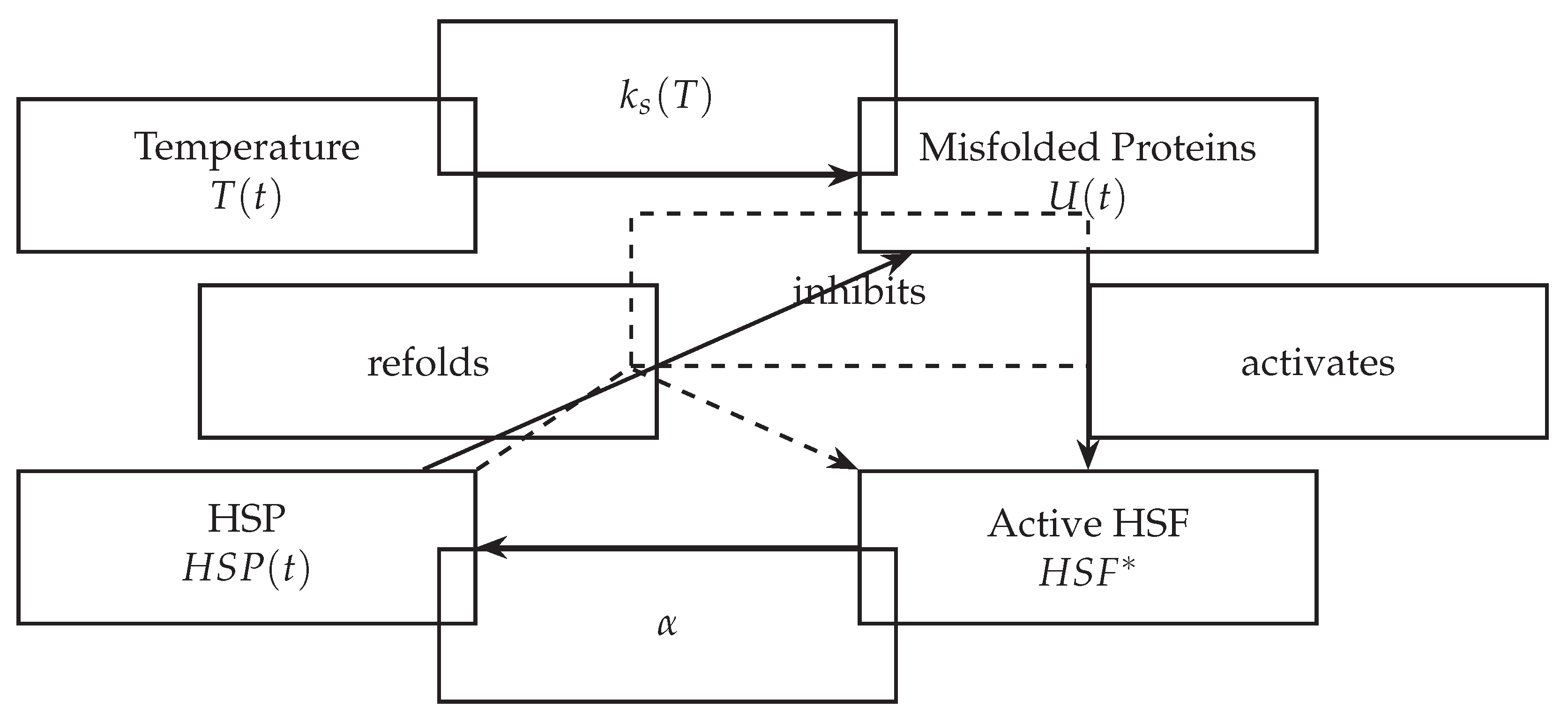
3. Modeling Framework
- –
- : the concentration of unfolded or misfolded proteins,
- –
- : the concentration of free heat shock proteins,
- –
- : the active form of the heat shock transcription factor.
3.1. Model Assumptions
- –
- The unfolding of the protein increases with increasing temperature.
- –
- HSPs bind misfolded proteins to assist in refolding or degradation.
- –
- When HSPs are titrated with misfolded proteins, HSF1 is activated, triggering the transcription of the HSP gene.
- –
- Active HSF1 degrades or deactivates over time in the absence of stress.
- –
- The system includes feedback inhibition, where increased HSP levels suppress further HSF1 activation.
3.2. Model Equations
- –
- is the temperature-dependent rate of misfolded protein production,
- –
- is the effective rate of HSP-mediated folding or clearance of misfolded proteins,
- –
- is the maximum transcription rate of HSPs induced by ,
- –
- is the degradation rate of HSPs,
- –
- governs the sensitivity of HSF activation to misfolded proteins,
- –
- is the deactivation or degradation rate of active HSF1,
- –
- is the half-saturation constant for HSF activation.
3.3. Temperature Profiles
- Gradual stress: A linearly increasing function representing slow heating over time:where is the initial temperature and r is the rate of increase.
- Acute shock: A step function to mimic a sudden rise in temperature:where is the time of shock, and is the post-shock temperature.
4. Model Calibration and Parameter Sensitivity
| Parameter | Description | Sensitivity Rank |
|---|---|---|
| Max HSP production rate | High | |
| HSF activation threshold | High | |
| HSF degradation rate | Medium | |
| Folding rate constant | High | |
| Half-saturation constant | Low |
5. Computational Implementation
6. Results and Analysis
6.1. Response to Gradual Heating
6.2. Response to Acute Heat Shock
6.3. Comparison with Experimental Data
- –
- The delayed but high-magnitude HSP expression in acute stress matches the observed transcriptional bursts in thermal shock experiments.
- –
- The anticipatory activation of HSP in gradual heating resembles physiological adaptation to sublethal thermal elevations.
7. Discussion
8. Applications and Future Extensions
- –
- Drug discovery: Simulating the effects of chaperone inhibitors or proteostasis modulators.
- –
- Synthetic biology: Designing temperature-sensitive genetic circuits using the HSR pathway.
- –
- Disease modeling: Investigating stress regulation failure in neurodegenerative diseases such as Alzheimer’s or Huntington’s disease [1].
- Spatial modeling using partial differential equations (PDEs) to represent intracellular gradients.
- In conjunction with the unfolded protein response (UPR) and oxidative stress pathways.
- Integration with single-cell transcriptomics data for personalized modeling.
9. Conclusion
References
- Morimoto, R.I. : Proteotoxic stress and inducible chaperone networks in neurodegenerative disease and aging. Genes & Development 2008, 22, 1427–1438. [Google Scholar] [CrossRef]
- Peper, D. , Grimbs, M., Kurths, J.: A mathematical model of hsf1 regulation of the heat shock response. Journal of Theoretical Biology 2017, 432, 44–58. [Google Scholar] [CrossRef]
- Rieger, J. , Körner, A.B., Buchner, B.: Modeling the heat shock response in eukaryotes: An integrative approach. PLoS Computational Biology 2005, 1, 343–355. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




