Submitted:
26 July 2025
Posted:
28 July 2025
You are already at the latest version
Abstract
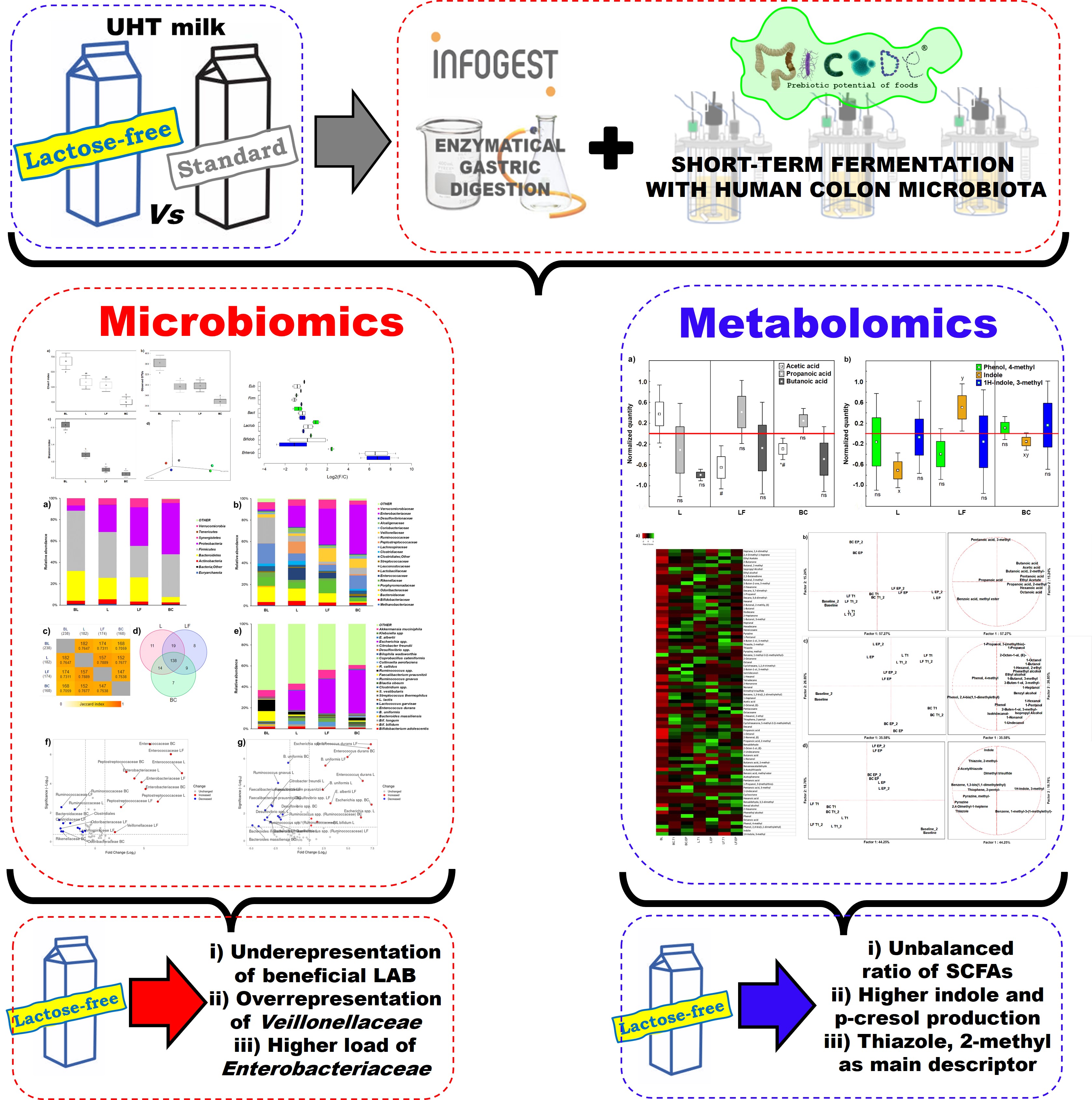
Keywords:
1. Introduction
2. Materials and Methods
2.1. Milk Samples
2.2. Experimental Workflow
2.3. Human Colon Microbiota
2.4. In Vitro Intestinal Model
2.5. Metabolomics
2.5.1. Volatilome Analysis
2.5.2. Quantification of Main Microbial VOCs
2.6. Microbiomics
2.6.1. Metataxonomy
2.6.2. Quantitation of Bacterial Groups by qPCR
2.7. Data Mining and Statistics
3. Results
3.1. Metabolomics
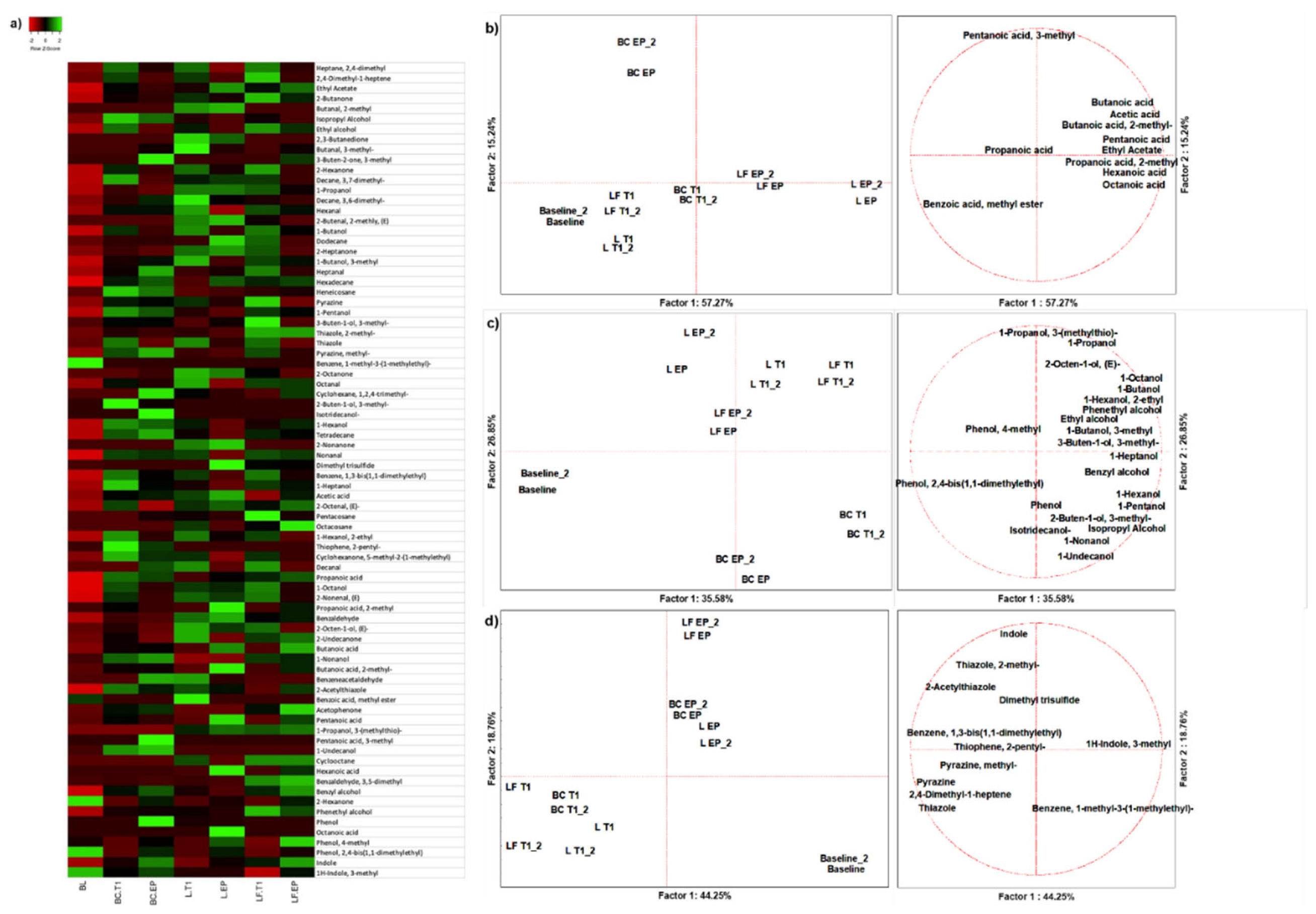
3.1.1. Volatilome Analysis
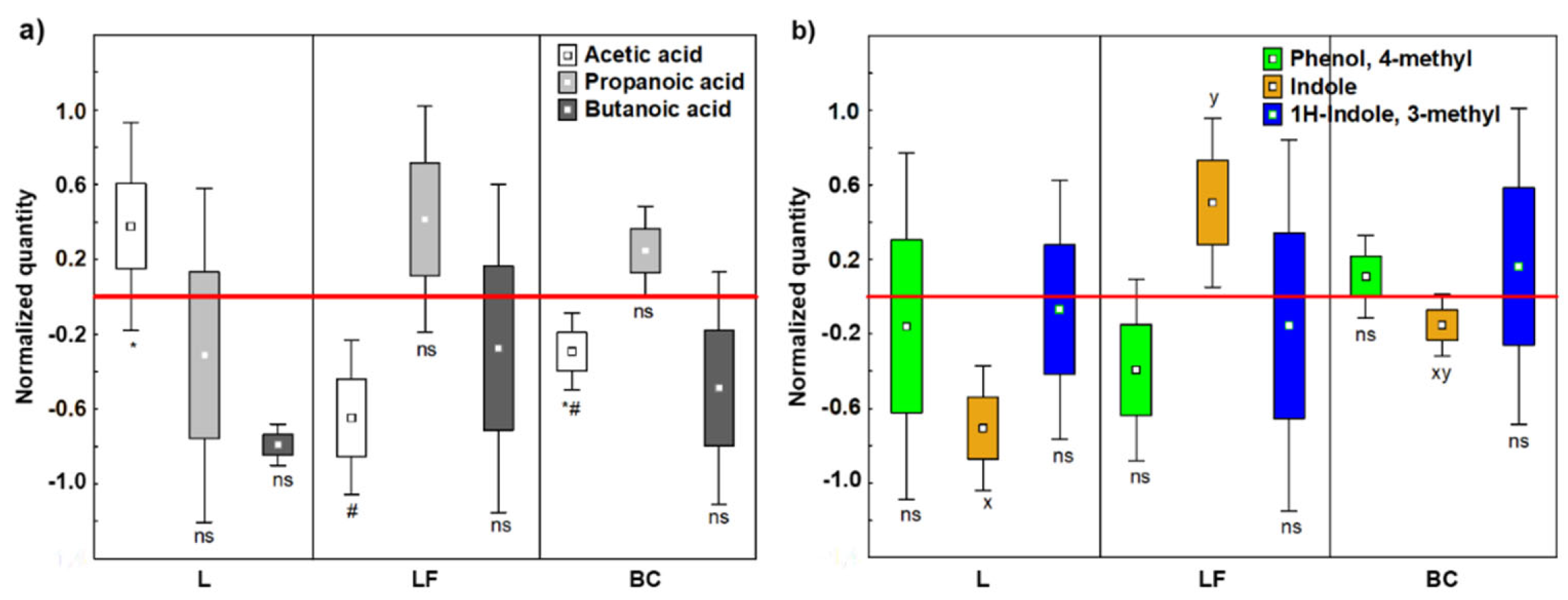
3.1.2. Short Chain Fatty Acids
3.1.3. Indoles and Phenols
3.2. Microbiomics
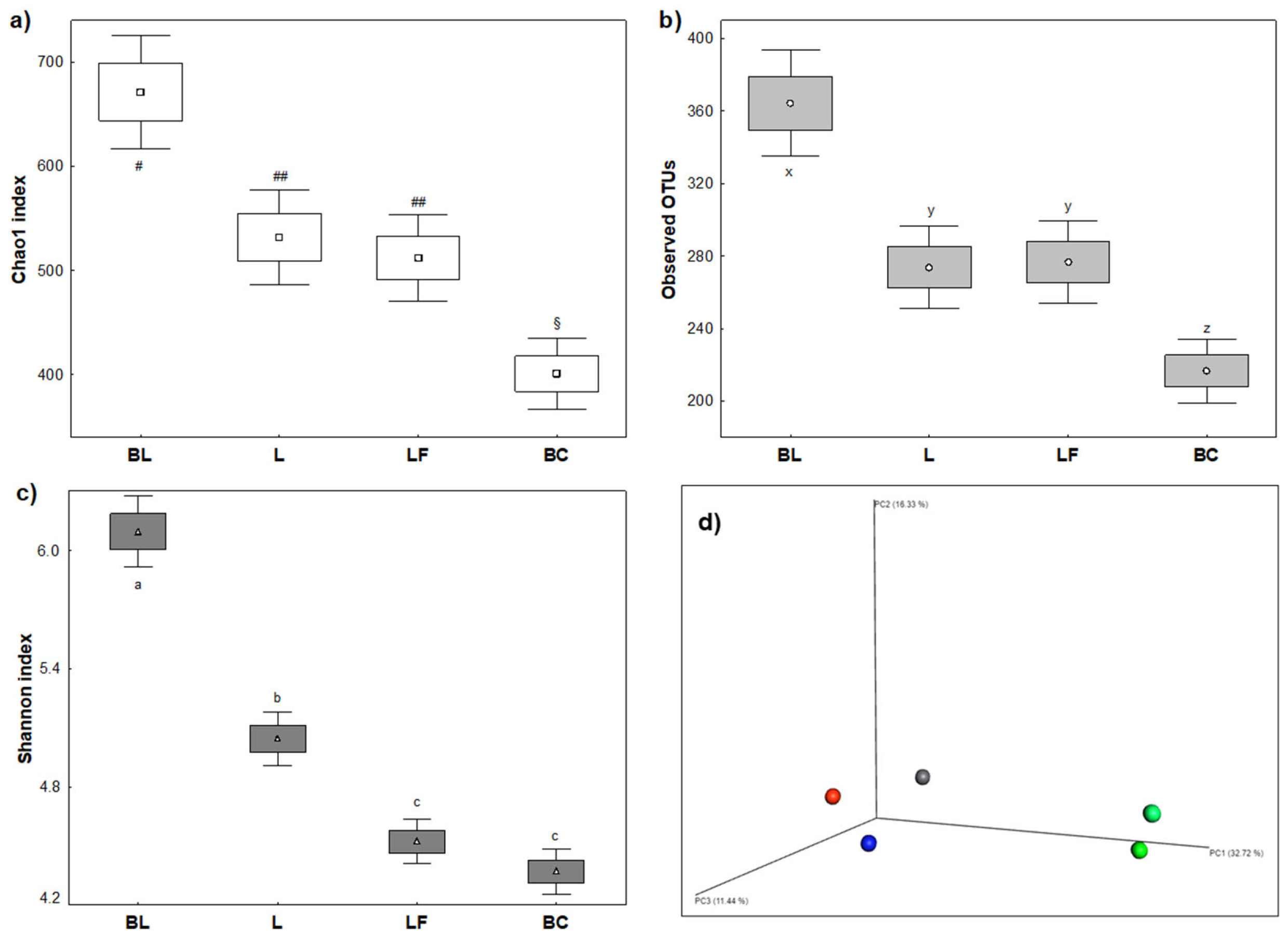
3.2.1. Ecological Biodiversity of Colonic Fermentations
3.2.2. Metataxonomy of Colonic Fermentations
3.2.3. Enumeration of Selected Bacterial Targets
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Forsgard, R.A. Lactose digestion in humans: Intestinal lactase appears to be constitutive whereas colonic microbiome is adaptable. American Journal of Clinical Nutrition 2019, 110, 273–279. [Google Scholar] [CrossRef] [PubMed]
- Campbell, A.K.; Waud, J.P.; Matthews, S.B. The molecular basis of lactose intolerance. Scientific Progress 2005, 88, 157–202. [Google Scholar] [CrossRef] [PubMed]
- Vesa, T.H.; Marteau, P.; Korpela, R. Lactose intolerance. Journal of the American College of Nutrition 2000, 19 (sup2), 165S–175S. [Google Scholar] [CrossRef] [PubMed]
- Canani, R.B.; Pezzella, V.; Amoroso, A.; Cozzolino, T.; Di Scala, C.; Passariello, A. Diagnosing and treating intolerance to carbohydrates in children. Nutrients 2016, 8, 157. [Google Scholar] [CrossRef]
- Misselwitz, B.; Butter, M.; Verbeke, K.; Fox, M.R. Update on lactose malabsorption and intolerance: pathogenesis, diagnosis and clinical management. Gut 2019, 68, 2080–2091. [Google Scholar] [CrossRef]
- Windey, K.; Houben, E.; Deroover, L.; Verbeke, K. Contribution of colonic fermentation and fecal water toxicity to pathophysiology of lactose-intolerance. Nutrients 2015, 7, 7505–7522. [Google Scholar] [CrossRef]
- Dekker, P.J.T.; Koenders, D.; Bruins, M.J. Lactose-Free Dairy Products: Market Developments, Production, Nutrition and Health Benefits. Nutrients 2019, 11, 551. [Google Scholar] [CrossRef]
- Gupta, R.S.; Warren, C.M.; Smith, B.M.; Jiang, J.; Blumenstock, J.A.; Davis, M.M.; Schleimer, R.P.; et al. Prevalence and Severity of Food Allergies Among US Adults. JAMA Netw. Open 2019, 2, e185630. [Google Scholar] [CrossRef]
- Li, X.; Yin, J.; Zhu, Y.; Wang, X.; Hu, X.; Bao, W.; Huang, Y.; et al. Effects of Whole Milk Supplementation on Gut Microbiota and Cardiometabolic Biomarkers in Subjects with and without Lactose Malabsorption. Nutrients 2018, 10, 1403. [Google Scholar] [CrossRef]
- Allied Market Research. Available online: https://www.alliedmarketresearch.com/lactose-free-dairy-market-A18453 (accessed on 9 February 2024).
- Slupsky, C.M.; He, X.; Hernell, O.; Andersson, Y.; Rudolph, C.; Lönnerdal, B.; et al. Postprandial metabolic response of breast-fed infants and infants fed lactose-free vs regular infant formula: A randomized controlled trial. Scientific Reports 2017, 7, 3640. [Google Scholar] [CrossRef]
- Abrams, S.A.; Griffin, I.J.; Davila, P.M. Calcium and zinc absorption from lactose-containing and lactose-free infant formulas. American Journal of Clinical Nutrition 2002, 76, 442–446. [Google Scholar] [CrossRef]
- Szilagyi, A. Lactose – a potential prebiotic. Aliment. Pharmacol. Ther 2002, 16, 1591–1602. [Google Scholar] [CrossRef]
- Szilagyi, A. Redefining lactose as a conditional prebiotic. Canadian Journal of Gastroenterology and Hepatology 2004, 18, 163–167. [Google Scholar] [CrossRef]
- Brodkorb, A.; Egger, L.; Alminger, M.; Alvito, P.; Assunção, R.; Ballance, S. . & Recio, I. INFOGEST static in vitro simulation of gastrointestinal food digestion. Nature protocols. 2019, 14, 991–1014. [Google Scholar]
- Gianotti, A.; Marin, V.; Cardone, G.; Bordoni, A.; Mancini, E.; Magni, M. . & Nissen, L. Personalized and precise functional assessment of innovative flatbreads toward the colon microbiota of people with metabolic syndrome: Results from an in vitro simulation. Food Research International. 2025, 209, 116197. [Google Scholar]
- Casciano, F.; Nissen, L.; Bordoni, A.; Gianotti, A. Colonic in vitro model assessment of effect of lactose-free milk on gut microbiota of lactose intolerant donors. International Journal of Food Science and Technology 2022, 58, 4485–4494. [Google Scholar] [CrossRef]
- Diotallevi, C.; Gaudioso, G.; Fava, F.; Angeli, A.; Lotti, C.; Vrhovsek, U.; Rinott, E.; et al. Measuring effect of Mankai® (Wolffia globosa) on the gut microbiota and its metabolic output using an in vitro colon model. Journal of Functional Foods 2021, 84, 104597. [Google Scholar] [CrossRef]
- Nissen, L.; Casciano, F.; Chiarello, E.; Di Nunzio, M.; Bordoni, A.; Gianotti, A. Sourdough process and spirulina-enrichment can mitigate the limitations of colon fermentation performances of gluten-free breads in non-celiac gut model. Food Chemistry 2024, 436, 137633. [Google Scholar] [CrossRef] [PubMed]
- Oba, S.; Sunagawa, T.; Tanihiro, R.; Awashima, K.; Sugiyama, H.; Odani, T.; Nakamura, Y.; et al. Prebiotic effects of yeast mannan, which selectively promotes Bacteroides thetaiotaomicron and Bacteroides ovatus in a human colonic microbiota model. Scientific Reports 2020, 10, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Nissen, L.; Casciano, F.; Babini, E.; Gianotti, A. Beneficial metabolic transformations, and prebiotic potential of hemp bran and its alcalase hydrolysate, after colonic fermentation in a gut model. Scientific Reports 2023, 13, 1552. [Google Scholar] [CrossRef]
- Wang, X.; Gibson, G.R.; Sailer, M.; Theis, S.; Rastall, R.A. Prebiotics inhibit proteolysis by gut bacteria in a host diet-dependent manner: a three-stage continuous in vitro gut model experiment. Applied and Environmental Microbiology. 2020, 86, 02730–19. [Google Scholar] [CrossRef] [PubMed]
- Cattivelli, A.; Nissen, L.; Casciano, F.; Tagliazucchi, D.; Gianotti, A. Impact of cooking methods of red-skinned onion on metabolic transformation of phenolic compounds and gut microbiota. Food and Function 2023, 14, 3509–3525. [Google Scholar] [CrossRef] [PubMed]
- Day-Walsh, P.; Shehata, E.; Saha, S.; Savva, G.M.; Nemeckova, B.; Speranza, J.; Kellingray, L.; et al. The use of an in vitro batch fermentation (human colon) model for investigating mechanisms of TMA production from choline, l-carnitine, and related precursors by the human gut microbiota. European Journal of Nutrition 2021, 60, 3987–3999. [Google Scholar] [CrossRef] [PubMed]
- Marino, M.; Dubsky de Wittenau, G.; Saccà, E.; Cattonaro, F.; Spadotto, A.; Innocente, N.; et al. Metagenomic profiles of different types of Italian high-moisture Mozzarella cheese. Food Microbiology 2018, 79, 123–131. [Google Scholar] [CrossRef]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nature Biotechnology 2019, 37, 852–857. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biology 2014, 15, 550. [Google Scholar] [CrossRef]
- Hill, C.J.; Lynch, D.B.; Murphy, K.; Ulaszewska, M.; Jeffery, I.B.; O'Shea, C.A.; Watkins, C.; et al. Evolution of gut microbiota composition from birth to 24 weeks in the INFANTMET Cohort. Microbiome 2017, 5, 4. [Google Scholar] [CrossRef]
- Modesto, M.; Stefanini, I.; D’Aimmo, M.R.; Nissen, L.; Tabanelli, D.; Mazzoni, M.; Bosi, P.; et al. Strategies to augment non-immune system based defence mechanisms against gastrointestinal diseases in pigs. NJAS - Wagening Journal of Life Science 2011, 58, 149–156. [Google Scholar] [CrossRef]
- Tanner, S.A.; Zihler Berner, A.; Rigozzi, E.; Grattepanche, F.; Chassard, C.; Lacroix, C. In vitro continuous fermentation model (PolyFermS) of the swine proximal colon for simultaneous testing on the same gut microbiota. PloS one 2014, 9, e94123. [Google Scholar] [CrossRef] [PubMed]
- Tsitko, I.; Wiik-Miettinen, F.; Mattila, O.; Rosa-Sibakov, N.; Seppänen-Laakso, T.; Maukonen, J.; Nordlund, E.; et al. A small in vitro fermentation model for screening the gut microbiota effects of different fiber preparations. International Journal of Molecular Science 2019, 20, 1925. [Google Scholar] [CrossRef]
- Granato, D.; de Araújo Calado, V.M.; Jarvis, B. Observations on the use of statistical methods in food science and technology. Food Research International 2014, 55, 137–149. [Google Scholar] [CrossRef]
- Vázquez-Baeza, Y.; Pirrung, M.; Gonzalez, A.; Knight, R. EMPeror: a tool for visualizing high-throughput microbial community data. Gigascience 2013, 2, 2047–217X. [Google Scholar] [CrossRef]
- Goedhart, J.; Luijsterburg, M.S. VolcaNoseR is a web app for creating, exploring, labeling and sharing volcano plots. Scientific Reports 2020, 10, 20560. [Google Scholar] [CrossRef]
- Spitzer, M.; Wildenhain, J.; Rappsilber, J.; Tyers, M. BoxPlotR: a web tool for generation of box plots. Nature Methods 2014, 11, 121–122. [Google Scholar] [CrossRef] [PubMed]
- Vitellio, P.; Celano, G.; Bonfrate, L.; Gobbetti, M.; Portincasa, P.; De Angelis, M. Effects of Bifidobacterium longum and Lactobacillus rhamnosus on gut microbiota in patients with lactose intolerance and persisting functional gastrointestinal symptoms: A randomised, double-Blind, cross-over study. Nutrients 2019, 11, 886. [Google Scholar] [CrossRef]
- Vipperla, K.; O’Keefe, S.J. The microbiota and its metabolites in colonic mucosal health and cancer risk. Nutrition in Clinical Practice 2012, 27, 624–635. [Google Scholar] [CrossRef]
- Tain, Y.L.; Chang, S.K.C.; Liao, J.W.; Chen, Y.W.; Huang, H.T.; Li, Y.L.; Hou, C.Y. Synthesis of short-chain-fatty-acid resveratrol esters and their antioxidant properties. Antioxidants 2021, 10, 420. [Google Scholar] [CrossRef]
- Xiong, R.G.; Zhou, D.D.; Wu, S.X.; Huang, S.Y.; Saimaiti, A.; Yang, Z.J.; Shang, A.; et al. Health benefits and side effects of short-chain fatty acids. Foods 2022, 11, 2863. [Google Scholar] [CrossRef]
- Xu, Y.; Zhu, Y.; Li, X.; Sun, B. Dynamic balancing of intestinal short-chain fatty acids: The crucial role of bacterial metabolism. Trends in Food Science and Technology 2020, 100, 118–130. [Google Scholar] [CrossRef]
- Ma, Q.; Meng, N.; Li, Y.; Wang, J. Occurrence, impacts, and microbial transformation of 3-methylindole (skole): A critical review. Journal of Hazardous Materials 2021, 416, 126181. [Google Scholar] [CrossRef]
- Diether, N.E.; Willing, B.P. Microbial fermentation of dietary protein: an important factor in diet-microbe-host interaction. Microorganisms 2019, 7, 19. [Google Scholar] [CrossRef]
- Bond, J.H. Quantitative measurement of lactose absorption. Gastroenterology 1976, 70, 1058–1062. [Google Scholar] [CrossRef] [PubMed]
- Starz, E.; Wzorek, K.; Folwarski, M.; Kazmierczak-Siedlecka, K.; Stachowska, L.; Przewlocka, K.; Stachowska, E.; et al. The Modification of the Gut Microbiota via Selected Specific Diets in Patients with Crohn’s Disease. Nutrients 2021, 13, 2125. [Google Scholar] [CrossRef]
- Kurilshikov, A.; Medina-Gomez, C.; Bacigalupe, R.; Radjabzadeh, D.; Wang, J.; Demirkan, A.; Le Roy, C.I.; et al. Large-scale association analyses identify host factors influencing human gut microbiome composition. Nature Genetics 2021, 53, 156–165. [Google Scholar] [CrossRef]
- Van den Abbeele, P.; Sprenger, N.; Ghyselinck, J.; Marsaux, B.; Marzorati, M.; Rochat, F.A. Comparison of the In vitro Effects of 2′Fucosyllactose and Lactose on the Composition and Activity of Gut Microbiota from Infants and Toddlers. Nutrients 2021, 13, 726. [Google Scholar] [CrossRef]
- Li, X.; Zhang, B.; Hu, Y.; Zhao, Y. New insights into gut-bacteria-derived indole and its derivatives in intestinal and liver diseases. Frontiers in Pharmacology 2021, 12. [Google Scholar] [CrossRef]
- Bonder, M.J.; Tigchelaar, E.F.; Cai, X.; Trynka, G.; Cenit, M.C.; Hrdlickova, B.; Zhong, H.; et al. The influence of a short-term gluten-free diet on the human gut microbiome. Genome Medicine 2016, 8, 45. [Google Scholar] [CrossRef]
- Valguarnera, E.; Wardenburg, J.B. Good gone bad: one toxin away from disease for Bacteroides fragilis. Journal of Molecular Biology 2020, 432, 765–785. [Google Scholar] [CrossRef] [PubMed]
- Centor, R.M.; Atkinson, T.P.; Xiao, L. Fusobacterium necrophorum oral infections - A need for guidance. Anaerobe 2022, 75, 102532. [Google Scholar] [CrossRef] [PubMed]
- Tennoune, N.; Andriamihaja, M.; Blachier, F. Production of Indole and Indole-Related Compounds by the Intestinal Microbiota and Consequences for the Host: The Good, the Bad, and the Ugly. Microorganisms 2022, 10, 930. [Google Scholar] [CrossRef]
- Sundar, A.; Kardes, F.R. The role of perceived variability and the health halo effect in nutritional inference and consumption. Psychology and Marketing 2015, 32, 512–521. [Google Scholar] [CrossRef]
- Castellini, G.; Savarese, M.; Graffigna, G. The role of free-from symbols on consumer perceptions of healthiness, quality and intention to buy baked food products. International Journal of Food Science and Nutrition 2023, 74, 395–402. [Google Scholar] [CrossRef]
- Bischoff, S.C.; Escher, J.; Hébuterne, X.; Kłęk, S.; Krznaric, Z.; Schneider, S.; Shamir, R.; et al. Forbes, ESPEN practical guideline: Clinical Nutrition in inflammatory bowel disease. Clinical Nutrition 2020, 39, 632–653. [Google Scholar] [CrossRef] [PubMed]
- Lane, D.J.; Harrison, Jr. A. P.; Stahl, D.; Pace, B.; Giovannoni, S.J.; Olsen, G.J.; Pace, N.R. Evolutionary relationship among sulfur- and iron-oxidizing eubacteria. Journal of Bacteriology 1992, 174, 269–278. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.; Xia, X.; Tang, R.; Zhou, J.; Zhao, H.; Wang, K. Development of a real-time PCR method for Firmicutes and Bacteroidetes in faeces and its application to quantify intestinal population of obese and lean pigs. Letters in Applied Microbiology 2008, 47, 367–73. [Google Scholar] [CrossRef]
- Walter, J.; Hertel, C.; Tannock, G.W.; Lis, C.M.; Munro, K.; Hammes, W.P. Detection of Lactobacillus, Pediococcus, Leuconostoc, and Weissella species in human feces by using group-specific PCR primers and Denaturing Gradient Gel Electrophoresis. Applied and Environmental Microbiology 2001, 67, 2578–2585. [Google Scholar] [CrossRef]
- Masco, L.; Ventura, M.; Zink, R.; X Huys, J.; Swings, V. Polyphasic taxonomic analysis of Bifidobacterium animalis and Bifidobacterium lactis reveals relatedness at the subspecies level: reclassification of Bifidobacterium animalis as Bifidobacterium animalis subsp. animalis subsp. nov. and Bifidobacterium lactis as Bifidobacterium animalis subsp. lactis subsp. nov. International Journal of Systematic and. Evolutionary Microbiology 2004, 54, 1137–1143. [Google Scholar]
- Bartosch, S.; Fite, A.; Macfarlane, G.T.; McMurdo, M.E. Characterization of bacterial communities in feces from healthy elderly volunteers and hospitalized elderly patients by using real-time PCR and effects of antibiotic treatment on the fecal microbiota. Applied and Environmental Microbiology 2004, 70, 3575–3581. [Google Scholar] [CrossRef]
| Outputs | Lactose tolerant | Lactose intolerant [17] | ||
| Metabolomics | UHT semi-skimmed milk (L) | UHT semi-skimmed lactose-free milk (LF) |
UHT semi- skimmed milk (L) |
UHT semi-skimmed lactose-free milk (LF) |
| Organic acids main descriptors | Pentanoic acid and Propanoic acid, 2-methyl | Butanoic acid | Pentanoic acid, Hexanoic acid, Octanoic acid | Butanoic acid |
| Alcohols main descriptors | 2-Octen-1-ol, (E) and 1-Propanol | 3-Buten-1-ol, 3-methyl-, Benzyl Alcohol, Phenethyl Alcohol and Phenol, 4-methyl | 1-Butanol, Phenol | Ethyl alcohol, 1-Octanol, 1-Hexanol, 2-ethyl |
| Other VOCs main descriptors | Dimethyl trisulfide | Thiazole, 2-methyl | 2-Hexanone | 2-Acetylthiazole |
| SCFA production | Increased Acetic acid | Unbalanced production | Increased Acetic acid | Increased Butanoic acid |
| Detrimental VOCs production | All decreased | Increased Indole | Increased p-cresol | All decreased |
| Outputs | Lactose tolerant | Lactose intolerant* | ||
| Microbiomics | UHT semi-skimmed milk (L) | UHT semi-skimmed lactose-free milk (LF) |
UHT semi- skimmed milk (L) |
UHT semi-skimmed lactose-free milk (LF) |
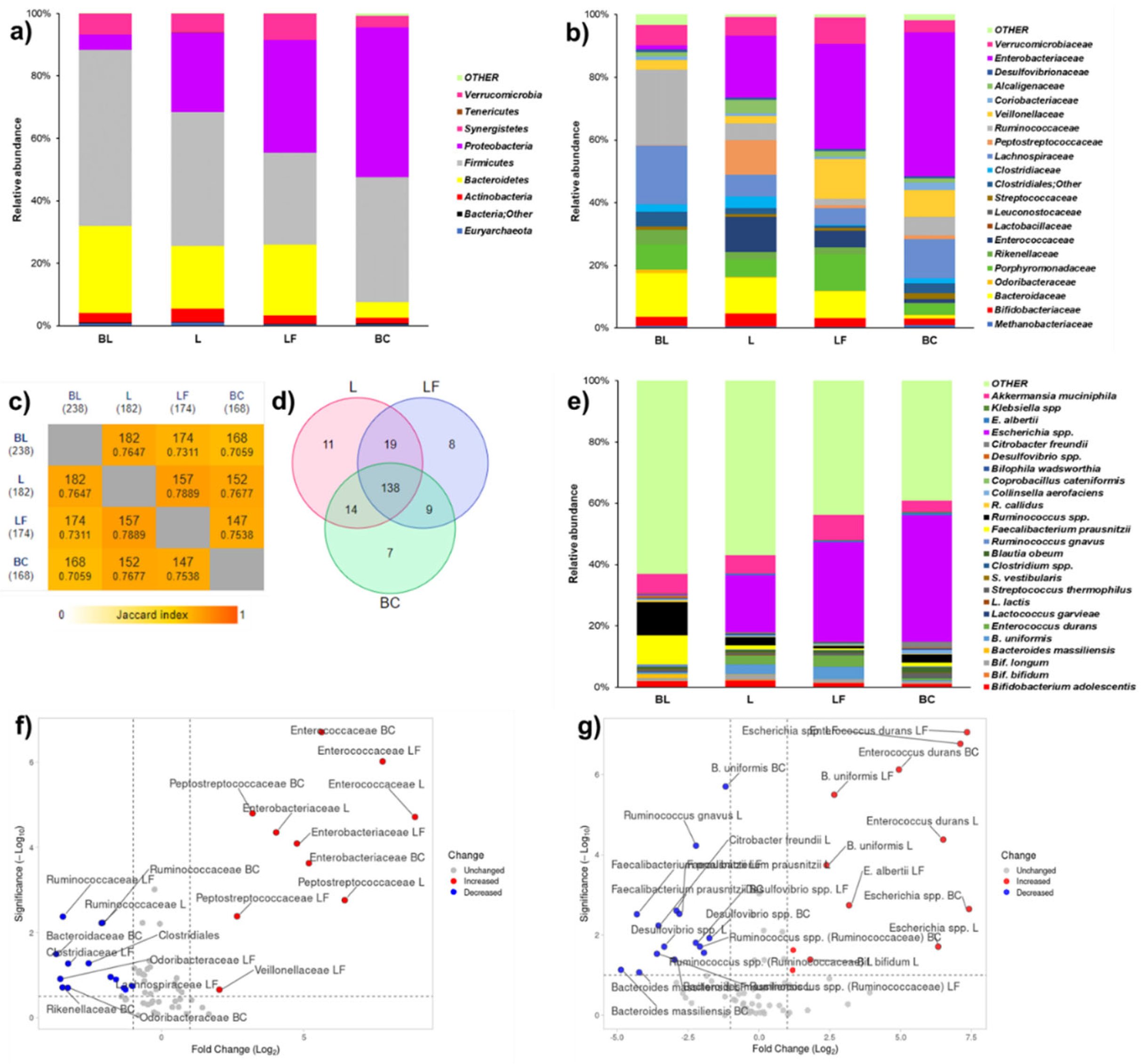
| Metataxonomy (16S-rRNA) |
Increased Bifidobacterium bifidum | Decreased Ruminococcaceae; increased Veillonellaceae and Peptostreptococcaceae |
Increased Klebsiella spp.; decreaed Faecalibacterium prausnitsii, Roseburia faecis. | Unchanged Verrucomicrobia phylum; decreased Peptostreptococcaceae. |
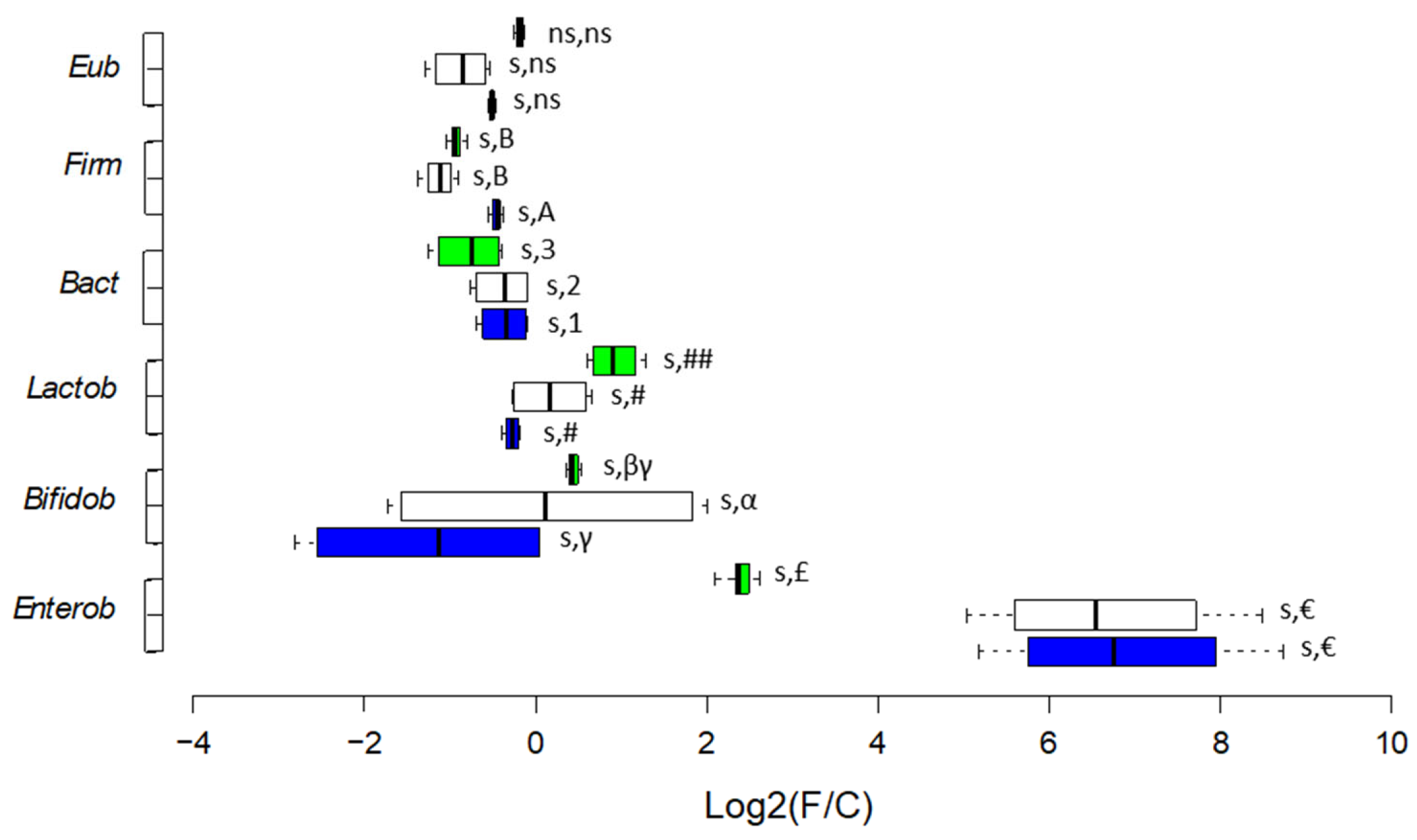
| Selected bacterial taxa (qPCR) |
Increased Lactobacillales and Bifidobacteriaceae | Decreased Lactobacillales and Bifidobacteriaceae | Increased Lactobacillales and Enterobacteriaceae | Decreased Bacteroidetes and Lactobacillales; increased Enterobacteriaceae. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




